Upper aerodigestive squamous cell carcinoma (UASCC) presents significant challenges in clinical management due to its aggressive nature. Here, we elucidate the role of MLL3 mutations as early, clonal genomic events in UASCC tumorigenesis, highlighting their role as foundational drivers of cancer development. Utilizing CRISPR-edited, cross-species organoid modeling, we demonstrate that loss of MLL3 contributes to early squamous neoplastic evolution. Furthermore, we identify an MLL3/GRHL2 protein complex that regulates the UASCC epigenome, particularly impacting immune response pathways. Notably, a novel MLL3/GRHL2-IRF1 axis promotes the expression of Th1 chemokines, enhancing anti-tumor immunity by facilitating T cell infiltration into the tumor microenvironment. Consequently, MLL3 regulates the in vivo efficacy of immune checkpoint blockade (ICB) therapy, corroborated by the strong association between MLL3 expression and human patients’ clinical response to ICB therapy. Our work underscores the significance of MLL3 in UASCC pathogenesis and highlights the interplay between MLL3/GRHL2 and immune response pathways as potential therapeutic targets for UASCC treatment.
Introduction
Squamous cell carcinoma (SCC) is the most common cancer type derived from the stratified epithelium of the upper aerodigestive tract, comprising oral cavity, larynx, oropharynx, and esophagus. These cancers are collectively known as upper aerodigestive SCC (UASCC). Close to one million new cases of UASCC are diagnosed worldwide yearly (∼950,000 in 2020) (Sung et al., 2021), with head and neck SCC (HNSCC) and esophageal SCC (ESCC) being the most common types (Siegel et al., 2020). UASCC is aggressive and difficult to treat, and the majority of its patients do not respond well to conventional cytotoxic therapies. Moreover, due to the vital functions of the organs along the upper aerodigestive tract, UASCC causes severe morbidity and greatly impairs the quality of life of its patients.
The early malignant transformation of UASCC is a sequential and multistep process: benign squamous epithelium first becomes dysplastic, then carcinoma in situ, and finally progresses to invasive SCC. However, biological research remains quite limited concerning the molecular mechanisms underlying this stepwise early neoplastic evolution of UASCC. Particularly, the most crucial question, the primary driver(s) for the initial malignant transformation of UASCC, remains unclear. Although recurrent genomic lesions affecting tumor protein p53 (TP53) and cyclin-dependent kinase inhibitor 2A (CDKN2A) have been identified in dysplastic precursor samples (Graveland et al., 2013; Farah et al., 2019), they independently fail to predict the progression risk of dysplastic lesions (Farah et al., 2019). Because of the poor understanding of premalignant biology, no therapeutic strategy is available for the prevention or early intervention of the precursor condition of UASCC. This shortcoming is partially due to difficulties in performing molecular research into the stepwise neoplastic evolution of UASCC, largely because viable and valid human models representing this unique, early pathological transition have been lacking.
We and others have previously demonstrated that the mixed-lineage leukemia protein 3 (MLL3) gene (also known as lysine N-methyltransferase 2C) is recurrently mutated (∼10%) in UASCC (Hao et al., 2016; Lin et al., 2014). A significant fraction of MLL3 mutations is frameshift/stop-gain, suggesting that it is likely tumor suppressive. Indeed, functional characterizations have shown that loss of MLL3 promoted cancer cell fitness, proliferation, and metastasis in murine tumor models and cancer cell line models (Wang et al., 2018; Chen et al., 2014; Lee et al., 2009; Zhang et al., 2016; Cui et al., 2023; Na et al., 2022). However, the significance of MLL3 in the context of premalignant biology and early neoplastic evolution has not been rigorously tested.
Here, in our analyses of UASCC clonal evolution, 58% of MLL3 mutations were clonal, early events during tumorigenesis. Notably, in our robust 3D organoid modeling of early development of UASCC, we showed that loss of MLL3 strongly promoted oral squamous neoplastic evolution. These data underscore MLL3 genomic inactivation as a common, early driver of UASCC malignant transformation.
In exploration of the functional contribution of MLL3 to the biology of UASCC, we identified an MLL3/GRHL2 protein complex in the regulation of UASCC epigenome. Unexpectedly, our bioinformatic analyses unbiasedly identified that MLL3/GRHL2-regulated pathways were all associated with immune responses in UASCC. We have validated this observation and further showed that MLL3/GRHL2 promoted the transcription and expression of Th1 chemokines, including chemokine (C-X-C motif) ligand 9 (CXCL9) and chemokine (C-X-C motif) ligand (CXCL10). This finding is of great interest, given the prominent and essential role of Th1 chemokines in anti-tumor immunity via recruitment of tumor-infiltrating T cells. Indeed, loss of Th1 chemokines causes both immune evasion and resistance to immunotherapy in multiple cancer types (Litchfield et al., 2021), such as melanoma (Gao et al., 2016; Tumeh et al., 2014; Zaretsky et al., 2016) and lung cancer (Hellmann et al., 2018). Substantiating this hypothesis, both our orthotopic and subcutaneous syngeneic UASCC mouse models demonstrated that the loss of MLL3/GRHL2 impaired the recruitment of T cells to the tumor microenvironment, which led to reduced anti-tumor efficacy of immune checkpoint immunotherapy. Clearly, investigation of the role of MLL3/GRHL2 in anti-tumor immunity is of both biological and clinical significance, considering that MLL3 mutations are early and common in UASCC patients.
Results
MLL3 mutations are early, clonal genomic lesions in UASCC tumors
We previously delineated the spatial heterogeneity and clonal evolution of 13 UASCC patients using multi-region, intratumoral whole exome sequencing (Hao et al., 2016). In a combined analysis of two other public datasets (Chen et al., 2017; Yan et al., 2019), we analyzed a total of 323 multi-region tumor samples from 72 UASCC patients. We detected MLL3 mutations in 12.7% cases, consistent with large-scale sequencing data from the the Cancer Genome Atlas (TCGA) (Cancer Genomic Atlas Network, 2015, 2017). Phylogenetic trees were then constructed on the basis of shared and private somatic mutations detected in these intratumoral regions, using both synonymous and non-synonymous ones. Importantly, over half of MLL3 mutations (58.9%) were identified in the trunks/branches of phylogenetic trees (Fig. 1, A–C), suggesting that they are early genomic lesions. We also observed convergent evolution of multiple independent MLL3 mutations starting from the trunk to different branches in one case (Fig. 1 B), indicating early and persistent selection pressure conferred by MLL3 mutations. Truncating and splice MLL3 mutations were commonly seen, suggestive of tumor suppressive properties. Supportively, low MLL3 expression is associated with both poor overall and disease-free survival of HNSCC patients (Fig. 1 D and Fig. S1 A).
MLL3 mutations occur early during UASCC tumorigenesis. (A and B) Phylogenetic trees of two representative UASCC cases, reproduced using published data from us (Hao et al., 2016) and others (Chen et al., 2017; Yan et al., 2019). T1/2/3/5 were 4 intratumoral samples of the primary tumor while L1/2/3 were 3 lymph node metastasis lesions. (C) Column charts summarizing the mutational status of all detected MLL3 mutations. (D) Kaplan–Meier curves of overall survival and disease-free survival upon MLL3 mRNA expression with TCGA HNSCC patient survival. (E) Co-occurrence of mutations/deletions of MLL3, TP53, and CDKN2A in the TCGA ESCC and HNSCC datasets, plotted by the cBioportal tool (Cerami et al., 2012).
MLL3 mutations occur early during UASCC tumorigenesis. (A and B) Phylogenetic trees of two representative UASCC cases, reproduced using published data from us (Hao et al., 2016) and others (Chen et al., 2017; Yan et al., 2019). T1/2/3/5 were 4 intratumoral samples of the primary tumor while L1/2/3 were 3 lymph node metastasis lesions. (C) Column charts summarizing the mutational status of all detected MLL3 mutations. (D) Kaplan–Meier curves of overall survival and disease-free survival upon MLL3 mRNA expression with TCGA HNSCC patient survival. (E) Co-occurrence of mutations/deletions of MLL3, TP53, and CDKN2A in the TCGA ESCC and HNSCC datasets, plotted by the cBioportal tool (Cerami et al., 2012).
Knockdown of MLL3 accelerates tumorigenesis. (A) Kaplan–Meier plots using the mean expression level of MLL3 as the statistical cutoff. (B and C) Sanger sequencing showing genomic mutations introduced to either human or murine TP53 (upper) and CDKN2A (lower) genes. (D) qRT-PCR data of MLL3 mRNA expression level following knockdown using shRNA in murine TP53/CDKN2ADKO organoids (n = 3 biological replicates). (E) Representative images of murine TP53/CDKN2ADKO organoids with and without MLL3 knockdown. Scale bar = 20 μm. (F) Quantification of organoid size, as measured by ImageJ. (G) Proliferation rate of organoids at the indicated time points (n = 3 biological replicates). (H and I) Representative images, scale bar = 50 μm, and (I) quantification of IF staining for Ki-67 in murine TP53/CDKN2ADKO organoids with and without MLL3 knockdown (n = 3 biological replicates). *P < 0.05; **P < 0.01; and ***P < 0.001.
Knockdown of MLL3 accelerates tumorigenesis. (A) Kaplan–Meier plots using the mean expression level of MLL3 as the statistical cutoff. (B and C) Sanger sequencing showing genomic mutations introduced to either human or murine TP53 (upper) and CDKN2A (lower) genes. (D) qRT-PCR data of MLL3 mRNA expression level following knockdown using shRNA in murine TP53/CDKN2ADKO organoids (n = 3 biological replicates). (E) Representative images of murine TP53/CDKN2ADKO organoids with and without MLL3 knockdown. Scale bar = 20 μm. (F) Quantification of organoid size, as measured by ImageJ. (G) Proliferation rate of organoids at the indicated time points (n = 3 biological replicates). (H and I) Representative images, scale bar = 50 μm, and (I) quantification of IF staining for Ki-67 in murine TP53/CDKN2ADKO organoids with and without MLL3 knockdown (n = 3 biological replicates). *P < 0.05; **P < 0.01; and ***P < 0.001.
The biological significance of MLL3 loss in a CRISPR-edited, cross-species organoid model of early oral neoplastic evolution
Considering that MLL3 mutations are early, clonal events in UASCC tumors, we next sought to understand the functional significance of MLL3 during the neoplastic evolution of UASCC. However, as mentioned earlier, there had been a lack of robust and valid model to recapitulate the dysplastic, premalignant transition from benign squamous epithelium to invasive SCC. To address this challenge, we leveraged 3D organoid modeling which recapitulates and maintains (epi)genetic, biological, and phenotypic characteristics of originating tissues of origin (Lo et al., 2020). As we have demonstrated previously (Zhao et al., 2022a, 2022b; Nam et al., 2022; Jiang et al., 2024a), 3D organoid modeling provides a robust and versatile platform to investigate premalignant biology and early tumorigenesis.
Specifically, we first developed normal squamous organoid models from both human and mouse oral tissues, followed by the knockout of both TP53 and CDKN2A genes using CRISPR/Cas9 genome editing, as we described recently (Zhao et al., 2022a). We chose to target these two genes because TP53 mutations and CDKN2A mutations/deletions are the most common drivers in UASCC (Cancer Genomic Atlas Network, 2015, 2017) (Fig. 1 E). Moreover, loss of these two drivers occurs early during the neoplastic evolution of UASCC. Indeed, the most common genomically inactivated genes in oral dysplasia are TP53 and CDKN2A (Graveland et al., 2013; Farah et al., 2019). Most importantly, MLL3 mutations/deletions almost always co-occur with the genomic inactivation of TP53/CDKN2A. These findings provide a compelling rationale to investigate the loss of MLL3 in the context of TP53/CDKN2A dual-knockout (DKO) organoid models.
After Sanger sequencing validation of biallelic frameshift mutations of the targeted TP53 exon and CDKN2A exon (Fig. S1, B and C; Tables S1 and S2), we performed loss-of-function experiments to knockdown MLL3 in these TP53/CDKN2ADKO organoids (Fig. 2, A and B; and Fig. S1 D). Notably, compared with control organoids, MLL3-knockdown organoids exhibited substantially larger size, more complex multicellular structures, and markedly enlarged, atypical nuclei (Fig. 2, C and D; and Fig. S1, E and F). Moreover, organoid growth rates and Ki-67 labeling index were increased significantly in MLL3-knockdown organoids versus control organoids (Fig. 2, E–G; and Fig. S1, G–I). Importantly, these observations were reproduced using independent shRNAs and siRNAs across both human (two independent samples) and murine organoid models.
Loss of MLL3 drives early neoplastic evolution in a TP53/CDKN2A DKO oral organoid model. (A) A schematic plot, generated using BioRender, showing the development of CRISPR-edited, cross-species organoid models for oral neoplastic evolution (TP53/CDKN2ADKO) and subsequent knockdown of MLL3 expression. (B) qRT-PCR verification of the knockdown of MLL3 mRNA expression (n = 3 biological replicates). (C) Representative images of organoids from indicated groups at 40× magnification, scale bar = 20 μm. (D) Quantification of organoid size, as measured by ImageJ. (E) WST-1 assay to detect cell proliferation rate of organoids (n = 3 biological replicates). (F and G) Representative images, scale bar = 50 μm (F) and quantification of IF staining for Ki-67 in human organoids (n = 3 biological replicates) (G). (H) Individual tumor growth curves of control (TP53/CDKN2ADKO + scramble sgRNA) versus MLL3-knockout organoids (TP53/CDKN2ADKO + MLL3 knockout) (n = 5 biological replicates). (I) Kaplan–Meier plot showing the survival of mice from the two groups (n = 5 biological replicates). *P < 0.05; **P < 0.01; ***P < 0.001.
Loss of MLL3 drives early neoplastic evolution in a TP53/CDKN2A DKO oral organoid model. (A) A schematic plot, generated using BioRender, showing the development of CRISPR-edited, cross-species organoid models for oral neoplastic evolution (TP53/CDKN2ADKO) and subsequent knockdown of MLL3 expression. (B) qRT-PCR verification of the knockdown of MLL3 mRNA expression (n = 3 biological replicates). (C) Representative images of organoids from indicated groups at 40× magnification, scale bar = 20 μm. (D) Quantification of organoid size, as measured by ImageJ. (E) WST-1 assay to detect cell proliferation rate of organoids (n = 3 biological replicates). (F and G) Representative images, scale bar = 50 μm (F) and quantification of IF staining for Ki-67 in human organoids (n = 3 biological replicates) (G). (H) Individual tumor growth curves of control (TP53/CDKN2ADKO + scramble sgRNA) versus MLL3-knockout organoids (TP53/CDKN2ADKO + MLL3 knockout) (n = 5 biological replicates). (I) Kaplan–Meier plot showing the survival of mice from the two groups (n = 5 biological replicates). *P < 0.05; **P < 0.01; ***P < 0.001.
To further test the impact of MLL3 loss in vivo, we first knocked out MLL3 using CRISPR/Cas9 editing in the TP53/CDKN2ADKO organoids, and then performed orthotopic injections into the cheeks of athymic nude mice. Importantly, the MLL3-knockout group exhibited significantly faster tumor formation and growth compared with the control organoids with scramble sgRNA (Fig. 2 H). Moreover, Kaplan–Meier analysis showed that mice bearing MLL3-knockout organoids had substantially shorter tumor-free survival than the control mice (Fig. 2 I). These results strongly suggest that loss of MLL3 functions as an early driver of squamous neoplastic evolution.
Identification of an MLL3/GRHL2 protein complex
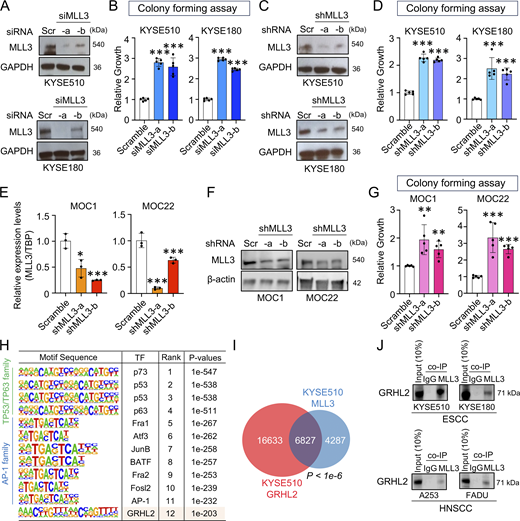
We next utilized cancer cell line models to interrogate the function of MLL3 in the setting of fully established UASCC tumors. Consistent with its tumor suppressive property, MLL3 knockdown led to increased colony formation in both human ESCC and HNSCC cell lines, which was validated by two independent siRNAs (Fig. 3, A and B; and Fig. S2 A) and two independent shRNAs (Fig. 3, C and D). Next, we silenced MLL3 in murine HNSCC cell lines, MOC1 and MOC22, derived from an HNSCC model of C57BL/6 mice (Chen et al., 2019). Consistently, MLL3 knockdown enhanced colony formation of these murine cells (Fig. 3, E–G), confirming its conserved function across species.
MLL3 inhibits cancer cell proliferation and interacts with GRHL2. (A) Protein levels of MLL3 following siRNA transfection for 48 h in KYSE510 and KYSE180 cell lines. (B) Quantification of colonies measured by absorbance rate of 480 nm (n = 5 biological replicates). (C) Protein levels of MLL3 after viral infection of shRNA into KYSE510 and KYSE180 cell lines. (D) Quantification of colonies in MLL3-silenced versus control KYSE510 and KYSE180 cell lines (n = 5 biological replicates). (E and F) Relative mRNA and protein levels of MLL3 following its knockdown in MOC1 and MOC22 cell lines, measured by qRT-PCR and western blot, respectively. (G) Quantification of colonies in MLL3-silenced MOC1 and MOC22 cell lines (n = 5 biological replicates). (H) Table of top enriched transcription factor–binding motifs in MLL3 ChIP-seq peak regions. (I) Venn diagram displaying the number of overlapped peaks between GRHL2 and MLL3 ChIP-seq in KYSE510 cell line. (J) Co-IP assay by pulling down using the anti-MLL3 antibody followed by detection of GRHL2 protein by western blot. *P < 0.05; **P < 0.01; ***P < 0.001. Co-IP: co-immunoprecipitation. Source data are available for this figure: SourceData F3.
MLL3 inhibits cancer cell proliferation and interacts with GRHL2. (A) Protein levels of MLL3 following siRNA transfection for 48 h in KYSE510 and KYSE180 cell lines. (B) Quantification of colonies measured by absorbance rate of 480 nm (n = 5 biological replicates). (C) Protein levels of MLL3 after viral infection of shRNA into KYSE510 and KYSE180 cell lines. (D) Quantification of colonies in MLL3-silenced versus control KYSE510 and KYSE180 cell lines (n = 5 biological replicates). (E and F) Relative mRNA and protein levels of MLL3 following its knockdown in MOC1 and MOC22 cell lines, measured by qRT-PCR and western blot, respectively. (G) Quantification of colonies in MLL3-silenced MOC1 and MOC22 cell lines (n = 5 biological replicates). (H) Table of top enriched transcription factor–binding motifs in MLL3 ChIP-seq peak regions. (I) Venn diagram displaying the number of overlapped peaks between GRHL2 and MLL3 ChIP-seq in KYSE510 cell line. (J) Co-IP assay by pulling down using the anti-MLL3 antibody followed by detection of GRHL2 protein by western blot. *P < 0.05; **P < 0.01; ***P < 0.001. Co-IP: co-immunoprecipitation. Source data are available for this figure: SourceData F3.
MLL3 and GRHL2 co-bind large numbers of genomic loci and regulate immune response pathways. (A) Quantification of colonies after knockdown of MLL3 (n = 5 biological replicates) in A253 and FADU cell lines. (B) Pie charts showing genomic distribution of MLL3- and GRHL2-binding region in KYSE510 cell line. (C) Heatmap of ChIP-seq peaks of MLL3, GRHL2, H3K27ac, and H3K4me1. (D and E) Gene set enrichment analysis (GSEA) of either MLL3 KD or GRHL2 KD RNA-seq. *P < 0.05; **P < 0.01; and ***P < 0.001.
MLL3 and GRHL2 co-bind large numbers of genomic loci and regulate immune response pathways. (A) Quantification of colonies after knockdown of MLL3 (n = 5 biological replicates) in A253 and FADU cell lines. (B) Pie charts showing genomic distribution of MLL3- and GRHL2-binding region in KYSE510 cell line. (C) Heatmap of ChIP-seq peaks of MLL3, GRHL2, H3K27ac, and H3K4me1. (D and E) Gene set enrichment analysis (GSEA) of either MLL3 KD or GRHL2 KD RNA-seq. *P < 0.05; **P < 0.01; and ***P < 0.001.
To understand the molecular mechanisms underlying the function of MLL3 in UASCC cells, we performed chromatin immunoprecipitation sequencing (ChIP-seq) using a validated ChIP antibody against MLL3 (Wang et al., 2018), considering its canonical role as a histone lysine 3 methyltransferase. We obtained a total of 11,114 MLL3-binding peaks, which were, as expected, distributed in non-promoter regions (Fig. S2 B). Transcription factor motif sequence enrichment analysis identified that top-ranking sequences are recognized by established squamous cell-specific transcription factors, including TP63, AP1 factors, and GRHL2 (Fig. 3 H). This result suggests that MLL3 co-occupies with lineage-specific transcription factors in squamous cells. We were particularly interested in GRHL2, given its reported tumor suppressive roles in cancer (Reese et al., 2019; Cieply et al., 2012). To validate this enrichment, we performed GRHL2 ChIP-seq in the same UASCC cells. Importantly, we found a strong and significant overlap in the occupancy of MLL3 and GRHL2 in H3K27ac+ putative enhancers (Fig. 3 I and Fig. S2 C). This co-occupancy was notably absent in promoter regions, consistent with the function of MLL3 in enhancer elements (Wang et al., 2021; Hu et al., 2013).
The genome-wide co-occupancy between MLL3 and GRHL2 indicates that these two epigenetic regulators may have protein–protein interactions, which were tested by co-immunoprecipitation experiments. Notably, across independent ESCC and HNSCC cell lines, we detected the GRHL2 protein in the MLL3-immunoprecipitated complex (Fig. 3 J), suggesting that these two factors may form protein complexes. The above data also indicate that MLL3 and GRHL2 may cooperatively regulate the biology of UASCC cells.
MLL3/GRHL2 co-regulate immune response pathways in UASCC
To explore MLL3/GRHL2-regulated functions in established UASCC tumors, we first performed RNA sequencing (RNA-seq) to compare gene expression changes upon silencing of either factor (Fig. 4 A). Following MLL3 knockdown, 865 and 1,194 genes were upregulated and downregulated, respectively (false discovery rate <0.05). 37.4% of downregulated genes contained MLL3-binding peaks, significantly higher than either all differentially expressed genes (29.2%) or all actively expressed genes (24.0%). In contrast, MLL3 binding was depleted in the group of upregulated genes, following MLL3 knockdown (17.9%) (Fig. 4 B). These data suggest that downregulated genes upon MLL3 silencing were significantly more likely to be under direct regulation of MLL3, consistent with the established epigenetic role of MLL3 in promoting transcription by enhancer priming and activation. A similar trend was also observed in the RNA-seq data following grainyhead-like transcription factor 2 (GRHL2) silencing (Fig. 4, A and B). We thus next focused on investigating the downregulated gene group upon the silencing of MLL3/GRHL2.
MLL3/GRHL2 co-regulate immune response genes. (A) A volcano plot of up- and downregulated genes from RNA-seq in MLL3 knockdown cells. Genes having binding peaks of MLL3 are marked as red dots, otherwise black dots. A similar volcano plot for GRHL2 knockdown is at the lower panel. (B) Bar graphs of the fraction of different groups of genes with binding peaks of MLL3 (left) or GRHL2 (right). (C) A Venn diagram illustrating shared downregulated genes between MLL3 knockdown and GRHL2 knockdown. A pie chart showing MLL3- and/or GRHL2-binding peaks on the 505 overlapped downregulated genes. (D) Top enriched signaling pathway of the 505 genes. (E) Gene set enrichment analysis (GSEA) plots showing the enrichment of IFN signaling in the RNA-seq upon knockdown of either MLL3 or GRHL2. P values were adjusted for multiple comparisons.
MLL3/GRHL2 co-regulate immune response genes. (A) A volcano plot of up- and downregulated genes from RNA-seq in MLL3 knockdown cells. Genes having binding peaks of MLL3 are marked as red dots, otherwise black dots. A similar volcano plot for GRHL2 knockdown is at the lower panel. (B) Bar graphs of the fraction of different groups of genes with binding peaks of MLL3 (left) or GRHL2 (right). (C) A Venn diagram illustrating shared downregulated genes between MLL3 knockdown and GRHL2 knockdown. A pie chart showing MLL3- and/or GRHL2-binding peaks on the 505 overlapped downregulated genes. (D) Top enriched signaling pathway of the 505 genes. (E) Gene set enrichment analysis (GSEA) plots showing the enrichment of IFN signaling in the RNA-seq upon knockdown of either MLL3 or GRHL2. P values were adjusted for multiple comparisons.
Importantly, there was a striking overlap between the downregulated genes following knockdown of MLL3 and GRHL2 (Fig. 4 C, P < 1e-10), further corroborating our prior observations that these two factors formed a protein complex and co-occupied a large number of genes. Indeed, among the 505 co-downregulated genes (Table S3), over one-third (35.3%) were co-occupied by both proteins (Fig. 4 C). We next performed pathway enrichment analysis of the co-downregulated genes. Surprisingly, all top five most significantly enriched pathways belong to immune response functions (Fig. 4 D). These immune response pathways include “cytokine signaling in immune system,” “IFN signaling,” “immune system,” etc. Each of these pathways was also significantly enriched in individual RNA-seq data (Fig. 4 E and Fig. S2, D and E), suggesting the converged regulation of MLL3 and GRHL2 on these immune response pathways.
The above data prompted us to explore the functional significance of MLL3/GRHL2 in regulating the anti-tumor immune response, initially using the aforementioned syngeneic murine HNSCC cell lines (MOC1 and MOC22). We first established stable murine lines with MLL3/GRHL2 knockdown, which were then implanted by subcutaneous injection in immune-competent C57BL/6 mice. In line with in vitro results, mice bearing MLL3-knockdown tumors had shorter survival than control mice (Fig. S3 A). After tumor formation and growth, we analyzed intratumoral immune cells by flow cytometry (Fig. 5 A and Fig. S3 B). Importantly, MLL3 knockdown led to markedly decreased infiltration of both CD4+ and CD8+ T lymphocytes in both MOC1 and MOC22 models (Fig. 5, B–E). The reduction of intratumoral CD8+ T cells was consistently observed in GRHL2-knockdown syngeneic models. Congruently, immunofluorescence (IF) staining of tumor slices demonstrated that MLL3 or GRHL2 knockdown reduced the infiltration of CD8+ T cells in the tumor microenvironment (Fig. 5 F). It was notable that CD8+ T cells exhibited such marked changes since they are not only the soldiers responsible for executing tumor killing but also the primary target of immune-checkpoint blockade (ICB) therapies. Thus, we next focused on analyzing further these T cells.
MLL3/GRHL2 knockdown accelerates tumor growth and reduces T cell infiltration. (A) Kaplan–Meier survival plots of control and MLL3-knockdown groups in MOC22 xenografts models (n = 3 biological replicates). Mice were sacrificed when tumor reached 1.5 cm. (B) Gating strategy for flow cytometry of Fig. 5, B and D. The cell population was first gated on SSC-A/FSC-A and FSC-H/FSC-A parameters, and then Zombie Violet dye was used to exclude dead cells. Next, CD45+CD3+-positive T cells were gated to isolate the T cell population. Finally, we measured the proportions of CD8+ and CD4+ T cells using the following equation: and .
MLL3/GRHL2 knockdown accelerates tumor growth and reduces T cell infiltration. (A) Kaplan–Meier survival plots of control and MLL3-knockdown groups in MOC22 xenografts models (n = 3 biological replicates). Mice were sacrificed when tumor reached 1.5 cm. (B) Gating strategy for flow cytometry of Fig. 5, B and D. The cell population was first gated on SSC-A/FSC-A and FSC-H/FSC-A parameters, and then Zombie Violet dye was used to exclude dead cells. Next, CD45+CD3+-positive T cells were gated to isolate the T cell population. Finally, we measured the proportions of CD8+ and CD4+ T cells using the following equation: and .
Loss of MLL3/GRHL2 attenuates T cell infiltration in the tumor microenvironment. (A) Schematic illustration of the workflow for the investigation of infiltrated T cell infiltration in syngeneic mouse models. (B and D) FACS plots of Zombie−CD45+CD3+CD4−CD8+T cells and Zombie−CD45+CD3+CD4+CD8− T cells in indicated samples. The left-most panel of B was also presented as the last gating plot (CD8 versus CD4) in Fig. S3 B. (C and E) Bar graph of the percentage of CD8+ T cells and CD4+ T cells among live cells (n = 5 for each group). (F) IF staining of infiltrated CD8+ T cells in the xenograft tumors and bar graph of quantification of CD8+ T cell (n = 3 biological replicates). Scale bar = 100 μm. (G) Schematic illustration of the workflow of T cell migration assay using transwell system. Sorted CD8+ T cells were cultured for 48 h using conditioned medium harvested from either MLL3 KD or GRHL2 KD cell line. (H and I) The number of migrated CD8+ T cells into the bottom counted by hemocytometer (n = 3 biological replicates). *P < 0.05; **P < 0.01; ***P < 0.001.
Loss of MLL3/GRHL2 attenuates T cell infiltration in the tumor microenvironment. (A) Schematic illustration of the workflow for the investigation of infiltrated T cell infiltration in syngeneic mouse models. (B and D) FACS plots of Zombie−CD45+CD3+CD4−CD8+T cells and Zombie−CD45+CD3+CD4+CD8− T cells in indicated samples. The left-most panel of B was also presented as the last gating plot (CD8 versus CD4) in Fig. S3 B. (C and E) Bar graph of the percentage of CD8+ T cells and CD4+ T cells among live cells (n = 5 for each group). (F) IF staining of infiltrated CD8+ T cells in the xenograft tumors and bar graph of quantification of CD8+ T cell (n = 3 biological replicates). Scale bar = 100 μm. (G) Schematic illustration of the workflow of T cell migration assay using transwell system. Sorted CD8+ T cells were cultured for 48 h using conditioned medium harvested from either MLL3 KD or GRHL2 KD cell line. (H and I) The number of migrated CD8+ T cells into the bottom counted by hemocytometer (n = 3 biological replicates). *P < 0.05; **P < 0.01; ***P < 0.001.
To substantiate these in vivo observations, we next performed an in vitro T cell migration assay, using conditioned medium from either control, MLL3- or GRHL2-silenced cells (Fig. 5 G). Knockdown of either MLL3 or GRHL2 consistently resulted in decreased T cell migration in vitro (Fig. 5, H and I). These data together suggest that MLL3 and GRHL2 promote T cell infiltration into the UASCC tumor microenvironment.
MLL3/GRHL2 promote IFN signaling via co-activating the transcription of IRF1
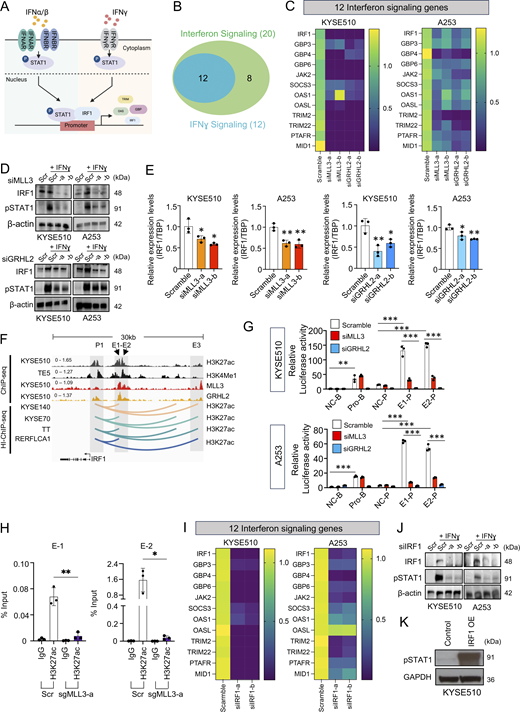
To understand how MLL3/GRHL2 promote T cell infiltration into the tumor microenvironment, we focused on the IFN signaling pathways (Fig. 6 A) because: (1) they ranked third and fifth most significantly downregulated pathways in MLL3/GRHL2-knockdown cells (Fig. 4 D) and (2) they play an essential role in recruiting T cells into the tumor microenvironment (Cheon et al., 2014). We identified 12 shared genes enriched in both IFN and IFNγ signaling pathways from our RNA-seq data (Fig. 6 B). Independent quantitative real-time PCR (qRT-PCR) assays of this 12-gene panel confirmed that silencing of either MLL3 or GRHL2 decreased the IFN signaling (Fig. 6 C) in ESCC and HNSCC cell lines.
MLL3/GRHL2 regulate IFN signaling through IRF1. (A) Schematic illustration of IFN signaling pathways. (B) Venn diagram of enriched genes between IFN signaling and IFNγ signaling in RNA-seq data following knockdown of either MLL3 or GRHL2. (C) Heatmap of mRNA expression of 12 overlapped genes upon knockdown of either MLL3 or GRHL2. (D) Protein levels of IRF1 and pSTAT1 in MLL3 or GRHL2 knockdown cells treated with IFNγ (10 ng/ml). (E) Relative mRNA levels of IRF1 upon knockdown of either MLL3 or GRHL2. (F) Integrative genomics viewer (IGV) tracks and line plots showing respective ChIP-seq profiles and H3K27ac high-throughput (HI)-ChIP loops at the locus of IRF1 gene. (G) Luciferase reporter assay after knockdown of either MLL3 or GRHL2. NC-B, negative control pGL3-basic vector; Pro-B, pGL3-basic with IRF1 promoter; NC-P, negative control pGL3-promoter vector; E1-P, pGL3-promoter with enhancer-1; and E2-P, pGL3-promoter with enhancer-2. (H) H3K27ac ChIP-qPCR of E1 and E2 regions in control and MLL3 knockout cells. (I) Heatmap of mRNA expression of 12 genes after silencing IRF1 in the presence of IFNγ (10 ng/ml). (J) Protein levels of IRF1 and pSTAT1 upon knockdown of IRF1 in the absence or presence of IFNγ (10 ng/ml). (K) Protein levels of pSTAT1 after overexpression of IRF1. n = 3 biological replicates for panels C, E, and G–I. *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData F6.
MLL3/GRHL2 regulate IFN signaling through IRF1. (A) Schematic illustration of IFN signaling pathways. (B) Venn diagram of enriched genes between IFN signaling and IFNγ signaling in RNA-seq data following knockdown of either MLL3 or GRHL2. (C) Heatmap of mRNA expression of 12 overlapped genes upon knockdown of either MLL3 or GRHL2. (D) Protein levels of IRF1 and pSTAT1 in MLL3 or GRHL2 knockdown cells treated with IFNγ (10 ng/ml). (E) Relative mRNA levels of IRF1 upon knockdown of either MLL3 or GRHL2. (F) Integrative genomics viewer (IGV) tracks and line plots showing respective ChIP-seq profiles and H3K27ac high-throughput (HI)-ChIP loops at the locus of IRF1 gene. (G) Luciferase reporter assay after knockdown of either MLL3 or GRHL2. NC-B, negative control pGL3-basic vector; Pro-B, pGL3-basic with IRF1 promoter; NC-P, negative control pGL3-promoter vector; E1-P, pGL3-promoter with enhancer-1; and E2-P, pGL3-promoter with enhancer-2. (H) H3K27ac ChIP-qPCR of E1 and E2 regions in control and MLL3 knockout cells. (I) Heatmap of mRNA expression of 12 genes after silencing IRF1 in the presence of IFNγ (10 ng/ml). (J) Protein levels of IRF1 and pSTAT1 upon knockdown of IRF1 in the absence or presence of IFNγ (10 ng/ml). (K) Protein levels of pSTAT1 after overexpression of IRF1. n = 3 biological replicates for panels C, E, and G–I. *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData F6.
IFN regulatory factor (IRF) and signal transducer and activator of transcription 1 (STAT) family members are central factors in mediating the IFN signaling (Parker et al., 2016). Notably, within IRF and STAT family genes, IFN regulatory factor 1 (IRF1) was among the 12 enriched IFN genes downregulated upon MLL3 or GRHL2 knockdown (Fig. 6 C and Fig. S4 A). We thus hypothesized that MLL3 and GRHL2 might promote the IFN signaling by controlling IRF1 expression. As expected, stimulation of cells with IFNγ potently elevated the levels of IRF1 and phosphorylation of STAT1 (Fig. 6 D and Fig. S4 B). We validated that silencing of either MLL3 or GRHL2 downregulated IRF1 expression, with an accompanying reduction of STAT1 phosphorylation. These changes were also confirmed at the mRNA levels (Fig. 6 E). Notably, among three putative H3k27ac+ enhancers (E1, E2, E3 in Fig. 6 F) of IRF1, two (E1 and E2) had MLL3 and GRHL2 co-binding peaks. These distal elements have direct contact with the IRF1 promoter, as shown by H3K27ac HiChIP loops. To test the transcriptional activities of these DNA regulatory elements, we cloned them into the PGL3-based reporter system and performed luciferase reporter assays. Both enhancers (E1 and E2) harbored significantly stronger reporter activities than the promoter (greater than three to fourfold) (Fig. 6 G). Moreover, MLL3 or GRHL2 knockdown potently reduced reporter activities of both enhancers. To further test the function of E1 and E2, we utilized the dCas9–KRAB system to inhibit the transcriptional activity of these enhancers. Importantly, we found that blocking either E1 or E2 significantly reduced the expression level of IRF1 and decreased secretion of CXCL9 and CXCL10 (Fig. S4, C–E), suggesting that MLL3 and GRHL2 regulate IRF1 transcription by directly activating these two distal enhancers. In addition, we performed ChIP-qPCR using an H3K27ac antibody to measure the H3K27ac intensity of both enhancers. Consistently, knockout of MLL3 by CRISPR/Cas9 strongly decreased the levels of H3K27ac in both E1 and E2 (Fig. 6 H).
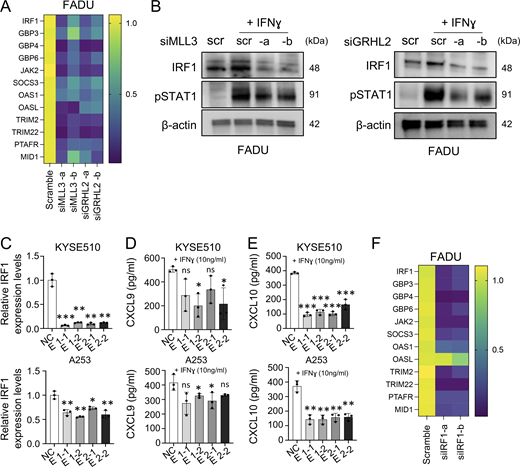
MLL3/GRHL2 directly regulates IRF1 expression levels. (A) Heatmap of 12 IFNγ-related genes after knockdown of either MLL3 or GRHL2 (n = 3 biological replicates). (B) Protein levels of IRF1 and p-STAT1 after silencing of either MLL3 or GRHL2 with treatment of IFNγ (10 ng/ml) for 48 h in FADU cell line. (C) Expression levels of IRF1 following dCas9-KRAB–mediated inhibition of either E1 or E2 in A253 and KYSE510 cell lines. Two independent sgRNAs were tested for each enhancer (n = 3 biological replicates). (D and E) Secretion levels of CXCL9 (D) and CXCL10 (E) were measured by the ELISA assay. NC, negative control; E1-1, sgRNA1 for enhancer-1; E1-2, sgRNA2 for enhancer-1; E2-1, sgRNA1 for enhancer-2; and E2-2, sgRNA2 for enhancer-2 (n = 3 biological replicates). (F) Heatmap of 12 IFN-related genes after silencing of IRF1 and treating IFNγ (10 ng/ml) in FADU cell lines (n = 3 biological replicates). *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData FS4.
MLL3/GRHL2 directly regulates IRF1 expression levels. (A) Heatmap of 12 IFNγ-related genes after knockdown of either MLL3 or GRHL2 (n = 3 biological replicates). (B) Protein levels of IRF1 and p-STAT1 after silencing of either MLL3 or GRHL2 with treatment of IFNγ (10 ng/ml) for 48 h in FADU cell line. (C) Expression levels of IRF1 following dCas9-KRAB–mediated inhibition of either E1 or E2 in A253 and KYSE510 cell lines. Two independent sgRNAs were tested for each enhancer (n = 3 biological replicates). (D and E) Secretion levels of CXCL9 (D) and CXCL10 (E) were measured by the ELISA assay. NC, negative control; E1-1, sgRNA1 for enhancer-1; E1-2, sgRNA2 for enhancer-1; E2-1, sgRNA1 for enhancer-2; and E2-2, sgRNA2 for enhancer-2 (n = 3 biological replicates). (F) Heatmap of 12 IFN-related genes after silencing of IRF1 and treating IFNγ (10 ng/ml) in FADU cell lines (n = 3 biological replicates). *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData FS4.
To validate the role of IRF1 in mediating the IFN signaling, we silenced the expression of IRF1, which downregulated the enriched genes of the IFN signaling (Fig. 6 I and Fig. S4 F). Moreover, silencing of IRF1 also decreased the phosphorylation of STAT1 (Fig. 6 J), further confirming the weakened IFN signaling. We next ectopically expressed IRF1, which increased the phosphorylation of STAT1 (Fig. 6 K). These results together identify a novel MLL3/GRHL2-IRF1 axis in UASCC cells, which regulates the IFN signaling.
The MLL3/GRHL2-IRF1 axis promotes the expression of Th1 chemokines
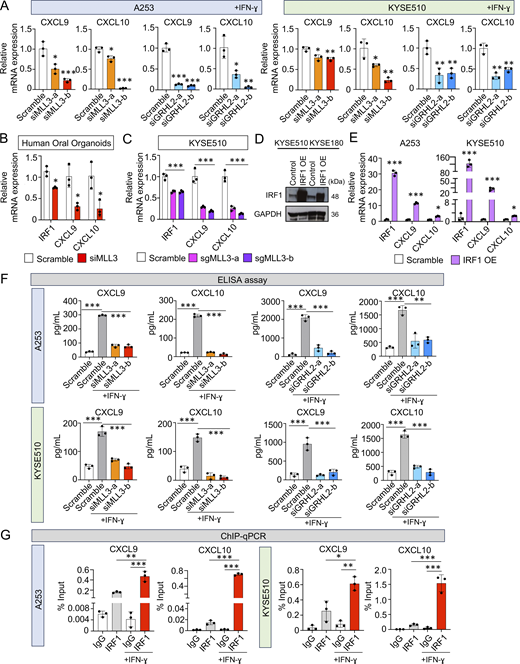
It is crucial to understand mechanistically how the MLL3/GRHL2-IRF1 axis regulates immune response, especially the intra-tumor T cell infiltration observed above (Fig. 5). T cell trafficking and recruitment are known to be predominantly regulated by Th1 chemokines, particularly CXCL9, CXCL10, and CXCL11 (Franciszkiewicz et al., 2012; Tokunaga et al., 2018), which are also canonical IFN signaling target genes. Since our data have shown that the MLL3/GRHL2-IRF1 axis positively regulates the IFN signaling (Fig. 6), we assessed whether this axis controls the expression of these Th1 chemokines. Notably, knockdown of either MLL3, GRHL2, or IRF1 consistently downregulated the mRNA levels of CXCL9 and CXCL10 (Fig. 7 A and Fig. S5, A and B). These results were reproduced in human oral organoids and MLL3 knockout cell lines (Fig. 7, B and C), as well as in the xenograft samples (Fig. S5 C). Moreover, overexpression of IRF1 elevated the mRNA expression of these Th1 chemokines by 10–20-fold (Fig. 7, D and E). To confirm the regulation at the protein level, we performed ELISA assays and verified that the amount of secreted proteins of Th1 chemokines was consistently reduced upon depletion of MLL3 or GRHL2 (Fig. 7 F and Fig. S5 D). We confirmed that these regulations were conserved across human and murine UASCC cells (Fig. S5 E). Furthermore, IRF1 overexpession promoted the secretion of CXCL9 and CXCL10 and increased T cell migration, which mitigated the suppressive effects induced by MLL3 knockdown (Fig. S5, F–H). We also found that IRF1 inhibited cell proliferation (Fig. S5 I), consistent with the antiproliferative function of MLL3.
The MLL3/GRHL2-IRF1 axis regulates the expression of chemokines CXCL9 and CXCL10. (A–C) Relative mRNA expression levels of CXCL9/10 in indicated cell line and organoid samples upon silencing of either MLL3 or GRHL2 in the presence of IFNγ (10 ng/ml). (D) Protein levels of IRF1 after overexpression of IRF1. (E) mRNA expression levels of CXCL9 and CXCL10 upon overexpression of IRF1. (F) ELISA assay of CXCL9/10 upon knockdown of either MLL3 or GRHL2 in the absence or presence of IFNγ (10 ng/ml). (G) IRF1 ChIP-qPCR in the absence or presence of IFNγ (10 ng/ml). n = 3 biological replicates for A–C and E–G. *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData F7.
The MLL3/GRHL2-IRF1 axis regulates the expression of chemokines CXCL9 and CXCL10. (A–C) Relative mRNA expression levels of CXCL9/10 in indicated cell line and organoid samples upon silencing of either MLL3 or GRHL2 in the presence of IFNγ (10 ng/ml). (D) Protein levels of IRF1 after overexpression of IRF1. (E) mRNA expression levels of CXCL9 and CXCL10 upon overexpression of IRF1. (F) ELISA assay of CXCL9/10 upon knockdown of either MLL3 or GRHL2 in the absence or presence of IFNγ (10 ng/ml). (G) IRF1 ChIP-qPCR in the absence or presence of IFNγ (10 ng/ml). n = 3 biological replicates for A–C and E–G. *P < 0.05; **P < 0.01; and ***P < 0.001. Source data are available for this figure: SourceData F7.
Downregulation of MLL3/GRHL2 decreases CXCL9/10 levels through regulation of IRF1 expression. (A) Relative mRNA expression levels of CXCL9/10 after knockdown of either MLL3 or GRHL2 in the presence of IFNγ (10 ng/ml) in murine cell lines (n = 3 biological replicates). (B) Relative mRNA expression of IRF1 and CXCL9/10 upon knockdown of IRF1 in A253, FADU, KYSE510, and KYSE180 cell lines (n = 3 biological replicates). All samples were treated with IFNγ (10 ng/ml). (C) Expression levels of IRF1, CXCL9, and CXCL10 in scramble and MLL3-knockdown xenograft samples. (n = 5 for each group). (D and E) ELISA assay of CXCL9 and CXCL10 levels in either MLL3 or GRHL2 silenced FADU, KYSE180, MOC1, and MOC22 cell lines (n = 3 biological replicates). (F and G) CXCL9 and CXCL10 protein levels were detected by ELISA assay (n = 3 biological replicates). (H) Quantification of migrated CD8+ T cells in the bottom well in a transwell assay (n = 3 biological replicates). (I) Cell proliferation rates upon IRF1 knockdown in murine oral organoids (TP53/CDKN2ADKO) or IRF1 overexpression in MOC1 cells (n = 3 biological replicates). (J) Quantification of migrated CD8+ T cell into bottom well in a transwell assay following anti-CXCR3 treatment (100 ng/ml) (n = 3 biological replicates). (K) Prediction of transcription factor (TF)-binding sites in CXCL9 and CXCL10 using JASPER. *P < 0.05; **P < 0.01; and ***P < 0.001.
Downregulation of MLL3/GRHL2 decreases CXCL9/10 levels through regulation of IRF1 expression. (A) Relative mRNA expression levels of CXCL9/10 after knockdown of either MLL3 or GRHL2 in the presence of IFNγ (10 ng/ml) in murine cell lines (n = 3 biological replicates). (B) Relative mRNA expression of IRF1 and CXCL9/10 upon knockdown of IRF1 in A253, FADU, KYSE510, and KYSE180 cell lines (n = 3 biological replicates). All samples were treated with IFNγ (10 ng/ml). (C) Expression levels of IRF1, CXCL9, and CXCL10 in scramble and MLL3-knockdown xenograft samples. (n = 5 for each group). (D and E) ELISA assay of CXCL9 and CXCL10 levels in either MLL3 or GRHL2 silenced FADU, KYSE180, MOC1, and MOC22 cell lines (n = 3 biological replicates). (F and G) CXCL9 and CXCL10 protein levels were detected by ELISA assay (n = 3 biological replicates). (H) Quantification of migrated CD8+ T cells in the bottom well in a transwell assay (n = 3 biological replicates). (I) Cell proliferation rates upon IRF1 knockdown in murine oral organoids (TP53/CDKN2ADKO) or IRF1 overexpression in MOC1 cells (n = 3 biological replicates). (J) Quantification of migrated CD8+ T cell into bottom well in a transwell assay following anti-CXCR3 treatment (100 ng/ml) (n = 3 biological replicates). (K) Prediction of transcription factor (TF)-binding sites in CXCL9 and CXCL10 using JASPER. *P < 0.05; **P < 0.01; and ***P < 0.001.
Considering that CXCL9/10 signal through binding to C-X-C motif chemokine receptor 3 (CXCR3) on T cells, we further performed loss-of-function experiments using the anti-CXCR3 antibody. The results demonstrated that CXCR3 blockade in T cells strongly inhibited their migration toward cancer cells (Fig. S5 J).
The above results demonstrate that MLL3 and GRHL2 promote the expression of CXCL9 and CXCL10 through IRF1. To connect the transcriptional activity of IRF1 to the expression of Th1 chemokines, we applied the JASPER (Castro-Mondragon et al., 2022) method and predicted direct binding sites of IRF1 on the promoters of both CXCL9 and CXCL10 (Fig. S5 K). To validate this prediction, we performed ChIP-qPCR using an IRF1 antibody, which revealed direct binding of IRF1 on their promoter regions (Fig. 7 G). Moreover, the occupancy of IRF1 was significantly enhanced by the stimulation of IFNγ. These results together demonstrate that the MLL3/GRHL2-IRF1 axis directly activates the transcription of CXCL9 and CXCL10 genes.
MLL3 regulates the efficacy of ICB therapy in murine SCC models
Given the prominent function of MLL3 in regulating the IFN pathway and CD8+ T cell infiltration in the tumor microenvironment, we asked whether MLL3 loss could affect the efficacy of ICB therapy, such as anti-PD-1 mAb therapy, which reinvigorates CD8+ T cells. Before addressing this, we recently confirmed significantly higher expression of CD274 (PD-L1) in UASCC tumors compared with matched normal tissues from TCGA cohorts (Jiang et al., 2024b). We orthotopically injected either scramble or shMLL3 MOC1 cells into the cheek of syngeneic immune-competent C57BL/6 mice transcutaneously, allowing for the characterization of the oral tumor microenvironment. These tumor-bearing mice were treated with either PD-1 mAb or vehicle control once the tumors reached an appropriate size (Fig. 8 A).
MLL3 regulates the efficacy of ICB therapy in HNSCC murine models. (A) Schematic illustration of the workflow of anti-PD1 treatment using a syngeneic orthotopic HNSCC model. (B) Kaplan–Meier survival plot of orthotopic models. Mice were euthanized due to either maximum tumor volume (a diameter of 15 mm in any direction) or tumor-associated morbidity. (C) Tumor growth curves for each group of mice (n = 5 mice in each group). (D) Flow cytometry measuring the fraction of tumor-infiltrated CD8+ T cells. (E) Box plots of the percentage of tumor-infiltrated CD8+ T cells and CD8+ IFNγ+ T cells among live cells in each group (n = 5 mice in each group). ***P < 0.001. (F) Kaplan–Meier survival plot of a subcutaneous syngeneic model with a similar treatment plan. (G) Individual tumor growth curves (n = 6 tumors in each group). (H and I) Survival analyses of the OAK/POPLAR cohorts of lung SCC patients treated with either ICB or chemotherapy. Patients were classified based on the mean value of MLL3 expression.
MLL3 regulates the efficacy of ICB therapy in HNSCC murine models. (A) Schematic illustration of the workflow of anti-PD1 treatment using a syngeneic orthotopic HNSCC model. (B) Kaplan–Meier survival plot of orthotopic models. Mice were euthanized due to either maximum tumor volume (a diameter of 15 mm in any direction) or tumor-associated morbidity. (C) Tumor growth curves for each group of mice (n = 5 mice in each group). (D) Flow cytometry measuring the fraction of tumor-infiltrated CD8+ T cells. (E) Box plots of the percentage of tumor-infiltrated CD8+ T cells and CD8+ IFNγ+ T cells among live cells in each group (n = 5 mice in each group). ***P < 0.001. (F) Kaplan–Meier survival plot of a subcutaneous syngeneic model with a similar treatment plan. (G) Individual tumor growth curves (n = 6 tumors in each group). (H and I) Survival analyses of the OAK/POPLAR cohorts of lung SCC patients treated with either ICB or chemotherapy. Patients were classified based on the mean value of MLL3 expression.
In agreement with the tumor-intrinsic role of MLL3, its loss increased the tumor volume compared with the scramble control group in the absence of the anti-PD-1 treatment. PD-1 mAb treatment impaired tumor growth and extended mouse survival in the scramble group (Fig. 8, B and C). However, the PD-1 mAb therapy produced almost no anti-tumor effects in the MLL3-knockdown group. We further analyzed intratumoral T cells from each group of mice using flow cytometry (Fig. 8, D and E). We confirmed that PD-1 mAb therapy increased the T cell abundance and enhanced their effector functionality, as indicated by the increased IFN-γ staining. Importantly, MLL3-loss tumors were refractory to the PD-1 mAb treatment, showing no changes in either T cell infiltration or activity. In an independent subcutaneous syngeneic model, we reproduced these results, showing that MLL3 loss impaired the anti-tumor effect of PD-1 mAb and reduced CD8+ T cell intratumoral infiltration and activity (Fig. 8, F and G).
To further explore the clinical significance of MLL3 in ICB therapy, we asked whether MLL3 expression was associated with response to ICB therapy in UASCC patients. In two lung squamous cancer cohorts (known as OAK [Rittmeyer et al., 2017] and POPLAR [Fehrenbacher et al., 2016] trials) treated with either ICB (anti-PD-L1) or chemotherapy (docetaxel), high MLL3 expression was strongly and specifically associated with better survival in the ICB arm but not the chemotherapy arm (Fig. 8, H and I). Concordant findings across these two independent clinical cohorts highlight a predictive value of MLL3 expression for the ICB response in squamous cancer.
Discussion
This study investigates the early neoplastic transformation of (UASCC and first bioinformatically identifies MLL3 mutations as crucial drivers of tumor initiation using phylogenetic analysis of multi-region whole exome sequencing (WES) data. To functionally pursue this computational finding, we developed a genetically engineered cross-species 3D oral organoid model, overcoming the scarcity of valid ex vivo models for studying the premalignant transitional characteristics of UASCC. The 3D organoid culture system recapitulates and maintains genetic, biological, and phenotypic characteristics of corresponding tissues of origin (Lo et al., 2020). Thus, organoid modeling provides a robust and powerful platform to investigate premalignant biology and early tumorigenesis. Moreover, organoid culturing has made it feasible and efficient to genetically manipulate cancer cells of origin directly from human samples. Leveraging this innovative model system, we were able to investigate key events occurring during the initiation and early stages of UASCC development, revealing the critical role of MLL3 loss in driving this cancer’s early pathogenesis.
In light of prior findings demonstrating the interaction between the pioneer transcription factor FOXA1 and GRHL2, which facilitates the recruitment of MLL3 to enhancer binding sites in MCF7 cells (Jozwik et al., 2016), our study adds further depth to the understanding of the functional interplay between MLL3 and GRHL2 in UASCC. While the interaction between GRHL2 and FOXA1 elucidates a mechanism for MLL3 recruitment to enhancer regions in breast cancer cells, our research reveals a novel protein complex formed by MLL3 and GRHL2 in the context of UASCC. This suggests a broader role for GRHL2 in facilitating the assembly of epigenetic regulatory complexes across different cancer types. Further investigations into the dynamic interactions between MLL3, GRHL2, and other regulatory factors may unveil additional layers of complexity in UASCC epigenomics.
Unexpectedly, our bioinformatic analyses unbiasedly identified that MLL3/GRHL2-regulated top pathways were all associated with immune responses in UASCC. Indeed, our cellular and animal experiments validate that this complex plays a critical role in anti-tumor immune responses by regulating immune pathways such as IFN signaling. Supportively, in a previous study on acute myeloid leukemia (Chen et al., 2014), gene ontology analysis revealed that genes downregulated in response to MLL3 knockdown were strongly enriched in cellular processes, including “immune response,” “T cell differentiation,” and “regulation of leukocyte activation.” While functional assays on immune regulation were not pursued in this prior study, these transcriptional signatures are in line with our present findings, indicating a conserved function of MLL3 in modulating immune responses. Moreover, KMT2D, a paralogous protein of MLL3, was recently shown to positively regulate immune signaling pathways, including the IFN signaling, in HNSCC (Callahan et al., 2021, Preprint). Intriguingly, in the context of regulating epithelial–mesenchymal transition in late-stage breast cancer, loss of MLL3 was found to lead to elevated IFNγ signaling, contributing to the induction of hybrid epithelial–mesenchymal transition cells and enhanced metastatic capacity (Cui et al., 2023). These findings suggest a context-dependent, multifaceted role of MLL3 in cancer progression and metastasis.
ICB therapies have been approved by the FDA for the treatment of HNSCC and ESCC. However, the majority (65–70%) of these patients do not exhibit meaningful clinical response (Burtness et al., 2019; Ferris et al., 2016). Therefore, further understanding of the regulation of anti-tumor immunity is critical to identify features associated with responsiveness, or the lack thereof, to current immunotherapies and will inform the development of effective single- and multi-agent immunotherapeutic regimens. Notably, the efficacy of ICB therapy depends crucially on tumor-infiltrating T cells (Litchfield et al., 2021; Gao et al., 2016; Zaretsky et al., 2016; Tumeh et al., 2014; Hellmann et al., 2018). In fact, the abundance of intratumoral T cells is among the most reliable predictors of effectiveness of ICB therapies. For example, one of the earlier studies revealed that the response of ICB therapy was directly related to the degree of intratumoral CD8+ T cells (Balatoni et al., 2018), a finding later reproduced across many different cancer types (Zheng et al., 2021; Tumeh et al., 2014). Based on our finding that MLL3/GRHL2 enhances intratumoral CD8+ T cell infiltration via activation of Th1 chemokines, we hypothesized that MLL3 regulates the efficacy of ICB therapy. Indeed, in both orthotopic and subcutaneous syngeneic models established using HNSCC cell lines, we found that MLL3 loss impairs the anti-tumor effect of PD-1 mAb by reducing CD8+T cell intratumoral infiltration and activity. Nevertheless, certain limitations of this regulation should be acknowledged. In particular, the evidence implicating the IRF1-CXCL9/CXCL10 axis as a key downstream mediator of MLL3 remains indirect, as the simplified in vitro system does not fully replicate the complexity of in vivo tumor–immune interactions. Future studies are required to address these limitations using in vivo perturbation experiments.
In conclusion, this study contributes to our understanding of UASCC, shedding light on the early genetic events that drive its development and progression. The role of MLL3 and its interaction with GRHL2 in regulating immune responses in UASCC has been elucidated. Furthermore, our results have implications for future therapeutic strategies, providing a foundation for further research in the field of upper aerodigestive squamous cancer.
Materials and methods
Human experimental guidelines approval statement
This research was performed in accordance with the principles of the Declaration of Helsinki. All procedures involving human samples were approved by the Institutional Review Board of the University of Southern California (study #HS-14-00267).
Cell lines and cell cultures
Human HNSCC cell lines, A253 and FADU, were obtained from the ATCC. Mouse HNSCC cell lines, MOC1 and MOC22, were purchased from Kerafast. A253 and FADU cell lines were cultured in McCoy’s 5A medium (#10-050-CV; Corning) and DMEM medium (#10-013-CV; Corning), respectively. Human ESCC cell lines, KYSE510 and KYSE180, were cultured in RPMI-1640 medium (#10-040-CV; Corning). MOC1 and MOC22 cell lines were cultured in IMDM medium (#SH30228.1; Hyclone). All media were supplemented with 10% FBS (#FB-02; Omega Scientific) and 1% penicillin-streptomycin sulfate (#30-002-CI; Thermo Fisher Scientific). Cell cultures were maintained in a 37°C incubator with 5% CO2. All cell lines were tested for mycoplasma contamination and verified using short tandem repeat analysis.
Antibodies and reagents
The following antibodies and reagents were utilized in the study: the anti-MLL3 antibody for ChIP-seq was kindly provided by Dr. Lu Wang at Northwestern University, Evanston, IL, USA, anti-MLL3 (ABE1851; Millipore Sigma, 1:500 for western blotting), anti-GRHL2 (#GTX109410; Genetex, 1:1,1000 for western blotting and 5 μg for ChIP), anti-IRF1 (#11335-1-AP; Proteintech, 1:1,000 for western blotting and 5 μg for ChIP-qPCR), anti-β-actin (#JLA20; Developmental Studies Hybridoma Bank, 1:5,000 for western blotting), anti-phospho STAT1 (#7649; Cell Signaling Technology, 1:1,000 for western blotting), anti-H3K27ac (#39034; Active Motif, 5 μg for ChIP-qPCR), goat anti-rabbit IgG, HRP conjugated (#12–348; Millipore Sigma, 1:5,000 for western blotting), goat anti-mouse IgG, HRP conjugated (#G21040; Invitrogen, 1:5,000 for western blotting), Lipofectamine RNAiMAX (#13778150; Thermo Fisher Scientific), Lipofectamine 3000 (#L3000015; Thermo Fisher Scientific), BioT transfection reagent (#B01-01; Bioland Scientific), Fc blocker (#553141; BD Pharmingen, 0.1 μg per sample), Brilliant violet 605-conjugated anti-mouse CD45 antibody (#103155; Biolegend), Percp/cyanine 5.5-conjugated anti-mouse CD3 antibody (#100217; Biolegend), FITC-conjugated anti-mouse CD4 antibody (#100405; Biolegend), PE-conjugated anti-mouse CD8a antibody (#100707; Biolegend), Alexa fluor 700-conjugated anti-mouse IFNγ (#505823; Biolegend), Zombie violet Fixable viability kit (#423113; Biolegend), rabbit polyclonal anti-Ki67 antibody (1:500, #ab15580; Abcam), rat monoclonal anti-CD8a antibody (1:500, #ab22378; Abcam), goat anti-rabbit IgG (H+L) secondary antibody conjugated with Alexa Fluor 568 conjugated (1:2,000, #ab175471; Abcam), goat anti-rat IgG (H+L) secondary antibody conjugated with Alexa Fluor 647 conjugated (1:2,000, #A-21245; Invitrogen), fluoroshield with DAPI (#F6057; Millipore Sigma), recombinant mouse IFNγ (#ab9922; Abcam), recombinant human IFNγ (#ab9659; Abcam), cell proliferation reagent WST-1 assay kit (#5015944001; Roche), MojoSort Mouse CD8+ T cell isolation kit (#480035; Biolegend), ELISA kits for human CXCL9 (#DY392-05; R&D Systems), human CXCL10 (#DY266-05; R&D Systems), mouse CXCL9 (#DY492-05; R&D Systems), mouse CXCL10 (#DY466-05; R&D Systems), Co-Immunoprecipitation Kit (#26149; Thermo Fisher Scientific), anti-mouse CXCR3 (#BE0249; InVivoMab), recombinant mouse CXCL9 (#492-MM-050/CF; R&D Systems), and recombinant mouse CXCL10 (#466-CR-050/CF; R&D Systems).
The following reagents were used for the culture of organoids: Cultrex RGF basement membrane extract type 2 (BME) (#3533005-02; R&D Systems), TrypLE Express (#12604013; Thermo Fisher Scientific), collagenase from Clostridium histolyticum (#C7657; Sigma-Aldrich), Dispase II (#17105041; Thermo Fisher Scientific), CHIR99021 (#SML1046; Sigma-Aldrich), PGE2 (P0409; Sigma-Aldrich), recombinant human FGF-10 (#100-26; PeproTech), hEGF (#E9644; Sigma-Aldrich), recombinant murine Noggin (#250-38; PeproTech), N-Acetyl-L-cysteine (#A9165; Sigma-Aldrich), Nicotinamide (#N0636; Sigma-Aldrich), A83-01 (#SML0788; Sigma-Aldrich), SB202190 (#S7067; Sigma-Aldrich), Y-27632 dihydrochloride (#Y0503; Sigma-Aldrich), Primocin (#ant-pm-1; InvivoGen), B-27 Supplement (50X) (#17504044; Thermo Fisher Scientific), HEPES (#15630080; Thermo Fisher Scientific), and Nutlin-3a (#SML0580; Sigma-Aldrich).
Construction of expression vectors
The pLentiCRISPRv2 vectors, containing sgMLL3-a or sgMLL3-b sequences, were generously provided by Dr. Lu Wang at Northwestern University. The dCas9-KRAB vector was purchased from Addgene (#71236; Addgene) and inserted sgRNAs were designed by CRISPR-ERA version 1.2. The shRNA sequences were designed by Biosettia shRNA design tool, and the siRNA sequences were purchased from IDT and annealed according to the manufacturer’s instructions. shRNA oligos for MLL3-a, MLL3-b, GRHL2-a, and GRHL2-b containing an overhang of Age1/EcoR1 were cloned into the pLKO.1-TRC lentiviral vector (#10878; Addgene). For ectopic expression of IRF1, the coding sequence region of IRF1 was amplified by PCR and inserted into the pcDNA3.1 vector. To constitutively express murine IRF1 in murine oral organoids and MOC cell lines, pLX304_zeo_mmirf1 vector was purchased from Addgene (#160098; Addgene). To generate viral particles for lentiviral vectors, lentiviral vectors and third generation of packaging vectors, including pMD2.G (#12259; Addgene), pMDLg/pRRE (#12251; Addgene), and pRSV-REV (#12253; Addgene), were co-transfected into 293T cells using a 2:1:1:1 ratio with BioT transfection reagent (#B01–01; Bioland Scientific). After 48 h of transfection, the supernatants were harvested and filtered with a 0.45-µm size pore. ESCC and HNSCC cells were infected with the collected virus supernatant in the presence of 10 mg/ml polybrene. Table S4 provides the sequences of siRNAs, shRNAs, and sgRNAs.
Subcutaneous syngeneic mouse models
All animal studies were approved by the ethical regulations of the Institutional Animal Care and Use Committee of the University of Southern California. 5-wk-old C57BL/6 mice were purchased from Charles River Laboratories. MOC1 and MOC22 murine HNSCC cell lines were transduced with lentiviral scramble shRNA, shMLL3, and shGRHL2 to establish stable cell lines. For subcutaneous injection, 3 × 106 cells were combined with 100 μl of plain DMEM medium and 100 μl of BME (#3533005-02; R&D Systems) and injected into both flanks of C57BL/6 mice. Tumor growth was monitored every 2 days until the tumors reached their maximum volume (a diameter of 15 mm in any direction). Mice were then euthanized using a CO2 chamber and spinal dislocation, and the tumor samples were collected. Half of each tumor was stored for IF while the remaining portion was dissociated into single cells for flow cytometry analysis.
Orthotopic, syngeneic mouse models
For orthotopic models, 5 × 105 MOC1 cells with scramble or MLL3 knockdown were combined with 50 μl of plain DMEM medium and 50 μl of BME (#3533005-02; R&D Systems) and injected into the right cheek of 5-wk-old C57BL/6 mice. After allowing 2 wk for tumor growth, anti-PD1 treatment (invivoMAB anti-mouse PD1, #BE0273; BioXcell) was initiated. Anti-PD1 mAb was diluted with PBS to a concentration of 10 mg/kg and administered intraperitoneally three times a week for 4 wk. Tumor growth was monitored every 3 days from the beginning of anti-PD1 mAb injection until reaching the maximum volume (a diameter of 15 mm in any direction). Mice with severe tumor-associated morbidity were euthanized. Murine tumors were collected following the aforementioned procedures. To investigate tumor growth in MLL3-deleted organoid models, 1.5 × 106 cells from mouse oral organoids (control: TP53/CDKN2A double knockout or MLL3 KO: TP53/CDKN2A/MLL3 triple knockout) were mixed with 50 μl of DMEM and 50 μl of BME and injected into the right cheek of 5-wk-old athymic nude mice. 1 wk after injection, tumor formation was monitored daily, and the tumor size was measured weekly until it reached the maximum allowable size.
T cell extraction
Isolation of fresh CD8+ T cells from mice was performed by MojoSort Mouse CD8+ T cell isolation kit (#480035; Biolegend). Initially, spleens from C57BL/6 mice were harvested and dissociated into single cells. To eliminate RBCs, 1× RBC Lysis Buffer (#00433-57; eBioscience) was added to the samples and incubated for 15 min at room temperature. Pellets were collected by centrifugation at 12,000 rpm for 10 min and resuspended in 1 ml of PBS. Next, 10 μl of Biotin-Antibody Cocktail was added to 1 × 108 cells/ml and incubated on ice for 15 min. Subsequently, 10 μl of Streptavidin Nanobeads was introduced to the samples with additional incubation on ice for 15 min. The samples were then placed in the Mojosort magnet (#480020; eBioscience) for 5 min. Subsequently, the liquid in the tube was carefully poured out, and the CD8+ T cells were collected by centrifugation at 12,000 rpm for 5 min.
RNA extraction, cDNA synthesis, and qRT-PCR
Total RNA was extracted using the RNeasy Mini Kit (#70106; QIAGEN) and concentration of RNA was measured by Nanodrop. cDNA synthesis was performed using the LunaScript RT Supermix Kit (#M3010L; New England Biolabs). qRT-PCR was carried out using the Powerup SYBR Green Master Mix (#A25918; Thermo Fisher Scientific). TATA-binding protein was used as the reference gene for normalization. The primers utilized in this study are listed in the Table S5.
ChIP assay
The ChIP assay was performed following the previously described (Jiang et al., 2020) protocol. Briefly, 3 × 107 cells were cultured in 10-cm dishes, washed twice with cold PBS, and fixed in 1 ml of 1% paraformaldehyde at room temperature for 10 min. Quenching was achieved with 125 mM glycine for 5 min. The collected samples were lysed twice with 1 ml lysis/wash buffer (150 mM NaCl, 0.5 M EDTA, pH 7.5, 1 M Tris, pH 7.5, and 0.5% NP-40) containing protease inhibitors. Cell pellets were then resuspended in 1 ml shearing buffer (1% SDS, 10 mM EDTA, pH 8.0, and 50 nM Tris, pH 8.0) and subjected to sonication. Following centrifugation at 12,000 × g for 10 min at 4°C, the supernatants were collected. The supernatants were then diluted five times with the dilution buffer (0.01% SDS, 1% Triton X-100, 1.2 mM EDTA, pH 8.0, and 150 nM NaCl) and incubated with primary antibodies (anti-rabbit-normal IgG as a negative control, anti-IRF1, anti-MLL3, anti-GRHL2, and anti-H3K27ac, all 5 μg) at 4°C overnight on a rotator. Dynabeads Protein G beads (#10004D; Thermo Fisher Scientific) were added, and the samples were incubated at 4°C for an additional 4 h on a rotator. After collection by centrifugation, Dynabeads were washed with a precooled wash buffer eight times and once with cold TE buffer. DNA samples were obtained after reverse cross-linking using 5 M NaCl at 65°C overnight and subsequent purification. The final products were subjected to either DNA library preparation and deep sequencing or qPCR quantification.
Flow cytometry
The populations of T cells in the tumors from syngeneic mouse models were analyzed with flow cytometry. In brief, to prevent nonspecific binding, dissociated tumor cells (1 × 106 cells in 100 μl) were incubated for 10 min at 4°C with 0.1 μg of Fc blocker (#553141; BD Pharmingen). Subsequently, the mixture of 1 test volume of Zombie violet dye (#423113; Biolegend), 0.1 µg of Brilliant Violet 605-conjugated anti-mouse CD45 antibody (#103155; Biolegend), 0.1 µg of Percp/cyanine 5.5-conjugated anti-mouse CD3 antibody (#100217; Biolegend), 0.1 µg of FITC-conjugated anti-mouse CD4 antibody (#100405; Biolegend), 0.1 µg of PE-conjugated anti-mouse CD8a antibody (#100707; Biolegend), and/or 0.1 µg of Alexa Fluor 700-conjugated anti-mouse IFNγ antibody (#505823; Biolegend) were added to the sample and incubated for 30 min at 4°C on a rotator. All flow cytometry analyses were performed on an Invitrogen flow machine (AttuneNXT). Data were processed by FlowJo software (Ver10.10; BD bioscience). Statistical analysis was performed to evaluate significant differences between the groups.
Luciferase reporter assay
Candidate enhancer and promoter elements were cloned via PCR amplification and subsequently inserted into pGL3-based luciferase reporter vectors (#E1751 and #E1761; Promega). The constructs were validated through Sanger sequencing to ensure the insertion of correct sequences. These vectors were then transfected into cell lines with transfection reagent BioT (#B01-01; Bioland Scientific). To normalize the transfection and cell number, a Renilla luciferase vector was co-transfected as an internal control. After 48 h of posttransfection, the cells were lysed by the Dual Luciferase Reporter Assay System (#E1960; Promega), and the luciferase activity levels were measured by the 20/20 Luminoritor (Promega).
Western blot assay
Cells were lysed with radioimmunoprecipitation assay buffer mixture containing cocktails of protease inhibitors (#SC24948A; Santa Cruz biotechnology) and mixed with Laemmli Sample Buffer (#161-0737; BioRad). Protein concentrations were measured by Bradford Reagent (#E530-1L; VMR) according to the manufacturer’s instructions. 20 μg of proteins were loaded on SDS-PAGE gels (#M00663 and #M00669; GenScript), which ran for 1 h at 120 V. Loaded proteins were then transferred to 0.45-μm polyvinylidene fluoride (PVDF) membranes (#IPVH00010; Millipore Sigma) with transfer buffer (formula: mixture of 25 mM Tris, 192 mM glycine, and 20% methanol) for 2 h on 270 mA. Membranes were blocked by 2% BSA buffer diluted in 1× PBST (1× PBS with 1% Tween 20) for 1 h at room temperature. The primary antibodies (anti-rabbit MLL3, 1:500; anti-rabbit GRHL2, 1:1,000; anti-rabbit pSTAT1, 1:1,000; anti-rabbit IRF1, 1:1,000; and anti-mouse Actin, 1:5,000) diluted in 1% BSA buffer were added to the membrane and incubated overnight at 4°C. Membranes were then washed three times for 5 min with 1× PBST. The secondary antibody, which is conjugated with HRP, was added to the membrane and incubated for 1 h at room temperature. Membranes were developed with ECL substrate (#A38554; Thermo Fisher Scientific) and detected by chemiluminescence image reader (#LAS-4000; Fujifilm). Broad multicolor pre-stained protein standard (#M00624-250; Genescript) was used for the protein marker.
Colony formation assay
5 × 102 cells of KYSE510, KYSE180, A253, and FADU cell lines were seeded into 6-well plates with 2 ml of corresponding medium, which were incubated for 1–2 wk at 37°C with 5% CO2. Colonies were fixed with 4% paraformaldehyde (#15711; Electron Microscopy Science) and subsequently stained with 1% crystal violet (#V5265; Sigma-Aldrich). After dissolving the colonies using 10% SDS treatment for 30 min, the absorbance rate on the 595-nm wavelength was detected by a microplate reader.
T cell transwell migration assay
T cell migration assays utilized 6.5-mm inserts with 8.0-µm polycarbonate membranes in a 24-well transwell system (#3422; COSTAR). A total of 2 × 105 CD8+ T cells were seeded in the top wells, while the bottom wells were prepared according to different experimental conditions. To investigate the role of CXCR3, anti-CXCR3 pretreated CD8+ T cells were seeded in top wells after plating 1 × 105 MOC1 or MOC22 cells treated with IFNγ in bottom wells for 24 h. After incubation, migrated CD8+ T cells in the bottom well were counted using a hemocytometer after removing the top well. Data from three independent chambers are presented as mean values.
ELISA assay
Using human cell lines, 2 × 105 cells were seeded in 6-well plates. 10 nM of siRNA targeting either MLL3 or GRHL2 were transfected to the cells. After 48 h incubation, 10 ng/ml IFNγ (#ab9659; Abcam) were treated to each well overnight. Murine HNSCC cell lines with scramble, MLL3 knockdown, GRHL2 knockdown, or IRF1 overexpression were seeded in equal numbers in 6-well plates and treated with or without 10 ng/mL IFNγ (#ab9922; Abcam). Supernatant was harvested for the analysis. ELISA kits for human CXCL9 (#DY392-05; R&D Systems), human CXCL10 (#DY266-05; R&D Systems), mouse CXCL9 (#DY492-05; R&D Systems), and mouse CXCL10 (#DY466-05; R&D Systems) were utilized according to the manufacturer’s protocols.
Oral organoid models
The establishment of organoids from the human and murine oral cavity was performed as previously described (Zhao et al., 2022a; Driehuis et al., 2019; Jiang et al., 2024a). Briefly, fresh oral tissue samples were collected, followed by 5× washing with conditioned PBS. Samples were then minced into fragments of <1 mm3 using micro-dissecting scissors. Tissue fragments were digested in DMEM containing 2.5% FBS, 1% penicillin/streptomycin, collagenase type IX (1 mg/ml), and dispase type II (120 μg/ml) at 37°C with 200 rpm shaking for 40–90 min. After centrifugation at 400 × g at 4°C for 3 min, the pellet was resuspended in 50 μl of BME. 2 × 103 cells were seeded on 24-well plates. After solidifying the BME, 500 μl of growth medium was added to each well. For passaging, organoids were washed with 1X PBS and digested with TrypLE (#12604013; Thermo Fisher Scientific) for 5 min at 37°C. After incubation, DMEM/F12 (#10-090-CV; Corning) was added to stop digestion. Organoids were mechanically dissociated by pipetting and centrifuged at 500 × g for 3 min. After resuspending the pellet in BME, 50–100 μl of organoids mixture was seeded on a new culture plate.
CRISPR-Cas9 genomic editing of organoids
Organoids were electroporated as previously described (Zhao et al., 2022a). Briefly, 2 days before the electroporation, organoids were dissociated by TrypLE (#12604013; Thermo Fisher Scientific) and maintained in the organoid culture medium without antibiotics. Organoids were dissociated into clusters of 10–15 cells, resuspended in 80 μl of Opti-MEM Buffer (#11058021; Thermo Fisher Scientific) containing 4 μM electroporation enhancer, and then mixed with 25 μl of RNP complex targeting TP53 and 25 μl of RNP complex targeting CDKN2A. The mixture was transferred into a precooled 2-mm electroporation cuvette (#12358-345; Bulldog Bio). Electroporation was performed by the NEPA electroporation instrument. After electroporation, 200 μl of prewarmed culture medium was immediately added to the electroporation cuvette. Cells were seeded after 40 min incubation at 37°C. After 2 days, transfection efficiency was measured by fluorescence microscopy. 10 μM Nutlin-3a was added to the organoids for the selection of TP53 mutant cells for 2–3 wk. The negative control RNP complex was electroporated to the organoids for the control group. To knockout MLL3, we introduced pLentiCRISPRv2 vectors containing sgMLL3 to these organoids and selected transduced cells with puromycin. To validate targeted mutations, genomic DNA was extracted from edited organoids, followed by PCR amplification and Sanger sequencing.
Organoid viability assay
To quantify proliferation of organoids, 2 × 104 organoids mixed with 100 μl of BME and organoid culture medium were seeded on 24-well plates and incubated at 37°C for a week. At the indicated time points, 100 μl of WST-1 (4-[3-(4-iodophenyl)-2-(4-nitro-phenyl)-2H-5-tetrazolio]-1,3-benzene sulfonate) mixed medium (1:10 ratios of WST1 and medium) was added to each well and incubated for 40 min at 37°C. The medium was collected and allotted to 96 wells, which was measured under the absorbance of 450-nm wavelength by a microplate reader. All experiments were performed in triplicate.
Measurement of organoid size
To measure the size of the organoids at specific time intervals, images of three different sites on the plate were collected using an Olympus microscope and software (Epview). 200× magnification was used for the representative images. The size of the organoids was determined using ImageJ software. A minimum of 90 organoids were measured for each sample.
IF staining
Both tissues and organoids were fixed with 4% buffered formalin solution and then dehydrated with 10% and 30% sucrose-PBS solution. Then, sample were mounted in optimal cutting temperature embedding compound and subsequently frozen at −80°C. Samples were sectioned at a thickness of 10 μm using a cryostat. The cryostat sections were thawed at room temperature for 30 min. The slides were rehydrated in PBS solution. The sections were incubated with primary antibodies, including rabbit polyclonal anti-KI67 antibody (1:500, #ab15580; Abcam) or rat monoclonal anti-CD8a antibody (1:500, #ab22378; Abcam). The goat anti-rabbit IgG (H+L) secondary antibody conjugated with Alexa Fluor 568 conjugated (1:2,000, #ab175471; Abcam) or goat anti-rat IgG (H+L) secondary antibody conjugated with Alexa Fluor 647 conjugated (1:2,000, #A-21245; Abcam) were applied. Finally, fluoroshield with DAPI (#F6057; Millipore Sigma) was added to the samples. Images were captured by a Keyence BZ-X710 fluorescence microscope.
ChIP-seq data analysis
Raw sequencing reads were first mapped to the hg19 (UCSC) genome using the BWA-MEM program with default options. Mapped reads were then sorted using the SAMtools program (version 1.3.1) (Li et al., 2009), followed by the removal of PCR duplicates and blacklist regions by Picard MarkDuplicates tool and bedtools (version 2.27.1). MACS2 (version 2.1.2) (Zhang et al., 2008) was applied to identify peaks with the default setting for transcription factors. Bigwig files were generated by deepTools (Ramírez et al., 2016) bamCompare function (version 3.1.3) with “--operation subtract --normalizeUsing CPM --extendReads 146 --binSize 20.” Transcription factor recognition motif sequence enrichment was performed by Homer findMotifs.pl function, and the enriched motifs with false discovery rate < 0.05 were identified.
RNA-seq data analysis
Paired-end reads were aligned to the hg19 (UCSC) genome using Hisat2 (version 7.2.0) (Kim et al., 2019). Reads were counted with htseq-count program (version 0.11.3) (Putri et al., 2022) and normalized to gene levels by fragments per kilobase per million mapped fragments. Differentially expressed genes were identified by the DESeq2 package with adjusted P value <0.05 and absolute log2 (fold change) >0.5. Pathway enrichment analysis was performed via KOBAS 3.0 (Bu et al., 2021) using the Reactome Pathway Database (Jassal et al., 2020), and the top 15 most significantly enriched pathways (adjusted P value <0.05) were shown in the bubble plot.
Prediction of IRF1-binding sites on CXCL9 and CXCL10
To search for potential IRF1-binding sites on CXCL9 and CXCL10 genes, we utilized the transcription factor–binding prediction method JASPAR (Castro-Mondragon et al., 2022). ChIP qRT-PCR primers were designed based on the predicted target sequences (Table S5).
Statistics
For comparisons of continuous variables between groups, either two-tailed Student t test or one-way ANOVA test was used. The values at P < 0.05 (*), P < 0.01 (**), and P < 0.001 (***) were considered statistically significant. Diagrams were created by GraphPad Prism software, and data were shown as the mean ± SD.
Illustration
Schematic illustrations (Fig. 2 A; Fig. 5, A and G; Fig. 6 A; and Fig. 8 A) in this article were created with https://BioRender.com.
Online supplemental material
Fig. S1 provides the additional information regarding Figs. 1 and 2 about survival probability and validation of knockout system in both human and mouse organoid models. Fig. S2 shows the additional results of both bulk RNA-seq and ChIP-seq analysis. Fig. S3 shares the survival plot of xenograft models using MOC22 cell lines and describes the gating strategy of flow cytometry analysis. Fig. S4 supports the direct regulation of IRF1 and IFNγ signaling by MLL3/GRHL2. Fig. S5 contains the additional results related to Fig. 7. Table S1 shows Sanger sequencing of individual alleles of TP53 following CRISPR/Cas9 editing. Table S2 shows Sanger sequencing of individual alleles of CDKN2A following CRISPR/Cas9 editing. Table S3 shows the list of overlapped genes that were downregulated in both MLL3 knockdown and GRHL2 knockdown. Table S4 shows the list of primer sequences of shRNA, siRNA, and sgRNA. Table S5 shows the list of primer sequences for qRT-PCR and ChIP-qPCR.
Data availability
Intratumoral multi-region whole exome sequencing data were collected from our previous work (Hao et al., 2016) and others (Yan et al., 2019; Chen et al., 2017). ChIP-seq data of H3K4me1 and H3K27ac in ESCC cell lines were from our previous studies (Jiang et al., 2018, 2020). H3K27ac HiChIP data in KYSE70, KYSE140, TT, and RERFCA1 cell lines were collected from Liu et al. (2021). The ChIP-seq and RNA-seq datasets generated in this study have been deposited in the Gene Expression Omnibus repository under the accession code GSE238119.
Acknowledgments
This work was supported by the National Institutes of Health under the award R37CA237022, R01DE033648, and 1R01DK135562 (D.-C. Lin), the National Cancer Institute under grant P30CA014089 (D.-C. Lin), the Ming Hsieh Institute for Research of Engineering-Medicine for Cancer (D.-C. Lin), the Ostrow School of Dentistry of USC Faculty Seed Grant (D.-C. Lin), the Concern Foundation for Cancer Research (D.-C. Lin and U.K. Sinha), the Wright Foundation Transformative Cancer Grant Program (U.K. Sinha and D.-C. Lin), the Watt Family Endowed Chair for Head and Neck Cancer Research (U.K. Sinha), and the Guangdong Natural Science Foundation (2024A1515013059, to G. Huang).
Author contributions: C. Nam: data curation, formal analysis, investigation, methodology, validation, and writing—original draft, review, and editing. G. Huang: investigation, methodology, resources, and visualization. Y. Zheng: formal analysis, investigation, methodology, and visualization. H. Zhao: investigation and methodology. Y. Pan: conceptualization, data curation, and investigation. B. Hu: investigation. T. Wenger: investigation, validation, and writing—review and editing. H.T. Van: resources. L.-Y. Xu: resources and supervision. E.-M. Li: resources and supervision. H.P. Koeffler: project administration, resources, and supervision. K. Ge: resources. Y. Dou: conceptualization. U.K. Sinha: conceptualization, project administration, supervision, and writing—review and editing. Y.M. Park: methodology, software, visualization, and writing—review and editing. D.-C. Lin: conceptualization, funding acquisition, project administration, supervision, and writing—original draft, review, and editing.
References
Author notes
C. Nam, G. Huang, and Y. Zheng contributed equally to this paper.
Disclosures: The authors declare no competing interests exist.