The dynamic addition and removal of long-chain fatty acids modulate protein function and localization. The α/β hydrolase domain–containing (ABHD) 17 enzymes remove acyl chains from membrane-localized proteins such as the oncoprotein NRas, but how the ABHD17 proteins are regulated is unknown. Here, we used cell-based studies and molecular dynamics simulations to show that ABHD17 activity is controlled by two mobile elements—an S-acylated N-terminal helix and a loop—that flank the putative substrate-binding pocket. Multiple S-acylation events anchor the N-terminal helix in the membrane, enabling hydrophobic residues in the loop to engage with the bilayer. This stabilizes the conformation of both helix and loop, alters the conformation of the binding pocket, and optimally positions the enzyme for substrate engagement. S-acylation may be a general feature of acyl-protein thioesterases. By providing a mechanistic understanding of how the lipid modification of a lipid-removing enzyme promotes its enzymatic activity, this work contributes to our understanding of cellular S-acylation cycles.
Introduction
S-acylation, which involves the posttranslational addition of a long-chain fatty acid to cysteine residues, is the only reversible lipid modification. Dynamic S-acylation has been shown to influence protein localization, stability, and activity and is important for cellular processes such as signaling and growth (reviewed in Chen et al. [2021] and Mesquita et al. [2024]). For example, NRas undergoes a cycle of acyl chain addition and removal that regulates its localization at the plasma membrane and is essential for oncogenic signaling (Remsberg et al., 2021; Xu et al., 2012). Altered acylation dynamics are implicated in conditions ranging from cancer and inflammation to neurodegenerative and cardiovascular diseases (Lin, 2021; Main and Fuller, 2022; Ramzan et al., 2023; Zhou et al., 2023).
Two classes of enzymes regulate S-acylation cycles: the protein-acyl transferase enzymes that attach acyl chains to cytosolically exposed cysteines via a thioester bond and the acyl-protein thioesterase (APT) enzymes that remove the lipid (Greaves and Chamberlain, 2010; Won et al., 2018). While an estimated 10–20% of all human proteins are S-acylated, predominantly with the 16-carbon lipid palmitate (Muszbek et al., 1999), only a small subset of these undergoes rapid deacylation (Martin et al., 2011). Cellular APT activity can also be enhanced by growth factor signaling (Kathayat et al., 2017), suggesting APT enzymes are both substrate-selective and stimulus-responsive. However, the mechanisms that regulate their substrate binding and activity are not well understood.
The best-studied cytosolic APTs are members of the metabolic serine hydrolase family, including APT1, APT2, and the α/β hydrolase domain-containing 17 proteins ABHD17A-C (collectively referred to here as ABHD17). ABHD17 is best characterized for its activity on NRas and synaptic PSD-95 (Lin and Conibear, 2015; Yokoi et al., 2016), though the list of targets is growing (Dixon et al., 2023, Preprint; Remsberg et al., 2021; Ulengin-Talkish et al., 2021). Activity of ABHD17 on the stress-regulated exon variant of the large conductance voltage- and calcium-activated potassium channel, nucleotide-binding domain, leucine-rich repeat, and pyrin domain-containing protein 3, and microtubule-associated protein 6 implicates dynamic acylation in physiological processes such as cellular excitability, inflammatory disease, and neuronal maturation and development (Tortosa et al., 2017; McClafferty et al., 2020; Zheng et al., 2023). Notably, a number of other enzymes have been proposed to have thioesterase activity and are under active investigation (Cao et al., 2019; Martin et al., 2011; Rosier et al., 2021; Savinainen et al., 2014; Xu et al., 2018; Yokoi et al., 2016).
From a structural and mechanistic point of view, an interesting feature of many APTs is that they themselves are S-acylated, which directs them to membranes where their substrates are located (Kong et al., 2013; Cao et al., 2019; Martin and Cravatt, 2009). Recent work suggests that acylation of APT2 has additional regulatory functions and occurs through a multistep process involving a membrane-targeting basic patch and hydrophobic β-tongue (Abrami et al., 2021). These structures are required for subsequent acylation of an N-terminal cysteine that confers stable membrane binding and deforms the bilayer, facilitating substrate extraction.
It is not known if these are general principles that apply to other APTs, including the ABHD17 thioesterases. The S-acylated ABHD17 N-terminus is important for localization and activity (Lin and Conibear, 2015; Yokoi et al., 2016), but it remains unclear whether other regions of the protein are important for membrane targeting and substrate extraction. Here, we show that ABHD17A undergoes an activation process involving two distinct structural features. Recruitment of an N-terminal helix followed by acylation is essential for the insertion of a flexible and conserved loop into the bilayer. Membrane binding of the N-terminal helix and loop orients ABHD17A for optimal substrate engagement and activity.
Results
The ABHD17A N-terminus is sufficient for S-acylation and plasma membrane localization
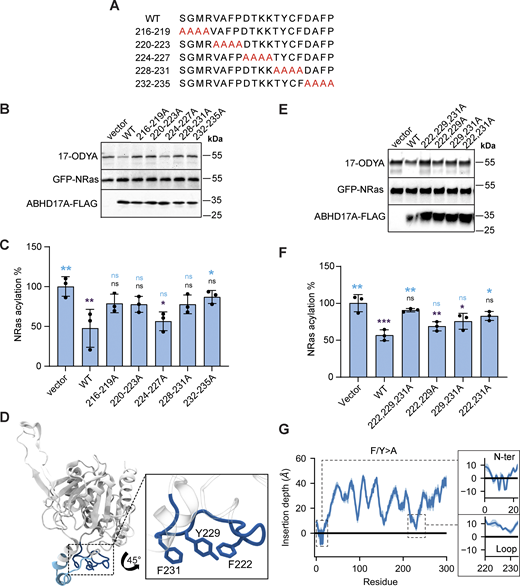
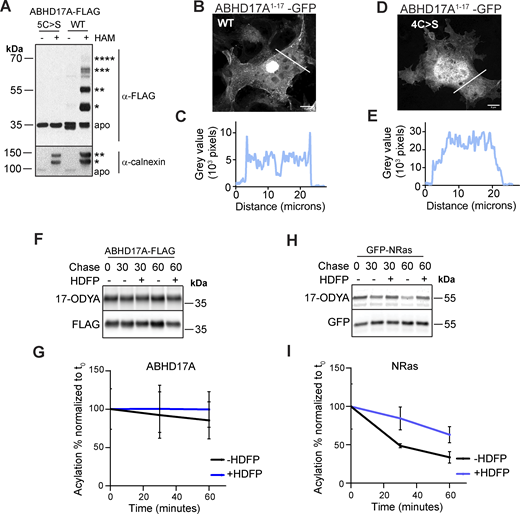
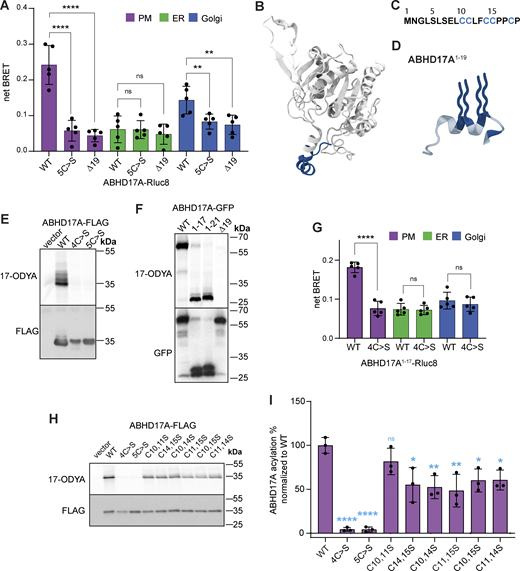
Prior studies showed that deletion of the ABHD17 N-terminus or mutation of all five cysteines in this region blocks S-acylation and plasma membrane targeting (Martin and Cravatt, 2009; Lin and Conibear, 2015; Yokoi et al., 2016). However, the number of modified cysteines remains unknown. To address this question, we focused on ABHD17A, which shows robust activity on NRas (Lin and Conibear, 2015). Using the acyl-PEG exchange (APE) assay, which adds a 5 kD maleimide-functionalized polyethylene glycol (mPEG-maleimide) mass tag to each S-acylation site (Percher et al., 2016), we detected four distinct high molecular weight bands of WT ABHD17A that were absent when the five N-terminal cysteines were mutated to serine (Fig. S1 A). While the fraction of ABHD17A labeled with four mass tags was relatively minor, similar incomplete labeling was observed for endogenous calnexin, a protein known to be dually S-acylated (Percher et al., 2016). This suggests either substoichiometric acylation at steady state, or that the APE reaction was incomplete. Together, these results indicate that at least four, and possibly all five, cysteines in the ABHD17A N-terminus are S-acylated.
It is unclear if other regions of ABHD17 mediate the initial membrane binding that is required for acylation, as reported for APT2 (Abrami et al., 2021). To identify the minimal acylation determinant, we used a sensitive bioluminescence resonance energy transfer (BRET) assay to quantitate its localization at different organelles (Lan et al., 2012). ABHD17A was tagged with the luciferase Rluc8 (donor) and expressed at low levels with different Venus-tagged organelle markers (acceptors). Proximity between the donor and acceptor constructs at organelle membranes results in a high net BRET ratio that reflects the subcellular localization of ABHD17A mutants. We found that replacing all five N-terminal cysteine residues with serine, or deleting the first 19 residues of ABHD17A, significantly decreased plasma membrane localization (Fig. 1 A), consistent with the results of previous immunofluorescence studies (Martin and Cravatt, 2009; Lin and Conibear, 2015). Both mutations also reduced the localization of ABHD17A at the Golgi (Fig. 1 A).
AlphaFold2 (AF2) modeling (Jumper et al., 2021; Varadi et al., 2022) indicates that the N-terminal region of ABHD17A contains an α-helix, with four of the five cysteine residues predicted to lie on or near one face of the helix (Fig. 1, B–D). By labeling cells with the “click-able” palmitate analog 17-octadecynoic acid (17-ODYA), we found that mutation of these four cysteine residues was sufficient to abolish ABHD17A acylation (Fig. 1 E). Moreover, a truncated form of ABHD17A consisting of only its first 17 residues, which includes the 4 cysteines, was acylated as efficiently as a longer fragment that contains all 5 cysteine residues (Fig. 1 F). This minimal N-terminal domain (ABHD17A1–17 GFP) was detected at the plasma membrane both by confocal microscopy (Fig. S1, B–E) and by the BRET-based localization assay (Fig. 1 G), and its targeting was abolished by mutating all four cysteine residues to serine. Together, this demonstrates that the region containing the N-terminal helix is necessary and sufficient for ABHD17A acylation and plasma membrane localization and that—in contrast to APT2—regions outside of the acylation motif are not required.
The acylated N-terminus has functions beyond membrane targeting
Reversible S-acylation of APT2 plays an important regulatory role: the acylated, membrane-bound enzyme is stable, but removal of the lipid modification results in ubiquitination and degradation (Abrami et al., 2021). We examined the acylation dynamics of ABHD17A by pulse-chase labeling cells with 17-ODYA in the presence or absence of the deacylase inhibitor hexadecylfluorophosphonate (HDFP; Martin et al., 2011) (Fig. S1, F and G). In contrast to the dynamically acylated protein NRas (Lin and Conibear, 2015) (Fig. S1, H and I), ABHD17A acylation showed little turnover, suggesting a distinct regulatory role.
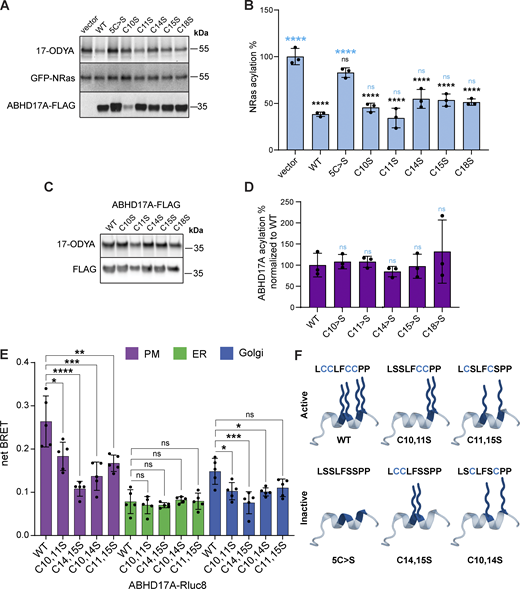
Our results suggest the minimal N-terminal domain has four acylation sites, yet two membrane anchors are considered sufficient to confer stable membrane association to proteins (Shahinian and Silvius, 1995; Conibear and Davis, 2010). When mutated individually, no single cysteine residue was required for enzyme activity (Fig. S2, A and B). Click labeling did not detect differences in acylation levels of single cysteine mutants (Fig. S2, C and D), whereas significant reductions were observed in most, but not all, double cysteine mutants (Fig. 1, H and I). Some discrepancies could result from incomplete acylation of the overexpressed protein, coupled with differences in the site-specific efficiency of S-acylation or de-acylation events.
Unexpectedly, specific pairwise cysteine substitutions revealed striking differences in enzyme activity despite similar reductions in acylation levels (Fig. 2, A and B). The C14,15S and C10,14S mutations largely abolished deacylation activity on NRas, while C11,15S mutations had minimal impact despite the comparable decrease in acylation. Plasma membrane targeting of these mutants, though reduced, was still observed (Fig. S2 E). Based on these results, we speculated that the dramatic effects on activity could be due to seemingly subtle differences in the position of acyl-modified sites (Fig. S2 F) and that the acylated N-terminus might serve functions beyond membrane targeting.
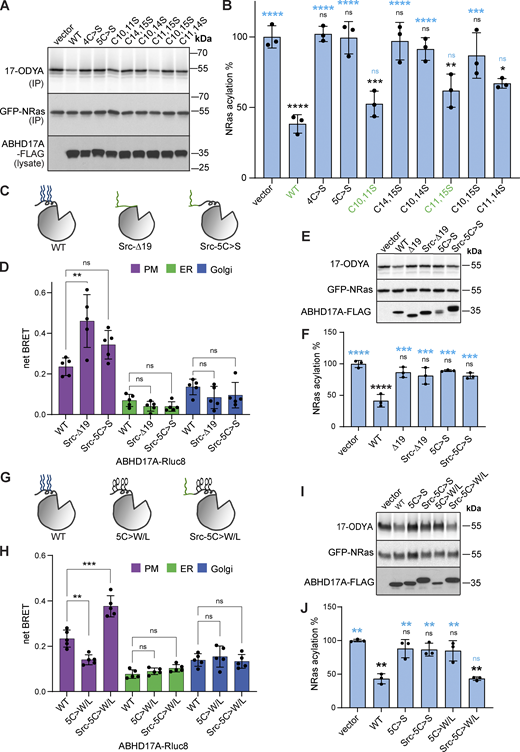
Replacing the acylated N-terminus of ABHD17B with a plasma membrane anchor based on the myristoylated N-terminus of Src did not restore activity on the known substrate PSD-95, highlighting the functional importance of the ABHD17 N-terminus (Yokoi et al., 2016). We hypothesized that this heterologous plasma membrane targeting strategy might circumvent the need for acylation if the N-terminal sequences of ABHD17 were retained. Appending the Src myristoylation motif to either the full-length, acyl-deficient version of ABHD17A (Src-5C>S-ABHD17) or to the N-terminal deletion mutant (Src-∆19-ABHD17) resulted in efficient plasma membrane targeting in a BRET assay (Fig. 2, C and D). However, neither construct showed significant deacylation activity on NRas (Fig. 2, E and F).
We hypothesized that acyl chains on one face of the helix might help embed it in the bilayer, thereby orienting ABHD17A optimally at the membrane. To test this, we replaced the cysteines with bulky hydrophobic residues (Trp/Leu) and added the Src motif (Src-5C>W/L-ABHD17; Fig. 2 G). This mutant not only localized to the plasma membrane (Fig. 2 H) but also showed significantly restored activity on NRas (Fig. 2, I and J), suggesting that proper membrane embedding of the N-terminal helix is essential for ABHD17A function.
ABHD17A is predicted to interact with membranes through two distinct domains
Our results indicate that the thioesterase activity of ABHD17A depends on a specific interaction between its N-terminal helix (“N-helix”) and the lipid bilayer. To understand how ABHD17A interacts with membranes, we turned to molecular dynamics (MD) simulations. First, to assess the quality of the AF2 model of ABHD17A, we analyzed its predicted local distance difference test (pLDDT) score. pLDDT for ABHD17A is very high (>90) for the entire protein, with only two notable exceptions: residues 1–25 (N-terminus) and 42–63 (unstructured loop), which are likely to be flexible in solution (Fig. S3 A). Since the N-terminus is functionally important (Figs. 1 and 2), we next performed all-atom (AA) MD simulations of ABHD17A in solution to investigate its structural and dynamic behavior. Root mean square fluctuation (RMSF) analysis highlights that the N-terminal domain exhibits a significant degree of flexibility along with unstructured linkers (Fig. S3 B). Despite this, the region does not undergo any major conformational changes throughout the simulation (Fig. S3, C and D). Furthermore, the N-terminal domain’s sequence includes multiple proline residues (Fig. 1 C), which are well-known for inducing kinks in α-helices. This characteristic explains the presence of an observed helix–turn–helix–coil structure, which remains stable during the entire AA-MD simulation (Fig. S3 C). This structural stability is confirmed by secondary structure timeline analysis and Ramachandran plots (Fig. S3, D and E), which illustrate that the overall fold of the region is maintained over time.
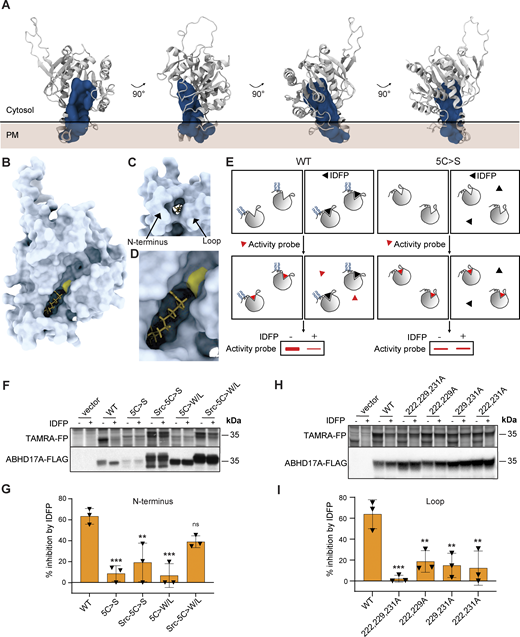
Next, coarse-grained (CG) simulations using the MARTINI force field (Souza et al., 2021) were employed to investigate the interaction between the predicted AF2 model of ABHD17A and a plasma membrane (PM)-like model membrane as this methodology has been shown to accurately predict protein–membrane interfaces for peripheral proteins (Srinivasan et al., 2021; Srinivasan et al., 2024) and has been successfully used to investigate the membrane-binding interface of the thioesterase APT2 (Abrami et al., 2021). We observed that the enzyme primarily interacts with the membrane through two regions: the N-helix and a structurally adjacent loop (residues 222–233, Fig. 3, A and B) that is conserved in all three ABHD17 isoforms and across metazoans (Fig. 3 C). Quantitative analysis of protein–membrane contacts and protein occupancy are shown in Fig. 3, D and E. Of note, analogous simulations started from a structure of ABHD17A with all four cysteines palmitoylated resulted in an identical binding interface with the lipid bilayer (Fig. S4).
To gain deeper insights into ABHD17A’s interaction with the PM-like membrane model, we next adopted a finer resolution by means of MD simulations at the atomistic level. To this end, a representative snapshot from the CG simulations was back-mapped to AA and later palmitoylated at four cysteine sites (Fig. 3 F). AA-MD simulations of this configuration show that the CG binding pose remains almost intact, with both the N-terminus (first 21 residues) and the adjacent loop (residues 222–233) penetrating the membrane to an average depth of ∼11 and 1 Å, respectively, with respect to the average level of the phosphate groups of the monolayer phospholipids (Fig. 3 G).
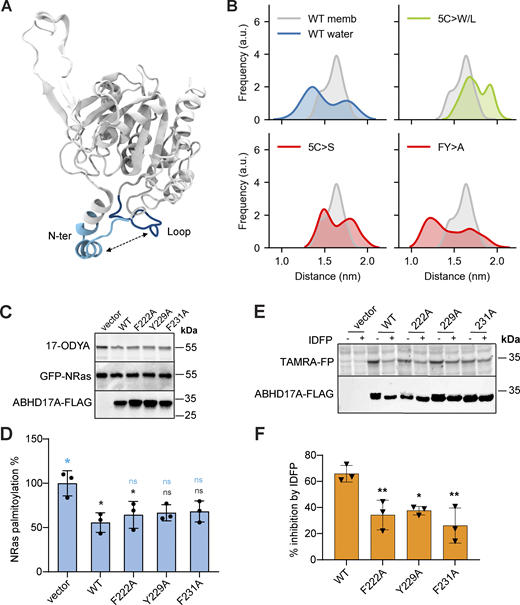
Our simulations predict that both N-helix and loop are important for the interaction of ABDH17A with the membrane. To understand the relative contribution of the two regions, we used AA-MD simulations to model different ABHD17A mutants. Upon mutation of the N-terminal cysteine residues to serine (5C>S), which renders the protein inactive (Fig. 2, A and B), the binding mode of ABHD17 was altered: the insertion depth of the N-terminus decreased significantly (to about 2 Å) and the loop was no longer inserted into the membrane (Fig. 3 H). In agreement with the functional assays (Fig. 2, I and J), mutation of the N-terminal cysteines to bulkier and more hydrophobic residues such as tryptophan (5C>W/L) partially restored the membrane insertion of both the N-helix (to about 5 Å) and loop (to about 0.2 Å; Fig. 3 I). Overall, our data suggest that membrane insertion of the palmitoylated N-helix allows the adjacent loop to insert into the bilayer, and that both interactions contribute to enzyme activity.
Hydrophobic loop residues are important for ABHD17A activity
To test the importance of the conserved loop structure observed in the MD simulations for deacylation activity, we created a series of mutants with four consecutive alanine substitutions spanning the loop (Fig. 4 A). Mutations in the central portion of the loop, which contains polar and charged residues, did not affect enzyme activity on NRas, but mutating outer regions of the loop abolished activity (Fig. 4, B and C). These outer regions contain three conserved hydrophobic residues (F222, Y229, and F231) that lie in a part of the loop that is predicted to insert into the membrane (Fig. 3 B) with protruding side chains that could contribute to membrane binding (Fig. 4 D).
To determine if these hydrophobic loop residues are required for activity, we further made alanine mutations in different combinations (Fig. 4 E). Most of the single and double mutations had relatively minor effects, with F222 and F231 having the greatest effect when mutated together. However, mutating all three hydrophobic residues abolished activity (Fig. 4 F). In AA-MD simulations using the palmitoylated form of ABHD17A, mutation of these hydrophobic loop residues (F/Y>A) resulted in a total loss of membrane insertion for the loop, without affecting the binding of the N-terminus (Fig. 4 G). Taken together, these results suggest that the integrity of both the N-helix and loop are important for membrane binding and activity of ABHD17A.
Mutation of the N-terminal helix and conserved loop alter binding pocket conformation
Many metabolic serine hydrolases have a lid domain that regulates access to the substrate-binding pocket (Devedjiev et al., 2000; Filippova et al., 2013). However, the substrate-binding site of ABHD17, and the mechanisms that regulate substrate binding, are not known. Using a binding pocket prediction algorithm relying on Voronoi tessellation, called FPocket (Le Guilloux et al., 2009), we identified a hydrophobic cavity that represents a potential substrate-binding site (Fig. 5 A). This cavity appears as a cleft in the AF2-predicted structure (Fig. 5 B), with the opening next to the membrane, framed by residues from the N-helix and loop (Fig. 5 C). The catalytic serine is located within the part of the cleft that is distal to the membrane (Fig. 5 D). Based on our top-scoring SwissDock model (Fig. 5, B–D; Grosdidier et al., 2011a), palmitate is predicted to dock in this cleft, with its carboxylate group adjacent to the catalytic serine (Ser190), as would be expected for substrate depalmitoylation. Through AA-MD simulations, we determined that the N-helix and loop regions of ABHD17A are flexible regions that can adopt a wide range of conformations in solution (Fig. S5, A and B), but upon membrane association, they appear to lock into a preferred conformation that could favor substrate insertion into the hydrophobic pocket. Thus, dynamic changes in the positions of the N-helix and/or loop have the potential to control access to the predicted substrate binding site.
Similar to other mSHs, acyl protein thioesterases are irreversibly inhibited by fluorophosphonate (FP) probes, which form a covalent linkage with the active site serine (Creighton, 1993; Piñeiro-Sánchez et al., 1997; Blankman et al., 2007; Martin et al., 2011). To examine the role of the N-helix and loop in regulating substrate binding, we used a competitive activity-based protein profiling (cABPP) assay that measures the ability of a pseudosubstrate FP inhibitor to bind at the active site and prevent subsequent labeling by the fluorescent FP probe TAMRA-FP (Leung et al., 2003; Fig. 5 E). We found that preincubation with the 12-carbon inhibitor isopropyl dodecylfluorophosphonate (IDFP) effectively inhibited TAMRA-FP binding to WT ABHD17A, but not to inactive forms of ABHD17A, such as 5C>S, or the plasma membrane–targeted form of this mutant (Src-5C>S-ABHD17A; Fig. 5, F and G). The altered substrate binding properties of these mutants suggest that changes to either the N-helix or loop that affect their membrane insertion alter the accessibility or conformation of the substrate binding pocket.
We hypothesized that substituting cysteines for bulky hydrophobic residues, which significantly enhanced activity (Fig. 2, I and J), would also restore substrate binding. Indeed, this mutant (Src-5C>W/L-ABHD17A) showed significantly restored IDFP inhibition relative to 5C>S-ABHD17A (Fig. 5, F and G; P ≤ 0.05), suggesting that the insertion of the N-helix in the bilayer is critical for binding pocket accessibility. Moreover, mutation of all three hydrophobic loop residues (F222, Y229, and F231), which blocked ABHD17A activity, strongly decreased IDFP binding (Fig. 5, H and I). This was not observed when hydrophobic loop residues were mutated individually, supporting the idea that their contribution is partially redundant (Fig. S5, C–F).
Taken together, these results show that mutations that reduce the membrane insertion of either the N-helix or loop alter the conformation of the substrate binding pocket and impair deacylation activity. Conversely, compensatory changes that restore membrane insertion restore pocket conformation and enzyme activity. This supports the model that ABHD17A uses both the N-terminal helix and a conserved loop to bind membranes and orient the hydrophobic pocket in a way that promotes substrate binding.
Discussion
We have shown that the activity of ABHD17A relies on the membrane embedding of two regions that flank the putative substrate binding pocket: the acylated N-terminal helix and a flexible loop. Both elements must be inserted into the membrane to position this binding pocket at the bilayer interface in a way that facilitates substrate entry. While our analysis focused on ABHD17A, all three ABHD17 proteins are predicted to have a similar structure that includes an acylated and flexible N-terminus and a conserved loop region, suggesting that they are likely to share the same regulatory mechanism.
Acylation and membrane binding of the N-terminal helix
To be acylated, APTs must first encounter membrane-localized palmitoyltransferases (DHHC-PATs). In the case of APT2, two different regions—a basic patch and a hydrophobic “tongue”—make this initial membrane contact, which leads to the acylation of a single site near the N-terminus (Abrami et al., 2021). In contrast, our work suggests that acylation of the ABHD17 N-terminal domain, which contains a cluster of cysteines on or near a predicted α-helix, does not require other domains. The amphipathic nature of this helix, together with the sulfhydryl side chains of cysteine residues, could favor insertion into the lipid core of the bilayer (Iyer and Mahalakshmi, 2019) and promote the initial membrane interaction needed for ABHD17 to encounter DHHC-PATs.
Because acylation is critical for membrane localization, the dynamic addition and removal of the acyl group was proposed to regulate the activity of APTs by altering access to membrane-localized substrates (Vartak et al., 2014), limiting their concentration by autodeacylation in trans (Kong et al., 2013) or exposing lysine residues that target them for ubiquitination and degradation (Abrami et al., 2021). However, the acylation of ABHD17 on a cluster of four or five cysteines is predicted to shield it from acyl protein thioesterases (Anwar and van der Goot, 2023), which would have to cleave multiple acyl groups to effect membrane release. We did not observe rapid turnover of ABHD17 acylation, thus ABHD17 is unlikely to be regulated by rapid cycling on and off membranes. Instead, our results suggested that acyl chain positioning on the helix was important for activity. Although the reason for this was unclear, the spacing of acylated cysteines could influence helix orientation, subcellular localization, or membrane domain association (Batrouni et al., 2022; El-Husseini et al., 2001).
While the acylated N-terminal helix of ABHD17 is important for membrane targeting, replacing it with a heterologous localization signal compromised activity (Yokoi et al., 2016), suggesting it has another role. We found that full deacylation activity requires the N-terminal helix to associate with the membrane through hydrophobic interactions. In molecular dynamics simulations, the acylated N-terminal helix inserts deeply into the bilayer, which in turn promotes the membrane insertion of the adjacent loop structure. While the insertion of the N-terminal helix is important for loop insertion, the converse is not true: mutations in the loop that block membrane binding and activity do not disrupt the embedding of the N-helix. This suggests that acylation of the N-helix is an initial step that positions ABHD17 for loop insertion and substrate engagement.
The opening of the predicted binding pocket faces the membrane, whereas the catalytic serine is at the base of the pocket, suggesting the lipid substrate must be extracted from the bilayer to enter the binding site. We hypothesize that the membrane insertion of both the N-helix and the loop, which flank this pocket, are important for orienting the enzyme so that the binding pocket is optimally positioned for lipid extraction to occur. Furthermore, the insertion of the acylated N-helix is expected to perturb the bilayer, facilitating this extraction. Indeed, palmitate, the predominant lipid used for S-acylation, decreases membrane fluidity and destabilizes membranes in liposomes (Leekumjorn et al., 2009). Because both tryptophan and phenylalanine insertion also destabilize membranes (Popova et al., 2002), loop binding could also contribute to substrate extraction. Similarly, membrane deformation by APT2 is proposed to facilitate substrate removal and subsequent hydrolysis (Abrami et al., 2021). Thus, acylation may be a common feature of APTs that not only serves as a membrane-targeting signal but also contributes to deacylation activity.
Binding pocket conformation is regulated at the membrane
It is unclear if access to the ABHD17 pocket is dynamically regulated. Many lipases have highly dynamic lid structures that occlude the binding pocket until interaction with a membrane causes the lid to open (Khan et al., 2017). Lid opening and substrate engagement can also cause conformational changes in the binding pocket that promote binding and position the active site residues for cleavage. The APT1/2-like protein FTT258 found in the bacteria Francisella tularensis has a flexible loop that acts like a lid to regulate membrane binding, binding pocket accessibility, and enzyme activity (Filippova et al., 2013; Smith et al., 2018). APT1 lacks a lid-like structure but does have a loop that forms part of the binding pocket and undergoes subtle structural rearrangements on substrate binding to accommodate the substrate, which provides isoform selectivity (Won et al., 2016; Harris et al., 2024). APT homologs in plants are also suggested to use sequences adjacent to their active site to select the chain length of substrates (Bürger et al., 2017).
Although ABHD17 lacks an obvious lid structure, we found the integrity of both the N-terminus and loop to be important for the conformation of the binding pocket. In MD simulations, the relative position of the N-helix and loop is highly variable in solution but uniform when bound to membranes. Based on these observations, we favor the hypothesis that the binding and insertion of the N-helix and loop elements into the lipid bilayer stabilizes a conformation that promotes substrate engagement. The location of the active site serine in a deep cleft might help accommodate the acylated peptide as it slides into the active site. Whether ABHD17 proteins undergo additional structural rearrangements in response to substrate binding will be the topic of future studies.
Different organelle membranes have distinct lipid compositions and lipid packing properties that can influence protein binding and insertion (Bigay and Antonny, 2012). It is tempting to speculate that the need for correct insertion of the N-terminal helix and loop regions limits ABHD17 activity to specific organelles or membrane subdomains. This could help explain why ABHD17 appears to act primarily on substrates at the plasma membrane (Remsberg et al., 2021). How ABHD17 localization affects activity and substrate specificity, and whether ABHD17 proteins are also regulated by additional posttranslational modification or expression level (Abazari et al., 2023; Wild et al., 2023) are important areas of future research. A complete understanding of ABHD17 regulation will be helpful in developing therapeutic approaches that target ABHD17 and will increase our understanding of related thioesterases.
Materials and methods
Plasmids
Venus-tagged KRas-tail, PTP1b-tail, Giantin, and Rluc8-N1 plasmids were gifts from Dr. Nevin Lambert (Augusta University, Augusta, GA, USA; Lan et al., 2012). ABHD17A mutant plasmids (plasmids 5–21 in Table S1) were made with NEBuilder HiFi DNA Assembly (New England Biolabs) using primers (Table S2) to amplify and mutate ABHD17A sequences that were inserted into EcoRI/Bsu36I-digested pABHD17A-FLAG (Lin and Conibear, 2015). The loop mutant plasmids (plasmids 22–33) were made by ligating two PCR products with SacII/BmgBI digested pABHD17A-FLAG. A similar approach was used to create pSrc-∆19-FLAG, ligating the PCR product into EcoRI/XhoI-digested pABHD17A(∆19)-FLAG.
To create pABHD17A-EGFP, the EGFP PCR product was ligated into EcoRi/SalI digested pABHD17A-FLAG. NEBuilder HiFi DNA Assembly was used to ligate 1–17 and 1–21 ABHD17A fragments with pABHD17A-EGFP that were digested with EcoRI/AgeI to remove ABHD17 sequences. pABHD17A-Rluc8 constructs were created by PCR amplifying ABHD17A mutants described above. Ligation of the PCR product with EcoRI/BamHI digested pRluc8-N1 was performed with NEBuider HiFi DNA Assembly.
Cell culture and cDNA transfection
COS-7 cells, a non-human primate male cell line from ATCC, were maintained and propagated in high-glucose Dulbecco’s Modified Eagle Medium (DMEM; Gibco) supplemented with 10% fetal bovine serum (FBS; Gibco), 4 mM L-glutamine, and 1 mM sodium pyruvate in a humidified incubator at 37°C, 5% CO2. Cells were checked for mycoplasma contamination every 6 mo by PCR-based analysis. COS-7 cells were grown on 60-mm plates and transfected at 90% confluence using Lipofectamine 2000 (Invitrogen) as per manufacturer’s instructions with 1 µg cDNA per plate of each DNA construct for 17-ODYA and cABPP labeling studies. For BRET, cells were grown in 6-well plates and transfected at 90% confluence with 0.5 μg cDNA of acceptor constructs and 0.05 μg cDNA of donor constructs. For fluorescence microscopy, cells were grown on 12-mm glass coverslips (Thermo Fisher Scientific) in 24-well plates and transfected at 90% confluence with 0.4 μg cDNA. Cells were transfected at 90% confluence with 11 μg cDNA for the APE assay.
BRET assays
Cells were washed in PBS, harvested with 0.25% EDTA-Trypsin (Gibco), resuspended in HBSS (Gibco), and transferred to Nunc F96 MicroWell White Polystyrene 96-well plates (Thermo Fisher Scientific). Fluorescence and luminescence measurements were made using a Tecan Spark multimode microplate reader (Tecan). Fluorescence emission was measured at 485 and 530 nm. Coelenterazine h (5 μM; Cayman) was added to cells immediately prior to making measurements. Raw BRET signals were calculated as the emission intensity at 530 nm divided by the emission intensity at 485 nm. Net BRET is the emissionacceptor/emissiondonor ratio minus the emissionacceptor/emissiondonor ratio measured from cells expressing only the BRET donor.
Acyl-PEG exchange assay
A modified version of the APE assay (Percher et al., 2016; Hurst et al., 2017) was used to assess S-acylation levels. 24 h after transfection, COS-7 cells were washed three times in PBS, frozen at −80°C for 24 h, and lysed in triethanolamine (TEA) buffer (50 mM triethanolamine pH 7.3, 150 mM NaCl) containing 4% SDS, 5 mM EDTA, 5 mM PMSF, and 1X Halt Protease Inhibitor Cocktail (Thermo Fisher Scientific). Clarified lysates were incubated with 25 mM N-ethylmaleimide (NEM; Sigma-Aldrich) for 2 h at RT to block free cysteines. 500 μg of protein in 135 ml TEA buffer was transferred to a new tube and samples were treated with 250 mM 2,3-dimethyl-1,3-butadiene (Sigma-Aldrich) for 1 h at RT with periodic vortexing to remove excess NEM. This was followed by the addition of 1/10th vol of chloroform, 1 min of vortexing, and centrifugation at maximum speed for 5 min. The supernatant was split into two aliquots, with one aliquot receiving 750 mM hydroxylamine treatment for 1 h at RT. A methanol–chloroform extraction was performed, after which the protein precipitates were resuspended in TEA buffer containing 1.33 mM mPEG-maleimide (Sigma-Aldrich), 4 mM EDTA, and 0.2% Triton X-100, and incubated for 2 h at RT. Following methanol–chloroform extraction to remove excess mPEG-maleimide, protein pellets were resuspended in Laemmli buffer (62.5 mM Tris-HCl pH 6.8, 2% SDS, 10% glycerol, 1% 2-mercaptoethanol, and 0.002% Bromophenol blue) and heated at 80°C for 5 min. Samples were analyzed by 10% SDS-PAGE followed by western blotting.
17-ODYA metabolic labeling
20 h following transfection, COS-7 cells were washed once in PBS and starved for 1 h in Base Labelling Media (1 mM L-Glutamine, 1 mM cysteine, and 1 mM sodium pyruvate in cysteine- and methionine-free DMEM). A 30 μM solution of saponified 17-ODYA (Cayman) was prepared by first incubating 6 mM ODYA in 7.2 mM sodium hydroxide at 65°C for 15 min, then adding this to Base Labeling Media containing 20% fatty acid-free BSA. After 2 h in the 17-ODYA solution, cells were washed three times in PBS and placed at −80°C for 24 h. Cells were lysed with TEA lysis buffer (50 mM TEA pH 7.4, 1% TritonX-100, 150 mM NaCl, 2×EDTA-free Halt Protease Inhibitor [Life Technologies]) and vigorously mixed by pipetting. Lysates were cleared by centrifugation at 16,000 × g for 15 min at 4°C, and the supernatant protein content was quantified using Bicinchoninic acid (BCA) assay (Life Technologies). 100–130 μg of supernatant was diluted in SDS-sample buffer (200 mM Tris-HCl pH 6.8, 8% SDS, 4% Bromophenol Blue, 40% Glycerol, 1% 2-mercaptoethanol). Samples were heated for 5 min at 95°C and kept at −20°C. The remaining lysate was used for immunoprecipitation and sequential on-bead CuAAC/click chemistry.
For pulse-chase metabolic labeling in the presence of inhibitors, following the 2 h 17-ODYA labeling, cells were washed once in PBS, and chase media (complete DMEM supplemented with 10% FBS and 300 mM palmitic acid) containing 10 μM HDFP (Cayman) in DMSO was added. At the indicated time points, cells were washed three times in PBS and placed at −80°C for 24 h. Cell lysis was carried out as described above.
Immunoprecipitation and sequential on-bead CuAAC/click chemistry
For immunoprecipitations, Protein A Sepharose beads (GE Healthcare) were washed three times in TEA lysis buffer and preincubated with rabbit anti-GFP antibody for 4 h at 4°C before the addition 500 μg–1 mg of transfected COS-7 cell lysates. Immunoprecipitations were carried out for 12–16 h on an end-to-end rotator at 4°C. Sepharose beads were then washed three times in modified radioimmunoprecipitation assay (RIPA) buffer (50 mM TEA pH 7.4, 150 mM NaCl, 1% sodium deoxycholate [wt/vol], 1% TX-100, and 0.1% SDS).
Sequential on-bead click chemistry of immunoprecipitated 17-ODYA–labeled proteins was carried out as previously described (Zhang et al., 2010) with minor modifications. Sepharose beads were washed three times in RIPA buffer and on-bead conjugation of AF647-azide (Invitrogen) to 17-ODYA was carried out for 1 h at RT in 50 μl of freshly mixed click chemistry reaction mixture containing 1 mM TCEP (Aldrich), 1 mM CuSO4·5H2O (Sigma-Aldrich), 100 µM TBTA (Aldrich), and 100 μM AF647-azide in water. Beads were washed three times with RIPA buffer and resuspended in SDS buffer (200 mM Tris-HCl pH 6.8, 150 mM NaCl, 12% SDS, 50 mM TEA pH 7.4, 4% Bromophenol Blue, 40% glycerol, and 1% 2-mercaptoethanol). Samples were heated for 5 min at 95°C and separated on 10% tris-glycine SDS-PAGE gels for subsequent in-gel fluorescence analyses and then transferred onto nitrocellulose membrane for western blotting. The percent of acylated NRas was calculated using the ratio of ODYA to total purified NRas, normalized to the average value of the vector control.
Lysate samples frozen in SDS-sample buffer were thawed, heated at 95°C for 2 min, sonicated gently, and run on 13% tris-glycine SDS-PAGE gels for western blot analyses.
Competitive activity-base protein profiling
24 h following transfection with ABHD17A constructs, COS-7 cells were washed once in PBS, incubated with 0.25% trypsin for 3 min at 37°C, detached with DMEM + FBS (10%), and collected by centrifugation at 1,000 × g for 3 min. The cells were resuspended in 300 μl 50 mM Tris and lysed with gentle sonication on ice. Protein was quantified by BCA assay and 30 μg protein was incubated with IDFP (20 μM; Cayman) in DMSO at RT for 30 min. TAMRA-FP (2 μM final concentration) labeling was carried out at RT for 1 h and quenched with 4× SDS-sample buffer heated to 95°C for 5 min. Samples were separated on SDS–PAGE, analyzed by in-gel fluorescence, and then transferred onto nitrocellulose membrane for western blotting. Inhibition was calculated by dividing the TAMRA-FP signal treated with IDFP by the DMSO control and subtracting this from 100.
In-gel fluorescence analyses
A Typhoon Trio scanner (GE Healthcare) was used to measure the in-gel fluorescence of SDS-PAGE gels. AF647 signals were acquired using the red laser (excitation 633 nm) with a 670BP30 emission filter, and rhodamine signals were acquired with the green laser (excitation 532 nm), with a 580BP30 emission filter. Signals were acquired in the linear range and quantified using Fiji (Schindelin et al., 2012).
Western blotting
Nitrocellulose membranes (50-206-3328; Thermo Fisher Scientific) were blocked in PBST (PBS with 0.1% Tween-20 [Sigma-Aldrich]) containing 3% BSA (Sigma-Aldrich) for 1 h. Membranes were then blotted with corresponding primary antibodies (mouse anti-GFP [11814460001; Roche; RRID:AB_390913] or rabbit anti-FLAG [701629; Thermo Fisher Scientific; RRID: AB_2532497]) in PBST + 3% BSA for 1 h followed by either horseradish peroxidase–conjugated polyclonal goat anti-mouse (115–035-146; Jackson ImmunoResearch Laboratories; RRID: AB_2307392) or horseradish peroxidase–conjugated polyclonal goat anti-rabbit (170-6515; Biorad; RRID: AB_11125142) in PBST + 3% BSA for 1 h. All blots were developed with ECL chemiluminescent reagents (RPN2106; Cytiva) and exposed to Amersham Hyperfilm (CA95-17; VWR). Developed films were scanned, and densitometry was performed in Fiji. The blots in Fig. S1, F and H; and Fig. S2, C and D were imaged on the Fusion FX (Vilber).
Weblogo
ABHD17A homologs were identified at the metazoan level with OrthoDB (Kuznetsov et al., 2023) and aligned with MUSCLE (Edgar, 2004). Weblogo (Crooks et al., 2004) was used to analyze amino acids 216–235 of Homo sapiens ABHD17A.
Fluorescence microscopy image acquisition and processing
Coverslips were mounted onto a microscopy slide using ProLong Gold Anti-Fade Mountant (Thermo Fisher Scientific). Microscopy images were acquired at RT on a Leica TCS SP8 Confocal Microscope (Leica Microsystems) with a high-contrast Plan Apochromat 63×/1.30 Glyc CORR CS objective (Leica Microsystems) and an ORCA-Flash4.0 digital camera (Hamamatsu Photonics). Confocal images were acquired with Leica Application Suite X (LASX) 3.5.7 software (Leica Microsystems), followed by deconvolution using Huygens Professional version 23.04 (Scientific Volume Imaging). Representative confocal images were individually adjusted for brightness and contrast using Fiji (Schindelin et al., 2012).
Statistical analysis
GraphPad Prism 10 was used to perform statistical analysis. One-way ANOVA was used to determine the P values of raw data with Tukey’s multiple comparison test. P values are reported in figure legends. ns = P > 0.05, * = P ≤ 0.05, ** = P ≤ 0.01, *** = P ≤ 0.001, **** = P ≤ 0.0001. Data distribution was assumed to be normal but this was not formally tested.
MD simulations
All MD simulations were carried out starting from the AF2 predicted model of ABHD17A (Uniprot accession number: Q96GS6) and a simple PM-like model membrane constituted of 40%POPC-30%POPS-30%CHOL. CG-MD simulations were carried out using the GROMACS package (version 2021.2) (Abraham et al., 2015) and employing the MARTINI3 forcefield (Souza et al., 2021). The initial system setup, consisting of ABHD17A positioned ∼5 nm above a PM-like membrane, solvated with a 0.15 NaCl solution, was built using the insane.py script (Wassenaar et al., 2015) and subsequently energy-minimized using the steepest descent algorithm. After a 10-ns equilibration conducted in the NPT ensemble, four replicas of 4 µs each were run with a timestep of 20 fs. Using semi-isotropic pressure coupling, the pressure was kept at 1 bar using the Parrinello–Rahman barostat (Parrinello and Rahman, 1981) and applied every 12.0 ps with a compressibility of 3 × 10−4 bar−1. Temperature was maintained constant at 310 K employing the V-rescale thermostat (Bussi et al., 2007). Palmitoylation was modeled using the protocol and parameters from Koukos et al. (2023).
AA-MD simulations were started using a representative snapshot of CG simulations, where ABHD17A was stably anchored to the membrane through both the N-terminus and the structurally adjacent loop. The system was first back-mapped from MARTINI3 CG model to the CHARMM36 AA model (Klauda et al., 2010), employing the CG2AT2 tool (Vickery and Stansfeld, 2021), and later palmitoylated at four sites (CYS10-11-14-15) using CHARMM-GUI tool PDB reader & manipulator (Jo et al., 2008; Jo et al., 2009). Systems containing WT and mutants were equilibrated following the classical CHARMM-GUI 6 step protocol and then simulated using GROMACS software and the CHARMM36m forcefield. Productions were repeated three times for 500 ns each with a timestep of 2 fs in the isothermal–isobaric ensemble (NPT). The Nosé-Hoover thermostat (Evans and Holian, 1985) was used to keep the temperature at 310 K, and the Parrinello–Rahman barostat (Parrinello and Rahman, 1981) was employed, applying a semi-isotropic pressure coupling, to maintain the pressure at 1 bar and a compressibility of 4.5 × 10−5 bar−1 every 5 ps. The Particle Mesh Ewald (Darden et al., 1993; Essmann et al., 1995) algorithm was utilized for long-range electrostatic interactions, with a cutoff of 1.2 nm. Lennard–Jones interactions were truncated at 1.2 nm. Bond constraints were treated using linear constraint solver algorithm (Hess et al., 1997).
To investigate the role of the flexibility of the N-terminus and the loop, AA simulations of WT-ABHD17A in solution were performed. The protein was put in a box of water with 0.15 M of NaCl and minimized using the steepest descent algorithm. Six-step CHARMM-GUI equilibration and 1.5 μs NPT production were performed utilizing the same temperature conditions as described before for AA simulations. The pressure was kept at 1 bar through the use of an isotropic barostat (Parrinello and Rahman, 1981) with a compressibility of 4.5 × 10−5 bar−1 using τP = 5 ps.
Protein–membrane contact analysis was conducted on CG systems using ProLint (Sejdiu and Tieleman, 2021), while the occupancy map was generated using GROMACS module gmx select. Insertion depth for AA systems was evaluated as in Rogers and Geissler (2023). Cavity pocket detection was performed utilizing the Fpocket software (Le Guilloux et al., 2009), giving as a reference structure for the pocket identification of the AF2 model of ABHD17A. Palmitic acid was docked onto the ABHD17A AF2 model using the SwissDock webserver (https://www.swissdock.ch/) using default parameters (Grosdidier et al., 2011a; Grosdidier et al., 2011b).
To investigate the flexibility of N-terminus and loop regions of ABHD17A WT and mutants in solution versus in membrane, the distance between the center of mass of the two regions was measured using the gmx pairdist, and the results were normalized and shown as a probability histogram. All the molecular images were rendered using Visual Molecular Dynamics software (Humphrey et al., 1996). Plots were generated using the python module matplotlib (Hunter, 2007).
Online supplemental material
Fig. S1 (complementary to Fig. 1) shows that ABHD17A has at least four acylated cysteine residues that do not show significant turnover and are important for plasma membrane localization. Fig. S2 (complementary to Fig. 1) examines the requirement for specific ABHD17A N-terminal cysteine residues for activity and plasma membrane localization. Fig. S3 shows the dynamic behavior of the ABHD17A N-terminus in AA-MD simulations (complementary to Fig. 3). Fig. S4 shows the effect of palmitoylation on membrane targeting by ABHD17A in CG simulations (complementary to Fig. 3). Fig. S5 shows the effect of mutants on ABHD17A conformation, activity, and binding pocket conformation (complementary to Fig. 5). Table S1 is a list of the plasmids used in this study. Table S2 is a list of the primers used in this study.
Data availability
The data underlying all biochemical experiments are available in the published article and its online supplemental material. MD simulations, all input files, initial configurations, and trajectories saved every 100 ns after water removal, as well as scripts used for analyses can be found at: https://doi.org/10.5281/zenodo.14702866.
Acknowledgments
We thank Dr. Nevin Lambert (Augusta University, Augusta, GA, USA) for the Venus-tagged organelle markers, Rluc8 plasmids (Lan e al., 2012), and guidance in quantification and statistical analysis of BRET. We also thank Dr. Alexis Shih for helping to develop the IDFP-cABPP assay and thank Dr. Gao Guang and the Life Sciences Institute Imaging Facility at the University of British Columbia for helping with confocal imaging and deconvolution.
This work was supported by funding from the Canada Foundation for Innovation (Leading Edge Fund 30636, to E. Conibear), the Canadian Institutes of Health Research (grant 162184 to E. Conibear), the European Research Council under the European Union’s Horizon 2020 Research and Innovation program (grant agreement no. 803952 to S. Vanni), and by grants from the Swiss National Supercomputing Centre under projects ID s1132, s1176, and s1221.
Author contributions: S. Holme: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Resources, Validation, Writing - original draft, J. Sapia: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Validation, Visualization, Writing - original draft, Writing - review & editing, M. Davey: Investigation, Methodology, Resources, S. Vanni: Conceptualization, Formal analysis, Funding acquisition, Resources, Supervision, Writing - original draft, Writing - review & editing, E. Conibear: Conceptualization, Data curation, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing - original draft, Writing - review & editing.
References
Author notes
S. Holme and J. Sapia contributed equally to this paper.
Disclosures: The authors declare no competing interests exist.


![ABHD17A-membrane interaction is predicted to occur through two distinct domains. (A) Mapping of full-length AA ABHD17A to CG MARTINI3.0 model. (B) Representative mechanism for the binding of ABHD17A (red) to a PM-like membrane (1-palmitoyl-2-oleoylphosphatidylcholine [POPC]: light yellow, 1-palmitoyl-2-oleoylphosphatidylserine [POPS]: orange, cholesterol [CHOL]: yellow) in CG simulations: the protein, initially randomly oriented at a minimum distance of 5 nm from the bilayer, interacts with the membrane via both its N-terminal helix and the adjacent loop. (C) Conservation of the loop residues in the three ABHD17 isoforms in metazoans as shown with Weblogo (Crooks et al., 2004). (D and E) Membrane interaction of the N-terminal helix and conserved loop shown in the contacts analysis (D) and the occupancy maps (E). (F) Snapshot from a representative AA simulation, showing the back-mapped atomistic structure of ABHD17A with the addition of palmitoyl-tails to cysteine residues at its N-terminus. (G) Insertion depth analysis of AA simulations confirms that ABHD17A inserts into the membrane via both the palmitoylated N-terminus (residues 1–21) and the adjacent loop (residues 222–233). (H) Insertion depth analysis of ABHD17A 5C>S with AA simulations shows mutation of the five N-terminal cysteine residues to serine decreases both N-terminal and loop membrane insertion. (I) Restoring N-terminal hydrophobicity of the non-acylated ABHD17A (5C>W/L) rescues insertion depth of the N-terminus and loop in AA simulations. The data shown in G–I are averaged over four replicates, and the error bars indicate STDEV. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jcb/224/4/10.1083_jcb.202405042/1/m_jcb_202405042_fig3.png?Expires=1781191626&Signature=2LNk5FDJ87z33-fNYhWddU86DIL97OsFHXqu1KT-rF9-g3Orp~3meFZuJJuFDanc42KPeNPKS-JQjVI-5wFKOw0D2BgMIizDh8mDSb8-kYDIYmCH6Uae2Jdq3X7LbS0e4PUtfty~3AMYT2-AliI03mfHd5Ri9rPSwA5V3Bk7FPYHXG00Pl4IyM-hzBBhEiu9unzVcg25oNoxOGdW~qB3wfquWZyBk1e0D0NpN2LzmWNF1Xjd5-FULR6qSwq5FbpdcCIAIkymibBNZ31extW3jYqUlqiw1gfwyANjgedYVtuGf9Qi388h4lzx6XO7SqM09MnkouRI0ETrUJV~O8fnmQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)