Calmodulin (CaM) plays a critical role in intracellular signaling and regulation of Ca2+-dependent proteins and ion channels. Mutations in CaM cause life-threatening cardiac arrhythmias. Among the known CaM targets, small-conductance Ca2+-activated K+ (SK) channels are unique, since they are gated solely by beat-to-beat changes in intracellular Ca2+. However, the molecular mechanisms of how CaM mutations may affect the function of SK channels remain incompletely understood. To address the structural and functional effects of these mutations, we introduced prototypical human CaM mutations in human induced pluripotent stem cell–derived cardiomyocyte-like cells (hiPSC-CMs). Using structural modeling and molecular dynamics simulation, we demonstrate that human calmodulinopathy-associated CaM mutations disrupt cardiac SK channel function via distinct mechanisms. CaMD96V and CaMD130G mutants reduce SK currents through a dominant-negative fashion. By contrast, specific mutations replacing phenylalanine with leucine result in conformational changes that affect helix packing in the C-lobe, which disengage the interactions between apo-CaM and the CaM-binding domain of SK channels. Distinct mutant CaMs may result in a significant reduction in the activation of the SK channels, leading to a decrease in the key Ca2+-dependent repolarization currents these channels mediate. The findings in this study may be generalizable to other interactions of mutant CaMs with Ca2+-dependent proteins within cardiac myocytes.
Introduction
Calmodulin (CaM) is a multifunctional Ca2+-binding protein that orchestrates a wide range of intracellular signaling and critical cellular processes (Crivici and Ikura, 1995; Chin and Means, 2000). CaM, a 17-kD protein, is encoded by three distinct genes in humans, namely CALM1, CALM2, and CALM3, each of which encodes an identical CaM protein. CaM is composed of N- and C-terminal lobes linked by a flexible helix (Babu et al., 1985; Heidorn and Trewhella, 1988). Each lobe contains two EF hands, canonical Ca2+-binding motifs, with the N-lobe having slightly lower Ca2+-binding affinity (Watterson et al., 1980; Klevit et al., 1984; Thulin et al., 1984). Ca2+ binding to the EF hands results in structural and functional changes in target molecules (Saimi and Kung, 2002).
Recent studies have provided genetic links between human heritable CaM mutations and several types of cardiac arrhythmia syndromes leading to sudden cardiac death, including long QT syndrome (LQTS; Crotti et al., 2013; Reed et al., 2015; Boczek et al., 2016), catecholaminergic polymorphic ventricular tachycardia (CPVT; Nyegaard et al., 2012), and familial idiopathic ventricular fibrillation (IVF; Hwang et al., 2014). N54I and N98S mutations in CaM are associated with cases of CPVT with altered CaM-ryanodine receptor 2 (RYR2) function; F90L has been shown to be linked to IVF; and D96V, D130G, and F142L are associated with LQTS.
LQTS mutations of CaM have been shown to reside at or near Ca2+ coordinating sites within the EF hands of the C-lobe of CaM (Limpitikul et al., 2014; Jensen et al., 2018). CALM1-D130G and CALM2-D96V mutations alter highly conserved aspartic acid residues that bind Ca2+ ions in EF-hand domains IV and III, respectively. The CALM1-F142L mutation results in an alteration of the energetic coupling of Ca2+ binding and the conformational change associated with CaM activation (Crotti et al., 2013). All three LQTS-associated mutations show reduced Ca2+ affinity in the C-lobe by 5-, 13-, and 53-fold for F142L, D96V, and D130G, respectively (Crotti et al., 2013). These mutations in CaM result in a significant reduction in Ca2+-dependent inactivation (CDI) of Cav1.2 L-type Ca2+ channel (LTCC; Peterson et al., 1999; Zühlke et al., 1999; Zühlke et al., 2000). In contrast, CPVT mutations in CaM impart little-to-mild reduction of Ca2+-binding affinity (Nyegaard et al., 2012).
Clearly, CaM exerts its function on a number of targets in cardiomyocytes, including RYR2 (Meissner, 1986; Fukuda et al., 2014; Hwang et al., 2014; Nomikos et al., 2014; Vassilakopoulou et al., 2015), Kv7.1 (or KCNQ1; Shamgar et al., 2006; Sachyani et al., 2014; Sun and MacKinnon, 2017), LTCCs (Cav1.2, Cav1.3; Liang et al., 2003; Halling et al., 2006; Findeisen and Minor, 2010), and small-conductance Ca2+-activated K+ (SK) channels (Xia et al., 1998; Schumacher et al., 2001). Among the CaM-targets, SK channels are unique because the channels are voltage independent and are gated solely by beat-to-beat changes in intracellular Ca2+ (Ca2+i). SK channels harbor a highly conserved CaM-binding domain (CaMBD); the gating of the channels is controlled by Ca2+ binding and unbinding to CaM associated with SK channels (Xia et al., 1998). Thus, CaM serves as the high affinity Ca2+ sensor for SK channels. Studies on human calmodulinopathy have provided evidence for the critical roles of CaM in the functional regulation of SK channels (Yu et al., 2016; Saljic et al., 2019). However, the mechanistic effects on SK channels by mutations in CaM remain incompletely understood.
Ca2+-activated K+ (KCa) channels can be divided into three main subfamilies based on their electrophysiological, pharmacological, and molecular phenotypes. These include large-conductance Ca2+- and voltage-activated K+ channels (BK), the intermediate-conductance KCa channels (IK or SK4), and the small-conductance KCa channels (SK1, 2, and 3; Köhler et al., 1996; Ishii et al., 1997; Joiner et al., 1997; Vergara et al., 1998). SK4 channels have intermediate single-channel conductance and are sensitive to charybdotoxin. In contrast, SK channels have small single-channel conductance and are sensitive to apamin. Recent cryo-EM structure of the SK4 channel–CaM complex (Lee and MacKinnon, 2018) reveals atomic details of the SK channel C-terminal interaction with the C-lobe of CaM, demonstrating that the C-lobe binds to the channel constitutively, while the N-lobe interacts with the S4-S5 linker in a Ca2+-dependent manner. The S4-S5 linker undergoes conformational changes upon CaM binding to open the channel pore (Lee and MacKinnon, 2018). CaM opens SK channels with a median effective intracellular concentration of Ca2+ of ∼100–400 nM (Köhler et al., 1996; Xia et al., 1998).
Here, we take advantage of the SK4 channel–CaM complex structure as a template for Rosetta molecular modeling, combined with MD simulation and biochemical and functional analyses to test the effects of the known CaM mutations on cardiac SK2 channel functions. Specifically, we tested the hypothesis that human CaM mutations linked to sudden cardiac death disrupt SK channel function by distinct mechanisms. Mutations in the Ca2+-binding domain of CaM (CALM1-D130G, CALM2-D96V) result in a dominant-negative (DN) effect, while specific mutations with phenylalanine to leucine (CALM1-F90L and -F142L) disrupt the interactions between apo-CaM with the CaMBD of SK channels. SK currents have been shown to be prominently expressed not only in atrial myocytes (Xu et al., 2003; Zhang et al., 2015) but also in cardiac Purkinje cells (Reher et al., 2017). Indeed, Purkinje cells have been shown to be the probable site of origin of cardiac arrhythmias, including in patients with heritable arrhythmia syndromes (Haïssaguerre et al., 2002; Wilde et al., 2019). Additionally, SK channels have been shown to be expressed and play important roles in pacemaking cells, including sinoatrial and atrioventricular nodes (Zhang et al., 2008; Torrente et al., 2017). Distinct mutant CaMs may result in a significant reduction in the activation of the SK channels, leading to a decrease in the key Ca2+-dependent repolarization currents mediated by SK channels, exacerbating the effects of CaM mutations in LQTS or IVF.
Materials and methods
Plasmid construction
Human filamin A (FLNA) in pREP4 vector (Life Technologies) was a kind gift from Dr. Paramita M. Ghosh (University of California Davis, Davis, CA). α-Actinin2 cDNA in pcDNA3 vector was a kind gift from Dr. David Fedida (University of British Columbia, Vancouver, BC, Canada; Cukovic et al., 2001). Construction of SK2 expression plasmids for heterologous expression in human embryonic kidney (HEK) 293 cells was as follows: full-length human cardiac SK2 cDNA was subcloned into pIRES2-EGFP (Takara Bio USA, Inc.) to obtain pSK2-IRES-EGFP plasmid. SK2 channel fusion constructs were generated harboring WT CaM (CaMWT; human SK2 [hSK2]-G4-CaMWT) or mutant CaMs (hSK2-G4 mutant CaM) to specifically occupy the CaMBD of SK2 channels with CaMWT or mutant CaMs using flexible linkers (glycine linkers) as previously described by Mori et al. (2004).
To study the subcellular localization of the SK2 channel subunit, modified human influenza hemagglutinin (HA) tag was inserted into the extracellular S1-S2 loop of the channel. Specifically, modified HA epitope was flanked with the ClC-5 chloride channel D1-D2 loop to increase accessibility and inserted in the end of the S1-S2 loop of the SK2 channel subunit as we described previously (Kim et al., 2011). The inserted amino acid sequence was NSEHYPYDVPDYAVTFEERDKCPEWNC. The epitope is shown in bold. Epitope tags were generated by recombination polymerase chain reaction and verified by automated sequencing.
HEK 293 cells and plasmid transfection
HEK 293 cells were maintained in Dulbecco’s modified Eagle’s medium supplemented with 10% FBS. Cell lines were maintained at 37°C in a humidified atmosphere containing 5% CO2. All cell culture reagents were purchased from Life Technologies. HEK 293 cells were transfected using the following plasmid compositions: pSK2-IRES-EGFP, in combination with pREP4-FLNA and pcDNA3–α-actinin2, with WT (CaMWT) or mutant CaM (1 µg for each plasmid), using Lipofectamine 2000 (Life Technologies) according to the manufacturer’s protocol. The 1:1 ratio of the plasmids was determined to be most optimal for SK2 current expression in our prior published studies (Rafizadeh et al., 2014).
Patch-clamp recordings
Whole-cell Ca2+-activated K+ current (IK,Ca) was recorded from transfected HEK 293 cells and human induced pluripotent stem cell (hiPSC)–derived cardiomyocyte-like cells (hiPSC-CMs) at room temperature using conventional patch-clamp techniques as previously described (Xu et al., 2003). For current recordings, the extracellular solution contained (in mM) 140 N-methylglucamine, 4 KCl, 1 MgCl2, 5 glucose, and 10 HEPES, pH 7.4, with HCl. The internal solution contained (in mM) 144 potassium gluconate, 1.15 MgCl2, 5 EGTA, 10 HEPES, and CaCl2 yielding a free cytosol Ca2+ concentration of 500 nM, using the Maxchelator software by C. Patton of Stanford University as we have previously described (Zhang et al., 2017). The pH was adjusted to 7.25 using KOH. To isolate apamin-sensitive SK currents, extracellular solution containing apamin (10 or 100 nM) was applied during the recordings, and the difference currents between the control and the apamin-containing solution were calculated to be the apamin-sensitive currents. The current was elicited from a holding potential of −55 mV using a voltage ramp protocol ranging from −120 to +60 mV with a 2-s duration.
For hiPSC-CM recordings, 0.006 mM CaMWT or mutant CaM peptides was added to the intracellular solution, and the recordings were performed 3 min after forming whole-cell recording for complete dialysis of the peptides.
All recordings were performed using 3 M KCl agar bridges. Cell capacitance was calculated as the ratio of total charge (the integrated area under the current transient) to the magnitude of the pulse (20 mV). Currents were normalized to cell capacitance to obtain the current density. Series resistance was compensated electronically. In all experiments, a series resistance compensation of ≥90% was obtained. The currents were recorded using Axopatch 200A amplifier (Molecular Devices), filtered at 1 kHz using a four-pole Bessel filter, and digitized at a sampling frequency of 5 kHz. Data acquisition and analysis were performed using pClamp 10 software (Molecular Devices) and Origin Software (OriginLab).
Immunofluorescence confocal laser scanning microscopy of HEK 293 cells
HEK 293 cells were cotransfected with human cardiac SK2-HA (SK2 channel with the extracellular HA tag) together with α-actinin, FLNA, and CaMWT or mutant CaM, using Lipofectamine (Life Technologies; catalog #11668-019) as we previously described (Rafizadeh et al., 2014). After blocking with 1% BSA (Sigma; catalog #A7030) under nonpermeabilization (NP) condition, SK2 channels localized on the cell membrane were labeled with monoclonal anti-HA antibody (Covance; catalog #MMS-101P; 1:100 dilution) by incubating overnight in the humidified chamber (4°C), followed by treatment with a chicken anti-mouse Alexa Fluor 555 secondary antibody (Life Technologies; catalog #A-21200; 1:500 dilution) for 1 h at room temperature. Cells were then permeabilized with 0.01% Triton X (Fisher) and blocked with 1% BSA again, and intracellular SK2 channels were labeled with anti-HA antibody (1:250 dilution) at 4°C overnight and a rabbit anti-mouse Alexa Fluor 633 secondary antibody (Life Technologies; catalog #21427; 1:500 dilution). Coverslips were mounted using mounting medium containing 4′,6-diamidino-2-phenylindole (VectaMount; Vector Laboratories, Inc.; catalog #H-5000) and imaged under a Zeiss LSM 700 confocal laser scanning microscope.
A fluorescence ratio of 555/633 then represents the ratio of SK2 channel numbers on cell membranes over those inside the cells, as we previously described (Rafizadeh et al., 2014). Cells cotransfected with CaMWT or mutant forms of CaM were immunolabeled in parallel, and all the microscopic settings were kept constant for all groups. Blind analysis was performed on unaltered images using Fiji ImageJ.
hiPSC-CMs
Feeder-free hiPSCs (iPS-D19-9-T7; WiCell) were cultured with mTeSRTM on hiPSC-qualified Matrigel. hiPSCs were differentiated into hiPSC-CMs following the protocols we previously reported (Sirish et al., 2016; Yamoah et al., 2018). We have observed ∼80–90% cardiomyocytes as shown previously (Yamoah et al., 2018). The hiPSC-CMs at day 40 (maturing cardiomyocytes) were used for experiments. CaMWT and mutant CaM peptides were used to test the functional changes of endogenous SK currents in hiPSC-CMs. The effects of WT compared with mutant CaMs were tested on the apamin-sensitive currents.
Generation of recombinant CaM proteins
WT and mutant CaM proteins were prepared as previously described (Hwang et al., 2014). Briefly, the recombinant CaM cDNA subcloned into a pET15b vector was mutated using QuikChange site-directed mutagenesis (Agilent Technologies). Proteins were expressed in Escherichia coli BL21 (DE3) cells and purified by hydrophobic chromatography using a phenyl sepharose column. Purified protein was dialyzed overnight at 4°C twice in 50 mM HEPES at pH 7.4, 100 mM KCl, and 5 mM EGTA and twice more with the same buffer except EGTA was lowered to 0.05 mM to remove Ca2+. The molecular mass of all proteins was confirmed using negative electrospray mass spectroscopy.
Structural modeling of CaM, CaM mutants, and the hSK2 channel
Structural modeling of CaMWT, CaMF90L, CaMF93L, and CaMF142L and the hSK2 channel was performed using Rosetta loop modeling and relax applications (Bonneau et al., 2001; Rohl et al., 2004; Yarov-Yarovoy et al., 2006; Wang et al., 2007; Yarov-Yarovoy et al., 2012; Bender et al., 2016; Alford et al., 2017) based on the hSK4 bound to the apo-CaM cryo-EM structure (PDB accession no. 6CNM; Lee and MacKinnon, 2018) as a template.
First, we directly compared the sequence between hSK2 (UniProt accession no. Q9H2S1) with the cryo-EM structure of hSK4 (PDB accession no. 6CNM) that was used as the template for molecular modeling. The sequence identity between the two structures is 46%, while the sequence similarity is 63%. We further compared the sequence for the CaMBD (amino acid residues 412–488 in hSK2, DTQLTKRVKNAAANVLRETWLIYKNTKLVKKIDHAKVRKHQRKFLQAIHQLRSVKMEQRKLNDQANTLVDLAKTQNI). The sequence identity between the two CaMBDs is 47% with a sequence identity of 68%.
The hSK2 channel sequence was threaded onto the hSK4 structure. 1,000 models of the hSK2 channel were generated using the Rosetta relax application and then clustered; the top model after clustering was selected as the best hSK2 model. The hSK2 model was used to replace the hSK4 peptide in the CaMBD based on hSK4–CaM complex structure (Lee and MacKinnon, 2018). 1,000 models of the hSK2–CaM complex were generated using the Rosetta relax application and then clustered, and the top model after clustering was selected as the best hSK2-CaM complex model, shown in Fig. 6. To determine the structure of the C-lobe of apo-CaM, CaM was taken from the 6CNM structure, mutated for the mutant models, relaxed, and finally clustered with the top models shown in Fig. 6. The mutant CaM models were created by mutating CaMWT and then using the Rosetta relax application to model potential conformational changes, induced by specific CaM mutations. 10,000 models were generated, the 1,000 lowest energy models were clustered, and the top models after clustering for each CaM mutant were selected as the best model. Molecular graphics and analyses were performed with the University of California San Francisco (UCSF) Chimera package (Pettersen et al., 2004).
MD simulations of WT and mutant CaM
Structures representing WT and mutant apo-CaM were obtained from Rosetta modeling as described above. The webserver CHARMM-GUI was used to prepare apo-CaM and create the solvent box (Jo et al., 2008; Brooks et al., 2009; Lee et al., 2016). The protein was immersed in a TIP3 water box with dimensions of 9 nm on all sides. Na and Cl ions were added to a final concentration of 150 mM to neutralize charge. In total, the system was composed of ∼15,000 (±1,000) atoms (Jo et al., 2008; Brooks et al., 2009; Lee et al., 2016). Periodic boundary conditions with dimensions equivalent to the water box were used (Phillips et al., 2005).
Equilibration and production runs were performed using NAMD 2.13 (Phillips et al., 2005) on a local GPU cluster using CHARMM36m force fields (Vanommeslaeghe et al., 2010; Huang et al., 2017). After 100,000 steps of steepest descent minimization, MD simulations started with a time step of 1 fs, with harmonic restraints initially applied to protein heavy atoms. These restraints were slowly released over 0.5 ns. These systems were equilibrated further for 0.2 ns with a time step of 2 fs. To use a 2-fs time step, all bonds to H atoms were constrained using the SHAKE algorithm. All simulations were performed in isothermal-isobaric ensemble. Temperature was maintained at 303.15 K using a Langevin thermostat, and constant pressure at 1 atm was maintained using a Nosé-Hoover barostat.
Electrostatic interactions were computed using the Particle Mesh Ewald method (Darden et al., 1993). Non-bonded pair lists were updated every 10 steps with a list cutoff distance of 16 Å and a real space cutoff of 12 Å, with energy switching starting at 10 Å. After equilibration of 1 ns, the simulation was continued for 1 µs. The full trajectory, including the equilibration and production steps, was used to calculate the RMSD. The 1-µs trajectory of the simulation during the production step was saved every 0.2 ns. This trajectory was then used for analysis.
RMSD
All measurements and renderings were performed using Visual Molecular Dynamics (Humphrey et al., 1996). RMSD was calculated every 0.2 ns over the equilibration, and production steps aligned to the first frame in the equilibration step. Specifically, for every 0.2 ns of the production simulation, the RMSD was calculated for apo-CaMWT and the mutant CaMs. The structures were clustered, and the cluster with the most decoys was selected as the most probable, with the center of the cluster being used as the representative model. Each mutant CaM structure was aligned individually to the CaMWT representative model using the matchmaker tool in UCSF Chimera, and the distance between the C-α of each residue was calculated. The averaged distance and the SD for each comparison for the full C-terminal domain (residues 81–145) and for Ca2+ binding site 4 (residues 131–138) was quantified (Table S1 and Table S2).
Root mean square fluctuation (RMSF)
Per residue RMSF was calculated over a sliding average window of 1 ns time and averaged over the 1-µs trajectory. The per-residue RMSF is presented as mean ± SD.
Ensembles were generated by clustering the MD run as described in the structural modeling section. The top 10 models were displayed and aligned with the top cluster model.
Data analysis
Data analysis from patch-clamp recordings was performed by using Origin software (OriginLab). Current density obtained from cells expressing SK2 with CaMWT or mutant CaM was compared by normalizing the currents with cell capacitance. Where appropriate, pooled data are presented as means ± SEM. Statistical comparisons were performed using one-way and two-way ANOVA combined with Tukey’s post hoc analyses. One-way ANOVA was used to determine statistically significant differences among three or more groups in Fig. 1, Fig. 2, Fig. 4, and Fig. 5. Two-way ANOVA was used to determine statistically significant differences with two independent variables among three or more groups in Fig. 3. Post hoc analyses were performed using Tukey’s test. Statistical analyses were performed using GraphPad Prism and Origin Software.
Online supplemental material
Fig. S1 shows summary data for fluorescence intensity under NP (left bars) and permeabilized (right bars) conditions for HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G). Fig. S2 shows RMSD graphs that include the equilibration and production stages. Fig. S3 shows the top 10 clustered models of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L aligned to represent an ensemble of possible states, illustrating the distribution of helix 3 and Ca2+-binding site 4 regions. Fig. S4 presents graphical comparisons from MD simulations of the C-lobe of apo-CaMs showing hydrogen bond in the apo-CaMWT. The side chain of Asn112 on helix 2 in apo-CaMWT maintains a stable hydrogen bond with the carboxyl oxygen of Ala89 residue on helix 1 during simulations of apo-CaMWT, which is not observed in the simulations of the CaM mutants. Fig. S5 shows comparison of the RMSF of Ca2+-CaMF142L from MD simulations (A) and N-H heteronuclear NOE saturated/unsaturated ratio (B; BMRB: 34262; Wang et al., 2018), with similar flexibility including identification of the most flexible amino acid being Lys116 residue. Video 1, Video 2, and Video 3 show morph videos from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta models of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L, respectively. Video 4, Video 5, and Video 6 show morph videos from the top MD model of the C-lobe of apo-CaMWT to the top MD models of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L, respectively. Table S1 shows quantification of the RMSD calculated from the production portion of MD presented in Fig. S2. Table S2 shows quantification of the differences in the RMSD among WT and mutant CaMs, derived from the C-terminal domain (residues 81–145) and Ca2+ binding site 4 (residues 131–138) presented in Fig. 8.
Results
Inhibitory effects of human CaM mutations on SK channels
We first coexpressed human cardiac SK2 channels with different human mutant CaMs compared with CaMWT in HEK 293 cells. Our previous studies showed that cytoskeletal proteins are critical for the proper membrane localization of SK2 channels; therefore, α-actinin2 and FLNA were also used for coexpression in all groups (Lu et al., 2009; Rafizadeh et al., 2014; Zhang et al., 2017).
As positive controls, we tested the roles of the N- and C-lobe mutant CaMs in the regulation of SK2 currents using CaM1,2 and CaM3,4, where the two Ca2+-binding sites in the N- and C-lobe are mutated, respectively, as well as a mutant CaM (CaM1,2,3,4) that abolishes the four high affinity Ca2+-binding sites (Xia et al., 1998). Apamin-sensitive IK,Ca is recorded at baseline (before apamin application, black traces) and after apamin application (10 nM, red traces, Fig. 1 A). Both EF hands in the N- and C-lobe of CaM play critical roles in activation of the apamin-sensitive current (Fig. 1 A). Moreover, CaM1,2,3,4 results in the most significant inhibition on the apamin-sensitive current compared with CaM1,2 and CaM3,4 (Fig. 1 A). Summary data for the density of apamin-sensitive currents at the test potentials of −120 and +60 mV are shown in Fig. 1 B. The results are distinct from those previously reported demonstrating the critical roles of Ca2+ binding to the N- but not the C-lobe of CaM in SK channel gating in oocytes (Keen et al., 1999). However, our experiments were performed in a mammalian expression system with coexpression of α-actinin2 and FLNA. Furthermore, our previous study documented binding of α-actinin2 to the CaMBD of SK2 channels (Lu et al., 2007; Lu et al., 2009; Rafizadeh et al., 2014).
Mutations in the N-lobe and C-lobe of CaM affect the currents of hSK2 expressed in HEK 293 cells. (A) hSK2 channels were coexpressed with CaMWT or mutant CaMs (CaM1,2, CaM3,4, or CaM1,2,3,4). Current was recorded before (black trace) and after (red trace) application of apamin (10 nM). (B) Summary data at +60 and −120 mV indicating significant reduction of the SK currents by mutant CaMs (n = 12 cells; *, P < 0.05). (C and D) hSK2 channels were coexpressed with CaMWT (black bars), CaMN54I (red bars), CaMF90L (green bars), CaMD96V (blue bars), CaMN98S (cyan bars), and CaMD130G (magenta bars). (C) Current was recorded before (black trace) and after (red trace) application of apamin (10 nM). (D) Summary data at −120 mV and +60 mV (n = 10–13 cells; *, P < 0.05). For B and D, the colors of the asterisks denote comparisons with the corresponding bar graphs of the same colors, using one-way ANOVA combined with Tukey's test (at −120 mV, P = 0.00001 for all pairwise comparisons except P = 0.04 for CaMN54I versus CaMD96V, and P = NS for CaMD96V versus CaMD130G and CaMWT versus CaMF90L; at +60 mV, P = 0.00001 for all pairwise comparisons except P= NS for CaMD96V versus CaMD130G). Data shown represents mean ± SEM.
Mutations in the N-lobe and C-lobe of CaM affect the currents of hSK2 expressed in HEK 293 cells. (A) hSK2 channels were coexpressed with CaMWT or mutant CaMs (CaM1,2, CaM3,4, or CaM1,2,3,4). Current was recorded before (black trace) and after (red trace) application of apamin (10 nM). (B) Summary data at +60 and −120 mV indicating significant reduction of the SK currents by mutant CaMs (n = 12 cells; *, P < 0.05). (C and D) hSK2 channels were coexpressed with CaMWT (black bars), CaMN54I (red bars), CaMF90L (green bars), CaMD96V (blue bars), CaMN98S (cyan bars), and CaMD130G (magenta bars). (C) Current was recorded before (black trace) and after (red trace) application of apamin (10 nM). (D) Summary data at −120 mV and +60 mV (n = 10–13 cells; *, P < 0.05). For B and D, the colors of the asterisks denote comparisons with the corresponding bar graphs of the same colors, using one-way ANOVA combined with Tukey's test (at −120 mV, P = 0.00001 for all pairwise comparisons except P = 0.04 for CaMN54I versus CaMD96V, and P = NS for CaMD96V versus CaMD130G and CaMWT versus CaMF90L; at +60 mV, P = 0.00001 for all pairwise comparisons except P= NS for CaMD96V versus CaMD130G). Data shown represents mean ± SEM.
Next, we tested the effects of LQTS (CaMD96V, CaMD130G), CPVT (CaMN54I, CaMN98S), and IVF (CaMF90L) CaM mutants compared with CaMWT. Fig. 1 C shows the most prominent inhibitory effects on the density of apamin-sensitive currents by the two LQTS mutations (CaMD96V and CaMD130G). CaMF90L resulted in the least inhibitory effects. The current densities at −120 mV and +60 mV are shown in Fig. 1 D.
No significant alteration in SK2 channel trafficking by human mutant CaMs
The effects of CaM mutations on SK2 current may result from a decrease in channel activation or trafficking to the cell membrane. We therefore tested if the reduction in the SK2 current density from coexpression of mutant CaMs results from a decrease in membrane trafficking. Immunofluorescence confocal microscopy was performed using SK2 channels harboring an HA tag in the extracellular S1-S2 loop (SK2-HA) by using NP compared with permeabilized (P) conditions as we previously described (Fig. 2 and Fig. S1; Rafizadeh et al., 2014). Our data suggest that the mutant CaM does not alter SK channel trafficking.
Trafficking of hSK2 channels after coexpression with CaMWT or mutant CaMs. (A) HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G) using NP and permeabilized (P) conditions. (B) Control represents incubation with secondary antibody alone. Scale bars are 10 µm. (C) Summary data showing fluorescence intensity under NP/P conditions (555/633 fluorescent intensity). Data shown represent mean ± SEM. Statistical analyses were performed using one-way ANOVA combined with Tukey’s test. n = 15–20 cells; P= NS. DAPI, 4′,6-diamidino-2-phenylindole.
Trafficking of hSK2 channels after coexpression with CaMWT or mutant CaMs. (A) HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G) using NP and permeabilized (P) conditions. (B) Control represents incubation with secondary antibody alone. Scale bars are 10 µm. (C) Summary data showing fluorescence intensity under NP/P conditions (555/633 fluorescent intensity). Data shown represent mean ± SEM. Statistical analyses were performed using one-way ANOVA combined with Tukey’s test. n = 15–20 cells; P= NS. DAPI, 4′,6-diamidino-2-phenylindole.
Trafficking of hSK2 channels after coexpression with CaMWT or mutant CaMs. HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G) using NP and permeabilized conditions. Summary data showing fluorescence intensity under NP (left bars) and permeabilized (right bars) conditions (555 and 633 fluorescent intensities in arbitrary units [AU] × 1,000; n = 15–20 cells; P = NS comparing NP with permeabilized for each group). Data shown represents mean ± SEM.
Trafficking of hSK2 channels after coexpression with CaMWT or mutant CaMs. HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G) using NP and permeabilized conditions. Summary data showing fluorescence intensity under NP (left bars) and permeabilized (right bars) conditions (555 and 633 fluorescent intensities in arbitrary units [AU] × 1,000; n = 15–20 cells; P = NS comparing NP with permeabilized for each group). Data shown represents mean ± SEM.
Inhibitory effects of mutant CaMs on SK2 channels revealed by SK2 fusion protein harboring CaMWT or mutant CaMs via a flexible glycine (Gn) linker
There are some possible caveats inherent in the experiments presented in Fig. 1 since mutations in CaMs may alter the efficiency of CaM expression in HEK 293 cells. To circumvent these possible confounding factors, we generated SK2 channel fusion constructs harboring CaMWT (hSK2-G4-CaMWT) or mutant CaMs (hSK2-G4-CaMMUT) to specifically occupy the CaMBD of SK2 channels with CaMWT or mutant CaMs using flexible linkers (Gn linker; Fig. 3 A), as previously described by Mori et al. (2004). This experimental paradigm enabled us to selectively test the contribution of the mutant CaMs compared with that of CaMWT on SK2 channels (Fig. 3 A).
hSK2-CaM fusion proteins exhibit similar inhibitory effects to that of coexpressed hSK2 channels with CaM. (A) Diagram illustrating experimental paradigms using SK2 fusion proteins with flexible Gn linker fused to CaMWT (upper panel) compared with mutant CaMs (lower panel). (B) Current recordings before (black trace) and after (red trace) apamin application (10 nM) for hSK2 expressed with CaMWT versus hSK2-G4-CaMWT versus SK2-G4-CaMWT expressed with DN CaM1,2,3,4. (C) Summary data at −120 mV and +60 mV from B. (D and E) Summary data for apamin-sensitive current density in HEK 293 cells coexpressing hSK2 with CaMWT or mutant CaM constructs (left bars) compared with SK2-G4-CaMWT or SK2-G4-CaMMUT (hashed bars on the right), where CaMMUT refers to CaMN54I (red bars), CaMF90L (green bars), CaMF93L (purple bars), CaMD96V (blue bars), CaMN98S (cyan bars), CaMD130G (magenta bars), and CaMF142L (yellow bars) at −120 mV (D) and +60 mV (E; n = 10–13 cells). Data represent mean ± SEM. Statistical analysis was performed using two-way ANOVA combined with Tukey’s test. The colors of the asterisks denote comparisons with the corresponding bar graphs of the same colors. *, P < 0.05. At −120 mV, P = 0.00001 for all pairwise comparisons except P = 0.04 for CaMN54I versus CaMF93L and CaMN54I versus CaMD96V, and P= NS for CaMWT versus CaMF90L, CaMF93L versus CaMD96V, CaMF93L versus CaMD130G, CaMD96V versus CaMD130G, and CaMN98S versus CaMF142L. At +60 mV, P = 0.00001 for all pairwise comparisons except P= NS for CaMD96V versus CaMD130G and CaMN98S versus CaMF142L. There were no significant differences between hSK2 coexpression with CaM compared with hSK2-G4-CaM for WT or mutant CaMs (P = NS).
hSK2-CaM fusion proteins exhibit similar inhibitory effects to that of coexpressed hSK2 channels with CaM. (A) Diagram illustrating experimental paradigms using SK2 fusion proteins with flexible Gn linker fused to CaMWT (upper panel) compared with mutant CaMs (lower panel). (B) Current recordings before (black trace) and after (red trace) apamin application (10 nM) for hSK2 expressed with CaMWT versus hSK2-G4-CaMWT versus SK2-G4-CaMWT expressed with DN CaM1,2,3,4. (C) Summary data at −120 mV and +60 mV from B. (D and E) Summary data for apamin-sensitive current density in HEK 293 cells coexpressing hSK2 with CaMWT or mutant CaM constructs (left bars) compared with SK2-G4-CaMWT or SK2-G4-CaMMUT (hashed bars on the right), where CaMMUT refers to CaMN54I (red bars), CaMF90L (green bars), CaMF93L (purple bars), CaMD96V (blue bars), CaMN98S (cyan bars), CaMD130G (magenta bars), and CaMF142L (yellow bars) at −120 mV (D) and +60 mV (E; n = 10–13 cells). Data represent mean ± SEM. Statistical analysis was performed using two-way ANOVA combined with Tukey’s test. The colors of the asterisks denote comparisons with the corresponding bar graphs of the same colors. *, P < 0.05. At −120 mV, P = 0.00001 for all pairwise comparisons except P = 0.04 for CaMN54I versus CaMF93L and CaMN54I versus CaMD96V, and P= NS for CaMWT versus CaMF90L, CaMF93L versus CaMD96V, CaMF93L versus CaMD130G, CaMD96V versus CaMD130G, and CaMN98S versus CaMF142L. At +60 mV, P = 0.00001 for all pairwise comparisons except P= NS for CaMD96V versus CaMD130G and CaMN98S versus CaMF142L. There were no significant differences between hSK2 coexpression with CaM compared with hSK2-G4-CaM for WT or mutant CaMs (P = NS).
We first demonstrated that the fusion proteins behave as expected by directly comparing apamin-sensitive IK,Ca recorded from HEK 293 cells expressing hSK2-G4-CaMWT versus hSK2 coexpressed with CaMWT (Fig. 3, B and C). Furthermore, coexpression of DN CaM (CaM1,2,3,4) with hSK2-G4-CaMWT failed to knock down apamin-sensitive current (Fig. 3, B and C) in contrast to hSK2 coexpressed with CaM1,2,3,4, as shown in Fig. 1 A.
Having determined that the fusion proteins function as expected, we compared the density of apamin-sensitive currents between hSK2-G4-CaMMUT and coexpression of the SK2 channel and mutant CaMs. The density is comparable, with the most significant inhibitory effects on IK,Ca by the two LQTS mutations tested (CaMD96V and CaMD130G), consistent with a DN effect from these mutations (Fig. 3, D and E). We further tested the effects of two additional mutant CaMs (F93L and F142L). CaMF142L is shown to be linked to LQTS, while the Phe93 residue is predicted from our molecular model to interact with CaMBD of SK2 channels (described in a later section). These two mutations produced intermediate effects on IK,Ca (Fig. 3, D and E).
Functional effects of mutant CaMs on SK currents in hiPSC-CMs
Thus far, the experiments have relied on a heterologous expression system. We took advantage of hiPSC-CMs. The rationale was to determine the functional effects of mutant CaMs on endogenous SK currents with accompanying ion channel interacting proteins within a human cardiomyocyte-like cell. Even though hiPSC-CMs are relatively immature, we demonstrated the expression of SK2 channels in hiPSC-CMs using immunofluorescence confocal microscopy (Fig. 4 A). Fig. 4 B shows apamin-sensitive current in hiPSC-CMs when the cells were dialyzed by WT or mutant CaM proteins. Dialysis of mutant CaMD96V proteins resulted in the most prominent knockdown of apamin-sensitive currents in hiPSC-CMs, consistent with its DN effects, compared with the effects of CaMF90L (Fig. 4, C and D).
Regulation of endogenous SK currents in hiPSC-CMs by intracellular CaM peptides. (A) hiPSC-CMs expressing SK2 channels (green) and α-actinin2 (red). 4′,6-Diamidino-2-phenylindole stain is shown in blue. Scale bar is 10 µm. (B) Effects of mutant CaM peptides (CaMN54I, CaMF90L, and CaMD96V) compared with CaMWT peptide on apamin-sensitive SK currents in hiPSC-CMs. The current was recorded before (black trace) and after (red trace) apamin application (100 nM). (C) Subtracted apamin-sensitive SK currents in the presence of CaMWT and mutant CaM peptides. (D) Comparisons of the apamin-sensitive current densities at −120 mV and +60 mV (n = 8–10 cells). Data represent mean ± SEM. Statistical analyses were performed using one-way ANOVA combined with Tukey’s test; *, P = 0.012 at −120 mV, and *, P = 0.037 at +60 mV between CaMWT and CaMD96V.
Regulation of endogenous SK currents in hiPSC-CMs by intracellular CaM peptides. (A) hiPSC-CMs expressing SK2 channels (green) and α-actinin2 (red). 4′,6-Diamidino-2-phenylindole stain is shown in blue. Scale bar is 10 µm. (B) Effects of mutant CaM peptides (CaMN54I, CaMF90L, and CaMD96V) compared with CaMWT peptide on apamin-sensitive SK currents in hiPSC-CMs. The current was recorded before (black trace) and after (red trace) apamin application (100 nM). (C) Subtracted apamin-sensitive SK currents in the presence of CaMWT and mutant CaM peptides. (D) Comparisons of the apamin-sensitive current densities at −120 mV and +60 mV (n = 8–10 cells). Data represent mean ± SEM. Statistical analyses were performed using one-way ANOVA combined with Tukey’s test; *, P = 0.012 at −120 mV, and *, P = 0.037 at +60 mV between CaMWT and CaMD96V.
Altered interaction between the SK2 channel and mutant CaMs
Coimmunoprecipitation was performed as we previously described (Lu et al., 2007) between SK2-HA channels (Fig. 5 A) and different mutant CaMs compared with CaMWT. Since native cells contain a large amount of CaM, we constructed a fusion protein of CaM with FLAG-tag (DYKDDDDK; CaMFLAG) to enable identification of transfected mutant CaMs versus endogenous CaMWT (Fig. 5 B). There was a significant decrease in SK2 channels complexed with mutant CaMF90L in the absence of Ca2+ with EGTA (Fig. 5 C). Negative control experiments were performed using nontransfected cells and cells transfected with SK2-HA alone, without CaMFLAG. The data suggest a significant reduction in SK2 channel complexed with apo-CaMF90L compared with other CaM mutants, supporting the lack of DN effects with relatively modest inhibition of CaMF90L on SK2 currents.
Interactions of mutant CaMs with SK2 channels revealed by coimmunoprecipitation assay. (A and B) Diagrams depicting the SK2 channel subunit with an HA tag in the S1-S2 extracellular loop and CaM with a FLAG tag in the C terminus under the cytomegalovirus (CMV) promoter. RBS, ribosome-binding site. (C) Coimmunoprecipitation using anti-Flag antibody (α-Flag) to pull down SK2 channels identified using anti–HA antibody (α-HA) in the presence of 2 mM EGTA. The right two lanes show the results from nontransfected HEK 293 cells (NT) and cells transfected with SK2 channels only (SK2) as negative controls. (D) Summary data of the findings in C. The background subtracted intensity for SK2 bands using anti–HA antibody was normalized to CaM bands using anti-Flag antibody (NIH ImageJ). Data shown represent mean ± SEM (n = 3). Statistical analyses were performed using one-way ANOVA combined with Tukey’s test; *, P < 0.05 for CaMWT compared with CaMF90L, CaMD96V, CaMN98S, and CaMD130G, and *, P < 0.05 for CaMF90L compared with CaMD96V, CaMN98S, and CaMD130G (the P values are as follows: CaMWT versus CaMF90L, 0.00003; CaMWT versus CaMD96V, 0.002; CaMWT versus CaMN98S, 0.004; CaMWT versus CaMD130G, 0.004; CaMF90L versus CaMD96V, 0.03; CaMF90L versus CaMN98S, 0.02; and CaMF90L versus CaMD130G, 0.02). IP, immunoprecipitation.
Interactions of mutant CaMs with SK2 channels revealed by coimmunoprecipitation assay. (A and B) Diagrams depicting the SK2 channel subunit with an HA tag in the S1-S2 extracellular loop and CaM with a FLAG tag in the C terminus under the cytomegalovirus (CMV) promoter. RBS, ribosome-binding site. (C) Coimmunoprecipitation using anti-Flag antibody (α-Flag) to pull down SK2 channels identified using anti–HA antibody (α-HA) in the presence of 2 mM EGTA. The right two lanes show the results from nontransfected HEK 293 cells (NT) and cells transfected with SK2 channels only (SK2) as negative controls. (D) Summary data of the findings in C. The background subtracted intensity for SK2 bands using anti–HA antibody was normalized to CaM bands using anti-Flag antibody (NIH ImageJ). Data shown represent mean ± SEM (n = 3). Statistical analyses were performed using one-way ANOVA combined with Tukey’s test; *, P < 0.05 for CaMWT compared with CaMF90L, CaMD96V, CaMN98S, and CaMD130G, and *, P < 0.05 for CaMF90L compared with CaMD96V, CaMN98S, and CaMD130G (the P values are as follows: CaMWT versus CaMF90L, 0.00003; CaMWT versus CaMD96V, 0.002; CaMWT versus CaMN98S, 0.004; CaMWT versus CaMD130G, 0.004; CaMF90L versus CaMD96V, 0.03; CaMF90L versus CaMN98S, 0.02; and CaMF90L versus CaMD130G, 0.02). IP, immunoprecipitation.
Structural modeling of CaMF90L, CaMF93L, and CaMF142L mutants and their interactions with SK2 channels
To explore potential structural changes induced by CaM mutations, we generated structural models of the C-lobe of apo-CaMWT, apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L using Rosetta structural modeling. The models of CaM reveal that Phe → Leu mutations resulted in conformational changes that affected helix packing in the C-lobe due to the substitution of a relatively large phenylalanine side chain to a smaller leucine side chain (Fig. 6, B–D). The Phe90 side chain is surrounded by the following residues in apo-CaMWT: Arg87 in helix 1; Ile101 in loop 1–2; Val137 in loop 3–4; and Tyr139 and Phe142 in helix 4. The Phe93 side chain is surrounded by the following residues in apo-CaMWT: Ile101 in loop 1–2; Leu106 and Val109 in helix 2; and Phe142, Met145, and Met146 in helix 4. The Phe142 side chain is surrounded by the following residues in apo-CaMWT: Ile86, Phe90, and Phe93 in helix 1; Ile101 in loop 1–2 and Leu106 in helix 2; Val137 in loop 3–4; and Tyr139, Met145, and Met146 in helix 4. Notably, Phe90, Phe93, and Phe142 are interacting with each other in apo-CaMWT. The conformational effects of the Phe → Leu mutations are explored in more detail using MD simulations.
Structural modeling of the interactions between CaM mutants and SK2 CaMBD. (A) Schematic of the four α-helices within the C-lobe of CaM. Location of mutations are shown by labels and colored markers. (B–D) Comparisons of CaMWT (green) and CaMF90L (red; B), CaMF93L (yellow; C), and CaMF142L (purple; D). Side chains of key amino acid residues are shown in stick representation using the color scheme shown in A. Conformational changes due to CaM mutation are indicated by black arrows in each panel. (E) C-lobe of apo-CaMWT (colored in green) bound to the C terminus of hSK2 channel (colored in light brown). Side chains of key amino acids are shown using space-filling representation. (F) Panel E rotated 90° to the left around the y axis. Molecular modeling was performed in Ca2+-free conditions.
Structural modeling of the interactions between CaM mutants and SK2 CaMBD. (A) Schematic of the four α-helices within the C-lobe of CaM. Location of mutations are shown by labels and colored markers. (B–D) Comparisons of CaMWT (green) and CaMF90L (red; B), CaMF93L (yellow; C), and CaMF142L (purple; D). Side chains of key amino acid residues are shown in stick representation using the color scheme shown in A. Conformational changes due to CaM mutation are indicated by black arrows in each panel. (E) C-lobe of apo-CaMWT (colored in green) bound to the C terminus of hSK2 channel (colored in light brown). Side chains of key amino acids are shown using space-filling representation. (F) Panel E rotated 90° to the left around the y axis. Molecular modeling was performed in Ca2+-free conditions.
Additionally, we generated a structural model of apo-CaMWT in complex with CaMBD from the SK2 channel based on the cryo-EM structure of the SK4 channel in complex with apo-CaM (PDB accession no. 6CNM; Lee and MacKinnon, 2018). The three phenylalanine residues (Phe90, Phe93, and Phe142) from the C-lobe of CaM can be seen interacting with hydrophobic residues from the CaMBD of SK2 channels (Fig. 6, E and F). Consistent with our hypothesis and the coimmunoprecipitation study (Fig. 5), we predict that substitution of the phenylalanine side chain may result in a significant decrease in the interaction of apo-CaM and SK2 CaMBD. Even though CaMF90L and CaMF142L were previously described to be linked to human arrhythmias, CaMF93L has not been described in patients. Our molecular modeling predicts that the Phe93 residue intimately interacts with SK2 CaMBD. Indeed, CaMF93L results in a significant decrease in SK2 current, supported by our functional analyses (Fig. 3, D and E).
MD simulations provide insights into the conformational changes of CaMF90L, CaMF93L, and CaMF142L
To further explore potential conformational changes induced by the Phe → Leu mutations in CaM, we performed MD simulations on the WT and three CaM mutants: F90L, F93L, and F142L. Each simulation was performed for 1 µs after equilibration. The simulation for apo-CaMWT was performed twice. The RMSDs demonstrated that the system equilibrated rapidly and that the RMSD had plateaued for the entire runs (Fig. S2 and Table S1).
Graphs of RMSD showing stability throughout the production run. RMSD graphs include the equilibration and prosecution stages. The RMSD from the C-lobe of apo-CaMWT (simulations [sims] 1 and 2), apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L are shown in blue, dark brown, red, yellow, and green, respectively. Runs are slightly different in length due to differences in equilibration time. The total simulation time is the same in all runs (1 µs).
Graphs of RMSD showing stability throughout the production run. RMSD graphs include the equilibration and prosecution stages. The RMSD from the C-lobe of apo-CaMWT (simulations [sims] 1 and 2), apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L are shown in blue, dark brown, red, yellow, and green, respectively. Runs are slightly different in length due to differences in equilibration time. The total simulation time is the same in all runs (1 µs).
Analyses of the RMSF show remarkable findings that the three Phe → Leu mutants affected the per-residue structure in similar manners (Fig. 7), even though the mutants are located in different helices of the C-lobe of CaM (helix 1 contains Phe90 and Phe93, and helix 4 contains Phe142). Compared with apo-CaMWT from the first simulation, all three CaM mutants exhibited more flexibility toward the C-terminal end of helix 2, while the C-terminal end of helix 3 and Ca2+-binding site 4 were less flexible. The largest shifts in RMSF were observed where the side chains of Phe90, Phe93, and Phe142 contact helices 2 and 3. This agrees with the notion that the Phe → Leu mutation creates a void volume, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions (Fig. 8, Fig. S3, Video 1, Video 2, Video 3, Video 4, Video 5, and Video 6). Differences in the RMSD between an equilibrated apo-CaMWT and mutant CaMs shown in Fig. 8 are quantified for the C-terminal domains as well as for Ca2+-binding site 4 (Table S2).
Graphs of RMSF showing regional changes in stability. RMSF from MD simulations of the C-lobe of apo-CaMWT is shown in blue (first simulation [sim]) and purple (second simulation), while RMSF from MD simulations of the C-lobe of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L mutants are shown in red, orange, and green, respectively. Diagrams above the graphs depict helices 1–4 from the C-lobe together with amino acid residues and are scaled to match the residue numbering of the graph. Locations of the three mutations are indicated by their respective colored dot on the graph and bars above the helices. Ca2+-binding sites are indicated with purple dots together with the numbers for the amino acid residues. Data shown are mean ± SD.
Graphs of RMSF showing regional changes in stability. RMSF from MD simulations of the C-lobe of apo-CaMWT is shown in blue (first simulation [sim]) and purple (second simulation), while RMSF from MD simulations of the C-lobe of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L mutants are shown in red, orange, and green, respectively. Diagrams above the graphs depict helices 1–4 from the C-lobe together with amino acid residues and are scaled to match the residue numbering of the graph. Locations of the three mutations are indicated by their respective colored dot on the graph and bars above the helices. Ca2+-binding sites are indicated with purple dots together with the numbers for the amino acid residues. Data shown are mean ± SD.
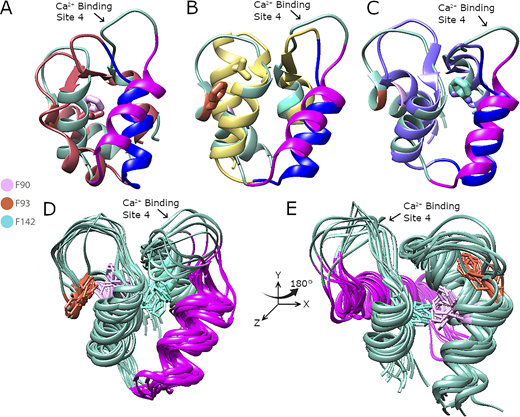
Graphical comparison from MD simulations of the C-lobe of apo-CaMs showing conformational changes in mutant CaMs with repacking of helices 2 and 3 and highly flexible Ca2+-binding site 4 in apo-CaMWT compared with the mutant CaMs, confirming the RMSF results.(A–C) Top clustered models of the C-lobe of apo-CaMF90L (red), apo-CaMF93L (yellow), and apo-CaMF142L (purple) are aligned with apo-CaMWT (green) in A, B, and C, respectively. Amino acid side chains of Phe90 (pink) and F90L (red) from apo-CaMWT and apo-CaMF90L (A), amino acid side chains of Phe93 (red) and F93L (yellow) from apo-CaMWT and apo-CaMF93L (B), and amino acid side chains of Phe142 (light blue) and F142L (light purple) from apo-CaMWT and apo-CaMF90L (C) are shown in the stick representation. Helix 3 from apo-CaMWT (magenta) resides at a larger angle in reference to the y axis compared with helices 3 from Phe → Leu mutant CaMs (dark blue) in A, B, and C, consistent with repacking of the structures. Quantification of the differences between apo-CaMWT and mutant CaMs shown in A, B, and C are summarized in Table S2. (D) Top 10 clustered models of apo-CaMWT aligned to represent an ensemble of possible states. Amino acid representation and color are conserved from A–C. Helix 3 is in purple. Helix 3 is mostly in a similar angle in reference to the y axis, with a much more extreme model present in the distribution. In addition, Ca2+-binding site 4 has a wide distribution of states, which does not align with those from the mutants (A–C). (E) Panel D rotated 180° to the right around the y axis.
Graphical comparison from MD simulations of the C-lobe of apo-CaMs showing conformational changes in mutant CaMs with repacking of helices 2 and 3 and highly flexible Ca2+-binding site 4 in apo-CaMWT compared with the mutant CaMs, confirming the RMSF results.(A–C) Top clustered models of the C-lobe of apo-CaMF90L (red), apo-CaMF93L (yellow), and apo-CaMF142L (purple) are aligned with apo-CaMWT (green) in A, B, and C, respectively. Amino acid side chains of Phe90 (pink) and F90L (red) from apo-CaMWT and apo-CaMF90L (A), amino acid side chains of Phe93 (red) and F93L (yellow) from apo-CaMWT and apo-CaMF93L (B), and amino acid side chains of Phe142 (light blue) and F142L (light purple) from apo-CaMWT and apo-CaMF90L (C) are shown in the stick representation. Helix 3 from apo-CaMWT (magenta) resides at a larger angle in reference to the y axis compared with helices 3 from Phe → Leu mutant CaMs (dark blue) in A, B, and C, consistent with repacking of the structures. Quantification of the differences between apo-CaMWT and mutant CaMs shown in A, B, and C are summarized in Table S2. (D) Top 10 clustered models of apo-CaMWT aligned to represent an ensemble of possible states. Amino acid representation and color are conserved from A–C. Helix 3 is in purple. Helix 3 is mostly in a similar angle in reference to the y axis, with a much more extreme model present in the distribution. In addition, Ca2+-binding site 4 has a wide distribution of states, which does not align with those from the mutants (A–C). (E) Panel D rotated 180° to the right around the y axis.
Top 10 clustered models of the C-lobe of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L aligned to represent an ensemble of possible states showing compact distribution of helix 3 and Ca2+-binding site 4 regions. Ensembles of the aligned top 10 clusters of the C-lobe of apo-CaMF90L (red; A), apo-CaMF93L (yellow; B), and apo-CaMF142L (purple; C). Helix 3 is shown in blue, and the Ca2+-binding site 4 is the loop immediately following helix 3 and is labeled. Lower panels are structures from the upper panels rotated 180° to the right around the y axis.
Top 10 clustered models of the C-lobe of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L aligned to represent an ensemble of possible states showing compact distribution of helix 3 and Ca2+-binding site 4 regions. Ensembles of the aligned top 10 clusters of the C-lobe of apo-CaMF90L (red; A), apo-CaMF93L (yellow; B), and apo-CaMF142L (purple; C). Helix 3 is shown in blue, and the Ca2+-binding site 4 is the loop immediately following helix 3 and is labeled. Lower panels are structures from the upper panels rotated 180° to the right around the y axis.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF90L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF90L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF93L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF93L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF142L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top Rosetta model of the C-lobe of apo-CaMWT to the top Rosetta model of apo-CaMF142L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF90L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF90L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF93L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF93L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF142L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
Morph video from the top MD model of the C-lobe of apo-CaMWT to the top MD model of apo-CaMF142L. The video illustrates the conformational changes that are required to accommodate a void volume that is created by the phenylalanine-to-leucine mutation, allowing helices 2 and 3 to repack their side chains and readjust their backbone positions. The coloring is the same as in Fig. 6 and Fig. 8 (residues Phe90, Phe93, and Phe142 are shown in pink, red, and light blue, respectively). Helices 2 and 3 are in the foreground, while helices 1 and 4 are in the background. Helix 3 is shown in magenta.
The RMSD from Simulation 2 of the apo-CaMWT exhibited a transient increase before returning to baseline toward the end of the simulation (Fig. S2). Closer examinations revealed that the major contributing regions of apo-CaMWT toward the transient increase in the RMSD in Simulation 2 are in helix 3, just proximal to Ca2+-binding site 4, similar to the conformational changes we observed in Simulation 1 but at a larger magnitude. Additionally, Simulation 2 of apo-CaMWT exhibited a similar per-residue RMSF pattern to Simulation 1, albeit lower in magnitude, as demonstrated in Fig. 7. This is consistent with the lower value of RMSD for Simulation 2 compared with Simulation 1 for the majority of the run. Finally, there was a return to the average RMSD after the transient increase, supporting the equilibration of the simulations.
Consistent with our observation, the side chain of Asn112 on helix 2 in apo-CaMWT maintained a stable hydrogen bond with the carboxyl oxygen of the Ala89 residue on helix 1 during the simulations of the apo-CaMWT, which was not observed in the simulations of the CaM mutants (Fig. S4). This may also, in part, influence flexibility toward the C-terminal end of helix 2 of the three mutant CaMs, as discussed above. We suggest that these observed changes may result in a decrease in the interaction of apo-CaM and SK2 CaMBD. Finally, to validate the MD simulations, we directly compared RMSF of Ca2+-CaMF142L from the MD simulation with the publicly available N-H heteronuclear NOE saturated/unsaturated ratio from a previously published NMR structure (BMRB accession no. 34262; Wang et al., 2018). Indeed, there is significant similarity in flexibility, including identification of the most flexible amino acid residue as Lys116 (Fig. S5).
Graphical comparison from MD simulations of the C-lobe of apo-CaMs showing the hydrogen bond in the apo-CaMWT. (A–D) Top clustered models of the C-lobe of apo-CaMWT (green; A), apo-CaMF90L (red; B), apo-CaMF93L (yellow; C), and apo-CaMF142L (purple; D) are shown. Helix 3 from apo-CaMWT is shown in magenta (A), while helices 3 from Phe → Leu mutant CaMs are shown in dark blue in B–D. The side chain of Asn112 on helix 2 in apo-CaMWT maintains a stable hydrogen bond with the carboxyl oxygen of Ala89 residue on helix 1 during simulations of the apo-CaMWT, which is not observed in the simulations of the CaM mutants.
Graphical comparison from MD simulations of the C-lobe of apo-CaMs showing the hydrogen bond in the apo-CaMWT. (A–D) Top clustered models of the C-lobe of apo-CaMWT (green; A), apo-CaMF90L (red; B), apo-CaMF93L (yellow; C), and apo-CaMF142L (purple; D) are shown. Helix 3 from apo-CaMWT is shown in magenta (A), while helices 3 from Phe → Leu mutant CaMs are shown in dark blue in B–D. The side chain of Asn112 on helix 2 in apo-CaMWT maintains a stable hydrogen bond with the carboxyl oxygen of Ala89 residue on helix 1 during simulations of the apo-CaMWT, which is not observed in the simulations of the CaM mutants.
Comparison of the RMSF of Ca2+-CaMF142L from MD simulations and N-H heteronuclear NOE saturated/unsaturated ratio. Related to BMRB accession no. 34262; Wang et al., 2018. (A and B) There is similar flexibility, including identification of the most flexible amino acid being Lys116 residue. Note: flexibility shown on graphs is inversed, with higher values in RMSF and lower values in heteronuclear NOE denoting higher flexibility.
Comparison of the RMSF of Ca2+-CaMF142L from MD simulations and N-H heteronuclear NOE saturated/unsaturated ratio. Related to BMRB accession no. 34262; Wang et al., 2018. (A and B) There is similar flexibility, including identification of the most flexible amino acid being Lys116 residue. Note: flexibility shown on graphs is inversed, with higher values in RMSF and lower values in heteronuclear NOE denoting higher flexibility.
Discussion
The gating of SK channels relies on Ca2+ binding to CaM. Therefore, mutations of CaM in human calmodulinopathy are predicted to have significant effects on SK channel function. In the current study, we demonstrate a significant reduction in SK currents in the presence of the LQTS CaM mutants CaMD96V and CaMD130G. The marked reduction in SK currents in the presence of LQTS CaM mutants is consistent with their DN effects. Immunofluorescence staining does not show significant changes in membrane-bound SK2 channels in the presence of WT or mutant CaM, suggesting that the effects of these CaM mutants are primarily on the Ca2+-CaM–dependent activation of the channels. The DN effect is predicted to result in a reduction in repolarization reserve. We confirmed these results using tandem SK2-CaM constructs to circumvent potential caveats from endogenous CaM in HEK 293 cells and/or changes in CaM expression from mutant constructs.
In contrast to the marked DN effects of CaMD96V and CaMD130G, CaMF90L results in only a modest reduction in IK,Ca, suggesting a distinct mechanism by disrupting the interaction between apo-CaM and the SK channel without the DN effects. Structural modeling and MD simulations reveals the atomistic mechanisms in conformational changes caused by the phenylalanine-to-leucine mutation, which affect helix packing in the C-lobe, disrupting interactions between apo-CaM with CaMBD of SK channels.
In Fig. 1, we showed that CaMD96V exhibits the most prominent inhibitory effect, while CaMF90L exerts only a mild inhibitory effect on apamin-sensitive current. Consistently, dialysis of mutant CaMD96V proteins in hiPSC-CMs results in the most prominent knockdown of apamin-sensitive currents in hiPSC-CMs compared with CaMF90L (Fig. 4). As expected, the endogenous SK currents in hiPSC-CMs (Fig. 4) are much smaller than the overexpressed SK currents in HEK 293 cells (Fig. 1). Therefore, the two systems were used as complementary studies to evaluate the effects of CaM mutants on SK currents.
There are six CaM alleles, and a mutation in one allele is expected to have only a partial effect on the overall CaM pool within the cell. Therefore, other possible mechanisms may play a role, including cell-specific differential expression of the different alleles.
Mechanisms of human calmodulinopathy
CaM serves as a Ca2+ sensor for many ion channels and transporters and consequently has been shown to play key roles in the regulation of cardiac excitability; mutations in CaM have been linked to LQTS, CPVT, and IVF (Crotti et al., 2013; George, 2015; Reed et al., 2015; Boczek et al., 2016). Studies on calmodulinopathy have provided evidence of the critical roles of CaM in the functional regulation of Cav1.2, RYR2, Kv7.1, and SK channels, among others (Nyegaard et al., 2012; Crotti et al., 2013; Fukuda et al., 2014; Hwang et al., 2014; George, 2015; Vassilakopoulou et al., 2015; Yu et al., 2016).
CaMD96V and CaMD130G occur in highly conserved aspartic acid residues responsible for chelating Ca2+ ions in the C-lobe (Crotti et al., 2013). Consequently, CaMD130G has been shown to have markedly decreased Ca2+-binding affinity (Hwang et al., 2014; Vassilakopoulou et al., 2015). In addition, CaMD130G was reported to have decreased interaction with RYR2, possibly through a DN effect. CaMD96V was previously shown to exhibit an increased Kd for Ca2+ binding compared with CaMWT, although to a lesser extent than CaMD130G (Vassilakopoulou et al., 2015). LQTS mutants (D96V, D130G, and F142L) have been reported to exhibit reduced Ca2+ affinity compared with that of CaMWT and CPVT (N54I and N98S) mutants (Hwang et al., 2014). In addition, the LQTS-associated mutants have been shown to suppress CDI of L-type Ca2+ currents (Limpitikul et al., 2014).
Potential roles of cardiac SK channels in calmodulinopathy
We previously identified the expression of SK channels (SK1, SK2, and SK3) in the heart and demonstrated the critical roles of the channels in mediating action potential repolarization in human atrial myocytes (Xu et al., 2003; Tuteja et al., 2005). SK2 channels physically coupled to α-actinin2, allowing a close association with LTCCs (Lu et al., 2007, 2009). The subcellular localization of SK channels in close proximity to LTCC and RYR2 provides an immediate Ca2+ source (Zhang et al., 2018), which, through binding of Ca2+ to CaM, triggers conformational changes to activate SK channels. SK channels may also play important roles in the feedback mechanism by regulating the activities of LTCCs and RYR2 to influence local and global Ca2+ signaling.
The roles of SK channels in the heart have been extensively studied in the past decade (Zhang et al., 2015). Single nucleotide polymorphisms of KCNN genes encoding for SK channels have been shown to be linked to human atrial fibrillation (AF; Roselli et al., 2018), and SK channels may represent a potential novel therapeutic target against atrial arrhythmias (Diness et al., 2015). Importantly, SK channels participate in the electrical remodeling in AF and heart failure. SK channels are significantly up-regulated in AF animal models (Ozgen et al., 2007; Qi et al., 2014) and failing ventricular myocytes (Chua et al., 2011; Chang et al., 2013). However, the mechanisms underlying SK channel remodeling in the diseased heart remain unclear.
SK channels are unique because the channels are solely gated by Ca2+i through a highly conserved CaMBD. Our data demonstrate the functional effects, structural basis, and molecular mechanisms of the disease-causing CaM mutants in the regulation of cardiac SK2 channels, highlighting the potential contribution of cardiac SK channels to human calmodulinopathy. Even though SK currents are more prominently expressed in atrial than in ventricular myocytes (Xu et al., 2003; Zhang et al., 2015), SK currents are known to be up-regulated in failing ventricular myocytes (Chua et al., 2011; Chang et al., 2013) and during hypokalemia (Chan et al., 2015). Recent studies have supported their critical roles in cardiac Purkinje cells (Reher et al., 2017), which are known to be the potential site of origin of cardiac arrhythmias, including in patients with heritable arrhythmia syndrome (Haïssaguerre et al., 2002; Wilde et al., 2019). Moreover, SK channels play critical roles in the pacemaker activity of sinoatrial and atrioventricular nodes (Zhang et al., 2008; Torrente et al., 2017). Therefore, mutations in CaMs may result in a significant reduction in the key Ca2+-dependent repolarization currents mediated by SK channels. Even though no SK channel variants have been reported to be associated with an inherited arrhythmia syndrome to date, a recent study reported a rare variant, c.1509C>G (p.F503L), in one KCNN2 allele in a patient who developed drug-induced LQTS (Ko et al., 2018). Indeed, this is a rapidly expanding area of research.
Additionally, SK2 channels colocalized within microdomains of LTCC and RYR2, which serve as the Ca2+ source for the activation of SK channels (Zhang et al., 2018). Activation of SK2 channels provides a direct link between beat-to-beat changes in Ca2+i and membrane potentials, and thus may serve as the feedback mechanisms to regulate the activities of the LTCC and RYR2 channels. Functional effects of CaM mutations on SK channel activation may indirectly disrupt this possible feedback mechanism on LTCC and RYR2 channels.
Recent studies have evaluated the effects of arrhythmogenic CaM mutations on SK2 and SK3 channel function in HEK 293 cells (Yu et al., 2016; Saljic et al., 2019). They similarly observed effects independent of SK2 expression and trafficking; however, our study provides new insights by taking advantage of computational analyses as well as native-like cells through our use of hiPSC-CMs.
Advancing the atomistic understanding of CaM and SK2 interaction by structural modeling and MD simulations
Apo-CaM binding to the CaMBD of SK channels requires the C-lobe, while binding of Ca2+ to the N-lobe is involved in SK channel gating (Keen et al., 1999). Several human CaM mutations affect the activation of SK channels (Yu et al., 2016). However, the exact mechanisms among the different CaM mutations remain incompletely understood. Our study shows a significant reduction of cardiac SK currents by CaMD96V and CaMD130G, not only in the hetero-expression system but also in endogenous SK currents in hiPSC-CMs, consistent with DN effects on channel function.
In contrast, phenylalanine mutations in CaM disrupt interaction between apo-CaM and the SK channel without the DN effects. Rosetta molecular modeling is used by taking advantage of the recently published cryo-EM structure of the SK4–apo-CaM complex (Lee and MacKinnon, 2018). We identified the interactions of three key phenylalanine residues (Phe90, Phe93, and Phe142) within the C-lobe of apo-CaM, with hydrophobic amino acids in the CaMBD within the C terminus of the SK2 channel. In addition, previous studies have shown reduced binding affinity to RYR2 by CaMF90L. Circular dichroism experiments in the same study suggest C-lobe destabilization and decreased Ca2+-binding affinity (Nomikos et al., 2014). Our molecular modeling of the CaMF90L mutant shows a significant deviation from CaMWT in the C-lobe structure in the Ca2+-free state (Fig. 6). Taken together with biochemical and functional analyses, the results suggest that the Phe mutations primarily disrupt apo-CaM interactions with the SK2 CaMBD. Indeed, despite significant conformational changes, there were only modest inhibitory effects on apamin-sensitive currents (Fig. 3), consistent with a lack of DN effects; in contrast to CaMD96V and CaMD130G mutants.
Structural modeling and MD simulations provide critical insights into the atomistic mechanisms of CaM mutants in human calmodulinopathy. Our findings suggest that all three Phe mutations in CaM result in similar conformational changes, including an increase in flexibility of the C-terminal end of helix 2, a decrease in flexibility of the C-terminal end of helix 3, and a decrease in the flexibility of the Ca2+-binding site 4 (Fig. 7). The findings support our conclusion that the three Phe → Leu mutations exhibit a similar mechanism in altering the regulation of the hSK2 channel, likely through a reduction in the interaction between apo-CaM and CaMBD in the SK2 channel. These critical insights into the distinct mechanisms whereby these CaM mutations exert their effects on Ca2+-dependent ion channels and cardiac action potential repolarization help to pave the way for understanding the genotype–phenotype correlations of human calmodulinopathy.
Limitations and future directions
RMSD as a metric is circumstantial to derive conclusions on the thermodynamic stability of WT and mutant CaMs. Nonetheless, we have provided additional analyses of RMSF in the current study. Future studies are needed to derive free-energy calculations of the mutations compared with CaMWT. The alteration in flexibility and the possible effects of the mutations on Ca2+-binding site 4 and their cooperativity with other Ca2+-binding sites require further studies as an underlying mechanism affecting CaM mutations. Future studies are needed to test the possible roles of hydrogen bonds between the side chain of Asn112 with the carboxyl oxygen of the Ala89 residue in apo-CaMWT, which is lost in all three Phe → Leu mutations based on MD simulations.
Additionally, studies evaluating the effects of CaM mutants in native cells would be ideal; however, the study of mutant proteins within endogenous cells is not without difficulties. To circumvent these issues, the study of CaM mutants in cardiomyocyte-like cells may yet yield valuable information. Consequently, patient-specific hiPSC-CMs may provide additional insights into the effects of CaM mutants on SK channels and action potential profiles. Finally, future studies using CRISPR/Cas9 gene editing would help to further elucidate the effects of CaM mutations on SK channel function.
Acknowledgments
Jeanne M. Nerbonne served as editor.
This work is funded in part by the American Heart Association Beginning Grant-in-Aid 14BGIA18870087 and National Institutes of Health (NIH) grants R56 HL138392 (to X.-D. Zhang); R01 HL085727, R01 HL085844, and R01 HL137228 (to N. Chiamvimonvat); R01 HL083374 (to A.L. George Jr.); R35 HL144980 (to B.C. Knollmann); and R01 DC015135, R01 DC015252, R01 DC016099, and P01 AG051443 (to E.N. Yamoah); and United States Department of Veterans Affairs Merit Review Grants I01 BX000576 and I01 CX001490 (to N. Chiamvimonvat). H.A. Ledford received a Predoctoral Fellowship from NIH/National Heart, Lung, and Blood Institute (NHLBI) Institutional Training Grant in Basic and Translational Cardiovascular Science (T32 HL086350 and F31 HL136120). R.L. Woltz received a Postdoctoral Fellowships from NIH/NHLBI (F32 HL151130). L. Ren received a Predoctoral Fellowship from the American Heart Association. P. Sirish received a Postdoctoral Fellowship from California Institute for Regenerative Medicine Training Grant to University of California Davis and NIH/NHLBI Institutional Training Grant in Basic and Translational Cardiovascular Science (T32 HL086350), an American Heart Association Career Development Award, and the Harold S. Geneen Charitable Trust Award. N. Chiamvimonvat is the holder of the Roger Tatarian Endowed Professorship in Cardiovascular Medicine and a part-time staff physician at VA Northern California Health Care System, Mather, CA.
The authors declare no competing financial interests.
Author contributions: H.A. Ledford, W. Wang, and X.-D. Zhang performed the electrophysiological experiments and analyzed and interpreted the data. H.A. Ledford, S. Park, L. Ren, and C.-R. Sihn performed the molecular cloning and biochemical experiments. R.L. Woltz, D. Muir, P.T. Nguyen, and V. Yarov-Yarovoy performed the structural modeling and MD simulations. H.A. Ledford performed immunofluorescence imaging. P. Sirish performed stem cell differentiation. H.A. Ledford, X.-D. Zhang, P.T. Nguyen, E.N. Yamoah, V. Yarov-Yarovoy, and N. Chiamvimonvat wrote the manuscript. A.L. George, Jr. and B.C. Knollman provided critical reagents for the study. H.A. Ledford, E.N. Yamoah, X.-D. Zhang, and N. Chiamvimonvat conceived the project, designed the experiments, interpreted the data, and wrote the manuscript.




![Trafficking of hSK2 channels after coexpression with CaMWT or mutant CaMs. HEK 293 cells expressing SK2 fusion protein with HA tag coexpressed with CaMWT or mutant CaMs (N54I, F90L, D96V, N98S, and D130G) using NP and permeabilized conditions. Summary data showing fluorescence intensity under NP (left bars) and permeabilized (right bars) conditions (555 and 633 fluorescent intensities in arbitrary units [AU] × 1,000; n = 15–20 cells; P = NS comparing NP with permeabilized for each group). Data shown represents mean ± SEM. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/12/10.1085_jgp.202012667/3/m_jgp_202012667_figs1.png?Expires=1779227526&Signature=Hx5rrBVnKEGBDjjkkfaAXsHWtr8RBKjtt6SuHGrL6MtSjT7jqNzo~0q9NtevOmAAw1dtqa8YKO~BU5X1k8Pjmj5kzNxdcIYTzTbwZ1zlJ2lEWSR9pALrIGhE9GmtTM8d0~q6DY2ej762l34zGy2mlxmYSR0Td344DfM3JClMVrkEpFFWlq9cki5iyc5a0HijeMNqCCDgbeV9hA2XtFzdRKAKGQVW9mamwgRWv57-SiwCovrHf92mbH80uCEzlyuBf-Tfj1~KlPlm-L9-6iC5FGmWCJYdWR3qAuErw8odjXLJjQLGxkTtsHtDt4fYhUOdjS6nQgy2Itsit6GjTL0xIQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)




![Graphs of RMSD showing stability throughout the production run. RMSD graphs include the equilibration and prosecution stages. The RMSD from the C-lobe of apo-CaMWT (simulations [sims] 1 and 2), apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L are shown in blue, dark brown, red, yellow, and green, respectively. Runs are slightly different in length due to differences in equilibration time. The total simulation time is the same in all runs (1 µs). Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/12/10.1085_jgp.202012667/3/m_jgp_202012667_figs2.png?Expires=1779227526&Signature=qWlYmB1vkx4YbWCehIp2cOl57jevUWG5lXPkOjkBTPpeqa9llW2QuvnRe-euNLZUToFM0xk-jCnYKP9oMGhepiEKNI7CzBJL-XbEOWszHFnCx4LcSn3Bxj~VgSD9EQ4yphD5NgnsIyoVhVq1autdLjCszD7QJMkQehUuEOlvShnylXIIfU1lloUHxGhaBxYgWwrs238NZlwOsg1MV4kWCYAlp79I4CXXXdhnFk0FW~1w65D8ZcL84oHNlKWcu4OyJ-cLrW8aeCV09pwHbhGiUQOrfLVAG6xgTi7iqwLaoFSAcEc5yMbp4gon11xj-~beVoq1YhDq6mbrditUpEatVw__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![Graphs of RMSF showing regional changes in stability. RMSF from MD simulations of the C-lobe of apo-CaMWT is shown in blue (first simulation [sim]) and purple (second simulation), while RMSF from MD simulations of the C-lobe of apo-CaMF90L, apo-CaMF93L, and apo-CaMF142L mutants are shown in red, orange, and green, respectively. Diagrams above the graphs depict helices 1–4 from the C-lobe together with amino acid residues and are scaled to match the residue numbering of the graph. Locations of the three mutations are indicated by their respective colored dot on the graph and bars above the helices. Ca2+-binding sites are indicated with purple dots together with the numbers for the amino acid residues. Data shown are mean ± SD. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/12/10.1085_jgp.202012667/3/m_jgp_202012667_fig7.png?Expires=1779227526&Signature=JsmtEkmUCdT3IOeMiWsVsSBzhYfm8AQGXvVw5XS1VxyJcPAqLTaHA3IqANiR8t7Jd48aMg7bScJIO4Kz2fwt-XTewWY9x6roCAcY1n9TdxOp-NdJFYAAcgYTsCLkTbdDWHRT0ju0ZUAcHzNELxU3n5ISGJLlFBIi7uIMwV8-kowgGV4NFyulu-rZRfRwXYXOq59JOOvbwX24dMVxmKf5T-LlfHMQcm2fEVtu9fDtXCwms1dEUQMJyVwG0TD8yR8IQkTQTXv2i4lxy4ANFLk4aA9GpGBI~ALMINTWqbrJvY3ZsIvSbhDvoV8npS2wvsZKP-eLTIRn-8UaQt5DzM0qeA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)