Exosomes are small vesicles that are secreted from cells to dispose of undegraded materials and mediate intercellular communication. A major source of exosomes is intraluminal vesicles within multivesicular endosomes that undergo exocytic fusion with the plasma membrane. An alternative fate of multivesicular endosomes is fusion with lysosomes, resulting in degradation of the intraluminal vesicles. The factors that determine whether multivesicular endosomes fuse with the plasma membrane or with lysosomes are unknown. In this study, we show that impairment of endolysosomal fusion by disruption of a pathway involving the BLOC-one-related complex (BORC), the small GTPase ARL8, and the tethering factor HOPS increases exosome secretion by preventing the delivery of intraluminal vesicles to lysosomes. These findings demonstrate that endolysosomal fusion is a critical determinant of the amount of exosome secretion and suggest that suppression of the BORC–ARL8–HOPS pathway could be used to boost exosome yields in biotechnology applications.
Introduction
Extracellular vesicles (EVs) are a heterogeneous group of membranous particles containing proteins, lipids, nucleic acids, and carbohydrates that are released from cells into the extracellular space (van Niel et al., 2018). Traditionally, EVs have been thought to enable the exocytic disposal of undegraded materials, thus contributing to the maintenance of cellular homeostasis (Vidal, 2019). Over the past two decades, however, EVs have been increasingly recognized as mediators of intercellular communication, with roles in various physiological and pathological processes (Couch et al., 2021). Furthermore, EVs are being intensely studied for their potential as disease biomarkers and therapeutic agents (Wiklander et al., 2019).
The heterogeneity of EVs arises, at least in part, from their origin in various cellular compartments, including the plasma membrane, and vesicular organelles such as endosomes, lysosomes, autophagosomes, amphisomes (i.e., fusion of autophagosomes with endosomes), and autolysosomes (i.e., fusion of autophagosomes with lysosomes) (Leidal and Debnath, 2021; van Niel et al., 2018; Xu et al., 2018). Cargos delivered to these organelles largely end up degraded by lysosomal hydrolases. However, variable amounts of cargo avoid degradation and are subsequently secreted from the cells as EVs. One of the best-characterized sources of EVs is late endosomal organelles known as multivesicular bodies or multivesicular endosomes (MVEs; the term used here; Gruenberg, 2020). MVEs are intermediaries in the endocytic transport of both soluble and membrane proteins from the plasma membrane to lysosomes (Gruenberg, 2020). After internalization into early endosomes, transmembrane proteins destined for lysosomal degradation undergo ubiquitination and concentration into endosomal membrane domains that invaginate to form intraluminal vesicles (ILVs), thus giving rise to MVEs (Raiborg et al., 2003). This process is mediated by a complex molecular machinery including ubiquitin ligases and “endosomal sorting complexes required for transport” (ESCRT; Henne et al., 2013). Cytosolic proteins and nucleic acids, and even some of the ESCRT proteins themselves, are captured by this process into the lumen of the ILVs. MVEs subsequently fuse with lysosomes, resulting in the degradation of the ILVs and their contents. Alternatively, MVEs can fuse with the plasma membrane, resulting in the release of ILVs as a particular type of small EVs known as “exosomes” (van Niel et al., 2018).
The factors that determine alternative fusion of MVEs with lysosomes or the plasma membrane remain poorly understood (van Niel et al., 2018). We hypothesize that these distinct fates are intimately tied to endolysosomal dynamics, a concept that includes the movement of endolysosomal organelles within the cytoplasm, and their tethering and fusion with other organelles. Unless otherwise specified, throughout this article, we use the term “endolysosome” to denote both late endosomes (including MVEs) and lysosomes because they share many components and use similar machineries for movement and tethering/fusion. A key regulator of endolysosome dynamics is the “BLOC-one-related complex” (BORC), an assembly of eight subunits named BORCS1–8 (Pu et al., 2015; Fig. 1 A). BORC associates with the cytosolic face of endolysosomes, at least in part through a myristoyl group at the N-terminus of the BORCS5 subunit (also known as myrlysin; Pu et al., 2015). BORC then recruits the small GTPase ARL8 (which in humans occurs as two isoforms named ARL8A and ARL8B) from the cytosol to endolysosomes (Bagshaw et al., 2006; Hofmann and Munro, 2006; Pu et al., 2015; Fig. 1 A); BORC does so likely acting as a guanine-nucleotide exchange factor for conversion of ARL8-GDP to ARL8-GTP (Niwa et al., 2017). In turn, ARL8-GTP recruits various effectors, including proteins that mediate endolysosome coupling to microtubule motors and tethering/fusion with other organelles.
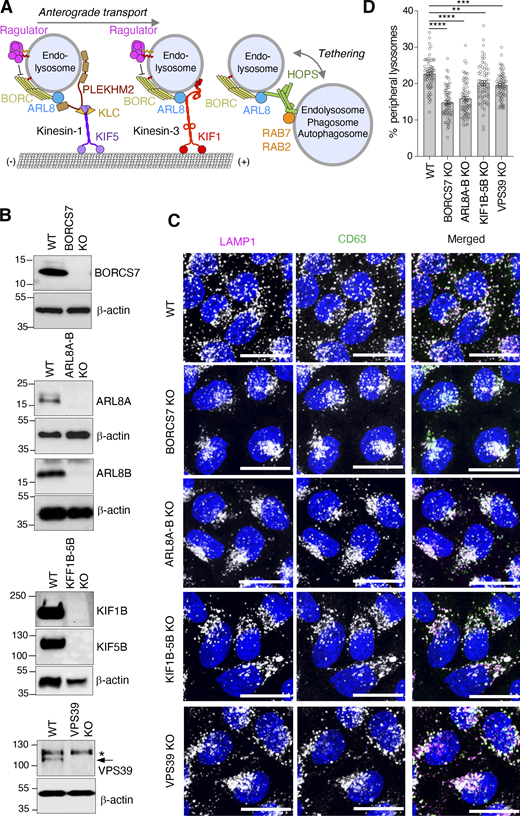
Lysosome distribution in cells with KO of BORCS7, ARL8A and ARL8B, KIF1Bβ and KIF5B, or VPS39. (A) Schematic representation of the roles of BORC and ARL8 in coupling endolysosomes to kinesin-1 (KIF52-KLC2) and kinesin-3 (KIF12) for anterograde transport along microtubules, and to the HOPS complex for tethering and fusion with other organelles. KIF5B and KIF1Bβ are the main isoforms of the kinesin-1 heavy chain and kinesin-3, respectively, expressed in non-neuronal cells. The Ragulator complex is shown to negatively regulate BORC-mediated anterograde lysosome transport, as previously reported (Filipek et al., 2017; Pu et al., 2017). (B) IB analysis of extracts from WT HeLa cells and from cells with KO of the BORCS7 subunit of BORC (BORCS7-KO), both ARL8A and ARL8B (ARL8A-B-KO), both KIF1Bβ and KIF5B (KIF1B-5B-KO), or the VPS39 subunit of HOPS (VPS39-KO). β-actin was used as a loading control. The positions of molecular mass markers (in kD) are indicated on the left. VPS39 is indicated with an arrow; the band marked with an asterisk is non-specific. (C) IF of the indicated WT and KO cell lines for the endogenous endolysosomal markers LAMP1 and CD63 (grayscale in single channels; magenta and green, respectively, in merged images). Nuclei were stained with DAPI. Images are maximum intensity projections of confocal sections. Scale bars: 20 μm. (D) Percentage of LAMP1 present in 1.7-micron outer shell of cell images such as those shown in panel C analyzed by shell analysis (Williamson et al., 2022). Individual data points from around ∼65 cells from three experiments are shown. Bars represent the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ***P < 0.001, ****P < 0.0001. Source data are available for this figure: SourceData F1.
Lysosome distribution in cells with KO of BORCS7, ARL8A and ARL8B, KIF1Bβ and KIF5B, or VPS39. (A) Schematic representation of the roles of BORC and ARL8 in coupling endolysosomes to kinesin-1 (KIF52-KLC2) and kinesin-3 (KIF12) for anterograde transport along microtubules, and to the HOPS complex for tethering and fusion with other organelles. KIF5B and KIF1Bβ are the main isoforms of the kinesin-1 heavy chain and kinesin-3, respectively, expressed in non-neuronal cells. The Ragulator complex is shown to negatively regulate BORC-mediated anterograde lysosome transport, as previously reported (Filipek et al., 2017; Pu et al., 2017). (B) IB analysis of extracts from WT HeLa cells and from cells with KO of the BORCS7 subunit of BORC (BORCS7-KO), both ARL8A and ARL8B (ARL8A-B-KO), both KIF1Bβ and KIF5B (KIF1B-5B-KO), or the VPS39 subunit of HOPS (VPS39-KO). β-actin was used as a loading control. The positions of molecular mass markers (in kD) are indicated on the left. VPS39 is indicated with an arrow; the band marked with an asterisk is non-specific. (C) IF of the indicated WT and KO cell lines for the endogenous endolysosomal markers LAMP1 and CD63 (grayscale in single channels; magenta and green, respectively, in merged images). Nuclei were stained with DAPI. Images are maximum intensity projections of confocal sections. Scale bars: 20 μm. (D) Percentage of LAMP1 present in 1.7-micron outer shell of cell images such as those shown in panel C analyzed by shell analysis (Williamson et al., 2022). Individual data points from around ∼65 cells from three experiments are shown. Bars represent the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ***P < 0.001, ****P < 0.0001. Source data are available for this figure: SourceData F1.
Among the motor-coupling ARL8 effectors is PLEKHM2 (also known as SKIP), which functions as an adaptor for the anterograde motor kinesin-1 (a KIF52-KLC2 heterotetramer) (Dumont et al., 2010; Rosa-Ferreira and Munro, 2011; Fig. 1 A). ARL8 also couples directly to some members of the kinesin-3 family (i.e., the brain-specific KIF1A and ubiquitous KIF1Bβ proteins) (Guardia et al., 2016; Hummel and Hoogenraad, 2021; Niwa et al., 2016; Fig. 1 A). Another ARL8 effector, RUFY3, mediates interaction with the retrograde motor dynein–dynactin (Keren-Kaplan et al., 2022; Kumar et al., 2022). Thus, BORC and ARL8 can promote both anterograde and retrograde transport of endolysosomes, although the effect on anterograde transport is dominant, since silencing of BORC/ARL8 or overexpression of ARL8 causes redistribution of endolysosomes toward the minus or plus ends of microtubules, respectively (Bagshaw et al., 2006; Guardia et al., 2016; Hofmann and Munro, 2006; Marwaha et al., 2017; Matsuo et al., 2004; Rosa-Ferreira et al., 2018).
ARL8 effectors that regulate fusion of lysosomes with other lysosomes (i.e., homotypic fusion) or with late endosomes, phagosomes, and/or autophagosomes (i.e., heterotypic fusion) include the tethering factors PLEKHM1 (Marwaha et al., 2017) and the “homotypic fusion and vacuole protein sorting” (HOPS) complex, composed of VPS11, VPS16, VPS18, VPS33, VPS39, and VPS41 subunits (Boda et al., 2019; Garg et al., 2011; Jia et al., 2017; Khatter et al., 2015; Marwaha et al., 2017; Rosa-Ferreira et al., 2018; Sindhwani et al., 2017; Fig. 1 A). Consequently, silencing of BORC or ARL8 impairs both homotypic and heterotypic fusion of endolysosomal organelles (Boda et al., 2019; Garg et al., 2011; Jia et al., 2017; Khatter et al., 2015; Sasaki et al., 2013; Schleinitz et al., 2023; Sindhwani et al., 2017). Changes in endolysosome motility/positioning and tethering/fusion upon silencing of BORC or ARL8 alters various cellular processes that depend on endolysosome dynamics (Pu et al., 2016).
In this study, we have investigated the influence of BORC, ARL8, and some of its effectors on exosome secretion. Specifically, we asked whether juxtanuclear clustering of endolysosomes by knockout (KO) of BORC, ARL8, or KIF5B and KIF1Bβ, or defective endolysosomal fusion by KO of BORC, ARL8, or HOPS alters exosome secretion. The results of our experiments show that KO of BORC, ARL8, or HOPS increases exosome secretion, and this is due to decreased HOPS-dependent fusion of MVEs with lysosomes. In contrast, double-KO of KIF5B and KIF1Bβ has no effect on the low level of exosome secretion in WT cells but reduces the increased exosome secretion in HOPS-KO cells. These findings demonstrate that disruption of BORC/ARL8/HOPS-dependent endolysosome fusion increases exosome secretion by impairing lysosomal targeting and degradation of ILVs. Disruption of anterograde endolysosome transport mediated by KIF5B and KIF1Bβ, on the other hand, reduces exosome secretion under conditions of increased exosome production (i.e., in HOPS-KO cells).
Results
Juxtanuclear clustering of endolysosomes in cells with KO of BORCS7, ARL8A-B, KIF1Bβ-KIF5B, or VPS39
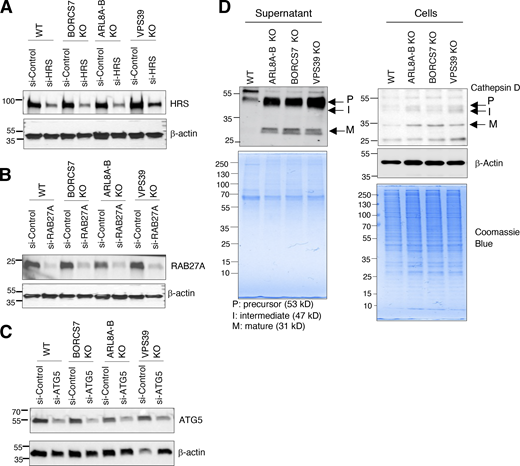
To examine the influence of endolysosomal dynamics on exosome secretion, we used HeLa cells with KO of genes encoding the BORCS7 subunit of BORC (also known as diaskedin or C10orf32), both the ARL8A and ARL8B isoforms of ARL8 (abbreviated ARL8A-B KO), both KIF1Bβ and KIF5B (abbreviated KIF1B-5B KO), or the VPS39 subunit of HOPS (Fig. 1, A–C). Deletion of one subunit of BORC or HOPS results in degradation of the other subunits (Pu et al., 2015; Snouwaert et al., 2018; Wartosch et al., 2015), so herein we refer to these conditions as BORC KO or HOPS KO. Double KO of ARL8A and ARL8B, or KIF1Bβ and KIF5B was needed because these proteins have partly redundant functions in endolysosome dynamics in HeLa cells (Guardia et al., 2016; Keren-Kaplan and Bonifacino, 2021). Immunoblot (IB) analysis confirmed the complete absence of these proteins in the corresponding KO cells (Fig. 1 B). All the KO cells were viable and exhibited similar growth rates relative to the parental WT cells (Fig. S1 A). Immunofluorescence microscopy (IF) of WT HeLa cells for the endogenous endolysosomal membrane proteins LAMP1 and CD63 showed the typical distribution of lysosomes throughout the cytoplasm, but with a higher concentration in the juxtanuclear area of the cells (Fig. 1 C; Jongsma et al., 2016; Korolchuk et al., 2011). In BORCS7-KO, ARL8A-B-KO, or KIF1B-5B-KO HeLa cells, LAMP1/CD63-positive endolysosomes became even more concentrated in the juxtanuclear area, with concomitant depletion from the peripheral cytoplasm (Fig. 1, C and D); this was consistent with the previously demonstrated role of BORC, ARL8, and KIF1Bβ/KIF5B in driving anterograde lysosome transport (Bagshaw et al., 2006; Bentley et al., 2015; Guardia et al., 2016; Hofmann and Munro, 2006; Hummel and Hoogenraad, 2021; Korolchuk et al., 2011; Matsushita et al., 2004; Nakata and Hirokawa, 1995; Niwa et al., 2016; Pu et al., 2015; Tanaka et al., 1998). VPS39-KO cells also exhibited more juxtanuclear LAMP1/CD63-positive vesicles (Fig. 1, C and D), which most likely corresponded to late endosomes/MVEs that failed to fuse with lysosomes (Pols et al., 2013).
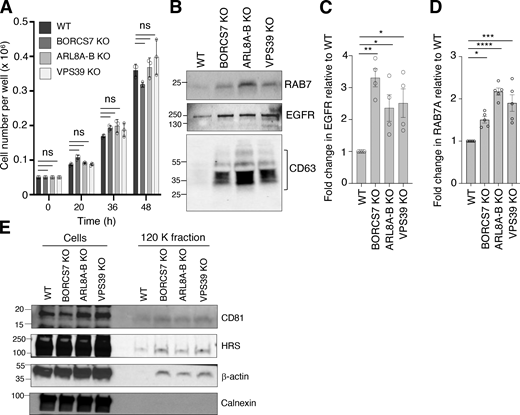
Growth rate of WT and KO cells, and characteristics of exosomes isolated from these cells. (A) Growth rates of WT and KO cells. Equal numbers of WT and KO cells were seeded, and growth rates calculated by harvesting cells and measuring cell number at different times. Individual data points and mean ± SEM from three independent experiments are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. (B) Increased secretion of RAB7A and EGFR in exosomes from BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells relative to WT cells. IB of exosome (120 K) fractions from WT and KO cells for EGFR, RAB7A, and CD63. The positions of molecular mass markers (in kD) are indicated on the left. (C) Quantification of EGFR from experiments such as that shown in panel B. The graph shows individual data points and the mean ± SEM from four independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01. (D) Quantification of RAB7A from experiments such as that shown in panel B. The graph shows individual data points and the mean ± SEM from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; ***P < 0.001; ****P < 0.0001. (E) IB analysis of cell and exosome (120 K) fractions from WT and KO cells for exosome-enriched proteins (CD81, HRS, and β-actin) and the ER protein calnexin. The positions of molecular mass markers (in kD) are indicated on the left. Source data are available for this figure: SourceData FS1.
Growth rate of WT and KO cells, and characteristics of exosomes isolated from these cells. (A) Growth rates of WT and KO cells. Equal numbers of WT and KO cells were seeded, and growth rates calculated by harvesting cells and measuring cell number at different times. Individual data points and mean ± SEM from three independent experiments are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. (B) Increased secretion of RAB7A and EGFR in exosomes from BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells relative to WT cells. IB of exosome (120 K) fractions from WT and KO cells for EGFR, RAB7A, and CD63. The positions of molecular mass markers (in kD) are indicated on the left. (C) Quantification of EGFR from experiments such as that shown in panel B. The graph shows individual data points and the mean ± SEM from four independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01. (D) Quantification of RAB7A from experiments such as that shown in panel B. The graph shows individual data points and the mean ± SEM from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; ***P < 0.001; ****P < 0.0001. (E) IB analysis of cell and exosome (120 K) fractions from WT and KO cells for exosome-enriched proteins (CD81, HRS, and β-actin) and the ER protein calnexin. The positions of molecular mass markers (in kD) are indicated on the left. Source data are available for this figure: SourceData FS1.
Increased exosome secretion from cells with KO of BORCS7, ARL8A-B, or VPS39, but not KIF1Bβ-KIF5B
We took advantage of the above-described cell lines to examine if inhibition of BORC/ARL8-dependent endolysosomal fusion or translocation to the cell periphery alters exosome secretion. To this end, we used a protocol for EV isolation from cell-conditioned medium involving differential centrifugation, filtration, and density gradient ultracentrifugation (Fig. 2 A; Kowal et al., 2016) This protocol yields several fractions that are enriched in apoptotic bodies (denoted as 2 K fraction, where K = 1,000 × g, and “g” represents the rate of change in velocity caused by gravitational force), large EVs (also known as microvesicles or ectosomes; 10 K fraction, for 10,000 × g fraction), and small EVs (also known as exosomes, the term used here; 120 K fraction, for 120,000 × g fraction) (Fig. 2 A). These fractions were analyzed by IB for proteins enriched in different EV subsets (Fig. 2 B). Exosome-enriched proteins included the tetraspanins CD63 and CD81, which localize to both ILVs and the limiting membrane of MVEs (Escola et al., 1998; Fordjour et al., 2022), and the cytoplasmic adaptor protein syntenin-1 (Théry et al., 2001). Ectosomes were detected by IB for the amino-acid transporter CD98 (specifically the SLC3A2 subunit) and the immunoglobulin superfamily member CD147 (also known as BSG), both of which are mostly localized to the plasma membrane (Mathieu et al., 2021).
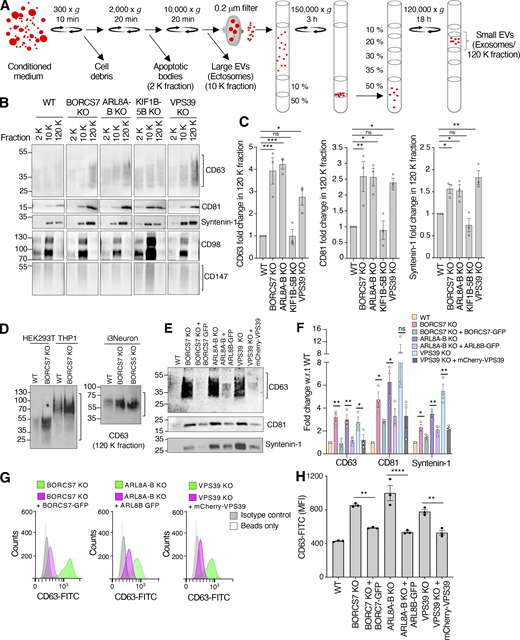
Increased exosome release in cells depleted of BORC, ARL8, or HOPS, but not KIF1Bβ and KIF5B. (A) Schematic representation of the isolation of extracellular particles such as apoptotic bodies (2 K fraction), large EVs/microvesicles/ectosomes (10 K fraction), and small EVs/exosomes (120 K fraction) using differential ultracentrifugation, filtration, and density gradient centrifugation (Kowal et al., 2016). Gradients were made with different percentages of iodixanol-sucrose-buffered solution. In all the experiments presented in this figure, fractions were isolated from 30 ml of medium after 48 h of culture. (B) IB analysis of exosome-enriched proteins (CD63, CD81, and syntenin-1), and ectosome-enriched proteins (CD98 and CD147), in 2, 10, and 120 K fractions from conditioned medium of WT and the indicated KO HeLa cells. The positions of molecular mass markers (in kD) are indicated on the left. CD63 migrates as smear reflective of glycosylation heterogeneity (Engering et al., 2003). (C) Quantification of CD63, CD81, and syntenin-1 fold change in the 120 K (exosome) fraction from three independent experiments such as that shown in panel B. Individual data points are shown with open circles. Bars represent the mean ± SEM of the ratio of CD63, CD81, and syntenin-1 in KO cells to CD63, CD81, and syntenin-1 WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, **P < 0.01, ***P < 0.001. (D) IB analysis of CD63 in the 120 K (exosome) fractions from conditioned medium of WT and BORCS7-KO HEK293T or THP-1 cells, and BORCS7-KO or BORCS5 KO iPSC-derived neurons (i3Neurons). The positions of molecular mass markers (in kD) are indicated on the left. Differences in CD63 mobility are due to varying degrees of glycosylation. (E) IB analysis of CD63 and CD81 in 120 K (exosome) fractions from conditioned medium of WT, BORCS7-KO, BORCS7-KO rescued with BORCS7-GFP, ARL8A-B-KO, ARL8A-B-KO rescued with ARL8B-GFP, VPS39-KO, and VPS39-KO rescued with VPS39-mCherry. The positions of molecular mass markers (in kD) are indicated on the left. (F) Quantification of CD63, CD81, and syntenin-1 fold change in the 120 K (exosome) fraction from three independent experiments such as that shown in panel E. Individual data points are shown with open circles. Bars represent the mean ± SEM of the ratio of CD63, CD81, and syntenin-1 in KO cells to CD63, CD81, and syntenin-1 in WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, **P < 0.01. (G) FACS analysis of CD63 in exosomes isolated from conditioned medium of BORCS7-KO, BORCS7-KO rescued with BORCS7-GFP, ARL8A-B-KO, ARL8A-B-KO rescued with ARL8B-GFP, VPS39-KO, and VPS39-KO rescued with mCherry-VPS39 cells. Analysis involved binding exosomes to magnetic beads coated with antibody to CD63, followed by labeling with FITC-conjugated antibody to CD63 or to isotype-matched immunoglobin G. (H) Quantification of CD63-FITC mean fluorescence intensity (MFI, in arbitrary units) in exosomes from three independent experiments such as that shown in panel F. Individual data points are shown. Bars represent the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ****P < 0.0001. Source data are available for this figure: SourceData F2.
Increased exosome release in cells depleted of BORC, ARL8, or HOPS, but not KIF1Bβ and KIF5B. (A) Schematic representation of the isolation of extracellular particles such as apoptotic bodies (2 K fraction), large EVs/microvesicles/ectosomes (10 K fraction), and small EVs/exosomes (120 K fraction) using differential ultracentrifugation, filtration, and density gradient centrifugation (Kowal et al., 2016). Gradients were made with different percentages of iodixanol-sucrose-buffered solution. In all the experiments presented in this figure, fractions were isolated from 30 ml of medium after 48 h of culture. (B) IB analysis of exosome-enriched proteins (CD63, CD81, and syntenin-1), and ectosome-enriched proteins (CD98 and CD147), in 2, 10, and 120 K fractions from conditioned medium of WT and the indicated KO HeLa cells. The positions of molecular mass markers (in kD) are indicated on the left. CD63 migrates as smear reflective of glycosylation heterogeneity (Engering et al., 2003). (C) Quantification of CD63, CD81, and syntenin-1 fold change in the 120 K (exosome) fraction from three independent experiments such as that shown in panel B. Individual data points are shown with open circles. Bars represent the mean ± SEM of the ratio of CD63, CD81, and syntenin-1 in KO cells to CD63, CD81, and syntenin-1 WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, **P < 0.01, ***P < 0.001. (D) IB analysis of CD63 in the 120 K (exosome) fractions from conditioned medium of WT and BORCS7-KO HEK293T or THP-1 cells, and BORCS7-KO or BORCS5 KO iPSC-derived neurons (i3Neurons). The positions of molecular mass markers (in kD) are indicated on the left. Differences in CD63 mobility are due to varying degrees of glycosylation. (E) IB analysis of CD63 and CD81 in 120 K (exosome) fractions from conditioned medium of WT, BORCS7-KO, BORCS7-KO rescued with BORCS7-GFP, ARL8A-B-KO, ARL8A-B-KO rescued with ARL8B-GFP, VPS39-KO, and VPS39-KO rescued with VPS39-mCherry. The positions of molecular mass markers (in kD) are indicated on the left. (F) Quantification of CD63, CD81, and syntenin-1 fold change in the 120 K (exosome) fraction from three independent experiments such as that shown in panel E. Individual data points are shown with open circles. Bars represent the mean ± SEM of the ratio of CD63, CD81, and syntenin-1 in KO cells to CD63, CD81, and syntenin-1 in WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, **P < 0.01. (G) FACS analysis of CD63 in exosomes isolated from conditioned medium of BORCS7-KO, BORCS7-KO rescued with BORCS7-GFP, ARL8A-B-KO, ARL8A-B-KO rescued with ARL8B-GFP, VPS39-KO, and VPS39-KO rescued with mCherry-VPS39 cells. Analysis involved binding exosomes to magnetic beads coated with antibody to CD63, followed by labeling with FITC-conjugated antibody to CD63 or to isotype-matched immunoglobin G. (H) Quantification of CD63-FITC mean fluorescence intensity (MFI, in arbitrary units) in exosomes from three independent experiments such as that shown in panel F. Individual data points are shown. Bars represent the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ****P < 0.0001. Source data are available for this figure: SourceData F2.
We observed that BORCS7-KO, ARL8A-B-KO, and VPS39-KO cells exhibited increased secretion of CD63, CD81, and syntenin-1 in the exosome fraction relative to WT cells (Fig. 2, B and C). We also observed that BORCS7-KO, ARL8A-B-KO, and VPS39-KO cells secreted higher amounts of the signaling receptor EGFR (epidermal growth factor receptor; Fig. S1, B and C) and the late endosomal small GTPase RAB7A (Fig. S1, B and D), both proteins that had been previously shown to be associated with exosomes (Keerthikumar et al., 2015; Welton et al., 2010; Wu et al., 2021; Yamashita et al., 2013). In contrast, KIF1B-5B-KO cells showed unchanged, low levels of CD63, CD81, and syntenin-1 in the exosome fraction relative to WT cells (Fig. 2, B and C). In WT cells, the exosome fraction was devoid of the ER chaperone calnexin (Fig. S1 E), demonstrating the purity of this preparation, as per published guidelines (Théry et al., 2001). BORCS7-KO, ARL8A-B-KO, and VPS39-KO cells showed no changes in secretion of the ectosome markers CD98 and CD147, but KIF1B-5B-KO cells did secrete more CD98 (Fig. 2 B). Other cell types previously used in studies of EVs, such as human embryonic kidney HEK293T (Shurtleff et al., 2016) and monocytic THP-1 cells (Aharon et al., 2008), as well as human-induce pluripotent stem cell (iPSC)–derived neurons (i3Neurons; Fernandopulle et al., 2018), also showed increased secretion of exosomal CD63 upon KO of BORCS7 or of another BORC subunit, BORCS5 (Fig. 2 D). The increased secretion of exosomal CD63 in BORCS7-KO, ARL8A-B-KO, or VPS39-KO HeLa cells could be reversed by transfection of the cells with plasmids encoding the respective proteins tagged with fluorescent proteins (Fig. 2, E and F). This rescue demonstrated that the effects of the KOs on exosome secretion were specifically caused by the absence of the target proteins. A FACS-based assay using magnetic beads coated with antibody to CD63 (Shelke et al., 2019) also showed increased secretion of CD63-containing exosomes in BORCS7-KO, ARL8A-B-KO, or VPS39-KO HeLa cells, which could be reversed by expression of the corresponding fluorescently tagged proteins (Fig. 2, G and H).
Negative-stain transmission electron microscopy (TEM) showed that exosomes isolated from BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells were on average ∼20–32% smaller than those isolated from WT cells (Fig. S2, A and B). However, nanoparticle tracking analysis (NTA; Gardiner et al., 2013) showed that the KO cells released ∼65–75% more particles than WT cells (Fig. S2 C). The fact that the increases in secreted exosomal CD63 in the KO cells were ∼2.5–4-fold relative to WT cells indicated that the exosomes contained a higher concentration of CD63 per particle.
Characteristics of exosomes isolated from WT and KO cells. (A) Negative-stain electron micrographs of exosomes isolated from the indicated WT and KO cells. Bars: 100 nm. (B) Quantification using Fiji of exosome size from electron micrographs such as those shown in panel A. The graph shows the individual data points and the mean ± SEM from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ****P < 0.0001. (C) NTA of the fold change in particle number in conditioned medium from WT and KO cells obtained after 2,000 and 10,000 × g centrifugations, and 0.2 μm filtration. Individual data points are shown with closed circles. Bars represent the mean ± SEM fold change in KO cells relative to WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01.
Characteristics of exosomes isolated from WT and KO cells. (A) Negative-stain electron micrographs of exosomes isolated from the indicated WT and KO cells. Bars: 100 nm. (B) Quantification using Fiji of exosome size from electron micrographs such as those shown in panel A. The graph shows the individual data points and the mean ± SEM from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ****P < 0.0001. (C) NTA of the fold change in particle number in conditioned medium from WT and KO cells obtained after 2,000 and 10,000 × g centrifugations, and 0.2 μm filtration. Individual data points are shown with closed circles. Bars represent the mean ± SEM fold change in KO cells relative to WT cells (defined as 1) from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01.
Taken together, the above experiments demonstrated that depletion of BORC, ARL8, or HOPS increased secretion of CD63-enriched exosomes, but not ectosomes. Since the BORC–ARL8–HOPS axis is involved in endolysosomal fusion (Boda et al., 2019; Garg et al., 2011; Jia et al., 2017; Khatter et al., 2015; Marwaha et al., 2017; Rosa-Ferreira et al., 2018; Sasaki et al., 2013; Schleinitz et al., 2023; Sindhwani et al., 2017), these findings indicate that endolysosomal fusion attenuates exosome production.
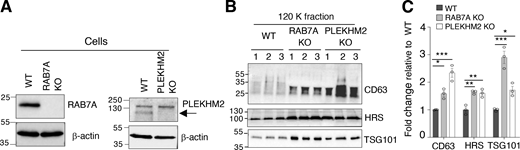
Further evidence for the influence of endolysosomal fusion on exosome secretion was obtained by depleting additional promoters of HOPS-dependent endolysosomal fusion, including the small GTPase RAB7A and the dual ARL8/RAB7A interactor PLEKHM2 (Jongsma et al., 2020; Marwaha et al., 2017). We observed that KO of RAB7A or PLEKHM2 increased the secretion of exosome-associated CD63, HRS, and TSG101 (Fig. S3, A–C), confirming that inhibition of endolysosome fusion enhances exosome secretion.
Effect of RAB7A KO and PLEKHM2 KO on exosome secretion. (A) IB analysis of RAB7A and PLEKHM2 in triplicate samples from WT, RAB7A-KO, and PLEKHM2-KO HeLa cells. Blots were probed with antibodies to the target proteins and to β-actin (loading control). The arrow points to PLEKHM2. (B) IB for exosome-associated proteins in RAB7A-KO and PLEKHM2-KO cell-derived exosomes (120 K fraction) isolated from 20 ml of medium after 24 h. The positions of molecular mass markers (in kD) are indicated on the left. (C) Quantification of CD63, HRS, and TSG101 in exosome fractions from panel B. The graph shows the individual data points and the mean ± SEM from the three samples. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01, ***P < 0.001. Source data are available for this figure: SourceData FS3.
Effect of RAB7A KO and PLEKHM2 KO on exosome secretion. (A) IB analysis of RAB7A and PLEKHM2 in triplicate samples from WT, RAB7A-KO, and PLEKHM2-KO HeLa cells. Blots were probed with antibodies to the target proteins and to β-actin (loading control). The arrow points to PLEKHM2. (B) IB for exosome-associated proteins in RAB7A-KO and PLEKHM2-KO cell-derived exosomes (120 K fraction) isolated from 20 ml of medium after 24 h. The positions of molecular mass markers (in kD) are indicated on the left. (C) Quantification of CD63, HRS, and TSG101 in exosome fractions from panel B. The graph shows the individual data points and the mean ± SEM from the three samples. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01, ***P < 0.001. Source data are available for this figure: SourceData FS3.
Increased exosome secretion is dependent on ILV formation and MVE fusion with the plasma membrane, but not on the autophagy protein ATG5
Exosome biogenesis and release involve ESCRT-dependent ILV formation in MVEs (Colombo et al., 2013; Tamai et al., 2010) and RAB27A-dependent fusion of MVEs with the plasma membrane (Ostrowski et al., 2010), respectively. To examine if these processes were required for the increased secretion of CD63-containing exosomes upon disruption of endolysosomal fusion, we tested the effect of knocking down the ESCRT-0 protein HRS or RAB27A in BORCS7-KO, ARL8A-B-KO, or VPS39-KO HeLa cells (Fig. S4, A and B; and Fig. 3, A and B). We observed that siRNA-mediated knockdown (KD) of HRS or RAB27A reduced the release of CD63-containing exosomes into the medium of the KO cells (Fig. 3, A and B). Components of the autophagy machinery such as ATG5 have also been shown to contribute to formation of at least some types of exosomes through autophagy-independent mechanisms (Guo et al., 2017). In our system, however, KD of ATG5 (Fig. S4 C) did not prevent the increased secretion of CD63-containing exosomes in BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells (Fig. 3, A and B). In addition, the sphingolipid ceramide is known to promote ESCRT-independent ILV formation and release, at least in some settings (Trajkovic et al., 2008). We observed that treatment with the neutral sphingomyelinase-2 inhibitor GW4869 decreased the release of CD63-containing exosomes in BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells (Fig. 3, C and D), indicating that ceramide also plays a role in this process.
KD efficiency and cathepsin D levels in WT and KO cells/supernatants. (A–C) IB analysis of HRS, RAB27A, and ATG5 in WT and KO Hela cells treated with non-targeting siRNA (si-Control) or siRNAs targeting HRS, RAB27A, or ATG5. Blots were probed with antibodies to the target proteins and to β-actin (loading control). The positions of molecular mass markers (in kD) are indicated on the left. (D) Increased levels of cathepsin D in KO cells relative to WT cells. WT and KO cells were incubated for 4 h in serum-free medium. Conditioned supernatant (6 ml) was collected and concentrated to 0.5 ml using a 10-kD spin column. Supernatant and cells were analyzed by SDS-PAGE and IB with antibody to cathepsin D or β-actin (loading control). Coomassie blue staining was done as an additional loading control. The positions of molecular mass markers (in kD) are indicated on the left. Source data are available for this figure: SourceData FS4.
KD efficiency and cathepsin D levels in WT and KO cells/supernatants. (A–C) IB analysis of HRS, RAB27A, and ATG5 in WT and KO Hela cells treated with non-targeting siRNA (si-Control) or siRNAs targeting HRS, RAB27A, or ATG5. Blots were probed with antibodies to the target proteins and to β-actin (loading control). The positions of molecular mass markers (in kD) are indicated on the left. (D) Increased levels of cathepsin D in KO cells relative to WT cells. WT and KO cells were incubated for 4 h in serum-free medium. Conditioned supernatant (6 ml) was collected and concentrated to 0.5 ml using a 10-kD spin column. Supernatant and cells were analyzed by SDS-PAGE and IB with antibody to cathepsin D or β-actin (loading control). Coomassie blue staining was done as an additional loading control. The positions of molecular mass markers (in kD) are indicated on the left. Source data are available for this figure: SourceData FS4.
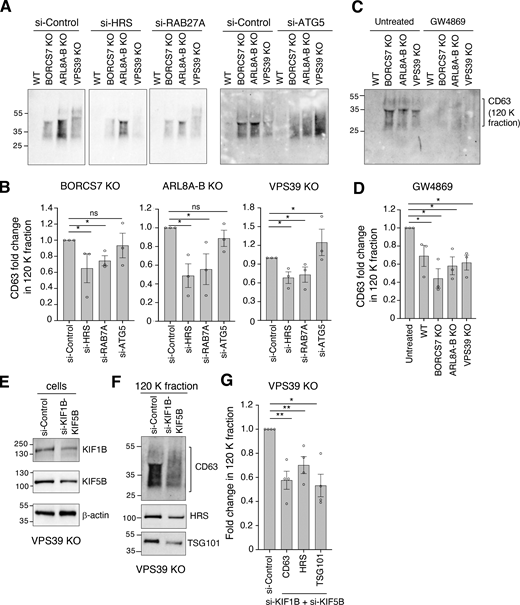
Requirement of HRS, RAB27A, and KIF1Bβ-KIF5B, but not ATG5, for increased exosome release in cells depleted of BORC, ARL8, or HOPS. (A) IB analysis of CD63 in 120 K (exosome) fractions isolated after 24 h of culture in 10 ml of medium from the indicated WT and KO HeLa cells treated with non-targeting siRNA (si-Control) or siRNAs targeting HRS, RAB27A, or ATG5. The positions of molecular mass markers (in kD) are indicated on the left. The efficiency of the knockdowns is shown in Fig. S4, A–C. (B) Quantification of the effect of siRNAs on the release of exosome-associated CD63 from WT and KO cells, calculated from three independent experiments such as those shown in panel A. Individual data points are indicated with open circles. Bars represent the mean ± SEM of the fold change in exosome-associated CD63 in siRNA-treated cells relative to cells treated with siRNA control (defined as 1). Statistical significance was calculated using unpaired t test. *P < 0.05. (C) IB analysis of CD63 in 120 K (exosome) fractions isolated after 16 h of culture in 10 ml of medium from WT and KO cells treated with 10 μM of the neutral sphingomyelinase-2 inhibitor GW4869 for 16 h. The positions of molecular mass markers (in kD) are indicated on the left. (D) Quantification of the effect of GW4869 treatment on the release of exosome-associated CD63, calculated from three independent experiments such as that shown in panel C. Individual data points are indicated with open circles. Bars represent the mean ± SEM of the fold change in exosome-associated CD63 in GW4869-treated cells relative to untreated cells (defined as 1). Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05. (E) IB analysis of KIF1B and KIF5B in VPS39-KO cells treated with non-targeting siRNA (si-Control), or siRNAs targeting both KIF1B and KIF5B, to determine the KD efficiency. The positions of molecular mass markers (in kD) are indicated on the left. (F) IB analysis of CD63, HRS and TGS101 in 120 K (exosome) fractions isolated after 24 h of culture in 10 ml of medium from VPS39-KO cells treated with siRNAs shown in panel E. The positions of molecular mass markers (in kD) are indicated on the left. (G) Quantification of the effect of siRNA to KIF1B and KIF5B on the release of exosome-associated proteins from VPS39-KO cells, calculated from three independent experiments such as those shown in panel F. Bars represent the mean ± SEM of the fold change in exosome-associated markers in VPS39-KO cells treated with siRNAs to KIF1B and KIF5B relative to cells treated with siRNA control (defined as 1). Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01. Source data are available for this figure: SourceData F3.
Requirement of HRS, RAB27A, and KIF1Bβ-KIF5B, but not ATG5, for increased exosome release in cells depleted of BORC, ARL8, or HOPS. (A) IB analysis of CD63 in 120 K (exosome) fractions isolated after 24 h of culture in 10 ml of medium from the indicated WT and KO HeLa cells treated with non-targeting siRNA (si-Control) or siRNAs targeting HRS, RAB27A, or ATG5. The positions of molecular mass markers (in kD) are indicated on the left. The efficiency of the knockdowns is shown in Fig. S4, A–C. (B) Quantification of the effect of siRNAs on the release of exosome-associated CD63 from WT and KO cells, calculated from three independent experiments such as those shown in panel A. Individual data points are indicated with open circles. Bars represent the mean ± SEM of the fold change in exosome-associated CD63 in siRNA-treated cells relative to cells treated with siRNA control (defined as 1). Statistical significance was calculated using unpaired t test. *P < 0.05. (C) IB analysis of CD63 in 120 K (exosome) fractions isolated after 16 h of culture in 10 ml of medium from WT and KO cells treated with 10 μM of the neutral sphingomyelinase-2 inhibitor GW4869 for 16 h. The positions of molecular mass markers (in kD) are indicated on the left. (D) Quantification of the effect of GW4869 treatment on the release of exosome-associated CD63, calculated from three independent experiments such as that shown in panel C. Individual data points are indicated with open circles. Bars represent the mean ± SEM of the fold change in exosome-associated CD63 in GW4869-treated cells relative to untreated cells (defined as 1). Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05. (E) IB analysis of KIF1B and KIF5B in VPS39-KO cells treated with non-targeting siRNA (si-Control), or siRNAs targeting both KIF1B and KIF5B, to determine the KD efficiency. The positions of molecular mass markers (in kD) are indicated on the left. (F) IB analysis of CD63, HRS and TGS101 in 120 K (exosome) fractions isolated after 24 h of culture in 10 ml of medium from VPS39-KO cells treated with siRNAs shown in panel E. The positions of molecular mass markers (in kD) are indicated on the left. (G) Quantification of the effect of siRNA to KIF1B and KIF5B on the release of exosome-associated proteins from VPS39-KO cells, calculated from three independent experiments such as those shown in panel F. Bars represent the mean ± SEM of the fold change in exosome-associated markers in VPS39-KO cells treated with siRNAs to KIF1B and KIF5B relative to cells treated with siRNA control (defined as 1). Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; **P < 0.01. Source data are available for this figure: SourceData F3.
From these experiments, we concluded that CD63-containing exosomes secreted by BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells are generated by both ESCRT-dependent and ceramide-dependent pathways and released into the medium by RAB27A-dependent fusion of MVEs with the plasma membrane.
Double KD of KIF1Bβ and KIF5B decreases exosome secretion in VPS39-KO cells
Although KIF1Bβ and KIF5B also function downstream of BORC and ARL8, double KO of these kinesins had no noticeable effect on exosome release in WT HeLa cells (Fig. 2, B and C). However, we could have missed a possible inhibitory effect of this KO because HeLa cells have low levels of exosome secretion under basal conditions (Fig. 2, B–F; and Fig. 3 A). We thus tested if KD of KIF1Bβ and KIF5B inhibited the increased exosome secretion in VPS39-KO cells (Fig. 3, E–G). Note that, under these conditions, ARL8A and ARL8B are still available to recruit KIF1Bβ and KIF5B to endolysosomes. Indeed, we found that double KD of KIF1Bβ and KIF5B to ∼30% of normal levels (Fig. 3 E) decreased the release of CD63, HRS, and TSG101 by 30–40% in the exosome fraction of VPS39-KO cells (Fig. 3, F and G). These experiments thus demonstrated that KIF1Bβ and KIF5B, and by extension lysosome translocation toward the cell periphery, promote exosome secretion when endolysosomal fusion is inhibited.
Increased frequency of fusion of CD63-containing MVEs with the plasma membrane in cells with KO of BORCS7, ARL8A-B, or VPS39
The increased quantities of exosomes and associated proteins released into the culture supernatant of BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells could be due to an increased number of MVEs fusing with the plasma membrane. To directly assess this possibility, we performed real-time, total internal reflection (TIRF) microscopy of live WT, BORCS7-KO, ARL8A-B-KO, VPS39-KO, or KIF1B-5B-KO cells expressing CD63-pHluorin, a pH-sensitive probe that is non-fluorescent at the acidic pH of MVEs but fluoresces green at the neutral pH of the extracellular medium (Miesenböck et al., 1998; Verweij et al., 2018). This methodology enables visualization of plasma membrane fusion events by imaging a TIRF field of ∼100 nm from the cell substrate (Axelrod, 1981). Imaging for 300 s showed two- to threefold more CD63-pHluorin fusion events per cell in BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells relative to WT cells (Fig. 4, A and B). KIF1B-5B-KO cells showed a trend toward fewer fusion events relative to WT cells, but the differences were not statistically significant (Fig. 4 B). After appearance at the plasma membrane, the CD63-pHluorin signal (shown as Δ intensity in Fig. 4 C) in WT cells decreased exponentially to baseline levels within ∼70 s (shown as Δ time in Fig. 4 C) due to release of CD63-pHluorin–containing exosomes and/or reinternalization of surface-bound CD63-pHluorin. The duration of the fluorescent signal was unaltered in BORCS7-KO, ARL8A-B-KO, VPS39-KO, or KIF1B-5B-KO cells relative to WT cells (Fig. 4 D). These results demonstrated that inhibition of intracellular endolysosomal fusion increased the number MVE–plasma membrane fusion events, though not the kinetics of CD63 disappearance from the plasma membrane.
Increased fusion of CD63-containing vesicles with the plasma membrane in cells depleted of BORC, ARL8, or VPS39. (A) Live-cell TIRF microscopy of WT or BORCS7-KO HeLa cells transfected with a plasmid encoding CD63-pHluorin. Images show cumulative fusion events over different times up to 300 s, with colored arrowheads indicating new fusion events at each time. Scale bars: 10 μm. (B) Quantification of the number of fusion events per cell over 300 s in the indicated WT and KO cells lines. Fusion events were defined by parameters established in a previous study (Verweij et al., 2018). The graph shows individual values and the mean ± SEM from six to 20 cells in two to three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; ***P < 0.001. (C) Representative profile of a single fusion event in WT cells showing the amplitude (Δ intensity, in arbitrary units) and duration (Δ time, in s) of the CD63-pHluorin signal. (D) The duration of fusion events (Δ time) in the indicated WT and KO cells lines was determined by plotting CD63-pHluorin intensity over time determined as in panel C. The graph shows individual values and the mean ± SEM from 20 events from two to three cells in two independent experiments. Statistical significance was tested using one-way ANOVA with Dunnett’s multiple comparisons test. All differences were not significant (ns).
Increased fusion of CD63-containing vesicles with the plasma membrane in cells depleted of BORC, ARL8, or VPS39. (A) Live-cell TIRF microscopy of WT or BORCS7-KO HeLa cells transfected with a plasmid encoding CD63-pHluorin. Images show cumulative fusion events over different times up to 300 s, with colored arrowheads indicating new fusion events at each time. Scale bars: 10 μm. (B) Quantification of the number of fusion events per cell over 300 s in the indicated WT and KO cells lines. Fusion events were defined by parameters established in a previous study (Verweij et al., 2018). The graph shows individual values and the mean ± SEM from six to 20 cells in two to three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05; ***P < 0.001. (C) Representative profile of a single fusion event in WT cells showing the amplitude (Δ intensity, in arbitrary units) and duration (Δ time, in s) of the CD63-pHluorin signal. (D) The duration of fusion events (Δ time) in the indicated WT and KO cells lines was determined by plotting CD63-pHluorin intensity over time determined as in panel C. The graph shows individual values and the mean ± SEM from 20 events from two to three cells in two independent experiments. Statistical significance was tested using one-way ANOVA with Dunnett’s multiple comparisons test. All differences were not significant (ns).
The above results differ from those reported by Verweij and colleagues, who showed that KD of ARL8B (∼75% at the mRNA level) decreases fusion of MVEs with the plasma membrane (Verweij et al., 2022). These differences could be due to the partial KD of ARL8B in the study by Verweij and colleagues, whereas we completely knocked out both ARL8A and ARL8B. Since ARL8A and ARL8B have redundant functions (Guardia et al., 2016; Keren-Kaplan and Bonifacino, 2021), these observations may reflect the requirement of different levels of ARL8 activity for translocation of MVEs to the cell periphery and endolysosomal fusion. Specifically, partial depletion of ARL8 activity could inhibit appearance of CD63-pHluorin at the cell surface but not endolysosomal fusion, whereas complete depletion of ARL8 activity could inhibit both. Inhibition of endolysosomal fusion could dramatically increase the pool of ILVs, overriding any inhibitory effects on translocation to the plasma membrane.
Increased number of ILV-containing MVEs in cells with KO of BORCS7, ARL8A-B, or VPS39
Our next set of experiments was aimed at examining changes in the morphology of MVEs upon inhibition of BORC/ARL8/HOPS-dependent endolysosomal fusion. This was done by allowing cells to internalize HRP for 4 h, followed by fixation, diaminobenzidine reaction, resin embedding, sectioning, and analysis by TEM (Fig. 5 A). In WT cells, we observed the presence of heterogeneous HRP-positive vesicles with the characteristic morphology of endosomes and lysosomes (Fig. 5 A). In BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells, HRP-positive vesicles were on average larger (Fig. 5, A and B) and contained more ILVs (Fig. 5, A and C) relative to those in WT cells. These findings indicated that depletion of BORC, ARL8, or HOPS resulted in accumulation of organelles with the characteristics of MVEs, consistent with inhibition of MVE–lysosome fusion and consequent delivery of cargo to lysosomes, as previously shown for ARL8-depleted cells (Garg et al., 2011; Michelet et al., 2015). FACS analysis also showed increases in total levels of cell-associated CD63 in BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells, but not KIF1B-5B-KO cells, relative to WT cells (Fig. 5, D and E), consistent with a decreased turnover of CD63. These experiments indicated that the increased release of CD63-containing exosomes correlates with a higher number of ILV-containing MVEs upon inhibition of their BORC/ARL8/HOPS-dependent fusion with lysosomes.
Increased number of endolysosomal vesicles in cells depleted of BORC, ARL8, or VPS39. (A) The indicated WT and KO cells were allowed to internalize HRP for 4 h at 37°C to label endolysosomal compartments. After fixation, DAB reaction, and embedding, 60–70-nm ultrathin sections were cut, mounted on EM grids, and imaged by TEM. Low-magnification images (left column; scale bars: 2 μm) and high magnification images of the dashed boxes (right column, scale bars: 1 μm) are shown. Dark structures correspond to endolysosomes. (B) Quantification of the size (area in μm2) of HRP-positive structures in TEM images from 20 WT and 20 each of the KO cells using Fiji software. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett's multiple comparisons test. ***P < 0.001. (C) Quantification of the percentage of HRP-positive structures that contain ILVs analyzed by TEM of 20 WT and 20 each of the KO cells using Fiji software. The graph shows individual values and the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, ****P < 0.0001. (D) FACS analysis of total CD63 levels in 5,000 WT and KO cells that were fixed and permeabilized before incubating them with FITC-conjugated antibody to CD63 (CD63-FITC) and isotype control immunoglobin (IgG-FITC). Profiles represent cell count vs. fluorescence intensity analyzed using FlowJo. (E) Quantification of fold change in total CD63 levels in KO relative to WT cells (defined as 1) in three independent experiments such as that shown in panel D. Mean values from each experiment, and mean ± SEM of the means, are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ****P < 0.0001.
Increased number of endolysosomal vesicles in cells depleted of BORC, ARL8, or VPS39. (A) The indicated WT and KO cells were allowed to internalize HRP for 4 h at 37°C to label endolysosomal compartments. After fixation, DAB reaction, and embedding, 60–70-nm ultrathin sections were cut, mounted on EM grids, and imaged by TEM. Low-magnification images (left column; scale bars: 2 μm) and high magnification images of the dashed boxes (right column, scale bars: 1 μm) are shown. Dark structures correspond to endolysosomes. (B) Quantification of the size (area in μm2) of HRP-positive structures in TEM images from 20 WT and 20 each of the KO cells using Fiji software. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett's multiple comparisons test. ***P < 0.001. (C) Quantification of the percentage of HRP-positive structures that contain ILVs analyzed by TEM of 20 WT and 20 each of the KO cells using Fiji software. The graph shows individual values and the mean ± SEM. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. *P < 0.05, ****P < 0.0001. (D) FACS analysis of total CD63 levels in 5,000 WT and KO cells that were fixed and permeabilized before incubating them with FITC-conjugated antibody to CD63 (CD63-FITC) and isotype control immunoglobin (IgG-FITC). Profiles represent cell count vs. fluorescence intensity analyzed using FlowJo. (E) Quantification of fold change in total CD63 levels in KO relative to WT cells (defined as 1) in three independent experiments such as that shown in panel D. Mean values from each experiment, and mean ± SEM of the means, are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ****P < 0.0001.
Decreased delivery of endocytic cargo to lysosomes in cells with KO of BORCS7, ARL8A-B, or VPS39
We have hitherto assumed that KO of BORCS7, ARL8A-B, or VPS39 impairs delivery of endocytic cargo to lysosomes, a fact that was previously demonstrated for silencing of ARL8 (Garg et al., 2011; Khatter et al., 2015; Michelet et al., 2015; Nakae et al., 2010) and HOPS (Galmes et al., 2015; Khatter et al., 2015; Pols et al., 2013; Schleinitz et al., 2023; Wartosch et al., 2015), but not BORC. Therefore, we used several fluorescence microscopy assays to monitor endocytic traffic in WT, BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells (Fig. 6). First, we examined the fate of internalized Alexa Fluor 488–conjugated EGF (EGF-AF488), which is taken up by receptor-mediated endocytosis for eventual degradation in lysosomes (Futter et al., 1996; Khatter et al., 2015). We observed that cells with KO of BORCS7, ARL8A-B, or VPS39 accumulated more EGF-AF488 than WT cells after 2 h of continuous uptake (Fig. 6, A and B). Next, we analyzed the endocytic transport of the fluid-phase probe DQ-Green-BSA, a quenched boron-dipyrromethene–labeled protein that becomes fluorescent upon degradation by lysosomal cathepsins (Bright et al., 2016; Marwaha et al., 2017). We observed that KO of BORCS7, ARL8A-B, or VPS39 inhibited the unquenching of DQ-Green-BSA relative to that in WT cells after 2 h of continuous uptake (Fig. 6, C and D). The inhibition of DQ-Green-BSA and EGF-AF488 degradation was likely due to impaired transport to lysosomes and not depletion of lysosomal proteases, as all the KO cells contained even higher levels of the mature lysosomal protease cathepsin D than the parental WT cells (Fig. S4 D), as previously reported (Anderson et al., 2022). Finally, we performed live-cell imaging to assess the endocytic delivery of another fluid-phase cargo, the pH-insensitive and non-degradable Alexa Fluor 488–conjugated dextran (dextran-AF488), to lysosomes labeled with the fluorogenic Magic Red cathepsin B substrate after 2 h of uptake and 1 h of chase (Bright et al., 2016). We observed that internalized dextran-AF488 was largely localized to Magic Red–positive lysosomes in WT cells but to Magic Red–negative vesicles in BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells (Fig. 6, E and F). Taken together, these experiments demonstrated that KO of BORCS7, ARL8A-B, or VPS39 impaired the delivery of various endocytic cargos to lysosomes, consistent with a block in MVE–lysosome fusion.
Defective endolysosomal fusion in cells depleted of BORC, ARL8, or VPS39. (A) Live WT and KO cells grown on coverslips were incubated with EGF-AF488 for 2 h at 37°C. Cells were subsequently washed, fixed, stained with DAPI (for nuclei), mounted on coverslips, and imaged by confocal fluorescence microscopy. Images are maximum intensity projections. EGF-AF488 is shown in grayscale and DAPI in blue. Scale bar: 20 μm. (B) Quantification using Fiji of raw integrated density of EGF-AF488 from 30 cells in three independent experiments such as that shown in panel A. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett's multiple comparisons test. ***P < 0.001. (C) Live WT and KO cells grown on coverslips were incubated with DQ-Green-BSA for 2 h at 37°C. Cells were subsequently washed, fixed, stained with DAPI (for nuclei), mounted on coverslips, and imaged by confocal fluorescence microscopy. Images are maximum intensity projections. DQ-Green-BSA is shown in grayscale and DAPI in blue. Scale bar: 20 μm. (D) Quantification using Fiji of raw integrated density of DQ-Green-BSA from 30 cells in three independent experiments such as that shown in panel A. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ****P < 0.0001. (E) Live WT and KO cells grown in chambered slides were incubated with dextran-AF488 for 2 h, chased for 1 h, and stained with Magic Red cathepsin B dye for 15 min, all at 37°C. Cells were imaged live by confocal fluorescence microscopy. Single-channel images are shown in grayscale. In merged images, dextran-AF488 is shown in green and Magic Red in magenta. White arrowheads indicate vesicles where both fluorescent signals co-localize. Scale bar: 10 μm. (F) The Pearson correlation coefficient of dextran-AF488 and Magic Red signals was determined using Fiji. The bar graph shows the individual data points and the mean ± SEM of points from two independent experiments, each quantifying >60 cells per condition. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ***P < 0.001.
Defective endolysosomal fusion in cells depleted of BORC, ARL8, or VPS39. (A) Live WT and KO cells grown on coverslips were incubated with EGF-AF488 for 2 h at 37°C. Cells were subsequently washed, fixed, stained with DAPI (for nuclei), mounted on coverslips, and imaged by confocal fluorescence microscopy. Images are maximum intensity projections. EGF-AF488 is shown in grayscale and DAPI in blue. Scale bar: 20 μm. (B) Quantification using Fiji of raw integrated density of EGF-AF488 from 30 cells in three independent experiments such as that shown in panel A. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett's multiple comparisons test. ***P < 0.001. (C) Live WT and KO cells grown on coverslips were incubated with DQ-Green-BSA for 2 h at 37°C. Cells were subsequently washed, fixed, stained with DAPI (for nuclei), mounted on coverslips, and imaged by confocal fluorescence microscopy. Images are maximum intensity projections. DQ-Green-BSA is shown in grayscale and DAPI in blue. Scale bar: 20 μm. (D) Quantification using Fiji of raw integrated density of DQ-Green-BSA from 30 cells in three independent experiments such as that shown in panel A. Individual values and mean ± SEM are shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ****P < 0.0001. (E) Live WT and KO cells grown in chambered slides were incubated with dextran-AF488 for 2 h, chased for 1 h, and stained with Magic Red cathepsin B dye for 15 min, all at 37°C. Cells were imaged live by confocal fluorescence microscopy. Single-channel images are shown in grayscale. In merged images, dextran-AF488 is shown in green and Magic Red in magenta. White arrowheads indicate vesicles where both fluorescent signals co-localize. Scale bar: 10 μm. (F) The Pearson correlation coefficient of dextran-AF488 and Magic Red signals was determined using Fiji. The bar graph shows the individual data points and the mean ± SEM of points from two independent experiments, each quantifying >60 cells per condition. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ***P < 0.001.
Increased regurgitation of fluid-phase cargo in cells with KO of BORCS7, ARL8A-B, or VPS39
To examine the fate of internalized fluid-phase cargo when endolysosomal fusion is impaired, we used a FACS-based dextran secretion assay shown in Fig. 7 A. The assay consisted of incubating live WT, BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells for 4 h with dextran-AF488, followed by a 20-h chase without dextran-AF488 (to allow the release of internalized dextran-AF488) and 2-h incubation with dextran-AF647 (to reload the endocytic pathway with a fluid-phase marker), all at 37°C, followed by analysis of AF488 vs. AF647 fluorescence intensity by FACS. These analyses showed a reduction of the AF488/AF647 ratio to 58% in BORCS7-KO, 44% in ARL8A-B-KO, and 52% in VPS39-KO cells relative to WT cells (Fig. 7, B and C). This was not due to differences in endocytosis as the levels of internalized dextran-AF647 between 5 and 60 min of uptake increased at similar rates in WT, BORCS7-KO, ARL8A-B-KO, or VPS39-KO cells (Fig. 7 D). Kinetic analysis of dextran-AF647 decrease after 4 h of uptake showed that all the KO cells exhibited higher rates in comparison with WT cells after a lag of 30 min and up to 20 h of chase (Fig. 7 E). These experiments thus demonstrated that KO of BORCS7, ARL8A-B, or VPS39 reduces accumulation of a fluid-phase cargo through increased regurgitation, consistent with the impaired fusion of MVEs with lysosomes in the KO cells.
Increased regurgitation of fluid-phase cargo in cells depleted of BORC, ARL8, or VPS39. (A) Experimental outline of dextran endocytosis-exocytosis (regurgitation) assay. Live WT and KO cells were incubated with dextran-AF488 for 4 h and chased for 18 h, all at 37°C in regular culture medium. Cells were next incubated with dextran-AF647 for 2 h under the same conditions. Cells were finally washed and analyzed by FACS for dextran-AF488 and dextran-AF647 fluorescence intensity. (B) FACS dot plots representing dextran-AF488 vs. dextran-AF647 fluorescence intensity in 5,000 WT and KO cells. Notice the downward shift of dextran-AF488 fluorescence but similar levels of dextran-AF647 in KO cells relative to WT cells, reflecting enhanced exocytosis of internalized dextran-AF488 but similar endocytosis of dextran-AF647 in the KO cells. (C) Quantification of the dextran-AF488 to dextran-AF647 ratio of MFI of WT and KO cells from experiments such as those described in panels A and B. Individual values and mean ± SEM from three independent experiments is shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ***P < 0.001. (D) Live WT and KO cells were incubated for the indicated times with dextran-A647, washed, and analyzed by FACS. Graphs show individual data points and mean ± SEM of dextran-A647 MFI from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ***P < 0.001. (E) Live WT and KO cells were incubated with dextran-AF647 for 4 h, chased for different times at 37°C, and analyzed by FACS. Values are the mean ± SEM of the fold change in cell-associated dextran-AF647 MFI at each time point relative to time 0. Statistical significance at each time point was calculated using two-way ANOVA with Tukey’s multiple comparisons test. ****P < 0.0001.
Increased regurgitation of fluid-phase cargo in cells depleted of BORC, ARL8, or VPS39. (A) Experimental outline of dextran endocytosis-exocytosis (regurgitation) assay. Live WT and KO cells were incubated with dextran-AF488 for 4 h and chased for 18 h, all at 37°C in regular culture medium. Cells were next incubated with dextran-AF647 for 2 h under the same conditions. Cells were finally washed and analyzed by FACS for dextran-AF488 and dextran-AF647 fluorescence intensity. (B) FACS dot plots representing dextran-AF488 vs. dextran-AF647 fluorescence intensity in 5,000 WT and KO cells. Notice the downward shift of dextran-AF488 fluorescence but similar levels of dextran-AF647 in KO cells relative to WT cells, reflecting enhanced exocytosis of internalized dextran-AF488 but similar endocytosis of dextran-AF647 in the KO cells. (C) Quantification of the dextran-AF488 to dextran-AF647 ratio of MFI of WT and KO cells from experiments such as those described in panels A and B. Individual values and mean ± SEM from three independent experiments is shown. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. **P < 0.01, ***P < 0.001. (D) Live WT and KO cells were incubated for the indicated times with dextran-A647, washed, and analyzed by FACS. Graphs show individual data points and mean ± SEM of dextran-A647 MFI from three independent experiments. Statistical significance was calculated using one-way ANOVA with Dunnett’s multiple comparisons test. ***P < 0.001. (E) Live WT and KO cells were incubated with dextran-AF647 for 4 h, chased for different times at 37°C, and analyzed by FACS. Values are the mean ± SEM of the fold change in cell-associated dextran-AF647 MFI at each time point relative to time 0. Statistical significance at each time point was calculated using two-way ANOVA with Tukey’s multiple comparisons test. ****P < 0.0001.
Discussion
The results of this study show that inhibition of endolysosomal fusion by KO of BORC, ARL8, or HOPS increases exosome secretion. This effect is likely due to loss of HOPS-mediated fusion of MVEs with lysosomes and consequent redirection of ILVs for secretion. Although KIF5B and KIF1Bβ also function downstream of BORC and ARL8 to promote endolysosome transport to the cell periphery, KO of both kinesins has no detectable impact on the low level of basal exosome secretion in HeLa cells. However, double KO of KIF5B and KIF1Bβ does reduce the increased exosome secretion in HOPS-deficient cells. From these observations, we conclude that BORC and ARL8 influence exosome secretion, negatively through their role in promoting HOPS-dependent endolysosomal fusion and positively through their role in driving kinesin-dependent endolysosomal transport to the cell periphery. Under basal conditions, the role of BORC and ARL8 in promoting endolysosomal fusion prevails and is thus a dominant factor in keeping the level of exosome secretion low. Previous studies showed that RAB7A and its interactors PLEKHM2 also participate in HOPS-dependent endolysosomal fusion (Jongsma et al., 2020). Indeed, we found that depletion of RAB7A or PLEKHM2 also enhances exosome secretion, further supporting the notion that endolysosomal fusion contributes to maintaining a low level of exosome secretion.
KO of BORC, ARL8, or HOPS increases not only the secretion of CD63-positive exosomes but also the levels of cell-associated CD63 and the percentage of endosomal structures containing ILVs. Therefore, the effect of the KOs is to increase the availability of ILVs, which are subsequently released as exosomes. It is worth noting that the efficiency of exosome release could have been even higher were it not for two counteracting processes: retrofusion of ILVs with the limiting membrane of MVEs (Perrin et al., 2021) and tethering of exosomes to the plasma membrane by BST-2/tetherin (Edgar et al., 2016). At present, we do not know if KO of BORC, ARL8, or HOPS affects these processes. It also remains to be determined if KO of BORC, ARL8, or HOPS enhances ILV and MVE biogenesis in addition to impairing their degradation.
Fig. 8 shows our interpretation of the data. In WT cells, BORC and ARL8 promote the recruitment of HOPS to lysosomes and/or MVEs. This enables tethering and eventual SNARE-dependent fusion of lysosomes with MVEs to form a hybrid organelle (Bright et al., 2016; Mullock et al., 1998). Within this hybrid, the contents of MVEs, including ILVs, are degraded, precluding their release as exosomes. Only a small fraction of MVEs and ILVs escape this fate, leading to the release of low amounts of exosomes into the medium (Fig. 8 A). In contrast, in BORC-KO, ARL8-KO, or HOPS-KO cells, lysosomes and MVEs fail to fuse, resulting in more MVEs fusing with the plasma membrane and releasing exosomes into the medium (Fig. 8 B). The enhanced release of exosomes in HOPS-KO cells is partially inhibited by KD of KIF5B and KIF1Bβ, indicating that translocation toward the cell periphery does promote exosome release under these conditions (Fig. 8 C). This activity of KIF5B-KIF1Bβ is undetectable in WT HeLa cells, which release few exosomes, but becomes noticeable in HOPS-KO cells, where exosome release is increased. The fact that exosomes are still released in BORC-KO and ARL8-KO cells despite the uncoupling of KIF5B and KIF1Bβ from endolysosomes could be explained by the juxtanuclear MVEs fusing with top and bottom regions of the plasma membrane adjacent to the nucleus, or by additional kinesins contributing to exosome translocation toward the cell periphery.
Model for the impact of lysosome–MVE fusion and kinesin-dependent transport on exosome biogenesis. (A) In WT cells, BORC, ARL8, and HOPS promote tethering of lysosomes to MVEs for eventual fusion into a lysosome–MVE hybrid organelle. The contents of MVEs, including ILVs, are then degraded by lysosomal hydrolases. A small fraction of MVEs that do not fuse with lysosomes undergo exocytosis and release of ILVs into the extracellular space as exosomes. The scheme depicts HOPS being anchored to the lysosomal and MVE membranes by ARL8 and RAB7, but the details of such anchoring, and the possible involvement of other small GTPases, have not been definitively established. (B) In BORC-KO, ARL8-KO, or HOPS-KO cells, lysosomes cannot tether and fuse with MVEs, leading to the accumulation of MVEs containing a larger number of ILVs. These MVEs undergo exocytosis, resulting in the release of more exosomes into the extracellular space. (C) Double KD of KIF1B and KIF5B in HOPS-KO cells partially impairs MVE exocytosis, diminishing the increased release of exosomes caused by HOPS KO.
Model for the impact of lysosome–MVE fusion and kinesin-dependent transport on exosome biogenesis. (A) In WT cells, BORC, ARL8, and HOPS promote tethering of lysosomes to MVEs for eventual fusion into a lysosome–MVE hybrid organelle. The contents of MVEs, including ILVs, are then degraded by lysosomal hydrolases. A small fraction of MVEs that do not fuse with lysosomes undergo exocytosis and release of ILVs into the extracellular space as exosomes. The scheme depicts HOPS being anchored to the lysosomal and MVE membranes by ARL8 and RAB7, but the details of such anchoring, and the possible involvement of other small GTPases, have not been definitively established. (B) In BORC-KO, ARL8-KO, or HOPS-KO cells, lysosomes cannot tether and fuse with MVEs, leading to the accumulation of MVEs containing a larger number of ILVs. These MVEs undergo exocytosis, resulting in the release of more exosomes into the extracellular space. (C) Double KD of KIF1B and KIF5B in HOPS-KO cells partially impairs MVE exocytosis, diminishing the increased release of exosomes caused by HOPS KO.
Our findings demonstrate that lysosome–MVE fusion limits the extent of exosome secretion, a concept that had been proposed but not yet experimentally addressed. This conclusion is in line with previous studies showing increased exosome production upon incubation with the lysosome-acidification inhibitors bafilomycin A1, chloroquine, or ammonium chloride, which prevent substrate degradation by lysosomal acid hydrolases (Alvarez-Erviti et al., 2011; Ortega et al., 2019; Yuyama et al., 2008). A more recent study, however, has challenged the notion that inhibition of lysosome acidification increases exosome secretion by demonstrating that bafilomycin A1 inhibits this process independently of changes in lysosomal pH and that chloroquine has no effect on exosome secretion (Cashikar and Hanson, 2019). These differences in sensitivity to lysosomal inhibitors could be explained by the different experimental systems used in these studies. Our findings are also consistent with the observation of increased exosome release upon disruption of lysosomal degradation by inhibition of phosphatidylinositol-3-phosphate synthesis (Miranda et al., 2018), an effect that may stem from the role of phosphatidylinositol-3-phosphate in recruiting HOPS to endolysosomes (Jeschke and Haas, 2018; Jiang et al., 2022). Finally, centrosome amplification was shown to promote exosome secretion because of lysosome disruption caused by increased reactive oxygen species (Adams et al., 2021). The key role of the BORC–ARL8–HOPS axis in endolysosomal fusion makes it a possible target for the regulation of exosome secretion under physiological and pathological conditions. To date, BORC and ARL8 are known to be subject to negative regulation by the Ragulator complex (Fig. 1 A) in response to nutrients or EGF (Filipek et al., 2017; Pu et al., 2017; Yordanov et al., 2019). In addition, the SARS-CoV-2 ORF3a protein was recently shown to target BORC–ARL8, in this case, to promote viral egress (Chen et al., 2021). It remains to be determined whether these or other upstream factors regulate the influence of BORC–ARL8–HOPS on exosome production.
We have shown that exosomes secreted by BORC-KO, ARL8-KO, or HOPS-KO cells are derived from ILVs produced by ESCRT-dependent and ceramide-dependent pathways, and released in a RAB27A-dependent manner. Accordingly, they contain typical exosome cargos such as CD63, CD81, HRS, and syntenin-1. In contrast, the secretion of plasma membrane–derived ectosomes, characterized by the presence of CD98 and CD147, is unchanged in BORC-KO, ARL8-KO, or HOPS-KO cells, indicating that the production of this type of EV is independent of lysosome–MVE fusion. Since BORC, ARL8, and HOPS also participate in fusion of lysosomes with autophagosomes (Boda et al., 2019; Jia et al., 2017; Jiang et al., 2014; Marwaha et al., 2017; Takáts et al., 2014), it is possible that silencing of these factors also enhances the release of autophagic cargos by secretory autophagy (Dupont et al., 2011; Zhang et al., 2015) or by alternative pathways such as LC3-dependent loading and secretion (Leidal et al., 2020) or secretory autophagy during lysosome inhibition (Solvik et al., 2022), which to some extent involves late endosomal intermediates. Further work will be required to determine whether the BORC–ARL8–HOPS ensemble regulates these alternative pathways of EV biogenesis.
Our findings also support the notion that BORC and ARL8 participate in fusion of late endosomes with lysosomes in general. This was known for ARL8 (Garg et al., 2011; Khatter et al., 2015; Marwaha et al., 2017; Michelet et al., 2015; Nakae et al., 2010) but not for BORC. In this study, we show that silencing of BORC, as well as ARL8 or HOPS, impairs the delivery of both membrane-bound (i.e., EGF) and soluble (i.e., BSA and dextran) cargos from late endosomes to lysosomes. Although other small GTPases, such as RAB2 (Fujita et al., 2017; Gillingham et al., 2014; Kajiho et al., 2016; Lőrincz et al., 2017; Schleinitz et al., 2023) and RAB7 (or its yeast ortholog Ypt7) (Cabrera et al., 2009; Lürick et al., 2017; Jongsma et al., 2020; Xing et al., 2021), bind HOPS and/or mediate its recruitment to late endosomes, autophagosomes, or autolysosomes, our observations are consistent with BORC and ARL8 playing an essential role in HOPS-dependent endolysosomal fusion events in mammalian cells. This conclusion applies to fusion of lysosomes with not only late endosomes carrying endocytic cargos but also TGN-derived carriers carrying biosynthetic cargos such as the cholesterol transport protein NPC2 (Anderson et al., 2022).
In addition to increased exosome secretion, the block in endolysosomal fusion imposed by BORC, ARL8, or HOPS depletion results in increased recycling of internalized fluid-phase cargos such as dextran. This process, referred to as “regurgitation” (Cupers et al., 1994), also reflects fusion of late endosomes with the plasma membrane instead of lysosomes. Like exosome release, endocytic regurgitation is dependent on RAB27A (Le Roux et al., 2012), suggesting that it could be used as a surrogate for further studies of the role of late endosome–lysosome fusion in exosome production.
A current limitation in the use of exosomes for therapeutic applications is the low yields obtained from cells under normal culture conditions, as exemplified by the WT HeLa cells used in our study. Several strategies have been employed to increase these yields, including overexpression of CD9 (Böker et al., 2018), treatment with lysosomotropic agents (Ortega et al., 2019) or other bioactive small molecules (Wang et al., 2020), and nutrient deprivation (Garcia et al., 2015; Haraszti et al., 2019). Our findings in HeLa, HEK293T, and THP1 cells, as well as iPSC-derived neurons, suggest a convenient alternative of using cells with KO of BORC, ARL8, or HOPS as sources of exosomes in biotechnology applications.
Materials and methods
Cell lines
HeLa, HEK293T, and THP-1 cell lines were obtained from the American Type Culture Collection. HeLa BORCS7-KO (Guardia et al., 2016), ARL8A-B-KO (Keren-Kaplan and Bonifacino, 2021), VPS39-KO (Anderson et al., 2022), and KIF5B-1B-KO (Jia et al., 2017) cell lines were previously described. RAB7A-KO HeLa cells were a gift from Morié Ishida (National Institute of Child Health and Human Development [NICHD], National Institutes of Health [NIH], Bethesda, MD, USA). PLEKHM2-KO HeLa cells were a gift from Tal Keren-Kaplan (NICHD, NIH, Bethesda, MD, USA). HeLa and HEK293T cells were cultured in DMEM with glutamine (#25-005-CI; Corning), supplemented with 10% FBS (#35-011-CV Corning), penicillin (100 IU/ml), and streptomycin (100 μg/ml; #30-002-CI; Corning). THP-1 cells were cultured in RPMI-1640 medium (#61870036; Gibco; Thermo Fisher Scientific) supplemented with 10% FBS and 0.05 mM 2-mercaptoethanol (#M3148; Millipore Sigma). FBS used in these cultures was depleted of EVs by centrifugation at 120,000 × g for 16 h (Shelke et al., 2019). Cells were cultured at 37°C in an incubator with a humidified atmosphere containing 5% CO2.
Differentiation of human iPSCs expressing doxycycline-inducible neurogenin 2 to neurons (referred to as i3Neurons) was performed as described (Fernandopulle et al., 2018). iPSCs were cultured in induction medium containing DMEM/F12 (#11330032; Gibco; Thermo Fisher Scientific) with 1 × non-essential amino acids (#11140050; Gibco; Thermo Fisher scientific), 1 × GlutaMAX (#35050061; Gibco; Thermo Fisher Scientific), 1 × N2A supplement (#17502048; Gibco; Thermo Fisher Scientific), 2 μg/ml doxycycline (#D9891; Millipore Sigma), and 10 μM Y-27632 dihydrochloride ROCK inhibitor (#S1049; Selleckchem). Upon differentiation, the medium was replaced with neuronal culture medium consisting of BrainPhys medium (#5790; Stem Cell Technology) supplemented with 10 ng/ml neurotrophin-3 (#450-03; PeproTech; Thermo Fisher Scientific), 10 ng/ml brain-derived neurotrophic factor (#450-02; PeproTech; Thermo Fisher Scientific), 1 × B-27 Supplement serum free (#17504044; Gibco; Thermo Fisher Scientific), 2 μg/ml doxycycline (#D9891; Millipore Sigma), and 1 μg/ml mouse laminin (#23017015; Gibco; Thermo Fisher Scientific).
Generation of CRISPR-Cas9 KO cell lines
Gene KOs in HEK293T, THP-1, and iPSC cells were made using the CRISPR-Cas9 system. For HEK293T and THP-1 KOs, single-guide RNAs (sgRNA) targeting BORCS7 (Table S1) were cloned separately into pSpCas9(BB)-2A-GFP (pX458). For iPSCs KOs, sgRNAs targeting BORCS5 or BORCS7 (Table S1) were cloned separately into pSpCas9(BB)-2A-GFP (pX458) plasmid. Briefly, sgRNA was designed using online software from Benchling (https://benchling.com/). Forward and reverse oligos for sgRNA were custom synthesized (Eurofins Scientific) and annealed to generate sticky ends. Plasmid (pX458) was cleaved with enzyme BbsI (#R3539; New England Biolabs) and ligated with annealed-sgRNA using a DNA ligation kit (#6022; Takara). Ligated plasmid was transformed in Escherichia coli (DH5-α), selected on LB-Agar ampicillin plate (#BPL-2200; KD Medicals), amplified, and isolated by using plasmid isolation kit (#12123; Qiagen). Around 0.5 × 106 cells were seeded onto a single well of a 6-well plate and after 24 h and transfected with 1.5 μg of the corresponding ligated plasmids using either Lipofectamine stem cell transfection reagent (#STEM00001; Thermo Fisher Scientific) for iPSCs or Lipofectamine 3000 (#L3000008; Thermo Fisher Scientific) for other cell lines. After 24 h, GFP-positive cells were single-cell cloned on 96-well plates using FACS Aria II flow cytometer. Culture medium was replaced with fresh medium after 24 h, and after 14 d, single-cell clones were analyzed by IB to confirm the KO of the corresponding gene.
Plasmid and siRNA transfection
Cells were seeded on culture plates at a density of 20,000 cells per cm2. After 24 h, plasmid transfection was performed using Lipofectamine 3000 according to the manufacturer’s instructions. A single well of a 6-well plate at 70% confluence was transfected with 1.5 μg of plasmid DNA. For gene KDs, siRNA pools targeting RAB27A, HRS, ATG5, KIF1B, and KIF5B were obtained from Dharmacon (ON-TARGETplus siRNA, SMARTpools, Horizon, PerkinElmer; Table S1). Cells were transfected with 20 nM siRNA pools twice at intervals of 24 h using Lipofectamine 3000 according to the manufacturer’s instructions. Briefly, for a single well of a 12-well plate, 1.5 μl Lipofectamine 3000 was mixed with 50 μl Opti-MEM medium (#31985070; Gibco; Thermo Fisher Scientific), and in a separate tube, 1 μl of siRNA pool was mixed with 50 μl Opti-MEM medium, and incubated for 5 min. Next, the contents of both tubes were mixed and incubated for 10 min before adding them to a single well. This step was repeated after 24 h. After 12 h, cells were replated in three wells of a 6-well plate with fresh medium (3 ml/well). After 24 h, the conditioned medium was collected for exosome isolation.
Antibodies
List of primary antibodies used: Rab27A (mouse, #ab55667; Abcam, 1:1,000 for IB), ATG5 (rabbit, #12994T; Cell Signaling Technology, 1:1,000 for IB), cathepsin D (rabbit, 1:1,000 for IB, #06-114; Millipore Sigma, 1:1,000 for IB), BORCS7 (rabbit, #PAB23142; Abnova, 1:1,000 for IB), ARL8A (rabbit, #17060-1-AP; Proteintech, 1:1,000 for IB), ARL8B (rabbit, 13049-1-AP; Proteintech, 1:1,000 for IB), KIF5B (rabbit, #ab5629; abcam, 1:2,000 for IB), KIF1B (rabbit, #15263-1-AP; Proteintech, 1:1,000 for IB), VPS39 (rabbit, #NBP1-76535; Novus Biologicals, 1:1,000 for IB). CD63 (H5C6, mouse, prepared in-house, 1:1,000 for IB, 1:500 for IF), LAMP1 (D2D11, rabbit, #9091S; Cell Signaling Technology, 1:1,000 for IB, 1:500 for IF), CD81 (454720, mouse, #FAB4615V; Novus Biologicals, 1:250 for IF), Syntenin-1 (EPR8102, rabbit, #ab133267; Abcam, 1:1,000 for IB), EGFR (rabbit, #18986-1-AP; Proteintech, 1:1,000 for IB), RAB7A (D95F2, Rabbit, #9367S; Cell Signaling Technology, 1:1,000 for IB), PLEKHM2 (rabbit, #HPA032304; Atlas Antibodies, 1:500 for IB), HRS (D7T5N, rabbit, #15087S; Cell Signaling Technology, 1:1,000 for IB), CD81 (D3N2D, rabbit, #56039S; Cell Signaling Technology, 1:1,000 for IB), CD98 (rabbit, #15193-1-AP; Proteintech, 1:10,000 for IB), CD147 (1G12B5, mouse, #66443-1-Ig; Proteintech, 1:10,000 for IB), TSG101 (51/TSG101, mouse, #612696; BD Biosciences, 1:1,000 for IB), β-actin (AC-15, mouse, #ab49900; Abcam, 1:10,000 for IB), CD63-FITC (H5C6, mouse, #557288; BD Biosciences, 20 μl/reaction), IgG-FITC (MOPC21, mouse, #400108; BioLegend, 4 μl/reaction), GM130 (D6B1, rabbit, #12480; Cell Signaling Technology, 1:1,000 for IB), and Calnexin (AF18, Mouse, MA3-027; Invitrogen; Thermo Fisher Scientific, 1:1,000 for IB).
List of secondary antibodies used: anti-rabbit IgG antibody labeled with HRP (goat, #NEF812001EA; Perkin Elmer, 1:5,000 for IB), anti-mouse IgG antibody labeled with HRP (goat, #NEF822001EA; Perkin Elmer, 1:5,000 for IB), anti-mouse IgG (H+L) secondary antibody Alexa Fluor 647 (goat, #A-21235; Invitrogen; Thermo Fisher Scientific, 1:500 for IF), anti-rabbit IgG (H+L) secondary antibody, Alexa Fluor 647, (donkey, #A-31573; Invitrogen; Thermo Fisher Scientific, 1:500 for IF), anti-rabbit IgG (H+L) secondary antibody, Alexa Fluor 555 (donkey, A-31572; Invitrogen; Thermo Fisher Scientific, 1:500 for IF), and anti-mouse IgG (H+L) secondary antibody, Alexa Fluor 488 (donkey, A-21202; Invitrogen; Thermo Fisher Scientific, 1:500 for IF).
IB
Equal number of cells (20,000 per cm2) was seeded for all the experiments. Cells were washed with ice-cold 1 × PBS (#RGF-3210; KD Medicals) and lysed in 1 × LDS loading buffer (#B0007; Thermo Fisher Scientific) with 5% 2-mercaptoethanol (#M3148; Millipore Sigma) under agitation for 15 min at room temperature. For preparing EV lysates, three parts of samples were mixed with one part of 4 × LDS loading buffer containing 5% 2-mercaptoethanol. To perform an IB analysis of CD63, samples were prepared as described above except that 2-mercaptoethanol was not added. Samples were heated at 95°C for 6 min and snap-chilled on ice. Samples were separated by SDS-PAGE on gels made with 10 or 15% acrylamide and transferred onto nitrocellulose membranes (#1620113; Bio-Rad Laboratories). The membranes were blocked for 1 h in blocking buffer composed of 5% non-fat milk (#706404XTU; Bio-Rad Laboratories) dissolved in 1 × Tris-buffered saline plus 0.01% Tween 20 (TBST; #P1379; Millipore Sigma). Primary antibodies were added in blocking buffer and incubated overnight at 4°C with rocking motion. Blots were washed three times (5 min each) with 1 × TBST and incubated for 1 h with the corresponding secondary antibodies added in blocking buffer. Blots were washed again three times in 1 × TBST, revealed with SuperSignal West Femto maximum sensitivity substrate (#34095; Thermo Fisher Scientific), and imaged using a ChemiDoc MP gel documentation system (#12003154; BioRad Laboratories).
FACS analysis
Exosomes
Crude exosomes derived from 10 ml of conditioned medium were incubated with anti-CD63-coated magnetic beads (30,000 beads per sample; #10606D; Thermo Fisher Scientific) overnight at 4°C with rotation. Beads were washed twice with FACS buffer (1% EV-depleted FBS in PBS) and incubated with human IgG (#I4506; Millipore Sigma) at 4°C for 20 min. After two washes with FACS buffer, the beads were incubated with phycoerythrin (PE)-labeled anti-CD63 or PE-isotype control at 1:30 dilution (#556020, #555749; BD Bioscience) for 45 min at room temperature with slow rotation mixing. The samples were washed twice and a total of 10,000 events were acquired on a LSRFortessa Cell Analyze (BD Bioscience) and analyzed using FlowJo Software version 10.8.0 (FlowJo, LLC).
Cells
Cells were collected by trypsinization (#25053CI; Corning), washed with PBS, and suspended in FACS buffer (1% FBS in PBS). Cells were kept at 4°C. A total of 5,000 events were acquired on a LSRFortessa Cell Analyzer (BD Bioscience) and analyzed using FlowJo Software version 10.8.0 (FlowJo, LLC).
IF
HeLa cells were seeded at 20,000 per cm2 on sterilized coverslips in a 24-well plate. After 24 h, cells were washed once with PBS and fixed with 4% paraformaldehyde (#30525-89-4; Thermo Fisher Scientific) for 20 min at room temperature. After fixation, cells were washed three times with PBS and blocked-permeabilized for 1 h at room temperature with blocking buffer (3% BSA [#9048-46-8; Millipore Sigma], 0.1% saponin [#47036; Millipore Sigma], and 0.02% sodium-azide [#26628-22-8; Millipore Sigma] in PBS). Next, cells were incubated with 1:250–1:500 dilution of primary antibody in blocking buffer overnight at 4°C, washed with PBS three times (5 min per wash), and incubated with a 1:500 dilution of secondary antibody for 1 h at room temperature. Finally, cells were washed three times with PBS (5 min per wash) and mounted on glass slides using Fluoromount-G reagent with DAPI (#00-4959-52; Invitrogen; Thermo Fisher Scientific). All incubations were performed in a humidified chamber to prevent drying of the samples. After 24 h, cells were imaged on a confocal microscope (LSM780; Carl Zeiss) with an oil-immersion 63×/1.40 NA Plan Apochromat Oil DIC M27 objective lens (Carl Zeiss). Settings (i.e., gain, laser power, and pinhole) were kept constant for images presented for comparison. Images were acquired using ZEN 2012 software (Carl Zeiss) and processed with Fiji software (https://fiji.sc).
Measurements of endolysosome position
To determine the position of endolysosomes for cells in Fig. 1 D, we performed a shell analysis (Williamson et al., 2022). Briefly, cells were plated on collagen I (#3442-005-01; R&D systems at 5 μg/cm2)–coated plates, imaged, and maximum intensity projections of z-stack confocal micrographs were subjected to analysis. Cells exhibiting morphologies where perinuclear clusters of lysosomes touched the plasma membrane were excluded from analysis. Cell outlines were traced in Fiji and the total fluorescence of LAMP1 signal was measured. Using the “enlarge” function in Fiji, the cell outline was shrunk by 1.7 μm and LAMP1 signal intensity was remeasured. The intensity of LAMP1 signal within the peripheral 1.7 μm shell was plotted as percentage of total cellular LAMP1 signal. One-way ANOVA, followed by multiple comparisons using the Dunnett’s test, was used for statistical analysis of data sets. Statistical analyses were done using Prism version 9 (GraphPad Software).
Extracellular vesicle isolation and purification
HeLa cells were seeded at 25,000 cells per cm2 in a 100-mm cell culture dish. EV isolation and purification were conducted using complete medium (10 ml or 30 ml, depending on the experiment) obtained from cells at 70–80% confluency cultured for 16–48 h (see schematic representation in Fig. 2 A). Supernatants were obtained from cultures of cells with >95% viability. Cell culture medium was fractionated by centrifugation using the following steps: (i) 300 × g for 10 min (intact cells and cell debris), (ii) 2,000 × g for 20 min (apoptotic bodies), (iii) 10,000 × g for 40 min (large EVs/ectosomes/microvesicles). The supernatant of this last centrifugation was passed through a 0.2-μm filter (#83.1826.001; Sarstedt). Various concentrations of iodixanol-sucrose buffered solution were prepared by mixing homogenization medium and 50% of iodixanol-containing working solution. Homogenization medium was composed of 0.25 M sucrose (#S0389; Millipore Sigma), 1 mM EDTA (#RGC5135; KD Medical, Inc), and 10 mM Tris-HCl (#10708976001; Millipore Sigma) at pH 7.4. Working solution with 50% of iodixanol was prepared by mixing one part of iodixanol (OptiPrep, #07820; Stem Cells Technologies) with five parts of solution containing 0.25 M sucrose, 6 mM EDTA, and 60 mM Tris-HCl at pH 7.4. These buffering conditions maintain the integrity of EVs. Filtered supernatant (10 ml) was layered on top of a density cushion made up of 500 μl 10 and 50% iodixanol-sucrose buffered solution and centrifuged at 150,000 × g for 3 h in a SW41-Ti rotor (k-factor = 172). The interphase between 10 and 50% iodixanol-sucrose buffered solution contained crude exosomes. These crude exosomes were collected and subjected to further purification on a density gradient. Briefly, crude exosomes preparation (∼1 ml) was mixed with 3 ml of 50% iodixanol-sucrose buffered solution to a final concentration of 45% and layered at the bottom of the tube. Layers of 1 ml of 30, 28, 20, 10, and 0% iodixanol-sucrose buffered solution were successively overlaid and tubes were centrifuged at 120,000 × g for 18 h in a SW41-Ti rotor (k-factor = 214). Purified exosomes were collected from fractions 2 and 3 corresponding to ∼10 and 20% iodixanol-sucrose buffered solution and pooled for further analysis. Depending on the downstream application, 2 ml exosome suspension was either used directly or concentrated in 0.4 ml of PBS using centrifugation at 150,000 × g for 2 h in a TLA-100.3 fixed angle rotor (k-factor = 45).
EGF-AF488 uptake
HeLa cells were seeded at 20,000 cells per cm2 onto a sterile coverslip placed in one well of a 24-well plate with complete medium. After 24 h, the coverslips were replaced with fresh medium containing 100 ng/ml Alexa Fluor-488 conjugated EGF (EGF-AF488; #13345; Thermo Fisher Scientific) for 2 h at 37°C. Finally, the coverslips were washed with PBS, fixed with 4% paraformaldehyde, and mounted onto glass slides as described in the “IF” section. Images were acquired with a confocal microscope (LSM780; Zeiss). The accumulation of EGF-AF488 was measured by averaging the raw integrated density from each cell using Fiji software.
DQ-Green-BSA uptake
HeLa cells were seeded at 20,000 per cm2 onto a sterile coverslip placed in one well of a 24-well plate with complete medium. After 24 h, cells were exposed to a medium supplemented with DQ-Green-BSA (10 μg/ml; #D12050; Thermo Fisher Scientific) for 2 h at 37°C. After a PBS wash and 4% paraformaldehyde-based fixation, coverslips were mounted onto glass slides and imaged in the AF488 channel using a confocal microscope (LSM780; Zeiss). DQ-Green-BSA fluorescence was quantified by averaging the raw integrated density from each cell using Fiji software.
Dextran-AF488 delivery to Magic Red cathepsin D compartments
To determine if internalized dextran is targeted to active cathepsin B–positive lysosomal compartments, cells were seeded at 20,000 per cm2 onto a 4-well chambered cover glass slide. After 24 h, cells were incubated with complete medium containing 1 mg/ml of 10,000 kD dextran-AF488 (anionic, fixable, #D22910; Thermo Fisher Scientific) for 2 h at 37°C. Cells were rinsed with PBS and chased for 1 h at 37°C in complete medium. Cells were then rinsed with PBS and incubated with complete medium containing Magic Red MR-(RR)2 cathepsin B substrate (1:2,500 dilution; #SKU:937; Immunochemistry Technologies, LLC). This non-fluorescent, cell-permeable dye becomes red fluorescent after it is acted upon by cathepsin B. After 10–15 min of incubation, the cells were placed in an imaging chamber with preheated (37°C), humidified atmosphere containing 5% CO2, and imaged live using an LSM780 confocal microscope for 3–5 min. The degree of colocalization of red (Magic Red) and green (dextran-AF488) channels corresponding to the fraction of endocytic cargo that arrived in Cathepsin B–positive compartment was measured using the Pearson colocalization coefficient in Fiji software and a custom macroscript for automation of analysis.
Dextran secretion assay
To determine the fate of fluid-phase cargo, cells were seeded in a 24-well plate at a density of 20,000 cells per cm2. After 24 h, cells were incubated with 100 μg/ml dextran-AF488 (10,000 kD anionic, fixable #D22910; Thermo Fisher Scientific) for 4 h at 37°C, washed with PBS, and chased for 20 h in complete medium. In the last 2 h of chase, cells were incubated with 100 μg/ml dextran-AF647 (10,000 kD anionic, fixable #D22914; Thermo Fisher Scientific) at 37°C. After 2 h of exposure, cells were washed, trypsinized, and analyzed by FACS, as described in a previous section. Mean fluorescence intensity (MFI) of dextran-AF488 and dextran-AF647 was measured for each cell and the ratio of both fluorescent signals was calculated. Lower ratios represent higher secretion and vice versa.
Particle measurement using NanoSight
Culture supernatant was processed as described in section “Extracellular vesicle isolation and purification” up to the filtration step. The number of particles was determined using NanoSight (#NS300; Malvern Panalytical). All experiments were performed on the same day; samples were kept at 4°C and not frozen to avoid particle damage. Small EV–enriched supernatant was brought to room temperature just before injection into the NanoSight device through a motorized pump to avoid turbulence-induced Brownian motion of particles. Three independent samples were used for each set of samples. Three measurements for each sample were performed and the average of this value was used to determine particle numbers. Sensitivity of the camera was kept the same for all measurements. Data were analyzed using the NanoSight software suite.
TIRF
To evaluate the fusion of MVEs with the plasma membrane, HeLa cells were transfected with pCMV-Sport6-CD63-pHluorin (gift from D.M. Pegtel, #130901; Addgene) and imaged by TIRF for CD63-pHluorin fluorescence upon exposure to the neutral pH of the medium. Imaging was conducted on a NIKON Eclipse Ti spinning disk microscope equipped with temperature-controlled stage module (37°C and CO2 at 5%), high-speed EM charge-coupled device camera (iXon DU897 from Andor), and NIS-Elements AR microscope imaging software. Images were acquired at 3 Hz for 3–5 min using an Apo TIRF 100× objective (numerical aperture 1.49) and iXon DU897 camera.
HRP uptake, diaminobenzidine (DAB) reaction, and TEM analysis
Cells were seeded on sterilized glass coverslips at 70% cell density. After 24 h, cells were incubated with 5 mg/ml HRP (#P8375; Millipore Sigma) for 4 h. Cells were fixed for 30 min with 1% glutaraldehyde in 0.1 M cacodylate/2 mM CaCl2 (CAC) buffer, pH 7.4, and washed three times for 2 min each with CAC. The coverslips were incubated for 1 min with 1.8 ml of 0.1% DAB, 200 μl 0.1% H2O2 was added, and incubation was continued for 2 min in the dark. Coverslips were washed three times for 2 min each with CAC, postfixed with 1% OsO4/1.5% K3Fe(CN)6 in CAC, and washed three times for 2 min each with CAC. After washing with 50 mM acetate buffer, pH 5.5, the cells were stained for 30 min with 1% uranyl acetate, washed with water, dehydrated with increasing concentrations of ethanol, and embedded in EMBed 812 (#14120; EMS). After polymerization, the coverslips were removed with hydrofluoric acid, and blocks of ∼1 × 1 mm were cut out and remounted on a carrier. Ultrathin sections of 60–70 nm were cut and mounted on EM grids.
Negative staining and TEM of exosomes
Exosomes were isolated from 30 ml of culture medium from HeLa cells seeded on a 100-mm dish at 20,000 cells per cm2 for 24 h at 37°C, as described in the section “Extracellular vesicle isolation and purification.” Purified exosomes were concentrated to 0.4 ml in PBS by ultracentrifugation of samples at 150,000 × g for 2 h. Concentrated exosome samples (3 μl) were applied on a freshly glow-discharged carbon-coated EM grid for 5 min followed by two washes of 30 s each with water, and negative staining with 0.75% uranyl formate twice for 20 s each. The staining solution was removed using filter paper and grids were air-dried.
EM
Samples were imaged on an FEI Tecnai 20 transmission electron microscope (Thermo Fisher Scientific) operated at 120 kV. Images were recorded using AMT XR81 CCD (AMT) camera and processed digitally.
Statistical analysis
Data are presented as mean ± SEM. As indicated in the figure legends, in most cases one-way ANOVA, followed by multiple comparisons using the Dunnett’s test, was used for statistical analysis. Tukey’s test was used for multiple comparisons in Fig. 7 E. Unpaired t-test was used for Fig. 3 B. For parametric tests, data distribution was assumed to be normal, but this was not formally tested. Prism version 9 (GraphPad Software) was used for these tests. Significant differences between control and treated samples are indicated (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001).
Online supplemental material
Figs. S1, S2, S3, and S4 show results from additional experiments. Fig. S1 shows the growth rate of WT and KO cells, and the presence of RAB7A, EGFR, CD81, HRS, β-actin, and calnexin in a 120,000 × g (120 K) fraction obtained from WT, BORCS7-KO, ARL8A-B-KO, and VPS39-KO HeLa cells. Fig. S2 shows electron micrographs and particle size analysis of 120-K fractions from WT, BORCS7-KO, ARL8A-B-KO, and VPS39-KO HeLa cells. Fig. S3 shows the presence of CD63, HRS, and TSG101 in 120-K fractions from WT, RAB7A-KO, and PLEKHM2-KO HeLa cells. Fig. S4 shows results from additional experiments supplementing Fig. 3. Table S1 includes a list of siRNAs, cell lines, plasmids, guide RNAs, and other reagents used in this study.
Data availability
Original IBs underlying Figs. 1 B, 2, B, D, and E, 3, A, C, E, and F, S1, B and E, S3, D and E, and S4, A–D are available in the supplementary source data files. All other data are available in the published article and its online supplemental material, or from the corresponding author upon reasonable request.
Acknowledgments
We thank Raffaella De Pace (NICHD, NIH, Bethesda, MD, USA) for i3Neuron culture supernatants, Morié Ishida (NICHD, NIH, Bethesda, MD, USA) for RAB7A-KO HeLa cells, Tal Keren-Kaplan (NICHD, NIH, Bethesda, MD, USA) for PLEKMH2-KO Hela cells, D.M. Pegtel (Amsterdam UMC Vrije Universiteit Amsterdam, Netherlands) for pCMV-Sport6-CD63-pHluorin (#130901; Addgene), Feng Zhang (Massachusetts Institute of Technology, Cambridge, MA, USA) for pSpCas9(BB)-2A-GFP (pX458) (#48138; Addgene), N. Altan-Bonnet (National Heart, Lung, and Blood Institute, NIH, Bethesda, MD, USA) for the use of the NanoSight NTA, and other members of the Bonifacino laboratory for helpful discussions.
This work was supported by the Intramural Program of NICHD, NIH, to J.S. Bonifacino (ZIA HD001607), the European Union’s Horizon 2020 Research and Innovation Programme under the Marie Skłodowska-Curie grant agreement No 754412, and a fellowship from the Swedish Society for Medical Research to G.V. Shelke.
Author contributions: G.V. Shelke performed most of the experiments, C.D. Williamson helped with TIRF microscopy and image analysis, M. Jarnik performed electron microscopy. G.V. Shelke and J.S. Bonifacino conceived the study, analyzed data, and wrote the manuscript. All authors read and edited the manuscript.
References
Author notes
Disclosures: The authors declare no competing interests exist.