Polyamines such as spermidine and spermine are found in nearly all cells, at concentrations ranging up to 0.5 mM. These cations are endogenous regulators of cellular K+ efflux, binding tightly in the pores of inwardly rectifying K+ (Kir) channels in a voltage-dependent manner. Although the voltage dependence of Kir channel polyamine blockade is thought to arise at least partially from the energetically coupled movements of polyamine and K+ ions through the pore, the nature of physical interactions between these molecules is unclear. Here we analyze the polyamine-blocking mechanism in the model K+ channel MthK, using a combination of electrophysiology and computation. Spermidine (SPD3+) and spermine (SPM4+) each blocked current through MthK channels in a voltage-dependent manner, and blockade by these polyamines was described by a three-state kinetic scheme over a wide range of polyamine concentrations. In the context of the scheme, both SPD3+ and SPM4+ access a blocking site with similar effective gating valences (0.84 ± 0.03 e0 for SPD3+ and 0.99 ± 0.04 e0 for SPM4+), whereas SPM4+ binds in the blocked state with an ∼20-fold higher affinity than SPD3+ (Kd = 28.1 ± 3.1 µM for SPD3+ and 1.28 ± 0.20 µM for SPM4+), consistent with a free energy difference of 1.8 kcal/mol. Molecular simulations of the MthK pore in complex with either SPD3+ or SPM4+ are consistent with the leading amine interacting with the hydroxyl groups of T59, at the selectivity filter threshold, with access to this site governed by outward movement of K+ ions. These coupled movements can account for a large fraction of the voltage dependence of blockade. In contrast, differences in binding energetics between SPD3+ and SPM4+ may arise from distinct electrostatic interactions between the polyamines and carboxylate oxygens on the side chains of E92 and E96, located in the pore-lining helix.
Introduction
Polyamine molecules, found in the cytosol of nearly all cells, can act as endogenous regulators of transmembrane ion fluxes (Lopatin et al., 1994, 1995; Shyng et al., 1996). These molecules, which are linear chains of cationic primary and secondary amines linked by 3- or 4-hydrocarbon spacers, are best known as limiters of K+ efflux through eukaryotic inwardly rectifying K+ (Kir) channels (Nichols and Lopatin, 1997). In addition, the polyamines spermidine (SPD3+) and spermine (SPM4+) are known to inhibit ion flow through other cation channels, including eukaryotic CNG channels and BK channels (Lu and Ding, 1999; Guo and Lu, 2000a; Zhang et al., 2006).
In Kir channels, polyamine blockade exhibits a steep voltage dependence corresponding to the movement of up to four elementary charges across the transmembrane electric field. This charge movement is thought to arise either from direct (slow) permeation of the polyamine through the pore or from a role of the polyamine in displacing up to four K+ ions from the pore during blockade, or possibly some combination of polyamine and K+ permeation (Lopatin et al., 1995; Guo and Lu, 2000b; Kurata et al., 2004; Shin and Lu, 2005; Kucheryavykh et al., 2007). Consequently, our understanding of the structural basis for control of physiological K+ efflux by these polyamines is incomplete. It is a matter of debate whether the polyamine binding site is contained within the vestibule of the K channel pore, or alternatively whether a polyamine can enter the narrow selectivity filter (SF) of the pore (Lopatin et al., 1995; Guo and Lu, 2000b; Kurata et al., 2004; Shin and Lu, 2005; Kucheryavykh et al., 2007). In addition, it is not known whether polyamines simply occlude a “passive” pore that is structurally inert during blockade, or whether interactions between a polyamine and the pore might have an impact on relative binding affinities for different species of polyamine.
In this work, we studied polyamine blockade of MthK, a model prokaryotic K+ channel, to gain insight toward the chemical and structural basis of this phenomenon. Similar to Kir channels, MthK exhibits inward rectification in the presence of physiological levels of free Mg2+ or polyamines (Thomson and Rothberg, 2012; Thomson et al., 2014). The structure of the MthK pore is known, and this structural knowledge is complemented by a quantitative framework defining the functional properties of the channel, including its gating by Ca2+, H+, and voltage, and blockade of the pore by Mg2+ and other ions (Jiang et al., 2002; Dong et al., 2005; Parfenova et al., 2006; Parfenova and Rothberg, 2006; Ye et al., 2006, 2010; Zadek and Nimigean, 2006; Kuo et al., 2007, 2008; Dvir et al., 2010; Pau et al., 2010, 2011; Thomson and Rothberg, 2010; Shi et al., 2011; Smith et al., 2012, 2013; Posson et al., 2013, 2015; Thomson et al., 2014). Comparison of the kinetics of SPD3+ versus SPM4+ blockade indicates that the polyamines block current with similar effective valences, whereas SPM4+ blockade exhibits an ∼20-fold higher affinity than SPD3+. Molecular simulations suggest that in MthK, the leading charged amine group of either polyamine can reach a binding site comprising in part the hydroxyl groups of T59, at the threshold of the SF, and that access to the threshold site is coupled with outward movement of K+.
Materials and methods
Electrophysiology
Recordings were obtained using methods described previously (Thomson and Rothberg, 2010; Thomson et al., 2014). Briefly, MthK channels were overexpressed and purified by metal-affinity and size-exclusion chromatography and reconstituted into proteoliposomes comprised of 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine:1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-(1′rac-glycerol; POPE:POPG; 3:1 ratio), which were stored at −80°C until use. The amount of protein used in reconstitution ranged from 5 to 25 µg protein/mg lipid.
For electrophysiological experiments, proteoliposomes were thawed and incorporated planar lipid bilayers of POPE:POPG (3:1) in a horizontal bilayer chamber. Unless otherwise specified, solution in the cis (top) chamber contained 200 mM KCl and 10 mM HEPES (pH 7.0). Solution in the trans (bottom) chamber contained 200 mM KCl, 10 mM HEPES (pH 8.1), and 100 µM CdCl2 to fully activate MthK channels (Smith et al., 2013; Thomson et al., 2014). Under these conditions, MthK channels are activated with open probabilities of >95% over a wide range of voltage (from −150 to 150 mV), and activation with Cd2+ precludes voltage-dependent fast blockade that can arise when activating the channels using high concentrations of Ca2+ (Thomson et al., 2014). Solutions in the trans chamber also contained the specified concentration of SPD3+, ranging from 10 to 1,000 µM, or SPM4+, ranging from 0.1 to 100 µM. Currents were analyzed using bilayers that contained 10 to 15 active MthK channels.
Currents were acquired in response to continuous voltage ramps from −150 to 150 mV. The ramp speed (200 mV/s) appeared slow enough to allow equilibration of rapid channel blockade by polyamines, as suggested by the observation that current–voltage relations elicited by ramps from negative to positive voltages were effectively the same as those elicited by ramps from positive to negative voltages. Currents were amplified using a Dagan PC-ONE patch clamp amplifier with low-pass filtering to give a final effective filtering of 333 Hz (dead time of 0.538 ms) and sampled by computer at a rate of 50 kHz using pClamp 9.2. Multiple ramp currents were acquired and averaged for each bilayer in each condition (i.e., polyamine concentration). Within each bilayer, multiple solution changes were performed using a gravity-fed perfusion system, and to ensure completeness of solution changes, the trans chamber was washed with a minimum of 10 ml (∼10 chamber volumes) of solution before recording under a given set of conditions. Current–voltage relations were averaged across bilayers following normalization, which was performed by dividing the ramp current trace by the current amplitude at −100 mV (at which it was observed that high concentrations of polyamine had almost no effect on current amplitude). Fractional currents were determined by dividing the current in the presence of polyamine by the current from the same bilayer in the absence of polyamine. Fractional currents close to the reversal potential (between −7 and +7 mV) tended to have a very high variance and were excluded from analysis.
Molecular dynamics simulations
Molecular dynamics simulations were performed using the MthK pore structure (Protein Database [PDB] accession no. 3LDC; Ye et al., 2010), by methods described previously (Barber et al., 2014). Briefly, molecular systems were assembled using the CHARMM-GUI web service (Woolf and Roux, 1994; Jo et al., 2008). The channel protein, with its symmetry axis aligned along the z axis, was embedded in a lipid bilayer of dipalmitoyl-phosphatidylcholine. The number of ions in the bulk was adjusted to reproduce experimental ionic concentrations (200 mM KCl) and to obtain electrical neutrality. The molecular system contained ∼59,170 atoms total. Calculations were performed using the NAMD computational code (Phillips et al., 2005), using the all-atom potential energy function CHARMM36 for protein and phospholipids, and the TIP3P potential for water molecules (Jorgensen et al., 1983; Mackerell et al., 2004). Periodic boundaries conditions were applied, and long-range electrostatic interactions were treated by the particle mesh Ewald algorithm (Essmann et al., 1995). The SPD3+ and SPM4+ molecules were modeled using CHARMM general force field (Vanommeslaeghe and MacKerell, 2012; Vanommeslaeghe et al., 2012). Before the initial equilibration, we added one polyamine molecule to the system (either SPD3+ or SPM4+, in the fully protonated state and in the extended conformation) by manually choosing the optimal rototranslations of the ligand molecule using the VMD software. The molecular systems were equilibrated for ∼2 ns with decreasing harmonic restraints applied to the protein atoms, the pore ions, and the water molecules localized in the filter. This was followed by an additional equilibration phase of 20 ns of unbiased molecular dynamics simulation. All trajectories were generated with a time step of 2 fs at constant normal pressure (1 atm) controlled by a Langevin piston and constant temperature (323 K; Nose, 1984; Hoover, 1985; Feller et al., 1995).
Generation of free energy profiles
Free energy calculations were performed using the metadynamics approach, as described previously (Stock et al., 2013; Delemotte et al., 2015). This approach uses a biasing potential that discourages the system from visiting the same regions of the configurational space repeatedly; thus, the approach is computationally efficient and yields results that are consistent with those obtained by other methods (such as umbrella sampling). Asymptotically, the biasing potential compensates exactly the underlying free energy surface, thereby providing a convenient way to reconstruct the free energy surface along a chosen set of collective variables (Laio and Parrinello, 2002). Two collective variables were used to bias the sampling. The first is the distance along the z axis between the center of mass of the polyamine and that of the SF. For the second collective variable, we first calculated the number of contacts between any of the potassium ions located in the SF and the centers of mass of the binding sites lined by residues 59 and 60. For each of these sites, the number of contacts is 1 when the site is occupied, 0 when the site is not occupied, or a number between 0 and 1 for any intermediate state. Finally, the collective variable was defined as the difference between these two numbers. Intuitively, this collective variable, which spans the range between −1 and 1, can be understood as reporting progress along the displacement of the ion between the two sites.
Generation of histograms
The reweighting procedure described above results in unbiasing of statistical estimators, and consequently in a normal distribution of the sample means. Moreover, we considered values of the dynamical variables separated by a time lag of 1 ns, to ensure that samples were uncorrelated. Thus, errors associated with histograms and related free energy profiles were estimated as the sample variance divided by the square root of the sample size.
Online supplemental material
Fig. S1 presents SPD3+ blockade described with an alternative three-state model (Scheme 4). Fig. S2 shows the time series of RMS fluctuations for SPD3+ and SPM4+ during simulations at 0 and 500 mV. Fig. S3 shows water density distributions along the z axis corresponding to the molecular dynamics simulations in Fig. 6. Fig. S4 shows representative snapshots of MthK, ions, and all water molecules within 3 Å of the selectivity, corresponding to the molecular dynamics trajectory in Fig. 6 A. Fig. S5 illustrates a scheme for charge movement coupled to polyamine binding at the entry to the MthK SF. Table S1 shows parameters of Eq.1 determined from fitting the fractional blockade data as a function of [SPM3+] and [SPM4+].
Results
Current through MthK channels is inhibited by polyamines
In previous work, MthK channel current was found to be sensitive to rapid blockade by Mg2+, with an apparent Kd at 0 mV (Kapp(0)) of ∼1.8 mM Mg2+, which is thought to be within the physiological range of Mg2+ found in Methanothermobacter thermoautotrophicum as well as eukaryotes (Sprott and Jarrell, 1981). This apparent affinity for Mg2+ blockade is intermediate between that observed in “weak” inward rectifying Kir channels (such as ROMK1) and “strong” inward rectifiers (such as IRK1), consistent with an electrostatic environment in the MthK pore that might favor cationic blockers (Lu and MacKinnon, 1994). We thus wondered whether MthK channels might share other functional properties with Kir channels, such as blockade by polyamines, which are found in the cytosol of both archaea and eukaryotes (Michael, 2018). We addressed this possibility by measuring MthK channel currents in the presence of polyamines SPD3+ and SPM4+, applied to the cytosolic side of the channels.
Effects of SPD3+ were assayed using voltage ramp currents from bilayers containing multiple (>10) channels. Fig. 1 A shows that SPD3+ blocked outward K+ current through MthK channels in a dose-dependent manner, such that 1 mM SPD3+ inhibited >90% of the outward current at depolarized voltages. The voltage dependence of blockade was quantified by first plotting the fractional current at each voltage as a function of [SPD3+] and fitting these data with a Eq. 1 to estimate Kapp at each voltage (Fig. 1 B and Table S1). These Kapp values are then plotted as a function of voltage and fitted with an exponential function to estimate the effective valence for SPD3+ blockade, as well as the Kapp(0) (Fig. 1 C; Woodhull, 1973). This analysis yielded mean Kapp(0) values of 860 ± 20 µM and zδ values of 0.67 ± 0.03 e0 (n = 4 bilayers). Performing this same analysis with the tetravalent polyamine SPM4+ yielded Kapp(0) values of 77 ± 6 µM and zδ values of 0.88 ± 0.05 e0 (n = 4 bilayers; Fig. 2). Thus by this analysis, the Kapp(0) for SPM4+ in MthK was ∼10-fold lower than that for SPD3+, whereas zδ was ∼30% greater.
MthK channel blockade by SPD3+.(A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPD3+] (≤1,000 µM) using a voltage-ramp protocol and normalized to the current amplitude at −100 mV for the same bilayer with 0 SPD3+. (B) Fractional current versus [SPD3+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages (parameters are in Table S1). (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 860 ± 20 µM, zδ = 0.67 ± 0.03 e0).
MthK channel blockade by SPD3+.(A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPD3+] (≤1,000 µM) using a voltage-ramp protocol and normalized to the current amplitude at −100 mV for the same bilayer with 0 SPD3+. (B) Fractional current versus [SPD3+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages (parameters are in Table S1). (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 860 ± 20 µM, zδ = 0.67 ± 0.03 e0).
MthK channel blockade by SPM4+. (A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPM4+] (≤100 µM) using a voltage-ramp protocol and normalized as described in Fig. 1. (B) Fractional current versus [SPM4+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages. (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 77 ± 6 µM, zδ = 0.88 ± 0.05 e0).
MthK channel blockade by SPM4+. (A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPM4+] (≤100 µM) using a voltage-ramp protocol and normalized as described in Fig. 1. (B) Fractional current versus [SPM4+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages. (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 77 ± 6 µM, zδ = 0.88 ± 0.05 e0).
By an analysis similar to the one described above, large quaternary amine ions were observed previously to block MthK currents with effective valences (zδ) ranging from 0.67 to 0.81 e0 (Posson et al. 2013). The range of effective valences observed for both quaternary amines and polyamines may thus arise from channel/blocker interactions attributable to properties of the blockers other than the total charges of the blocker. In other words, although SPD3+ and SPM4+ have total valences that differ from one another by one charge, each molecule is essentially a “string” of spaced point charges. It has been hypothesized that the “leading charge” likely interacts with the SF (where most of the transmembrane voltage drop occurs), whereas the other charges may interact either with water molecules or with polar or charged side chains lining the pore. Thus the relatively low zδ values for blockade by SPD3+ and SPM4+ are consistent with the idea that only a single leading charge from either of these molecules crosses a fraction of the electric field during blockade, although this charge movement could also arise from movement of K+ ions or other charges through the field in response to polyamine binding.
Although our initial analysis is consistent with polyamine blockade involving interactions with the electric field, we wondered whether a simple two-state model based on this idea could quantitatively predict the voltage dependence of fractional current over a wide range of voltage (−150 to 150 mV). To test this, we calculated fractional current directly from MthK voltage-ramp currents over a range of [SPD] and fitted the currents using Scheme 1 to estimate values of Kd and zδ.
Kinetic parameters for SPD3+ blockade from Scheme 1
| . | Patch 1 . | Patch 2 . | Patch 3 . | Patch 4 . | Mean ± SEM . |
|---|---|---|---|---|---|
| Kd (μm) | 1130.8 | 1110.3 | 468.6 | 1251.2 | 990.2 ± 176 |
| zδ (e0) | 0.662 | 0.685 | 0.490 | 0.660 | 0.62 ± 0.05 |
| . | Patch 1 . | Patch 2 . | Patch 3 . | Patch 4 . | Mean ± SEM . |
|---|---|---|---|---|---|
| Kd (μm) | 1130.8 | 1110.3 | 468.6 | 1251.2 | 990.2 ± 176 |
| zδ (e0) | 0.662 | 0.685 | 0.490 | 0.660 | 0.62 ± 0.05 |
SPD3+ and SPD4+ blockade described with Scheme 1 . (A) Mean fractional MthK current over a range of [SPD3+], with Scheme 1 predictions superimposed (solid red curves). This model predicts that MthK currents approach 100% blockade with very strong depolarization, and thus provides a poor description of the data over the observed range of voltage and [SPD3+]. (B) Mean fractional MthK current over a range of [SPM4+], with Scheme 1 predictions superimposed. As with SPD3+, Scheme 1 predicts that MthK currents approach 100% blockade with very strong depolarization and is thus inconsistent with the data over the observed range of voltage and [SPM4+].
SPD3+ and SPD4+ blockade described with Scheme 1 . (A) Mean fractional MthK current over a range of [SPD3+], with Scheme 1 predictions superimposed (solid red curves). This model predicts that MthK currents approach 100% blockade with very strong depolarization, and thus provides a poor description of the data over the observed range of voltage and [SPD3+]. (B) Mean fractional MthK current over a range of [SPM4+], with Scheme 1 predictions superimposed. As with SPD3+, Scheme 1 predicts that MthK currents approach 100% blockade with very strong depolarization and is thus inconsistent with the data over the observed range of voltage and [SPM4+].
Kinetic parameters for SPM4+ blockade from Scheme 1
| . | Patch 5 . | Patch 6 . | Patch 7 . | Patch 8 . | Mean ± SEM . |
|---|---|---|---|---|---|
| Kd (μm) | 89.04 | 69.77 | 57.32 | 35.78 | 63.0 ± 11 |
| zδ (e0) | 0.835 | 0.808 | 0.683 | 0.648 | 0.74 ± 0.05 |
| . | Patch 5 . | Patch 6 . | Patch 7 . | Patch 8 . | Mean ± SEM . |
|---|---|---|---|---|---|
| Kd (μm) | 89.04 | 69.77 | 57.32 | 35.78 | 63.0 ± 11 |
| zδ (e0) | 0.835 | 0.808 | 0.683 | 0.648 | 0.74 ± 0.05 |
The experimental results from both SPD3+ and SPM4+ blockade instead appear to be consistent with a model in which at least one step in the blocking mechanism is voltage dependent, and at least one step beyond the voltage-dependent step exhibits a much weaker voltage dependence, to reach a plateau in the blocking effect. We next explored mechanisms to account for polyamine blockade over a wide range of voltages.
Description of polyamine blockade by a three-state kinetic scheme
The fractional blockade from SPD3+ predicted by Scheme 2 is illustrated in Fig. 4 A. By comparing the predictions of Scheme 1 and Scheme 2, it is evident that Scheme 2 provides a better description of the plateau effect observed with voltages >100 mV. Estimates of the parameters for Scheme 2 fitted with the SPD3+ block data yielded J0 = 0.027 ± 0.0013, zδ = 0.84 ± 0.032 e0, and Kd = 28.1 ± 3.1 µM (Table 3). Because Scheme 1 and Scheme 2 are nonnested models, there is no direct comparison between the parameters used in each model to describe the data. In terms of the underlying mechanism, however, Scheme 2 predicts that at very depolarized voltages, all channels will be in equilibrium between O’ and B’ that depends largely on polyamine concentration and consequently determines the fractional current at those voltages.
SPD3+ and SPM4+ blockade described with Scheme 2 . (A) Fractional MthK current over a range of [SPD3+] with Scheme 2 predictions superimposed. Scheme 2 predicts that depolarization drives the channel toward a receptive open state (O’) with enhanced blocker affinity, accounting well for the experimental data. Fit parameters for individual experiments are provided in Table 3. (B) Fractional MthK current over a range of [SPM4+] with Scheme 2 predictions superimposed. Scheme 2 accounts for the major features of SPM4+ blockade over a 1,000-fold range of [SPM4+]. Fit parameters for individual experiments are provided in Table 4.
SPD3+ and SPM4+ blockade described with Scheme 2 . (A) Fractional MthK current over a range of [SPD3+] with Scheme 2 predictions superimposed. Scheme 2 predicts that depolarization drives the channel toward a receptive open state (O’) with enhanced blocker affinity, accounting well for the experimental data. Fit parameters for individual experiments are provided in Table 3. (B) Fractional MthK current over a range of [SPM4+] with Scheme 2 predictions superimposed. Scheme 2 accounts for the major features of SPM4+ blockade over a 1,000-fold range of [SPM4+]. Fit parameters for individual experiments are provided in Table 4.
Kinetic parameters for SPD3+ blockade from Scheme 2
| . | Patch 1 . | Patch 2 . | Patch 3 . | Patch 4 . | Mean ± SEM . |
|---|---|---|---|---|---|
| J0 | 0.028 | 0.025 | 0.030 | 0.024 | 0.027 ± 0.0013 |
| Kd (μm) | 33.8 | 28.7 | 19.4 | 30.5 | 28.1 ± 3.1 |
| zδ (e0) | 0.90 | 0.88 | 0.76 | 0.81 | 0.84 ± 0.032 |
| . | Patch 1 . | Patch 2 . | Patch 3 . | Patch 4 . | Mean ± SEM . |
|---|---|---|---|---|---|
| J0 | 0.028 | 0.025 | 0.030 | 0.024 | 0.027 ± 0.0013 |
| Kd (μm) | 33.8 | 28.7 | 19.4 | 30.5 | 28.1 ± 3.1 |
| zδ (e0) | 0.90 | 0.88 | 0.76 | 0.81 | 0.84 ± 0.032 |
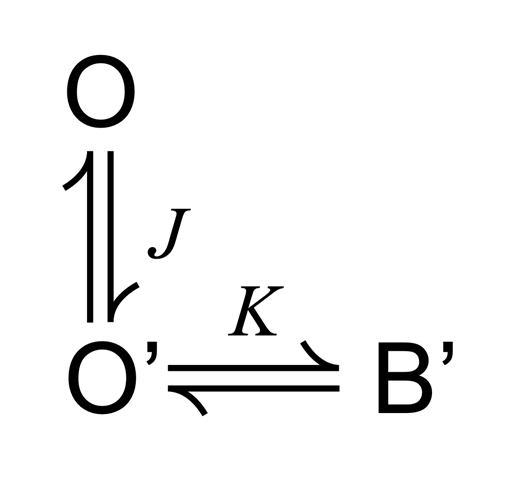
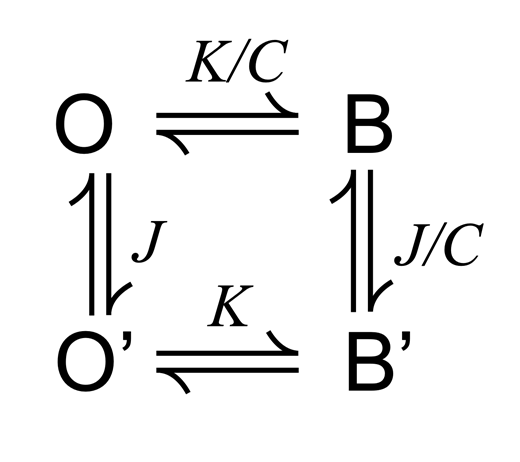
SPD3+ blockade described with an alternative three-state model (Scheme 4 ). Mean fractional MthK current (I/I0) over a range of [SPD3+], with Scheme 4 predictions superimposed (solid red curves). (A) Scheme 4 (left), with equations describing fractional current (right). (B) Mean fractional MthK current (I/I0) over a range of [SPD3+] as indicated, with Scheme 4 predictions superimposed (solid red curves). The best fit with this model (χ2 = 35.4) predicts that MthK currents will ultimately reach 100% blockade with very strong depolarization, although the apparent voltage dependence of blockade is less steep than that of Scheme 1. (C) An alternative fit representing a local minimum (χ2 = 97.6) predicts that MthK currents can increase with very strong depolarization.
SPD3+ blockade described with an alternative three-state model (Scheme 4 ). Mean fractional MthK current (I/I0) over a range of [SPD3+], with Scheme 4 predictions superimposed (solid red curves). (A) Scheme 4 (left), with equations describing fractional current (right). (B) Mean fractional MthK current (I/I0) over a range of [SPD3+] as indicated, with Scheme 4 predictions superimposed (solid red curves). The best fit with this model (χ2 = 35.4) predicts that MthK currents will ultimately reach 100% blockade with very strong depolarization, although the apparent voltage dependence of blockade is less steep than that of Scheme 1. (C) An alternative fit representing a local minimum (χ2 = 97.6) predicts that MthK currents can increase with very strong depolarization.
Scheme 2 predicts that depolarization drives the channel away from O, which effectively does not bind SPD3+, toward O’, which is receptive to SPD3+ with an effective affinity of 28 µM. We next tested whether Scheme 2 could further describe SPM4+ blockade and found that the scheme provided a very good description of the data over a 300-mV range of voltages and a 1,000-fold range of [SPM4+], including the plateau effect observed with voltages >100 mV. Kinetic parameters for SPM4+ fitted with Scheme 2 were J0 = 0.018 ± 0.0019, zδ = 0.99 ± 0.044 e0, and Kd = 1.28 ± 0.20 µM (Fig. 4 B and Table 4).
Kinetic parameters for SPM4+ blockade from Scheme 2
| . | Patch 5 . | Patch 6 . | Patch 7 . | Patch 8 . | Mean ± SEM . |
|---|---|---|---|---|---|
| J0 | 0.013 | 0.016 | 0.022 | 0.019 | 0.018 ± 0.0019 |
| Kd (μm) | 1.29 | 1.11 | 1.83 | 0.87 | 1.28 ± 0.20 |
| zδ (e0) | 1.08 | 1.02 | 1.00 | 0.87 | 0.99 ± 0.044 |
| . | Patch 5 . | Patch 6 . | Patch 7 . | Patch 8 . | Mean ± SEM . |
|---|---|---|---|---|---|
| J0 | 0.013 | 0.016 | 0.022 | 0.019 | 0.018 ± 0.0019 |
| Kd (μm) | 1.29 | 1.11 | 1.83 | 0.87 | 1.28 ± 0.20 |
| zδ (e0) | 1.08 | 1.02 | 1.00 | 0.87 | 0.99 ± 0.044 |
It is worth noting that there is a relatively small difference (∼50%) in the fitted values for J0 for SPD3+ and SPM4+ datasets, consistent with a free energy difference (ΔΔG) of ∼0.2 kcal/mol for this transition (RT × ln[0.018/0.027]). This small difference supports the idea that the voltage-dependent transition is intrinsic to the channel and is not likely to directly depend on the presence of polyamine. In contrast, the 20-fold difference in the channel’s affinity for SPM4+ over SPD3+ is consistent with an increased energetic stabilization of the SPM4+-blocked state of ∼1.8 kcal/mol (RT × ln[28.1/1.28]) over the SPD3+-blocked state. Thus this difference is strongly linked to the polyamine structure.
Given a relatively simple model that provides a good description of the functional properties of polyamine blockade over a wide range of conditions, we wondered what might be the structural correlates of the kinetic states, and how we might begin to account for the apparent difference in polyamine affinities. We explored this using computational methods, as described below.
Binding modes of polyamines
To gain insight toward the structural correlates of the kinetic states proposed in the blocking model, we performed molecular simulations of the MthK pore embedded in a lipid bilayer with either SPD3+ or SPM4+ in the pore. Bulk solutions in the simulations consisted of water and K+ and Cl− ions to approximate 200 mM KCl at both sides of the membrane. The crystal structure of the MthK pore (from PDB accession no. 3LDC), shown in Fig. 5, illustrates the mapping of z-axis coordinates in subsequent figures onto the pore structure. Each simulation was initiated with K+ ions in the S0, S2, and S4 positions along the SF (i.e., at t = 0 in Fig. 6, A–D), in which S0 corresponds to the partially hydrated K+ ion at the extracellular threshold of the SF coordinated by carbonyl oxygens from Y62; S2 corresponds to the K+ ion coordinated between carbonyl oxygens from G61 and V60; and S4 corresponds to the K+ ion coordinated between carbonyl and side chain oxygens from T59. The molecular system was observed to maintain its integrity over the courses of simulations with either 0 or 500 mV across the membrane, as indicated by plots of the overall RMS of the protein and water density within the system (Fig. S2 and Fig. S3).
Structure and dimensions of the MthK pore. Two of the four pore-forming subunits from the crystal structure of the MthK pore (PDB accession no. 3LDC) are shown side by side, to illustrate the transmembrane ion conduction pathway. Distances represented in the z-axis scale in subsequent computational experiments (corresponding to Figs. 5, 6, 7, 8, and 9) are shown at the left. K+ ions in the SF are represented by purple spheres, and selected main chain and side chain atoms are shown as indicated (red, oxygen; blue, nitrogen; gray, carbon).
Structure and dimensions of the MthK pore. Two of the four pore-forming subunits from the crystal structure of the MthK pore (PDB accession no. 3LDC) are shown side by side, to illustrate the transmembrane ion conduction pathway. Distances represented in the z-axis scale in subsequent computational experiments (corresponding to Figs. 5, 6, 7, 8, and 9) are shown at the left. K+ ions in the SF are represented by purple spheres, and selected main chain and side chain atoms are shown as indicated (red, oxygen; blue, nitrogen; gray, carbon).
Correlated movement of K+ ions and polyamine in the MthK pore. (A) Simulation trajectory showing the positions of the NL of SPD3+ (blue) and K+ ions in the SF (shades of purple), relative to oxygen atoms of SF residues (red, as indicated to the right) with 0 mV transmembrane voltage. (B) Simulation trajectory for SPD3+ with 500 mV transmembrane voltage. (C) Simulation trajectory for SPM4+ with 0 mV transmembrane voltage. (D) Simulation trajectory for SPM4+ with 500 mV transmembrane voltage. These data illustrate that movement of NL to positions within hydrogen-bonding distance of T59 oxygen atoms depends on outward displacement of K+ ions.
Correlated movement of K+ ions and polyamine in the MthK pore. (A) Simulation trajectory showing the positions of the NL of SPD3+ (blue) and K+ ions in the SF (shades of purple), relative to oxygen atoms of SF residues (red, as indicated to the right) with 0 mV transmembrane voltage. (B) Simulation trajectory for SPD3+ with 500 mV transmembrane voltage. (C) Simulation trajectory for SPM4+ with 0 mV transmembrane voltage. (D) Simulation trajectory for SPM4+ with 500 mV transmembrane voltage. These data illustrate that movement of NL to positions within hydrogen-bonding distance of T59 oxygen atoms depends on outward displacement of K+ ions.
Time series of RMS fluctuations for SPD3+ and SPM4+ during simulations at 0 and 500 mV. (A–D) For each condition, the RMS fluctuations varied by <1 Å over the course of the simulation.
Time series of RMS fluctuations for SPD3+ and SPM4+ during simulations at 0 and 500 mV. (A–D) For each condition, the RMS fluctuations varied by <1 Å over the course of the simulation.
Water density distributions along the z-axis for MD simulations. (A–D) Solid lines represent water density as mean number of oxygen atoms per Å3 in a 1-Å–thick layer centered around z; dashed lines represent the variance of the distribution. Water distributions within the simulation were similar under each condition, consistent with stability of the membrane. Plots were obtained using software from Giorgino (2014).
Water density distributions along the z-axis for MD simulations. (A–D) Solid lines represent water density as mean number of oxygen atoms per Å3 in a 1-Å–thick layer centered around z; dashed lines represent the variance of the distribution. Water distributions within the simulation were similar under each condition, consistent with stability of the membrane. Plots were obtained using software from Giorgino (2014).
Fig. 6 shows the positions of K+ ions in the SF (shades of purple), with the positions of oxygens from SF residues (red) and the “leading nitrogen” (NL) of the polyamine (blue) along the pore axis (z) during the time course of simulations performed with either 0 mV (Fig. 6 A) or 500 mV (Fig. 6 B) across the membrane. Under the conditions of these simulations, NL exhibited fluctuations within the vestibule (cavity) of the pore. With 0 mV, little direct interaction between SPD3+ NL and the SF oxygens is observed until the K+ ion that primarily occupied S4 moves outward to the S3 or S2 position (e.g., at t > 400 ns, Fig. 6 A). This outward K+ movement was followed by apparent interaction between NL and the side chain hydroxyl group of T59 (T59-OH), which forms the threshold of the SF. Initially NL appears to be centered at the “cytosolic” side of T59-OH (referred to as the “S5” site), although, after further outward displacement of K+ to S2 (Fig. 6 A, at t > 450 ns), NL could apparently occupy the S4 site. With 500 mV, the S4 K+ was observed to rapidly move to the S3 position (t = ∼40 ns, Fig. 6 B), resulting in less fluctuation of SPD3+ NL and greater interaction with T59-OH (at the S5 site). Qualitatively similar results were observed with SPM4+ with respect to the NL fluctuations (Fig. 6, C and D), in which there was an apparent correlation between the position of NL and occupancy of S4 by a K+ ion, and outward movement of the K+ ion was followed by outward movement of NL. As with SPD3+, movement of a K+ ion from S3 to S2 was followed by movement of NL into the S4 position in the SF.
To quantify these correlated movements, we constructed histograms of the positions of K+ ions along the pore, and then performed potential of mean force calculations of the polyamine to estimate the energetics of NL movements as a function of K+ ion position (Fig. 7). Overall, the results illustrate that NL can occupy a relatively broad energy well that corresponds to the MthK pore cavity as long as a K+ ion is coordinated in part by T59-OH. However, the energetics exhibit a shift to a narrow energy well centered on the S4 position, correlated with K+ ion occupancy of S3 (Fig. 7 A). This pattern is also observed in simulations with 500 mV; depolarization primarily acts to shift K+ occupancies, based on the histograms, whereas the free energy of NL along the MthK pore axis is correlated with the occupancies (Fig. 7 B). Similar results were observed with SPM4+ at 0 and 500 mV (Fig. 8).
Distributions of K+ and amine energetics of SPD3+ in the MthK pore. (A) Fractional occupancy of the bottommost K+ ion versus position along the pore axis from simulations with SPD3+, at 0 mV (top row). Panels in the middle row contain the free energy profiles for the NL, conditioned on the position of the bottommost K+ ion, superimposed on histogram of the corresponding K+ occupancy. Snapshots of structures corresponding to the global minima of the free energy plot are shown below; two of the MthK pore-forming subunits are shown for clarity (light blue); SF main chain, polyamine, and SF-bound water molecules are shown as sticks, and SF-bound K+ ions are shown as purple spheres. (B) Corresponding histograms, free energy profiles (middle), and structure snapshots from simulations with SPD3+, at 500 mV. Whereas conditional free energy profiles are similar with 0 or 500 mV, the bottommost K+ ion is driven from the pore cavity toward the SF with depolarization, thus driving NL toward increased occupancy at the entry to the SF, to occlude K+ permeation. Dashed lines in A and B correspond to SEM, computed by considering points from the variable sampled every 1 ns.
Distributions of K+ and amine energetics of SPD3+ in the MthK pore. (A) Fractional occupancy of the bottommost K+ ion versus position along the pore axis from simulations with SPD3+, at 0 mV (top row). Panels in the middle row contain the free energy profiles for the NL, conditioned on the position of the bottommost K+ ion, superimposed on histogram of the corresponding K+ occupancy. Snapshots of structures corresponding to the global minima of the free energy plot are shown below; two of the MthK pore-forming subunits are shown for clarity (light blue); SF main chain, polyamine, and SF-bound water molecules are shown as sticks, and SF-bound K+ ions are shown as purple spheres. (B) Corresponding histograms, free energy profiles (middle), and structure snapshots from simulations with SPD3+, at 500 mV. Whereas conditional free energy profiles are similar with 0 or 500 mV, the bottommost K+ ion is driven from the pore cavity toward the SF with depolarization, thus driving NL toward increased occupancy at the entry to the SF, to occlude K+ permeation. Dashed lines in A and B correspond to SEM, computed by considering points from the variable sampled every 1 ns.
Distributions of K+ and amine energetics of SPM4+. Histograms, free energy profiles, and structure snapshots from simulations with SPM4+, at 0 mV (A) and 500 mV (B). As with SPD3+, conditional free energy profiles with SPM4+ are similar at 0 and 500 mV; because the bottommost K+ ion is driven outward with depolarization, NL is driven to occupy a site at the entry to the SF, leading to blockade. Dashed lines on the plots indicate SEM estimates.
Distributions of K+ and amine energetics of SPM4+. Histograms, free energy profiles, and structure snapshots from simulations with SPM4+, at 0 mV (A) and 500 mV (B). As with SPD3+, conditional free energy profiles with SPM4+ are similar at 0 and 500 mV; because the bottommost K+ ion is driven outward with depolarization, NL is driven to occupy a site at the entry to the SF, leading to blockade. Dashed lines on the plots indicate SEM estimates.
Collectively, these results are consistent with the idea that voltage-dependent outward K+ movement is energetically coupled with coordination of polyamines within the SF. However, these results on their own do not seem to provide an explanation for the higher apparent affinity of SPM4+ compared with SPD3+, and they imply that the difference in affinities arises in part from interactions outside of the binding of NL within the SF.
Roles of pore-lining side chains in polyamine binding
It was previously hypothesized that the greater apparent affinity for SPM4+ over Mg2+ in blockade of BK channels arose from stabilization of SPM4+ by a ring of negatively charged glutamate side chains (E321 and E324) at the cytosolic end of the BK channel pore (Zhang et al. 2006). MthK also contains glutamate side chains (E92 and E96) at the cytosolic end of the pore. Mutation of these glutamates individually, however, results in channels with greatly reduced open probability (Parfenova et al. 2006; Shi et al. 2011).
To explore the possible roles of interactions with E92 and E96 that may contribute to polyamine binding energetics, we plotted the positions of all polyamine nitrogens along with the positions of the side chain oxygens of T59 (T59-OH, at the entry to the SF), and with the side chain oxygens of E92 and E96, during the simulation time courses (Fig. 9). Although the side chains of E92 and E96 for each of the four MthK subunits are close to the central axis of the pore, their distances from one another appear to be sufficient such that their pKa values were not shifted by carboxylate–carboxylate interactions, and simulations were performed with side chain oxygens deprotonated (and thus one carboxylate oxygen from each side chain having a charge of −1). This is supported by calculations using DelPhiPKa based on the 3LDC atomic coordinates (Wang et al. 2016; Pahari et al. 2018). From these plots, one can observe that when SPD3+-N1 is coordinated by T59-OH, SPD3+-N3 may be coordinated by an E92 side chain oxygen. However, when SPM4+-N1 is coordinated by T59-OH, SPM4+-N3 may be coordinated an E92 side chain oxygen, in addition to coordination of SPM4+-N4 by side chain oxygens from either E92 or E96. The likelihood of overlap between the amine and carboxylates is further illustrated in histograms derived from the trajectories (Fig. 10). Thus in our working hypothesis, the structure of SPM4+, along with its additional charged amine group, may form key electrostatic interactions that augment its binding energetics within the MthK channel.
Distinct SPD3+ and SPM4+ coordination at the cytosolic end of the MthK pore. Positions of each polyamine nitrogen (in shades of blue) for SPD3+ (top panels) and SPM4+ (bottom panels) as a function of time, superimposed on positions of glutamate side chain oxygens (corresponding to centers of mass for the pairs of oxygens) and hydroxyl oxygen of T59 are shown in shades of red. With strong depolarization (panels at right), SPD3+ N atoms may simultaneously interact with T59-OH and E92, whereas SPM4+ N atoms may interact additionally with E96.
Distinct SPD3+ and SPM4+ coordination at the cytosolic end of the MthK pore. Positions of each polyamine nitrogen (in shades of blue) for SPD3+ (top panels) and SPM4+ (bottom panels) as a function of time, superimposed on positions of glutamate side chain oxygens (corresponding to centers of mass for the pairs of oxygens) and hydroxyl oxygen of T59 are shown in shades of red. With strong depolarization (panels at right), SPD3+ N atoms may simultaneously interact with T59-OH and E92, whereas SPM4+ N atoms may interact additionally with E96.
Distributions of polyamine nitrogen atoms and glutamate side chain oxygen atoms at the cytosolic end of the MthK pore, when NL is interacting with T59-OH. Atom occupancy histograms corresponding to the trajectories in Fig. 8, with polyamine and voltage as indicated. These illustrate that SPD3+ is typically within hydrogen bonding distance of only E92 at either 0 mV (top left) or 500 mV (top right), based on differences in the peak positions (<3 Å). In contrast, SPM4+ is typically within hydrogen bonding distance of both E92 and E96 at either 0 mV (bottom left) or 500 mV (bottom right). These results are consistent with electrostatic bonds between MthK pore-lining side chains and SPM4+ that are greater than those with SPD3+. Dashed lines indicate error estimates.
Distributions of polyamine nitrogen atoms and glutamate side chain oxygen atoms at the cytosolic end of the MthK pore, when NL is interacting with T59-OH. Atom occupancy histograms corresponding to the trajectories in Fig. 8, with polyamine and voltage as indicated. These illustrate that SPD3+ is typically within hydrogen bonding distance of only E92 at either 0 mV (top left) or 500 mV (top right), based on differences in the peak positions (<3 Å). In contrast, SPM4+ is typically within hydrogen bonding distance of both E92 and E96 at either 0 mV (bottom left) or 500 mV (bottom right). These results are consistent with electrostatic bonds between MthK pore-lining side chains and SPM4+ that are greater than those with SPD3+. Dashed lines indicate error estimates.
Discussion
In this work, we sought to better understand the structural basis for interactions between polyamines and K+ channels using the model K+ channel, MthK. Although little is known of how MthK is gated under native conditions, polyamines are thought to be nearly ubiquitous among archaea, including M. thermoautotrophicum (with ∼250 µM SPD3+ and 500 µM SPM4+; Hamana et al., 2007). Whereas polyamines are known to be critical for life in many organisms primarily because of their role in stabilizing nucleic acids, these molecules are also well known to interact with ion channels and are especially important in the functional role of Kir channels in stabilizing membrane potential. The relatively high apparent affinity of MthK for SPD3+ and SPM4+ would thus be consistent with a physiological role for MthK in M. thermoautotrophicum that may be similar to the role of a Kir channel in eukaryotes, although MthK appears to be additionally gated by divalent ions and modulated by protons (Jiang et al., 2002; Zadek and Nimigean, 2006; Li et al., 2007; Pau et al., 2010, 2011; Smith et al., 2013; Posson et al., 2015).
We have analyzed polyamine blockade using kinetic models, including a two-state model in which voltage and concentration dependence are defined by a single voltage-dependent equilibrium constant (Scheme 1). Although Scheme 1 may be adequate to describe blockade over a limited range of depolarized voltages, it fails to capture blocking kinetics over a wide range of voltages, where it becomes clear that blockade reaches a plateau at depolarized voltages (Fig. 3). For this reason, we explored slightly more complex models and found that a relatively simple three-state model, Scheme 2, can account for polyamine blocking kinetics over a 300-mV range of transmembrane voltage (Fig. 4). Whereas Scheme 2 provides a good description of the fractional MthK currents over a wide range of conditions, it should be noted that the model predictions deviate from the experimental data at low [SPM4+] (<1 µM), indicating either that Scheme 2 may be too simple to account for blocking interactions under these conditions, or possibly that the free [SPM4+] may differ slightly from the total [SPM4+] under the conditions of our experiments.
Scheme 2 contains a transition that precedes blockade, defined by a voltage-dependent equilibrium constant, followed by an transition from the open to the blocked state, which depends only on blocker concentration. In physical terms, the voltage-dependent O-O’ transition in Scheme 2 may correspond to movement of K+ ions within the pore while the channel is in a conducting state. In the absence of polyamine, voltage would drive the O-O’ transition, resulting in unobstructed permeation. In the presence of polyamine, however, the channel is driven from O’ to B. In this sense, O’ might be viewed more specifically as being in a configuration with the S4 site unoccupied by K+, and thus receptive to polyamine blockade. It is important to note, however, that even with polyamine occupying the cavity region of the MthK pore, K+ ions may enter the pore from the bulk solution (Fig. 6 and Fig. S4). If complete dissociation of the polyamine from the channel is not obligatory for current flow, then this may account in part for the observation of incomplete fractional blockade in the presence of relative high polyamine concentrations, with moderately depolarized voltages (Fig. 4 A, 200–1,000 µM SPD3+).
Representative snapshots of MthK, ions, and all water molecules within 3 Å of the SF, during an MD trajectory. (A) Time series of the simulation with SPD3+ at 0 mV from Fig. 6 A. (B) SPD3+ with NL coordinated by T59-OH groups at the entry to the SF. (C) SPD3+ released from the SF, followed by a water molecule entering the SF. (D) Entry of a K+ ion from the bulk solution into the pore, past a SPD3+ molecule. (E) Entry of the K+ into the SF. Thus under these conditions, ions and water can enter the pore and SF with SPD3+ in the pore.
Representative snapshots of MthK, ions, and all water molecules within 3 Å of the SF, during an MD trajectory. (A) Time series of the simulation with SPD3+ at 0 mV from Fig. 6 A. (B) SPD3+ with NL coordinated by T59-OH groups at the entry to the SF. (C) SPD3+ released from the SF, followed by a water molecule entering the SF. (D) Entry of a K+ ion from the bulk solution into the pore, past a SPD3+ molecule. (E) Entry of the K+ into the SF. Thus under these conditions, ions and water can enter the pore and SF with SPD3+ in the pore.
Scheme 2 indicates a valence of ∼1 e0 to describe the voltage dependence of polyamine blockade in MthK. In eukaryotic strongly Kir channels (such as Kir2.1), polyamine blockade exhibits a steep voltage dependence with a valence of up to four elementary charges, which has been attributed to either direct (slow) permeation of the polyamine through the pore or a role of the polyamine in displacing up to four K+ ions from the pore during blockade, or possibly some combination of polyamine and K+ permeation (Lopatin et al., 1995; Guo and Lu, 2000b; Kurata et al., 2004; Shin and Lu, 2005; Kucheryavykh et al., 2007). In our computational studies of MthK, we have not observed polyamine permeation through the SF beyond entry of NL into the S4 site of the SF (Fig. 7). Although we cannot entirely rule out a polyamine permeation mechanism, our computational studies indicate that outward K+ movement is a prerequisite for NL binding to T59-OH groups and subsequent entry to the S4 site (Figs. 8 and 9, and illustrated in Fig. S5). Crystal structures of a KirBac1.3-Kir3.1 chimera, as well as Kir2.2 and Kir3.1, are consistent with the idea that the “long pore” common within the Kir family results in alignment of as many as seven ions along the permeation pathway (Nishida et al., 2007; Tao et al., 2009; Xu et al., 2009). Just as the outward displacement of several ions may underlie steep voltage dependence for polyamine blockade in Kir channels, the net displacement of one ion across the SF provides a plausible explanation for the valence of 1 e0 observed with MthK.
Illustration of the scheme for charge movement coupled to polyamine binding at the entry to the SF. (A) K+ ions (purple spheres) occupy the canonical S1 and S3 sites within the SF, coordinated by carbonyl oxygen atoms (from Y62-G61, and V60-T59), as well as the S5 site, in which a partially hydrated K+ ion can be coordinated by side chain oxygens from T59. A hydrated polyamine (blue) can occupy the cavity, electrostatically repelled from the SF by the K+ ion at S5. (B) K+ ions occupying more extracellular S2 and S4 positions, coordinated by oxygen atoms (carbonyls from G61-V60, and carbonyl/side chain from T59-T59OH), as well as the S0 site, in which a partially hydrated K+ ion can be coordinated by carbonyl oxygens from Y62. The polyamine remains electrostatically repelled from the SF by the K+ ion at S4. (C) Further extracellular movement of K+ (to S1/S3) permits coordination of the polyamine by T59-OH, with dissociation of K+ to the bulk solution. Thus, polyamine blockade is coupled to net translocation of one electronic charge through the SF.
Illustration of the scheme for charge movement coupled to polyamine binding at the entry to the SF. (A) K+ ions (purple spheres) occupy the canonical S1 and S3 sites within the SF, coordinated by carbonyl oxygen atoms (from Y62-G61, and V60-T59), as well as the S5 site, in which a partially hydrated K+ ion can be coordinated by side chain oxygens from T59. A hydrated polyamine (blue) can occupy the cavity, electrostatically repelled from the SF by the K+ ion at S5. (B) K+ ions occupying more extracellular S2 and S4 positions, coordinated by oxygen atoms (carbonyls from G61-V60, and carbonyl/side chain from T59-T59OH), as well as the S0 site, in which a partially hydrated K+ ion can be coordinated by carbonyl oxygens from Y62. The polyamine remains electrostatically repelled from the SF by the K+ ion at S4. (C) Further extracellular movement of K+ (to S1/S3) permits coordination of the polyamine by T59-OH, with dissociation of K+ to the bulk solution. Thus, polyamine blockade is coupled to net translocation of one electronic charge through the SF.
In terms of energetics, the fitted parameters for Scheme 2 suggest that the O-O’ transition is similar for channels with SPD3+ or SPM4+, with a free energy difference of 0.2 kcal/mol. This supports the idea that the voltage-dependent transition is largely intrinsic to the channel and does not depend strongly on the species of polyamine present, but does not rule out a small contribution of the polyamine to the energetics of the transition. In contrast, the transition from O’ to B in Scheme 2 depends strongly on the species of polyamine, and the free energy difference between SPD3+ and SPM4+ of 1.8 kcal/mol for this transition is consistent with the observation of additional electrostatic interactions that are permitted with SPM4+ in the MthK pore compared with SPD3+ (Figs. 9 and 10).
In a broader sense, however, the transition from O’ to B in Scheme 2 must be viewed as a collective transition, as it must incorporate the movement of polyamine from the bulk solution into the pore, followed by movement to a high-affinity binding site. Likewise, the transition from O to O’ must be considered as a collection of all possible transitions that lead to the polyamine-receptive conformation.
Bacteria and other prokaryotes are known to synthesize polyamines (Morris and Jorstad, 1973; Tabor and Tabor, 1976; Michael, 2018). KirBac1.1, which is among the closest bacterial relatives to eukaryotic Kir channels, is observed to be insensitive to SPM4+, whereas the pore mutation I138D can impart SPM4+ sensitivity in the channel, which along with studies of eukaryotic Kir and BK channels as well as the present study, is consistent with the importance of pore electrostatic interactions in potentially determining polyamine affinity (Kurata et al., 2006; Zhang et al., 2006; Cheng et al., 2009). In the present study, we have performed analysis of polyamine blockade in a relatively simple archaeal system. This system, in turn, may be further developed to account quantitatively for the microscopic steps in polyamine blockade, and also be compared with similar phenomena that occur in more complex bacterial and mammalian systems.
Acknowledgments
Merritt C. Maduke served as editor.
This work was supported by grants from the National Science Foundation (MCB-1243803 to B.S. Rothberg) and the National Institutes of Health (R01 GM126581 to B.S. Rothberg and R01 GM093290 to V. Carnevale). This research includes calculations carried out on Temple University's High Performance Computing resources and thus was supported in part by the National Science Foundation through major research instrumentation grant number 1625061, by the Army Research Laboratory under contract number W911NF-16-2-0189, and by the National Institutes of Health through grant S10 OD020095.
The authors declare no competing financial interests.
Author contributions: A. Suma designed experiments, acquired and analyzed data, and assisted with manuscript preparation. D. Granata assisted with acquiring and analyzing data. A.S. Thomson acquired and analyzed data and assisted with manuscript preparation. V. Carnevale supervised the project, designed experiments, analyzed data, and reviewed and edited the manuscript. B.S. Rothberg supervised the project, designed experiments, analyzed data, wrote the first draft of the manuscript, and reviewed and edited the manuscript.
References
Author notes
V.Carnevale and B.S. Rothberg contributed equally to this paper.
A.S. Thomson’s current address is GlaxoSmithKline, Upper Merion, PA.


![MthK channel blockade by SPD3+.(A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPD3+] (≤1,000 µM) using a voltage-ramp protocol and normalized to the current amplitude at −100 mV for the same bilayer with 0 SPD3+. (B) Fractional current versus [SPD3+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages (parameters are in Table S1). (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 860 ± 20 µM, zδ = 0.67 ± 0.03 e0). Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/7/10.1085_jgp.201912527/3/m_jgp_201912527_fig1.png?Expires=1779282171&Signature=GTlMbve-BZDM2LFkJG0phy9FbJw8YGKqzWQdhwzb8co~XVD8iDK5zjbeOQDjiWwmA69n1eQKnJwJItNIsrQsZbThfe~IecFdiTpvSDmUxTaspZoelM5ODOYdqG642D82IHsKX~UrvhjnmGDJSXhB2dsO7fLFVd4PzJhC7pekZftRUeQed8ZCkygnS8xoBFxsbQAd~yNETLApjEpUhJtTE0EPjV3JVyt9TQ-HYZHFP6j1pwS~W-5iai3EE1JRlLWGVJRBT8CAyLL~0ZDeREfdt7rbA22taVngzEZIuMYWRSJqJrCUCBwwBV0o9aAhr4l4M-1CB1gyLSvRveeP-tgCMw__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![MthK channel blockade by SPM4+.(A) Mean normalized macroscopic MthK currents. Currents were obtained over a range of [SPM4+] (≤100 µM) using a voltage-ramp protocol and normalized as described in Fig. 1. (B) Fractional current versus [SPM4+] plotted at five representative voltages. Data were fitted with Hill equations (smooth curves) to estimate Kapp over a range of voltages. (C)Kapp versus voltage, fitted with Eq. 1 (solid blue line; Kapp(0) = 77 ± 6 µM, zδ = 0.88 ± 0.05 e0). Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/7/10.1085_jgp.201912527/3/m_jgp_201912527_fig2.png?Expires=1779282171&Signature=fKAmRXPIgmF-OQ8Q~qoHnM9puCRlSf72jeh3aHPGn1N4pqtKawdZSVVtbyBW0u7u3mdCzNeYEYuwSROEJjF~t7PGdNm0KH3e3nSBqH1SQ2Xuj29Y7zUFlRSC3B6J6UQCKNwbA5s3msLXvXoJ9fV6J4LP0adcsV-ptM7E2bqqMFU3yIkeFbtrrjjoVF3i5cj~YdG9SS2dt3tgSLuLETAv429RYFN-ms608NnBDdoCklFFUPxHeY3D5dyI8YhaCJtGB9e6CeeOOkXtyeBh44-LglbtwAj1nB6UElNgGpDjWfGX~BGiWjloGoZ~-R0xBHe6dUzDb~BPuVtCf6R9dx9SDg__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)

![SPD3+ and SPD4+ blockade described withScheme 1.(A) Mean fractional MthK current over a range of [SPD3+], with Scheme 1 predictions superimposed (solid red curves). This model predicts that MthK currents approach 100% blockade with very strong depolarization, and thus provides a poor description of the data over the observed range of voltage and [SPD3+]. (B) Mean fractional MthK current over a range of [SPM4+], with Scheme 1 predictions superimposed. As with SPD3+, Scheme 1 predicts that MthK currents approach 100% blockade with very strong depolarization and is thus inconsistent with the data over the observed range of voltage and [SPM4+]. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/7/10.1085_jgp.201912527/3/m_jgp_201912527_fig3.png?Expires=1779282171&Signature=LrQpW7uMTj4-s6Wiegtfsypt~RwGzAk-8UXI8PanfYMRsRHdYxNXfVzVcsiqNlk5-s01by9ZD4IQStuY7pyHN6Zfw~kwITYUyRtGXHBZ6O4dMTysLiiCrRG3DjVzJ4arOajV8I1tsJoh2pye4vBNL4ZO0~AJiK~6rBO6J6RKa-BJ4liTeGHJmtzGkYGdXQJ50nYqXxgf1DN526FYRYsIJKTAyrer4DQQcstwBUY9MN1JrdqSVkXD0iz5KR99zC90fFMuCy1eepWx4AZfMOUhAUQt6VPuN9hMyVLQq~cV-wQugalI5AaA-sCSj6MgACZjdE62AE05TcBX~gfanoNk1g__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![SPD3+ and SPM4+ blockade described withScheme 2.(A) Fractional MthK current over a range of [SPD3+] with Scheme 2 predictions superimposed. Scheme 2 predicts that depolarization drives the channel toward a receptive open state (O’) with enhanced blocker affinity, accounting well for the experimental data. Fit parameters for individual experiments are provided in Table 3. (B) Fractional MthK current over a range of [SPM4+] with Scheme 2 predictions superimposed. Scheme 2 accounts for the major features of SPM4+ blockade over a 1,000-fold range of [SPM4+]. Fit parameters for individual experiments are provided in Table 4. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/7/10.1085_jgp.201912527/3/m_jgp_201912527_fig4.png?Expires=1779282171&Signature=dMzIBV0BP3loEvbDPaEZRFZ3NPtHhcz~TUBIhErPlWugrBCzPSJUP9M24BxPzRkPz1VcTpM-0rztUTpcYQV6siKXJFVI4IPbWf4jul0hGGhDmA4r011jkcd6Jfn6PgJXWZDhnPDFPxxH1Vw2i9uUgew4Ze0Z9Jydq07qwbtapqTtBY9tEWpekzyWRRJGC4YKYsgYGyrX8-a0iSzE9Me690zxQjuOtpWCaY44XmEj3BEVvR0pjUQvn~376nXOnE2q0okc6mCSq9MSGenc0U6A81ysggiwFt8cAp2Jyvx6u4s39XZb4~KmIEHN6Dhb8SRVqRgbJPdrsfGUWo1iwPh46Q__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)

![SPD3+ blockade described with an alternative three-state model (Scheme 4). Mean fractional MthK current (I/I0) over a range of [SPD3+], with Scheme 4 predictions superimposed (solid red curves). (A) Scheme 4 (left), with equations describing fractional current (right). (B) Mean fractional MthK current (I/I0) over a range of [SPD3+] as indicated, with Scheme 4 predictions superimposed (solid red curves). The best fit with this model (χ2 = 35.4) predicts that MthK currents will ultimately reach 100% blockade with very strong depolarization, although the apparent voltage dependence of blockade is less steep than that of Scheme 1. (C) An alternative fit representing a local minimum (χ2 = 97.6) predicts that MthK currents can increase with very strong depolarization. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jgp/152/7/10.1085_jgp.201912527/3/m_jgp_201912527_figs1.png?Expires=1779282171&Signature=g7pf075tGdKefWhm0-ycJFcreAVZalpdjEGOMsvurv5R2x0dsv7LMB616chB68~4RAzLuPcGgWQ06hJyrUsHG~oklRN32432GWEmll25l9yXLz7wKr16XJw~EvwqxjRAkt6vjlBRNg6xc~jFBZSqEJC2QzFcuK6NES2LP1NuqQWW99ctImkSpnI66K~gB5lfzQkEcU6Xh0LMh8la5Qb84o4-TWcdfxMMw3FVso2mp1fZUfa1dD~Iv3eDO4cK~rDTNrVoo9niS~jMBtJjhTZoZAciUnjyI4KX6yT8u14NJmF4HDmtlg9h0cMQSHyoIbWIVXjs9i3Sp3ThP98xQYcG0A__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)












