Volume electron microscopy is an important imaging modality in contemporary cell biology. Identification of intracellular structures is a laborious process limiting the effective use of this potentially powerful tool. We resolved this bottleneck with automated segmentation of intracellular substructures in electron microscopy (ASEM), a new pipeline to train a convolutional neural network to detect structures of a wide range in size and complexity. We obtained dedicated models for each structure based on a small number of sparsely annotated ground truth images from only one or two cells. Model generalization was improved with a rapid, computationally effective strategy to refine a trained model by including a few additional annotations. We identified mitochondria, Golgi apparatus, endoplasmic reticulum, nuclear pore complexes, caveolae, clathrin-coated pits, and vesicles imaged by focused ion beam scanning electron microscopy. We uncovered a wide range of membrane–nuclear pore diameters within a single cell and derived morphological metrics from clathrin-coated pits and vesicles, consistent with the classical constant-growth assembly model.
Introduction
Three-dimensional, high-resolution imaging provides a snapshot of the internal organization of a cell at a defined time point and in a defined physiological state. Focused ion beam scanning electron microscopy (FIB-SEM) yields nearly isotropic, nanometer-level resolution, and three-dimensional images by sequential imaging of the surface layer of a sample, which is then etched away with an ion beam to reveal the layer beneath (Knott et al., 2008; Xu et al., 2017). FIB-SEM technology continues to develop, and it can be a particularly valuable contemporary tool for imaging the complete volume of a cell, but segmentation of the three-dimensional datasets and subsequent analysis of the results are substantial hurdles as the images are far too large to interpret by inspection (Heinrich et al., 2021).
The widespread success of machine learning in bioimage analysis has recently inspired the application of deep learning approaches to automated segmentation. Examples using deep convolutional networks for data with anisotropic resolution include DeepEM3D (Zeng et al., 2017) and CDeep3M (Haberl et al., 2018), for segmentation of mitochondria and Golgi apparatus with extensive post-processing (Žerovnik Mekuč et al., 2020; Žerovnik Mekuč et al., 2022), as well as cell organelle segmentation in quasi-isotropic FIB-SEM data of beta cells (Müller et al., 2021). CDeep3M was the first project using cloud computing and achieves good results on mostly large-size organelles or clusters of smaller-size vesicles. However, it was conceived for anisotropic data and therefore has conceptual limitations when applied to 3D isotropic FIBSEM data. A pipeline created by the COSEM project (Heinrich et al., 2021) enables automated whole-cell simultaneous segmentation of up to 35 organelles from relatively sparse but very precise 3D ground truth annotations from FIB-SEM data of cells prepared by high-pressure freezing and freeze substitution (HPFS), obtained at 3–5 nm voxel size with approximately isotropic resolution. The most common strategy used by the COSEM project involved training with ground truth annotations from multiple classes of objects at the same time, typically at a high computational cost (500,000 or more training iterations; Heinrich et al., 2021).
The current approaches all suffer from a demand for substantial computational resources, and they generally require a large set of precise manual annotations. Both requirements limit their practical applicability. We describe here the development and use of a new deep-learning pipeline called automated segmentation of intracellular substructures in electron microscopy (ASEM), which can detect structures of a wide range in size and complexity using deep neural networks trained on a limited number of loosely marked, i.e., not necessarily pixel-precise, ground truth annotations whose object boundaries could be off by 1–2 voxels. ASEM includes a semiautomated graph-cut procedure we developed to assist in the tedious task of ground truth preparation and a computationally efficient transfer-learning approach with a fine-tuning protocol that can be used without the need for high-end specialized CPU/GPU workstations.
We illustrate here the utility of ASEM by describing the results of its application to data from several types of cells, including FIB-SEM datasets made publicly available by the COSEM Project (Heinrich et al., 2021). We note that while cellular samples have traditionally been processed by chemical fixation (CF) and staining at room temperature, HPFS at cryogenic temperatures (as was the case for the COSEM Project data) yields a substantial increase in the preservation of many complex cellular features. We applied ASEM to three-dimensional FIB-SEM images of cells prepared by either CF or HPFS. We validated our approach by segmenting mitochondria, ER, and Golgi apparatus, as these organelles had been studied previously in similar efforts (Žerovnik Mekuč et al., 2020; Žerovnik Mekuč et al., 2022; Heinrich et al., 2021; Liu et al., 2020), and then used ASEM to recognize much smaller structures, nuclear pores, and clathrin-coated pits and vesicles. For nuclear pores in interphase, we can segment nearly all the pores in the nuclear membrane. We can therefore directly analyze the range of membrane-pore diameters, even for a single cell in a particular physiological state. For clathrin-coated pits, we show that a relatively restricted training set leads to an accurate segmentation of coated pits at all stages of their maturation as well as coated vesicles, the final step after fission from the originating membrane, and we can derive morphological metrics consistent with the classical constant-growth assembly model (Ehrlich et al., 2004; Kirchhausen, 1993; Kirchhausen, 2009; Willy et al., 2021).
All datasets (https://open.quiltdata.com/b/asem-project), models, and code (https://github.com/kirchhausenlab/incasem) are open-source so that other users working with images acquired with the same or somewhat different imaging conditions can generate their own predictive models and benefit from our pretrained models, either directly or by adapting them by fine-tuning, without the need for specialized CPU/GPU workstations.
Results
FIB-SEM imaging of cells
We obtained three-dimensional focused ion beam scanning electron microscopy (FIB-SEM) datasets for different types of adherent mammalian cells grown in culture (Table S1). The samples we imaged were prepared either by conventional chemical fixation and staining with osmium and uranyl acetate at room temperature (CF) or by fixation and similar staining at very low temperatures using high-pressure freezing and freeze substitution (HPFS), a protocol that substantially increases sample preservation (Hoffman et al., 2020; Studer et al., 2008; Xu et al., 2021). To image the volume of a cell, we used a block-face crossbeam FIB-SEM with a nominal isotropic resolution of 5 or 10 nm per voxel; each image stack, obtained during 1–2 d of continuous FIB-SEM operation, was about 15–20 GB in size, contained ∼2,000 registered sequential TIFF files, and spanned a volume of roughly 2,0003 voxels corresponding to large parts of each cell. These volume datasets were used to train the deep learning pipeline for automated segmentation of intracellular structures and to explore the effects of different fixation and staining procedures on the outcome of the segmentation tasks.
We also tested the performance of our deep learning models with a small number of FIB-SEM images from HPFS preparations of complete cells (Xu et al., 2021), obtained from the publicly available OpenOrganelle initiative (Heinrich et al., 2021; Xu et al., 2021; Table S1). They were acquired by the COSEM team at the Janelia Research Campus at a nominal resolution of 4 × 4 × 3–5 nm per voxel with a custom-modified FIB-SEM as part of their concurrent efforts to develop a methodology for automated organelle segmentation aided by deep learning.
As described below, using the specific models generated with our deep learning pipeline (Fig. 1), we could reliably identify intracellular structures ranging in size and complexity from mitochondria, ER, and Golgi apparatus to nuclear pores, clathrin-coated pits, clathrin-coated vesicles, and caveolae.
Pipelines used for training and deep-learning neural network prediction. Schematic representation of the deep-learning approach for recognizing intracellular structures in FIB-SEM volume images using a 3D U-net encoder-decoder neural network. (A) For training, three-dimensional stacks containing FIB-SEM data, augmented as described in Materials and methods, are provided as input images to the 3D U-Net; in this example, the stack includes a limited number of three-dimensional ground truth annotations for the ER in the form of binary masks (yellow). The ER predicted by the 3D U-net model is a 3D probability map, whose error is calculated by comparing the ground truth annotation with the cross-entropy loss. The model parameters are iteratively updated during training until convergence of the cross-entropy loss is achieved. (B) For prediction, small 3D stacks with data not used for training covering the complete FIB-SEM volume image of a naïve cell (or from the remaining regions of the cell used for training) are provided as input to the 3D U-net model trained in A. In this example, the predicted ER is a thresholded 3D probability map for the entire cell volume.
Pipelines used for training and deep-learning neural network prediction. Schematic representation of the deep-learning approach for recognizing intracellular structures in FIB-SEM volume images using a 3D U-net encoder-decoder neural network. (A) For training, three-dimensional stacks containing FIB-SEM data, augmented as described in Materials and methods, are provided as input images to the 3D U-Net; in this example, the stack includes a limited number of three-dimensional ground truth annotations for the ER in the form of binary masks (yellow). The ER predicted by the 3D U-net model is a 3D probability map, whose error is calculated by comparing the ground truth annotation with the cross-entropy loss. The model parameters are iteratively updated during training until convergence of the cross-entropy loss is achieved. (B) For prediction, small 3D stacks with data not used for training covering the complete FIB-SEM volume image of a naïve cell (or from the remaining regions of the cell used for training) are provided as input to the 3D U-net model trained in A. In this example, the predicted ER is a thresholded 3D probability map for the entire cell volume.
Ground truth annotation
The first step in most common machine learning segmentation procedures is to create pixelwise “ground truth” annotations—to be used for training a specific segmentation model. In the present work, we used a modest number of manually annotated segmentations of the intracellular structure of interest (see Materials and methods for details). These segmentations came from arbitrarily chosen, diverse regions from one or more cells (Table S2; Shorten and Khoshgoftaar, 2019).
We obtained ground truth annotations for mitochondria and Golgi apparatus, portions of the ER, 19 endocytic clathrin-coated pits at the plasma membrane, and 10 nuclear pores on the nuclear envelope (Table S4).
The choice of annotation tool for a given organelle was empirically determined to minimize the total time (semiautomated analysis/editing) required. We found that available tools like Ilastik (Berg et al., 2019) and Volume Annotation and Segmentation Tool (VAST; Berger et al., 2018) were sufficient for simpler structures like mitochondria, nuclear pores, and clathrin-coated structures. For more complicated structures like Golgi and ER, they, however, were less suitable, necessitating the development of a dedicated annotation tool (see below). We annotated mitochondria using the carving module in Ilastik (Berg et al., 2019), and if required, edited the annotation manually using VAST (Berger et al., 2018), a volume annotation and segmentation tool for manual and semiautomatic labeling of large 3D image stacks (see example in Fig. S1 and Video 1). We annotated the more complex Golgi apparatus and ER with a custom graph-cut-assisted, semiautomated annotation tool, which we developed and have described in the Materials and methods, that accelerated the annotation time by 5- to 10-fold; when needed, we corrected the annotation locally with VAST (see example in Fig. S2). We generated manually, also with VAST, the ground truth annotations for nuclear pores (Fig. 6 A) and clathrin-coated structures (Fig. 7 A).
Ground truth annotation workflow for mitochondria. (A) Example to illustrate the sequential steps used with Ilastik Carving module to generate the ground truth annotation for a mitochondrion in Cell 1 HEK293A prepared by chemical fixation and visualized with ∼5 nm isotropic resolution. Coarse annotations for background (yellow) and object (blue) drawn in broadly spaced consecutive planes of the stack were used to seed the Ilastik Carving module from which a binary mask spaced along adjacent planes spaced 5 nm in the z-stack and corresponding to the mitochondria ground annotation was generated (magenta). Manual corrections using VAST are used as needed, to remove incorrectly assigned pixels, in this example corresponding to an adjacent ER (white arrow). Scale bar, 500 nm. (B) Volume rendering corresponding to the ground truth annotation of the mitochondrion shown in A.
Ground truth annotation workflow for mitochondria. (A) Example to illustrate the sequential steps used with Ilastik Carving module to generate the ground truth annotation for a mitochondrion in Cell 1 HEK293A prepared by chemical fixation and visualized with ∼5 nm isotropic resolution. Coarse annotations for background (yellow) and object (blue) drawn in broadly spaced consecutive planes of the stack were used to seed the Ilastik Carving module from which a binary mask spaced along adjacent planes spaced 5 nm in the z-stack and corresponding to the mitochondria ground annotation was generated (magenta). Manual corrections using VAST are used as needed, to remove incorrectly assigned pixels, in this example corresponding to an adjacent ER (white arrow). Scale bar, 500 nm. (B) Volume rendering corresponding to the ground truth annotation of the mitochondrion shown in A.
Ground truth annotations for mitochondria, ER, and Golgi apparatus. Passing through a FIB-SEM volume with contrast equalized using CLAHE. Image is from Cell 1 HEK293A prepared by CF and imaged at ∼5 nm isotropic resolution. The video shows ground truth annotations for mitochondria (cyan), ER (red), and Golgi apparatus (green). The annotations were generated for all mitochondria and Golgi apparatus within the FIB-SEM volume, and all ER within the highlighted 8 × 3 × 3 µm block (orange box).
Ground truth annotations for mitochondria, ER, and Golgi apparatus. Passing through a FIB-SEM volume with contrast equalized using CLAHE. Image is from Cell 1 HEK293A prepared by CF and imaged at ∼5 nm isotropic resolution. The video shows ground truth annotations for mitochondria (cyan), ER (red), and Golgi apparatus (green). The annotations were generated for all mitochondria and Golgi apparatus within the FIB-SEM volume, and all ER within the highlighted 8 × 3 × 3 µm block (orange box).
Ground truth annotation workflow for ER and Golgi apparatus. (A and B) Example of graph-cut assisted segmentation used to generate the ground truth annotation for ER (A) or Golgi apparatus (B) in Cell 1 HEK293A prepared by chemical fixation and visualized with ∼5 nm isotropic resolution. Coarse annotations for background (lines, solid areas in pink) and object (dotted lines in yellow) drawn in the indicated broadly spaced planes of the stack were used as seeds to generate the ground truth annotations with the graph-cut assisted segmentation program.
Ground truth annotation workflow for ER and Golgi apparatus. (A and B) Example of graph-cut assisted segmentation used to generate the ground truth annotation for ER (A) or Golgi apparatus (B) in Cell 1 HEK293A prepared by chemical fixation and visualized with ∼5 nm isotropic resolution. Coarse annotations for background (lines, solid areas in pink) and object (dotted lines in yellow) drawn in the indicated broadly spaced planes of the stack were used as seeds to generate the ground truth annotations with the graph-cut assisted segmentation program.
To increase the number of ground truth annotations, we applied randomized voxel-wise as well as geometric transformations to each of the manual segmentations (see Materials and methods and Table S3). Such data augmentation is common for training deep neural networks (Shorten and Khoshgoftaar, 2019) and was crucial for our raw FIB-SEM images since the contrast and textural appearance can vary substantially based on sample preparation and imaging conditions.
Deep-learning segmentation pipeline
Our general training strategy (schematically represented in Fig. 1 A) relied on a 3D convolutional neural network (CNN) architecture based on a 3D U-Net (Çiçek et al., 2016; Fig. S3 A); this approach has been used previously for segmenting intracellular structures in electron microscopy data (Guay et al., 2021; Heinrich et al., 2021; Wei et al., 2020). For each organelle class, we used a single, dedicated deep neural network, trained on augmented ground truth annotations generated from a small number of annotations contained within subvolumes (∼2–80 µm3) of the FIB-SEM data (Table S4). During training, we used binary cross entropy as a loss function. Overfitting was avoided by validating the evolution of the model periodically during the training session by monitoring the loss between the model prediction and the subset of ground truth annotations in validation blocks of the FIB-SEM image not used for training. Each model was trained until the validation loss converged to a stable value (Fig. S3, B–D), which corresponds to roughly 100,000 iterations on a single GPU (∼23 h). The final model yielded a predicted map that assigned to each voxel a probability of belonging to the structure (Fig. 1 B), from which we derived a final binary map by setting a threshold value of 0.5. These models, unique for a given organelle or structure, were then used to find the specific cellular structure of interest in the FIB-SEM images of regions excluded from training or of “naïve” cells that had not been used for training at all.
3D U-net architecture, examples of network behavior during training, and F1 as a metric to compare ground truth annotations with model predictions. (A) Schematic representation of the steps used to train the 3D U-net encoder-decoder neural network. The input for the neural network mode are 3D blocks consisting of a stack of consecutive FIB-SEM images (size 204 × 204 × 204 voxels). The 3D block is subjected in the encoder to three cycles, each consisting of consecutive 3 × 3 × 3 convolutions without padding (purple) and downsampling operators with 2 × 2 × 2 max-pooling (pink). The feature maps from the encoder are then upsampled in the decoder by a factor of 2 (yellow), followed by concatenation with previous feature maps from the downsampling branch that had been exposed to central cropping and finally subjected to consecutive 3 × 3 × 3 convolutions without padding (purple); these steps are repeated three times. The output of the neural network model is a probability map (size 110 × 110 × 110 voxels) of two channels, representing the foreground (FG) and background (BG = 1- FG) probability maps, respectively. Number of featured maps are denoted in red, spatial dimensions at the indicated steps in the neural network in black. Figure designed based on PlotNeuralNet (https://github.com/HarisIqbal88/PlotNeuralNet; adapted from Sheridan et al., 2022). (B–D) Examples of plots showing validation cross entropy loss used to evaluate the predicting behavior of the indicated neural network models for (B) mitochondria, (C) Golgi, or (D) ER periodically obtained during training using FIB-SEM volume data of cells prepared by chemical fixation obtained at ∼5 nm resolution. Cross entropy values were obtained using hold-out ground truth annotations from the training set not used during training or from naïve cells, respectively. The gray area shows the first appearance of relatively stable cross-entropy loss and absence of major spikes obtained by the models during 20,000 consecutive training iterations; these models were then used for prediction. (E) Ground truth annotations consist of true positive (TP) and false negatives (FN) voxels and define the presence or absence of a perfect match with the subcellular structure of interest. The output of the model consists of true (TP) and false positives (FP) voxels, depending on whether the predicted voxels are part or not of the ground truth. F1, as defined in the figure, is used as a practical metric to evaluate the prediction accuracy of the neural network to identify the structure of interest. A perfect model prediction would yield F1 = 1 with FP = 0, FN = 0.
3D U-net architecture, examples of network behavior during training, and F1 as a metric to compare ground truth annotations with model predictions. (A) Schematic representation of the steps used to train the 3D U-net encoder-decoder neural network. The input for the neural network mode are 3D blocks consisting of a stack of consecutive FIB-SEM images (size 204 × 204 × 204 voxels). The 3D block is subjected in the encoder to three cycles, each consisting of consecutive 3 × 3 × 3 convolutions without padding (purple) and downsampling operators with 2 × 2 × 2 max-pooling (pink). The feature maps from the encoder are then upsampled in the decoder by a factor of 2 (yellow), followed by concatenation with previous feature maps from the downsampling branch that had been exposed to central cropping and finally subjected to consecutive 3 × 3 × 3 convolutions without padding (purple); these steps are repeated three times. The output of the neural network model is a probability map (size 110 × 110 × 110 voxels) of two channels, representing the foreground (FG) and background (BG = 1- FG) probability maps, respectively. Number of featured maps are denoted in red, spatial dimensions at the indicated steps in the neural network in black. Figure designed based on PlotNeuralNet (https://github.com/HarisIqbal88/PlotNeuralNet; adapted from Sheridan et al., 2022). (B–D) Examples of plots showing validation cross entropy loss used to evaluate the predicting behavior of the indicated neural network models for (B) mitochondria, (C) Golgi, or (D) ER periodically obtained during training using FIB-SEM volume data of cells prepared by chemical fixation obtained at ∼5 nm resolution. Cross entropy values were obtained using hold-out ground truth annotations from the training set not used during training or from naïve cells, respectively. The gray area shows the first appearance of relatively stable cross-entropy loss and absence of major spikes obtained by the models during 20,000 consecutive training iterations; these models were then used for prediction. (E) Ground truth annotations consist of true positive (TP) and false negatives (FN) voxels and define the presence or absence of a perfect match with the subcellular structure of interest. The output of the model consists of true (TP) and false positives (FP) voxels, depending on whether the predicted voxels are part or not of the ground truth. F1, as defined in the figure, is used as a practical metric to evaluate the prediction accuracy of the neural network to identify the structure of interest. A perfect model prediction would yield F1 = 1 with FP = 0, FN = 0.
As previously noted by others, we also observed that the image contrast and texture of FIB-SEM data can vary substantially between different acquisitions, depending not only on cell type and mode of sample preparation (CF, HPFS) but unexpectedly also between adjacent cells of the same type in the same Epon block (Fig. S4). We found empirically that while the neural network could be trained to segment organelles successfully from samples prepared by the same mode of preparation, a model trained with ground truth annotations from HPFS cells failed when applied to CF-treated cells and vice versa (cross-domain prediction, Fig. 5 A). Although routinely implemented in our pipeline, contrast normalization by contrast-limited adaptive histogram equalization (CLAHE; Pizer et al., 1987; Zuiderveld, 1994) of FIB-SEM datasets from different cells failed to improve the predictions (Table S5). The use of recently proposed local shape descriptors as an auxiliary learning task (Sheridan et al., 2022), calculated from the ground truth annotations and representing high-level morphological notions such as object size and distance to object boundary, also did not improve model prediction. As described below in detail, we addressed the problem of substantial differences in image contrast and texture between different cells by combining ground truth annotations from multiple cells for training.
Use of CLAHE to equalize the contrast of FIB-SEM images. (A–D) Single plane views of FIB-SEM volume data after contrast equalization using CLAHE with a clip limit of 0.02. The samples were prepared by CF (A and B) or HFFS (C and D) and imaged at ∼5 nm isotropic resolution.
Use of CLAHE to equalize the contrast of FIB-SEM images. (A–D) Single plane views of FIB-SEM volume data after contrast equalization using CLAHE with a clip limit of 0.02. The samples were prepared by CF (A and B) or HFFS (C and D) and imaged at ∼5 nm isotropic resolution.
Automated segmentation of organelles
We first applied ASEM to perform automated segmentation of FIB-SEM images from cells prepared by CF with a nominal 5 nm isotropic resolution and relatively high contrast (Fig. 2 and Video 2); the summary shown in Table S6 illustrates the predictive performance obtained for models specific for mitochondria, ER, and Golgi apparatus. For mitochondria, we selected from Cell 1 a training block of about 462 × 106 voxels (1,200 × 700 × 550 voxels) and used semiautomated annotations as ground truth annotations for the mitochondria contained within this volume, representing ∼8% of all voxels (Table S6). Model performance was assessed every 1,000 iterations during the training phase by calculating the cross-entropy loss between the current prediction and the mitochondria ground truth within a validation block (not used during training). Additional smaller validation blocks (Table S4) containing mitochondria ground truth from naïve Cells 2, 3, and 6 were used to avoid overfitting during the training phase and to validate the model performance by measuring the validation losses. Validation losses rapidly converged within 20,000–40,000 training iterations, resulting in a relatively high F1 score (0.91) for Cell 1 and lower values for the data from naïve Cells 2, 3, and 6 (0.47, 0.66, 0.81, cf. Table S6). Similar results were obtained when training with ground truth annotations from Cell 2 instead of Cell 1 (Table S6); the validation losses also converged within 20,000–40,000 training iterations with good F1 scores for Cell 2 (0.87) and naïve Cells 1 and 3 (0.89, 0.74), and a slightly lower score for Cell 6 (0.70), with no further improvement with additional training iterations. To calculate adequate F1 scores even for slightly inaccurate ground truth annotations, we followed previous work (Haberl et al., 2018) and defined a thin metric exclusion zone at the boundary of the ground truth annotations according to the specific structure, ranging from a maximum of eight voxels for mitochondria to a minimum of two voxels for ER (see Materials and methods).
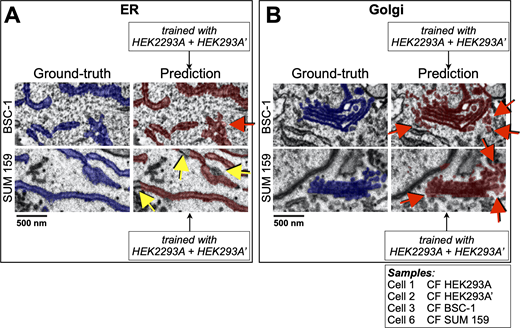
Performance of the deep-learning network to predict in naïve cells. (A and B) Visual comparisons between predictions (crimson) by 3D U-net models trained using combined data from two HEK293A cells to recognize (A) ER or (B) Golgi apparatus and corresponding ground truth annotations (blue) in the naïve BSC-1 and SUM 159 cells not used for training (Table S1). The representative images of single plane views from FIB-SEM volume data are from cells prepared by CF isotropically acquired at a 5 nm resolution; red and yellow arrows highlight small regions containing voxels of false positive and false negative assignments. Scale bar, 500 nm (see Videos 1 and 2).
Performance of the deep-learning network to predict in naïve cells. (A and B) Visual comparisons between predictions (crimson) by 3D U-net models trained using combined data from two HEK293A cells to recognize (A) ER or (B) Golgi apparatus and corresponding ground truth annotations (blue) in the naïve BSC-1 and SUM 159 cells not used for training (Table S1). The representative images of single plane views from FIB-SEM volume data are from cells prepared by CF isotropically acquired at a 5 nm resolution; red and yellow arrows highlight small regions containing voxels of false positive and false negative assignments. Scale bar, 500 nm (see Videos 1 and 2).
Predictions of mitochondria. Passing through the FIB-SEM volume with contrast was equalized using CLAHE. The video shows image and predictions from naïve Cell 1 HEK293A (not used for training using the model trained with ground truth annotations for mitochondria from Cell 2 HEK293A. Both cells were prepared by CF and imaged at ∼5 nm isotropic resolution. The model identified all mitochondria; comparison of the ground truth annotations and predictions shows correct voxel assignments (true positives, yellow), missed assignments (false negatives, cyan), incorrect assignments (false positives, magenta). The small fraction of false positive assignments predicted by the model are associated with unidentified tubular and spherical structures of small size (Chou et al., 2021).
Predictions of mitochondria. Passing through the FIB-SEM volume with contrast was equalized using CLAHE. The video shows image and predictions from naïve Cell 1 HEK293A (not used for training using the model trained with ground truth annotations for mitochondria from Cell 2 HEK293A. Both cells were prepared by CF and imaged at ∼5 nm isotropic resolution. The model identified all mitochondria; comparison of the ground truth annotations and predictions shows correct voxel assignments (true positives, yellow), missed assignments (false negatives, cyan), incorrect assignments (false positives, magenta). The small fraction of false positive assignments predicted by the model are associated with unidentified tubular and spherical structures of small size (Chou et al., 2021).
To find additional ways to enhance the generalization ability of the model, we modified the training pipeline to combine the ground truth annotations from Cells 1 and 2. We first tested the performance of the mitochondria model using the validation blocks in naïve Cells 3 and 6. In this case, the new model had a significantly improved performance (Table S6), reflected by even higher F1 scores for naïve Cells 3 and 6 (0.75, 0.88), but only after 95,000–115,000 iterations (Fig. S5). A similar improvement in model performance was observed for ER predictions when we first combined ground truth annotations of Cells 1 and 2. The new ER model, which was used to predict ER in Cells 1 and 2 and naïve Cells 3 and 6, led to generally improved F1 scores of 0.95, 0.90, 0.92, and 0.77, respectively (Table S6). Consistent with F1 scores smaller than the optimal value of 1, we observed by visual inspection a small number of false negative (yellow arrows) or false positive (red arrows) assignments, as highlighted in Fig. 2 A (see also Video 2). Combining ground truth annotations from Cells 1 and 2 during training to predict the more complex Golgi apparatus in naïve Cells 3 or 6 marginally outperformed the models trained with either Cell 1 or Cell 2 (Table S6), which was also illustrated with one example of visual inspection of ground truth annotations and predictions showing instances of false positive assignments (red arrows, Fig. 2 B). Thus, the predictive performance of a model could often be improved by using a model obtained by jointly training with ground truth annotations from two cells instead of training with data from one cell or the other.
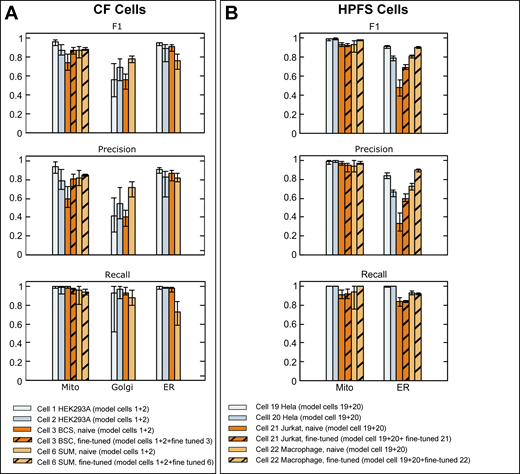
Comparison of validation metrics used to evaluate the prediction accuracy of neural models predicting mitochondria, ER, and Golgi apparatus. (A and B) Ground truth annotations from FIB-SEM volume data from the indicated cells at ∼5 nm isotropic resolution prepared by CF (A) or HPFS (B) were used for training to generate models for mitochondria, ER, and Golgi apparatus. The bar plots show F1, precision, and recall metrics obtained using ground truth annotations not used for training. These values are shown as averages from 20 training iterations spaced at 1,000 intervals, with respective error bars representing maximum and minimal values, calculated after ∼100,000 training iterations. The results also show metrics obtained after fine-tuning with a small number of additional training iterations using ground truth annotations from the naïve cell. Details of datasets, ground truth annotations, and models are summarized in Tables S4, S5, and S2.
Comparison of validation metrics used to evaluate the prediction accuracy of neural models predicting mitochondria, ER, and Golgi apparatus. (A and B) Ground truth annotations from FIB-SEM volume data from the indicated cells at ∼5 nm isotropic resolution prepared by CF (A) or HPFS (B) were used for training to generate models for mitochondria, ER, and Golgi apparatus. The bar plots show F1, precision, and recall metrics obtained using ground truth annotations not used for training. These values are shown as averages from 20 training iterations spaced at 1,000 intervals, with respective error bars representing maximum and minimal values, calculated after ∼100,000 training iterations. The results also show metrics obtained after fine-tuning with a small number of additional training iterations using ground truth annotations from the naïve cell. Details of datasets, ground truth annotations, and models are summarized in Tables S4, S5, and S2.
We also tested the performance of ASEM using FIB-SEM images and ground truth annotations acquired by the OpenOrganelle initiative (Xu et al., 2021; Table S1). These cells were prepared by HPFS and imaged with higher isotropic resolution (4 × 4 × 3–5 nm) but lower contrast. We examined the ability of our training pipeline to segment these datasets and focused on mitochondria and ER but not Golgi due to the lack of a sufficient number of ground truth annotations for Golgi objects in the available OpenOrganelle datasets (Table S7). We generated independent models for mitochondria and ER by training with corresponding combined ground truth annotations from Hela Cells 19 and 20, followed by model performance verification using unseen ground truth annotations from the same Hela cells or different types of naïve cells not used for training (Cell 21 Jurkat-1 and Cell 22 Macrophage-2, Table S7). Our pipeline performed well after ∼100,000 training cycles and managed to segment mitochondria in unseen data from each of the two Hela cells used for training (F1 scores of 0.99, Table S7) and from unseen data from each of the naïve Cell 21 Jurkat-1 or Cell 22 Macrophage-2 (F1 scores of 0.94 and 0.93; Table S7). Automated segmentation of the ER was less efficient, requiring ∼200,000 training cycles to reach the highest model performance (F1 scores of 0.91, 0.80, 0.48, and 0.81, respectively; Table S7). These first results indicate that our training strategy can create predictive models for the successful identification of mitochondria, ER, and Golgi apparatus in cells prepared by CF and of mitochondria and ER in samples prepared by HPFS.
Next, we explored the tolerance of the training pipeline to modest variations of image resolution in naïve cells. The results shown for the representative FIB-SEM images in Fig. 3 and Fig. 4 A, and Video 3 were obtained for a naïve Cell 15 SVG-A prepared by HPFS acquired at an isotropic 5 × 5 × 5 nm (Table S4); visual inspection of the images show successful predicted segmentations for mitochondria, ER, and Golgi apparatus using models obtained by combined training with ground truth annotations from Hela cells 19 and 20, also prepared by HPFS, and whose FIB-SEM images were acquired with mixed resolutions of 4 × 4 × 5.2 and 4 × 4 × 3.2 nm, respectively.
Network predictions of mitochondria, ER, and Golgi apparatus. Single plane view from a FIB-SEM volume image from naïve cell 15 (SVG-A) not used for training prepared by HPFS and visualized during interphase at 5 nm isotropic resolution. The small region contains representative model predictions for mitochondria (cyan), ER (yellow), and Golgi apparatus (magenta) obtained from three 3D U-net models, each trained with organelle-specific ground truth annotations, without fine-tuning, from interphase cells 19 (Hela-2) and 20 (Hela-3) prepared by HPFS. Scale bar, 2 μm.
Network predictions of mitochondria, ER, and Golgi apparatus. Single plane view from a FIB-SEM volume image from naïve cell 15 (SVG-A) not used for training prepared by HPFS and visualized during interphase at 5 nm isotropic resolution. The small region contains representative model predictions for mitochondria (cyan), ER (yellow), and Golgi apparatus (magenta) obtained from three 3D U-net models, each trained with organelle-specific ground truth annotations, without fine-tuning, from interphase cells 19 (Hela-2) and 20 (Hela-3) prepared by HPFS. Scale bar, 2 μm.
Predictive ER model resolves the structural complexity of the ER network during different stages of the cell cycle. (A) Representative examples of ER predictions in naïve cell 15 (SVG-A) processed during interphase as described in Fig. 3 showing the characteristic network of ER sheets connected at branch points to ER tubules. ER tubules were more abundant toward the periphery of this cell and ER sheets were more abundant closer to the nucleus. For clarity, manual VAST editing was used to eliminate pixels of false positive predictions associated with the nuclear envelope. Scale bar, 1 μm. (B) Representative examples of ER predictions from a mitotic naïve cell 8 (SUM 159) prepared by CF and imaged isotropically at 10 nm; the ER model was trained with ER ground truth annotations from interphase cells 1 and 2 (HEK293A) prepared by CF visualized isotropically at 5 nm resolution and downsampled to 10 nm. It shows successful recognition of an extensive network of fenestrated ER sheets (red arrow heads) connected to ER tubules, characteristic of mitotic cells. Ground truth annotations used to train the interphase ER model did not contain ER fenestrations, as they are barely present during stage of the cell cycle. Darker regions corresponding to chromosomes are outlined with yellow dotted lines. Scale bar, 3 μm (see Video 3).
Predictive ER model resolves the structural complexity of the ER network during different stages of the cell cycle. (A) Representative examples of ER predictions in naïve cell 15 (SVG-A) processed during interphase as described in Fig. 3 showing the characteristic network of ER sheets connected at branch points to ER tubules. ER tubules were more abundant toward the periphery of this cell and ER sheets were more abundant closer to the nucleus. For clarity, manual VAST editing was used to eliminate pixels of false positive predictions associated with the nuclear envelope. Scale bar, 1 μm. (B) Representative examples of ER predictions from a mitotic naïve cell 8 (SUM 159) prepared by CF and imaged isotropically at 10 nm; the ER model was trained with ER ground truth annotations from interphase cells 1 and 2 (HEK293A) prepared by CF visualized isotropically at 5 nm resolution and downsampled to 10 nm. It shows successful recognition of an extensive network of fenestrated ER sheets (red arrow heads) connected to ER tubules, characteristic of mitotic cells. Ground truth annotations used to train the interphase ER model did not contain ER fenestrations, as they are barely present during stage of the cell cycle. Darker regions corresponding to chromosomes are outlined with yellow dotted lines. Scale bar, 3 μm (see Video 3).
Prediction of mitochondria, ER, Golgi apparatus, nuclear pores, and clathrin-coated pits and vesicles. Passing through the raw FIB-SEM volume from naïve Cell 15 SVG-A prepared by HPFS and imaged at ∼5 nm isotropic resolution. The video shows predictions as surface renderings for mitochondria (cyan), ER (yellow), Golgi apparatus (magenta). For simplicity, only predictions in a block of 3 × 3 × 3 µm (block size: 664 × 586 × 572 voxels) are shown. A small number of false positive pixels generated by the Golgi model and located within a 323 × 271 × 230 voxel block were removed using VAST. One identified Golgi apparatus is highlighted (light pink). Predictions for all nuclear pores (yellow) and clathrin-coated pits and vesicles (red) within the imaged volume are also shown. Visual inspection confirmed that the models trained with ground truth annotations from Cell 19 Hela and Cell 20 Hela prepared by HPFS and imaged at ∼5 nm isotropic resolution correctly predicted all the intracellular structures in Cell 15 SVG-A.
Prediction of mitochondria, ER, Golgi apparatus, nuclear pores, and clathrin-coated pits and vesicles. Passing through the raw FIB-SEM volume from naïve Cell 15 SVG-A prepared by HPFS and imaged at ∼5 nm isotropic resolution. The video shows predictions as surface renderings for mitochondria (cyan), ER (yellow), Golgi apparatus (magenta). For simplicity, only predictions in a block of 3 × 3 × 3 µm (block size: 664 × 586 × 572 voxels) are shown. A small number of false positive pixels generated by the Golgi model and located within a 323 × 271 × 230 voxel block were removed using VAST. One identified Golgi apparatus is highlighted (light pink). Predictions for all nuclear pores (yellow) and clathrin-coated pits and vesicles (red) within the imaged volume are also shown. Visual inspection confirmed that the models trained with ground truth annotations from Cell 19 Hela and Cell 20 Hela prepared by HPFS and imaged at ∼5 nm isotropic resolution correctly predicted all the intracellular structures in Cell 15 SVG-A.
Since models trained on mitochondria or ER ground truth annotations from cells prepared by CF performed poorly on cells prepared by HPFS and vice versa, as judged by a qualitative visual assessment of the outcomes (cross-domain prediction, Fig. 5 A), we explored the possibility of combining training data from both sample preparation protocols to create generalist models using the same training datasets from HEK293A Cells 1 and 2 prepared by CF, and from Cells 19 and 20 Hela prepared by HPFS. On these cells, the generalist mitochondria and ER models performed nearly as well as with models obtained using samples prepared with either one of the protocols on almost all validation datasets for either sample preparation protocol (Fig. 5 A and Table S8).
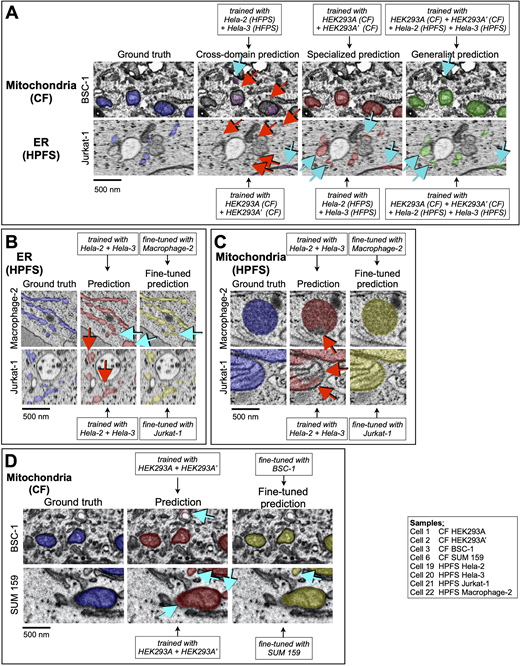
Effects of extensive combination of datasets and fine-tuning during training. (A–D) Examples to highlight the effect on the predictive performance of (A, C, and D) mitochondria and (A and B) ER and models trained with data from cells prepared by CF or HPFS, with substantial differences in general appearance and contrast. The images show several comparisons between ground truth annotations and predictions from models trained as described in the insets with data obtained from cells prepared by different sample preparation protocols. Details of the cell and training protocols are in Tables S1, S2, and S8. Voxels corresponding to false positive (cyan arrows) and false negative (red arrows) predictions are indicated. Scale bar, 500 nm. (A) Predictions from cross-domain models, for which the training data and predictions were done using cells prepared with different sample preparation protocols, were less accurate than those obtained from the specialized models, for which training and predictions were done using cells prepared with the same sample preparation protocol. Predictions from the generalist models, obtained by training using ground truth annotations from cells prepared by CF and HPFS, performed only slightly worse than the predictions from the specialized models. (B–D) Effect on the predictive performance of the models by fine-tuning during training.
Effects of extensive combination of datasets and fine-tuning during training. (A–D) Examples to highlight the effect on the predictive performance of (A, C, and D) mitochondria and (A and B) ER and models trained with data from cells prepared by CF or HPFS, with substantial differences in general appearance and contrast. The images show several comparisons between ground truth annotations and predictions from models trained as described in the insets with data obtained from cells prepared by different sample preparation protocols. Details of the cell and training protocols are in Tables S1, S2, and S8. Voxels corresponding to false positive (cyan arrows) and false negative (red arrows) predictions are indicated. Scale bar, 500 nm. (A) Predictions from cross-domain models, for which the training data and predictions were done using cells prepared with different sample preparation protocols, were less accurate than those obtained from the specialized models, for which training and predictions were done using cells prepared with the same sample preparation protocol. Predictions from the generalist models, obtained by training using ground truth annotations from cells prepared by CF and HPFS, performed only slightly worse than the predictions from the specialized models. (B–D) Effect on the predictive performance of the models by fine-tuning during training.
We also evaluated the performance of ASEM to predict mitochondria, ER, and Golgi apparatus imaged with FIB-SEM data at 5 nm isotropic resolution but processed at a lower resolution of 10 nm. This test was done by using datasets from Cells 1 and 2 isotropically downsampled to 10 nm to train new models for mitochondria, ER, and Golgi apparatus and then used to predict the validation data from Cells 1, 2, 3, and 6 isotropically downsampled to 10 nm (Table S9). These results showed that while the mitochondria and ER models performed similarly at both resolutions, the performance for the Golgi apparatus model notably decreased (Table S9), presumably explained by the relatively larger spatial complexity of the Golgi apparatus.
Fine-tuning
To improve the predictive performance with images from naïve cells, we explored the effect of fine-tuning a pre-existing model, a simple implementation of transfer learning (Weiss et al., 2016). As described in the Materials and methods, we started with an already trained model and resumed model training for a low number of iterations (15,000) using only the new ground truth annotations from the naïve cell; the new ground truth annotations, although resembling those used for the first training, would typically have slightly different characteristics.
The following examples illustrate the range of results obtained upon implementation of fine-tuning using HPFS FIB-SEM data. The ER model, first obtained after ∼180,000 training cycles using ground truth annotations from Hela Cells 19 and 20, was then fine-tuned for additional 12,000 or 6,000 training cycles with small amounts of ground truth data from either naïve Cell 21 Jurkat-1 or Cell 22 Macrophage-2; both fine-tuning cases showed a significant improvement in the F1 precision scores, from 0.48 to 0.69 and from 0.81 to 0.90, without affecting recall (Fig. S5 and Table S10). In other words, the model learned to correctly classify ER while at the same time reduced the number of false positives by rejecting structures that appeared similar but did not belong to the same semantic class (Fig. 5 B). The next two cases of fine-tuning illustrate little or no improvement in predictive model performance for mitochondria in cells prepared by HPFS or CF (Fig. 5 C, Fig. S5, and Table S10). The model obtained after 95,000 training cycles using HPFS FIB-SEM data from Hela Cells 19 and 20 showed similar F1 scores (0.93) for naïve Cell 21 Jurkat-1 or Cell 22 Macrophage-2 before or after fine-tuning for 7,000 cycles.
Similarly, a mitochondria model obtained after 95,000 training iterations using CF FIB-SEM data from Cells 1 and 2 and then fine-tuned for additional 6,000 fine-tuning training steps using ground truth annotations from Cells 3 or 6 showed either a significant increase (from 0.75 to 0.88) or no increase at all (0.88) in F1 scores, respectively (Fig. 5 D and Table S10). Fine-tuning had minimal or no effect for situations in which the pretrained model produced a prediction of naïve cells with a high F1 score, such as in mitochondria with an F1 score of around 0.9. We conclude that fine-tuning can be beneficial for segmenting relatively large membrane-bound organelles, particularly in cases where the pretrained model behaved poorly in naïve cells, but it could not resolve situations in which the staining characteristics of the samples were extremely different, even though they had been prepared by the same staining procedures.
Automated segmentation of nuclear pores
To test whether our pipeline can automatically identify and segment small intracellular structures, we trained the neural network with ground truth annotations from nuclear pores, structures embedded in the double-membrane nuclear envelope with membrane pore openings of ∼100–120 nm in diameter. We used FIB-SEM data with a nominal 5 nm isotropic resolution from interphase SVG-A and Hela cells imaged using HPFS to ensure minimal perturbations in the structural organization of the nuclear pores and their surrounding inner and outer nuclear membranes.
We used VAST to generate ground truth annotations for ten nuclear pores from Cell 13a–SVG-A (5 × 5 × 5 nm isotropic resolution; Table S4). The segmentations, representing the inner and outer nuclear membrane envelope contours immediately adjacent to nuclear pores, also included five additional pixels (∼25 nm) of inner and outer nuclear membrane extending away from the nuclear pore opening (Fig. 6 A). The training was performed with the augmented data generated from only eight nuclear pores (with two additional objects for validation), resulting in a nuclear pore model that performed well after 100,000 training iterations (F1 = 0.52, Precision = 0.35, Recall = 0.99, Table S8). In all cases, the high recall score was consistent with a perfect correspondence to all the voxels that defined the ground truth annotations. The relatively low F1 and precision scores reflected “fatter” predictions due to voxels assigned to positions immediately adjacent to the “single row” of voxels overlapping the nuclear pores in the ground truth annotations. Visual inspection confirmed accurate identification of all nuclear pores in naïve SVG-A cells 15 (Video 4) and 17 (5 × 5 × 5 nm isotropic resolution) and Cell 19 Hela (4 × 4 × 5.2 nm) not used for training (Fig. 6 B). Because of the high predictive accuracy attained with this simple nuclear pore model (Video 4), it was not necessary to improve the model using our more extended training pipelines, such as fine-tuning.
Identification of nuclear pores and variations in their membrane pore diameter. A nuclear pore model was generated by training on ground truth annotations of nuclear pores from cell 13a SVG-A prepared by HPFS and imaged at 5 nm isotropic resolution. (A) Orthogonal views of a representative nuclear pore not used for training show ground truth annotations and model prediction. Scale bar, 50 nm. (B) Nuclear pore predictions for all the pores on the nuclear envelope of naïve cell 19 (Hela-2) prepared by HPFS and visualized during interphase at 4 × 4 × 5.3 nm isotropic resolution (left panel); the inset highlights the characteristic doughnut shape of the nuclear pore. Scale bar, 2 μm. Representative orthogonal views (right panels) of a nuclear pore and model prediction. Scale bar, 50 nm. (C) Histogram of nuclear membrane pore diameters measured in naïve cells 15, 17, and 19 (N = 934) identified by the nuclear pore model. Each membrane pore diameter determined in the raw image represents the average value from 18 measurements spaced 10° apart (see inset and Materials and methods). The Gaussian fit (blue) shows the expected size distribution if the data had come from membrane pores of a single diameter centered on the experimentally determined median (d = 100 nm, most abundant species); a standard deviation of 6 nm corresponds to the expected error of the measurements (see Materials and methods). (D) Three-dimensional distribution of nuclear pores on the nuclear envelope of cell 19, color-coded as a function of membrane pore diameter.
Identification of nuclear pores and variations in their membrane pore diameter. A nuclear pore model was generated by training on ground truth annotations of nuclear pores from cell 13a SVG-A prepared by HPFS and imaged at 5 nm isotropic resolution. (A) Orthogonal views of a representative nuclear pore not used for training show ground truth annotations and model prediction. Scale bar, 50 nm. (B) Nuclear pore predictions for all the pores on the nuclear envelope of naïve cell 19 (Hela-2) prepared by HPFS and visualized during interphase at 4 × 4 × 5.3 nm isotropic resolution (left panel); the inset highlights the characteristic doughnut shape of the nuclear pore. Scale bar, 2 μm. Representative orthogonal views (right panels) of a nuclear pore and model prediction. Scale bar, 50 nm. (C) Histogram of nuclear membrane pore diameters measured in naïve cells 15, 17, and 19 (N = 934) identified by the nuclear pore model. Each membrane pore diameter determined in the raw image represents the average value from 18 measurements spaced 10° apart (see inset and Materials and methods). The Gaussian fit (blue) shows the expected size distribution if the data had come from membrane pores of a single diameter centered on the experimentally determined median (d = 100 nm, most abundant species); a standard deviation of 6 nm corresponds to the expected error of the measurements (see Materials and methods). (D) Three-dimensional distribution of nuclear pores on the nuclear envelope of cell 19, color-coded as a function of membrane pore diameter.
Prediction of mitotic ER. Passing through the FIB-SEM volume with contrast equalized. Image is from naïve prometaphase Cell 8 SUM 159 imaged at ∼10 nm isotropic resolution. The video shows ER predictions (yellow) generated with ground truth annotations from interphase Cell 1 HEK293A and Cell 2 HEK293A imaged at ∼5 nm isotropic resolution. All cells were prepared by CF. Visual inspection confirmed that the model correctly predicted all the ER, including the fenestrations characteristic of the mitotic ER sheets; fenestrations were not included in the ground truth annotations used for training, as they are mostly absent in the ER of interphase cells (Chou et al., 2021).
Prediction of mitotic ER. Passing through the FIB-SEM volume with contrast equalized. Image is from naïve prometaphase Cell 8 SUM 159 imaged at ∼10 nm isotropic resolution. The video shows ER predictions (yellow) generated with ground truth annotations from interphase Cell 1 HEK293A and Cell 2 HEK293A imaged at ∼5 nm isotropic resolution. All cells were prepared by CF. Visual inspection confirmed that the model correctly predicted all the ER, including the fenestrations characteristic of the mitotic ER sheets; fenestrations were not included in the ground truth annotations used for training, as they are mostly absent in the ER of interphase cells (Chou et al., 2021).
Based on ensemble cryo-EM data from thousands of nuclear pores that provide a unique atomic model per dataset (Schuller et al., 2021), combined with more selective images of single nuclear pores obtained using cryo tomography of yeast cells in different physiological states (Zimmerli et al., 2021), it is now believed that the diameter of the nuclear pore varies in response to the physiological state of the cell. It is not known, however, as to what extent this size variability occurs within a single cell in a unique physiological state. Taking advantage of our automated segmentation pipeline that makes it practical to analyze hundreds of single nuclear pores, we explored the extent to which their membrane pore diameters varied within a single cell. Inspection of the nuclear membrane surrounding the pores viewed along the axis normal to the nuclear envelope confirmed the radial symmetry of the pore (Fig. 6 B) with a relatively broad and continuous variation in membrane pore diameter, ranging from 60 to 130 nm (median 92 nm, with 75–108 nm 10–90 percentile range: n = 934; 305, 135, and 494 pores from SVG-A Cells 15 and 17, and Hela Cell 19, respectively; Fig. 6 C); these values were obtained by measuring the distances between the peak signals at opposite ends of the nuclear membrane pore (see Materials and methods and Fig. S6, A–D) in the raw images. The membrane pore sizes did not follow a normal distribution but instead had a slight asymmetry contributed by smaller species. They were also distinct from the Gaussian fit (blue, Fig. 6 C) corresponding to the expected size distribution if the data would have originated from a single pore size centered on the most abundant species (d = 100 nm). We found no evidence to suggest the presence of spatial correlation between pore diameter and different regions of the nuclear envelope within the cell, for example, away from the coverslip or normal to this surface, nor did we find evidence of local clustering of pores with a favored size (Fig. 6 D and Fig. S7).
Steps to determine the diameter of the nuclear pore membrane. (A) Nuclear pore predictions for all the pores on the nuclear envelope of naïve interphase cell 19 (Hela-2) prepared by HPFS and visualized at 4 × 4 × 5.3 nm isotropic resolution. The nuclear pore predictions were obtained using a model trained without fine tuning with ground truth annotations for Cell 13 (Hela) prepared by HPFS and imaged at ∼5 nm isotropic resolution. (B) Volume location of the centroid of each of the predicted nuclear pore, color coded according to their relative position along the Z-axis (top panel) and surface rendition of the nuclear envelope (green) obtain by alpha-shape triangulation of the centroids (see Materials and methods). Orthonormal vectors associated with each triangle are shown (red). (C) Example of realignment of a nuclear pore from its acquisition orientation in the FIB-SEM volume image to a new view with the nuclear envelope orthogonal to the Z-axis; side views and volume rendition of the nuclear pore prediction are shown. (D) Single plane on the face and orthogonal views of a nuclear pore centered on the middle of the nuclear envelope (left panels) and examples of the intensity plots used to estimate the membrane pore diameters by determining the distance separating the two intensity minima along the indicated axis (right panels). The nuclear pore diameter is reported as the average of 18 values obtained 10° apart (inset in left panel).
Steps to determine the diameter of the nuclear pore membrane. (A) Nuclear pore predictions for all the pores on the nuclear envelope of naïve interphase cell 19 (Hela-2) prepared by HPFS and visualized at 4 × 4 × 5.3 nm isotropic resolution. The nuclear pore predictions were obtained using a model trained without fine tuning with ground truth annotations for Cell 13 (Hela) prepared by HPFS and imaged at ∼5 nm isotropic resolution. (B) Volume location of the centroid of each of the predicted nuclear pore, color coded according to their relative position along the Z-axis (top panel) and surface rendition of the nuclear envelope (green) obtain by alpha-shape triangulation of the centroids (see Materials and methods). Orthonormal vectors associated with each triangle are shown (red). (C) Example of realignment of a nuclear pore from its acquisition orientation in the FIB-SEM volume image to a new view with the nuclear envelope orthogonal to the Z-axis; side views and volume rendition of the nuclear pore prediction are shown. (D) Single plane on the face and orthogonal views of a nuclear pore centered on the middle of the nuclear envelope (left panels) and examples of the intensity plots used to estimate the membrane pore diameters by determining the distance separating the two intensity minima along the indicated axis (right panels). The nuclear pore diameter is reported as the average of 18 values obtained 10° apart (inset in left panel).
Three-dimensional distribution of nuclear pores on the nuclear envelope. Three-dimensional distribution of nuclear pores on the nuclear envelopes of Cells 15 and 17 color-coded by a heat map as a function of membrane pore diameter.
Three-dimensional distribution of nuclear pores on the nuclear envelope. Three-dimensional distribution of nuclear pores on the nuclear envelopes of Cells 15 and 17 color-coded by a heat map as a function of membrane pore diameter.
Automated segmentation of clathrin-coated pits, coated vesicles, and caveolae
As a further test of ASEM with relatively small structures, we chose clathrin-coated pits, 30–100 nm membrane invaginations in the plasma membrane, and the trans-Golgi network (TGN) involved in selective cargo traffic (Kirchhausen, 2000). We trained the model with ground truth annotations from 15 endocytic plasma membrane–coated pits of different sizes and shapes, thus representing different stages of clathrin coat assembly. While the resolution of the FIB-SEM was insufficient to discern the familiar spikes or the hexagonal and pentagonal facets of a clathrin coat as seen in samples imaged by TEM, the presence of strong membrane staining, which we attribute to clathrin and associated proteins (Fig. 7 A), made these invaginations recognizably distinct from caveolae, which are smaller (50–100 nm) flask-shaped invaginations that lack enhanced membrane staining (Fig. 7 B). None of the cells had recognizable regions of strongly stained, flat membrane, often found on the coverslip-attached surface of cells in culture and other specialized locations (Akisaka et al., 2008; Grove et al., 2014; Heuser, 1980; Maupin and Pollard, 1983; Saffarian et al., 2009; Signoret et al., 2005). We used VAST to generate the clathrin-coated pit ground truth annotations, which were simply a collection of single traces loosely overlapping the endocytic membrane invagination (Fig. 7 A, blue).
Identification of clathrin-coated pits, coated vesicles and caveolae. A coated pit model was generated by training with ground truth annotations from Cell 12 (Hela-2) prepared by HPFS and imaged at ∼5 nm isotropic resolution. (A) Orthogonal views of a representative endocytic clathrin-coated pit (CCP) not used for training showing ground truth annotations and model prediction. Scale bar, 50 nm. (B) Orthogonal views of a caveola, an endocytic clathrin-coated pit (CCP) and a clathrin-coated vesicle (CCV) at the plasma membrane, and a coated pit (CCP) and vesicle (CCV) associated with membranes from the secretory pathway. Each panel shows the ground truth annotation and the model prediction. An example of a COPI vesicle not predicted by the coated pit model is also shown. Views are from naïve cell 17 SVGA prepared by HPFS and imaged with ∼5 nm isotropic resolution.
Identification of clathrin-coated pits, coated vesicles and caveolae. A coated pit model was generated by training with ground truth annotations from Cell 12 (Hela-2) prepared by HPFS and imaged at ∼5 nm isotropic resolution. (A) Orthogonal views of a representative endocytic clathrin-coated pit (CCP) not used for training showing ground truth annotations and model prediction. Scale bar, 50 nm. (B) Orthogonal views of a caveola, an endocytic clathrin-coated pit (CCP) and a clathrin-coated vesicle (CCV) at the plasma membrane, and a coated pit (CCP) and vesicle (CCV) associated with membranes from the secretory pathway. Each panel shows the ground truth annotation and the model prediction. An example of a COPI vesicle not predicted by the coated pit model is also shown. Views are from naïve cell 17 SVGA prepared by HPFS and imaged with ∼5 nm isotropic resolution.
The coated pit model obtained after 80,000–100,000 training iterations used six ground truth annotations from Cell 12 Hela and nine from Cell 13 Hela. Visual inspection of the predictions generated by this relatively simple training in parts of Hela cells 12 and 13, cells that had not been used for training, showed accurate recognition of all endocytic coated pits (representative example in Fig. 7 B); we obtained similar results from naïve SVG-A cells 15 (Video 4) and 17 and Hela cell 19. The model also identified all coated pits in the TGN (Fig. 7 B), incorrectly identified caveolae as coated pits (Fig. 7 B), and we could detect no other incorrect predictions anywhere in the cell volume. Since caveolae were easy to filter out by a combination of size and appearance, we chose not to train another model that could have, for example, included a high and disproportionate amount of caveolae as the background (in these cells, caveolae are significantly less abundant than coated pits captured at different stages of assembly). A sharply invaginating curvature of the stained membrane outline thus appears to be an important component of the pattern the model learned to recognize.
We used our additional annotated ground truth annotations from Hela Cells 12 and 13 that had not been included in the training set to calculate F1, recall, and precision scores (Table S8). In all cases, the high recall score (0.99) demonstrated the almost perfect reconstruction of all voxels belonging to the ground truth annotations. The relatively low F1 and precision scores (∼0.65 and 0.51) were due to incorrect voxel predictions immediately adjacent to the “single row” of true voxel assignments overlapping the invaginated membrane in the ground truth annotations (Fig. 7, A and B).
The model also recognized vesicles near the plasma membrane and the TGN that an expert human observer would have interpreted from their staining to be clathrin-coated vesicles, even though training of the model did not include ground truth annotations representing them (Fig. 7 B). We confirmed that the model recognized all the presumptive coated vesicles in a cell by visual inspection across the full volumes of Hela cells 12 and 13, as well as of three cells that did not contribute at all to the training set, SVG-A cells 15 and 17 and Hela cell 19. Training on endocytic-coated pits thus also allowed recognition of endocytic-coated vesicles and TGN-coated pits. In contrast, the model did not recognize vesicles associated with the Golgi apparatus or the ER that had been interpreted by their staining as COPI or COPII.
We took advantage of the large, combined set of three-dimensional image data from coated pits and vesicles to analyze assembly stages using the metrics depicted in Fig. S8. We determined the depths and widths at half a depth for each of the membrane invaginations in SVG-A cell 17 (Fig. 8 A). Caveolae, recognized by the absence of an enhanced membrane signal, were relatively small, with narrow distributions of depths and widths centered on 61 and 81 nm (Fig. 8 A), respectively. Endocytic-coated pits, identified by their enhanced membrane signals, were generally larger than caveolae and had wider distributions of depths and widths, which clustered into two groups. Coated pits with open necks (>40 nm) had shallow, ∼50-nm invaginations; those with narrower necks (∼10–40 nm) had deep, ∼100–130 nm invaginations (Fig. 8 B, left and central panel, and Fig. 8 C, right panel). Endocytic-coated pits and vesicles were also larger than the corresponding secretory structures emanating from internal membranes associated with the TGN (Fig. 8 B, left panel).
Definition of metrics used to characterize clathrin-coated structures. (A) Schematic representation of the timeline to describe the formation of a clathrin-coated pit mediated by the assembly of the clathrin coat (Kirchhausen et al., 2014). The last step mediated by fission of the membrane neck connecting the mature coated pit from the originating membrane results in formation of the fully formed coated vesicle. Metrics of neck width, pit height, full width at half maximum, and major and minor axis of the fitted ellipse used to morphologically describe the clathrin-coated pits are shown. (B) Metrics used to characterize clathrin-coated vesicles. (C) Example of a single plane from a selected endocytic clathrin-coated pit in a cell prepared by HPFS and imaged by FIB-SEM at ∼5 nm isotropic resolution. The darker voxels corresponding to the deformed membrane and the coat surrounding the pit (left panel) were segmented using an Otsu-based intensity threshold approach (Otsu, 1979) to generate a skeletonized binary mask (central panel) which was then used to fit the ellipse (right panel).
Definition of metrics used to characterize clathrin-coated structures. (A) Schematic representation of the timeline to describe the formation of a clathrin-coated pit mediated by the assembly of the clathrin coat (Kirchhausen et al., 2014). The last step mediated by fission of the membrane neck connecting the mature coated pit from the originating membrane results in formation of the fully formed coated vesicle. Metrics of neck width, pit height, full width at half maximum, and major and minor axis of the fitted ellipse used to morphologically describe the clathrin-coated pits are shown. (B) Metrics used to characterize clathrin-coated vesicles. (C) Example of a single plane from a selected endocytic clathrin-coated pit in a cell prepared by HPFS and imaged by FIB-SEM at ∼5 nm isotropic resolution. The darker voxels corresponding to the deformed membrane and the coat surrounding the pit (left panel) were segmented using an Otsu-based intensity threshold approach (Otsu, 1979) to generate a skeletonized binary mask (central panel) which was then used to fit the ellipse (right panel).
Dimensions of clathrin-coated pits, coated vesicles, and caveolae. (A) Violin plots of width and height for caveolae (CAV) and endocytic clathrin-coated pits (CCP) in the raw images of the structures identified by the coated pit model in cell 17 (see also Fig. S9, and Clathrin-coated pits and vesicles, Materials and methods). (B) Violin plots of the major and minor axis and eccentricity of the fitted ellipse of all pits and vesicles in the raw images of the structures identified by the coated pit model in cell 17 (see also Fig. S9, and Clathrin-coated pits and vesicles, Materials and methods). (C) The left-hand panel shows the distribution of height versus neck width for endocytic clathrin-coated pits in cell 17, identified by the coated pit model. The plot shows two clusters, which correspond to early and late coated pits, respectively, as illustrated by the schematics (see also Fig. S9, and Clathrin-coated pits and vesicles in Materials and methods). The right-hand panel shows histograms for height and major axis of the fitted ellipse for late endocytic coated pits and coated vesicles, respectively.
Dimensions of clathrin-coated pits, coated vesicles, and caveolae. (A) Violin plots of width and height for caveolae (CAV) and endocytic clathrin-coated pits (CCP) in the raw images of the structures identified by the coated pit model in cell 17 (see also Fig. S9, and Clathrin-coated pits and vesicles, Materials and methods). (B) Violin plots of the major and minor axis and eccentricity of the fitted ellipse of all pits and vesicles in the raw images of the structures identified by the coated pit model in cell 17 (see also Fig. S9, and Clathrin-coated pits and vesicles, Materials and methods). (C) The left-hand panel shows the distribution of height versus neck width for endocytic clathrin-coated pits in cell 17, identified by the coated pit model. The plot shows two clusters, which correspond to early and late coated pits, respectively, as illustrated by the schematics (see also Fig. S9, and Clathrin-coated pits and vesicles in Materials and methods). The right-hand panel shows histograms for height and major axis of the fitted ellipse for late endocytic coated pits and coated vesicles, respectively.
The eccentricity of the assembling pit, defined as the ratio of major and minor axes of the ellipsoid that fit best to a given membrane profile, showed a relatively narrow and overlapping distribution (Fig. 8 B, right panel), ranging from 1 (symmetric) to 1.6 (less symmetric) for endocytic pits and vesicles, respectively. Most of the pits and vesicles associated with internal membranes in SVG-A Cell 17 (Fig. 8 B, right panel) had eccentricities close to 1; in those cases, the major axis of most pits was orthogonal to the plane from which the pits invaginated. Similar results were obtained for SVG-A cell 15 and Hela cells 12 and 13 (Fig. S9, A–C). These results are consistent with a budding mechanism in which the stepwise growth of the clathrin coat drives invagination of the membrane, ultimately creating a constriction, as the curved clathrin lattice approaches closure that is narrow enough for dynamin to pinch off the nascent vesicle (Kirchhausen et al., 2014).
Characterization of clathrin-coated pits and coated vesicles. Data shown in this figure for Cells 12, 13, 15, and 17 were generated using the coated pit model employed in Fig. 7 obtained by training with ground truth annotations from Cell 12 prepared by HPFS and imaged at ∼5 nm isotropic resolution. (A) Violin plots of the major and minor axis and eccentricity of the fitted ellipse of all pits and vesicles in the raw images of the structures identified by the coated pit model. (B) Scatter plot of height versus neck width of endocytic clathrin-coated pits clustered in two groups associated with early and late stages of pit formation (left panel). The histogram compares the height and major axis for the fitted ellipse of late endocytic coated pits and coated vesicles, respectively. (C) Scatter plot of height versus neck width of “secretory” clathrin-coated pits associated with internal membranes.
Characterization of clathrin-coated pits and coated vesicles. Data shown in this figure for Cells 12, 13, 15, and 17 were generated using the coated pit model employed in Fig. 7 obtained by training with ground truth annotations from Cell 12 prepared by HPFS and imaged at ∼5 nm isotropic resolution. (A) Violin plots of the major and minor axis and eccentricity of the fitted ellipse of all pits and vesicles in the raw images of the structures identified by the coated pit model. (B) Scatter plot of height versus neck width of endocytic clathrin-coated pits clustered in two groups associated with early and late stages of pit formation (left panel). The histogram compares the height and major axis for the fitted ellipse of late endocytic coated pits and coated vesicles, respectively. (C) Scatter plot of height versus neck width of “secretory” clathrin-coated pits associated with internal membranes.
Discussion
The automated 3D image segmentation pipeline embodied in ASEM overcomes three critical hurdles for making FIB-SEM more practical and more broadly useful than currently available procedures. (1) Our graph-cut-based annotation approach facilitates and simplifies the manual stages of ground truth annotation for convoluted structures like the Golgi apparatus by minimizing the number of hand-curated annotations. Between 8 and 15 annotated structures encompassing the complete volumes of smaller objects (nuclear pores, clathrin-coated pits) or partial volumes of larger ones (mitochondria, ER, Golgi apparatus) were generally enough when augmented as described. We used rough annotations that could be off by one to two voxels rather than voxel-precise labeling to delineate the outline of the intracellular structures for which we were training. While this strategy was effective for our training pipeline, it was much less time-consuming than the precise delineation efforts used by COSEM (see Materials and methods). We could then readily correct any erroneous voxel detections, either by manual intervention or by automatic postprocessing. (2) For the applications described here, ASEM requires far less computational effort than COSEM or other approaches, largely because we restrict the training to a single type of structure and thus create a separate model for each type. Consequently, we found that about ∼100,000–150,000 training iterations were sufficient for accurate prediction, whereas COSEM required five times as many. (3) We can substantially improve the success rate in a completely naïve cell by using a model trained on ground truth annotations from another cell and retraining by a simplified transfer learning approach with a very small number of ground truth annotations from the new cell, thereby adapting the model to a cell with slightly different imaging characteristics at the cost of modest additional segmentation and computational effort. In the examples here, just 5,000–10,000 training iterations were enough to increase prediction accuracy throughout the rest of that cell.
To test the robustness and flexibility of ASEM, we used the model trained with ER ground truth annotations from cells in interphase for identifying and segmenting ER in an early anaphase mitotic cell. The model, which had correctly identified and segmented the complete ER in a naïve interphase cell imaged at ∼5-nm isotropic resolution, also accurately identified and segmented the ER in the mitotic cell imaged at ∼10 nm isotropic resolution with a model trained with ground truths from the same cells in interphase but computationally downsampled to 10 nm (Fig. 4 B and Video 4). The result is nontrivial because relatively extended, fenestrated, and double-membrane sheets with small interconnecting tubules dominate the morphology of the mitotic ER, while tubules of varying lengths, connecting much smaller sheets, are the principal structures in the interphase ER. Segmenting the mitotic ER within the imaged cell volume (∼29 voxels, voxel size 10 × 10 × 10 nm) required less than an hour; it would have taken a human annotator several months. Previous analyses were limited to small cell volumes precisely because of this constraint. We further showed that automatic segmentation of the Golgi apparatus with ASEM confirmed the results described by the COSEM Project team (Heinrich et al., 2021). The Golgi is not a stack of closely packed, uniform cisternae, as often diagrammed in textbooks. Rather, each member of the stack is a complex, perforated structure with variable shapes surrounded by many small vesicles.
We used automatic segmentation of mitochondria, ER, and Golgi apparatus primarily for comparison with published results from other methods to validate the features of ASEM designed to accelerate and simplify the entire pipeline. We turned to smaller intracellular structures as tests of new and potentially more challenging applications. Nuclear pores are more homogenous than larger organelles, and thus in principle, easier to recognize, and while any single pore has a much less distinct substructure than does a Golgi stack or a mitochondrion, we have found that the diameter of the membrane pore varies even across the nucleus of single cells at a fixed time point, despite the likely invariance of much of the nuclear pore complex protein assembly. Clathrin-coated pits are both small (on the scale of the ER and Golgi apparatus) and variable in size and as well as in the assembly stage. In both cases, by training ASEM with a large set of ground truth annotations generated by data augmentation from a very small number of hand-annotated objects, we could automatically identify essentially all the objects in the cell, despite the variable diameter of the nuclear membrane pore and the variable size and stage of completion of a clathrin-coated pit. Moreover, a model trained on plasma-membrane-coated pits identified coated pits in the Golgi and free clathrin-coated vesicles in the cytosol.
The osmium-uranyl staining in current FIB-SEM sample preparation, for both CF and HPFS, preferentially marks lipid headgroups, proteins, and nucleic acids. Although with the training set used here, the model did not distinguish between clathrin-coated pits and caveolae, the eye clearly picks up the much heavier staining of the former (Fig. 7). The model correctly retrieved clathrin-coated vesicles, as well as coated pits in the TGN, and distinguished them from COPI and COPII vesicles that carry cargo between the Golgi apparatus and the ER, perhaps because they are smaller structures, of substantially sharper curvature than the clathrin-coated structures the model had learned to recognize. How well the model will find protein-dominated structures—e.g., virus assembly intermediates—remains to be determined.
Imaging the entire volume of a single cell at ∼5 nm resolution can answer questions that are much harder to tackle by methods such as cryo-tomography that access at somewhat higher resolution only a small slice of a cell. One example is our finding that nuclear pores vary in size across the nuclear membrane and hence that the variability identified by cryo-tomography is present at an arbitrary time point in a single cell. The deep learning protocols we have developed and the readily implemented and freely accessible analysis tools we provide form an experimental pipeline that will run entirely on commercially available workstations. We suggest that EM volume imaging will prove to be a powerful complement to fluorescence volume imaging afforded by lattice light-sheet microscopy (Chen et al., 2014; Gao et al., 2019; Liu et al., 2018).
Material and methods
Chemical fixation, dehydration, and embedding
Cells plated on glass coverslips were processed for chemical fixation (CF) by incubation for 30 min at room temperature with 0.2% glutaraldehyde (Cat.16220; Electron Microscopy Science) and 2.5% paraformaldehyde (PFA, Cat. 15700; Electron Microscopy Science) dissolved in 0.1 M PIPES buffer (pH 7.4, Cat. P6757; Sigma-Aldrich), followed by a rinse with 0.1 M PIPES buffer. A 2% OsO4 aqueous solution (Electron Microscopy Sciences) dissolved in 0.1 M PIPES, pH 7.4 was used to stain the cells for 1 h at RT, followed by incubation for another 1 h at RT in a solution containing 2.5% potassium ferrocyanide (Sigma-Aldrich) in 0.1 M PIPES, pH 7.4. The cells were rinsed three times at 5-min intervals with deionized ultrapure water followed by a 30-min incubation at RT with a filtered (Whatman, 0.2 μm) freshly prepared solution of 1% thiocarbohydrazide (Electron Microscopy Sciences) made by dissolving it at 60°C for 15 min. The cells were again rinsed three times at 5-min intervals, followed by another incubation with 2% OsO4 aqueous solution for 1 h at RT. The cells were again rinsed three times at 5-min intervals with ultrapure water, followed by two rinses with 0.05 M maleate buffer, pH 5.15 (Sigma-Aldrich), and finally incubated with 1% uranyl acetate (Electron Microscopy Sciences) dissolved in 0.05 M maleate buffer pH 5.15 for 12 h at 4°C.
A resin mixture containing methylhexahydrophthalic anhydride (J&K Scientific) and cycloaliphatic epoxide (ERL 4221; Electron Microscopy Sciences) at a weight ratio of 1.27:1, mixed with the catalyzing agent (Hishicolin PX-4ET; Nippon Chemical Industrial) at a 1:100 ratio by volume, was prepared in a water bath sonicator at RT for 15 min.
During the same period, the glass coverslips with the attached CF samples were placed face up on wet ice and then rinsed twice for 5 min with ultrapure water, followed by dehydration using a graded series of ethanol solutions (30, 50, 70, 90%) each step lasting 3 min, then three washes in 100% absolute ethanol for 10 min, ending with three washes with anhydrous acetone (Sigma-Aldrich) for 10 min at RT.
The glass coverslips with the attached, dehydrated CF samples immersed in anhydrous acetone were placed in a wide-mouth glass jar, mixed with the resin at a 1:1 volumetric ratio, and gently rocked on a plate rocker for 12 h at room temperature. The resin mixture was then removed by aspiration and replaced with 10 ml of freshly prepared resin mixture and further incubated with gentle rocking for another 2 h; this step was repeated thrice, each time with freshly prepared resin. Finally, the glass coverslips with the attached cells were placed on top of cut-off caps from 1.5 ml Eppendorf tubes containing a freshly prepared resin that was oriented with the cells toward the cap, and the resin was allowed to polymerize for 12 h at 100°C. Upon resin hardening, the caps were immersed in boiling water for 5 min and then quickly transferred into liquid nitrogen leading to the separation of the glass coverslip from the resin and the retention of the cells in the polymerized resin.
High-pressure freezing, freeze-substitution, and embedding
Cells were plated on 6 × 0.1 mm sapphire disks in MEM (Corning 10009CV) supplemented with 10% fetal bovine serum (S11150; Atlanta Biologicals). Two sapphire discs (616-100; Technotrade international), one or both containing attached cells facing inward, were separated by a 100-μm stainless steel spacer (1,257-100; Technotrade international) and processed for high-pressure freezing on a Leica EM ICE high-pressure freezer (Leica Microsystems). Following high-pressure freezing, the sapphire discs were placed under liquid nitrogen and transferred into the top of cryotubes placed in liquid nitrogen and containing frozen 2% OsO4, 0.1% uranyl acetate, and 3% water in acetone; freeze-substitution (FS) was carried using an EM AFS2 automatic freeze substitution device (Leica Microsystems) according to a preprogrammed FS schedule (−140 to −90°C for 2 h, −90 to −90°C for 24 h, −90–0°C for 12 h and 0–22°C for 1 h). Samples were then removed from the AFS2 device; rinsed three times in anhydrous acetone, three times in propylene oxide (Electron Microscopy Sciences), and three times in 50% resin (24 g Embed 812, 9 g DDSA, 15 g NMA, 1.2 g BDMA; 14121; Electron Microscopy Sciences); dissolved in propylene oxide; and finally transferred into embedding molds (EMS 70900) containing 100% resin; the resin was then allowed to polymerize for 48 h at 65°C. The sapphire disc was then separated from the resin block by sequential immersion in liquid nitrogen and boiling water.
FIB-SEM imaging
The polymerized resin blocks were cut from the molds and glued, with the free face facing away, onto the top of aluminum pin mount stubs (Ted Pella) using conductive silver epoxy adhesive (EPO-TEK H20S; Electron Microscopy Sciences). The free face was then coated with carbon (20 nm thickness) generated from a high-purity carbon cord source (Electron Microscopy Sciences) using a Quorum Q150R ES sputter coater (Quorum Technologies), and the resin block was loaded on the microscope specimen stage of a Zeiss crossbeam 540 microscope for FIB-SEM imaging. After eucentric correction, the stage was tilted to 54° with a working distance of 5 mm for the coincidence of the ion and electron beams. A cell of interest was located on the free face of the resin block by SEM, after which a thin layer of platinum was deposited using the gas injection system. A coarse trench was then milled adjacent to the cell using the 30 kV/30 nA gallium ion beam. This block face was polished with a 30 kV/7 nA gallium beam before starting the interlaced sequence of FIB milling with a 30 kV/3 nA gallium beam and SEM imaging with a 1.5 kV/400 pA electron beam advanced in 5-nm steps. The X/Y pixel size was 5 nm to create isotropic voxels. For samples prepared by HPFS, we added registration marks on top of the platinum layer generated with a 1.5 kV/50 pA gallium beam, followed by contrast enhancement of the marks by irradiation with a 1.5 kV/5 nA electron beam, and a final deposition of a second platinum layer. FIB-SEM images were collected using the Inlens detector with a pixel dwell time of 10–15 μs. The FIB-SEM images were aligned after acquisition with the Fiji plugin Register Virtual Stack Slices https://imagej.net/plugins/register-virtual-stack-slices using the translation (Feature extraction model and Registration model) and shrinkage constraint options (Schroeder et al., 2021).
FIB-SEM data at 10 nm were acquired using a backscatter electron detector (EsB) with a grid voltage set to 808 V to filter out scattered secondary electrons, with a dwell time of 3 μs, line averaging of 8, and a pixel size of 10 × 10 nm (X/Y). FIB milling was performed with the 30 kV/30 nA gallium ion beam in 10-nm steps to create isotropic 10 × 10 × 10 nm (XYZ) voxels. The sequential FIB-SEM images were registered using the Fiji plugin StackReg with Rigid Body transformation.
Ground truth annotation
All our ground truth annotations were binary masks located at least 47 voxels away from the boundaries of the 3D FIB-SEM image. This ensured that training of the neural network was done with a sufficient semantic context within the image, resulting in improved model predictions.
Ground truth annotations for mitochondria, Golgi apparatus, and ER were generated by using the carving module of Ilastik (Berg et al., 2019) or our graph-cuts-based semiautomated annotation tool, and when needed, by further manual editing using VAST (Berger et al., 2018) to remove voxels that did not belong to the structure of interest or to add voxels for regions that had not been included in the original binary mask.
Ground truth annotations for the relatively complex three-dimensional substructure of the Golgi apparatus included membrane boundaries and the lumen for the characteristic three to six closely stacked fully enclosed membrane lamellae, the fenestrated and somewhat swollen trans-Golgi network stack, and a variable number of small vesicles clustered next to the Golgi apparatus. They were created with our semiautomated graph-cut annotation tool.
Ground truth annotations for endocytic clathrin-coated pits and for caveolae were manually generated in consecutive planes using VAST by drawing along the contours following the plasma membrane invaginations characteristic of these structures.
Ground truth annotations for nuclear pores were manually generated plane by plane using VAST by drawing along the contours of the nuclear outer and inner membrane adjacent to the nuclear pore.
Graph-cut annotation tool
We developed a new semiautomated tool to aid an expert annotator in marking sparse and coarse labels in a subvolume, one plane at a time so that the annotator can define high-level seeds required to generate ground truth annotations for a chosen organelle and separate it from the background (see Fig. S2). Once the seeds are marked, the tool, at the push of a button, uses mathematical techniques (detailed below) to combine them with the grayscale values in the volume to infer the structure of the organelle, thus resulting in a semiautomated segmentation.
Our tool operates on the full 3D volume (rather than 2D slices). We observed that manual annotation with only a few sparse 2D brush strokes (seeds) in only a few 2D arbitrarily spaced planes still results in satisfying volumetric annotations. Furthermore, this tool allows the annotator to visualize the volume plane-by-plane and define seed voxels in a plane as either part of the organelle or the background. Scrolling through, the annotator can seed a few planes at arbitrary intervals over the entire stack. It also allows views along all three axes so that the annotator can look down the z-axis to mark the xy-plane, and similarly for the y- and x-axes.
The technique of graph-cuts-based segmentation was adopted to generate segmentations from these seeds. The seeds were defined on manually extracted volumes from cells. Even though the seeds were coarse, the annotator took care not to mislabel any voxel. The maxflow algorithm (Boykov et al., 2001) was then employed to segment organelles based on these annotations. This tool was written in Python and adapted to suit the annotation needs associated with 3D FIB-SEM data. It is publicly available at https://github.com/kirchhausenlab/gc_segment, accompanied by detailed usage instructions and best practices.
The next step of the semiautomated segmentation is a reduction of problem complexity. To get a segmentation based on the seeding, each voxel must be assigned either an organelle label or a background label. At this point, we note that the volumes with which we work are characteristically large—a volume of 1 cubic micrometer contains 8 million voxels at a resolution of 5 × 5 × 5 nm. We also observe that as organelles are contiguous objects in the volume, a group of nearby voxels has a high chance of belonging to the same organelle. We hence merge nearby voxels to form supervoxels using the SLIC algorithm (Achanta et al., 2012), so that we can work with ∼103 supervoxels instead of ∼106 voxels—a reduction of three orders of magnitude. Under this strategy, supervoxels were formed by grouping adjacent voxels together based on a similarity criterion, which for our problem setting was chosen as the agreement in grayscale values.
A postprocessing step has been included in this tool that can modify the resulting segmented foreground voxels by fitting to the foreground voxels a univariate Gaussian mixture model based on its grayscale values. This optional postprocessing can help remove outlier voxels with the minimal additional overhead of computation time.
The aim of the graph-cuts-based strategy is therefore to define and optimize an objective energy function over the space of labels , where the labels 0 and 1 represent organelle and background respectively and is the number of supervoxels in the volume. Each point in this space determines whether a supervoxel is considered an organelle or a background and hence represents a segmentation of the volume. The objective energy function for our problem was formulated based on early work on graph-cut segmentation in computer vision (Boykov and Kolmogorov, 2004).
Here, δ(Au, Av) is set equal to 1 if and 0 otherwise. denotes the set of supervoxels adjacent to each other and hence deemed neighbors. Adjacency implies a shared boundary between the two supervoxels. , , and are weights given to the individual terms. and represent unary terms of the energy function, as they depend only on one supervoxel, while represents pair-wise terms.
The unary terms are defined as follows. The terms are determined by the aggregate grayscale values of the supervoxels and their agreement with the seeded foreground and background voxels (the vector ), as in earlier work (Boykov and Kolmogorov, 2004; Boykov et al., 2001), and the terms are determined by the distance of the voxels from the nearest seeding of organelle and background. The pair-wise terms are also defined according to earlier work, based on the distance between the two supervoxels, defined as the distance between the arithmetic center of the two supervoxels. As the defined energy function is submodular, it can be optimized using graph cuts (Kolmogorov and Zabin, 2004). An efficient algorithm to optimize such functions, maxflow (Boykov and Kolmogorov, 2004), was used to find the optimum vector . There are hyperparameters in our formulation, namely , , and . These were empirically defined as β = 1, γ = 1, and λ = 10. It should be noted that these values can be tuned according to the user’s needs and observations.
Data preprocessing for deep learning
Cell image stacks underwent the following steps before they were ready for training: (1) Conversion from TIFF format to the block-wise storage format ZARR. The size of a FIB-SEM dataset corresponding to a stack of registered TIFF files (∼2,000 planes) was about 20 GB. These TIFF stacks were converted into a ZARR 3-D compressed array (https://zenodo.org/record/7115955) to increase the efficiency for further preprocessing steps and, most importantly, for neural network training. (2) Cropping of the dataset to exclude empty regions outside the cell and to speed up all further preprocessing steps. (3) Block-wise adjustment of brightness and contrast with 3D contrast-limited adaptive histogram equalization (CLAHE, [Zuiderveld, 1994]) using scikit-image.exposure.equalize_adapthist with kernel size 128 and clip limit 0.02 (see Fig. S4). (4) Application of morphological operations to automatically clean up ground truth annotations based on biological assumptions was implemented with the python libraries scikit-image.morphology (van der Walt et al., 2014) and scipy.ndimage (Virtanen et al., 2020). For mitochondria and Golgi apparatus, small objects were removed (<27 voxels); for ER, holes were removed (<20,000 voxels, corresponding to 0.0025 µm3); no clean-up was required for nuclear pores or clathrin-coated pits. (5) Automatic creation of a coarse voxel-wise mask to mark voxels outside of the cell using a combination of operations from the python libraries scikit-image.morphology and scipy.ndimage. The parameters and combination of operations were adapted visually to each dataset. Operations included intensity thresholding, binary opening and closing, filling small holes, and removing small objects. (6) Optional: Correction for systematic biases in annotations. We observed that our semiautomatic annotations carry biases that can be corrected automatically. For mitochondria and Golgi apparatus, most of the annotations did not include the membrane, which we wanted to consider as part of the organelle. Note that this correction depended on the characteristics of a specific dataset (e.g., the contrast of membranes); mitochondria annotations were dilated by three voxels (15 nm); Golgi apparatus annotations were dilated by one voxel (5 nm); and ER, nuclear pores, and clathrin-coated pits annotations were not dilated. (7) Defining a metric exclusion zone. Although step (6) allowed us to add most of the organelles’ membranes to the annotation, the ground truth was often not voxel accurate at the organelle boundaries. A neural network model trained with such data cannot produce voxel-accurate predictions at the organelle boundaries, leading to misleading evaluation scores (e.g., F1, see Fig. S3 E). Following previous works (Haberl et al., 2018; Lucchi et al., 2012), we avoided this issue by defining an exclusion zone around our semiautomatic imprecise annotations created by dilating and eroding the annotations and taking the logical difference between the two outcomes. The size of both dilation and erosion depends on the specific structure, as follows: four voxels for mitochondria, two voxels for Golgi apparatus, one voxel each for ER, and three voxels for dilation plus one voxel for erosion for clathrin-coated pits and nuclear pores.
All operations required only local context, meaning that they could be applied block-wise, and the computation could be parallelized to multiple CPU cores. To avoid artefacts at the block boundary, we provided sufficient spatial context to each block with the python library DAISY (https://github.com/funkelab/daisy), which was used for multiprocess computation on all cores of a CPU. These computations were performed on Intel Xeon workstation processors with 20–40 physical cores. Detailed instructions on the use of the preprocessing pipeline are provided at https://github.com/kirchhausenlab/incasem#Prepare-your-own-ground-truth-annotations-for-fine-tuning-or-training.
Deep learning
Model architecture
A 3D U-Net (Çiçek et al., 2016) based on the architecture used in Funke et al. (2019) was defined, with three downsampling layers with a factor of two, and two convolutional layers on each downsampling level. Refer to Fig. S3 A for details. In total, the network had ∼six million parameters. It was implemented in PyTorch (Paszke et al., 2019).
Training: Overview of pipeline
The pipeline to feed blocks to the neural network was based on Buhmann et al. (2021) and implemented using GUNPOWDER (https://github.com/funkey/gunpowder), a library that facilitates machine learning on large multidimensional arrays.
We trained one model per organelle, i.e., for model training data, foreground refers to voxels corresponding to only one type of organelle. For each iteration during the training phase, a block of 204 × 204 × 204 voxels was randomly sampled from the electron microscopy dataset, together with the corresponding block of ground truth. The blocks were augmented by voxel-wise transformations, e.g., random intensity shifts, and geometric transformations, e.g., random rotations and deformations. The blocks were processed through the network, which returned as an output block a 3D probability map of 110 × 110 × 110 voxels, centered with respect to the larger input block. The input blocks contained an additional 47 voxels per side to provide the context required by our convolutional neural network architecture. The output probability map was compared with the ground truth using cross-entropy loss, which was minimized by iteratively updating the model parameters by using the Adam optimizer (Kingma and Ba, 2014).
Training: Data sampling
Our dataset annotations were highly imbalanced. As our structures of interest were small, only a few voxels formed the so-called foreground (FG), with a large portion of the dataset consisting of arbitrary background (BG = 1-FG; e.g., cytosol, nucleus, other organelles). Imbalanced datasets are known to be problematic for convergence of neural network training, and we confirmed this empirically while working with our datasets. As a rule of thumb, it is desirable to sample blocks having a foreground to background ratio roughly equivalent to the global ratio of the two. To make use of all available training data, while keeping the number of unbalanced training blocks as low as possible, we implemented the following scheme (using operations available in GUNPOWDER): (1) Reject blocks that contain more than 25% out-of-sample voxels. (2) Calculate the FG/BG ratio for each incoming block. (3) Reject a block with probability 0.9 if fewer than 5% of the voxels in it (ratio in step [2]) are labeled as foreground.
Training: Data augmentation
It was impractical to process and store tens of thousands of augmented FIB-SEM blocks required to train the 3D neural network model. Instead, during each training cycle, we augmented the number of ground truth annotations by randomly applying the transformations listed in Table S8 to the training block.
Training: Pipeline details
After data augmentation, we shifted the scale of the data in the input block (204 × 204 × 204 voxels) such that the input intensities were in (−1, 1). Each block accepted by the neural network was then propagated through the network leading to outputs of spatial dimensions 110 × 110 × 110 voxels, centered with respect to the larger input block. The neural network assigned complementary FG and BG probability to each voxel. The probability map was then compared to the ground truth annotations with the binary cross-entropy loss. We balanced the loss contribution of foreground and background voxels inversely proportional to their occurrence, clipped at a value of 1:100. The training loss was backpropagated, and the network parameters were updated using the Adam optimizer (Kingma and Ba, 2014) with 0.00003 learning rate and 0.00003 weight decay. The network parameters were saved at the end of every 1,000 training iterations.
Training: Computational requirements
Eight CPU cores were used in parallel for data fetching and augmentation, while a single GPU (A100; Nvidia on a DGX-A100 system) was used for training. Typically, a training iteration lasted 1 s, and 100,000–150,000 iterations (28–42 h, including periodic validation tests) were sufficient to train our 3D neural network model. A training session could also be done with a standard GPU workstation equipped with an Nvidia GPU with 12-GB GPU memory and 500-GB CPU memory. Presumably, workstations with 64-GB CPU memory can also be used since our training pipeline processes out-of-memory datasets.
Validation: Procedure
To avoid overfitting, we assessed the model’s performances during the training phase on both the training dataset and validation dataset, where the latter dataset was not used to update the model parameters. We saved every 1,000 iterations of the training model, froze their weights, and calculated the loss on a small set of ground truth blocks. These were hold-out blocks from the cells that contained the training data or blocks originating from naïve cells not represented in the training data. By comparing training and validation losses (see plots in Fig. S3, B and C), we usually identified three different regimes: under-fit, fit, and over-fit. When the model was under-fit, both training and validation loss decreased with the training iteration. In the fit regime (Fig. S3, B and C, in gray), typically starting after more than 20,000 training iterations, the validation loss was approximately constant, while the training loss was slightly reduced. In the over-fit regime, the training loss continued to drop, but the validation loss started to rise. We considered the model saved at the training iteration in the middle of the fit regime to be the one that could best generalize, i.e., make optimal predictions on previously unseen data. This is a standard procedure in Machine Learning, known as “early stopping.”
Validation: Performance metrics
The performance of the models obtained by the 3D U-net neural networks was determined by comparing the predicted binary segmentation with respect to ground-truth using the following three metrics: (1) precision (percentage of voxels predicted as intracellular structure that is the substructure), (2) recall (percentage of substructure voxels correctly predicted as substructure), and (3) F1 index score (harmonic mean of precision and recall, see Fig. S3 E). These metrics were also used by other state-of-the-art supervised learning methods, such as COSEM, allowing for a quantitative comparison.
Prediction: Data preprocessing
As for training, the segmentation of new FIB-SEM datasets required certain preprocessing steps (refer to Data preprocessing for deep learning for details): (1) Conversion from TIFF format to block-wise storage format ZARR. (2) Crop the dataset to exclude empty regions outside the cell. (3) Create an approximate voxel-wise mask to mark voxels outside of the cell. (4) Image data normalization with CLAHE (see Fig. S4).
Prediction: Pipeline
As the first step, the trained model at the iteration determined by early stopping (see Validation: Procedure, above) was loaded with frozen weights. The dataset to segment was scanned block by block and fed into the model, without performing data augmentation. Since the architecture of the 3D U-Net neural network is fully convolutional and since each predicted voxel has access to sufficient context, we could produce the predictions block-wise independently and in parallel.
The model performed a forward pass, producing as output a 3D voxel-wise probability map, saved to disk in ZARR format. A threshold of 0.5 was applied to the predicted probability map to extract a binary segmentation map (organelle/background). Although our pipeline allowed the user to set this threshold to an arbitrary value from zero to one, we set it to the default value of 0.5 in every experiment to avoid introducing any postprocessing bias. The segmentation map was later visualized in neuroglancer (https://github.com/google/neuroglancer) and superimposed on the electron microscopy image. Finally, we converted the predicted segmentations back to TIFF format and reverted the initial dataset cropping to obtain a segmentation that was globally aligned with the originally acquired image stack from the microscope.
Prediction: Computational requirements
A prediction was performed on a single GPU (A100; Nvidia on a DGX-A100 system), backed by multiple CPU cores and employed to parallelly load and preprocess the data. When using five CPU cores, we achieve a prediction throughput of ∼1.6 M voxels per second, roughly corresponding to the size of 1153 voxels for one block of actual prediction as depicted in our Unet architecture in Fig. S3. Hence the prediction of one cell image stack, acquired at 5 nm isotropic resolution, typically took between 30 and 90 min, depending on its volume. A similar prediction could also be done with a standard GPU workstation equipped with a Nvidia GPU with 12 GB GPU memory and 500 GB CPU memory. Presumably, workstations with 32-GB CPU memory can also be used since our training pipeline processes out-of-memory datasets.
Fine-tuning
To fine-tune a trained model on a naïve cell, we performed the following steps: (1) One block of ground-truth block (minimum 204 × 204 × 204 voxels) within the cell to fine-tune was annotated. (2) A model previously trained on the same organelle, whose weights were frozen at the first train iteration of the fit region, was loaded for continued training. (3) A training of 15,000 iterations was launched, using as training data only the newly prepared ground-truth block. All fine-tuning training hyperparameters were set identical to the original training. (4) The model was saved every 1,000 iterations. The best model iteration was picked based on the original validation dataset from the fine-tuning target cell.
Details to perform fine-tuning training using our pipeline are provided at https://github.com/kirchhausenlab/incasem#Fine-tuning.
Brief example of fine-tuning
We started by preparing a small ground-truth block of the cell to fine-tune. As the volume of the additional ground truth was much smaller than the volume of the ground truth fed to the pretrained model (from ∼1/69 to ∼1/3, Table S10), the additional annotation effort was not very demanding. In the case of CF datasets, for the fine-tuning of mitochondria in Cell 3 BSC-1 and Cell 6 SUM 159, we loaded the segmentation map performed by the pretrained model (model 1847) and refined it by manual editing with VAST. Conversely, to fine-tune HPFS OpenOrganelle datasets in the prediction of mitochondria and ER, we picked one additional COSEM ground-truth crop, previously excluded from the training and validation data. We loaded the pretrained model at the first training iteration within the fit regime (i.e., the iteration after which the validation loss was stable, Fig. S3, B and C); for example, in Fig. S5 A, for the CF Mitochondria model, we loaded the model at the training iteration 95,000 (Fig. S3 B). We resumed training on the model by using only the additional ground truth block and data augmentation. Typically, after a few thousand training iterations (about ∼1/15 of the training iterations needed to produce the pretrained models), the fine-tuned model learned to segment the fine-tune dataset more precisely, with an increased overall F1 score (Table S10). We noticed that fine-tuning was beneficial mainly when the initially trained model behaved poorly, such as in the case of HPFS ER and Cell 21 Jurkat-1, for which F1 increased by 0.21.
To understand how fine-tune training helped to improve segmentation, we compared the segmentation performed by the pretrained model versus the one produced by the fine-tuned model (Fig. 5 D). In the case of CF, mitochondria, only a few portions of mitochondria were occasionally missed by the pretrained model, but they were eventually retrieved by the fine-tuned models. In HPFS ER, fine-tuning reduced the number of false positives, resulting in a neater segmentation map, suitable for further biological studies.
To quantify the impact of fine-tune training, we calculated precision and recall in addition to F1 (Fig. S5). Typically, fine-tuning enhanced precision without affecting recall: the fine-tuned model learned to classify cell components that looked like the organelle under study, but did not actually belong to the same semantic class, reducing the number of false positives.
At the baseline, we compared the fine-tuned model with one having randomly initialized weights and trained using the same (small) ground truth volume. We found that for most datasets and organelles (ER Cell 21 Jurkat-1, ER Cell 22 Macrophage-2, mitochondria Cell 21 Jurkat-1, mitochondria Cell 3 BSC-1, and mitochondria Cell 6 SUM 159), the models trained using randomly initialized weight reached substantially lower F1 scores, even when trained much longer, with up to 200,000 training iterations. Only in one case (mitochondria Macrophage-2),the model achieved approximately the same F1 score, but only after training for 160,000 iterations, 20 times more than the number required by the corresponding fine-tuned model.
We concluded that fine-tune training is a useful tool to apply whenever the segmentation of a naïve cell falls short. By taking advantage of pretrained models and preparing a small ground truth volume, one can train more accurate models at a small fraction of the ground truth annotation and computational costs usually required.
Estimate of time effort required to annotate ground truths
We evaluated our annotation times for all cells and organelle type (Table S4). The total annotation time for the given volumes was 120 h for 176 μm3 (mitochondria), 100 h for 166 μm3 (Golgi apparatus), and 120 h for 92 μm3 (ER). On average, this equates to roughly 0.8 h per cubic micron per organelle for ASEM. In contrast, the COSEM paper (Heinrich et al., 2021) states that annotating a block of 1 cubic micron with 35 organelle classes took one annotator 2 wk (∼100 h), which equates to 2.8 h per cubic micron per organelle, thus being ∼3.5 times slower than ASEM.
Analysis of nuclear pores
Automated orientation of nuclear pores on the nuclear envelope
We determined the nuclear pore membrane diameters from top-down FIB-SEM views toward the nuclear envelope of each of the nuclear pores. This orientation process was automated with the following steps. First, we generated a 3D binary mask corresponding to the predicted probability with a threshold of 0.5 for each one of the nuclear pores identified by the 3D U-net nuclear pore model (Fig. S6 A). Second, we determined the volume, principal axis, and centroid coordinates for each mask. A filtering step was included to eliminate masks with a small volume or short axis due to incompleteness of the predicted mask. Third, we used median filtering of the 3D point cloud data to remove outliers, thus creating a virtual “low resolution” 3D surface of the nuclear envelope by alpha-shape triangulation of the centroids (Akkiraju et al. 1995; Fig. S6 B, top). Fourth, we obtained a vector normal to each triangle within the triangulation (Fig. S6 B, bottom). Finally, we used the angular information of this vector to reorient the coordinates of the raw image of the nuclear pore closest to the triangulation to position the nuclear membrane on a view normal to the observer (Fig. S6 C).
Determination of nuclear pore membrane diameter
The diameter of the membrane pore, defined by the contact between the nuclear membrane and the pore opening, was determined from the distance separating the two peak signals measured in the FIB-SEM image along a line transecting the middle of the nuclear envelope immediately surrounding the nuclear pore (d1 and d2 in Fig. S6 D). The nuclear pore membrane diameter was expressed as the median of 18 radial measurements 10° apart. This calculation increased the precision of the measurement by taking advantage of the known radial symmetry of the nuclear pore and the surrounding nuclear envelope on the axis normal to the nuclear envelope; the standard deviation for each pore diameter measurement (i.e., experimentally determined uncertainty) was 6 nm.
Analysis of clathrin-coated pits and coated vesicles
The model trained on endocytic clathrin-coated pits in cell 12 and cell 13 was used to predict clathrin-coated structures in cells 12, 13, 15, and 17. The predictions were gated at a probability of 0.5 and the corresponding masks were then used to locate the clathrin-coated structures. These structures were classified as endocytic clathrin-coated pits and coated vesicles if they were located at the plasma membrane or within 400 nm, respectively; “secretory” coated pits and coated vesicles denote the remaining similar structures located in the cell interior. Each prediction was confirmed by the visual inspection of the corresponding image along the three orthogonal directions.
Measurements of neck, height, and width from the pits (Fig. S8 A), and major and minor axis of the ellipses best fitting the pits and coated vesicles (Fig. S8, A and B) were determined by the following sequential steps: (1) select the view displaying the largest outline by inspection of nine consecutive planes along each of the three orthogonal views centered on the centroid of the pit or vesicles; (2) manually measure the neck height and widths of the pits; (3) establish the outline of the pit or vesicle in the section chosen in the first step; the darker pixels where the pit or vesicle was present were selected manually (Fig. S8 C, white square in the left panel), segmented into a binary mask with an Otsu intensity threshold (Otsu, 1979), and skeletonized (Fig. S8 C, middle panel); (4) establish the ellipse best fitting the skeletonized outline of the pit or vesicle (Fig. S8 C, right panel); and (5) obtain major and minor axis of the ellipse.
Statistical analysis
The normality of the nuclear pore size distribution was examined using the Shapiro–Wilk test (Fig. 6 C). The comparison of size distributions between nuclear-pore diameters from values determined experimentally and from simulated values based on the experimental median value with an uncertainty of 6 nm using the nonparametric Kolmogorov–Smirnov test showed they were statistically different (P < 0.0001).
Online supplemental material
Fig. S1 shows the ground truth annotation workflow for mitochondria. Fig. S2 shows the ground truth annotation workflow for ER and Golgi apparatus. Fig. S3 shows the 3D-Unet architecture, examples of network behavior during training, and F1 as a metric to compare ground truth annotations with model predictions. Fig. S4 shows the use of CLAHE to equalize the contrast of FIB-SEM images. Fig. S5 shows the comparison of validation metrics of neural models predicting mitochondria, ER, and Golgi apparatus. Fig. S6 shows the steps to determine the diameter of the nuclear pore membrane. Fig. S7 shows the three-dimensional distribution of nuclear pores on the nuclear envelopes of Cells 15 and 17. Fig. S8 shows the definition of metrics used to characterize clathrin-coated structures. Fig. S9 shows the characterization of clathrin-coated pits and coated vesicles. Table S1 is the list of cells used in this study. Table S2 shows the size of hold-out volumes containing ground truth annotations and their use for model training, validation, or prediction. Table S3 shows the types of data augmentation used in this study. Table S4 lists the procedures used to generate ground truth annotations. Table S5 shows the effect of CLAHE on prediction performance. Table S6 compares examples of predictive performance by models trained with data from one or two cells. Table S7 compares model performances using the ASEM (this study) and COSEM pipelines. Table S8 compares examples of predictive performance by models trained with data from cells prepared with the same or different fixation protocols. Table S9 shows the effect of resolution on the predictive performance. Table S10 is a summary of experiments to test the effect of fine-tuning.
Data availability
The datasets of raw and normalized FIBSEM cells images, ground truth annotations, probability maps predicted by the models, and corresponding segmentation masks are publicly available at the AWS ASEM bucket (https://open.quiltdata.com/b/asem-project).
The software and step-by-step instructions to use it are publicly available at https://github.com/kirchhausenlab/gc_segment (Graph-cut annotation tool) and https://github.com/kirchhausenlab/incasem (Deep-learning pipeline).
Trained neural network models are available at https://open.quiltdata.com/b/asem-project with usage instructions at https://github.com/kirchhausenlab/incasem.
Acknowledgments
We thank Jan Funke for initial guidance on the use of deep learning and for maintaining together with members of his laboratory the gunpowder and funlib libraries; Jose Inacio Da Costa Filho for testing our publicly available software; Arlo Sheridan for helpful discussions; Teresa Ribeiro-Rodrigues and Henrique Girão for providing Cells 1 and 2; Giovanni de Nola and Teresa Rodriguez for preparing Cells 1–4 for FIB-SEM imaging; Rasmus Herlo and Max Page for preparing Cell 5 for FIB-SEM imaging; Justin Houser for preparing Cell 6a-9 for FIB-SEM imaging; Gleb Shtengel and C. Shan Xu for preparing Cells10-13 for FIB-SEM imaging; S.C. Harrison for extensive editorial help; and members of the Kirchhausen laboratory for help and encouragement.
The research was supported by a National Institute of General Medical Sciences Maximizing Investigators’ Research Award GM130386 and a generous grant from SANA to T. Kirchhausen, B. Gallusser, and G. Maltese were supported in part by discretionary funds available to T. Kirchhausen. Acquisition of the FIB-SEM microscope was supported by a generous grant from Biogen to T. Kirchhausen. Acquisition of the computing hardware including the DGX’s GPU-based computers, CPU clusters, fast access memory, archival servers, and workstations that made possible this study were supported by generous grants from the Massachusetts Life Sciences Center to T. Kirchhausen and an equipment supplement to the National Institute of General Medical Sciences Maximizing Investigators’ Research Award GM130386. Construction of the server room housing the computing hardware was made possible with generous support from the PCMM Program at Boston Children’s Hospital. B. Gallusser and M. Weigert were supported by the EPFL School of Life Sciences and by generous funding from CARIGEST SA.
The authors declare no competing financial interests.
Author contributions: B. Gallusser, M. Sahasrabudhe, and T. Kirchhausen conceptualized the computational aspects of the project; B. Gallusser, G. Maltese, G. Di Caprio, and M. Sahasrabudhe created, implemented, and used the computational pipeline to train the neural network, obtain models, and make predictions; J. O’Connor helped set up and maintain the computing infrastructure; T.J. Vadakkan, B. Gallusser, G. Maltese, and G. Di Caprio generated and curated the ground truth annotations used for training. G. Di Caprio analyzed the data on nuclear pores and coated structures. A. Sanyal generated the cell lines used to image Cells 13a-17; A. Sanyal and E. Somerville prepared Cells 13a-17 for FIB-SEM imaging. T.J. Vadakkan used FIB-SEM to image most cells used in this study (Cells 1–17); T. Kirchhausen supervised the imaging experiments; B. Gallusser, G. Di Caprio, G. Maltese, M. Sahasrabudhe, M. Weigert, and T. Kirchhausen participated drafting the manuscript; and T. Kirchhausen contributed to the final manuscript in consultation with the authors.
References
Author notes
B. Gallusser, G. Maltese, and G. Di Caprio contributed equally to this paper.
Supplementary data
shows cells used in this study.