Mesenchymal cells such as fibroblasts are weakly polarized and reorient directionality by a lamellipodial branching mechanism that is stabilized by phosphoinositide 3-kinase (PI3K) signaling. However, the mechanisms by which new lamellipodia are initiated and directed are unknown. Using total internal reflection fluorescence microscopy to monitor cytoskeletal and signaling dynamics in migrating cells, we show that peripheral F-actin bundles/filopodia containing fascin-1 serve as templates for formation and orientation of lamellipodia. Accordingly, modulation of fascin-1 expression tunes cell shape, quantified as the number of morphological extensions. Ratiometric imaging reveals that F-actin bundles/filopodia play both structural and signaling roles, as they prime the activation of PI3K signaling mediated by integrins and focal adhesion kinase. Depletion of fascin-1 ablated fibroblast haptotaxis on fibronectin but not platelet-derived growth factor chemotaxis. Based on these findings, we conceptualize haptotactic sensing as an exploration, with F-actin bundles directing and lamellipodia propagating the process and with signaling mediated by adhesions playing the role of integrator.
Introduction
The importance of cell migration in development, immunity, wound repair, and cancer progression has long been appreciated. Unifying these various physiological and pathological contexts is a common design principle: the ability of migrating cells to change or maintain directionality as they monitor their microenvironment for spatial cues (Petrie et al., 2009; Bear and Haugh, 2014). However, different cell types use fundamentally distinct mechanisms to achieve this objective. Whereas amoeboid cells such as leukocytes exhibit a robustly polarized and excitable cytoskeleton, which only needs to be subtly perturbed by soluble cues to bias cell movement (i.e., in chemotaxis; Xu et al., 2003; Arrieumerlou and Meyer, 2005; Iglesias and Devreotes, 2012), mesenchymal cells such as fibroblasts exhibit weakly polarized migration phenotypes and respond to both chemical and physical cues (Lara Rodriguez and Schneider, 2013; Bear and Haugh, 2014). The latter include spatial gradients of immobilized, adhesive ligand density (haptotaxis) and of mechanical stiffness (durotaxis).
The most prominent cytoskeletal structure that drives cell locomotion is the lamellipod, a broad, fan-shaped protrusion with an F-actin–rich leading edge. The dendritic architecture of the leading-edge F-actin array is formed by integration of the Arp2/3 complex, which nucleates assembly of new actin filaments from existing ones and thus largely controls the rate of actin polymerization that drives lamellipodial protrusion (Rotty et al., 2013). This activity is in turn controlled by a host of signaling molecules, most notably the small GTPase Rac and the phospholipid phosphatidylinositol (3,4,5)-trisphosphate (PIP3), which cooperate to activate the SCAR–WAVE regulatory complex upstream of Arp2/3 (Lebensohn and Kirschner, 2009). PIP3 is produced by type I phosphoinositide 3-kinases (PI3Ks) and, like GTP-bound Rac, is focally enriched in protruding lamellipodia (Kraynov et al., 2000; Weiger et al., 2009). The weakly polarized morphology of fibroblasts is typically characterized by multiple lamellipodia, which exhibit intermittent protrusion and signaling and compete with one another to determine the overall direction of migration (Petrie et al., 2009; Weiger et al., 2010).
Previously, we characterized a mechanism by which fibroblasts execute large-scale changes in orientation by extension of nascent lamellipodia, which most often form by bifurcation of the dominant lamellipod; if the two branches successfully propagate to their fullest extent, a 90° turn is achieved (Welf et al., 2012). Our experiments revealed a specific role for PI3K signaling in lamellipodial spreading, which is required to maintain the propagation of the branches, whereas initiation of branching is PI3K independent. Accordingly, we found that increases in local PI3K signaling lag behind the acceleration of protrusion. These findings established the macroscopic morphodynamics of fibroblast migration that allow efficient reorientation of directionality, e.g., in response to external cues, but they also spurred a new set of questions aimed at the subcellular level. How do newly branched lamellipodia form? What determines the distinct directions of lamellipodial extension? Here, we show that F-actin bundles containing fascin-1, which often manifest as filopodia, seed the formation and set the orientations of nascent lamellipodia.
Filopodia are narrow, dynamic, finger-like protrusions with established roles in neuronal communication and development (Teddy and Kulesa, 2004), epithelial cell–cell adhesion (Vasioukhin et al., 2000; Wood et al., 2002), and cell motility, yet the precise contexts in which filopodia affect cell migration are largely unknown. The concept that filopodia generally serve as sensing organelles has been broadly speculated (Ridley et al., 2003; Mattila and Lappalainen, 2008), yet there is scant evidence supporting this notion except in the neuronal context (Davenport et al., 1993; Dent et al., 2011). In fibroblasts, it has been observed that filopodia seed the formation of distinct lamellipodia during the transition from isotropic to anisotropic spreading on an adhesive surface (Guillou et al., 2008), but the functional and mechanistic connections to random or directed locomotion are not yet established.
Critical to the assembly and stability of filopodia is the cross-linking of parallel actin filaments. Fascin is one of the major actin-bundling proteins in filopodia, though it has been shown to promote filopodia independent of its cross-linking function (Zanet et al., 2012). Among the three isoforms of fascin, only fascin-1 is broadly expressed in mesenchymal cells. Depletion of fascin-1 or impairment of fascin–actin binding significantly reduces the number of filopodia, whereas the constitutively active (S39A) actin-binding mutant increases the number and mean length of filopodia (Vignjevic et al., 2006; Li et al., 2010).
In this work, we used high-resolution imaging and analysis of subcellular dynamics, combined with directed migration assays, to elucidate a key function of F-actin bundles/filopodia in orchestrating lamellipodia during both random and directed migration of mesenchymal cells. Our results implicate both structural and signaling roles of fascin-containing bundles in this process and thus connect the subcellular and macroscopic morphodynamics of cell locomotion.
Results
Filopodia direct formation of newly branched lamellipodia
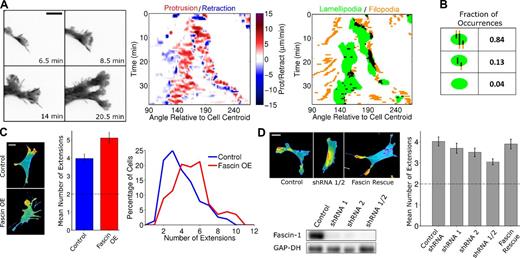
To study the formation of nascent lamellipodia, high-resolution total internal reflection fluorescence (TIRF) videos of cells randomly migrating on fibronectin were acquired and analyzed. Owing to the exquisite z resolution of TIRF illumination, adherent filopodia/microspikes of migrating NIH 3T3 fibroblasts were readily visible. We found that such filopodia typically preceded the emergence and oriented the protrusion of new lamellipodia, which most often branch off from existing ones (Fig. 1). Accordingly, spatiotemporal maps of protrusion and retraction activity show that large-scale branching of lamellipodia coincides with protrusion over filopodia (Fig. 1 A and Video 1). A method for identifying and tracking filopodia was devised (Fig. S1), allowing a clearer visualization of these structures in relation to cell shape dynamics. Lamellipodia were also identified automatically, based on a combination of morphological and protrusion criteria, and spatiotemporal overlap of lamellipodia and filopodia was assessed. Of the lamellipodia identified (n = 1,005, in 21 cells analyzed), 72% overlapped with filopodia; 55% of the lamellipodia overlapped with preexisting filopodia (Fig. 1 B).
Time-lapse video of filopodia directing nascent lamellipodia, as depicted in Fig. 1 A. TIRF microscopy images of an NIH 3T3 cell expressing EGFP-AktPH (pseudocolored) were acquired every 15 s for 132 min. Time stamp is shown in minutes, seconds, and milliseconds. Bar, 20 µm.
Time-lapse video of filopodia directing nascent lamellipodia, as depicted in Fig. 1 A. TIRF microscopy images of an NIH 3T3 cell expressing EGFP-AktPH (pseudocolored) were acquired every 15 s for 132 min. Time stamp is shown in minutes, seconds, and milliseconds. Bar, 20 µm.
Filopodia direct nascent lamellipodia. (A, left) Pseudocolored TIRF images of a NIH 3T3 cell expressing the EGFP-AktPH biosensor, representative of 28 cells in four independent experiments. The montage shows that filopodia template nascent lamellipodia, which typically branch from existing ones during random migration on fibronectin (see also Video 1). (center) The corresponding spatiotemporal map of protrusion/retraction velocity, overlaid on regions of morphological extension (gray; see Materials and methods) indicates the timing and angular locations of these branching events relative to the cell centroid (clockwise from the negative x axis). The colored arrowheads on both the montage and map show these events. (right) The accompanying lamellipod/filopod overlap map indicates pixels associated with putative lamellipodia, filopodia, or overlap of the two (black). (B) Analysis of lamellipod/filopod overlap. Indicated are the percentages of lamellipodial regions that emerged overlapping a preexisting filopod, emerged without a preexisting filopod but overlapped with at least one filopod thereafter, or never overlapped with a filopod. (C) TIRF montage, representative of 67 cells showing that local photoactivation of Rac (red spot) in NIH 3T3 cells induces protrusion and PI3K signaling preferentially over actin bundles (red arrowheads). Frames labeled with red circles show the cell immediately before activation of Rac at the indicated position. The other frames show the cell right before activation was ceased at that location and allowed to recover until the next “activation start” (see also Video 2). (D) Montage showing a time course after microinjection of Arp2/3 complex into Arp2/3-depleted IA32 MEFs (Wu et al., 2012). Red arrowheads indicate filopodia where protrusion initiated. Bars: (A) 10 µm; (C and D) 20 µm.
Filopodia direct nascent lamellipodia. (A, left) Pseudocolored TIRF images of a NIH 3T3 cell expressing the EGFP-AktPH biosensor, representative of 28 cells in four independent experiments. The montage shows that filopodia template nascent lamellipodia, which typically branch from existing ones during random migration on fibronectin (see also Video 1). (center) The corresponding spatiotemporal map of protrusion/retraction velocity, overlaid on regions of morphological extension (gray; see Materials and methods) indicates the timing and angular locations of these branching events relative to the cell centroid (clockwise from the negative x axis). The colored arrowheads on both the montage and map show these events. (right) The accompanying lamellipod/filopod overlap map indicates pixels associated with putative lamellipodia, filopodia, or overlap of the two (black). (B) Analysis of lamellipod/filopod overlap. Indicated are the percentages of lamellipodial regions that emerged overlapping a preexisting filopod, emerged without a preexisting filopod but overlapped with at least one filopod thereafter, or never overlapped with a filopod. (C) TIRF montage, representative of 67 cells showing that local photoactivation of Rac (red spot) in NIH 3T3 cells induces protrusion and PI3K signaling preferentially over actin bundles (red arrowheads). Frames labeled with red circles show the cell immediately before activation of Rac at the indicated position. The other frames show the cell right before activation was ceased at that location and allowed to recover until the next “activation start” (see also Video 2). (D) Montage showing a time course after microinjection of Arp2/3 complex into Arp2/3-depleted IA32 MEFs (Wu et al., 2012). Red arrowheads indicate filopodia where protrusion initiated. Bars: (A) 10 µm; (C and D) 20 µm.
We next asked whether or not the tendency to initiate protrusion around filopodia is a general property of lamellipodia. To test this, we used photoactivatable Rac (Wu et al., 2009) to locally induce Rac and Arp2/3 activation in NIH 3T3 cells. Although the spots of photoactivation were aimed near and not centered over visibly adhered filopodia, the ensuing protrusions occurred preferentially along those filopodia (Fig. 1 C and Video 2). This protrusion bias was seen 92% of the time (66/72), and PI3K signaling was enriched along the bundle in all but one of those instances. In a related experiment reported previously (Wu et al., 2012), Arp2/3-depleted fibroblasts were microinjected with purified Arp2/3 complex to induce lamellipodial protrusion globally. Analysis of this sequence confirms that the protrusions originated along filopodia before coalescing to form broad lamellipodia (Fig. 1 D).
Time-lapse video of PI3K signaling and protrusion during photoactivation of Rac near filopodia, as depicted in Fig. 1 C. TIRF microscopy images of an NIH 3T3 cell coexpressing mCerulean-PA-Rac and mCherry-AktPH (pseudocolored) were acquired every minute for 79 min. Time stamp shows hours and minutes. The red spots indicate the timing and area of photoactivation. Bar, 20 µm.
Time-lapse video of PI3K signaling and protrusion during photoactivation of Rac near filopodia, as depicted in Fig. 1 C. TIRF microscopy images of an NIH 3T3 cell coexpressing mCerulean-PA-Rac and mCherry-AktPH (pseudocolored) were acquired every minute for 79 min. Time stamp shows hours and minutes. The red spots indicate the timing and area of photoactivation. Bar, 20 µm.
Expression of the actin-bundling protein fascin-1 tunes cell morphology
To monitor filopod/lamellipod dynamics in better detail, we used fluorescent protein (FP)–tagged fascin-1 as a live-cell marker of F-actin bundles. In FP-fascin–expressing fibroblasts, we observed the same templating of branched protrusions as before, but here, we were not limited by the appearance of filopodia and could readily see F-actin bundles embedded behind the leading edge (Fig. 2 A, Video 3, and Fig. S2 A). The protrusions that emerge around bundles are labeled by FP-Lifeact, indicating F-actin network at the leading edge, characteristic of lamellipodia (Fig. S2 B). Analysis of lamellipodial overlap with F-actin bundles and filopodia in FP-fascin–expressing fibroblasts showed that 84% of the lamellipodia (n = 1,429, from 31 cells) overlapped with preexisting F-actin bundles or filopodia (Fig. 2 B). Conversely, most F-actin bundles/filopodia never overlap with lamellipodia during their lifetime (Fig. S2 C), indicating that emergence of lamellipodia is rare compared with the appearance and disappearance of adherent filopodia.
Time-lapse video of F-actin bundles directing lamellipodial protrusion, as depicted in Fig. 2 A. TIRF microscopy images of an NIH 3T3 cell expressing EGFP-fascin (inverted grayscale) were acquired every 30 s for 21 min. Time stamp shows minutes and seconds. Bar, 10 µm.
Time-lapse video of F-actin bundles directing lamellipodial protrusion, as depicted in Fig. 2 A. TIRF microscopy images of an NIH 3T3 cell expressing EGFP-fascin (inverted grayscale) were acquired every 30 s for 21 min. Time stamp shows minutes and seconds. Bar, 10 µm.
Modulation of fascin-1 expression tunes cell morphology. (A) TIRF montage (inverted grayscale) of a branching event in a NIH 3T3 cell expressing EGFP-fascin to mark F-actin bundles (left; see also Video 3), representative of 31 cells in 13 independent experiments (see Fig. S2 A for another example). The corresponding protrusion/retraction and lamellipod/filopod overlap maps are shown at right. (B) Lamellipod/filopod overlap analysis as shown in Fig 1 B; here, both putative filopodia and submembranous bundles labeled by FP-fascin were included in the analysis. (C) Representative TIRF images of NIH 3T3 cells expressing EGFP-AktPH or coexpressing mCherry AktPH and EGFP-fascin are shown. The bar graph shows the mean numbers of morphological extensions for the control (blue, n = 152) and fascin-overexpressing (red, n = 113) cells; error bars show 95% confidence intervals. The dashed line indicates that two extensions is the de facto minimum. The histogram shows the corresponding distributions. OE, overexpression. (D) NIH 3T3 cells were depleted of fascin-1 by two targeting shRNAs, expressed separately or in combination. The immunoblot confirms shRNA-mediated loss of fascin-1 expression levels relative to a nontargeting control shRNA, with GAPDH as a loading control. Representative TIRF images of these cells as well as of cells in which fascin-1 expression was rescued are shown. The bar graph shows the mean number of extensions quantified for each population of cells (n ≥ 140 for each condition); error bars show 95% confidence intervals. The dashed line indicates that two extensions is the de facto minimum. Bars: (A) 10 µm; (C and D) 20 µm.
Modulation of fascin-1 expression tunes cell morphology. (A) TIRF montage (inverted grayscale) of a branching event in a NIH 3T3 cell expressing EGFP-fascin to mark F-actin bundles (left; see also Video 3), representative of 31 cells in 13 independent experiments (see Fig. S2 A for another example). The corresponding protrusion/retraction and lamellipod/filopod overlap maps are shown at right. (B) Lamellipod/filopod overlap analysis as shown in Fig 1 B; here, both putative filopodia and submembranous bundles labeled by FP-fascin were included in the analysis. (C) Representative TIRF images of NIH 3T3 cells expressing EGFP-AktPH or coexpressing mCherry AktPH and EGFP-fascin are shown. The bar graph shows the mean numbers of morphological extensions for the control (blue, n = 152) and fascin-overexpressing (red, n = 113) cells; error bars show 95% confidence intervals. The dashed line indicates that two extensions is the de facto minimum. The histogram shows the corresponding distributions. OE, overexpression. (D) NIH 3T3 cells were depleted of fascin-1 by two targeting shRNAs, expressed separately or in combination. The immunoblot confirms shRNA-mediated loss of fascin-1 expression levels relative to a nontargeting control shRNA, with GAPDH as a loading control. Representative TIRF images of these cells as well as of cells in which fascin-1 expression was rescued are shown. The bar graph shows the mean number of extensions quantified for each population of cells (n ≥ 140 for each condition); error bars show 95% confidence intervals. The dashed line indicates that two extensions is the de facto minimum. Bars: (A) 10 µm; (C and D) 20 µm.
Based on the apparent role of F-actin bundles in cell morphodynamics, and considering that fascin-1 promotes the stability of filopodia, we hypothesized that cell shape is sensitive to fascin-1 expression level. To quantify cell morphology, we determined the number of extended structures (e.g., lamellipodia) in single images of many cells. When fascin-1 was overexpressed in NIH 3T3 fibroblasts, the mean number of morphological extensions was significantly increased relative to control cells, consistent with indications in other cell types (Yamashiro et al., 1998). The incidence of cells with six or more extensions was dramatically increased, along with reduced incidence of cells exhibiting three or fewer extensions, in the fascin-overexpressing population (Fig. 2 C); in contrast with cell morphology, overall migration speed was not significantly altered by fascin overexpression (Fig. S2 D). Conversely, depletion of fascin-1 by shRNA interference had the opposite effect on cell morphology. Expression of two shRNAs, which yielded ∼90% knockdown of fascin-1 each and >97% knockdown when combined, progressively reduced the mean number of morphological extensions (Fig. 2 D). To confirm the specificity of this approach, expression of human fascin-1 (which does not contain the sequences targeted in murine fascin-1) in the double-knockdown background fully rescued the normal phenotype (Fig. 2 D).
PI3K signaling colocalizes with F-actin bundles and is enriched in filopodia
Our previous work showed that PI3K signaling locally increases after the onset of protrusion, stabilizing nascent lamellipodia to affect macroscopic changes in cell morphology (Welf et al., 2012). With the present insight that F-actin bundles/filopodia initiate and orient this process, we asked whether or not these structures also harbor PI3K signaling and thus serve as primers for actin polymerization. Consistent with this idea, in NIH 3T3 fibroblasts coexpressing FP-fascin and FP-AktPH (Akt pleckstrin homology domain; a translocation biosensor for 3′ phosphoinositides), the two typically colocalize near the leading edge and especially at the bases of filopodia during the early phase of lamellipod formation, when protrusion accelerates (Fig. 3 A and Video 4).
Time-lapse video of PI3K signaling and fascin localization during protrusion over an F-actin bundle, as depicted in Fig. 3 A. TIRF microscopy images of an NIH 3T3 cell coexpressing mCherry-AktPH (left, pseudocolored) and EGFP-fascin (right, inverted grayscale) were acquired every 30 s for 67 min. Time stamp is shown in hours, minutes, and seconds. Bar, 10 µm.
Time-lapse video of PI3K signaling and fascin localization during protrusion over an F-actin bundle, as depicted in Fig. 3 A. TIRF microscopy images of an NIH 3T3 cell coexpressing mCherry-AktPH (left, pseudocolored) and EGFP-fascin (right, inverted grayscale) were acquired every 30 s for 67 min. Time stamp is shown in hours, minutes, and seconds. Bar, 10 µm.
F-actin bundles/filopodia harbor PI3K signaling. (A) Montage of PI3K signaling (pseudocolored) and fascin-1 localization (inverted grayscale) during protrusion over actin bundles in an NIH 3T3 cell (see also Video 4), representative of 31 cells viewed. Bar, 10 µm. (B) NIH 3T3 cells coexpressing mCherry-AktPH and TFP as a volume marker were used for ratiometric analysis. Mean enrichment ratios (means ± 95% confidence interval) of PI3K signaling in filopodia (left) and the associated histograms (right) for untreated cells (blue, n = 724 filopodia) versus cells treated with 1 µM PI3K-α inhibitor IV (red, n = 119 filopodia) are shown. (C, left) Montage of mCherry-AktPH/TFP ratio (pseudocolored) showing PI3K signaling enrichment preceding lamellipodial protrusion over an actin bundle. Bar, 5 µm. The percentage of instances in which PI3K signaling first increased in filopodia was determined for each cell (right; n = 27 cells, 194 protrusions total over actin bundles). The red dot indicates the mean ± 95% confidence interval across cells.
F-actin bundles/filopodia harbor PI3K signaling. (A) Montage of PI3K signaling (pseudocolored) and fascin-1 localization (inverted grayscale) during protrusion over actin bundles in an NIH 3T3 cell (see also Video 4), representative of 31 cells viewed. Bar, 10 µm. (B) NIH 3T3 cells coexpressing mCherry-AktPH and TFP as a volume marker were used for ratiometric analysis. Mean enrichment ratios (means ± 95% confidence interval) of PI3K signaling in filopodia (left) and the associated histograms (right) for untreated cells (blue, n = 724 filopodia) versus cells treated with 1 µM PI3K-α inhibitor IV (red, n = 119 filopodia) are shown. (C, left) Montage of mCherry-AktPH/TFP ratio (pseudocolored) showing PI3K signaling enrichment preceding lamellipodial protrusion over an actin bundle. Bar, 5 µm. The percentage of instances in which PI3K signaling first increased in filopodia was determined for each cell (right; n = 27 cells, 194 protrusions total over actin bundles). The red dot indicates the mean ± 95% confidence interval across cells.
Is PI3K signaling enriched within filopodia as well? This question is not straightforward, given the size of these structures: ∼100 nm in diameter, below the resolution limit and comparable to the depth of TIRF illumination. Therefore, we assessed enrichment of PI3K signaling in filopodia by ratio imaging. The mean TIRF intensity of the FP-AktPH domain was normalized by that of an FP cytoplasmic volume marker; this fluorescence ratio, measured for each filopodium, was normalized again by the fluorescence ratio of a region near the center of the contact area (see Materials and methods and Fig. S3 for technical notes). PI3K signaling in filopodia is considered significant if the resulting quantity, the enrichment ratio E, is significantly >1. Averaged over the lifetime of each filopodium (identified by the aforementioned segmentation algorithm; Fig. S1), the mean value of E = 3.3, whereas pharmacological inhibition of PI3K yielded a mean value of E = 1.1 (Fig. 3 B). Thus, the ratio imaging approach was validated and indicates that filopodia harbor substantial PI3K signaling.
Having devised a method for detecting PI3K signaling in filopodia, we asked whether or not this activity was present before the emergence of lamellipodia. In time-lapse videos of 27 cells subjected to ratio imaging, we identified those instances when lamellipodia emerged over filopodia without marked preexisting enrichment of PI3K signaling in the proximal region of the plasma membrane (i.e., as would be indicated by inspection of the FP-AktPH channel alone). In 78% of those instances (151/194), PI3K signaling was enriched in the filopodial structures beforehand and was thereafter elevated in the lamellipod (Fig. 3 C). These findings establish filopodia as both structural guides and signaling hubs that prime the emergence of lamellipodia.
Nascent adhesions form along F-actin bundles in tandem with propagation of PI3K signaling
Integrin-mediated adhesion is responsible for attachment of filopodia to ECM (Partridge and Marcantonio, 2006; Galbraith et al., 2007), whereas in lamellipodia, it is established that adhesion complexes foster activation of PI3K and Rac, leading to actin polymerization mediated by the Arp2/3 complex (Chen and Guan, 1994; Cox et al., 2001). To test whether or not adhesions nucleate beneath F-actin bundles with the appropriate timing relative to lamellipod formation, we imaged NIH 3T3 fibroblasts coexpressing FP-fascin and FP-paxillin, a marker of adhesion complexes. Adhesions were found along the length of fascin-containing bundles, with the highest density of paxillin typically at the base (Fig. 4 A and Video 5). Most often, such adhesions at the cell periphery lack zyxin, a marker for mature adhesions and stress fibers (Fig. S4, A and B; Zaidel-Bar et al., 2003; Yoshigi et al., 2005). Small adhesions often appeared at the tips of filopodia followed by adhesion growth along the length of the bundle as the nascent lamellipod protruded over it. In cells coexpressing FP-paxillin and FP-AktPH, the large adhesions present at the bases of filopodia colocalize (overlap) with the emergence of PI3K signaling, during the early phase of lamellipodial protrusion (Fig. 4 B and Video 6). Overlap was detected in 42/49 (86%) of protrusions over actin bundles (Fig. S4 C).
Time-lapse video of adhesion dynamics during protrusion over F-actin bundles, as depicted in Fig. 4 A. TIRF microscopy images of an NIH 3T3 cell expressing mCherry-Fascin (left, grayscale) and EGFP-paxillin (center, pseudocolored) were acquired every 30 s for 20 min. Time is given in minutes and seconds. An overlay of the two channels is shown at right. Bar, 5 µm.
Time-lapse video of adhesion dynamics during protrusion over F-actin bundles, as depicted in Fig. 4 A. TIRF microscopy images of an NIH 3T3 cell expressing mCherry-Fascin (left, grayscale) and EGFP-paxillin (center, pseudocolored) were acquired every 30 s for 20 min. Time is given in minutes and seconds. An overlay of the two channels is shown at right. Bar, 5 µm.
Time-lapse video showing adhesion formation and PI3K signaling during protrusion over actin bundles, as depicted in Fig. 4 B. TIRF microscopy images of an NIH 3T3 cell expressing mCherry-AktPH (left) and EGFP-paxillin (right) were acquired every 30 s for 10 min. Time is given in minutes and seconds. An overlay of the two channels is shown at right. Bar, 5 µm.
Time-lapse video showing adhesion formation and PI3K signaling during protrusion over actin bundles, as depicted in Fig. 4 B. TIRF microscopy images of an NIH 3T3 cell expressing mCherry-AktPH (left) and EGFP-paxillin (right) were acquired every 30 s for 10 min. Time is given in minutes and seconds. An overlay of the two channels is shown at right. Bar, 5 µm.
Nascent adhesions form underneath F-actin bundles and colocalize with maximal PI3K signaling. (A) TIRF montages of mCherry-fascin (grayscale) and EGFP-paxillin (pseudocolored) coexpressed in a NIH3T3 cell, along with the overlay of the two channels (red, fascin; green, paxillin; see also Video 5). Bar, 10 µm. The sequence is representative of nine cells viewed and shows the formation of adhesions along the fascin-containing actin bundle during protrusion. (B) TIRF montages of mCherry-AktPH and EGFP-paxillin (both pseudocolored) coexpressed in a NIH 3T3 cell, along with the overlay of the two channels (red, AktPH; green, paxillin; see also Video 6). Bar, 5 µm. The sequence is representative of 13 cells viewed and shows the emergence of PI3K signaling colocalized with nascent adhesions formed along the bundle. (C) TIRF images of FP-AktPH–expressing NIH 3T3 cells, plated on either fibronectin or poly-lysine, before and after FAK inhibition (Inhib; 10 µM FAK inhibitor II; see also Video 7). Bars, 20 µm. For each cell, the after/before ratio compares the difference between the mean intensity of the morphological extension with the highest intensity and the mean intensity of the center region of the contact area; a value significantly <1 indicates that the pattern became more uniform after FAK inhibition. Values are reported as means ± 95% confidence interval for the fibronectin (n = 19) and poly-lysine (n = 24) data.
Nascent adhesions form underneath F-actin bundles and colocalize with maximal PI3K signaling. (A) TIRF montages of mCherry-fascin (grayscale) and EGFP-paxillin (pseudocolored) coexpressed in a NIH3T3 cell, along with the overlay of the two channels (red, fascin; green, paxillin; see also Video 5). Bar, 10 µm. The sequence is representative of nine cells viewed and shows the formation of adhesions along the fascin-containing actin bundle during protrusion. (B) TIRF montages of mCherry-AktPH and EGFP-paxillin (both pseudocolored) coexpressed in a NIH 3T3 cell, along with the overlay of the two channels (red, AktPH; green, paxillin; see also Video 6). Bar, 5 µm. The sequence is representative of 13 cells viewed and shows the emergence of PI3K signaling colocalized with nascent adhesions formed along the bundle. (C) TIRF images of FP-AktPH–expressing NIH 3T3 cells, plated on either fibronectin or poly-lysine, before and after FAK inhibition (Inhib; 10 µM FAK inhibitor II; see also Video 7). Bars, 20 µm. For each cell, the after/before ratio compares the difference between the mean intensity of the morphological extension with the highest intensity and the mean intensity of the center region of the contact area; a value significantly <1 indicates that the pattern became more uniform after FAK inhibition. Values are reported as means ± 95% confidence interval for the fibronectin (n = 19) and poly-lysine (n = 24) data.
These colocalization experiments suggest that adhesions mediate activation of PI3K signaling and other pathways that promote Arp2/3-based, lamellipodial protrusion around F-actin bundles. To test this hypothesis, we assessed the role of FAK, a known intermediate in integrin-mediated activation of PI3K (Chen and Guan, 1994; Reiske et al., 1999). Pharmacological inhibition of FAK activity, which prevents FAK autophosphorylation of the tyrosine residue that engages type IA PI3Ks, ablated the localization of PI3K signaling to putative lamellipodia (Fig. 4 C and Video 7). Whereas FAK inhibition significantly reduced PI3K localization in the cells plated on fibronectin as before, this was not the case for cells plated on poly-lysine, which promotes integrin-independent adhesion of cell membranes (Fig. 4 C). Inhibition of the Src family of protein tyrosine kinases, which are recruited to adhesion complexes as well as other signaling complexes (Plopper et al., 1995; Cuevas et al., 2001), also ablated localization of PI3K signaling in cells plated on fibronectin, and this treatment partially reduced PI3K signaling in cells plated on poly-lysine (Fig. S4 D).
Time-lapse video showing ablation of localized PI3K signaling by FAK inhibition in cells plated on fibronectin but not on poly-lysine, as depicted in Fig. 4 C. TIRF microscopy images of NIH 3T3 cells expressing FP-AktPH (pseudocolored), plated on either fibronectin (left) or poly-lysine (right), were acquired every 30 s for 17 min in separate experiments. At the indicated time, FAK activity was inhibited by addition of 10 µM FAK inhibitor II. Time is given in minutes and seconds. Bar, 20 µm.
Time-lapse video showing ablation of localized PI3K signaling by FAK inhibition in cells plated on fibronectin but not on poly-lysine, as depicted in Fig. 4 C. TIRF microscopy images of NIH 3T3 cells expressing FP-AktPH (pseudocolored), plated on either fibronectin (left) or poly-lysine (right), were acquired every 30 s for 17 min in separate experiments. At the indicated time, FAK activity was inhibited by addition of 10 µM FAK inhibitor II. Time is given in minutes and seconds. Bar, 20 µm.
Arp2/3-driven protrusion amplifies adhesion-based signaling by positive feedback
The observation that photoactivation of Rac induces redistribution of PI3K signaling (Fig. 1 C), by a mechanism that requires actin polymerization (Welf et al., 2012), suggested that the signaling circuit controlling Arp2/3 activation in fibroblasts operates under positive feedback. This is plausible because lamellipodial protrusion results in formation of nascent adhesions that are in transient contact with the dendritic F-actin network (Vicente-Manzanares et al., 2009). Consistent with such a mechanism, after Arp2/3 was pharmacologically inhibited in NIH 3T3 cells coexpressing FP-fascin and FP-AktPH, most of the numerous lamellipodia retracted, in concert with loss of PI3K signaling (Fig. 5 A). FP-paxillin labeling confirmed that small adhesions near the leading edge were also lost after Arp2/3 inhibition (Fig. S5 A) as well as in other contexts in which lamellipodia are impaired: after PI3K inhibition and in fascin-depleted cells (Fig. S5, B and C). The effect of Arp2/3 inhibition on PI3K signaling was not observed in cells plated on poly-lysine (Fig. 5 B), linking the phenomenon to integrin-mediated adhesion. To rule out off-target effects of the drug on PI3K signaling, we performed similar experiments with a different fibroblast line, in which conditional knockout of the Arpc2 (p34) subunit of the Arp2/3 complex was established (Rotty et al., 2015). Whereas uninduced (wild type) cells treated with Arp2/3 inhibitor showed the same ablation of PI3K signaling seen in NIH 3T3 cells, cells treated with tamoxifen to induce Cre-based silencing of Arpc2 showed no marked changes in distribution of PI3K signaling upon Arp2/3 inhibition (Fig. 5 C and Video 8).
Time-lapse video showing ablation of localized PI3K signaling by Arp2/3 inhibition in uninduced cells, but not in cells with tamoxifen-induced knockout of Arpc2, as depicted in Fig. 5 C. Fibroblasts with conditional knockout of the Arpc2 (p34) subunit of the Arp2/3 complex, transfected with FP-AktPH (pseudocolored), were either uninduced (top) or tamoxifen-induced (bottom). TIRF images were acquired every 30 s for 19 min in separate experiments. At the indicated time, the Arp2/3 complex was inhibited by addition of 100 µM CK666. Time is given in minutes and seconds. Bar, 20 µm.
Time-lapse video showing ablation of localized PI3K signaling by Arp2/3 inhibition in uninduced cells, but not in cells with tamoxifen-induced knockout of Arpc2, as depicted in Fig. 5 C. Fibroblasts with conditional knockout of the Arpc2 (p34) subunit of the Arp2/3 complex, transfected with FP-AktPH (pseudocolored), were either uninduced (top) or tamoxifen-induced (bottom). TIRF images were acquired every 30 s for 19 min in separate experiments. At the indicated time, the Arp2/3 complex was inhibited by addition of 100 µM CK666. Time is given in minutes and seconds. Bar, 20 µm.
Arp2/3-driven protrusion amplifies adhesion-based signaling. (A) TIRF images of a NIH 3T3 cell coexpressing mCherry-AktPH and EGFP-fascin before and after inhibition of Arp2/3 complex by 50 µM CK666 (representative of 15 cells in six independent experiments). (B) CK666 treatment (100 µM) does not ablate localized PI3K signaling in FP-AktPH–expressing NIH 3T3 cells plated on poly-lysine (representative of 42 cells). (C) FP-AktPH–transfected fibroblasts with conditional knockout of Arpc2 (p34) were either uninduced (wild type [WT]) or induced with tamoxifen (Arpc2−/−). TIRF images acquired before and after CK666 treatment are shown (see also Video 8; representative of 13 cells each). (D, left) Protrusion dilation waves as shown by pseudocolored mCherry-Akt-PH (see also Video 9), representative of 11 such sequences viewed in six cells. (right) The protrusion width and FP-AktPH intensity are plotted for the same sequence on kymograph-like plots. A.U., arbitrary unit. Bars: (A–C) 20 µm; (D, left) 10 µm.
Arp2/3-driven protrusion amplifies adhesion-based signaling. (A) TIRF images of a NIH 3T3 cell coexpressing mCherry-AktPH and EGFP-fascin before and after inhibition of Arp2/3 complex by 50 µM CK666 (representative of 15 cells in six independent experiments). (B) CK666 treatment (100 µM) does not ablate localized PI3K signaling in FP-AktPH–expressing NIH 3T3 cells plated on poly-lysine (representative of 42 cells). (C) FP-AktPH–transfected fibroblasts with conditional knockout of Arpc2 (p34) were either uninduced (wild type [WT]) or induced with tamoxifen (Arpc2−/−). TIRF images acquired before and after CK666 treatment are shown (see also Video 8; representative of 13 cells each). (D, left) Protrusion dilation waves as shown by pseudocolored mCherry-Akt-PH (see also Video 9), representative of 11 such sequences viewed in six cells. (right) The protrusion width and FP-AktPH intensity are plotted for the same sequence on kymograph-like plots. A.U., arbitrary unit. Bars: (A–C) 20 µm; (D, left) 10 µm.
A hallmark of positive feedback is the phenomenon of traveling waves. We observed recurring, traveling waves of protrusion/retraction directed along a subset of filopodia (n = 11, in 6 cells analyzed this way; Fig. 5 D and Video 9). In all instances, PI3K signaling was most intense at the base of the structure. We developed an automated analysis to plot the width of protrusions as a kymograph, i.e., as a function of time and position parallel to protrusion. From these kymographs, we estimated a mean wave velocity of ∼3 µm/min (Fig. S5 D).
Time-lapse video of protrusion dilation waves, as depicted in Fig. 5 D. TIRF microscopy images were of an NIH 3T3 cell expressing mCherry-AktPH (pseudocolored) were acquired every 10 s for 30 min. Time is given in minutes and seconds. Bar, 10 µm.
Time-lapse video of protrusion dilation waves, as depicted in Fig. 5 D. TIRF microscopy images were of an NIH 3T3 cell expressing mCherry-AktPH (pseudocolored) were acquired every 10 s for 30 min. Time is given in minutes and seconds. Bar, 10 µm.
Fascin-1 depletion blocks fibroblast haptotaxis but not chemotaxis
Having elucidated the dynamic relationship between fascin-containing F-actin bundles and formation of lamellipodia that affects large-scale changes in cell morphology, we sought to test the functional role of fascin-1 in different modes of directed cell migration. For this, we used IA32 mouse fibroblasts, which exhibit robust tactic migration in response to a gradient either soluble PDGF (chemotaxis) or immobilized fibronectin (haptotaxis) established in a microfluidic chamber (Wu et al., 2012). In this cell line, we confirmed the basic findings that fascin-containing bundles serve as templates for formation of lamellipodia and activation of PI3K signaling (Fig. 6 A and Video 10). As in NIH 3T3 cells, we depleted fascin-1 expression by shRNA interference and achieved rescue in this background by expression of homologous fascin-1 (Fig. 6 B). In PDGF chemotaxis assays, fascin-1 depletion yielded a modest reduction in the mean forward migration index (FMI); however, the difference is not statistically significant (Fig. 6 C). In contrast, fascin-1 depletion ablated haptotaxis, a phenotype that was reversed by rescue of fascin-1 expression (Fig. 6 D). Under both chemotaxis and haptotaxis conditions, metrics of unbiased cell migration efficiency—mean speed and persistence (D/T ratio)—were not significantly affected by fascin-1 depletion. We conclude that fascin-1 performs an important function in directed migration of fibroblasts but in a context-dependent manner.
Time-lapse video of multiple F-actin bundles directing protrusions that coalesce to form a broad lamellipod, as depicted in Fig. 6 A. TIRF microscopy images of an IA32 MEF coexpressing EGFP-fascin (inverted grayscale) and mCherry-AktPH (pseudocolored) were acquired every 30 s for 21 min. Time is given in minutes and seconds. Bar, 10 µm.
Time-lapse video of multiple F-actin bundles directing protrusions that coalesce to form a broad lamellipod, as depicted in Fig. 6 A. TIRF microscopy images of an IA32 MEF coexpressing EGFP-fascin (inverted grayscale) and mCherry-AktPH (pseudocolored) were acquired every 30 s for 21 min. Time is given in minutes and seconds. Bar, 10 µm.
Depletion of fascin-1 blocks haptotaxis but not chemotaxis. (A) TIRF montage, representative of 13 cells, showing filopodia-templated protrusion in IA32 MEFs coexpressing EGFP-fascin (inverted grayscale) and mCherry Akt-PH (pseudocolored). Bar, 10 µm. In the example shown, protrusions over filopodia coalesce to form a broad lamellipod (see also Video 10). Arrowheads mark filopodia that preceded lamellipodial protrusion. (B) Representative immunoblot showing relative fascin-1 expression levels in the nontargeting shRNA, fascin shRNA (#1), and fascin rescue IA32 cells used in directed migration experiments. (C) Chemotaxis assay results: wind-rose plots showing the distributions of overall cell migration directionality, expressed as an angle relative to an external gradient of PDGF, are shown for control (n = 74) and fascin-1 shRNA (n = 131). Mean FMI and velocity and persistence (D/T) are displayed ±95% confidence intervals. (D) Haptotaxis assay results are shown for control (n = 128), fascin-1 shRNA (n = 124), and fascin rescue (n = 57) tracks. Mean FMI and velocity and persistence (D/T) are displayed ±95% confidence intervals.
Depletion of fascin-1 blocks haptotaxis but not chemotaxis. (A) TIRF montage, representative of 13 cells, showing filopodia-templated protrusion in IA32 MEFs coexpressing EGFP-fascin (inverted grayscale) and mCherry Akt-PH (pseudocolored). Bar, 10 µm. In the example shown, protrusions over filopodia coalesce to form a broad lamellipod (see also Video 10). Arrowheads mark filopodia that preceded lamellipodial protrusion. (B) Representative immunoblot showing relative fascin-1 expression levels in the nontargeting shRNA, fascin shRNA (#1), and fascin rescue IA32 cells used in directed migration experiments. (C) Chemotaxis assay results: wind-rose plots showing the distributions of overall cell migration directionality, expressed as an angle relative to an external gradient of PDGF, are shown for control (n = 74) and fascin-1 shRNA (n = 131). Mean FMI and velocity and persistence (D/T) are displayed ±95% confidence intervals. (D) Haptotaxis assay results are shown for control (n = 128), fascin-1 shRNA (n = 124), and fascin rescue (n = 57) tracks. Mean FMI and velocity and persistence (D/T) are displayed ±95% confidence intervals.
Discussion
Cooperation of bundled and dendritic F-actin in formation and orientation of lamellipodia
Based on the evidence provided here, together with previous work, we construct the following sequence of processes that link subcellular dynamics to macroscopic changes in cell shape (Fig. 7). (a) F-actin bundles form at the periphery of the cell, often manifest as filopodia. Larger adhesions are typically present at the bases of these actin bundles, whereas smaller adhesions typically form at the tip and eventually along the length of the bundle (Fig. 4 A), consistent with published studies (Nemethova et al., 2008; Schäfer et al., 2010). (b) These structures seed and orient new lamellipodia (Figs. 1 and 2), characterized by Arp2/3-mediated actin polymerization, and a positive feedback loop involving adhesion formation and a FAK–PI3K signaling pathway sustains the spreading (extension and dilation) of the nascent lamellipod. (c) As it does so, new F-actin bundles/filopodia emerge/adhere (Fig. 2 A), consistent with previous indications that Arp2/3-mediated protrusion promotes bundle formation (Korobova and Svitkina, 2008; Yang and Svitkina, 2011); this spatial relationship explains the apparent branching of lamellipodia that results in large-scale reorientation of overall cell migration direction (Welf et al., 2012). Whether before or after lamellipodial branching, the protrusions can retract, and the F-actin bundles are left behind to template future waves of lamellipodial protrusion.
Conceptual model of morphodynamics directed by F-actin bundles. Peripheral actin bundles, e.g., filopodia, template clustering of nascent adhesions and harbor PI3K signaling. The bundles direct lamellipodia, formed by Arp2/3-mediated actin polymerization, by setting the preferred sites and directions of protrusion initiation and propagation, respectively. An adhesion-based positive feedback loop (inset) amplifies PI3K signaling as the lamellipod spreads, promoting additional activation of Arp2/3 complex and formation of new actin bundles. The new bundles can direct initiation of distinct lamellipodia, seemingly branched from existing ones. Propagation of lamellipodia is self-limiting or metastable, and therefore, protrusion and PI3K signaling cease, and the lamellipodia at least partially retract but typically leave the F-actin bundles intact. Hence, the cycle can repeat, sometimes as a series of traveling protrusion waves.
Conceptual model of morphodynamics directed by F-actin bundles. Peripheral actin bundles, e.g., filopodia, template clustering of nascent adhesions and harbor PI3K signaling. The bundles direct lamellipodia, formed by Arp2/3-mediated actin polymerization, by setting the preferred sites and directions of protrusion initiation and propagation, respectively. An adhesion-based positive feedback loop (inset) amplifies PI3K signaling as the lamellipod spreads, promoting additional activation of Arp2/3 complex and formation of new actin bundles. The new bundles can direct initiation of distinct lamellipodia, seemingly branched from existing ones. Propagation of lamellipodia is self-limiting or metastable, and therefore, protrusion and PI3K signaling cease, and the lamellipodia at least partially retract but typically leave the F-actin bundles intact. Hence, the cycle can repeat, sometimes as a series of traveling protrusion waves.
Priming of PI3K signaling by actin bundles
Ratiometric TIRF imaging revealed substantial PI3K signaling in adherent filopodia (Fig. 3 B), following a precedent reported for dendritic filopodia in neurons (Luikart et al., 2008). We detected this signaling activity even before the emergence of nascent lamellipodia (Fig. 3 C) and the more readily visible increases in PI3K signaling that follow (Welf et al., 2012). As shown in Fig. S3, the apparent enrichment of the PIP3 biosensor FP-AktPH in filopodia is partially attributed to geometric considerations; it is estimated that the entire membrane sheath of a filopodium is within the field of TIRF illumination. Considering also the uncertainties about partitioning of the cytoplasmic FP-AktPH pool in/out of filopodia and the net diffusion of PIP3 from the filopodial sheath to the adjacent region of the plasma membrane, the present data do not allow accurate quantification of the extent of PI3K activity enrichment in these structures. The possibility that the density of PIP3 is enhanced by reduction of its turnover (Schneider et al., 2005), i.e., by regulation of lipid dephosphorylation, must also be considered. If achieved by specific localization of PI3K, our colocalization, FAK inhibition, and poly-lysine control experiments implicate the adhesion complexes that form underneath adherent F-actin bundles/filopodia as both the structural and signaling linchpins.
Our results further suggest that PI3K and other signaling activities associated with F-actin bundles prime a positive feedback loop in which adhesions and FAK/PI3K signaling both promote and respond to Arp2/3-mediated actin polymerization and membrane protrusion (Fig. 5, A–C). Excitability and traveling waves are hallmarks of systems with positive feedback and have been characterized in other migrating cell types (Allard and Mogilner, 2013; Huang et al., 2013). The traveling waves of protrusion and PI3K signaling that we sometimes observe (Fig. 5 D) suggest that there is negative regulation as well. Candidates for such regulation mechanisms include mechanical feedback involving Rho signaling to Myosin II (Guilluy et al., 2011; Welf et al., 2013) and an incoherent feedforward loop wherein active Rac both promotes and inhibits the Arp2/3 complex, through WAVE and Arpin, respectively (Dang et al., 2013).
The role of fascin-containing bundles in directed cell migration
Our functional data demonstrate that fascin-1 depletion impairs lamellipodia formation and haptotactic sensing of fibroblasts on ECM. Previous work showed that depletion of the Arp2/3 complex ablated dendritic actin arrays associated with lamellipodia and likewise impaired haptotaxis (Wu et al., 2012). These results concordantly indicate the importance of actin-based protrusion in ECM haptotaxis, with adhesion complexes serving both mechanical and signaling roles. By the same token, depletion of neither fascin-1 nor of the Arp2/3 complex impaired PDGF chemotaxis, at least not significantly so, suggesting that mesenchymal cells use diverse, context-dependent mechanisms for achieving asymmetric force generation (Bear and Haugh, 2014). This does not discount the role of lamellipodia and lamellipodial branching in exploratory motility that allows fibroblasts to align their direction of migration with a PDGF gradient (Welf et al., 2012). Furthermore, considering that our PDGF chemotaxis assay was conducted under near-optimal gradient conditions, our results do not exclude the possibility that coordination of bundled and dendritic F-actin plays a more important role in sensing suboptimal gradients.
Beyond this level of understanding, our results suggest a more refined hypothesis of mesenchymal taxis. A distinct aspect of haptotaxis is that the cell must actively protrude to encounter immobilized cues, whereas soluble chemoattractants are encountered passively, i.e., by diffusion. The search for adhesive ligands is inherently an exploration, with F-actin bundles apparently directing, and lamellipodia propagating, the process. Our observations of adhesions forming beneath the bundles, even before the emergence of lamellipodia, lead us to speculate that the two F-actin structures dynamically cooperate in the haptotactic sensing of ECM ligands and that integrin-mediated signaling integrates those dynamics. Although this is an appealing model, this mechanism might be only one of multiple ways that fascin influences random and haptotactic cell migration. For example, fascin has been shown to regulate tension in stress fibers and thus the dynamics of mature adhesions (Elkhatib et al., 2014).
Fascin expression has been implicated in cancer progression and associated with poor prognoses in many cancers, making it a promising therapeutic target or biomarker (Machesky and Li, 2010; Arjonen et al., 2011). Fascin is associated with the epithelial-to-mesenchymal transition (Li et al., 2014) and is thus linked to acquisition of tumor cell invasiveness. Our results implicate fascin and F-actin bundles in exploratory morphodynamics that are generally important for directed cell migration, haptotaxis especially. ECM gradients are encountered during collective fibroblast invasion of wounds and likewise during tumor cell invasion of the surrounding matrix, and it will be interesting to investigate the relative roles of haptotactic and chemotactic cues in those invasion processes (Lara Rodriguez and Schneider, 2013; Bear and Haugh, 2014). Another important aspect of haptotaxis in need of characterization is how it is affected by the structure of ECM in 3D versus 2D and contact guidance of cell migration that is a manifestation of ECM remodeling by mesenchymal cells (Even-Ram and Yamada, 2005).
Materials and methods
Cell culture, DNA constructs, and other reagents
NIH 3T3 mouse fibroblasts (ATCC), IA32 mouse embryonic fibroblasts (MEFs; Cai et al., 2008), and Arpc2 conditional knockout fibroblasts (Rotty et al., 2015) were cultured in DMEM supplemented with 10% FBS and 1% PSG (penicillin-streptomycin-glutamine; Life Technologies). Transient transfection by lipofection was performed using either Lipofectamine Plus (Life Technologies) or NanoJuice (EMD Millipore). The construct containing EGFP fused to human fascin-1 was a gift from J. Adams (University of Bristol, Bristol, England, UK; Adams and Schwartz, 2000) and was constructed by inserting GenBank accession no. U09873 into the pEGFP-C1 backbone. The EGFP-Akt-PH vector was also constructed using the pEGFP-C1 backbone. To construct the mCherry–fascin-1 plasmid, the mCherry sequence was substituted for EGFP by standard methods and verified by sequencing. The tdTomato-glycophosphatidylinositol (GPI) was created by inserting tdTomato into the pMCP-GPI vector (New England Biolabs, Inc.). The tdTomato-Lifeact construct was cloned into the pLL7.0 vector. The GFP-paxillin vector was obtained from Addgene (plasmid 15233; Laukaitis et al., 2001), as was RFP-zyxin construct (plasmid 26720; Bhatt et al., 2002). The C2-EGFP-MyoX fusion construct was gift from R. Cheney (University of North Carolina [UNC]-Chapel Hill, Chapel Hill, NC; Berg and Cheney, 2002) and the pTriEX-mCerulean-PA-Rac (N-term2) was a gift from the laboratory of K. Hahn (UNC-Chapel Hill; Wu et al., 2009). All constructs were transformed into chemically competent DH5α cells (Life Technologies) and use a cytomegalovirus promoter except the shRNA vectors, which use a U6 promoter. The pharmacological inhibitors CK666, PP2, and FAK inhibitor II (PF-573228) were purchased from EMD Millipore; PI3Kα inhibitor IV was purchased through EMD. Antibodies for immunoblotting were obtained from EMD Millipore (mouse anti-fascin mAb clone 55K2) and Cell Signaling Technology (rabbit anti-GAPDH; mAb 5174).
RNA interference and conditional knockout
Two shRNAs directed against murine fascin-1 (#1, 5′-CCGTTCCAGTTACGATGTCTT-3′, and #2, 5′-CCTCCTGTTATCCTTACTCAT-3′), expressed in the pLKO-puro vector (a lentiviral vector containing a puromycin [puro] resistance gene), were obtained from the Lenti-shRNA Core Facility at UNC-Chapel Hill; efficacy was tested by standard immunoblotting. To coexpress both of the shRNAs, puromycin resistance was swapped for hygromycin resistance in the shRNA #2 plasmid. The shRNAs were packaged in 293T cells cotransfected with pCMV–VSV-G (8454; Addgene) and pCMV DR8.91 plasmids. The control plasmid containing a nontargeting shRNA sequence (5′-TTATCGCGCATATCACGCG-3′), pLKO-shNEG-puro (Everett, 2010), and pCMV DR8.91 were gifts from R. Everett (Medical Research Council-University of Glasgow Centre for Virus Research, Glasgow, Scotland, UK). Virus was harvested, and target cells were infected by the standard method with the addition of 5 µg/ml polybrene. Cells expressing shRNAs were selected using 2 µg/ml puromycin and, in the case of cotransfection, 60 µg/ml hygromycin.
Arpc2−/− fibroblasts were generated as previously described (Rotty et al., 2015). In brief, adult tail fibroblasts were harvested from a conditional knockout mouse with exon 8 of the gene encoding the p34 subunit of the Arp2/3 complex (Arpc2) flanked by LoxP sites in an Ink4a/Arf−/− background. Knockout of p34 was induced by subculturing cells in cell culture media supplemented with 2 µg/ml puromycin and 2 µM tamoxifen for 5 d. The cells were used for up to 10 d thereafter.
Live-cell TIRF microscopy
Prism-based TIRF microscopy was performed as previously described (Schneider and Haugh, 2004, 2005; Johnson and Haugh, 2013). The microscope uses a modified microscope (Axioskop 2 FS; Carl Zeiss) equipped with 40×, 0.8 NA and 60×, 1.0 NA Achroplan water dipping objectives (Carl Zeiss) and a charge-coupled device camera (ORCA ER; Hamamatsu Photonics). Coverslips or glass-bottom dishes (MatTek Corporation) were coated with 10 µg/ml fibronectin (BD), except where noted that the surface was coated with poly-d-lysine (70–150 kD; 1 mg/ml; Sigma Aldrich). Cells were briefly trypsinized and resuspended in imaging buffer (20 mM Hepes, 125 mM NaCl, 5 mM KCl, 1.5 mM MgCl2, 1.5 mM CaCl2, 10 mM glucose, and 2 mg/ml fatty acid–free BSA, pH 7.4) supplemented with 1% PSG and either 1% (NIH 3T3s) or 10% FBS (IA32s). The cells, transiently transfected 1 d before the experiment, were allowed to spread for 2 h (NIH 3T3s) or overnight (IA32s). FPs were excited using the following lasers: 442 nm (TFP), 488 nm (EGFP), and 561 nm (mCherry and tdTomato); emission filters were 480/40, 515/30, and 630/60 nm, respectively. Photoactivation of Rac was performed using a mercury arc lamp as previously described (Welf et al., 2012). In brief, a 50-W mercury arc lamp passed through a 436/20-nm excitation filter and focused to create a small spot. Diffuse light was blocked, and the spot was imaged using fluorescent dextran. A threshold was applied to this image to determine the area of excitation. Image acquisition was programmed using MetaMorph software (Universal Imaging). The frequency of image acquisition varied from 1–12 frames/min (most commonly, 2 frames/min); acquisition for the protrusion dilation wave experiments (Fig. 5 D) was consistently at the upper end of the range (6 or 12 frames/min). All experiments were performed at 37°C. ImageJ (National Institutes of Health) was used for basic image preparation for presentation to crop, convert, adjust contrast, or smooth images for clarity as described in detail previously (Johnson and Haugh, 2013).
Morphodynamic analysis
All image analysis was performed using MATLAB (MathWorks). Images were segmented either automatically by k-means clustering or manually via thresholding. Spatiotemporal mapping and the identification of morphological extensions were performed as described in detail previously (Welf et al., 2012; Johnson and Haugh, 2013). In brief, protrusion/retraction for the spatiotemporal maps was calculated by the difference in overlap between consecutive frames. The angle of each protruded/retracted pixel relative to the cell centroid is determined, and these are binned according to 1° increments. The sum of the pixel counts in each bin for the time interval is converted to a velocity, which is plotted as a function of angle and time and smoothed with a moving mean filter. To identify morphological extensions, the distances of all pixels on the cell contour from the centroid are first measured; portions of the contour that exceeded the moving mean distance taken over 100-pixel windows by ≥0.5 µm for >13-µm contour length, and which contained ≥50 pixels in the extended region, were designated as extensions. These parameters were set so that the structures thus identified are clearly extended from the cell body and exclude filopodia (at this resolution, 0.25 µm/pixel, even very long filopodia will not contain 50 pixels above the threshold).
Analysis of lamellipod/filopod overlap
To identify putative filopodia in each cell, the area containing the cell was cropped and binarized by a manually set threshold. This threshold was determined by starting from a low value and iteratively increasing it until either the cell outline becomes sharp or filopodia begin to be lost; at that point, the threshold is returned to the lowest value that gives a reasonably crisp cell outline. A 7-pixel-wide (1.75 µm at 40×), square top-hat filter was applied to the binary cell mask to segment filopodia. Pixels were filled in between segmented structures within a few pixels of each other to “heal” discontinuities. Then, regions below a certain area cutoff (<15 pixels at 40×) were removed. By subtracting the resulting mask of segmented filopodia from the whole-cell mask, a mask of the rest of the cell was also generated. The first several frames of each cell were checked to ensure reasonably accurate segmentation. A sensitivity analysis was performed on a few cells, varying the chosen threshold and the constant top-hat filter size. Increasing or decreasing the threshold by as much as 20% of the chosen value or varying the filter size by ±1 pixel altered the mean number of filopodia identified by no more than 15% across all frames for the cells tested. Errors in segmentation are primarily attributable to fragmentation of a single filopod into multiple regions or merging of multiple filopodia into a single region.
Based on the morphodynamic characteristics of lamellipodia as seen in cell migration videos, and as analyzed using protrusion maps (Machacek and Danuser, 2006), we defined these structures in NIH 3T3 cells as continuous regions showing estimated protrusion velocity > 1.25 µm/min and spanning ≥8° for ≥1 min. This is performed using the mask excluding filopodia; therefore, the pixels associated with the identified lamellipodia and filopodia cannot overlap. Rather, we define the regions as “overlapping” if they contain pixels in the same angular bin or in adjacent bins (for the same frame of the video or in adjacent frames).
For the FP-fascin–expressing cells, submembranous bundles were included in addition to the putative filopodia. Those bundles were segmented from the FP-fascin channel by applying a 1-µm top-hat filter and taking the highest k-means bin of that image. Bundles not within 5 µm of the cell edge were excluded. At least in protruding lamellipodia, this would exclude regions associated with stress fibers, for example.
Ratiometric analysis of PI3K signaling in filopodia
The volume marker (TFP) channel was used to make masks of the putative filopodia as described in the previous section. For each filopod, the base was identified as the end connected or closest to the cell body. Tracking of filopodia across frames of a video was performed as follows. If the base of a filopod was within a few pixels of one found in the previous two frames, it was counted as the same; otherwise, it was counted as a new filopod. Filopodia that appeared for fewer than three frames were considered nonadherent and removed. The aforementioned mask of the cell excluding filopodia was eroded by 5 µm to identify a central region of the cell. The mean, background-subtracted TIRF intensities, F, from the TFP and AktPH channels were calculated for both the filopodia and the central region. For each filopod, the enrichment ratio, E, was calculated and averaged over its lifetime.
A value of E > 1 is interpreted to mean that AktPH is enriched in the filopod relative to a cytosolic volume marker.
Quantification of change in PI3K signaling pattern
To quantify the change in PI3K signaling in response to FAK inhibition, we first identified morphological extensions as described under Morphodynamic analysis. Subtracting these regions from the whole-cell mask yields a mask of the cell body. The mean AktPH intensity in the cell body region was used as the background level, which was subtracted from the highest AktPH mean intensity among the morphological extensions. The net intensity thus calculated, averaged over five frames beginning 5 min after adding FAK inhibitor, was compared as a ratio to the net intensity for the five-frame mean just before the inhibitor was added (after/before). Complete ablation of localization (AktPH intensity becomes spatially uniform) by the inhibitor returns a value close to 0, whereas no change in the pattern of localization returns a value of 1.
Analysis of PI3K signaling/adhesion colocalization
To quantify overlap enrichment of PI3K signaling with adhesions at the base of filopodia, we visually determined when protrusion over an actin bundle first began and then located where increasing FP-AktPH recruitment was first visible. We next identified the most intense pixels in the associated FP-AktPH image by k-means segmentation, using a bin size of 4; if there was no focal region of FP-AktPH enrichment, the protrusion was scored as having no overlap. The corresponding paxillin channel was likewise segmented, and size cutoffs were applied to filter out noise. The hotspot was then checked for overlap with all adhesions (the two highest bins were 3 and 4; ≥15 contiguous pixels in size) and with those containing the highest intensity of FP-paxillin labeling (bin 4 only; ≥7 contiguous pixels in size). A segmented example is shown in Fig. S4 C.
Chemotaxis and haptotaxis assays
Microfluidic devices were prepared as previously described (Wu et al., 2012). In brief, transparency masks were printed using a high-resolution printer (Fineline Imaging), and the pattern for the chamber was fabricated on 4-inch silicon wafers by a two-step photolithography process. The silicon wafer was exposed to silane overnight after developing and after baking. Polymethylsiloxane (PDMS) was then poured on the wafer and cured overnight at 70°C. Individual PDMS devices were cut out from the wafer, and ports were punched out. The devices were washed with water and then with ethanol, blow dried, and plasma cleaned. The PDMS device was placed into contact with a glass dish bottom immediately, ensuring that an irreversible seal was formed. To establish a PDGF gradient, 27.5-gauge needles were attached to gas tight 100-µl Hamilton glass syringes (81020,1710 TLL 100 µl SYR) and connected via tubing to the entrance ports of the device. Both syringes were filled with DMEM, and the source was supplemented with120 ng/ml PDGF and 1–5 µg/ml TRITC/Cy5-dextran to visualize the gradient. Using a syringe pump, the flow rate was set to 20 nl/min, and a stable gradient was established within 30 min. Haptotactic gradients were formed by the addition of 250 µg/ml Cy5-labeled fibronectin to the source channel. After washing with sterile PBS, the gradient of immobilized fibronectin was visualized by epifluorescence before seeding cells in the culture chamber.
Co-cultures of control and fascin shRNA-containing cells were used in each chamber. The control cells were dyed using CellTracker green (Life Technologies), with the exception of the rescue experiments, in which the rescue cells expressed EGFP-tagged human fascin-1 (isolated by fluorescence-activated cell sorting). Differential interference contrast images were acquired every 10 min for 12–24 h using an IX81 (Olympus), VivaView FL (Olympus), or Biostation (Nikon) microscope, each equipped with motorized stages and a 20× S Apochromat, 0.75 NA (0.5 magnification) or a 20× Plan Fluor DL, 0.5 NA objective. All experiments were performed at 37°C in a humidified environment.
Cells were tracked using the manual tracking plugin (Ibidi) in ImageJ. Intervals during which cells were touching or dividing were excluded. The cell centroid tracks were analyzed in MATLAB to calculate FMI and speed and persistence (D/T ratio) and to construct wind-rose plots. Each wind-rose plot shows a histogram of the angles between endpoints of each track, smoothed by a moving mean. The FMI is defined as the total translocation in the direction of the gradient divided by the overall path length (Foxman et al., 1999). The D/T ratio is defined as the Euclidian distance between endpoints divided by the overall path length.
Online supplemental material
Fig. S1 illustrates the method for segmentation of adherent filopodia. Fig. S2 includes an additional example of fascin-containing bundles templating branch formation, images of Lifeact-labeled F-actin during formation of lamellipodia over actin bundles, quantification of the fates of individual filopodia relative to lamellipodia formation, and quantification of cell migration speeds for wild-type and fascin-overexpressing cells. Fig. S3 illustrates and provides further details regarding the method for calculating the filopodial enrichment ratio. Fig. S4 shows the distinction of mature, zyxin-containing focal adhesions from those at the base of actin bundles that seed lamellipodia, the analysis of PI3K signaling/adhesion colocalization, and results of Src inhibition. Fig. S5 shows the loss of lamellipodia and associated nascent adhesions when the PI3K-Arp2/3 adhesion feedback loop is disrupted and also the quantification of dilation wave velocities. Video 1 shows a time-lapse TIRF video of a randomly migrating GFP-Akt-PH–expressing cell forming lamellipodia over filopodia as depicted in Fig. 1 A. Video 2 shows a time-lapse TIRF video of the directing of lamellipodia during protrusion stimulated by photoactivation of Rac as depicted in Fig. 1 C. Video 3 shows a time-lapse TIRF video of lamellipodial protrusion directed by fascin-containing bundles as depicted in Fig. 2 A. Video 4 shows a time-lapse TIRF video of the localization of fascin and PI3K signaling during protrusion over actin bundles as depicted in Fig. 3 A. Video 5 shows a time-lapse TIRF video of the formation of focal adhesions under actin bundles during protrusion as depicted in Fig. 4 A. Video 6 shows a time-lapse TIRF video of the activation of PI3K at the base and during protrusion over actin bundles as depicted in Fig. 4 B. Video 7 shows a time-lapse TIRF video of PI3K signaling during FAK inhibition on both fibronectin and poly-d-lysine, as depicted in Fig. 4 C. Video 8 shows a time-lapse TIRF video of PI3K signaling during addition of 100 µM CK666 in induced and uninduced Cre-lox Arpc2 fibroblasts, as depicted in Fig. 5 C. Video 9 shows a time-lapse TIRF video of protrusion waves as depicted in Fig. 5 D. Video 10 shows a time-lapse TIRF video of lamellipodia formation over actin bundles in IA32 MEFs as depicted in Fig. 6 A.
Acknowledgments
We are grateful to colleagues who provided plasmids as indicated under the Materials and methods and the UNC Lenti-shRNA Core Facility for the fascin shRNAs. We also wish to thank Erik Welf and Keefe Chan for helpful discussions and Elizabeth Haynes for providing Arp2/3 microinjection videos.
This work was supported by National Science Foundation grant 1133476 (J.M. Haugh) and National Institutes of Health grant GM083035 (J.E. Bear).
The authors declare no competing financial interests.





![Figure 5. Arp2/3-driven protrusion amplifies adhesion-based signaling. (A) TIRF images of a NIH 3T3 cell coexpressing mCherry-AktPH and EGFP-fascin before and after inhibition of Arp2/3 complex by 50 µM CK666 (representative of 15 cells in six independent experiments). (B) CK666 treatment (100 µM) does not ablate localized PI3K signaling in FP-AktPH–expressing NIH 3T3 cells plated on poly-lysine (representative of 42 cells). (C) FP-AktPH–transfected fibroblasts with conditional knockout of Arpc2 (p34) were either uninduced (wild type [WT]) or induced with tamoxifen (Arpc2−/−). TIRF images acquired before and after CK666 treatment are shown (see also Video 8; representative of 13 cells each). (D, left) Protrusion dilation waves as shown by pseudocolored mCherry-Akt-PH (see also Video 9), representative of 11 such sequences viewed in six cells. (right) The protrusion width and FP-AktPH intensity are plotted for the same sequence on kymograph-like plots. A.U., arbitrary unit. Bars: (A–C) 20 µm; (D, left) 10 µm.](https://cdn.rupress.org/rup/content_public/journal/jcb/208/4/10.1083_jcb.201406102/7/m_jcb_201406102_fig5.jpeg?Expires=1779195099&Signature=bbY4DyHvQe7xlnLZu7roTFAyNvnCInKWCZHGELo9v6Y22NGKVRfPIUJJEbDbtr2E4sLs2bKSrKWp8zxW89i0YvRsB7tXCCmDz-IiX~hKfKNhXZvEJANKNS32odxfaPuwItcE1VAo619Kh3~0oSK1HVWK0iVI9oP7egi2LZw-Rg1PTgU2Hg89AoXv9DxtzSoQ-YbiMKLzWkbvhMVO9GzXKIji3nPYtOrUaPBfuBYs2W3kIfBsZJckod1NkJjUqdB-WyNA9zDGpALSrWTRSsSCKpOFmMUiKTRTOIj4nE5yk~hanEF7~7x7N78RzhpIloPY1OvsBzTrsUEXbYlMSfE1qQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)




