Oncogenic Ras induces cell transformation and promotes an invasive phenotype. The tumor suppressor p53 has a suppressive role in Ras-driven invasion. However, its mechanism remains poorly understood. Here we show that p53 induces activation of the mitochondrial protease high-temperature requirement A2 (HtrA2; also known as Omi) and prevents Ras-driven invasion by modulating the actin cytoskeleton. Oncogenic Ras increases accumulation of p53 in the cytoplasm, which promotes the translocation of p38 mitogen-activated protein kinase (MAPK) into mitochondria and induces phosphorylation of HtrA2/Omi. Concurrently, oncogenic Ras also induces mitochondrial fragmentation, irrespective of p53 expression, causing the release of HtrA2/Omi from mitochondria into the cytosol. Phosphorylated HtrA2/Omi therefore cleaves β-actin and decreases the amount of filamentous actin (F-actin) in the cytosol. This ultimately down-regulates p130 Crk-associated substrate (p130Cas)-mediated lamellipodia formation, countering the invasive phenotype initiated by oncogenic Ras. Our novel findings provide insights into the mechanism by which p53 prevents the malignant progression of transformed cells.
Introduction
The tumor suppressor p53 typically triggers apoptosis, cell cycle arrest, or senescence in response to the activation of oncogenes such as ras. This ultimately prevents tumorigenesis (Meek, 2009). In addition to its role in blocking tumorigenesis, p53 hampers malignant phenotypes such as tumor invasion and metastasis (Muller et al., 2011). In some cases, even in the presence of wild-type TP53—the gene that encodes p53—transformation is induced when the ability of p53 to suppress tumorigenesis is somehow compromised. Mutations in TP53 are observed in ∼50% of human cancers and frequently occur in late stages of a variety of cancers, including colorectal, pancreatic, and breast cancers (Rivlin et al., 2011). Missense mutations in Ras oncogenes, resulting in their aberrant activation, are observed in ∼30% of human cancers. Oncogenic Ras not only induces cellular transformation but also promotes tumor cell invasion and metastasis (Campbell and Der, 2004). Although much is known about the mechanism by which p53 prevents Ras transformation, less is known about how p53 suppresses Ras-driven invasion.
Integrins are essential for cell invasion, not only because they mediate adhesion to the ECM, but also because they initiate intracellular signals that regulate actin and adhesion dynamics (Guo and Giancotti, 2004). Upon binding to the ECM, integrins recruit a large number of signaling proteins such as p130 Crk-associated substrate (p130Cas) and FAK to form integrin adhesion complexes. p130Cas is a major substrate of Src at integrin adhesion complexes and Src-mediated tyrosine phosphorylation of p130Cas initiates Rac activation to promote lamellipodia formation (Sharma and Mayer, 2008). Moreover, p130Cas confers an invasive phenotype in cancer cells (Cabodi et al., 2010). The integrin adhesion complexes, of which p130Cas is a component, activate a range of Rho GTPases, including Rho, Rac, and Cdc42. Rho GTPases drive changes in cell shape during cell movement by controlling actin polymerization and actomyosin contraction.
High-temperature requirement A2 (HtrA2)/Omi is a serine protease and chaperone protein (Vande Walle et al., 2008) that has a mitochondrial targeting sequence and transmembrane domain in the N-terminal region. It is initially synthesized as a precursor protein with a predicted molecular weight of 49 kD. The mature 36-kD form of HtrA2/Omi is generated after its posttranslational translocation into mitochondria and subsequently resides in the mitochondrial intermembrane space (IMS). HtrA2/Omi maintains mitochondrial homeostasis including mitochondrial respiration; however, under stress conditions, HtrA2/Omi promotes apoptosis. In the presence of apoptotic stimuli, HtrA2/Omi is released from mitochondria into the cytosol, where it binds to and cleaves inhibitor of apoptosis proteins (IAPs), such as X-IAP, and activates caspases. Overexpression of mature HtrA2/Omi also induces cell rounding through its protease activity (Suzuki et al., 2001). This occurs without caspase activation. Proteomic analysis identified cytoskeletal proteins including β-actin as substrates of HtrA2/Omi (Vande Walle et al., 2007), raising the possibility that the morphological change induced by HtrA2/Omi is due to proteolysis of cytoskeletal proteins.
Mitochondria are dynamic organelles that undergo continuous cycles of fission and fusion (Westermann, 2010). Mitochondrial fission results from the constriction of mitochondria by the dynamin-related GTPase dynamin-related protein 1 (Drp1), and mitochondrial fusion is mediated by other dynamin-related GTPases such as mitofusin 1 (Mfn1) and mitofusin 2 (Mfn2). This dynamic behavior enables daughter cells to inherit mitochondria after cell division and maintains the metabolic functions of mitochondria, including ATP synthesis. Furthermore, mitochondrial fission is often promoted during the early stages of apoptosis. This is commonly observed before caspase activation, and is known to induce mitochondrial outer membrane permeabilization (MOMP; Martinou and Youle, 2011). MOMP leads to the release of IMS proteins such as cytochrome c and HtrA2/Omi into the cytosol. Cytosolic cytochrome c induces caspase activation and further promotes MOMP, resulting in a robust release of IMS proteins and apoptosis (Tait and Green, 2010). Mitochondrial dynamics therefore play an integral role in cell homeostasis, and indeed perturbation of the mitochondrial fission–fusion balance results in mitochondrial fragmentation, which is associated with many disease conditions (Westermann, 2010).
Oncogenic Ras promotes actin reorganization to promote cell invasion (Campbell and Der, 2004). Concurrently, oncogenic Ras induces a p53-mediated alteration of the actin cytoskeleton via HtrA2/Omi, which hampers cell invasion. In this study, we show the opposing effects of oncogenic Ras on cell invasion. Ras-induced disruption of mitochondrial integrity is required for the p53-mediated suppression of cell invasion, and this reflects a novel role of p53 in suppressing the malignancy of transformed cells.
Results
p53 is involved in Ras-induced down-regulation of p130Cas phosphorylation
Oncogenic Ras transforms immortal rodent cells expressing wild-type p53, as well as p53-deficient cells, but induces senescence or apoptosis in primary rodent cells expressing wild-type p53 (Sherr, 1998). To investigate the mechanism by which p53 prevents invasion of Ras-transformed cells, we transformed p53-deficient (p53−/−) mouse embryonic fibroblasts (MEFs), NIH3T3 cells, and immortalized wild-type MEFs (iMEFs) by introducing the activated form of Ha-Ras (Ha-RasV12). NIH3T3 cells and iMEFs are spontaneously immortalized MEFs and express wild-type p53. The invasiveness of Ras-transformed p53−/− MEFs into 3D matrices was higher than that of Ras-transformed NIH3T3 cells or iMEFs (Fig. 1, A and B). The expression of a cancer-associated mutant p53 (p53R175H), which acts as a dominant-negative form, increased the invasiveness of Ras-transformed NIH3T3 cells (Fig. 1, C and D). These results indicate that p53 prevents the invasiveness of Ras-transformed cells.
p53 suppresses Ras-driven invasion. (A and B) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (C and D) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (A and C) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (B and D) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.05.
p53 suppresses Ras-driven invasion. (A and B) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (C and D) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (A and C) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (B and D) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.05.
Reorganization of the actin cytoskeleton and focal adhesions (FAs) is commonly observed during cell transformation (Pawlak and Helfman, 2001). We therefore compared the actin architecture and FAs between the Ras-transformed cells. NIH3T3 cells and iMEFs, after transformation by oncogenic Ras, exhibited poorly spread morphologies (Fig. S1 A) and decreased formation of actin stress fibers and FAs (Fig. 2, A and B). However, in p53−/− MEFs, FA formation remained upon Ras transformation while actin stress fiber formation decreased. We then examined the status of FA proteins and showed that phosphorylation of p130Cas was reduced in Ras-transformed NIH3T3 cells and iMEFs, but not p53−/− MEFs (Fig. 2, B and C). In contrast, all cells irrespective of p53 expression showed a slight decrease in the phosphorylation of FAK, which is another FA protein, upon Ras transformation (Fig. 2 C). Furthermore, all cells showed an increase in the activity of Src, suggesting that the decreased phosphorylation of p130Cas is unlikely due to a down-regulation of Src activity. The expression of p53R175H restored both FA formation and p130Cas phosphorylation (Fig. 2, D and E), which were decreased after oncogenic Ras transformation in NIH3T3 cells (Fig. 2, B and C). These results show that p53 is involved in the down-regulation of p130Cas phosphorylation and FA formation after Ras transformation. Because p130Cas is known to promote cancer cell invasion (Defilippi et al., 2006), we wondered whether p130Cas is involved in the enhanced invasiveness of Ras-transformed cells when wild-type p53 function is lost. We found that knockdown of p130Cas suppressed the invasion of Ras-transformed NIH3T3 cells expressing p53R175H (Fig. 2, F–H) and the invasion of Ras-transformed p53−/− MEFs (Fig. 2, I–K), suggesting that p53 suppresses Ras-driven invasion by down-regulating p130Cas phosphorylation.
Oncogenic Ras decreases phosphorylation of p130Cas in a p53-dependent manner. (A–C) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (D and E) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (A, B, and D) Confocal images of cells stained for F-actin (A) or for paxillin and phosphorylated p130Cas (p-p130Cas; B and D). Bars, 20 µm. (C and E) Levels of phosphorylated p130Cas, phosphorylated FAK (p-FAK), and phosphorylated Src (p-Src) were evaluated by immunoblotting. Blots of phosphorylated p130Cas and p130Cas were quantified and the relative values of p130Cas phosphorylation are shown. (F–H) NIH3T3 cells were infected with Ha-RasV12– and p53R175H-expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (I–K) p53−/− MEFs were infected with a Ha-RasV12–expressing retrovirus together with a control or p130Cas shRNA–expressing retrovirus. (F and I) The level of p130Cas expression was evaluated by immunoblotting. (G and J) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (H and K) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 5 (H) or n = 3 (K). *, P < 0.05.
Oncogenic Ras decreases phosphorylation of p130Cas in a p53-dependent manner. (A–C) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (D and E) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (A, B, and D) Confocal images of cells stained for F-actin (A) or for paxillin and phosphorylated p130Cas (p-p130Cas; B and D). Bars, 20 µm. (C and E) Levels of phosphorylated p130Cas, phosphorylated FAK (p-FAK), and phosphorylated Src (p-Src) were evaluated by immunoblotting. Blots of phosphorylated p130Cas and p130Cas were quantified and the relative values of p130Cas phosphorylation are shown. (F–H) NIH3T3 cells were infected with Ha-RasV12– and p53R175H-expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (I–K) p53−/− MEFs were infected with a Ha-RasV12–expressing retrovirus together with a control or p130Cas shRNA–expressing retrovirus. (F and I) The level of p130Cas expression was evaluated by immunoblotting. (G and J) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (H and K) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 5 (H) or n = 3 (K). *, P < 0.05.
p53 is required for HtrA2/Omi-mediated β-actin cleavage after Ras transformation
Phosphorylation of FA proteins is affected by actin cytoskeleton dynamics (Vuori and Ruoslahti, 1995; Parsons et al., 2010). Treatment of NIH3T3 cells with blebbistatin, a myosin II inhibitor, reduced phosphorylation of FAK, but did not affect phosphorylation of p130Cas (Fig. 3 A). By contrast, when actin polymerization was inhibited by treatment with cytochalasin D or latrunculin A, phosphorylation of both p130Cas and FAK was diminished. These results suggest that the formation of F-actin is important for phosphorylation of p130Cas. We found that oncogenic Ras decreased the amount of F-actin and the ratio of F-actin to globular actin (G-actin) in NIH3T3 cells and iMEFs but not in p53−/− MEFs (Fig. 3, B–D; Fig. S2 A). Introducing p53R175H increased the amount of F-actin and the F/G-actin ratio in Ras-transformed NIH3T3 cells (Fig. 3, E–G; Fig. S2 B). Phosphorylation of p130Cas increased when F-actin was stabilized by treatment with jasplakinolide in Ras-transformed NIH3T3 cells (Fig. 3 H). Collectively, these results suggest that p53 mediates the Ras-induced decrease in F-actin and thereby suppresses phosphorylation of p130Cas.
Oncogenic Ras decreases the amount of F-actin in a p53-dependent manner. (A) NIH3T3 cells were treated with 0.5 µM cytochalasin D, 0.2 µM latrunculin A, or 50 µM blebbistatin for 30 min. The levels of phosphorylated p130Cas (p-p130Cas) and phosphorylated FAK (p-FAK) were evaluated by immunoblotting. (B–D) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (E–G) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (B and E) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bars, 20 µm. (C, D, F, and G) Intensity ratios of F/G-actin (C and F) or F-actin (D and G) were quantified from images in B and E. Values were normalized with the mean value of the control cells. Data represent the mean ± SD of >28 cells. *, P < 0.01. (H) Ras-transformed NIH3T3 cells were treated with 50 nM jasplakinolide for 2 h. The level of phosphorylated p130Cas was evaluated by immunoblotting.
Oncogenic Ras decreases the amount of F-actin in a p53-dependent manner. (A) NIH3T3 cells were treated with 0.5 µM cytochalasin D, 0.2 µM latrunculin A, or 50 µM blebbistatin for 30 min. The levels of phosphorylated p130Cas (p-p130Cas) and phosphorylated FAK (p-FAK) were evaluated by immunoblotting. (B–D) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (E–G) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (B and E) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bars, 20 µm. (C, D, F, and G) Intensity ratios of F/G-actin (C and F) or F-actin (D and G) were quantified from images in B and E. Values were normalized with the mean value of the control cells. Data represent the mean ± SD of >28 cells. *, P < 0.01. (H) Ras-transformed NIH3T3 cells were treated with 50 nM jasplakinolide for 2 h. The level of phosphorylated p130Cas was evaluated by immunoblotting.
We found that oncogenic Ras induced the cleavage of β-actin in NIH3T3 cells and iMEFs but not p53−/− MEFs (Fig. 4, A and B). The cleavage of β-actin in Ras-transformed NIH3T3 cells was diminished by expressing p53R175H (Fig. 4, C and D). We therefore hypothesized that p53-dependent cleavage of β-actin causes a decrease in the amount of F-actin and consequently decreases phosphorylation of p130Cas in Ras-transformed cells. Previous studies have shown that β-actin is cleaved by the proteases caspase-3 and HtrA2/Omi, yielding fragments of ∼30 kD from its C-terminal end (Enoksson et al., 2007; Vande Walle et al., 2007). However, caspase-3 was unlikely to be involved in Ras-induced cleavage of β-actin because oncogenic Ras did not significantly activate caspase-3 and treatment with the DNA-damaging agent Adriamycin activated caspase-3 but did not increase β-actin cleavage in nontransformed cells (Fig. S3 A). We therefore examined whether HtrA2/Omi is involved in p53-dependent cleavage of β-actin, which results in down-regulation of p130Cas phosphorylation upon Ras transformation. Knockdown of HtrA2/Omi did not affect phosphorylation of p130Cas in nontransformed cells (Fig. 4 E). In contrast, knockdown of HtrA2/Omi restored down-regulated p130Cas phosphorylation induced by oncogenic Ras and diminished the cleavage of β-actin (Fig. 4, E and F). The HtrA2/Omi inhibitor Ucf-101 also diminished the cleavage of β-actin and increased phosphorylation of p130Cas in Ras-transformed cells (Fig. 4, G and H). These results suggest that HtrA2/Omi cleaves β-actin after oncogenic Ras transformation.
Knockdown of HtrA2/Omi attenuates the cleavage of β-actin and restores p130Cas phosphorylation in Ras-transformed cells. (A and B) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (C and D) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (E) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus together with a control or HtrA2/Omi shRNA–expressing retrovirus. (F, and I–M) NIH3T3 cells were infected Ha-RasV12–expressing retrovirus and a control or HtrA2/Omi shRNA–expressing retrovirus. (G and H) NIH3T3 cells infected with Ha-RasV12–expressing retrovirus were treated with 50 µM Ucf-101 for 2 h. (N and O) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus and a control or HtrA2/Omi shRNA–expressing retrovirus. (P–R) NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (A, C, F, and H) The cleavage of β-actin was evaluated by immunoblotting. Black arrowheads indicate full-length β-actin, and white arrowheads indicate cleaved fragments. (B and D) The amount of cleaved fragments of β-actin relative to the total amount of β-actin was evaluated. Data represent the mean ± SD; n = 4; *, P < 0.01. (E and G) The levels of phosphorylated p130Cas (p-p130Cas; E and G) and HtrA2/Omi (E) were evaluated by immunoblotting. Blots of phosphorylated p130Cas and p130Cas were quantified and the relative values of p130Cas phosphorylation are shown. (I) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bar, 20 µm. (J and K) F/G-actin ratio (J) or F-actin (K) values were normalized with the mean value of the control cells. Data represent the mean ± SD of more than 30 cells. *, P < 0.01. (L and M) Confocal images of cells stained for F-actin (L) or for paxillin and phosphorylated p130Cas (M). Enlarged images of top panel were shown in bottom panel. Arrowheads indicate lamellipodia. Bars, 20 µm. (N and Q) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (O and R) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.01. (P) The expression level of p130Cas was evaluated by immunoblotting.
Knockdown of HtrA2/Omi attenuates the cleavage of β-actin and restores p130Cas phosphorylation in Ras-transformed cells. (A and B) NIH3T3 cells, iMEFs, and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (C and D) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (E) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus together with a control or HtrA2/Omi shRNA–expressing retrovirus. (F, and I–M) NIH3T3 cells were infected Ha-RasV12–expressing retrovirus and a control or HtrA2/Omi shRNA–expressing retrovirus. (G and H) NIH3T3 cells infected with Ha-RasV12–expressing retrovirus were treated with 50 µM Ucf-101 for 2 h. (N and O) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus and a control or HtrA2/Omi shRNA–expressing retrovirus. (P–R) NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (A, C, F, and H) The cleavage of β-actin was evaluated by immunoblotting. Black arrowheads indicate full-length β-actin, and white arrowheads indicate cleaved fragments. (B and D) The amount of cleaved fragments of β-actin relative to the total amount of β-actin was evaluated. Data represent the mean ± SD; n = 4; *, P < 0.01. (E and G) The levels of phosphorylated p130Cas (p-p130Cas; E and G) and HtrA2/Omi (E) were evaluated by immunoblotting. Blots of phosphorylated p130Cas and p130Cas were quantified and the relative values of p130Cas phosphorylation are shown. (I) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bar, 20 µm. (J and K) F/G-actin ratio (J) or F-actin (K) values were normalized with the mean value of the control cells. Data represent the mean ± SD of more than 30 cells. *, P < 0.01. (L and M) Confocal images of cells stained for F-actin (L) or for paxillin and phosphorylated p130Cas (M). Enlarged images of top panel were shown in bottom panel. Arrowheads indicate lamellipodia. Bars, 20 µm. (N and Q) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (O and R) Cells invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.01. (P) The expression level of p130Cas was evaluated by immunoblotting.
Because cleavage of β-actin seems to correlate with p130Cas phosphorylation, we wondered whether knockdown of HtrA2/Omi has an impact on F-actin and lamellipodia formation. Knockdown of HtrA2/Omi increased the amount of F-actin and the ratio of F/G-actin (Fig. 4, I–K; Fig. S2 C) and facilitated the formation of lamellipodia and FAs, but not stress fibers, in Ras-transformed NIH3T3 cells (Fig. 4, I [middle], L, and M), as was the case with the introduction of p53R175H (Fig. 2 D; Fig. 3 E, middle). Taken together, these results suggest that, upon Ras transformation, HtrA2/Omi cleaves β-actin in a p53-dependent manner and diminishes F-actin formation, resulting in down-regulation of p130Cas phosphorylation.
HtrA2/Omi suppresses invasion of Ras-transformed cells
We next examined role of HtrA2/Omi in the invasion of Ras-transformed cells. The invasiveness of Ras-transformed NIH3T3 cells was increased by knockdown of HtrA2/Omi (Fig. 4, N [bottom] and O). This was abolished by knockdown of p130Cas (Fig. 4, P–R), suggesting that HtrA2/Omi prevents Ras-driven invasion via down-regulation of p130Cas phosphorylation. Even though phosphorylation of p130Cas in nontransformed cells was higher than that in Ras-transformed cells (Fig. 2 C), nontransformed NIH3T3 cells were not invasive (Fig. 4 N, top) as previously reported (Albini et al., 1987) and Ras-transformed cells invaded to a modest though significant extent (Fig. 4 N, middle). Oncogenic Ras promotes cell invasion by increasing matrix degradation and migration regulated by multiple downstream molecules including Rho GTPases (Campbell and Der, 2004). In this context, our results suggest that p130Cas has an integral role to play in Ras-driven invasion, though it is not the sole molecule responsible for this process.
Because the formation of lamellipodia, which are key structures for cancer cell invasion (Olson and Sahai, 2009; Petrie and Yamada, 2012), is facilitated by p53R175H expression or HtrA2/Omi knockdown in Ras-transformed cells (Fig. 3 E, middle; Fig. 4, I [middle] and L), we next examined role of p130Cas in lamellipodia formation. We found that knockdown of p130Cas hampered lamellipodia formation in Ras-transformed cells with HtrA2/Omi knockdown and instead filopodia formation was promoted in these cells (Fig. 5, A–C, middle; Videos 1–3), without any alteration in the amount of F-actin or the ratio of F/G-actin (Fig. 5, C–E; Fig. S2, D and E). Similar effects of p130Cas knockdown were observed in Ras-transformed NIH3T3 cells expressing p53R175H (Fig. 5, F–J; Videos 4 and 5). In addition, a decrease in the cell-spreading area upon oncogenic Ras transformation in a p53- or HtrA2/Omi-dependent manner was not affected by knockdown of p130Cas (Fig. S1, B–D). These results suggest that HtrA2/Omi decreases F-actin and hampers lamellipodia formation through down-regulation of p130Cas signaling, resulting in restricted Ras-driven invasion.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi-shRNA. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a HtrA2/Omi shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi-shRNA. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a HtrA2/Omi shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi- and p130Cas-shRNA. NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a p130Cas shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi- and p130Cas-shRNA. NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a p130Cas shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min
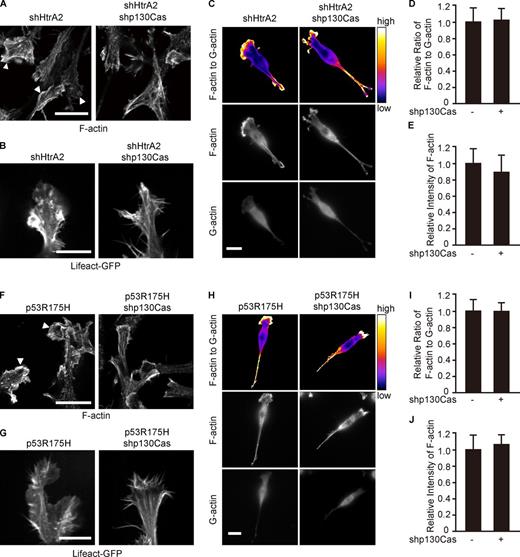
Knockdown of p130Cas suppresses lamellipodia formation in Ras-transformed cells. (A–E) NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (F–J) NIH3T3 cells were infected with Ha-RasV12– and p53R175H-expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (A and F) Confocal images of cells stained for F-actin. Arrowheads indicate lamellipodia. Bar, 20 µm. (B and G) Cells were transfected with a Lifeact-GFP expression vector to label F-actin. Single section of protrusions from Videos 2 and 3 (B) or Videos 4 and 5 (G). Bars, 10 µm. (C and H) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bars, 20 µm. (D, E, I, and J) Intensity ratios of F/G-actin (D and I) or F-actin (E and J) were quantified from images in C and H. Values were normalized with the mean value of the control cells. Data represent the mean ± SD; n = 28 cells.
Knockdown of p130Cas suppresses lamellipodia formation in Ras-transformed cells. (A–E) NIH3T3 cells were infected with Ha-RasV12– and HtrA2/Omi shRNA–expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (F–J) NIH3T3 cells were infected with Ha-RasV12– and p53R175H-expressing retroviruses together with a control or p130Cas shRNA–expressing retrovirus. (A and F) Confocal images of cells stained for F-actin. Arrowheads indicate lamellipodia. Bar, 20 µm. (B and G) Cells were transfected with a Lifeact-GFP expression vector to label F-actin. Single section of protrusions from Videos 2 and 3 (B) or Videos 4 and 5 (G). Bars, 10 µm. (C and H) Cells were stained for F-actin (middle) and G-actin (bottom). Ratio images of F/G-actin (top). Bars, 20 µm. (D, E, I, and J) Intensity ratios of F/G-actin (D and I) or F-actin (E and J) were quantified from images in C and H. Values were normalized with the mean value of the control cells. Data represent the mean ± SD; n = 28 cells.
Oncogenic Ras induces the release of HtrA2/Omi from mitochondria into the cytosol by promoting mitochondrial fragmentation
Given the cleavage of β-actin by HtrA2/Omi, we hypothesized that the release of HtrA2/Omi into the cytosol is induced by oncogenic Ras. We found that the amount of HtrA2/Omi in the cytosolic fraction was increased upon Ras transformation in NIH3T3 cells (Fig. 6 A). The release of IMS proteins, including HtrA2/Omi and cytochrome c, requires MOMP (Tait and Green, 2010). Unlike with MOMP induced by apoptotic stimuli, a robust release of cytochrome c was not observed in Ras-transformed cells regardless of the presence or absence of the pan-caspase inhibitor Q-VD-OPh (Fig. S3 B). MOMP after activation of Ras may therefore be mechanistically different from that resulting from apoptotic stimuli. Because mitochondrial fission promotes MOMP (Martinou and Youle, 2011), we examined the effect of oncogenic Ras on mitochondrial morphology in NIH3T3 cells. In control cells, mitochondria were frequently observed as long, snake-like tubules whereas in Ras-transformed cells, mitochondria mostly appeared small and spherical (Fig. 6 B). Overexpression of Mfn2 promotes mitochondria fusion and attenuated both the release of HtrA2/Omi from mitochondria and cleavage of β-actin in Ras-transformed cells (Fig. 6, C–E). These results suggest that oncogenic Ras promotes mitochondrial fission to induce the release of HtrA2/Omi into the cytosol.
Oncogenic Ras enhances the release of HtrA2/Omi from mitochondria into the cytosol. (A and B) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus. (C–E) Ras-transformed NIH3T3 cells were transfected with a control or Flag-tagged Mfn2 expression vector and cultured for 36 h. (A and D) After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of HtrA2/Omi was evaluated by immunoblotting. COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (B and C) Cells were transfected with a Mito-DsRed expression vector to visualize mitochondria. Images were obtained by three-dimensional structured illumination microscopy (top). Bars (main panels and insets), 10 µm. Colors indicate the lengths of mitochondria. Insets show the raw fluorescence images. Data represent the mean ± SEM (bottom); n = 5. (E) The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment.
Oncogenic Ras enhances the release of HtrA2/Omi from mitochondria into the cytosol. (A and B) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus. (C–E) Ras-transformed NIH3T3 cells were transfected with a control or Flag-tagged Mfn2 expression vector and cultured for 36 h. (A and D) After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of HtrA2/Omi was evaluated by immunoblotting. COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (B and C) Cells were transfected with a Mito-DsRed expression vector to visualize mitochondria. Images were obtained by three-dimensional structured illumination microscopy (top). Bars (main panels and insets), 10 µm. Colors indicate the lengths of mitochondria. Insets show the raw fluorescence images. Data represent the mean ± SEM (bottom); n = 5. (E) The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment.
p53 is involved in Ras-induced translocation of p38 MAPK into mitochondria and phosphorylation of HtrA2/Omi
Overexpression of p53 is known to induce the release of HtrA2/Omi from mitochondria (Marabese et al., 2008). However, in Ras-transformed p53−/− MEFs, mitochondria appeared small and spherical (Fig. 7 A) and HtrA2/Omi was released from mitochondria into the cytosol (Fig. 7, B–D, top), as was observed in Ras-transformed NIH3T3 cells. These results indicate that p53 is not required for Ras-induced release of HtrA2/Omi.
Oncogenic Ras increases mitochondrial translocation of p38 MAPK and phosphorylation of HtrA2/Omi in a p53-dependent manner. (A–E) NIH3T3 cells and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (F) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (G and H) Ras-transformed NIH3T3 cells were treated with 10 µM BIRB796 for 16 h. (A) p53−/− MEFs were transfected with a Mito-DsRed expression vector to visualize mitochondria. Images were obtained by three-dimensional structured illumination microscopy (top). Bars (main panel and inset), 10 µm. Colors indicate the lengths of mitochondria. Insets show the raw fluorescence images. Data represent the mean ± SEM (bottom); n = 5. (B) Confocal images of cells stained for HtrA2/Omi. Bars (top), 10 µm. The square region in the top panel is shown magnified in the bottom panel. Bars (bottom), 1 µm. (C) Relative fluorescence intensities in the cytosol were calculated from the images in B. Data represent the mean ± SD of more than 15 cells. *, P < 0.01. (D) The distribution of HtrA2/Omi and p38 MAPK, and the levels of phosphorylated HtrA2/Omi (p-HtrA2) and phosphorylated p38 MAPK (p-p38), were evaluated by immunoblotting after subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito). COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (E and F) The levels of phosphorylated HtrA2/Omi and phosphorylated p38 MAPK were evaluated by immunoblotting. Black arrowheads indicate phosphorylated HtrA2/Omi. The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was also evaluated. (G) The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was evaluated by immunoblotting. (H) The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment. The relative amount of the cleaved fragment of β-actin to the total amount of β-actin was quantified.
Oncogenic Ras increases mitochondrial translocation of p38 MAPK and phosphorylation of HtrA2/Omi in a p53-dependent manner. (A–E) NIH3T3 cells and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (F) NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. (G and H) Ras-transformed NIH3T3 cells were treated with 10 µM BIRB796 for 16 h. (A) p53−/− MEFs were transfected with a Mito-DsRed expression vector to visualize mitochondria. Images were obtained by three-dimensional structured illumination microscopy (top). Bars (main panel and inset), 10 µm. Colors indicate the lengths of mitochondria. Insets show the raw fluorescence images. Data represent the mean ± SEM (bottom); n = 5. (B) Confocal images of cells stained for HtrA2/Omi. Bars (top), 10 µm. The square region in the top panel is shown magnified in the bottom panel. Bars (bottom), 1 µm. (C) Relative fluorescence intensities in the cytosol were calculated from the images in B. Data represent the mean ± SD of more than 15 cells. *, P < 0.01. (D) The distribution of HtrA2/Omi and p38 MAPK, and the levels of phosphorylated HtrA2/Omi (p-HtrA2) and phosphorylated p38 MAPK (p-p38), were evaluated by immunoblotting after subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito). COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (E and F) The levels of phosphorylated HtrA2/Omi and phosphorylated p38 MAPK were evaluated by immunoblotting. Black arrowheads indicate phosphorylated HtrA2/Omi. The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was also evaluated. (G) The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was evaluated by immunoblotting. (H) The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment. The relative amount of the cleaved fragment of β-actin to the total amount of β-actin was quantified.
HtrA2/Omi is phosphorylated at Ser 142 by p38 MAPK, which increases the protease activity of HtrA2/Omi (Plun-Favreau et al., 2007). We then examined whether oncogenic Ras induces the activation of HtrA2/Omi in a p53-dependent manner. Oncogenic Ras increased phosphorylation of HtrA2/Omi at Ser 142 in NIH3T3 cells but not in p53−/− MEFs (Fig. 7 E, top). Moreover, the increased phosphorylation in Ras-transformed NIH3T3 cells was diminished by expressing p53R175H (Fig. 7 F, top). Collectively, these results indicate that p53 is required for activation of HtrA2/Omi induced by oncogenic Ras.
Mitochondrial serine/threonine protein kinase PTEN-induced putative kinase 1 (PINK1) has been shown to be essential for p38 MAPK-induced HtrA2/Omi phosphorylation (Plun-Favreau et al., 2007). Oncogenic Ras increased HtrA2/Omi phosphorylation in both the cytosol and mitochondria of NIH3T3 cells but not in p53−/− MEFs (Fig. 7, D and E), whereas levels of phosphorylated p38 MAPK—the active form of p38 MAPK—were comparable between NIH3T3 cells and p53−/− MEFs (Fig. 7 E). We therefore hypothesized that oncogenic Ras induces the translocation of p38 MAPK into mitochondria in a p53-dependent manner. Furthermore, we believed the subsequent phosphorylation of HtrA2/Omi occurred in mitochondria, before its release into the cytosol. In line with this, we found that in NIH3T3 cells but not p53−/− MEFs, oncogenic Ras increased the amount of p38 MAPK and its phosphorylated form in the mitochondrial fraction (Fig. 7 D). These results suggest that oncogenic Ras induces phosphorylation of HtrA2/Omi by increasing the translocation of p38 MAPK into mitochondria in a p53-dependent manner. Consistent with this, p53R175H diminished Ras-induced phosphorylation of HtrA2/Omi in the mitochondrial fraction (Fig. 7 F). Furthermore, we showed that treatment with BIRB796, an inhibitor of p38 MAPK, diminished Ras-induced phosphorylation of HtrA2/Omi in mitochondria and cleavage of β-actin (Fig. 7, G and H). These results suggest that p53 mediates β-actin cleavage via activation of HtrA2/Omi, which is induced by mitochondrial p38 MAPK after oncogenic Ras transformation.
Next, we examined the direct effects of HtrA2/Omi on the cleavage of β-actin. The exogenous mature form of HtrA2/Omi, which is diffusely distributed in cells (Suzuki et al., 2001), increased the cleavage of β-actin in Ras-transformed cells expressing p53R175H (Fig. 8 A). The phosphomimetic mutant of mature HtrA2/Omi (mHtrA2/Omi-S142D) was more potent in cleaving β-actin (Fig. 8 A). To further examine whether HtrA2/Omi affects the stability of F-actin, we incubated fluorescently labeled F-actin with a purified mature form of HtrA2/Omi (Fig. 8 B). Wild-type HtrA2/Omi induced disassembly of F-actin, with mHtrA2/Omi-S142D showing an even more prominent effect (Fig. 8, C [left and right] and E; Videos 6 and 8). The protease-dead mutant of HtrA2/Omi (HtrA2/Omi-S306A) did not disassemble F-actin (Fig. 8, C [middle] and E; Video 7), indicating that HtrA2/Omi-induced disassembly of F-actin depends on its protease activity. Because mHtrA2/Omi-S142D induced fragmentation of F-actin (Fig. 8 D; Video 8), activated HtrA2/Omi is likely to cleave actin molecules in F-actin, thereby inducing its disassembly. Furthermore, we showed that mHtrA2/Omi-S142D abolished the invasiveness of Ras-transformed p53−/− MEFs (Fig. 8, F and G). These results support the notion that invasion of Ras-transformed cells is attenuated by the cleavage of β-actin through activation of HtrA2/Omi.
Images of F-actin incubated with HtrA2/Omi-S306A. Alexa Fluor 488–labeled F-actin was incubated with purified HtrA2/Omi-S306A. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Images of F-actin incubated with HtrA2/Omi-S306A. Alexa Fluor 488–labeled F-actin was incubated with purified HtrA2/Omi-S306A. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Images of F-actin incubated with HtrA2/Omi-S142D. Alexa Fluor 488–labeled F-actin was incubated with purified HtrA2/Omi-S142D. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Images of F-actin incubated with HtrA2/Omi-S142D. Alexa Fluor 488–labeled F-actin was incubated with purified HtrA2/Omi-S142D. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Expression of the phosphomimetic form of HtrA2/Omi enhances β-actin cleavage and suppresses Ras-driven cell invasion. (A) Ras-transformed p53R175H-expressing cells were transfected with a Flag-tagged wild-type (WT) or S142D mature HtrA2/Omi (mHtrA2) expression vector. The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment. The relative amount of the cleaved fragment of β-actin to the total amount of β-actin was evaluated. Anti-Flag and anti-HtrA2/Omi antibodies were used to examine the expression level of exogenous HtrA2/Omi. Black arrows indicate exogenous HtrA2/Omi, and white arrow indicates endogenous HtrA2/Omi. N.S., nonspecific band. (B) Purified proteins of wild-type (WT), S306A, or S142D mature HtrA2/Omi were subjected to SDS-PAGE, followed by Coomassie brilliant blue (CBB) staining or anti-Flag immunoblotting. (C) Purified HtrA2/Omi proteins in B were incubated with F-actin for indicated time. Images were obtained by TIRF microscopy. Bar, 5 µm. (D) Serial time-lapse images of the actin filament in the red rectangle in C. Numbers indicate the time in seconds. Bar, 1 µm. (E) Fluorescence intensities after incubation with each form of purified HtrA2/Omi relative to fluorescence intensities before the incubation were calculated. Data represent the mean ± SD; n = 3. *, P < 0.05. (F and G) Ras-transformed p53−/− MEFs were transfected with a control or Flag-tagged S142D mature HtrA2/Omi (mHtrA2) expression vector. (F) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (G) Cells that had invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.01.
Expression of the phosphomimetic form of HtrA2/Omi enhances β-actin cleavage and suppresses Ras-driven cell invasion. (A) Ras-transformed p53R175H-expressing cells were transfected with a Flag-tagged wild-type (WT) or S142D mature HtrA2/Omi (mHtrA2) expression vector. The cleavage of β-actin was evaluated by immunoblotting. Black arrowhead indicates full-length β-actin, and white arrowhead indicates a cleaved fragment. The relative amount of the cleaved fragment of β-actin to the total amount of β-actin was evaluated. Anti-Flag and anti-HtrA2/Omi antibodies were used to examine the expression level of exogenous HtrA2/Omi. Black arrows indicate exogenous HtrA2/Omi, and white arrow indicates endogenous HtrA2/Omi. N.S., nonspecific band. (B) Purified proteins of wild-type (WT), S306A, or S142D mature HtrA2/Omi were subjected to SDS-PAGE, followed by Coomassie brilliant blue (CBB) staining or anti-Flag immunoblotting. (C) Purified HtrA2/Omi proteins in B were incubated with F-actin for indicated time. Images were obtained by TIRF microscopy. Bar, 5 µm. (D) Serial time-lapse images of the actin filament in the red rectangle in C. Numbers indicate the time in seconds. Bar, 1 µm. (E) Fluorescence intensities after incubation with each form of purified HtrA2/Omi relative to fluorescence intensities before the incubation were calculated. Data represent the mean ± SD; n = 3. *, P < 0.05. (F and G) Ras-transformed p53−/− MEFs were transfected with a control or Flag-tagged S142D mature HtrA2/Omi (mHtrA2) expression vector. (F) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (G) Cells that had invaded 30 µm or more were quantified. Data represent the mean ± SD; n = 3. *, P < 0.01.
Taken together, these results suggest that p53 mediates Ras-induced translocation of p38 MAPK into mitochondria and induces activation of HtrA2/Omi, resulting in down-regulation of p130Cas signaling via the disassembly of F-actin.
Cytoplasmic p53 promotes the translocation of p38 MAPK into mitochondria
We next investigated the mechanism by which p53 mediates Ras-induced translocation of p38 MAPK into mitochondria. Several studies show that p38 MAPK is present in mitochondria under stress conditions (Grethe et al., 2004; Kong et al., 2005). It is known that p53 mediates a stress-induced reduction of the inner mitochondrial membrane potential (ΔΨm; Vousden and Lane, 2007; Vaseva and Moll, 2009), which is usually generated by the respiratory chain. We therefore examined ΔΨm using JC-1. Cancer cells exhibit decreased oxidative phosphorylation, but maintain ΔΨm (Fantin and Leder, 2006). Consistent with this, ΔΨm in most mitochondria was maintained in Ras-transformed NIH3T3 cells (Fig. 9, A and B). However, ΔΨm in mitochondria at the cell periphery was decreased and this decrease was blocked by expressing p53R175H and abolished in p53−/− MEFs, indicating that oncogenic Ras decreases ΔΨm in a subset of mitochondria within a cell in a p53-dependent manner. We showed that treatment with CCCP, which uncouples the proton gradient and dissipates ΔΨm, promoted the translocation of p38 MAPK into mitochondria in Ras-transformed p53−/− MEFs (Fig. 9 C). These results suggest that p53 mediates Ras-induced translocation of p38 MAPK into mitochondria by decreasing ΔΨm.
Oncogenic Ras decreases mitochondrial membrane potential in a p53-dependent manner. (A and B) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (A) Confocal images of cells stained with JC-1. The ratio of red fluorescence (J-aggregate) to green fluorescence (monomer) is correlated with mitochondrial membrane potential. Bars, 20 µm. (B) The ratio of red fluorescence to green fluorescence at peripheral mitochondria was quantified from images in A. Data represent the mean ± SD of more than 20 cells. *, P < 0.01. (C) Ras-transformed p53−/− MEFs were treated with CCCP (20 µM) for 4 h. After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of p38 MAPK was evaluated by immunoblotting. COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively.
Oncogenic Ras decreases mitochondrial membrane potential in a p53-dependent manner. (A and B) NIH3T3 cells were infected with a control or Ha-RasV12–expressing retrovirus together with a control or p53R175H-expressing retrovirus. p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. (A) Confocal images of cells stained with JC-1. The ratio of red fluorescence (J-aggregate) to green fluorescence (monomer) is correlated with mitochondrial membrane potential. Bars, 20 µm. (B) The ratio of red fluorescence to green fluorescence at peripheral mitochondria was quantified from images in A. Data represent the mean ± SD of more than 20 cells. *, P < 0.01. (C) Ras-transformed p53−/− MEFs were treated with CCCP (20 µM) for 4 h. After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of p38 MAPK was evaluated by immunoblotting. COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively.
It is well known that p53 induces MOMP by inducing expression of its target genes, such as PUMA and BAX, resulting in a loss of ΔΨm (Miyashita and Reed, 1995; Nakano and Vousden, 2001; Vousden and Lane, 2007). Oncogenic Ras did not increase the expression of PUMA and BAX in both NIH3T3 cells and in p53−/− MEFs (Fig. S4, A and B), suggesting that the decrease in ΔΨm induced by oncogenic Ras may not be dependent on transactivation of p53 targets. In addition to its role as a transcription factor in the nucleus, p53 has functions in the cytoplasm (Green and Kroemer, 2009; Vaseva and Moll, 2009). Under stress conditions, p53 translocates to mitochondria, contributing to the decrease in ΔΨm. We found that p53 accumulated at mitochondria and in the cytosol in Ras-transformed NIH3T3 cells (Fig. 10 A). We then addressed whether cytoplasmic p53 is involved in the translocation of p38 MAPK into mitochondria after Ras transformation. In Ras-transformed p53−/− MEFs, the nuclear localization signal (NLS) mutant of p53 (p53KRKKK) localized at mitochondria and in the cytosol and enhanced the translocation of p38 MAPK into mitochondria (Fig. 10, B and C), without the induction of PUMA and BAX expression (Fig. S4 C). p53KRKKK increased phosphorylation of HtrA2/Omi in mitochondria and decreased the invasion of Ras-transformed p53−/− MEFs (Fig. 10, D–F). Altogether, these results suggest that cytoplasmic p53 restricts Ras-induced cell invasion through increasing p38 MAPK–induced HtrA2/Omi activation in mitochondria.
Expression of p53KRKKK enhances mitochondrial translocation of p38 MAPK and phosphorylation of HtrA2/Omi in Ras-transformed cells. (A) NIH3T3 cells and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of p53 was evaluated by immunoblotting. Black arrowhead indicates nonspecific band (N.S.). COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (B–F) p53−/− MEFs were infected with a Ha-RasV12–expressing retrovirus together with a control or p53KRKKK-expressing retrovirus. (B, top) DIC images of cells. Cell was stained for nucleus with DAPI (middle) or for p53 (bottom). Bar, 20 µm. (C) After subcellular fractionation of the cytosol and mitochondria, the distribution of p38 MAPK was evaluated by immunoblotting. (D) The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was evaluated by immunoblotting. (E) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (F) Cells invaded 30 µm or more were quantified from image in E. Data represent the mean ± SD; n = 3. *, P < 0.01. (G) Schematic of p53-mediated suppression of Ras-driven invasion via HtrA2/Omi. After Ras transformation, concurrent events of mitochondria fission and p53-mediated activation of HtrA2/Omi induced by p38 MAPK in mitochondria leads to the release of activated HtrA2/Omi. Consequentially, actin polymerization and p130Cas phosphorylation are inhibited, resulting in the suppression of invasion.
Expression of p53KRKKK enhances mitochondrial translocation of p38 MAPK and phosphorylation of HtrA2/Omi in Ras-transformed cells. (A) NIH3T3 cells and p53−/− MEFs were infected with a control or Ha-RasV12–expressing retrovirus. After subcellular fractionation of the cytosol (Cyto) and mitochondria (Mito), the distribution of p53 was evaluated by immunoblotting. Black arrowhead indicates nonspecific band (N.S.). COX IV and α-tubulin were used as mitochondrial and cytosolic markers, respectively. (B–F) p53−/− MEFs were infected with a Ha-RasV12–expressing retrovirus together with a control or p53KRKKK-expressing retrovirus. (B, top) DIC images of cells. Cell was stained for nucleus with DAPI (middle) or for p53 (bottom). Bar, 20 µm. (C) After subcellular fractionation of the cytosol and mitochondria, the distribution of p38 MAPK was evaluated by immunoblotting. (D) The level of phosphorylated HtrA2/Omi in the mitochondrial fraction was evaluated by immunoblotting. (E) Inverted invasion assays were performed. Images of serial Z-sections (10-µm intervals). Bars, 100 µm. (F) Cells invaded 30 µm or more were quantified from image in E. Data represent the mean ± SD; n = 3. *, P < 0.01. (G) Schematic of p53-mediated suppression of Ras-driven invasion via HtrA2/Omi. After Ras transformation, concurrent events of mitochondria fission and p53-mediated activation of HtrA2/Omi induced by p38 MAPK in mitochondria leads to the release of activated HtrA2/Omi. Consequentially, actin polymerization and p130Cas phosphorylation are inhibited, resulting in the suppression of invasion.
Discussion
We have revealed a novel molecular mechanism by which p53 suppresses Ras-driven invasion (Fig. 10 G). Upon Ras transformation, accumulation of p53 in the cytoplasm is induced, resulting in the activation of HtrA2/Omi via an increased translocation of p38 MAPK into mitochondria. Concurrently, oncogenic Ras induces mitochondrial fragmentation regardless of p53 expression. As a result, activated HtrA2/Omi is released from mitochondria into the cytosol, where it induces F-actin disassembly. This leads to a down-regulation of p130Cas signaling, thereby suppressing cell invasion through the impairment of lamellipodia formation. This p53-mediated pathway will play an important role in the suppression of malignancy when tumorigenesis is induced by oncogenic Ras.
Mitochondrial dysfunction induced by small molecule drugs including the F0 proton channel inhibitor oligomycin triggers mitochondrial fragmentation without cell death (Fig. S5 A; De Vos et al., 2005). In addition, treatment with oligomycin induces the release of HtrA2/Omi from mitochondria (Fig. S5 B). Because Ras transformation leads to mitochondrial dysfunction (Hu et al., 2012), it is likely that oncogenic Ras induces HtrA2/Omi release through mitochondrial fragmentation caused by mitochondrial dysfunction.
Members of the pro-apoptotic Bcl2 family, including Bax and Bak, are major activators of MOMP and apoptosis (Tait and Green, 2010). After apoptotic stimuli, Bax is recruited from the cytosol to the outer mitochondrial membrane, where Bak is already present. Oligomerization of the two proteins promotes MOMP, leading to caspase activation, which follows the release of IMS proteins involving HtrA2/Omi. Oncogenic Ras induces the release of HtrA2/Omi without caspase activation (Fig. 7, B–D; Fig. S3), implying that MOMP after Ras transformation is mechanistically different from MOMP induced by apoptotic stimuli. Bax shuttles between the cytosol and mitochondrial fission sites and oligomerizes at the fission sites to induce MOMP (Martinou and Youle, 2011). GFP-tagged Bax (GFP-Bax) was diffusely distributed in nontransformed cells (Fig. S5 C), in line with a previous study (Perumalsamy et al., 2010), whereas it clearly localized to mitochondria in Ras-transformed cells or oligomycin-treated cells (Fig. S5, C and D). Bak is constitutively down-regulated by oncogenic Ras (Liu et al., 2006). This suggests that oncogenic Ras-induced MOMP and HtrA2/Omi release involves Bax, but not Bak, and that oncogenic Ras functions to suppress Bax/Bak-induced MOMP after apoptotic stimuli. Indeed, oncogenic Ras suppresses ECM detachment–induced anoikis, a form of apoptosis, via inhibition of HtrA2/Omi release, which is mediated by Bax and Bak (Gilmore et al., 2000; Liu et al., 2006). Thus, oncogenic Ras controls the release of HtrA2/Omi through the regulation of a different type of MOMP.
Our results suggest that the mitochondrial translocation of p38 MAPK, which is induced by cytoplasmic p53 to activate HtrA2/Omi, is caused by a decrease in ΔΨm (Fig. 9 C). This decrease in ΔΨm is dispensable for HtrA2/Omi release from mitochondria (Fig. S5, B and E). In response to stress signals, p53 is known to rapidly translocate to mitochondria and decrease in ΔΨm (Vaseva and Moll, 2009). When p53 accumulates in the mitochondrial outer membrane, p53 binds to Bcl-2 family members and promotes Bax/Bak-induced MOMP. This will not occur after Ras transformation because neither the release of cytochrome c nor activation of caspase is observed in Ras-transformed cells (Fig. S3). p53 also accumulates in the mitochondrial matrix, thereby inhibiting cyclophilin D, which is a regulator of the mitochondrial permeability transition pore (MPTP), and triggering the opening of MPTP (Vaseva et al., 2012). A decrease in ΔΨm was observed in mitochondria at the cell periphery in Ras-transformed cells expressing p53 (Fig. 9 A). The subcellular heterogeneity of ΔΨm is usually due to mitochondria interacting with other organelles, including the ER and acidic vesicular organelles (Keil et al., 2011). p53 is also known to localize at the ER or lysosome (Yamasaki et al., 2007; Johansson et al., 2010). Cytoplasmic p53 seems to contribute to the decrease in ΔΨm by modulating these interactions in Ras-transformed cells. In addition, cytosolic p53 is known to inhibit the autophagic elimination of depolarized mitochondria (Hoshino et al., 2013) and may facilitate the accumulation of mitochondria with low ΔΨm in Ras-transformed cells.
The cleavage of β-actin after Ras transformation was observed in both NIH3T3 cells and iMEFs expressing wild-type p53 (Fig. 4, A and B). The amount of β-actin cleavage in NIH3T3 cells is higher than that in iMEFs and this is correlated with a decrease in the F-actin amount; however, the decrease in F/G-actin ratio was comparable between NIH3T3 cells and iMEFs (Fig. 3 C). Because HtrA2/Omi cleaves nonpolymerized actin (Vande Walle et al., 2007) as well as polymerized actin (Fig. 8, C–E), changes in the F-actin to G-actin ratio after Ras transformation appears to be associated with a decrease in not only F-actin, but also G-actin. As actin filament formation plays an important role in p130Cas phosphorylation, correlation between decreased p130Cas phosphorylation and the amount of F-actin, rather than the F-actin to G-actin ratio, is reasonable. Because immortal cells have genes with mutations, this may influence the cleavage of β-actin in response to oncogenic Ras. However, results using primary p53−/− MEFs or p53R175H-introduced NIH3T3 cells support our provided model that p53 hampers Ras-driven invasion via HtrA2/Omi-mediated cleavage of β-actin when the ability of p53 to suppress tumorigenesis is somehow compromised.
p130Cas and FAK are localized at FAs and cooperate to promote cancer cell invasion (Wang and McNiven, 2012). Increase in cellular tension is known to induce activation of FAK, which subsequently induces autophosphorylation at Tyr397, leading to the recruitment of various proteins to FAs, including p130Cas (Parsons et al., 2010). Interestingly, their regulation in Ras-transformed cells appears to be different. FAK phosphorylation was slightly decreased in Ras-transformed cells irrespective of p53 expression, whereas p130Cas phosphorylation was decreased only in Ras-transformed cells expressing p53 (Fig. 2 C). Phosphorylation of both FAK and p130Cas was significantly decreased by the inhibition of actin polymerization in nontransformed cells (Fig. 3 A). Importantly, the amount of F-actin in p53−/− MEFs was not affected by transformation (Fig. 3 D) and therefore oncogenic Ras-mediated down-regulation of FAK phosphorylation is not caused by the inhibition of actin polymerization. A Ras-induced reduction in traction force exertion, which is associated with actomyosin contractility (Moore et al., 2010), was observed in both NIH3T3 cells and p53−/− MEFs (Fig. S1, E and F), and may therefore, at least in part, be involved in the decrease in FAK phosphorylation. In addition, FAK phosphorylation is regulated independently of focal adhesion regulation (Zheng et al., 2009). Taken together, p130Cas promotes cell invasion of Ras-transformed cells, without a correlation in FAK activity.
Here, we propose a novel role of p53 in dictating the malignant phenotype of oncogenic Ras-transformed cells. The evidence presented in this study highlights the importance of p53 in the regulation of the actin cytoskeleton via disruption of mitochondrial integrity.
Materials and methods
Cell culture and retroviral infection
Cells were cultured in DMEM supplemented with 10% FBS (Biowest). Immortalized wild-type MEFs were described elsewhere (Su et al., 1999). p53−/− MEFs were prepared as described previously (Kawauchi et al., 2008) and infected after two passages. We performed all experiments within two weeks after selection to avoid genetic abnormalities acquired during prolonged culture. Retroviral infection was performed as described previously (Kawauchi et al., 2008). Infected cells were selected using 300 µg/ml hygromycin and 1.5 µg/ml puromycin for 3 d, and 500 µg/ml zeocin and 500 µg/ml neomycin for 7 d.
Plasmids
To generate retroviruses encoding shRNAs against HtrA2/Omi and p130Cas, the HtrA2/Omi target sequence, 5′-CACACTGAGGATTCAAACCAA-3′, and p130Cas target sequence, 5′-GCATGACATCTACCAAGTT-3′, were cloned into a pSuper puro (Oligoengine) or pSINsi (Takara Bio Inc.) vector. pBabe Ha-RasV12 with a hygromycin selection marker and pBabe p53R175H with a zeocin selection marker were used (Kawauchi et al., 2008). pBabe p53KRKKK, which contains K305A, R306A, K319A, K320A, and K321A substitutions in NLS (O’Keefe et al., 2003), was generated from pBabe p53 with a puromycin selection marker. The mature forms of mouse HtrA2/Omi cDNA were obtained by PCR from NIH3T3 cDNA pools and cloned into a pCMV-Flag vector. pCMV-Flag-mature HtrA2/Omi was then generated with either the S142D or S306A mutation, where Ser 142 or Ser 306 of HtrA2/Omi was replaced by aspartic acid or alanine, respectively. The Mito-DsRed expression vector was obtained from Takara Bio Inc.
Antibodies and materials
Anti-p130Cas rabbit polyclonal (αCas3), anti-phospho-FAK (Tyr 397) mouse monoclonal (EMD Millipore), anti-FAK mouse monoclonal (12G4; Abcam), anti–phospho-Src (Tyr 416) rabbit monoclonal (D49G4; Cell Signaling Technology), anti-Src mouse monoclonal (GD11; EMD Millipore), anti–phospho-p38 (Thr 180/Tyr 182) rabbit monoclonal (3D7; Cell Signaling Technology), anti-p38 rabbit polyclonal (Cell Signaling Technology), anti–phospho-HtrA2/Omi (Ser 142) rabbit polyclonal, anti-HtrA2/Omi goat polyclonal (V-17; Santa Cruz Biotechnology, Inc.), anti-p53 mouse monoclonal (1C12; Cell Signaling Technology), anti–β-actin mouse monoclonal (ACTBD11B7; Santa Cruz Biotechnology, Inc.), anti-COX IV rabbit polyclonal (ab16056; Abcam), anti–caspase-3 rabbit polyclonal (Cell Signaling Technology), anti-cleaved caspase-3 rabbit monoclonal (5A1E; Cell Signaling Technology), anti-Flag mouse monoclonal (M2; Sigma-Aldrich), anti–α-tubulin mouse monoclonal (DM1A; Sigma-Aldrich), and anti-GAPDH mouse monoclonal (6C5; Life Technologies) antibodies were used for immunoblot analysis. Anti–phospho-p130Cas (Tyr 165) rabbit polyclonal (Cell Signaling Technology) and anti-p53 mouse monoclonal (DO-1; Santa Cruz Biotechnology, Inc.) antibodies were used for immunoblot and immunofluorescence analyses. Anti-HtrA2/Omi rabbit polyclonal (R&D Systems), anti-paxillin mouse monoclonal (BD), and anti–cytochrome c mouse monoclonal (6H2; Santa Cruz Biotechnology, Inc.) antibodies were used for immunofluorescence analysis. Anti-p130Cas (αCas3) antibody was generated against a bacterially expressed fragment of p130Cas as described elsewhere (Sakai et al., 1994). Anti–phospho-HtrA2/Omi (Ser 142) antibody was generated against the phosphopeptide SPPPA(pS)PRSQYC (a gift from Hélène Plun-Favreau, University College London, London, England, UK). Ucf-101, blebbistatin, and Adriamycin were purchased from EMD Millipore. BIRB796 was purchased from LC Laboratories. Cytochalasin D, latrunculin A, jasplakinolide, Q-VD-OPh, carbonyl cyanide 3-chlorophenylhydrazone (CCCP), carbonyl cyanide 4-trifluoromethoxyphenylhydrazone (FCCP), and oligomycin were purchased from Sigma-Aldrich.
Immunoblot analysis
Cells were solubilized with ice-cold lysis buffer (50 mM Tris, pH 8.0, 150 mM NaCl, 1% Triton X-100, 0.5% SDS, 10 mM EDTA, 1 mM Na3VO4, 10 mM NaF, and protease inhibitor cocktail [Nacalai Tesque]). Lysates were sonicated and centrifuged at 20,000 g for 20 min. The supernatants were then subjected to SDS-PAGE. For immunoblotting against β-actin, cells were solubilized with SDS sample buffer. For immunoblotting against phosphorylated HtrA2/Omi for total cell lysates, cells were solubilized with 1% CHAPS lysis buffer (1% CHAPS, 20 mM Tris, pH 7.4, 137 mM NaCl, 1 mM EDTA, and 10% glycerol).
Live-cell imaging
Protrusions of cells expressing Lifeact-GFP (a gift from Roland Wedlich-Söldner, University of Munster, Munster, Germany; Riedl et al., 2008) in cell culture medium were observed with a spinning-disk confocal microscopy system (UltraVIEW VoX; PerkinElmer) with an inverted microscope (model IX81; Olympus), a 100× NA 1.40 oil immersion UPlanSApo objective (Olympus), and an electron multiplying charge-coupled device 14-bit 1K × 1K camera (ImagEM-1K C9100-50; Hamamatsu Photonics) at 37°C in 5% CO2. A 488-nm laser was used for imaging. Images were acquired at an interval of 10 s for 5 min and then exported as TIFF files using Volocity software (version 6.0.1; PerkinElmer). All images were cropped and converted to AVI files using ImageJ (version 1.45f; National Institutes of Health). To observe localization of Bax, GFP-Bax, and DsRed-Mito were simultaneously visualized with the same microscope setup as that for Lifeact-GFP except that an electron multiplying charge-coupled device 16-bit 512 × 512 camera (ImagEM-Enhanced C9100-13; Hamamatsu Photonics) was used.
Immunofluorescence
Immunofluorescence staining for p130Cas or paxillin was performed as described previously (Kawauchi et al., 2012). Cells were fixed with 4% PFA, permeabilized with 0.2% Triton X-100, and then blocked with 2% BSA in cytoskeletal stabilizing buffer (137 mM NaCl, 5 mM KCl, 1.1 mM sodium phosphate [dibasic], 0.4 mM monopotassium phosphate, 4 mM sodium bicarbonate, 2 mM MgCl2, 5.5 mM glucose, 2 mM EGTA, and 5 mM Pipes, pH 6.1). Alexa Fluor 488– or 546–conjugated goat anti–mouse and anti–rabbit IgG (Molecular Probes) were used as secondary antibodies. Alexa Fluor 488– or 594–conjugated phalloidin (Molecular Probes) was used to stain F-actin. For immunofluorescence against cytochrome c, cells were cultured in the presence or absence of 100 µM Q-VD-OPh for 12 h, fixed with 4% PFA, permeabilized with 100% methanol, and then blocked with 2% BSA in PBS. Alexa Fluor 546–conjugated goat anti–mouse IgG was used as a secondary antibody. Immunofluorescence staining for exogenous p53 was performed as for p130Cas, except that PBS was used instead of cytoskeletal stabilizing buffer. Alexa Fluor 488–conjugated goat anti–mouse IgG was used as secondary antibody. Images of cells in PBS were acquired using a confocal microscopy system (A1Rsi; Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 100× NA 1.40 Plan Apochromat VC objective (Nikon), and an electron multiplying charge-coupled device 12-bit 512 × 512 camera (DU897; Andor Technology) at RT. Both 488- and 561-nm lasers were used for imaging. Images were acquired using NIS-Elements imaging software (version 4.10.00; Nikon). Acquired images were cropped using ImageJ software (version 1.45f). Immunofluorescence staining for HtrA2/Omi was performed as for p53. Cells in PBS were observed with a spinning-disk confocal microscopy system (UltraVIEW VoX; PerkinElmer) with an inverted microscope (model IX81; Olympus), a 100× NA 1.40 oil immersion UPlanSApo objective (Olympus), and an electron multiplying charge-coupled device 14-bit 1K × 1K camera (ImagEM-1K C9100-50; Hamamatsu Photonics) at 37°C with 5% CO2. Both 488- and 561-nm lasers were used for imaging. Images were acquired and then exported as TIFF files using Volocity software (version 6.0.1). Images were cropped and analyzed using ImageJ software (version 1.45f). To evaluate the release of HtrA2/Omi from mitochondria into cytosol, fluorescence intensities of three areas (1 µm in diameter) devoid of Mito-DsRed fluorescence in each cell were quantified.
Super-resolution imaging of mitochondria
The spatial organization of mitochondria in live cells expressing Mito-DsRed in cell culture medium was analyzed using a three-dimensional structure illumination microscopy system (N-SIM; Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 100× NA 1.49 oil immersion CFI Plan Apo IR objective (Nikon), and an electron multiplying charge-coupled device 16-bit 512 × 512 camera (iXon3 897; Andor Technology) at 37°C with 5% CO2. A 561-nm laser was used for imaging. Images were acquired using NIS-Elements imaging software (version 4.11.00). Reconstructed z-stack images were segmentalized and analyzed using Imaris (Bitplane AG). Each mitochondrion was identified and fitted with a single ellipsoidal object. The length (along the long-axis) of each ellipsoid-fitted mitochondrion was measured (defined as mitochondria length) and then used to compare differences in mitochondrial shape. The cumulative normalized distribution of mitochondria length was analyzed and plotted with the SEM in Prism (GraphPad Software).
Subcellular fractionation
Mitochondria were isolated using a mitochondria isolation kit (QIAGEN) according to the manufacturer’s protocol. The isolated mitochondrial fraction was solubilized with ice-cold lysis buffer (50 mM Tris, pH 8.0, 150 mM NaCl, 1% Triton X-100, 10 mM EDTA, 1 mM Na3VO4, 10 mM NaF, and protease inhibitor cocktail).
Analysis of F/G-actin ratio
Analysis of the F/G-actin ratio was performed as described previously (Kawauchi et al., 2012). F-actin and G-actin were stained with Alexa Fluor 488–phalloidin and Alexa Fluor 594–DNase I, respectively. Epifluorescence images of cells in PBS were acquired using a confocal microscopy system (A1Rsi; Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 60× NA 1.40 CFI Plan Apochromat VC oil immersion objective (Nikon), and an electron multiplying charge-coupled device 16-bit 512 × 512 camera (DU897; Andor Technology) at RT. Both 488- and 561-nm lasers were used for imaging. Images were acquired using NIS-Elements imaging software (version 4.10.00). Acquired images were analyzed using ImageJ software (version 1.45f). The intensity ratios of F/G-actin were calculated for each pixel and the calculated values were averaged within individual cell regions.
Inverted invasion assay
Matrigel (BD) supplemented with 25 µg/ml fibronectin (EMD Millipore) was polymerized in 8-µm pore-sized Transwell inserts (Corning) for 1 h at 37°C. The Transwell inserts were inverted before 3 × 104 cells labeled with carboxyfluorescein diacetate succinimidyl ester (CFDA-SE; Molecular Probes) were seeded onto the base of each insert. After incubating for 6 h at 37°C, the Transwell inserts were placed in serum-free medium. To establish a chemotactic gradient, medium supplemented with 10% FBS and 50 ng/ml EGF was added to the upper chamber. Cells were allowed to invade into the Matrigel for 72 h at 37°C. Images of cells in the Transwell inserts were acquired using a confocal microscopy system (A1Rsi; Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 20× NA 0.75 Plan Apochromat VC objective (Nikon), and an electron multiplying charge-coupled device 16-bit 512 × 512 camera (DU897; Andor Technology) at RT. A 488-nm laser was used for imaging. Images were acquired at 10-µm intervals using NIS-Elements imaging software (version 4.10.00). Cell areas were measured using ImageJ software (version 1.45f). The ratio of cells that had invaded 30 µm or more was calculated.
Purification of HtrA2/Omi proteins
HEK293T cells were transfected with pCMV-Flag-mature HtrA2/Omi and solubilized with ice-cold lysis buffer (50 mM Tris, pH 7.4, 150 mM NaCl, 1% Triton X-100, 10 mM EDTA, 1 mM Na3VO4, 10 mM NaF, and protease inhibitor cocktail). Lysates were centrifuged at 20,000 g for 20 min at 4°C. The Flag-mature HtrA2/Omi were captured on anti-Flag M2 affinity gel (Sigma-Aldrich) and eluted by competition with free 3× Flag peptide (Sigma-Aldrich) dissolved in TBS buffer (50 mM Tris, pH 7.4, and 150 mM NaCl) for 3 h at 4°C.
In vitro F-actin severing
Chamber slides were coated with 1% 3-triethoxysilylpropylamine in methanol for 1 h and 0.11% biotin-polyethylene glycol (PEG) and 4.4% mPEG (Laysan Bio) in 100 mM Na2CO3 for 2 h at room temperature. The coated chamber slides were blocked with 4% BSA in PBS overnight and incubated with 10 ng/ml NeutrAvidin (Thermo Fisher Scientific) for 5 min at room temperature. 0.6 µM actin, 0.03 µM biotin-labeled actin (Cytoskeleton, Inc.), and 0.14 µM Alexa Fluor 488–conjugated actin (Molecular Probes) were converted into Mg2+-actin in Mg2+ buffer (2 mM Tris, pH 8.0, 0.1 mM MgCl2, 1 mM DTT, 0.4 mM ATP, and 0.01% NaN3) and polymerized in 1× KMEI (50 mM KCl, 1 mM MgCl2, 1 mM EGTA, and 10 mM imidazole, pH 7.0). We included 3 mg/ml glucose and catalase/glucose oxidase (Sigma-Aldrich) to prevent photobleaching and 20 mM phosphoenol (Sigma-Aldrich), 100 U/ml pyruvate kinase (Sigma-Aldrich), and 10 mM KCl to regenerate ATP. The polymerized actin was attached to the chamber slides through biotin-NeutrAvidin interactions and incubated with 0.2 µM purified HtrA2/Omi at room temperature. Images of the polymerized actin in 4% BSA containing PBS were acquired using a total internal reflection fluorescence (TIRF) microscopy system (Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 100× NA 1.49 oil immersion CFI Plan Apochromat TIRF objective (Nikon), and an electron multiplying charge-coupled device 16-bit 512 × 512 camera (iXon3 897; Andor Technology) at room temperature. A 488-nm laser was used for imaging. Images were acquired at an interval of 15 s for 1 h using NIS-Elements imaging software (version 4.13.01). Fluorescence intensities of images were quantified and converted to AVI files using ImageJ software (version 1.45f).
Traction force microscopy
Polyacrylamide substrates were prepared on 25-mm glass coverslips. 40 µl of mixed acrylamide solution containing 5% acrylamide, 0.1% N,N-methylene-bis-acrylamide (BIS), 0.2% TEMED, 0.1% wt/vol APS, and a 1:500 dilution of latex microspheres (Fluoresbrite Plain YG 0.2-µm microspheres; Polysciences) was applied onto a coverslip silanized with 3-methacryloxypropyltrimethoxysilane and allowed to polymerize evenly by covering with another untreated coverslip. Type-I collagen (Koken Co.) was covalently attached to the surface of the gel using the photoactivatable cross-linker sulfo-SANPAH (Thermo Fisher Scientific). Cells were seeded onto polyacrylamide substrates in cell culture medium. Images were acquired using a spinning-disk confocal microscopy system (UltraVIEW VoX; PerkinElmer) with an inverted microscope (model IX81; Olympus), a 20× NA 0.70 water immersion UApoN340 objective (Olympus), and an electron multiplying charge-coupled device 14-bit 1K × 1K camera (ImagEM-1K C9100-50; Hamamatsu Photonics) at 37°C with 5% CO2. Images of a cell and latex microspheres before and after cell detachment with trypsin were acquired with differential interference contrast (DIC) and 440-nm laser, respectively, using Volocity software (version 6.0.1). Once the entire displacement field was calculated from images of latex microspheres using MATLAB software (version R2012a), the traction stress field was obtained as the solution to the inverse Boussinesq problem as described previously (Yip et al., 2013).
F-actin sedimentation assay
F-actin and G-actin were isolated as described previously with a slight modification (Kiuchi et al., 2007; Akin and Mullins, 2008). Cells were solubilized with ice-cold lysis buffer (50 mM Hepes, pH 7.4, 100 mM NaCl, 1 mM MgCl2, 0.2 mM CaCl2, 1 mM DTT, 1% NP-40, 2 µM phalloidin [Sigma-Aldrich], and 2 µM latrunculin B [EMD Millipore]). Lysates were centrifuged at 100,000 g for 30 min at 4°C. Equal amounts of pellet (F-actin) and supernatant (G-actin) were subjected to SDS-PAGE. Blots against actin (Cytoskeleton, Inc.) were quantified using ImageJ software.
Monitoring mitochondrial membrane potential
Cells were cultured on glass-bottom dishes in the presence of JC-1 (2 µM; Molecular Probes), a cationic dye that accumulates in energized mitochondria, for 30 min. Images of cells in PBS were acquired using a confocal microscopy system (A1Rsi; Nikon) with an inverted microscope (Eclipse Ti-E; Nikon), a 40× NA 0.95 Plan Apochromat objective (Nikon), and an electron multiplying charge-coupled device 16-bit 512 × 512 camera (DU897; Andor Technology) at 37°C with 5% CO2. Both 488- and 561-nm lasers were used for imaging monomers and J-aggregate forms, respectively. Images were acquired using NIS-Elements imaging software (version 4.10.00). Acquired images were cropped and fluorescence intensities were quantified using ImageJ software (version 1.45f).
Quantitative real-time PCR
Quantitative real-time PCR analysis was performed as described previously (Kawauchi et al., 2012). The following primers were used. Mouse PUMA forward 5′-ACGACCTCAACGCGCAGTACG-3′, reverse 5′-GAGGAGTCCCATGAAGAGATTG-3′; mouse BAX forward 5′-CAGGATGCGTCCACCAAGAA-3′, reverse 5′-AGTCCGTGTCCACGTCAGCA-3′; and mouse cyclophilin A forward 5′-CCTTGGGCCGCGTCTCCTT-3′, reverse 5′-CACCCTGGCACATGAATCCTG-3′. After normalization against cyclophilin A, mRNA expression levels relative to control expression levels were calculated for each experiment.
Statistical analysis
Data were analyzed by unpaired Student’s t test.
Online supplemental material
Fig. S1 shows that oncogenic Ras decreases cell-spreading area in a p130Cas-independent manner. Fig. S2 confirms that oncogenic Ras decreases the ratio of F/G-actin in a p53-dependent manner. Fig. S3 shows that caspase-3 is not involved in Ras-induced cleavage of β-actin. Fig. S4 suggests that the transcriptional activity of p53 is dispensable for Ras-induced translocation of p38 MAPK to mitochondria. Fig. S5 suggests roles for Bax and a decrease in oxidative phosphorylation in Ras-induced release of HtrA2/Omi. Video 1 shows actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell. Video 2 shows actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi-shRNA. Video 3 shows actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing HtrA2/Omi- and p130Cas-shRNA. Video 4 shows actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing p53R175H. Video 5 shows actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing p53R175H with p130Cas-shRNA. Video 6 shows images of F-actin incubated with wild-type HtrA2/Omi. Video 7 shows images of F-actin incubated with HtrA2/Omi-S306A. Video 8 shows images of F-actin incubated with HtrA2/Omi-S142D.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing mutant p53. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a p53 R175H-expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing mutant p53. NIH3T3 cells were infected with a Ha-RasV12–expressing retrovirus together with a p53 R175H-expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing mutant p53 with p130Cas-shRNA. NIH3T3 cells were infected with Ha-RasV12– and p53 R175H–expressing retroviruses together with a p130Cas shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Live imaging of actin filament dynamics in cell membrane protrusions from a Ras-transformed NIH3T3 cell expressing mutant p53 with p130Cas-shRNA. NIH3T3 cells were infected with Ha-RasV12– and p53 R175H–expressing retroviruses together with a p130Cas shRNA–expressing retrovirus and transfected with a Lifeact-GFP expression vector to label F-actin. Live imaging of protrusion was performed with an inverted microscope (model IX81; Olympus) equipped with a spinning-disk confocal unit (PerkinElmer), an oil-immersion objective (100×, NA 1.40, PL FL; Olympus), and an EMCCD camera (model C9100-50; Hamamatsu Photonics). Frames were taken every 10 s for 5 min.
Images of F-actin incubated with wild-type HtrA2/Omi. Alexa Fluor 488–labeled F-actin was incubated with purified wild-type HtrA2/Omi. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Images of F-actin incubated with wild-type HtrA2/Omi. Alexa Fluor 488–labeled F-actin was incubated with purified wild-type HtrA2/Omi. Images were analyzed by TIRF microscopy using a TIRF microscope (N-STORM; Nikon) equipped with an oil-immersion objective (100×, NA 1.49, CFI Plan Apochromat TIRF; Nikon). Frames were taken every 15 s for 55 min.
Acknowledgments
We thank L. Kenney, M. Sheetz, G.V. Shivashankar, B.C. Low, Y. Ito, K. Kohu, A. Bershadsky, S. Wolf, H. Machiyama, M. Sokabe, and Y. Nishimura for discussion; F.B. Abu Bakar for technical support; I. Uehara for preparation of p53–/–MEFs; R. Wedlich-Soldner for Lifeact-GFP expression vector; S. Hirose for pcDNA3-Flag Mfn2; and H. Plun-Favreau for anti–phospho-HtrA2/Omi antibody. All microscopy was carried out at the microscopy core facility, MBI, National University of Singapore.
This work was supported by the National Research Foundation, Singapore, and the Ministry of Education, Singapore, through the Mechanobiology Institute; and a Kurata Grant from the Kurata Memorial Hitachi Science and Technology Foundation, Japan.
The authors declare no competing financial interests.
References
Author notes
S. Yamauchi’s present address is Faculty of Medical Sciences, University of Fukui, Fukui 910-1193, Japan.
K. Kawauchi’s present address is Frontiers of Innovative Research in Science and Technology (FIRST), Konan University, Kobe 650-0047, Japan.