The membrane remodeling events required for autophagosome biogenesis are still poorly understood. Because PX domain proteins mediate membrane remodeling and trafficking, we conducted an imaging-based siRNA screen for autophagosome formation targeting human PX proteins. The PX-BAR protein SNX18 was identified as a positive regulator of autophagosome formation, and its Drosophila melanogaster homologue SH3PX1 was found to be required for efficient autophagosome formation in the larval fat body. We show that SNX18 is required for recruitment of Atg16L1-positive recycling endosomes to a perinuclear area and for delivery of Atg16L1- and LC3-positive membranes to autophagosome precursors. We identify a direct interaction of SNX18 with LC3 and show that the pro-autophagic activity of SNX18 depends on its membrane binding and tubulation capacity. We also show that the function of SNX18 in membrane tubulation and autophagy is negatively regulated by phosphorylation of S233. We conclude that SNX18 promotes autophagosome formation by virtue of its ability to remodel membranes and provide membrane to forming autophagosomes.
Introduction
Autophagy is important for human health and development through protection against neurodegeneration and cancer, removal of invading pathogens, and promotion of life-span extension (Mizushima and Komatsu, 2011). Moreover, autophagy ensures cellular quality control at basal levels and recycles nutrients to permit cellular survival during stresses like starvation (Levine and Kroemer, 2008). Macroautophagy (here referred to as autophagy) is characterized by the sequestration of cytoplasmic material through expansion and closure of a phagophore membrane, forming double-membrane vesicles called autophagosomes. The autophagosomes mature by fusion with endosomes and finally fuse with lysosomes, where the contents are degraded and the products recycled to the cytosol for reuse.
The process of forming an autophagosome requires membrane remodeling and trafficking, and is still poorly understood. The origin of the autophagic membrane is a subject of debate, with the ER, Golgi, mitochondria, plasma membrane, and recycling endosomes as suggested sources (Axe et al., 2008; Hayashi-Nishino et al., 2009; Hailey et al., 2010; Ravikumar et al., 2010; Guo et al., 2012; Longatti et al., 2012). It is also unclear how the membrane curvature is generated. More than 30 proteins assisting this process have been described as Autophagy-related (Atg) proteins, which function in a hierarchical order to mediate autophagosome biogenesis (Xie and Klionsky, 2007; Itakura and Mizushima, 2010). Four multiprotein complexes have been found to be required for autophagosome formation, including the Atg1/ULK1 complex, the class III phosphatidylinositol 3-kinase (PI3K) complex with the associated subunit Atg14L, the Atg9 trafficking system, and finally, the two ubiquitin-like proteins Atg12 and Atg8/LC3 and their conjugation systems. In brief, Atg12 is conjugated to Atg5 through the activity of the Atg7 (E1-like) and Atg10 (E2-like) enzymes, followed by their interaction with membrane-bound Atg16L. Atg8/LC3 becomes conjugated to phosphatidylethanolamine in a reaction that requires Atg7 and the E2-like enzyme Atg3, and is facilitated by the Atg5–Atg12–Atg16L1 complex (Hanada et al., 2007).
PX domain proteins are known to mediate membrane remodeling and trafficking dependent on phosphoinositide binding (Seet and Hong, 2006). There are 47 human PX domain proteins, including the sorting nexins (SNXs), and many also contain Bin/Amphiphysin/Rvs homology (BAR) domains, which are sensors and inducers of membrane curvature (Itoh and De Camilli, 2006). To better understand the phosphoinositide signaling, membrane remodeling, and trafficking events in autophagy, we performed an siRNA screen targeting human PX domain proteins. Depletion of the PX-BAR protein SNX18 strongly inhibited the formation of GFP-LC3–positive autophagosomes, whereas overexpression of SNX18 promoted GFP-LC3 spot formation, dependent on its membrane binding and tubulating ability. SNX18 localizes to structures containing early autophagic markers and interacts with LC3 family members and Atg16L1. We propose a role for SNX18 in promoting LC3 lipidation on tubovesicular structures from recycling endosomes, thereby facilitating membrane delivery to the expanding phagophore. The role of SNX18 in autophagy is conserved, as the Drosophila melanogaster SNX18 homologue SH3PX1 is required for efficient autophagosome formation in larval fat body.
Results
siRNA screen reveals SNX18 as a positive regulator of autophagy
To uncover a role of PX domain proteins in autophagy, we performed an imaging-based siRNA-screen where HEK cells stably transfected with GFP-LC3 (HEK GFP-LC3; Chan et al., 2007) were transfected with pools of siRNA targeting PX domain proteins, using ULK1 and TSG101 as controls for reduced and increased autophagosome levels, respectively. The cells were starved or not starved before imaging and high-content image analysis where the total intensity (not depicted) and number of GFP-LC3 spots per cell (Fig. 1 A, Table S1, and Fig. S1 A) were quantified as a measure for autophagosome formation. SNX18 was a promising candidate because its silencing strongly inhibited GFP-LC3 spot formation in both fed and starved cells (Fig. 1 A).
siRNA screen for PX domain proteins in autophagy. (A) HEK GFP-LC3 cells were transfected with siRNA pools targeting PX domain proteins and starved (large circles) or not starved (small circles) for 2 h, followed by fixation and counterstaining of the nuclei by Hoechst. The images were processed for high-content image analysis to quantify the number of GFP-LC3 spots per cell. The graph shows the average of three independent experiments in triplicate with a total of 30,000 cells. siRNA against ULK1 and TSG101 were used as controls (blue circles). Green, negative controls; red, PX domain proteins. (B) Selected PX domain proteins were depleted by transfection of HEK GFP-LC3 with the four individual siRNA oligos of the pools used in A. The cells were treated, imaged, and analyzed as in A. The area of each circle corresponds to the relative amount of target mRNA as measured by qPCR. The graph shows the average of two independent experiments in triplicate with a total number of 18,000 cells. Coloring is as in A. (C) Micrographs of starved HEK GFP-LC3 cells treated with nontargeting or siSNX18-3 siRNA. (left and middle) 3 × 3 fields of cells. (right) Close-up of a few cells from the respective population. The images in the right panels are shown again in Fig. S1 B alongside cells treated with other SNX18-targeting siRNAs. Bars, 10 µm. See also Table S1 and Table S2.
siRNA screen for PX domain proteins in autophagy. (A) HEK GFP-LC3 cells were transfected with siRNA pools targeting PX domain proteins and starved (large circles) or not starved (small circles) for 2 h, followed by fixation and counterstaining of the nuclei by Hoechst. The images were processed for high-content image analysis to quantify the number of GFP-LC3 spots per cell. The graph shows the average of three independent experiments in triplicate with a total of 30,000 cells. siRNA against ULK1 and TSG101 were used as controls (blue circles). Green, negative controls; red, PX domain proteins. (B) Selected PX domain proteins were depleted by transfection of HEK GFP-LC3 with the four individual siRNA oligos of the pools used in A. The cells were treated, imaged, and analyzed as in A. The area of each circle corresponds to the relative amount of target mRNA as measured by qPCR. The graph shows the average of two independent experiments in triplicate with a total number of 18,000 cells. Coloring is as in A. (C) Micrographs of starved HEK GFP-LC3 cells treated with nontargeting or siSNX18-3 siRNA. (left and middle) 3 × 3 fields of cells. (right) Close-up of a few cells from the respective population. The images in the right panels are shown again in Fig. S1 B alongside cells treated with other SNX18-targeting siRNAs. Bars, 10 µm. See also Table S1 and Table S2.
The candidates that affected GFP-LC3 spot formation the most were further validated in a secondary screen where HEK GFP-LC3 cells were transfected with the four individual siRNA oligonucleotides from the pool used in the primary screen (Fig. 1 B and Table S2). To be selected for further analysis, three out of four individual siRNA oligos had to reproduce the effect from the primary screen and correlate with the level of knockdown, as measured by quantitative real-time PCR (qPCR; Fig. 1 B, spot size correlating to mRNA level). SNX18 and HS1BP3 passed our selection criteria, and in the present study we characterized the role of SNX18 in autophagy.
SNX18 is required for autophagosome formation
SNX18 is a member of the SNX9 family of PX-BAR proteins (Håberg et al., 2008). The membrane tubulation activity of these proteins made SNX18 an interesting candidate for a protein involved in autophagosome biogenesis. In line with the decreased number of GFP-LC3 spots seen in the secondary screen (Fig. 1, B and C: Fig. S1; and Table S2), SNX18 depletion also inhibited the conjugation of GFP-LC3 to phosphatidylethanolamine, as measured by quantification of lipidated LC3 (LC3-II) versus nonlipidated LC3 (LC3-I; Kabeya et al., 2000), in correlation with the level of SNX18 mRNA and protein knockdown (Fig. 2 A, Fig. S1 C, and Fig. S2 A). In the following experiments we used the best oligo from the pool (siSNX18-3) as well as another (siSNX18-5), which also efficiently depleted SNX18 protein and decreased GFP-LC3-II levels and GFP-LC3 spot formation upon starvation (Figs. 2 A and S2 B). SNX18 depletion did not decrease LC3 mRNA levels (Fig. S1 D). As a decrease in GFP-LC3 spots and GFP-LC3-II levels during starvation could result from inhibited formation or increased turnover of autophagosomes, we measured the autophagic flux in starved cells in the absence or presence of Bafilomycin A1 (BafA1), which inhibits lysosomal degradation. SNX18-depleted cells treated with BafA1 showed decreased accumulation of GFP-LC3-II compared with control cells (Fig. 2 A), confirming that depletion of SNX18 inhibits autophagosome formation. In line with this, depletion of SNX18 inhibited the starvation-induced degradation of long-lived proteins (Fig. 2 B). SNX18 levels did not change during starvation, nor accumulate upon BafA1 treatment (Fig. 2 A), which suggests that SNX18 itself is not an autophagic cargo. The inhibitory effect of siSNX18 on GFP-LC3 spot formation was rescued by siRNA-resistant myc-SNX18, confirming that the effect is specific (Fig. 2 C and Fig. S3 H). Interestingly, expression of myc-SNX18 in SNX18-depleted cells increased the number of GFP-LC3 spots compared with control cells (Fig. 2 C), which indicates that SNX18 overexpression may stimulate biogenesis of autophagosomal membranes. Indeed, expression of myc-SNX18 caused increased lipidation of endogenous LC3 (Fig. 2 D), as well as an increased number of GFP-LC3 spots, dependent on the core autophagy machinery (Fig. 2 E and Fig. S2 D). However, myc-SNX18 expression did not increase degradation of long-lived proteins (Fig. 2 F), indicating that although SNX18 overexpression stimulates LC3 lipidation and GFP-LC3 spot formation, it is insufficient for increased autophagic flux. Collectively, these results establish SNX18 as a positive regulator of autophagosome formation.
SNX18 is a positive regulator of autophagy. (A) HEK GFP-LC3 cells transfected with control (siCtrl) or two different SNX18 siRNA oligos (siSNX18-3 and siSNX18-5) were starved or not starved for 2 h in the presence or absence of BafA1. GFP-LC3 lipidation and SNX18 protein knockdown were monitored by immunoblotting. The graph shows the average GFP-LC3-II relative to actin normalized to siCtrl-1 fed ± SEM (error bars), n = 4. (B) The degradation of long-lived proteins in HeLa cells transfected with control or SNX18 siRNA was quantified after 4 h of starvation in the absence or presence of 3-MA and normalized to the degradation in fed cells (mean ± SEM [error bars], n = 3). (C) HEK GFP-LC3 cells were transfected with control or SNX18 siRNA and then with a myc control or siRNA-resistant myc-SNX18 plasmid. The number of GFP-LC3 spots per cell was quantified (graph shows mean ± SEM [error bars], n = 3). (D) HEK GFP-LC3 cells were transfected with myc-SNX18 or a myc control plasmid and treated as in A. The graph shows the average LC3-II levels relative to actin normalized to myc-transfected fed cells, ±SEM (error bars). n = 3. (E) HEK GFP-LC3 cells were transfected with control or Atg7 siRNA and then with myc-SNX18 or a myc control plasmid, followed by quantification of the number of GFP-LC3 spots per cell (graphs show mean ± SEM [error bars], n = 3). Representative images are shown. Bars, 5 µm. (F) The degradation of long-lived protein in HeLa cells transfected with myc-SNX18 or a myc control plasmid was quantified as in B (graph shows mean ± SEM [error bars], n = 4). *, P < 0.05; **, P < 0.01. See also Figs. S1 and S2.
SNX18 is a positive regulator of autophagy. (A) HEK GFP-LC3 cells transfected with control (siCtrl) or two different SNX18 siRNA oligos (siSNX18-3 and siSNX18-5) were starved or not starved for 2 h in the presence or absence of BafA1. GFP-LC3 lipidation and SNX18 protein knockdown were monitored by immunoblotting. The graph shows the average GFP-LC3-II relative to actin normalized to siCtrl-1 fed ± SEM (error bars), n = 4. (B) The degradation of long-lived proteins in HeLa cells transfected with control or SNX18 siRNA was quantified after 4 h of starvation in the absence or presence of 3-MA and normalized to the degradation in fed cells (mean ± SEM [error bars], n = 3). (C) HEK GFP-LC3 cells were transfected with control or SNX18 siRNA and then with a myc control or siRNA-resistant myc-SNX18 plasmid. The number of GFP-LC3 spots per cell was quantified (graph shows mean ± SEM [error bars], n = 3). (D) HEK GFP-LC3 cells were transfected with myc-SNX18 or a myc control plasmid and treated as in A. The graph shows the average LC3-II levels relative to actin normalized to myc-transfected fed cells, ±SEM (error bars). n = 3. (E) HEK GFP-LC3 cells were transfected with control or Atg7 siRNA and then with myc-SNX18 or a myc control plasmid, followed by quantification of the number of GFP-LC3 spots per cell (graphs show mean ± SEM [error bars], n = 3). Representative images are shown. Bars, 5 µm. (F) The degradation of long-lived protein in HeLa cells transfected with myc-SNX18 or a myc control plasmid was quantified as in B (graph shows mean ± SEM [error bars], n = 4). *, P < 0.05; **, P < 0.01. See also Figs. S1 and S2.
SNX18 functions downstream of the class III PI3K complex in autophagosome biogenesis
SNX18 has been reported to localize to the plasma membrane and endosomes, and is also called SNAG1 (SNX-associated Golgi protein 1; Håberg et al., 2008; Park et al., 2010; Willenborg et al., 2011). We detected SNX18 in both the cytosol and membrane fractions of HeLa cells (Fig. S2 E), and when the membrane fraction was further separated according to density by gradient centrifugation (Lundmark and Carlsson, 2003, 2005), the majority of SNX18 banded at medium density in both fed and starved cells, partially co-migrating with markers of the ER (calnexin), Golgi (GM130), endosomes (Transferrin receptor [TfR], Rab11), and plasma membrane (SNX9; Fig. 3 A). SNX18 did not cofractionate with the majority of LC3-II–positive membranes, which supports the finding that it is not itself an autophagy cargo. Interestingly, a minor part of SNX18 was found on dense membranes together with the majority of membrane-associated LC3-I and GABARAP-I (Fig. 3 A). The identity of the LC3 and GABARAP bands were verified by siRNA and Western blotting (Fig. S2 F). A striking comigration was found between SNX18 and the core autophagy proteins Atg14L and Atg16L1 in the medium-density fractions, and with the latter also in the dense fractions, in both fed and starved cells (Fig. 3 A), which further suggests a role for SNX18 in autophagosome biogenesis. The SNX18-related protein SNX9 was heterogeneously distributed in heavier membrane fractions, and depletion of SNX9 did not inhibit autophagosome formation (Fig. 1 A and Fig. S2 C), which suggests that the role of SNX18 in autophagosome formation is not shared by SNX9.
SNX18 colocalizes with autophagic markers in the perinuclear area. (A) The total membrane fractions from fed or starved HeLa cells were analyzed by density gradient centrifugation and probed for the indicated proteins by immunoblotting. Migration of different organelles is indicated, based on the positions in the gradient of calnexin (ER), GM130 (Golgi stack/CGN), SNX9 (plasma membrane), AP-1, TfR and Rab11a (Endosomes/TGN), and LC3-II and GABARAP-II (autophagosomes). (B) HeLa cells were starved or not starved for 2 h and immunostained for SNX18 and GM130. Panels to the right show enlarged views of the boxed regions. (C) HeLa cells were transfected with GFP-Atg14L and immunostained for SNX18 and calnexin. The boxed regions are enlarged on the top and bottom. (D and E) HEK GFP-DFCP1 cells were starved or not starved for 50 min and immunostained for SNX18 and LC3 (D) or WIPI2 (E). Results shown are mean ± SEM (error bars), n = 3 with 30 cells in each experiment. *, P < 0.05. (F) HeLa cells were transfected with control or SNX18 siRNA, starved for 2 h, and immunostained against SNX18 and Atg16L1. (G) HEK GFP-DFCP1 cells were transfected with control or SNX18 siRNA, starved for 50 min, and immunostained for Atg16L1. All images are from confocal microscopy. Inset panels show enlarged views of the boxed regions. Bars, 5 µm. See also Fig. S3.
SNX18 colocalizes with autophagic markers in the perinuclear area. (A) The total membrane fractions from fed or starved HeLa cells were analyzed by density gradient centrifugation and probed for the indicated proteins by immunoblotting. Migration of different organelles is indicated, based on the positions in the gradient of calnexin (ER), GM130 (Golgi stack/CGN), SNX9 (plasma membrane), AP-1, TfR and Rab11a (Endosomes/TGN), and LC3-II and GABARAP-II (autophagosomes). (B) HeLa cells were starved or not starved for 2 h and immunostained for SNX18 and GM130. Panels to the right show enlarged views of the boxed regions. (C) HeLa cells were transfected with GFP-Atg14L and immunostained for SNX18 and calnexin. The boxed regions are enlarged on the top and bottom. (D and E) HEK GFP-DFCP1 cells were starved or not starved for 50 min and immunostained for SNX18 and LC3 (D) or WIPI2 (E). Results shown are mean ± SEM (error bars), n = 3 with 30 cells in each experiment. *, P < 0.05. (F) HeLa cells were transfected with control or SNX18 siRNA, starved for 2 h, and immunostained against SNX18 and Atg16L1. (G) HEK GFP-DFCP1 cells were transfected with control or SNX18 siRNA, starved for 50 min, and immunostained for Atg16L1. All images are from confocal microscopy. Inset panels show enlarged views of the boxed regions. Bars, 5 µm. See also Fig. S3.
To further investigate the cellular localization of SNX18 and its role in autophagy, cells costained for endogenous SNX18 and autophagy markers were analyzed by confocal microscopy. SNX18 was mainly present in a perinuclear area, colocalizing with the Golgi marker GM130, in both fed and starved HeLa cells (Fig. 3 B). Moreover, we observed a partial colocalization of SNX18 and GFP-Atg14L in areas outside the ER, although Atg14L also colocalized with the ER marker calnexin (Fig. 3 C). Atg14L has been shown to recruit the class III PI3K complex to the ER for localized production of phosphatidylinositol 3-phosphate (PI3P) at sites of autophagosome formation (called omegasomes; Axe et al., 2008) and can also promote membrane curvature through its BATS domain (Fan et al., 2011). GFP–double FYVE containing protein 1 (DFCP1), a marker of omegasomes, was found together with SNX18 and LC3 in the perinuclear area in both fed and starved cells; however, SNX18 did not colocalize with peripheral GFP-DFCP1– and LC3-positive spots formed upon starvation (Fig. 3 D). Moreover, whereas omegasome formation was strongly inhibited by depletion of the class III PI3K complex subunit Beclin 1, it was not significantly affected by SNX18 depletion (Fig. S3 B), which suggests that SNX18 functions downstream of the class III PI3K complex. In line with this, depletion of Atg14L and ULK1 prevented myc-SNX18–induced GFP-LC3 spot formation (Fig. S2 D). Depletion of SNX18 caused puncta of the PI3P-binding protein WIPI2 to accumulate (Fig. 3 E), which suggests that aberrant intermediate structures accumulate in the absence of SNX18. Endogenous Atg16L1 (or flag-Atg16L1) also colocalized specifically with SNX18 in the perinuclear region (Fig. 3 F and Fig. S3, C–E), and depletion of SNX18 prevented recruitment of Atg16L1 to this region (Fig. 3 F) and to omegasomes (Fig. 3 G), without affecting Atg16L1 protein levels (not depicted). Together, our data indicate that SNX18 functions downstream of DFCP1, Atg14L, and WIPI2 through recruitment of Atg16L1 to a perinuclear region to facilitate remodeling of LC3-positive membranes as a membrane source for autophagosome formation from omegasomes.
Binding of SNX18 to phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2)-containing membranes is needed for Atg16L1 spot formation and for SNX18-induced GFP-LC3 spots
To further address the molecular mechanisms underlying the role of SNX18 in autophagy, we took advantage of the fact that SNX18 overexpression stimulates LC3 lipidation and GFP-LC3 spot formation in an autophagy-specific manner (Fig. 2, D and E), and induces formation of membrane tubules containing GFP-LC3 and flag-Atg16L1 (Fig. 4, A and B). Endogenous Atg16L1 was also associated with myc-SNX18–induced tubules (unpublished data). Membrane tubules induced by SNX9 were negative for GFP-LC3 (Fig. S4 C). To investigate whether the membrane binding and tubulation activity of SNX18 are important for its role in autophagosome formation, we introduced mutations in the SNX18 PX-BAR region (Fig. 4 C), based on the resolved structure of the SNX9 PX-BAR unit (Pylypenko et al., 2007). The positively charged concave surface of the BAR domain dimer allows SNX9 to sense membrane curvature and provides affinity to the negatively charged membrane surface, and the SNX9 K366E/R367E mutation or mutation of the phosphoinositide-binding pocket, R286Q/Y287A/K288A, was found to abolish membrane binding (Pylypenko et al., 2007; Yarar et al., 2008). The corresponding mutations were introduced in mCherry-SNX18 (mCh-SNX18 K394E/R395E and R312Q/Y313A/K314A, designated KR and RYK, respectively) and expressed in HEK GFP-LC3 cells at similar levels (Fig. S3 F). The SNX18 KR and RYK mutants were defective in membrane binding (Fig. 4 D) and unable to induce GFP-LC3 spots (Fig. 4 E and Fig. S3 H) and LC3 lipidation (Fig. 4 F and Fig. S3 G) as efficiently as wild-type (WT) SNX18, which indicates that SNX18 membrane binding is important for its function in autophagy. SNX18 was previously found to bind PI(4,5)P2 (Håberg et al., 2008), and depletion of PI(4,5)P2 by ionomycin (Várnai and Balla, 1998; Fig. 4 G) or by rapalog-induced recruitment of a phosphoinositide 5-phosphatase to the plasma membrane (Varnai et al., 2006; Zoncu et al., 2007; Fig. 4 H) strongly inhibited the induction of GFP-LC3 spots seen in myc-SNX18–transfected cells. Inhibition of GFP-LC3 spot formation by ionomycin was also observed after starvation without SNX18 overexpression (Fig. S3 I). Furthermore, SNX18 membrane binding was important for starvation-induced formation of Atg16L1 spots, and although the total intensity of endogenous Atg16L1 spots was increased by expression of myc-SNX18, it was strongly decreased by myc-SNX18 KR compared with control-transfected cells (Fig. 4 I). Moreover, Atg16L1 depletion prevented SNX18-induced GFP-LC3 spots (unpublished data). The close colocalization and interdependence of SNX18 and Atg16L1 prompted us to investigate if they interact, and we found myc-SNX18 full-length or the PX-BAR domain, but not the SH3-LC region, to coimmunoprecipitate with flag-Atg16L1 (Fig. 4 J). Thus, binding of SNX18 to PI(4,5)P2-containing membranes is important for its role as a positive regulator of autophagy.
SNX18 binding to PI(4,5)P2 membranes is required for its function in autophagy. (A) HeLa GFP-LC3 cells were transfected to express myc-SNX18, immunostained against myc, and analyzed by confocal imaging. Asterisks indicate a nontransfected cell. Bars: (main panels) 20 µm; (inset) 10 µm. (B) HeLa cells transfected with flag-Atg16L1 and myc-SNX18 were starved or not starved for 2 h before immunostaining with anti-myc and anti-flag antibodies and confocal microscopy. Bars, 5 µm. (C) SNX18 domain structure. An N-terminal SH3 domain is followed by a low-complexity region (LC). The membrane-remodeling part consists of an amphipathic helix (helix 0), a PX domain that is flanked by sequences forming a Yoke domain (Y), and a BAR half-domain. The functional BAR domain is formed by dimerization. Truncated constructs and point mutations used are indicated. (D) HeLa cells were transfected with the indicated myc-SNX18 plasmids for 6 h and starved for an additional 2 h before harvest. Equal parts of cytosol (C) and membrane (M) fractions were analyzed by Western blotting. The graph shows the average membrane-bound myc-SNX18 ± SEM (error bars), n = 3. (E and F) HEK GFP-LC3 cells were transfected with the indicated mCherry-SNX18 (E) or mycSNX18 (F) constructs for 16 h. The number of GFP-LC3 spots per transfected cell (E; graph shows mean ± SEM [error bars], n = 5) or ratio of LC3-II to actin determined by Western blotting (F) was quantified. (G and H) HEK GFP-LC3 cells transfected with myc-SNX18 or a myc control plasmid alone (G) or together with CFP-FRB and mRFP-FKBP or mRFP-FKBP 5pase (H) for 6 h were treated or not treated with 5 µM ionomycin (G) or 2.5 µM A/C heterodimerizer (rapalog; H) for the last 3 h before analysis of the total intensity of GFP-LC3 spots per transfected cell (graph shows mean ± SEM [error bars], n = 3). (I) MEFs were transfected with myc, myc-SNX18 WT, or KR for 16 h, starved or not starved for 2 h, and immunostained for endogenous Atg16L1. Insets show enlarged views of the boxed regions. Bars, 5 µm. (J) HeLa cells were transfected for 12 h with the indicated constructs and starved for 90 min. Cells were solubilized and immunoprecipitated with anti-flag or an unrelated antibody of the same isotype (mock ab). Immunoprecipitates and 2% of the lysates were analyzed by immunoblotting against myc and flag. *, P < 0.05; **, P < 0.01. See also Fig. S3.
SNX18 binding to PI(4,5)P2 membranes is required for its function in autophagy. (A) HeLa GFP-LC3 cells were transfected to express myc-SNX18, immunostained against myc, and analyzed by confocal imaging. Asterisks indicate a nontransfected cell. Bars: (main panels) 20 µm; (inset) 10 µm. (B) HeLa cells transfected with flag-Atg16L1 and myc-SNX18 were starved or not starved for 2 h before immunostaining with anti-myc and anti-flag antibodies and confocal microscopy. Bars, 5 µm. (C) SNX18 domain structure. An N-terminal SH3 domain is followed by a low-complexity region (LC). The membrane-remodeling part consists of an amphipathic helix (helix 0), a PX domain that is flanked by sequences forming a Yoke domain (Y), and a BAR half-domain. The functional BAR domain is formed by dimerization. Truncated constructs and point mutations used are indicated. (D) HeLa cells were transfected with the indicated myc-SNX18 plasmids for 6 h and starved for an additional 2 h before harvest. Equal parts of cytosol (C) and membrane (M) fractions were analyzed by Western blotting. The graph shows the average membrane-bound myc-SNX18 ± SEM (error bars), n = 3. (E and F) HEK GFP-LC3 cells were transfected with the indicated mCherry-SNX18 (E) or mycSNX18 (F) constructs for 16 h. The number of GFP-LC3 spots per transfected cell (E; graph shows mean ± SEM [error bars], n = 5) or ratio of LC3-II to actin determined by Western blotting (F) was quantified. (G and H) HEK GFP-LC3 cells transfected with myc-SNX18 or a myc control plasmid alone (G) or together with CFP-FRB and mRFP-FKBP or mRFP-FKBP 5pase (H) for 6 h were treated or not treated with 5 µM ionomycin (G) or 2.5 µM A/C heterodimerizer (rapalog; H) for the last 3 h before analysis of the total intensity of GFP-LC3 spots per transfected cell (graph shows mean ± SEM [error bars], n = 3). (I) MEFs were transfected with myc, myc-SNX18 WT, or KR for 16 h, starved or not starved for 2 h, and immunostained for endogenous Atg16L1. Insets show enlarged views of the boxed regions. Bars, 5 µm. (J) HeLa cells were transfected for 12 h with the indicated constructs and starved for 90 min. Cells were solubilized and immunoprecipitated with anti-flag or an unrelated antibody of the same isotype (mock ab). Immunoprecipitates and 2% of the lysates were analyzed by immunoblotting against myc and flag. *, P < 0.05; **, P < 0.01. See also Fig. S3.
The tubulation activity of SNX18 is important for its role in autophagy and is regulated by phosphorylation
A short conserved sequence just upstream of the PX domain is suggested to form an amphipathic helix upon membrane contact (Pylypenko et al., 2007; referred to as helix 0), and is, together with the PX-BAR, required for efficient membrane tubulation by SNX9 (Lundmark and Carlsson, 2009). The PhosphoSitePlus database (http://www.phosphosite.org/) reports a phosphorylation in the corresponding SNX18 helix 0 at serine-233, which increased in cells treated with the autophagy-inducing drug rapamycin (Chen et al., 2009). We raised a phosphospecific antibody against this site (Fig. S3, J and K) and found that GFP-SNX18 showed a time-dependent increase in SNX18 S233 phosphorylation during starvation, correlating with increased levels of LC3-II, that was reversed by re-feeding of starved cells (Fig. 5 A). To elucidate the role of SNX18 S233 phosphorylation in autophagy, we made myc-SNX18 S233 nonphosphorylatable (S233A) or phosphomimicking (S233D) mutants. The S233A or S233D mutants bound as efficiently to membranes as WT myc-SNX18 (Fig. 4 D). However, the PX-BAR S233A mutant was less efficient at, and the S233D mutant completely incapable of, inducing membrane tubulation as compared with WT PX-BAR (Fig. 5 B). The two phospho mutants were also much less efficient at inducing GFP-LC3 spots (Fig. 5 C and Fig. S3 H) when expressed at similar levels to SNX18 WT (Fig. S3 L). Moreover, neither the membrane binding (KR and RYK)– nor the membrane tubulation (S233D)–deficient mutants were able to induce formation of GFP-LC3–positive tubules as efficiently as myc-SNX18 WT (Fig. 5 D). Interestingly, although expression of SNX18 PX-BAR did induce extensive membrane tubulation, these tubules were devoid of GFP-LC3 (Fig. 5 D), and the number of GFP-LC3 spots in these cells was reduced (Fig. 5 E), which indicates that the N-terminal part of SNX18 is required for recruitment of LC3 onto SNX18-induced tubules and for autophagosome formation. We further noticed that SNX18 overexpression induced a strong perinuclear localization of GFP-LC3 (Figs. 4 A and 5 D) that was neither seen in cells expressing the membrane-binding deficient SNX18 mutants (Fig. 5 D), nor upon depletion of Atg16L1 (Fig. 5 F), which suggests that SNX18 facilitates membrane recruitment of LC3 dependent on Atg16L1. Together these results indicate that the membrane-remodeling capability of SNX18 is required for autophagosome formation, and that this activity is negatively regulated by a phosphorylation in the amphipathic helix 0.
The tubulation activity of SNX18 is important for its role in autophagy and is regulated by phosphorylation. (A) HeLa GFP-SNX18 cells were starved for the indicated time, followed by incubation in full medium (Refeed) where indicated. Cell lysates were analyzed for LC3 by immunoblotting, or immunoprecipitated with GFP-Trap before immunoblotting against SNX18(pS233) and GFP. The ratio of pS233 to GFP, and the ratio of LC3-II to LC3-I, was quantified, and normalized to 1 for the zero-time sample in both cases (graph shows mean ± SEM [error bars], n = 4; *, P < 0.05). (B) The indicated myc-SNX18 PX-BAR constructs were transfected into HeLa cells, and their tubulation efficiency was measured by scoring the percentage of cells displaying more than three elongated structures longer than 5 µm (n = 300). The graph shows the mean of two independent experiments ± range (error bars). Bars, 10 µm. (C) The indicated myc-tagged SNX18 constructs were transfected into HEK GFP-LC3 cells, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 5. (D) HeLa GFP-LC3 cells were transfected with the indicated myc-tagged SNX18 constructs for 16 h, starved for 2 h, and analyzed by confocal imaging. Bars, 10 µm. (E) HEK GFP-LC3 cells were transfected with myc-SNX18 WT or myc-SNX18 PX-BAR constructs, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 3. (F) HeLa GFP-LC3 cells were transfected with siRNA against Atg16L1 and later with myc-SNX18 constructs before immunostaining and confocal imaging. Bar, 5 µm.
The tubulation activity of SNX18 is important for its role in autophagy and is regulated by phosphorylation. (A) HeLa GFP-SNX18 cells were starved for the indicated time, followed by incubation in full medium (Refeed) where indicated. Cell lysates were analyzed for LC3 by immunoblotting, or immunoprecipitated with GFP-Trap before immunoblotting against SNX18(pS233) and GFP. The ratio of pS233 to GFP, and the ratio of LC3-II to LC3-I, was quantified, and normalized to 1 for the zero-time sample in both cases (graph shows mean ± SEM [error bars], n = 4; *, P < 0.05). (B) The indicated myc-SNX18 PX-BAR constructs were transfected into HeLa cells, and their tubulation efficiency was measured by scoring the percentage of cells displaying more than three elongated structures longer than 5 µm (n = 300). The graph shows the mean of two independent experiments ± range (error bars). Bars, 10 µm. (C) The indicated myc-tagged SNX18 constructs were transfected into HEK GFP-LC3 cells, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 5. (D) HeLa GFP-LC3 cells were transfected with the indicated myc-tagged SNX18 constructs for 16 h, starved for 2 h, and analyzed by confocal imaging. Bars, 10 µm. (E) HEK GFP-LC3 cells were transfected with myc-SNX18 WT or myc-SNX18 PX-BAR constructs, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 3. (F) HeLa GFP-LC3 cells were transfected with siRNA against Atg16L1 and later with myc-SNX18 constructs before immunostaining and confocal imaging. Bar, 5 µm.
SNX18 interacts with LC3/GABARAP
Our findings that SNX18 cofractionates with LC3 (Fig. 3 A), and recruits GFP-LC3 to the perinuclear area and onto SNX18-induced membrane tubules (Fig. 5 D), indicated that SNX18 might interact with LC3. Indeed, endogenous SNX18 coimmunoprecipitated with GFP-LC3, and even more so with the lipidation-defective GFP-LC3 G120A mutant (Kabeya et al., 2000; Fig. 6 A), which suggests that SNX18 preferentially interacts with the nonlipidated form of LC3 (LC3-I). LC3-I is generally considered to be cytosolic, but cell fractionation (Fig. S2 E) and density gradient separation of the membrane fraction (Fig. 3 A) clearly show that LC3-I and GABARAP-I can associate with membranes. Interestingly, we observed a shift in the LC3-I/GABARAP-I distribution from heavier fractions to the medium-density LC3-II– and SNX18-positive membranes upon starvation, which might suggest that LC3 lipidation occurs on SNX18-positive membranes (Fig. 3 A). GFP-GABARAP was also detected on SNX18-induced membrane tubules (Fig. S4 B). Endogenous SNX18 from both fed and starved MDBK cell lysates bound strongly to recombinant GST-GABARAP and GST-LC3 (Fig. 6 B), further indicating that SNX18 interacts with nonlipidated Atg8 homologues.
SNX18 interacts with LC3 family members. (A) GFP, GFP-LC3, or GFP-LC3 G120A were immunoprecipitated from transfected HeLa cells. The immunoprecipitates and 2% of the lysate were analyzed by immunoblotting against SNX18. The GFP proteins were detected by Ponceau staining. (B) Lysates from fed or starved MDBK cells were incubated with the indicated glutathione-Sepharose–bound GST-tagged proteins. SNX18 was detected by immunoblotting. Ponceau staining shows the GST proteins. (C) In vitro translated GFP-p62 or GFP-SNX18 were incubated with the indicated GST-tagged Atg8 family proteins. Bound proteins were detected by autoradiography and GST proteins by Coomassie blue staining.
SNX18 interacts with LC3 family members. (A) GFP, GFP-LC3, or GFP-LC3 G120A were immunoprecipitated from transfected HeLa cells. The immunoprecipitates and 2% of the lysate were analyzed by immunoblotting against SNX18. The GFP proteins were detected by Ponceau staining. (B) Lysates from fed or starved MDBK cells were incubated with the indicated glutathione-Sepharose–bound GST-tagged proteins. SNX18 was detected by immunoblotting. Ponceau staining shows the GST proteins. (C) In vitro translated GFP-p62 or GFP-SNX18 were incubated with the indicated GST-tagged Atg8 family proteins. Bound proteins were detected by autoradiography and GST proteins by Coomassie blue staining.
SNX18, like p62, an autophagy receptor known to interact with Atg8 proteins (Pankiv et al., 2007), was found to interact with all Atg8 family members, although little interaction was observed with GABARAPL2 (Fig. 6 C). The interaction site was mapped to the SNX18 SH3-LC region (Fig. 7 A) and the exact sequence was determined by a peptide array covering the entire SNX18 SH3-LC sequence, where GST-GABARAP bound to all peptides containing 154-WDDEW-158 (Fig. 7 B), the same motif previously found to mediate binding of SNX18 to AP-1 (Håberg et al., 2008). Mutation of one or both of the tryptophan residues (W154S and/or W158S) abolished the interaction with LC3/GABARAP (Fig. 7, C and D). Furthermore, although tubules were still formed in cells expressing the myc-SNX18 W154S/W158S mutant, these were negative for GFP-LC3 (Fig. 7 E), and the number of GFP-LC3 spots was also significantly reduced compared with SNX18 WT (Fig. 7 F). In summary, SNX18 interacts with Atg8 proteins through a WDDEW sequence in its SH3-LC region, and this interaction is required for localization of GFP-LC3 to SNX18-induced tubules and for GFP-LC3 spot formation.
A WDDEW sequence in the SNX18 LC region mediates the interaction with LC3/GABARAP. (A) GFP-p62 or -SNX18 full-length, SH3-LC, or PX-BAR regions were in vitro translated and incubated with GST-LC3B or -GABARAP. The resulting pulldowns were separated by SDS-PAGE. Bound proteins were detected by autoradiography and GST proteins by Coomassie blue staining. (B) 18-mer peptides covering the entire sequence of the SNX18 SH3-LC region were spotted on a membrane that was incubated with GST or GST-GABARAP, which were detected by immunoblotting against GST. The peptide sequences that specifically bound GST-GABARAP are shown with the common WDDEW motif in bold. (C) Peptides with the sequence YGGYQASQGSDDDWDDEWDDSSTVADEPGAL (SNX18 WT) or with the first (SNX18 W154S), the second (SNX18 W158S), or both (SNX18 W154S/W158S) W mutated to S were spotted on membranes that were incubated with GST, GST-LC3B, or GST-GABARAP. Binding was analyzed as in B. (D) GFP-SNX18 WT and the indicated mutants were in vitro translated and incubated with GST-LC3B or -GABARAP, and their binding was analyzed as in A. (E) HeLa GFP-LC3 cells were transfected with myc-SNX18 W154S/W158S mutant, starved for 2 h, immunostained against myc, and analyzed by confocal imaging. Bar, 10 µm. (F) Indicated myc-SNX18 constructs were transfected into HEK GFP-LC3 cells, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 3. *, P < 0.05.
A WDDEW sequence in the SNX18 LC region mediates the interaction with LC3/GABARAP. (A) GFP-p62 or -SNX18 full-length, SH3-LC, or PX-BAR regions were in vitro translated and incubated with GST-LC3B or -GABARAP. The resulting pulldowns were separated by SDS-PAGE. Bound proteins were detected by autoradiography and GST proteins by Coomassie blue staining. (B) 18-mer peptides covering the entire sequence of the SNX18 SH3-LC region were spotted on a membrane that was incubated with GST or GST-GABARAP, which were detected by immunoblotting against GST. The peptide sequences that specifically bound GST-GABARAP are shown with the common WDDEW motif in bold. (C) Peptides with the sequence YGGYQASQGSDDDWDDEWDDSSTVADEPGAL (SNX18 WT) or with the first (SNX18 W154S), the second (SNX18 W158S), or both (SNX18 W154S/W158S) W mutated to S were spotted on membranes that were incubated with GST, GST-LC3B, or GST-GABARAP. Binding was analyzed as in B. (D) GFP-SNX18 WT and the indicated mutants were in vitro translated and incubated with GST-LC3B or -GABARAP, and their binding was analyzed as in A. (E) HeLa GFP-LC3 cells were transfected with myc-SNX18 W154S/W158S mutant, starved for 2 h, immunostained against myc, and analyzed by confocal imaging. Bar, 10 µm. (F) Indicated myc-SNX18 constructs were transfected into HEK GFP-LC3 cells, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 3. *, P < 0.05.
Recycling endosomes provide membrane for SNX18-mediated autophagosome biogenesis
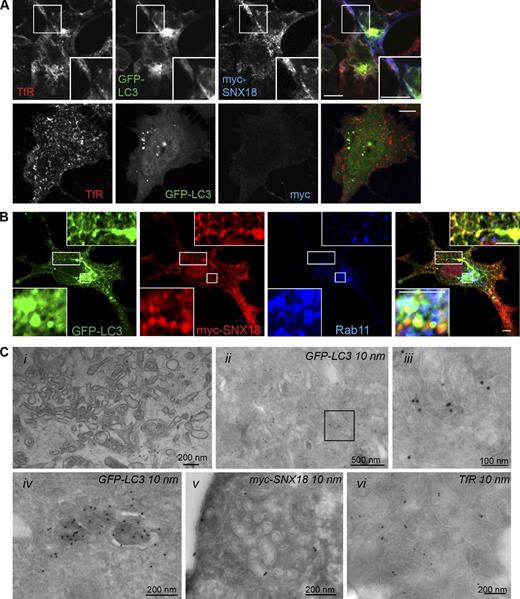
Our results thus far indicate that SNX18 is involved in membrane remodeling and recruitment of Atg16L1 and LC3 to membranes contributing to autophagosome biogenesis. We asked whether these were endosomal membranes, and found TfR and the small GTPase Rab11, involved in trafficking from recycling endosomes, to colocalize with perinuclear GFP-LC3 in myc-SNX18–expressing cells (Fig. 8, A and B), whereas no colocalization with early endosomal antigen 1 (EEA1), LAMP-1, or lyso-bisphosphatidic acid (LBPA) was detected (Fig. S4 D). Interestingly, TfR (Fig. 8 A), but not Rab11 (Fig. 8 B), was detected on the GFP-LC3–positive SNX18-induced tubules. SNX18-induced membrane tubules were also apparent at the ultrastructural level, and clusters of tubulo-vesicular structures together with small multivesicular body–like vesicles were typically seen (Fig. 8 C, i). By immuno-EM these tubules were found to be positive for myc-SNX18, GFP-LC3, and TfR (Fig. 8 C). GFP-LC3–labeled structures that resembled more typical autophagosomal vesicles were also found (Fig. 8 C, iv), although at similar levels in myc-SNX18– and myc-transfected cells. Also in nontransfected cells, Rab11 was strongly recruited to the SNX18-positive perinuclear area upon starvation, where it partially colocalized with TfR (Fig. 9 A), but not with EEA1 (Fig. S3 A). SNX18 also partially cofractionated with Rab11 and TfR (Fig. 3 A).
SNX18 tubules are positive for TfR. HeLa GFP-LC3 cells were transfected with myc-SNX18 or a control myc plasmid and then immunostained for myc and TfR (A) or Rab11 (B). Bars: (A) 10 µm; (B) 5 µm. Insets show enlarged views of the boxed regions. (C, i) Clusters of tubulo-vesicular structures together with small multivesicular body–like vesicles were observed upon plastic embedding of HeLa GFP-LC3 cells overexpressing myc-SNX18. (ii–vi) Immuno-EM showed that similar clusters labeled strongly for GFP-LC3 (ii, and enlarged in iii from the boxed region), myc-SNX18 (v), and TfR (vi). Strong GFP-LC3 labeling in structures resembling more typical autophagosomal vesicles was also detected (iv). See also Fig. S4.
SNX18 tubules are positive for TfR. HeLa GFP-LC3 cells were transfected with myc-SNX18 or a control myc plasmid and then immunostained for myc and TfR (A) or Rab11 (B). Bars: (A) 10 µm; (B) 5 µm. Insets show enlarged views of the boxed regions. (C, i) Clusters of tubulo-vesicular structures together with small multivesicular body–like vesicles were observed upon plastic embedding of HeLa GFP-LC3 cells overexpressing myc-SNX18. (ii–vi) Immuno-EM showed that similar clusters labeled strongly for GFP-LC3 (ii, and enlarged in iii from the boxed region), myc-SNX18 (v), and TfR (vi). Strong GFP-LC3 labeling in structures resembling more typical autophagosomal vesicles was also detected (iv). See also Fig. S4.
Recycling endosomes provide membranes for SNX18-mediated GFP-LC3 membrane trafficking and remodeling. (A) HeLa cells were starved or not starved for 2 h before fixation and immunostaining against endogenous SNX18, Rab11, and TfR. (B) Cells were transfected to express flag-Atg16L1, starved, and immunostained against Rab11 or TfR and flag. (C) Cells were starved and immunostained against TfR, Atg16L1, and SNX18. (D) Cells were transfected with control or SNX18 siRNA, starved, and immunostained against SNX18 and Rab11. Inset panels show enlarged views of the boxed regions. (E) Cells were transfected with control or Rab11 a+b siRNA, starved, and immunostained against SNX18 and Atg16L1. Cells were imaged by confocal microscopy. Bars, 5 µm.
Recycling endosomes provide membranes for SNX18-mediated GFP-LC3 membrane trafficking and remodeling. (A) HeLa cells were starved or not starved for 2 h before fixation and immunostaining against endogenous SNX18, Rab11, and TfR. (B) Cells were transfected to express flag-Atg16L1, starved, and immunostained against Rab11 or TfR and flag. (C) Cells were starved and immunostained against TfR, Atg16L1, and SNX18. (D) Cells were transfected with control or SNX18 siRNA, starved, and immunostained against SNX18 and Rab11. Inset panels show enlarged views of the boxed regions. (E) Cells were transfected with control or Rab11 a+b siRNA, starved, and immunostained against SNX18 and Atg16L1. Cells were imaged by confocal microscopy. Bars, 5 µm.
Interestingly, Atg16L1 colocalized extensively with Rab11 and TfR, both in peripheral spots (Fig. 9 B) and in the SNX18-positive perinuclear area (Fig. 9 C). To delineate the mechanistic relationship among these proteins, we depleted each one and observed the localization of the others. Depletion of SNX18 inhibited the perinuclear localization of Rab11 during starvation (Fig. 9 D) and depletion of Rab11 inhibited the perinuclear localization of Atg16L1, but not SNX18 (Fig. 9 E). We previously showed that SNX18 depletion prevents the perinuclear localization of Atg16L1 (Fig. 3 F) and that Atg16L1 depletion prevents the SNX18-induced perinuclear localization of GFP-LC3 (Fig. 5 F). Collectively, our results indicate that upon induction of autophagy, Atg16L1-positive recycling endosomes redistribute in a Rab11- and SNX18-dependent manner to a perinuclear SNX18-positive region where Atg16L1 facilitates recruitment of LC3, followed by SNX18-mediated membrane remodeling that provides membranes for autophagosome biogenesis (Fig. 10 A).
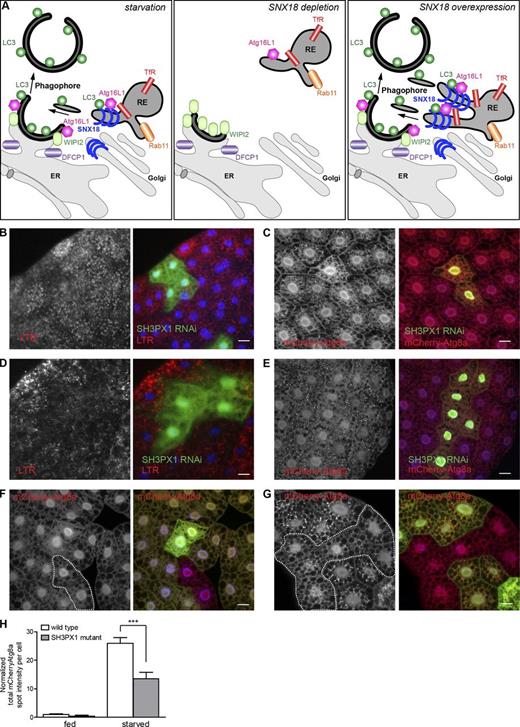
SNX18 is a positive regulator of autophagy and its function is conserved in D. melanogaster fat body cells. (A) During starvation, Rab11-positive recycling endosomes are recruited to the SNX18-positive perinuclear area dependent on SNX18 itself. SNX18 and Rab11 are required for the perinuclear localization of Atg16L1 to omegasomes marked by DFCP1. LC3 lipidation and autophagosome formation proceeds. In the absence of SNX18, recruitment of recycling endosomes to the perinuclear site is inhibited, Atg16L1 fails to localize to DFCP1-positive omegasomes, and WIPI2 accumulates. LC3 lipidation and autophagosome formation is inhibited. Upon SNX18 overexpression, Rab11-, TfR-, and LC3-positive membranes accumulate at a perinuclear site, and extensive membrane tubules with SNX18, Atg16L1, LC3, and TfR form from perinuclear recycling endosomes. LC3 lipidation and autophagosome formation is increased. Based on the experimental observations summarized in this model, we propose that SNX18 facilitates LC3 lipidation via Atg16L1 and provides membrane from recycling endosomes to forming autophagosomes. (B and C) RNAi against SH3PX1 (GFP-positive clones) decreases formation of LTR punctae in response to 4 h of starvation in 20% sucrose (B) and inhibits formation of mCherry-Atg8a-positive autophagosomes in response to 2 h starvation (C). (D and E) RNAi against SH3PX1 inhibits formation of LTR (D) and mCherry-Atg8a–positive structures (E) at early stages of developmental autophagy in fat bodies of 110-h-old larvae. (F) There was no change in mCherry-Atg8a in SH3PX1 mutant clones (GFP negative, outlined) in well-fed larvae. (G) Decreased formation of mCherry-Atg8a–positive autophagosomes in SH3PX1 mutant clones (GFP negative, outlined) in fat bodies from larvae starved in 20% sucrose for 3 h. Bars, 20 µm. (H) The graph shows quantification of total mCherry-Atg8a spot intensity per cell for WT and SH3PX1 mutant cells. The graph shows mean ± SEM (error bars); ***, P < 0.001. Genotypes: (B and D) hs-flp; UAS-dicer/+; Act>CD2>GAL4 UAS-GFPnls/UAS-SH3PX1 RNAi; (C and E) hs-flp; UAS-dicer/+; r4:mCherry-Atg8a Act>CD2>GAL4 UAS-GFPnls/UAS-SH3PX1 RNAi; (F and G) hs-flp; Cg-GAL4 UAS-mChAtg8a/+; FRT80B UAS-2XeGFP/FRT80B SH3PX1HK62b. See also Fig. S5.
SNX18 is a positive regulator of autophagy and its function is conserved in D. melanogaster fat body cells. (A) During starvation, Rab11-positive recycling endosomes are recruited to the SNX18-positive perinuclear area dependent on SNX18 itself. SNX18 and Rab11 are required for the perinuclear localization of Atg16L1 to omegasomes marked by DFCP1. LC3 lipidation and autophagosome formation proceeds. In the absence of SNX18, recruitment of recycling endosomes to the perinuclear site is inhibited, Atg16L1 fails to localize to DFCP1-positive omegasomes, and WIPI2 accumulates. LC3 lipidation and autophagosome formation is inhibited. Upon SNX18 overexpression, Rab11-, TfR-, and LC3-positive membranes accumulate at a perinuclear site, and extensive membrane tubules with SNX18, Atg16L1, LC3, and TfR form from perinuclear recycling endosomes. LC3 lipidation and autophagosome formation is increased. Based on the experimental observations summarized in this model, we propose that SNX18 facilitates LC3 lipidation via Atg16L1 and provides membrane from recycling endosomes to forming autophagosomes. (B and C) RNAi against SH3PX1 (GFP-positive clones) decreases formation of LTR punctae in response to 4 h of starvation in 20% sucrose (B) and inhibits formation of mCherry-Atg8a-positive autophagosomes in response to 2 h starvation (C). (D and E) RNAi against SH3PX1 inhibits formation of LTR (D) and mCherry-Atg8a–positive structures (E) at early stages of developmental autophagy in fat bodies of 110-h-old larvae. (F) There was no change in mCherry-Atg8a in SH3PX1 mutant clones (GFP negative, outlined) in well-fed larvae. (G) Decreased formation of mCherry-Atg8a–positive autophagosomes in SH3PX1 mutant clones (GFP negative, outlined) in fat bodies from larvae starved in 20% sucrose for 3 h. Bars, 20 µm. (H) The graph shows quantification of total mCherry-Atg8a spot intensity per cell for WT and SH3PX1 mutant cells. The graph shows mean ± SEM (error bars); ***, P < 0.001. Genotypes: (B and D) hs-flp; UAS-dicer/+; Act>CD2>GAL4 UAS-GFPnls/UAS-SH3PX1 RNAi; (C and E) hs-flp; UAS-dicer/+; r4:mCherry-Atg8a Act>CD2>GAL4 UAS-GFPnls/UAS-SH3PX1 RNAi; (F and G) hs-flp; Cg-GAL4 UAS-mChAtg8a/+; FRT80B UAS-2XeGFP/FRT80B SH3PX1HK62b. See also Fig. S5.
The D. melanogaster SNX18 homologue SH3PX1 is also required for autophagy
Whereas the vertebrate SNX9 family consists of three proteins, insects and nematodes have only one. To test if the D. melanogaster SNX18 homologue SH3PX1 is also involved in autophagy, we used mosaic analysis of SH3PX1 RNAi cells (GFP positive) surrounded by WT cells (GFP negative) in the larval fat body, where autophagy is robustly induced in response to nutrient limitation or at late larval stages as part of the developmental program (Rusten et al., 2004; Scott et al., 2004; Neufeld, 2008). Immunostaining of SH3PX1 confirmed that SH3PX1 levels were decreased by SH3PX1 RNAi (Fig. S5 A). Autophagic compartments were virtually absent from well-fed third instar larvae fat body cells as visualized by LysoTracker red (LTR; not depicted) or mCherry-Atg8a (Fig. S5 B); they were, however, readily observed in WT fat body cells, but not in cells depleted of SH3PX1 after larval starvation (Fig. 10, B and C). After longer starvation, fewer and smaller autophagosomes were observed in SH3PX1-depleted cells compared with the larger autophagosomes in WT cells (Fig. S5 C). Moreover, LTR (Fig. 10 D)- or mCherry-Atg8a (Fig. 10 E)–positive compartments were easily detected in WT fat body cells from 110-h-old well-fed larvae undergoing developmental autophagy, whereas very few were visible in the SH3PX1 RNAi–expressing cells. These compartments were larger in the WT fat body cells from older larvae (113 h), and were fewer and smaller in SH3PX1 RNAi cells (Fig. S5, D and E). Collectively, this indicates that the D. melanogaster SNX18 homologue SH3PX1 is also required for efficient autophagosome formation, both during starvation-induced and developmental autophagy.
To verify the results obtained with the SH3PX1 RNAi line, we generated a fly strain lacking the first exon and start codon of SH3PX1 (SH3PX1HK62b), which expressed no detectable SH3PX1 protein (Fig. S5, I–K) and was viable, which is consistent with the viability of other Atg mutants (Scott et al., 2004; Juhász et al., 2007). No autophagosomes were observed in fat bodies from well-fed 96-h-old larvae (Fig. 10, F and H), whereas mCherry-Atg8a–positive structures were abundant in cells heterozygous for the SH3PX1 deletion (GFP positive) after starvation, but fewer and smaller in cells homozygous for the SH3PX1 deletion (GFP negative; Fig. 10, G and H), which supports the finding that SH3PX1 is necessary for normal autophagosome formation.
Finally, although SH3PX1 was efficiently overexpressed (Fig. S5 F), and able to interact with DmAtg8a (Fig. S5 L), autophagosome formation was not induced by overexpression of SH3PX1 in the fat body of well-fed third instar larvae (Fig. S5 G), nor did SH3PX1 overexpression affect the formation of autophagosomes during starvation (Fig. S5 H). Collectively, these results show that SH3PX1 is necessary, but not sufficient, for normal formation of autophagosomes in vivo.
Discussion
We identified SNX18 as a PX domain protein involved in autophagy by a siRNA-mediated screen and validated SNX18 as a positive regulator of autophagy dependent on its membrane remodeling activity and binding to PI(4,5)P2-containing membranes. SNX18 interacts with Atg16L1 and is required for recruitment of Atg16L1-positive recycling endosomes to a perinuclear region in a Rab11-dependent manner. Importantly, SNX18 interacts directly with LC3 family proteins, and its overexpression was shown to induce perinuclear recruitment of GFP-LC3 in an Atg16L1-dependent manner. The membrane binding and tubulation activity of SNX18, as well as its binding to LC3, is required for formation of GFP-LC3–, Atg16L1-, and TfR-positive tubules that we propose contribute to the formation of transport vesicles that provide membrane to the forming autophagosome (Fig. 10 A).
Because autophagy depends on the generation of PI3P, it was surprising to find a protein with specificity for PI(4,5)P2 (Håberg et al., 2008) as a prime candidate in a screen for autophagy-related PX-domain proteins. Indeed, the observed inhibitory effect on SNX18-induced GFP-LC3 spot formation by rapalog-induced recruitment of a phosphoinositide 5-phosphatase to the plasma membrane (known to deplete PI(4,5)P2), as well as the use of PX domain mutants, indicate that binding of SNX18 to PI(4,5)P2-containing membranes is essential for its function in autophagy. PI(4,5)P2 has previously been shown to be important for autophagy through the formation of Atg16L1-positive autophagosome precursors from the plasma membrane (Ravikumar et al., 2010; Moreau et al., 2012). Interestingly, we find that Atg16L1 extensively colocalizes with markers of recycling endosomes (Rab11 and TfR) and localizes to an SNX18-positive perinuclear area in a Rab11- and SNX18-dependent manner, and that Atg16L1 and SNX18 PX-BAR interact, which suggests that the SNX18-dependent perinuclear recruitment of Atg16L1-positive recycling endosomes could depend on SNX18 binding to PI(4,5)P2.
We found membrane tubulation by SNX18 to be regulated by phosphorylation at S233, a residue located within the proposed amphipathic helix 0 just upstream of the PX domain. Interestingly, increased phosphorylation of this residue was previously reported upon rapamycin treatment (Chen et al., 2009), and we found a similar increase upon starvation. However, S233 mutations inhibited formation of GFP-LC3 spots, which suggests that SNX18 S233 phosphorylation negatively regulates its function in autophagy. It is possible that the membrane tubulation activity of SNX18 is regulated through cycles of phosphorylation and dephosphorylation to prevent excessive delivery of membranes to forming autophagosomes, and we speculate that S233 dephosphorylation might stimulate dissociation of SNX18 from the forming LC3-positive tubules.
Interaction partners of LC3 family members have been found to bind through a conserved X−3X−2X−1W0X1X2LX motif, requiring an aromatic residue in the W and a hydrophobic residue in the L position, and with acidic amino acids often found in the X positions (Noda et al., 2008; Johansen and Lamark, 2011). The SNX18 WDDEW motif found to interact with LC3/GABARAP is a noncanonical motif because of the presence of the W and several acidic residues, and the lack of the hydrophobic residue in the L position. Interestingly, this motif is also required for the SNX18 interaction with AP-1, an adaptor protein implicated in autophagosome biogenesis from the trans-Golgi network (Guo et al., 2012). The dynamic interplay of the SNX18 interaction with LC3/GABARAP and AP-1 in relation to autophagy remains to be studied. Furthermore, our results suggest that SNX18 interacts mainly with the nonlipidated LC3-I. Although LC3-I is generally considered to be cytosolic, we clearly show that SNX18 is found in membrane fractions containing both LC3-I and –II, as well as Atg16L. Membrane-associated LC3-I has previously been largely ignored, but can be detected if physiological buffers are used in cell fractionations (see, e.g., Polson et al., 2010).
SNX18 affects the localization of Atg16L1, which can stimulate LC3 lipidation (Hanada et al., 2007; Fujita et al., 2008). We propose that the SNX18 interaction with Atg16L1 and LC3 facilitates LC3 lipidation on membranes to allow their subsequent fusion with the growing phagophore. Because Rab11 relocalized to the perinuclear region during starvation in a SNX18-dependent manner and SNX18-induced membrane tubules were positive for TfR, we propose that SNX18 provides membrane input from recycling endosomes to forming autophagosomes (Fig. 10 A). This is in line with recent findings showing that vesicular transport from recycling endosomes contributes to starvation-induced autophagy (Longatti et al., 2012). Overexpression of the Rab11-binding protein TBC1D14 was found to cause tubulation of recycling endosomes and inhibit autophagosome formation (Longatti et al., 2012). Furthermore, it was shown that TBC1D14 relocates to the Golgi upon starvation and that Rab11 is required for formation of autophagosomes. These results are complementary to ours and indicate that tubulation of recycling endosome by TBC1D14 and SNX18 has opposite effects on autophagosome formation.
Materials and methods
Antibodies
The antibodies were from the following providers: rabbit anti-Atg16L1 (for Western blotting), rabbit anti-LC3 and rabbit anti-myc (Cell Signaling Technology), rabbit anti-Atg16L1 (for immunofluorescence), rabbit anti-Atg14L, rabbit anti-GABARAP and rabbit anti-LC3 (MBL), mouse anti–γ-adaptin (AP-1), mouse anti-flag M2, mouse anti–β-actin and mouse anti-GFP (Sigma-Aldrich), guinea pig anti-p62 (Progen), mouse anti-p62 (BD), mouse anti-tubulin, rabbit anti-GM130 (Abcam), rabbit anti-Rab11 (Invitrogen), mouse anti-TfR (Boehringer Mannheim clone B3/25), mouse anti-GM130 (BD), anti-calnexin (Santa Cruz Biotechnology, Inc.), mouse anti-LAMP1 (Developmental Studies Hybridoma Bank), rabbit anti-Atg16L1 (for Western blotting; Cosmo Bio Co.), HRP- or Cy2/3/5-conjugated secondary antibodies (Jackson ImmunoResearch Laboratories), and Alexa Fluor–conjugated secondary antibodies (Invitrogen). The rabbit anti-SH3PX1 antibody (Worby et al., 2001) was a gift from J.E. Dickson (University of Michigan Medical School, Ann Arbor, MI). The mouse anti-WIPI2 antibody was a gift from S. Tooze (Cancer Research UK, London, England, UK). Mouse monoclonal anti–lyso-bisphosphatidic acid (LBPA) was provided by J. Gruenberg (University of Geneva, Geneva, Switzerland) and human anti-EEA1 antiserum was from B.-H. Toh (Monash University, Melbourne, Australia). Affinity-purified rabbit antibodies against SNX9 (Lundmark and Carlsson, 2003) and SNX18 (Håberg et al., 2008) were produced by immunizing with recombinant full-length proteins fused to GST, and subsequently affinity purified using the respective antigen without GST. Chicken anti-SNX18 (used for immunofluorescence) was produced by immunizing with recombinant SH3 domain of SNX18, and affinity purified from egg yolk using the same antigen. In some experiments, rabbit anti-SNX18 from a commercial source was also used (Prestige Antibodies; Sigma-Aldrich). The phosphospecific SNX18 pS233 antibody was raised in rabbits against a phospho-peptide corresponding to the helix 0 sequence in SNX18 (CRNLNRFpSTFVKSGG; pS233; Genscript USA Inc.).
Constructs
See Table S3.
Cell lines, media, and inhibitors
Cells were maintained in DMEM (Gibco) supplemented with 10% fetal bovine serum, 5 U/ml penicillin, and 50 µg/ml streptomycin. The HEK 293A GFP-LC3 cell line (Chan et al., 2007) was a gift from S. Tooze. The HEK GFP-DFCP1 cell line (Axe et al., 2008) was a gift from N. Ktistakis (Babraham Institute, Cambridge, England, UK). HeLa cells inducibly expressing GFP, GFP-SNX18, GFP-LC3, or GFP-GABARAP (see Fig. S4 A) were made with the TRex FlpIN system (Invitrogen) with the TRex HeLa FlpIn cell line (Tighe et al., 2008), which was obtained as a gift from A. Thige and S.S. Taylor (University of Manchester, Manchester, England, UK). Expression was induced by addition of 10–500 ng/ml Tetracycline or Doxycycline (Sigma-Aldrich). Mouse embryonic fibroblasts (MEFs) were a gift from M. Komatsu (Tokyo Metropolitan Institute of Medical Science, Tokyo, Japan). Glass support was coated by 20 µg/ml fibronectin (Sigma-Aldrich) before plating HEK cell lines. For starvation in nutrient-deplete medium, the cells were incubated in Earls Balanced Salt Solution (EBSS; Invitrogen), with the exception of the HEK GFP-DFCP1 cells, which were starved as described previously (Axe et al., 2008) in 140 mM NaCl, 1 mM CaCl2, 1 mM MgCl2, 5 mM glucose, and 20 mM Hepes, pH 7.4. BafA1 (AH Diagnostics) was used at 100–200 nM and ionomycin at 5 µM (Sigma-Aldrich).
Transfection of plasmids or siRNA oligonucleotides and Western blotting
Plasmids were transfected using FuGene (Roche) or Lipofectamine 2000 (Invitrogen). The siRNAs used in the primary screen were Dharmacon SMARTpool ON-TARGET plus (Thermo Fischer Scientific). In the secondary screen, the corresponding individual Dharmacon ON-TARGET plus oligonucleotides were used. See Tables S1 and S2 for Dharmacon siRNA numbers and siRNA sequences. siSNX18-1, -2, -3, and -4 correspond to Dharmacon J-013438-09, -10, -11, and -12. The following siRNA oligonucleotides were obtained from Invitrogen (Stealth): siSNX18-5, 5′-CAGGAUCGCUGUAACACUAUUUCUU-3′; siSNX9-1, 5′-AAGAGAGUCAGCAAUCAUGUCU-3′; and siSNX9-2, 5′-AACCUACUAACACUAAUCGAU-3′. Other siRNA sequences used were from Thermo Fisher Scientific: siAtg14L, 5′-GCAAAUCUUCGACGAUCCCAUAUUU-3′; siAtg7, 5′-GCCCACAGAUGGAGUAGCA-3′; siULK1, 5′-UCACUGACCUGCUCCUUAA-3′; siAtg16L1-1, 5′-UGUGGAUGAUUAUCGAUUA-3′; siAtg16L1-2, 5′-GUUAUUGAUCUCCGAACAA-3′; siRab11a, 5′-GUAGGUGCCUUAUUGGUUU-3′; and siRab11b, 5′-CAAGAGCGAUAUCGAGCUA-3′. SMARTpools (Thermo Fisher Scientific) were used against: siLC3A, M-013579-00; siLC3B, M-012846-01; siGAB, M-012368-01; and siGAB L1, M-014715-01. The siRNA was delivered to cells by Lipofectamine RNAi max (Invitrogen). For rescue experiments, siRNA-resistant plasmids were delivered to cells by transfection with Extremegene (Roche) the day after siRNA transfection. For SDS-PAGE, the cells were lysed in 25 mM Hepes, pH 7.5, 125 mM K-Acetate, 2.5 mM Mg-Acetate, 5 mM EGTA, 1 mM DTT, and 0.5% NP-40 supplemented with Complete protease inhibitor (Roche). Protein concentration was measured by a protein assay (Bio-Rad Laboratories) to run equal amounts of cell lysate on SDS-PAGE, followed by Western blotting using specified primary antibodies and HRP-conjugated secondary antibodies detected by the Supersignal West Dura Extended Duration Substrate kit (Pierce). Imaging and quantification of protein levels were performed using the Syngene gel documentation unit, GeneSnap acquisition, and GeneTools analysis software (all from Syngene). In some cases, Western blotting was performed with far red fluorophore-conjugated secondary antibodies, and detection and analysis were performed with the Odyssey imaging system (LI-COR).
High-content immunofluorescence microscopy and confocal microscopy
For the screen, siRNA-treated HEK GFP-LC3 cells growing in glass-bottom 96-well plates were starved or not starved for 2 h and pre-permeabilized on ice with 0.05% saponin in 80 mM K-Pipes, pH 6.8, 5 mM EGTA, and 1 mM MgCl2 before fixation in 3% PFA. The nuclei were counterstained with 1 µg/ml Hoechst in PBS. The number of GFP-LC3 spots was quantified from a total of ∼30,000 cells from three independent experiments in triplicate using the automated ScanR microscope (Olympus) equipped with a ULS-Apochromat 40× objective lens (NA 0.95) and the corresponding analysis program. For other experiments, cells were grown on glass coverslips and after the described treatments, fixed in 3% PFA for 15 min on ice or in methanol for 10 min at −20°C before immunostaining with indicated antibodies and mounting in Mowiol or Prolong Gold mounting media (Life technologies) containing 1 µg/ml Hoechst or DAPI. The cells were imaged on an automated Cell Observer equipped with a 40× EC Plan Neofluar objective (NA 0.75) and a camera (AxioCam MRm) using Axiovision software (all from Carl Zeiss). The number of GFP-DFCP1, WIPI2, Atg16L1, or GFP-LC3 spots was quantified from ∼300 cells using the Physiology module of the Assaybuilder software (Carl Zeiss). Confocal images were acquired with a confocal microscope (LSM780; Carl Zeiss) equipped with an Ar laser multiline (458/488/514 nm), a DPSS-561 10 (561 nm), a laser diode 405-30 CW (405 nm), and a HeNe laser (633 nm). The objective used was a Plan-Apochromat 63×/1.4 NA oil DIC III objective lens (Carl Zeiss). Image processing was performed with basic software (zen 2010; Carl Zeiss) and Photoshop CS4 (Adobe). Some confocal images were acquired on a laser scanning confocal microscope (FluoView 1000; Olympus) based on an inverted microscope (IX81; Olympus) fitted with a Super Apochromat 60×/1.35 NA oil objective lens and the following laser lines: UV laser diode (405 nm, 6 mW), multiline Ar laser (457 nm, 488 nm, 515 nm, and 30 mW), green helium-neon laser (561 nm, 10 mW), and a red helium-neon laser (633 nm, 10 mW). Images were acquired using the dedicated FV1000 software (Olympus).
EM
Cells for conventional plastic embedding were fixed with 2% glutaraldehyde in 0.1 M Cacodylate buffer, followed by postfixation in 2% OsO4 and 1.5% KFeCN, dehydration in graded ethanol series, and epon embedding. Ultrathin sections were poststained with Uranyl acetate followed by Pb-citrate. Samples for immuno-EM were fixed in 4% formaldehyde and 0.1% glutaraldehyde, scraped and embedded in 10% gelatin, infused with 2.3 M sucrose, and mounted and frozen in liquid nitrogen. Sectioning was performed at −110°C, and 70–100 nm sections were collected with a 50:50 mixture of 2.3 M sucrose and 2% methyl cellulose. Labeling was performed with the following antibodies: rabbit anti-GFP (T. Johansen, University of Tromsø, Tromsø, Norway), mouse anti-myc (Sigma-Aldrich), and mouse anti-TfR (Boehringer Mannheim). Monoclonal antibodies were followed by secondary rabbit anti–mouse (Dako) antibodies and finally protein A gold 10 nm (University Medical Center, Utrecht, Netherlands). Sections were observed in a transmission electron microscope (JEM 1230; JEOL) at 80 kV and images were recorded with a Morada camera (SIS; Olympus), using iTEM software (Olympus). Further image processing was done with Photoshop CS5 (Adobe).
Rapalog-induced depletion of PI(4,5)P2
Experiments were performed as described previously (Zoncu et al., 2007), with some modifications: In brief, HEK GFP-LC3 cells were triple transfected with plasmids encoding plasma membrane–targeted FRB domain of human mTOR (CFP-FRB), FKBP alone (mRFP-FKBP), or the phosphoinositide 5-phosphate (5pase) fused to FKBP (mRFP-FKBP-5pase; (provided by P. Di Camilli, Yale University School of Medicine, New Haven, CT; and T. Balla, National Institutes of Health, Bethesda, MD) and myc or myc-SNX18. Addition of 2.5 µM rapalog (A/C heterodimerizer; Takara Bio Inc.) results in the heterodimerization of FRB and FKBP, leading to the recruitment of 5pase to the plasma membrane and depletion of PI(4,5)P2. The cells were fixed before immunostaining against myc and analysis of GFP-LC3 spots per cell.
qPCR
siRNA-transfected cells grown in 96-well plates were frozen dry at −80°C, RNA was isolated with an RNeasy plus kit (QIAGEN), cDNA was synthesized by reverse transcription (iScript; Bio-Rad Laboratories), and qPCR was performed using SYBR green (QIAGEN) and primer sets for the described targets relative to SDHA as a housekeeping gene on a Lightcycler 480 (Roche). All primer sets were predesigned and bought from QIAGEN (Quantitect), with the exception of SNX18, where a primer set was designed separately: 5′-GGACCTATTAGCGCTGTATCAG-3′ and 5′-CACGTGTCGCCTACTCTC-3′.
In vitro translation and GST pulldown
Indicated GFP fusion proteins were in vitro translated in TNT T7–coupled reticulocyte lysate (L4610; Promega) in the presence of [35S]methionine (PerkinElmer) and precleared on glutathione-Sepharose before incubation in NETN buffer (50 mM Tris, pH 8, 100 mM NaCl, 6 mM EDTA, 6 mM EGTA, 0.5% NP-40, 1 mM DTT, and Roche Complete protease inhibitor) together with glutathione-Sepharose–bound recombinant GST-tagged Atg8 proteins expressed in and purified from Escherichia coli according to manufacturer’s instructions. The resulting pulldowns were separated by SDS-PAGE. The gels were Coomassie blue stained, and the in vitro translated copurified proteins were detected by autoradiography on a Typhoon phosphorimaging scanner (GE Healthcare). For GST pulldown from cell lysate, cells were lysed in 10 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.5 mM EDTA, 0.5% NP-40, protease inhibitor (Roche), and phosphatase inhibitor (Sigma-Aldrich), and the cell lysate was incubated with recombinant glutathione-Sepharose–bound GST proteins. The resulting pulldowns were analyzed by immunoblotting.
Long-lived protein degradation
To measure the degradation of long-lived proteins by autophagy, proteins were first labeled with 0.25 µCi/ml l-[14C]valine (PerkinElmer) for 24 h in GIBCO-RPMI 1640 medium (Invitrogen) containing 10% FBS. The cells were washed and then chased for 3 h in nonradioactive DMEM (Invitrogen) containing 10% FBS and 10 mM valine (Sigma-Aldrich) to allow degradation of short-lived proteins. The cells were washed twice with EBSS (Invitrogen), and starved or not starved for 4 h in the presence or absence of 10 mM 3-methyladenine (3MA; Sigma-Aldrich). 10% TCA was added to the cells before incubation at 4°C to precipitate radioactive proteins. Ultima Gold LSC cocktail (PerkinElmer) was added to the samples and protein degradation was determined by measuring the ratio of TCA-soluble radioactivity relative to the total radioactivity detected by a liquid scintillation analyzer (Tri-Carb 3100TR; PerkinElmer), counting 3 min per sample.
Immunoprecipitation
GFP, GFP-LC3 WT, GFP-LC3 G120A, and GFP-SNX18 were immunoprecipitated by GFP trap (ChromoTek) according to the manufacturer’s instructions. To determine the specificity of the phosphospecific SNX18 pS233 antibody, a sample of GFP-Trap–enriched GFP-SNX18 was either mock-treated or treated with Lambda phosphatase (LPP; New England Biolabs, Inc.) for 30 min at 30°C, and analyzed by immunoblotting. For immunoprecipitation of GFP-SNX18, the HeLa GFP-SNX18 cells were harvested and lysed in 1% NP-40 in PBS containing PhosSTOP (Roche) and protease inhibitors (EMD Millipore). After centrifugation at 70,000 g for 30 min, the supernatants were immunoprecipitated with GFP-Trap. For flag-Atg16L1 immunoprecipitations, HeLa cells were transfected to express flag-Atg16L1 and myc-SNX18 variants followed by solubilization of the cells, and were immunoprecipitated with anti-flag or an unrelated antibody of the same isotype.
Cell fractionation and density-gradient centrifugation
The cells were harvested by trypsinization and resuspended in 2–5 volumes of KSHM buffer (100 mM potassium acetate, 85 mM sucrose, 20 mM HEPES-KOH, pH 7.4, and 1 mM magnesium acetate) containing 1 mM PMSF and protease inhibitor cocktail (EMD Millipore) before a quick freeze/thaw cycle to open up the plasma membrane (Lundmark and Carlsson, 2003). Cells were centrifuged at 1,500 g for 5 min, and the supernatant was centrifuged at 70,000 g for 30 min, then collected (cytosol). The pellets from the high- and low-speed centrifugations were combined and solubilized in KSHM containing 1% NP-40 and protease inhibitors, and centrifuged at 70,000 g for 30 min to obtain a supernatant (membranes). Equal proportions of cytosol and membrane were analyzed by immunoblotting.
For density gradient separation of total membranes, HeLa cells grown in full medium or in nutrient-deficient medium in the presence of BafA1 for 2 h were trypsinized and washed in PBS. After resuspension in 10 vol of 15 mM Hepes-KOH, pH 7.4, 1 mM EDTA, and 0.25 M sucrose (HES buffer) containing protease and phosphatase inhibitors, the cells were gently homogenized by passage through a ball-bearing device (Lundmark and Carlsson, 2002). Nuclei were pelleted by centrifugation at 800 g for 10 min, and organelles and membranes in the supernatant were collected by centrifugation at 70,000 g for 30 min. The organelle/membrane pellet was resuspended in 200 µl HES buffer and layered on top of a 4-ml 0–50% continuous Nycodenz gradient prepared in HES buffer. Centrifugation was performed at 70,000 g for 16 h after which 250-µl fractions were collected from the top. Equal volumes of each fraction were analyzed by SDS-PAGE followed by immunoblotting.
Peptide array
Peptide arrays were synthesized at the Biotechnology Centre of Oslo, University of Oslo, on cellulose membranes using a MultiPep automated synthesizer (INTAVIS Bioanalytical Instruments AB) as described previously (Frank, 1992).
Drosophila genetics
The SH3PX1EY08084 and SH3PX1 RNAi (TRiP.JF02730) lines were obtained from the Bloomington Drosophila Stock Center at Indiana University. Clonal analysis was performed using the methods and fly stocks described in Arsham and Neufeld (2009). In brief, SH3PX1 was depleted in fat body clones by crossing SH3PX1JF02730 to y,w,hs-flp; UAS-dicer; act>CD2>GAL4, UAS-GFPnls/TM6B, Hu, Tb or y,w,hs-flp; UAS-dicer; r4:mCherryAtg8a, act>CD2>GAL4, UAS-GFPnls/TM6B, Hu, Tb. Clones of fat body cells overexpressing SH3PX1 were generated in crosses of SH3PX1EY08084 to y,w,hs-flp; r4:mCherryAtg8a, act>CD2>GAL4, UAS-GFPnls/TM6B, Hu, Tb or y,w,hs-flp; act>CD2>GAL4, UAS-GFPnls/TM6B, Hu, Tb. Clones of homozygous SH3PX1 mutant cells were generated from the cross SH3PX1HK62b FRT80B/TM6B, Hu, Tb χ y,w,hs-flp; Cg-GAL4, UAS-mChAtg8a; UAS-2XeGFP, FRT80B. The resulting progeny were subjected to a 1-h heat shock (37°C) at 0–6 h after egg deposition.
Fat body autophagy assays
72-h-old larvae were moved to fresh food with yeast paste. 24 h later, the larvae were starved or not starved for the indicated times in 20% sucrose. The larvae were bisected, inverted, and fixed in 3.7% formaldehyde. After washing in PBS with 0.1% Triton X-100 (PBS-T) and counterstaining with 1 µg/ml DAPI, the fat bodies were dissected out and mounted in Fluoroguard. Fat bodies for antibody staining were blocked in 5% goat serum in PBS-T before staining with the indicated antibodies. For LTR (Invitrogen) staining, freshly dissected fat bodies were incubated in 100 nM LTR and 1 µg/ml DAPI for 3 min and imaged immediately. All larval samples were imaged on a confocal microscope equipped (LSM510) with a 40× objective lens (both from Carl Zeiss). Total mCherryAtg8a spot intensity was quantified by the Carl Zeiss AssayBuilder Physiology module.
Generation of SH3PX1 deletion mutant
The P element SH3PX1EY08084 located in the 5′ UTR of the SH3PX1 gene was remobilized, and lines where part of the gene region was deleted were screened by PCR using primers 5′-ACGAAAACAAAGCGCAGCGC-3′ and 5′-ACGCCTCGCATTTGGTGCAA-3′. One such line missing 1,198 bp of the gene region was identified (SH3PX1HK62b), and sequencing showed that the first exon of SH3PX1 including the start codon was deleted (1,198 nucleotides removed from the EY08084 insertion site (3L:9,706,877; FlyBase release FB2012_02).
Lysate of larval fat bodies
Fat bodies were dissected out of 96-h-old larvae and immediately boiled for 5 min at 95°C in SDS sample buffer before SDS-PAGE and immunoblotting.
Statistics
For the primary siRNA screen, data were collected from three experiments with three repetitions each. In a quality control process, plates were inspected with respect to the number of cells in the wells and the consistency of results in wells with equal siRNA. Based on this, some plates were excluded, leaving 6–8 plates of each type. The estimated GFP-LC3 count for each siRNA was then computed as a weighted mean over all experiments and repetitions to eliminate systematic differences in the level of GFP-LC3 spots between plates. The weights were adjusted in such a way that plates with equal siRNA layout had an equal mean count over all wells, and plates with different layout had equal mean count over the control wells. A similar procedure was used for the secondary screen. The P-values of other analysis were derived from statistical tests using a two-sided t test from Excel (Microsoft), and considered statistically significant at P < 0.05.
Online supplemental material
Fig. S1 shows how the Olympus ScanR software detects the GFP-LC3 spots as well as images showing GFP-LC3 spots for all SNX18 siRNAs and the corresponding relative expression of SNX18 and LC3 mRNA upon SNX18 knockdown. Fig. S2 shows the effect of siSNX18 compared with siSNX9 and includes controls relating to membrane fractionation and induction of GFP-LC3 spots. Fig. S3 complements the characterization of endogenous SNX18 localization and provides further controls for the perinuclear recruitment of Atg16L1 and induction of GFP-LC3 spots. Fig. S4 is a characterization of the TRex FlpIN HeLa GFP-LC3 and -GABARAP cells including data on myc-SNX18 and endocytic markers. Fig. S5 shows that SH3PX1 RNAi results in formation of smaller mCherry-Atg8a–marked autophagosomes, whereas SH3PX1 overexpression does not induce or change formation of mCherry-positive autophagosomes in larval fat bodies. It also describes how the SH3PX1 deletion mutant was generated by remobilization of the P element SH3PX1EY08084 located in the 5′ UTR of the SH3PX1 gene, and that SH3PX1 binds to DmAtg8a. Table S1 is a comprehensive list of the gene targets, siRNA numbers, number of cells analyzed, and GFP-LC3 spots per cell from the primary screen. Table S2 lists the gene targets, siRNA numbers and sequences, percent knockdown of mRNA for the specified target, number of cells analyzed, and GFP-LC3 spots per cell from the secondary screen. Table S3 lists the plasmids used in this study.
Acknowledgments
We thank S. Tooze, N. Ktistakis, I. Dikic, P. de Camilli, T. Balla, T. Johansen, M. Komatsu, and J.E. Dickson for generously sharing reagents, A. Engen for assistance with cell culture, C. Knudsen for help with Fig. 10 A, and the confocal microscopy core facility at the Institute for Cancer Research, Oslo, for providing access to confocal microscopes. We received materials through the Drosophila Genomics Resource Center.
This work was supported by grants from the Molecular Life Science program of the University of Oslo, the Swedish Research Council, and the Kempe Foundation.
References
- 5pase
phosphoinositide 5-phosphate
- BafA1
Bafilomycin A1
- BAR
Bin/Amphiphysin/Rvs homology
- DFCP1
double FYVE containing protein 1
- EBSS
Earls Balanced Salt Solution
- LTR
LysoTracker red
- PI3K
phosphatidylinositol 3-kinase
- PI3P
phosphatidylinositol 3-phosphate
- PI(4,5)P2
phosphatidylinositol 4,5-bisphosphate
- qPCR
quantitative real-time PCR
- SNX
Sorting nexin
- TfR
Transferrin receptor
- WT
wild type
Author notes
H. Knævelsrud and K. Søreng contributed equally to this paper.


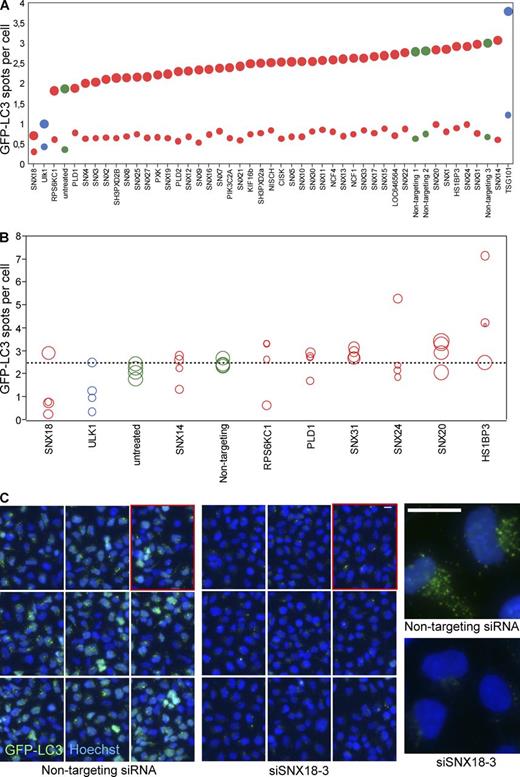
![Figure 2. SNX18 is a positive regulator of autophagy. (A) HEK GFP-LC3 cells transfected with control (siCtrl) or two different SNX18 siRNA oligos (siSNX18-3 and siSNX18-5) were starved or not starved for 2 h in the presence or absence of BafA1. GFP-LC3 lipidation and SNX18 protein knockdown were monitored by immunoblotting. The graph shows the average GFP-LC3-II relative to actin normalized to siCtrl-1 fed ± SEM (error bars), n = 4. (B) The degradation of long-lived proteins in HeLa cells transfected with control or SNX18 siRNA was quantified after 4 h of starvation in the absence or presence of 3-MA and normalized to the degradation in fed cells (mean ± SEM [error bars], n = 3). (C) HEK GFP-LC3 cells were transfected with control or SNX18 siRNA and then with a myc control or siRNA-resistant myc-SNX18 plasmid. The number of GFP-LC3 spots per cell was quantified (graph shows mean ± SEM [error bars], n = 3). (D) HEK GFP-LC3 cells were transfected with myc-SNX18 or a myc control plasmid and treated as in A. The graph shows the average LC3-II levels relative to actin normalized to myc-transfected fed cells, ±SEM (error bars). n = 3. (E) HEK GFP-LC3 cells were transfected with control or Atg7 siRNA and then with myc-SNX18 or a myc control plasmid, followed by quantification of the number of GFP-LC3 spots per cell (graphs show mean ± SEM [error bars], n = 3). Representative images are shown. Bars, 5 µm. (F) The degradation of long-lived protein in HeLa cells transfected with myc-SNX18 or a myc control plasmid was quantified as in B (graph shows mean ± SEM [error bars], n = 4). *, P < 0.05; **, P < 0.01. See also Figs. S1 and S2.](https://cdn.rupress.org/rup/content_public/journal/jcb/202/2/10.1083_jcb.201205129/5/m_jcb_201205129_fig2.jpeg?Expires=1781754571&Signature=wna1Lw9weunW00skCfi1HGtxK~EZ190TgSewLIKx8Yy1ZypEdcQKJHJtWo5qmj5FJz9RDgFu4MPRk92hjaGcuobr9bvuwes04rh3Iexncub4KwfMK0wpHD5h4sE8xODRMRSTwBHzGcs1I76npOHRimxKjAvtO-RU6wEEDrAvP6oRMT04w~WcpnVMqBb8MIdLinX00VrGdrTJM3i8hFO0E8nWbY28mwxvYXPXzBLeo-P3dYHSDZJC2vsnBt1Q7lakZScMsJ9Wg2UU2hEjWT9UwnNOqfLnbOTDfnFJCdKlpJRbdjQsXYgU2AiQVM9AKNFVgyBmx9SZAOaEQP7ZkoQOMQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
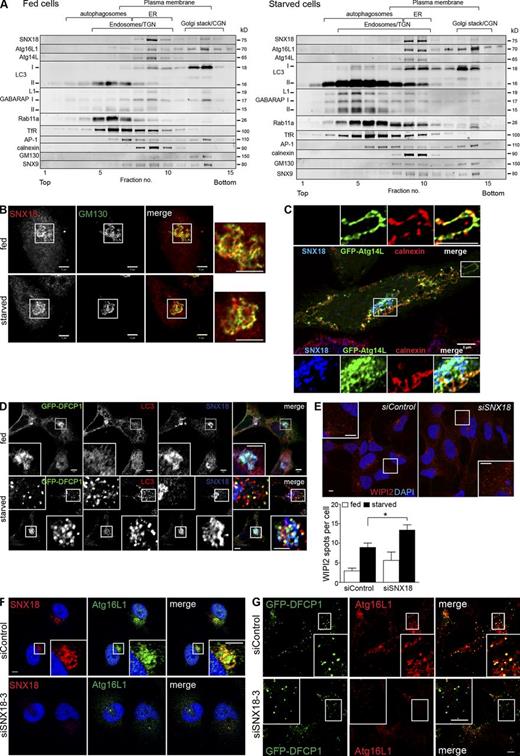
![Figure 4. SNX18 binding to PI(4,5)P2 membranes is required for its function in autophagy. (A) HeLa GFP-LC3 cells were transfected to express myc-SNX18, immunostained against myc, and analyzed by confocal imaging. Asterisks indicate a nontransfected cell. Bars: (main panels) 20 µm; (inset) 10 µm. (B) HeLa cells transfected with flag-Atg16L1 and myc-SNX18 were starved or not starved for 2 h before immunostaining with anti-myc and anti-flag antibodies and confocal microscopy. Bars, 5 µm. (C) SNX18 domain structure. An N-terminal SH3 domain is followed by a low-complexity region (LC). The membrane-remodeling part consists of an amphipathic helix (helix 0), a PX domain that is flanked by sequences forming a Yoke domain (Y), and a BAR half-domain. The functional BAR domain is formed by dimerization. Truncated constructs and point mutations used are indicated. (D) HeLa cells were transfected with the indicated myc-SNX18 plasmids for 6 h and starved for an additional 2 h before harvest. Equal parts of cytosol (C) and membrane (M) fractions were analyzed by Western blotting. The graph shows the average membrane-bound myc-SNX18 ± SEM (error bars), n = 3. (E and F) HEK GFP-LC3 cells were transfected with the indicated mCherry-SNX18 (E) or mycSNX18 (F) constructs for 16 h. The number of GFP-LC3 spots per transfected cell (E; graph shows mean ± SEM [error bars], n = 5) or ratio of LC3-II to actin determined by Western blotting (F) was quantified. (G and H) HEK GFP-LC3 cells transfected with myc-SNX18 or a myc control plasmid alone (G) or together with CFP-FRB and mRFP-FKBP or mRFP-FKBP 5pase (H) for 6 h were treated or not treated with 5 µM ionomycin (G) or 2.5 µM A/C heterodimerizer (rapalog; H) for the last 3 h before analysis of the total intensity of GFP-LC3 spots per transfected cell (graph shows mean ± SEM [error bars], n = 3). (I) MEFs were transfected with myc, myc-SNX18 WT, or KR for 16 h, starved or not starved for 2 h, and immunostained for endogenous Atg16L1. Insets show enlarged views of the boxed regions. Bars, 5 µm. (J) HeLa cells were transfected for 12 h with the indicated constructs and starved for 90 min. Cells were solubilized and immunoprecipitated with anti-flag or an unrelated antibody of the same isotype (mock ab). Immunoprecipitates and 2% of the lysates were analyzed by immunoblotting against myc and flag. *, P < 0.05; **, P < 0.01. See also Fig. S3.](https://cdn.rupress.org/rup/content_public/journal/jcb/202/2/10.1083_jcb.201205129/5/m_jcb_201205129_fig4.jpeg?Expires=1781754571&Signature=SGicSlNsvq9bp6nlAG-5FFO5~q8fETO67yhFv3VQZuIUh4GUlx5yV5J7XD7NIpinf3ecRq4wytgQEzzXm-pL6v1fRSjQlTQaMDKaBvlodoswcbCKTxVQHNUyFdZfUQLfMFwI-ek-26sHCtGMpP52Rx6xgwk8T6Nyhpvy6gKAcsBJ80UYXZwjGdE-hJ5FNdXJnK1C2axMxavUq6L6ss0ssaHUqhXrUX3TF2Ani5nOE2l~Y5aR-YqgQ-somDdntjkCJ0vmc5qDM5PPnSWUgYWLQtuwPOPjWst2Wo0gi48VfBAH2mpRfgXevxkRD2yfGkbkwuw6U9vnUihEFJqO69j9xA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![Figure 5. The tubulation activity of SNX18 is important for its role in autophagy and is regulated by phosphorylation. (A) HeLa GFP-SNX18 cells were starved for the indicated time, followed by incubation in full medium (Refeed) where indicated. Cell lysates were analyzed for LC3 by immunoblotting, or immunoprecipitated with GFP-Trap before immunoblotting against SNX18(pS233) and GFP. The ratio of pS233 to GFP, and the ratio of LC3-II to LC3-I, was quantified, and normalized to 1 for the zero-time sample in both cases (graph shows mean ± SEM [error bars], n = 4; *, P < 0.05). (B) The indicated myc-SNX18 PX-BAR constructs were transfected into HeLa cells, and their tubulation efficiency was measured by scoring the percentage of cells displaying more than three elongated structures longer than 5 µm (n = 300). The graph shows the mean of two independent experiments ± range (error bars). Bars, 10 µm. (C) The indicated myc-tagged SNX18 constructs were transfected into HEK GFP-LC3 cells, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 5. (D) HeLa GFP-LC3 cells were transfected with the indicated myc-tagged SNX18 constructs for 16 h, starved for 2 h, and analyzed by confocal imaging. Bars, 10 µm. (E) HEK GFP-LC3 cells were transfected with myc-SNX18 WT or myc-SNX18 PX-BAR constructs, and the number of GFP-LC3 spots per cell was quantified. The graph shows mean ± SEM (error bars), n = 3. (F) HeLa GFP-LC3 cells were transfected with siRNA against Atg16L1 and later with myc-SNX18 constructs before immunostaining and confocal imaging. Bar, 5 µm.](https://cdn.rupress.org/rup/content_public/journal/jcb/202/2/10.1083_jcb.201205129/5/m_jcb_201205129_fig5.jpeg?Expires=1781754571&Signature=3iMQoQm0f18-SrhWf0r9s28j6Gz2CIMxsTcLmo9pkr4eqqTzpXe1b79gsAFbSkyc4gMsCkim0Y2dLzN42gEgYfbk-gpX3z52eWhL2ZIrJnp6A7EVInZeHFPbdJ6glk2~2ecVEgFLAwn93HVo9O9hejvbIAKwKmzBctpm8VYgt-NWPMGilM3VgD26XtyjEdbeBB0TiFLHhdYGrNKNd0VfcrHKcyu307Rp3CsixrUhOt2JFKL7DhO41n4Acwu0ctpqm0oSpqJ8WbocrSS6y3KRepCv98pJdXCw1SeA~LOYeFCFmY-NwXee0xMpYFDmoHccX3ofqbbmuPji3jvyTN-4QA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)