Myo4p, one of two class V myosins in budding yeast, continuously transports messenger RNA (mRNA) cargo in the cell but is nonprocessive when characterized in vitro. The adapter protein She3p tightly binds to the Myo4p rod, forming a single-headed motor complex. In this paper, we show that two Myo4p–She3p motors are recruited by the tetrameric mRNA-binding protein She2p to form a processive double-headed complex. The binding site for She3p was mapped to a single α helix that protrudes at right angles from She2p. Processive runs of several micrometers on yeast actin–tropomyosin filaments were observed only in the presence of She2p, and, thus, motor activity is regulated by cargo binding. While moving processively, each head steps ∼72 nm in a hand-over-hand motion. Coupling two high-duty cycle monomeric motors via a common cargo-binding adapter protein creates a complex with transport properties comparable with a single dimeric processive motor such as vertebrate myosin Va.
Introduction
A feature once thought to be a hallmark of all class V myosins was their ability to move processively, that is, to take multiple steps on actin tracks without dissociating. This concept arose from the fact that myosin Va, the most well-studied class V myosin from vertebrates, steps along actin for several micrometers, a feature well suited for transporting cargo in the cell. However, several class V myosins involved in intracellular cargo transport have been characterized as nonprocessive under in vitro conditions. These include human myosin Vc (Takagi et al., 2008; Watanabe et al., 2008), Drosophila melanogaster myosin V (Tóth et al., 2005), and both class V myosins (Myo2p and Myo4p) from the budding yeast Saccharomyces cerevisiae (Reck-Peterson et al., 2001; Dunn et al., 2007; Hodges et al., 2008).
How can nonprocessive motors function as cargo transporters? Processivity is only necessary to achieve long continuous runs if a single motor is attached to its cargo. Multiple nonprocessive motors may collectively be as effective at transport as a single processive one. Alternatively, most in vitro studies have been performed using bare actin filaments in the absence of any bound cargo, and, thus, motors characterized as nonprocessive might be processive when assayed under more physiologically relevant conditions. Here, we focus on understanding potential mechanisms by which Myo4p, one of the two nonprocessive class V myosins from budding yeast, can continuously transport and asymmetrically localize >20 different mRNAs (Shepard et al., 2003; Jambhekar et al., 2005) to the bud tip via actin cables. Asymmetric localization of mRNA is a widely used mechanism to allow cells to spatially and temporally control protein function by determining their sites of synthesis.
Myo4p has some unusual features for a class V myosin. Although it has a high duty cycle motor domain and a long lever arm (Krementsova et al., 2006), it is single headed and thus cannot move processively as a single motor (Fig. 1 A; Dunn et al., 2007; Hodges et al., 2008). Instead of forming an α-helical coiled-coil homodimer, as do all other class V myosin heavy chains, the rod region of Myo4p tightly binds to the cargo adapter protein She3p. The two proteins copurify, and She3p is, in essence, a subunit of the motor complex (Figs. 1 A and S1 A; Dunn et al., 2007; Hodges et al., 2008). Based on the fact that both proteins have coiled-coil motifs, the rod of Myo4p and She3p might form a hetero–coiled-coil, which would explain the inability of Myo4p to form a homodimer. Although most class V myosins have multiple adapter proteins to allow binding to a variety of different cargoes, Myo4p uses She3p as the sole adapter protein for its two types of cargo, mRNA and cortical ER (Estrada et al., 2003; Shepard et al., 2003; Jambhekar et al., 2005; Schmid et al., 2006). Incorporating She3p as a tight-binding subunit of Myo4p thus makes biological sense.
Sedimentation velocity and crystal structure of She2p. (A) Diagram showing the domain structure of Myo4p and its association with She3p. The motor domain contains the actin and ATP-binding sites followed by the lever arm, which binds six light chains/CaM. The rod of Myo4p binds to She3p. The C terminus of the Myo4p heavy chain forms a globular tail domain. (B) Crystal structure of She2p (Protein Data Bank accession no. 1XLY). The two chains of the dimer are shown in green and cyan. The location of the two point mutants, S120Y and L130Y, are shown in space-filling representation. S120Y is at the interface of the crystallographic dimer, whereas L130Y is at the proposed interface for tetramer formation (Müller et al., 2009). The four differences between WT and WT* are shown in yellow space-filling representation. The Cys to Ser mutations were made to prevent formation of higher oligomers of She2p (see C). A single α helix that protrudes from the side of She2p (red) is identified as all or part of the binding site for She3p. The mRNA-binding site is indicated in tan. (C) WT She2p exhibited a polydisperse profile when analyzed by sedimentation velocity in the analytical ultracentrifuge. In contrast, when four Cys residues were mutated to Ser (C14S/C68S/C106S/C180S; yellow spheres in B), the protein showed a single homogeneous peak that sedimented at 5.4 S. This construct, called WT*, was used as the backbone for further mutants described in this paper. Protein concentration was 8 µM She2p tetramer. A representative centrifuge run is illustrated (n = 3). OD, optical density.
Sedimentation velocity and crystal structure of She2p. (A) Diagram showing the domain structure of Myo4p and its association with She3p. The motor domain contains the actin and ATP-binding sites followed by the lever arm, which binds six light chains/CaM. The rod of Myo4p binds to She3p. The C terminus of the Myo4p heavy chain forms a globular tail domain. (B) Crystal structure of She2p (Protein Data Bank accession no. 1XLY). The two chains of the dimer are shown in green and cyan. The location of the two point mutants, S120Y and L130Y, are shown in space-filling representation. S120Y is at the interface of the crystallographic dimer, whereas L130Y is at the proposed interface for tetramer formation (Müller et al., 2009). The four differences between WT and WT* are shown in yellow space-filling representation. The Cys to Ser mutations were made to prevent formation of higher oligomers of She2p (see C). A single α helix that protrudes from the side of She2p (red) is identified as all or part of the binding site for She3p. The mRNA-binding site is indicated in tan. (C) WT She2p exhibited a polydisperse profile when analyzed by sedimentation velocity in the analytical ultracentrifuge. In contrast, when four Cys residues were mutated to Ser (C14S/C68S/C106S/C180S; yellow spheres in B), the protein showed a single homogeneous peak that sedimented at 5.4 S. This construct, called WT*, was used as the backbone for further mutants described in this paper. Protein concentration was 8 µM She2p tetramer. A representative centrifuge run is illustrated (n = 3). OD, optical density.
Here, we begin to increase the complexity of the characterization of the Myo4p–She3p motor complex in vitro by introducing the mRNA-binding protein She2p. She2p is the middleman that binds to both the motor (Myo4p–She3p) and the cargo (mRNA). We show by EM that She2p recruits two Myo4p–She3p motors. Unlike a single Myo4p–She3p motor, this complex supports long-range continuous transport on actin. Surprisingly, the two motors linked via She2p show hand-over-hand stepping indistinguishable from vertebrate myosin Va, which is dimerized by an α-helical coiled-coil. The ability to show processive motion only when bound to the cargo adapter protein She2p provides an elegant mechanism by which this motor can be regulated. The results suggest that motors that have been characterized as nonprocessive in vitro may achieve the continuous motion expected of a cargo transporter under conditions that more closely mimic the cellular situation.
Results
She2p is a tetramer
She2p links the Myo4p–She3p motor complex to its mRNA cargo. The crystal structure of She2p showed that it was a homodimer related by a twofold axis of symmetry (Fig. 1 B; Niessing et al., 2004). The She2p monomer consists of a five–α helix bundle with a basic helical hairpin motif that binds mRNA. Analysis of She2p by sedimentation velocity in the analytical ultracentrifuge showed a pronounced tendency to form multiple higher oligomers, even in the presence of DTT, as also observed by others (Fig. 1 C; Müller et al., 2009). Because higher-order oligomers would complicate interpretation of our further studies, we adopted some of the strategy used to crystallize She2p, namely mutation of four Cys residues (14, 68, 106, and 180) to Ser (Fig. 1 B). This effectively abolished the tendency of She2p to aggregate via formation of disulfide bonds (Fig. 1 C), resulting in a homogeneous species that sedimented at 5.4 S (Table I). This homogeneous She2p construct (called wild type* [WT*]) was used for all experiments and is the backbone for subsequent mutations.
Oligomeric state of WT and mutant She2p
| She2p | Sedimentation value | MM | Oligomer |
| S | kD | ||
| WT* | 5.4 | 98.7 ± 3.9 | Tetramer |
| L130Y | 4.1 | 54.4 ± 1.0 | Dimer |
| S120Y | 4.0 | 51.7 ± 1.3 | Dimer |
| Δhelix | 5.4 | ND | Tetramer |
| She2p | Sedimentation value | MM | Oligomer |
| S | kD | ||
| WT* | 5.4 | 98.7 ± 3.9 | Tetramer |
| L130Y | 4.1 | 54.4 ± 1.0 | Dimer |
| S120Y | 4.0 | 51.7 ± 1.3 | Dimer |
| Δhelix | 5.4 | ND | Tetramer |
The calculated molecular mass (MM) of a She2p monomer is 28 kD.
The ability of She2p WT* to functionally replace native She2p in living she2Δ yeast cells was tested. The most well-studied mRNA in budding yeast is ASH1, which is moved by Myo4p to the bud tip to repress mating-type switching in the daughter cell (Haarer et al., 1994; Sil and Herskowitz, 1996; Long et al., 1997; Takizawa et al., 1997). Mutations in any of the five genes SHE1–SHE5 cause defects in ASH1 mRNA transcript localization. Actively budding yeast cells that contained one fluorescent ASH1 mRNA particle were scored according to whether the particle was localized in the bud tip (correct localization) or in the mother cell (incorrect localization; Fig. 2 A). Transformation of the she2Δ cells with WT She2p showed 97% of the particles being correctly localized to the bud tip compared with 96% for She2p WT* (Fig. 2 B). Mutation of the four Cys residues to Ser thus did not affect the cellular function of She2p.
Ability of various She2p constructs to correctly localize ASH1 mRNA particles in budding yeast. (A) Representative images of correctly and incorrectly localized ASH1 mRNA. Correctly localized mRNA is found in the bud, whereas particles that are not transported remain in the mother cell. (left) Differential interference contrast [DIC]. (middle) Epifluorescence to visualize GFP-labeled ASH1 mRNA. (right) Merged images. (B) Percentage of correctly localized ASH1 mRNA particles for various She2p constructs. The indicated constructs were transformed into a she2Δ strain. Constructs lacking She2p do not localize ASH1 mRNA. WT* is the WT construct with four Cys residues (14, 68, 106, and 180) mutated to Ser. All other constructs are based on the WT* backbone. Addition of a fluorescent protein to either the N or C terminus of She2p had no effect on cellular function. Data are shown for a C-terminal fluorescent protein, but the data are identical for YFP at the N terminus. The negative control (neg) is a plasmid lacking an insert. The number of cells counted is indicated above each bar. Data were derived from at least two independent transformations per construct.
Ability of various She2p constructs to correctly localize ASH1 mRNA particles in budding yeast. (A) Representative images of correctly and incorrectly localized ASH1 mRNA. Correctly localized mRNA is found in the bud, whereas particles that are not transported remain in the mother cell. (left) Differential interference contrast [DIC]. (middle) Epifluorescence to visualize GFP-labeled ASH1 mRNA. (right) Merged images. (B) Percentage of correctly localized ASH1 mRNA particles for various She2p constructs. The indicated constructs were transformed into a she2Δ strain. Constructs lacking She2p do not localize ASH1 mRNA. WT* is the WT construct with four Cys residues (14, 68, 106, and 180) mutated to Ser. All other constructs are based on the WT* backbone. Addition of a fluorescent protein to either the N or C terminus of She2p had no effect on cellular function. Data are shown for a C-terminal fluorescent protein, but the data are identical for YFP at the N terminus. The negative control (neg) is a plasmid lacking an insert. The number of cells counted is indicated above each bar. Data were derived from at least two independent transformations per construct.
The molecular mass of the 5.4 S She2p WT*, obtained by sedimentation equilibrium in the analytical ultracentrifuge, was 98.7 ± 3.9 kD (Table I). Given that the monomer molecular mass of She2p is 28 kD, this result is most consistent with She2p being a tetramer, which is in agreement with recent results from two other groups (Müller et al., 2009; Chung and Takizawa, 2010). The dimeric She2p structure observed by crystallography (Niessing et al., 2004) is therefore not the state of oligomerization that exists in solution.
Myo4p–She3p binds She2p in the absence of mRNA
To determine whether the Myo4p–She3p complex requires mRNA to bind to She2p, we expressed a truncated complex of Myo4p that lacked the motor domain and neck bound to She3p (Δhead-She3p). The major reason for using this construct was its homogeneity by sedimentation velocity (4.5 S), thus allowing for unambiguous detection of a potential complex with the 5.4 S She2p. When the motor construct and She2p were mixed together, a new complex at 7.5 S was observed, demonstrating that mRNA is not essential to form a ternary complex at micromolar concentration (Fig. 3). Complex formation was also demonstrated by a pull-down of She2p with the FLAG-tagged Δhead–She3p complex (Fig. S1 B). An actin-pelleting assay showed that She2p also cosedimented with full-length Myo4p–She3p (Fig. 4).
Complex formation between Myo4p–She3p and She2p in the absence of mRNA. Analytical ultracentrifuge runs of She2p (red), a headless Myo4p–She3p construct (green), and the two proteins mixed together (black). She2p sedimented at 5.4 S, the motor construct at 4.5 S, and the complex at 7.5 S. Protein concentration was 7 µM Myo4p–She3p and 7 µM tetrameric She2p. When mixed, each concentration was halved. OD, optical density.
Complex formation between Myo4p–She3p and She2p in the absence of mRNA. Analytical ultracentrifuge runs of She2p (red), a headless Myo4p–She3p construct (green), and the two proteins mixed together (black). She2p sedimented at 5.4 S, the motor construct at 4.5 S, and the complex at 7.5 S. Protein concentration was 7 µM Myo4p–She3p and 7 µM tetrameric She2p. When mixed, each concentration was halved. OD, optical density.
Actin-pelleting assays to determine which She2p constructs are able to bind to Myo4p–She3p. SDS gels showing the supernatant (S) and pellet (P) after high-speed centrifugation of actin, Myo4p–She3p, and various She2p constructs. Myo4p–She3p cosediments with actin in the absence of MgATP; thus, She2p mutants able to bind Myo4p–She3p appear in the pellet. Molecular masses of the proteins are indicated. HC, heavy chain.
Actin-pelleting assays to determine which She2p constructs are able to bind to Myo4p–She3p. SDS gels showing the supernatant (S) and pellet (P) after high-speed centrifugation of actin, Myo4p–She3p, and various She2p constructs. Myo4p–She3p cosediments with actin in the absence of MgATP; thus, She2p mutants able to bind Myo4p–She3p appear in the pellet. Molecular masses of the proteins are indicated. HC, heavy chain.
Deletion of the protruding α helix on She2p abolishes She3p binding
The binding site on She2p for She3p has not been previously mapped. The She2p structure shows a prominent α helix protruding at right angles from the middle of each monomer (Fig. 1 B). This helix, and its connection to the five–α helix bundle, is conserved when compared with nine other yeast species (Niessing et al., 2004). Based on this observation and given that single α helices are generally not stable in solution except when bound to a partner, we speculated that this helix in She2p constitutes a major part of the binding site for She3p. When residues V174–K179 of She2p were deleted (She2p-Δhelix), Myo4p–She3p did not bind, as assessed by an actin-pelleting assay (Fig. 4), nor was it pulled down with the FLAG-tagged Δhead–She3p complex (Fig. S1 B). However, deletion of these residues had no effect on the homogeneity or S value of She2p (Table I). When the She2p-Δhelix construct was transformed into she2Δ yeast cells, only 8% of the particles was correctly localized, similar to the value obtained with a negative control (Fig. 2). No mRNA localization is seen when the connection between the motor and She2p is disrupted (Bookwalter et al., 2009), and, thus, we conclude that this helix constitutes all or part of the binding site for She3p.
Point mutants of She2p disrupt tetramer formation
She2p with various point mutations has been tested for mRNA binding in vitro and mRNA localization in situ (Niessing et al., 2004). Two of these mutations also affect the oligomeric state of She2p. She2p(S120Y), located at the interface of the crystallographic dimer (Fig. 1 B), showed a reduced sedimentation coefficient compared with WT* (4.0 vs. 5.4 S) and a reduced molecular mass by sedimentation equilibrium (51.7 ± 1.3 kD), consistent with a dimer (Table I). The mutant She2p(S120Y) tended to form large aggregates that could be removed by high-speed centrifugation, as might be expected given that a dimeric interface with a large surface area was disrupted. Little or no She2p(S120Y) bound to Myo4p–She3p, as assessed by an actin-pelleting assay (Fig. 4). We also confirmed the finding that She2p(S120Y) does not support mRNA localization in vivo (Fig. 2; Niessing et al., 2004). The markedly reduced affinity of She2p(S120Y) both for mRNA (<20% of WT; Niessing et al., 2004) and for She3p accounts for its inability to localize ASH1 mRNA in the cell.
A second mutation, L130Y, located at the conserved upper region of She2p (Fig. 1 B), is at the interface of the proposed elongated tetramer (Müller et al., 2009). We confirmed that She2p(L130Y) was a dimer with a molecular mass of 54.4 ± 1.0 kD and an S value of 4.1 S (Table I). Unlike She2p(S120Y), an actin-pelleting assay showed that dimeric She2p(L130Y) binds to the Myo4p–She3p complex (Fig. 4). When tested in vivo, this mutant supported correct localization of mRNA to the same high extent as WT and WT* She2p (Fig. 2).
Two Myo4p–She3p motors bind to a She2p tetramer and form a processive complex
EM was used to determine the number of Myo4p–She3p motors that bind to a She2p tetramer. Metal-shadowed images of Myo4p–She3p alone showed an elongated structure with a globular domain at one end, which is presumably the myosin motor domain (Fig. 5). Two-headed structures were not observed. In contrast, when Myo4p–She3p was incubated with She2p, 25 ± 7% of the images (mean ± SD; n = 186; five fields) showed a V-shaped image, consistent with two Myo4p–She3p motors bound to a She2p tetramer. In the majority of images, the only contact between the two motors appears to be at the base where She3p binds to She2p. We conclude that a She2p tetramer recruits two Myo4p–She3p motor complexes. Two-headed structures were not observed with either the S120Y or the L130Y mutant She2p dimers.
Two Myo4p–She3p motor complexes bind per She2p tetramer. (A) A field of metal-shadowed images of Myo4p–She3p alone. (B) A field of metal-shadowed images of Myo4p–She3p in the presence of She2p. Note the prevalent V-shaped structures, which show that two motor complexes bind per She2p tetramer. (C) Gallery of Myo4p–She3p alone. (D) Gallery of images showing two Myo4p–She3p motors bound to the She2p tetramer.
Two Myo4p–She3p motor complexes bind per She2p tetramer. (A) A field of metal-shadowed images of Myo4p–She3p alone. (B) A field of metal-shadowed images of Myo4p–She3p in the presence of She2p. Note the prevalent V-shaped structures, which show that two motor complexes bind per She2p tetramer. (C) Gallery of Myo4p–She3p alone. (D) Gallery of images showing two Myo4p–She3p motors bound to the She2p tetramer.
Total internal reflection fluorescence (TIRF) microscopy was used to determine whether the complex of two Myo4p–She3p motors bound to She2p can move processively on actin (Fig. 6 A). To follow motion of the ternary complex, a She2p-YFP fusion construct was created. Control experiments in budding yeast confirmed that the YFP tag on She2p did not affect the ability of the motor complex to bind because >98% correct localization of ASH1 mRNA was obtained in she2Δ yeast cells transformed with She2p-YFP (Fig. 2).
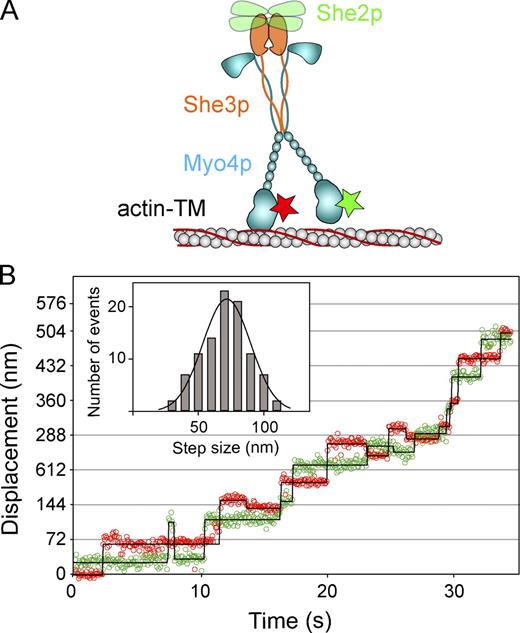
Myo4p–She3p moves processively when bound to tetrameric She2p. (A) Diagram of the experimental setup for the TIRF processivity assay. The motor complex is tracked by the YFP tag on She2p. The yellow stars represent YFP. TM, tropomyosin. (B) Myo4p–She3p bound to She2p-YFP (green) moves processively on yeast actin–tropomyosin (Tpm1p) filament (red) in 1 mM MgATP. Yellow arrowheads indicate the moving YFP particle. (C) Representative run length histogram at 1 mM Mg ATP on yeast actin–tropomyosin. A total of 67 processive runs was observed (n = 3).
Myo4p–She3p moves processively when bound to tetrameric She2p. (A) Diagram of the experimental setup for the TIRF processivity assay. The motor complex is tracked by the YFP tag on She2p. The yellow stars represent YFP. TM, tropomyosin. (B) Myo4p–She3p bound to She2p-YFP (green) moves processively on yeast actin–tropomyosin (Tpm1p) filament (red) in 1 mM MgATP. Yellow arrowheads indicate the moving YFP particle. (C) Representative run length histogram at 1 mM Mg ATP on yeast actin–tropomyosin. A total of 67 processive runs was observed (n = 3).
We previously showed that Myo4p–She3p is nonprocessive in the absence of She2p (Hodges et al., 2008), which is in agreement with other laboratories (Reck-Peterson et al., 2001; Dunn et al., 2007). When Myo4p–She3p was mixed with She2p-YFP, YFP particles were observed to move processively along single filaments of yeast actin–tropomyosin (Tpm1p) for distances of up to 6 µm (Fig. 6 [B and C] and Video 1). The mean speed was 1.50 ± 0.49 µm/s (mean ± SD) in 1 mM MgATP, consistent with previous Myo4p ensemble motility measurements (Reck-Peterson et al., 2001; Dunn et al., 2007; Hodges et al., 2008). A mean run length of 1.38 µm was obtained by fitting the distribution of measured run lengths (x) to a single exponential equation, y = Ae−x/λ, in which λ is the mean run length, and A is a constant (Fig. 6 C). The intensity of the majority of particles was consistent with approximately four YFPs from the tetrameric She2p (Fig. S2). Occasionally, aggregates with an intensity more than 10 times that of the mean particle were also observed to move processively but at a much slower speed (0.36 ± 0.19 µm/s in 1 mM MgATP; n = 6). Thus, the vast majority of moving particles are composed of tetrameric She2p with two Myo4p–She3p motors bound.
A Myo4p–She3p-She2p motor complex moving processively on an actin filament. Myo4p–She3p was mixed with She2p-YFP and visualized by TIRF microscopy. YFP particles (green) were observed to move processively along Alexa Fluor 594–labeled actin filaments (red) in 1 mM MgATP. The field of view is 5.3 × 7.4 µm. Data were collected at 10 frames per second and played back at 30 frames per second.
A Myo4p–She3p-She2p motor complex moving processively on an actin filament. Myo4p–She3p was mixed with She2p-YFP and visualized by TIRF microscopy. YFP particles (green) were observed to move processively along Alexa Fluor 594–labeled actin filaments (red) in 1 mM MgATP. The field of view is 5.3 × 7.4 µm. Data were collected at 10 frames per second and played back at 30 frames per second.
Processivity of Myo4p–She3p-She2p is seen with both yeast actin–tropomyosin and bare skeletal actin, but the run frequency is 11.2-fold higher on yeast actin–tropomyosin. For comparison, the run frequency of processive dimeric mammalian myosin Va on bare skeletal actin is three orders of magnitude higher than that of the Myo4p complex. Thus, it is likely that not all of the Myo4p–She3p is complexed with She2p at nanomolar concentrations, and more stable binding may require mRNA.
When the tetrameric mutant lacking the protruding α helix was tested in this assay, no processive runs were observed, which is consistent with results from our in vitro and in vivo characterization of this mutant. Occasional processive runs were observed with the two dimeric She2p point mutants (S120Y and L130Y), but the frequency of events was greatly reduced relative to WT*-She2p (Table II). Although the point mutations shift the equilibrium from tetramer to dimer, it is possible that a small fraction of tetramers remains for both mutants. In addition, She2p(S120Y) showed a tendency to form larger nonspecific aggregates, likely accounting for the observed processive runs with this mutant.
Frequency of processive runs by WT and mutant She2p
| She2p | Number of events | Event frequency |
| mm−2 s−1 | ||
| WT* | 59 | 56.4 |
| S120Y | 17 | 10.7 |
| L130Y | 5 | 1.3 |
| Δhelix | 0 | 0 |
| She2p | Number of events | Event frequency |
| mm−2 s−1 | ||
| WT* | 59 | 56.4 |
| S120Y | 17 | 10.7 |
| L130Y | 5 | 1.3 |
| Δhelix | 0 | 0 |
The values in the second column were obtained with chicken skeletal actin. Event frequency for WT* is 11.2 times higher on yeast actin–tropomyosin.
The Myo4p–She3p-She2p complex steps hand over hand
To determine whether the two Myo4p heads are coordinated during processive stepping, streptavidin-coated quantum dots (Qdots) were bound to an N-terminally biotinated Myo4p construct. An equimolar mixture of red (655 nm) and green (565 nm) Qdots was used to label the motor domains (Fig. 7 A). No processive runs were observed in the absence of She2p. She2p was then added to recruit two Myo4p–She3p motors per She2p tetramer. Most complexes were labeled with only one Qdot, allowing us to track the motion of one of the two motor domains. A histogram of step sizes showed a mean step of 71.8 ± 18 nm (Fig. 7 B, inset), equal to twice the 36-nm semirepeat of F-actin. This implies that the center of mass moves 36 nm each step, which is very similar to that of the more well-studied dimeric vertebrate myosin Va molecule. The stepping pattern of two representative singly labeled complexes is shown in Fig. S3. A few complexes contained a different colored Qdot on each head. The stepping pattern of the dual-labeled motor complex showed a hand-over-hand stepping pattern, with the lead and trailing head changing position with each step on actin, which is again very similar to myosin Va (Fig. 7 B).
Hand-over-hand stepping of Myo4p-She3p bound to She2p. (A) Diagram of the experimental setup for the TIRF processivity assay. Steps are tracked by Qdots bound to the N terminus of the motor domain. For the trace shown in B, one head is labeled with a 565-nm Qdot (green star) and the other with a 655-nm Qdot (red star). TM, tropomyosin. (B) Myo4p processive run with heads labeled with different colored Qdots showing a hand-over-hand stepping pattern, with lead and trailing heads exchanging position with each step. The displacement versus time for the 565- and 655-nm Qdots is shown by green and red open circles, respectively. Qdot positions were determined by the ImageJ plugin SpotTracker (Sage et al., 2005). Steps were identified using the Kerssemakers step-finding algorithm (solid lines; Kerssemakers et al., 2006). The inset shows a histogram of the step sizes, which averaged 71.8 ± 18 nm. Experiments were performed at 1 µM MgATP. The occasional back steps in this trace are not representative (see also Fig. S3). The green/red displacement versus time data is a single representative trace. A total of 15 traces from three experiments was analyzed to generate the histogram (inset).
Hand-over-hand stepping of Myo4p-She3p bound to She2p. (A) Diagram of the experimental setup for the TIRF processivity assay. Steps are tracked by Qdots bound to the N terminus of the motor domain. For the trace shown in B, one head is labeled with a 565-nm Qdot (green star) and the other with a 655-nm Qdot (red star). TM, tropomyosin. (B) Myo4p processive run with heads labeled with different colored Qdots showing a hand-over-hand stepping pattern, with lead and trailing heads exchanging position with each step. The displacement versus time for the 565- and 655-nm Qdots is shown by green and red open circles, respectively. Qdot positions were determined by the ImageJ plugin SpotTracker (Sage et al., 2005). Steps were identified using the Kerssemakers step-finding algorithm (solid lines; Kerssemakers et al., 2006). The inset shows a histogram of the step sizes, which averaged 71.8 ± 18 nm. Experiments were performed at 1 µM MgATP. The occasional back steps in this trace are not representative (see also Fig. S3). The green/red displacement versus time data is a single representative trace. A total of 15 traces from three experiments was analyzed to generate the histogram (inset).
Discussion
Here, we show that Myo4p–She3p, the single-headed class V myosin from budding yeast, supports processive motion on actin only when bound to the tetrameric adapter protein She2p. This is possible because the motor domain of Myo4p has a high-duty cycle (Hodges et al., 2008) and because two Myo4p–She3p motors are recruited per She2p tetramer. The two motors are coupled only by virtue of being bound to the same She2p tetramer in contrast to mammalian myosin Vs, which are dimerized via an α-helical coiled-coil. Given this fundamental difference, it was surprising that the two Myo4p–She3p motors bound to She2p take ∼72-nm steps in a hand-over-hand fashion, the same size and stepping pattern exhibited by myosin Va (Yildiz et al., 2003; Warshaw et al., 2005). The main determinants of the common step size of myosin Va and Myo4p–She3p are the long neck or lever arm that binds six CaMs and/or light chains and the pseudorepeat of the actin helix rather than the exact geometry by which the two heads are joined.
The long run lengths of myosin Va are believed to be possible because of gating, meaning that the two heads communicate to keep their kinetic cycles out of phase. This communication is thought to occur via strain transmitted through the two lever arms, which predominantly acts to prevent premature dissociation of the leading head from actin (Purcell et al., 2005; Veigel et al., 2005; Oguchi et al., 2008). Electron microscope images are consistent with the idea that myosin Va is distorted and under tension when both heads are attached to actin (Walker et al., 2000; Oke et al., 2010). Although we cannot rule out that the two Myo4p–She3p motors bound to She2p form a four-stranded coiled-coil, it appears likely that any strain between Myo4p heads is mediated indirectly via She2p. It is unclear whether the two Myo4p heads are gated, but the ∼1.4-µm run length of the Myo4p–She3p–She2p complex compares favorably with run lengths of 0.8–1.8 µm determined for myosin Va (Baker et al., 2004; Hodges et al., 2007; Ali et al., 2008). It has recently been argued that the detailed tuning of structure and intramolecular communication that ensures gating is not essential for processivity, provided that the motor heads have a sufficiently high-duty ratio and that multiple motor heads simultaneously interact with the actin network (Elting et al., 2011). The 1.4-µm run length of Myo4p corresponds to 39 steps of 36 nm. According to a simple two-state model, two heads cycling independently with no gating could achieve this run length if they had a duty ratio of ∼87% (Veigel et al., 2002). However, gating undoubtedly improves the efficiency of the motor by preventing the wasting of ATP on unproductive cycles and plays a role in minimizing backward motion under load by preventing the front head from releasing from the actin filament.
Features of She2p
Our conclusion that She2p is a tetramer, which is in agreement with two other groups (Chung and Takizawa, 2010; Heuck et al., 2010), is inconsistent with crystallographic experiments, which concluded that She2p is dimeric (Niessing et al., 2004). To reconcile the crystallographic experiments with the solution experiments, one must assume either that the N- and C-terminal residues that were truncated to obtain crystals are necessary for tetramerization or that the crystallization conditions abolished interactions necessary for tetramer formation while retaining the large and presumably stronger dimer interface.
Here, we show that the conserved helix that protrudes at right angles from the She2p structure provides all or part of the binding site for She3p (Fig. 1 B). Müller et al. (2011) also demonstrated that a mutant similar to our Δhelix does not bind She3p but assigned the primary function of this region to mRNA binding based on UV cross-linking. The major mRNA-binding site on She2p has been mapped to a basic helical hairpin RNA–binding motif consisting of two antiparallel α helices separated by a loop (residues 36–63; Fig. 1 B, tan-colored region; Niessing et al., 2004). The only other conserved region of She2p that has not been assigned a function is its C terminus, and that is dispensable for She3p binding (Müller et al., 2011). Collectively, it is thus likely that the primary role of the protruding helix on She2p is to bind She3p. Because only two motors are recruited per She2p tetramer, it is possible that two of the protruding helices are involved in binding one Myo4p–She3p motor.
Two point mutants convert the She2p tetramer into a dimer. S120Y lies at the interface of the crystallographic dimer, whereas L130Y lies at the interface of the proposed elongated tetrameric interface (Müller et al., 2009). Mutation of these residues might produce different dimers, i.e., a longitudinal versus a lateral dimer. Indeed, the two mutations had very different effects on ASH1 mRNA localization in the cell. She2p(L130Y) was indistinguishable from WT, whereas She2p(S120Y) showed no correct ASH1 mRNA localization. The lack of ASH1 mRNA localization with She2p(S120Y) is consistent with its weak binding to both Myo4p–She3p and mRNA (Niessing et al., 2004), which likely compromises the coupling between motor and cargo (Table III). But how can the She2p(L130Y) mutant function like WT in the cell, as it did not support processive motion in vitro? One possibility is that multiple motors are recruited when multiple mRNAs are cotransported (Lange et al., 2008). Alternatively, it is possible that the presence of mRNA in the cell acts to stabilize some or all of the She2p(L130Y) as a tetramer.
Summary of properties of WT and mutant She2p constructs
| She2p | Oligomeric state in solution | Binds to She3p in vitro | Processive | Binds to mRNA in vitro | Correct localization of ASH1 mRNA in yeast |
| WT* | Tetramer | ++++ | + | ++++a | + |
| Kd = 0.1 µMb | |||||
| S120Y | Dimer | + | − | +a | − |
| L130Y | Dimer | +++ | − | ++a | + |
| Kd = 1.0 µMb | |||||
| Δhelix | Tetramer | − | − | ND | − |
| She2p | Oligomeric state in solution | Binds to She3p in vitro | Processive | Binds to mRNA in vitro | Correct localization of ASH1 mRNA in yeast |
| WT* | Tetramer | ++++ | + | ++++ | + |
| Kd = 0.1 µM | |||||
| S120Y | Dimer | + | − | + | − |
| L130Y | Dimer | +++ | − | ++ | + |
| Kd = 1.0 µM | |||||
| Δhelix | Tetramer | − | − | ND | − |
Relative ability of 1 µM She2p to bind to mRNA using filter-binding experiments (Niessing et al., 2004).
Values from mRNA filter-binding experiments using the ASH1 mRNA E3 zip code element (Müller et al., 2009).
Role of mRNA in complex formation
Our data show that baculovirus-expressed Myo4p–She3p can bind She2p at nanomolar concentration in the absence of mRNA. However, the relatively low run frequency compared with dimeric mammalian myosin Va suggests that not all of the Myo4p–She3p is likely to be bound to She2p in the absence of mRNA. This observation is consistent with several other lines of evidence. A 20-S complex isolated from budding yeast contained Myo4p, She3p, and She2p only when intact mRNA was present (Chung and Takizawa, 2010). RNase treatment caused the complex to dissociate based on sucrose gradient sedimentation assays. Another recent study showed that mRNA substantially stabilizes the She3p–She2p interaction, caused in part by the ability of She3p to directly bind to mRNA. It was concluded that the mRNA itself triggers joining of all components into the high-specificity mature transport complex (Müller et al., 2011). An attractive feature of this idea is that only She2p that has bound and escorted ASH1 mRNA out of the nucleus would form a high affinity and specific complex with cytoplasmic Myo4p–She3p. We are currently examining the single molecule properties of Myo4p–She3p-She2p in the presence of ASH1 mRNA and speculate that the number of processive events will increase substantially in the presence of mRNA.
Strategies for single-headed motors to move processively
Other mechanisms by which single-headed motors can achieve processive motion within the cell have been proposed. Full-length myosin VI isolated from cells or expressed in baculovirus is monomeric (Sweeney and Houdusse, 2010). It was suggested that when myosin VI is bound to the adapter proteins optineurin or Dab2, an internal dimerization sequence in the rod is exposed, which triggers coiled-coil formation (Phichith et al., 2009). Interestingly, even the monomeric adapter protein Dab2 is capable of dimerizing myosin VI because it contains two discrete binding sites for the tail (Yu et al., 2009). A preexisting oligomer is thus not required for dimerization. Myo4p–She3p differs from myosin VI in that it is not internally dimerized but simply joined via She2p.
Another mechanism involves clustering of monomeric motors on membrane vesicles, which then favors internal dimerization. Monomeric myosin VIIA required forced dimerization to transport cargo to the filopodial tip (Sakai et al., 2011). Because the adapter proteins MyRip/Rab27a did not directly dimerize myosin VIIA, it was proposed that the motors dimerize when closely clustered on the vesicle. Deletion of a predicted short coiled-coil domain did not diminish localization, and, so, the dimerization domain in myosin VIIA is unknown.
An alternative mechanism for monomeric motors to achieve long-range transport is to work in ensembles. The motion of endosome-sized nanospheres coated with multiple monomeric myosin VI motors on a skinned keratocyte lamellipodium was tracked (Sivaramakrishnan and Spudich, 2009). Multiple monomers interacting with multiple actin filaments could cooperate to move continuously for long distances under low load. Similarly, we previously showed that robust continuous movement was observed when multiple (approximately three to four) monomeric Myo4p–She3p motors were bound to a Qdot cargo (Hodges et al., 2008). Here, we show that only two Myo4p–She3p motors are necessary to support long-range processive transport when coupled by She2p.
Implications for cellular cargo transport
Our findings imply that a mechanism for regulating cargo motility by monomeric motors is to help or hinder their transition to a functional dimer. In budding yeast, She2p interacts with ASH1 mRNA in the nucleus and shuttles it into the cytoplasm (Fig. 8; Du et al., 2008; Shen et al., 2009). Based on our studies, once in the cytoplasm, the She2p–mRNA complex should be able to recruit two Myo4p–She3p motors per zip code, the localization element on the mRNA that binds She2p. Although She2p can bind to Myo4p–She3p in the absence of mRNA, in the cell, there needs to be a checkpoint to ensure that She2p is coupled to mRNA before it encounters the motor. This is likely facilitated by the fact that She3p is cytoplasmic, but She2p binds to mRNA in the nucleus and then escorts it into the cytoplasm. A recent study suggests an additional level of control, namely that a high-affinity, specific complex with the correct mRNA is formed only in the presence of She3p (Müller et al., 2011). This latter step would ensure that only complexes containing mRNA will get transported on actin cables in the proposed manner.
Cartoon showing how Myo4p–She3p supports continuous motion of ASH1 mRNA. She2p binds to mRNA and escorts it out of the nucleus. In the cytoplasm, two single-headed Myo4p–She3p complexes are recruited by the She2p tetramer. The two-headed complex steps in a hand-over-hand fashion, with each head moving ∼72 nm. The processive complex transports mRNA to the bud tip. Once there, the mRNA is anchored, and the complex dissociates into nonprocessive monomers and free She2p.
Cartoon showing how Myo4p–She3p supports continuous motion of ASH1 mRNA. She2p binds to mRNA and escorts it out of the nucleus. In the cytoplasm, two single-headed Myo4p–She3p complexes are recruited by the She2p tetramer. The two-headed complex steps in a hand-over-hand fashion, with each head moving ∼72 nm. The processive complex transports mRNA to the bud tip. Once there, the mRNA is anchored, and the complex dissociates into nonprocessive monomers and free She2p.
In vivo, redundancies are also built into the system. Many mRNAs, including ASH1 mRNA, have multiple zip codes and should thus be able to recruit multiple two-headed motor complexes. In addition, the single mRNA particle that gets translocated to the bud tip contains multiple copies of mRNA (Lange et al., 2008). Although a single pair of motors is sufficient to walk in vitro for over a micrometer, in the cell, multiple pairs of motors may be beneficial for enhancing run lengths and for overcoming any resistive loads. In agreement with this, increasing the number of Myo4p–She3p motors directly bound to a reporter mRNA (which bypasses She2p) increased the efficiency of mRNA transport to the bud (Chung and Takizawa, 2010).
Once at its destination, a switch is necessary to dissociate the complex and unload cargo. One potential regulatory mechanism could involve phosphorylation of She3p, which has been shown to be phosphorylated at S343, S348, and S361 (Landers et al., 2009). It has been shown that She3p possesses a novel activity required for ASH1 mRNA localization that is independent of its binding to either Myo4p or She2p. This function is negatively regulated by phosphorylation and may involve association with mRNA directly. This possibility is supported by a recent study that shows that She3p can bind mRNA (Müller et al., 2011).
Unlike higher eukaryotes that use microtubules and kinesin/dynein for long-range transport, budding yeast uses motion of two class V myosins on actin cables for cargo transport. Here, we show that the mechanism used by one of these motors, Myo4p–She3p, is a variation on a theme of cargo-mediated dimerization that is also used by the monomeric vertebrate class VI and VII motors to ensure regulated transport over micrometer-long distances.
Materials and methods
Constructs
Sequence-encoding full-length Myo4p (1,472 amino acids) was cloned into pVL1392 with a C-terminal FLAG tag to facilitate purification by affinity chromatography. A second full-length Myo4p construct contained a biotin tag at the N terminus for attachment to streptavidin Qdots (Invitrogen). The biotin tag is an 88–amino acid sequence segment from the Escherichia coli biotin carboxyl carrier protein, which, when expressed in Sf9 cells, is biotinated at a single Lys residue (Cronan, 1990; Li and Cronan, 1992). A truncated Myo4p construct lacking the motor domain and neck (K924 to the end of MYO4) with C-terminal FLAG tag was also cloned into pAcSG2 for baculovirus expression. Yeast CaM and the yeast light chain Mlc1p were separately cloned into the pAcUW51 plasmid under the polyhedrin promoter. A C-terminally HIS-tagged Mlc1p was cloned into pAcSG2 (BD) and a C-terminal Myc-tagged yeast CaM in pacUW51. These constructs were used to facilitate identification of the bound light chains by Western blotting. For bacterial expression, yeast CaM and Mlc1p were each cloned into pNew (a kanamycin-resistant plasmid with T7 promoter). SHE3 with a C-terminal His tag was cloned into the baculovirus vector pAcSG2 as previously described (Hodges et al., 2008).
The SHE2 gene was amplified by PCR from S. cerevisiae genomic DNA and verified as the coding sequences found at the Saccharomyces genome database. For cellular studies, SHE2 was cloned into a pRS314 derivative with ADE2 selection behind the inducible GAL1 promoter. For bacterial expression, the SHE2 gene was cloned into pGEX-2T. A mutant She2p called WT* had four Cys residues mutated to Ser (C14S, C68S, C106S, and C180S). All additional variations of She2p were made on the WT* backbone for both the cellular and in vitro experiments. This includes two point mutants (S120Y and L130Y), a Δhelix construct (lacking residues 174–179; VQFAIK), and a construct containing a C-terminal YFP (She2p-GSG-YFP-FLAG). All changes were verified by DNA sequencing. She2p constructs cloned with fluorescent proteins at either the N or C terminus were shown to be completely functional in Δshe2 yeast cells.
S. cerevisiae actin was obtained from genomic DNA and cloned into the pAcSG2 baculovirus vector following a FLAG and recombinant Tobacco Etch Virus (rTEV) cleavage site (DYKDDDDK-ENLYFQ-MDSEV …). This construct allowed the actin to be purified on a FLAG column followed by cleavage of the tag with rTEV protease (between Q and M), removing all nonnative residues before the start codon of the yeast gene. Met was included in the actin sequence because actin retains its initiator Met in yeast (Cook et al., 1991).
Budding yeast tropomyosin (Tpm1p) was a PCR product made using S. cerevisiae genomic DNA. To mimic acetylation, an Ala-Ser was added at the N terminus before the start codon (MASMDSEV …) (Maytum et al., 2001). The construct was cloned into the pET3 vector.
Protein expression and purification
Myosin constructs were expressed in Sf9 cells using the baculovirus system and purified on a FLAG affinity column essentially as previously described (Fig. S1; Krementsova et al., 2006). In brief, Sf9 cells were infected with recombinant viruses for the Myo4p heavy chain, She3p, yeast CaM, and MLC1p and incubated with shaking for ∼72 h at 27°C. Cells were pelleted and resuspended in 10 mM imidazole, pH 7.4, 0.3 M NaCl, 5 mM MgCl2, 7% sucrose, 1 mM EGTA, 2 mM DTT, 25 µg/ml yeast CaM, 25 µg/ml MLC1p, and protease inhibitors (0.5 mM 4-(2-aminoethyl)benzenesulfonyl fluoride, 5 µg/ml leupeptin, and 0.78 mg/ml benzamidine). The cells were lysed by sonication, and the lysate was centrifuged for 15 min at 250,000 g in the presence of 2 mM MgATP. After incubation with FLAG affinity resin (Sigma-Aldrich) for 30 min, the resin was sedimented for 5 min at 1,000 rpm and washed with buffer (10 mM imidazole, pH 7.4, 0.3 M NaCl, and 1 mM EGTA). Bound protein was eluted using a 0.1-mg/ml solution of FLAG peptide in the same buffer. Fractions were pooled, concentrated in 50% glycerol, and stored at −20°C.
All She2p variants were expressed as GST fusion proteins in E. coli BL21(DE3). A 5-ml overnight culture grown in Lennox-L broth was diluted 50 times into enriched media and grown for 3.5 h at 37°C. After induction with 50 µM IPTG, the cells were grown overnight at 25°C. Cells were lysed by sonication in PBS with protease inhibitors and loaded onto a 20-ml glutathione Sepharose 4B column (GE Healthcare). The column was washed with 5 vol PBS and drained. 90 U thrombin (Haematologic Technologies, Inc.) in 5 ml PBS was added to the column, and the slurry was incubated for 1 h at room temperature with occasional mixing. The column was drained and washed with ∼25 ml PBS. 1 mM DTT was added followed by 2 µg/ml leupeptin to inactivate the thrombin. Thrombin was removed by passing the mixture over a benzamidine Sepharose 6B column (GE Healthcare). The protein was concentrated in a 50-ml Centricon (Millipore) to ∼5 ml and stored in 50% glycerol, 10 mM imidazole, pH 7.4, 0.2 M NaCl, and 1 mM DTT.
Chicken skeletal actin was prepared from acetone powder (Pardee and Spudich, 1982). Yeast actin was expressed using the baculovirus/insect cell expression system. Sf9 cells were infected with recombinant baculovirus encoding yeast actin and harvested 72 h later. Cells were lysed using 1 M Tris-HCl, pH 7.5, 0.6 M KCl, 0.5 mM MgCl2, 0.5 mM NaATP, 1 mM DTT, 4% Triton X-100, 1 mg/ml Tween 20, 0.5 mM AEBSF, 0.5 mM tosyllysine chloromethyl ketone, and 5 µg/ml leupeptin and stirred for 1.5 h at 4°C. The lysate was clarified for 1 h at 177,700 g. The supernatant was dialyzed overnight against a buffer containing 300 mM NaCl, 10 mM imidazole, pH 7.5, 0.2 mM CaCl2, 0.5 mM NaATP, 0.2 mM DTT, and 1 µg/ml leupeptin. After clarifying at 25,000 g, the supernatant was incubated with 3 ml FLAG resin for 1 h. The resin was washed with the same buffer as above but without NaATP and DTT. Actin was eluted using 100 µg/ml FLAG peptide in 300 mM NaCl, 10 mM imidazole, pH 7.5, 0.2 mM CaCl2, 0.5 mM NaATP, and 1 µg/ml leupeptin. Actin was then dialyzed against G buffer overnight (5 mM Tris, pH 8.3, 0.2 mM CaCl2, 0.2 mM DTT, 0.2 mM NaATP, and 1 µg/ml leupeptin), clarified for 30 min at 400,000 g, and concentrated. The actin concentration was determined by the Bradford assay, and rTEV protease (Miller and Trybus, 2008) was added at a 1:8 molar ratio (rTEV protease/actin) and incubated overnight. G-actin was polymerized by adding 50 mM KCl, 4 mM MgCl2, and 1 mM MgATP.
Tpm1p was expressed in E. coli BLR(DE3) cells grown in Lennox-L broth. They were induced with 0.4 mM IPTG and grown overnight at 27°C before being harvested and frozen. Tpm1p was purified using a protocol similar to that in Maytum et al. (2000). In brief, cells were lysed by sonication. The clarified supernatant was boiled for 10 min and clarified for 10 min at 10,000 rpm. The soluble Tpm1p was precipitated by lowering the pH to 4.5. The Tpm1p was further purified using a 1-ml MonoQ column with 0–1 M NaCl gradient with the protein eluting at ∼250 mM NaCl. Only fractions that showed a single band by SDS-PAGE and that had a 260:280-nm absorbance ratio below 0.8 were kept. Tpm1p was concentrated and stored in 50% glycerol with 0.2 M NaCl, 10 mM imidazole, pH 7.4, 1 mM EGTA, 1 µg/ml leupeptin, and 1 mM DTT at −20°C.
Yeast CaM and Mlc1p were expressed in E.coli BL21(DE3) grown in enriched media overnight. Cells were lysed by sonication in 50 mM Tris-Cl, pH 7.5, 2 mM EDTA, 0.2 mM AEBSF, and 1 µg/ml leupeptin and clarified for 30 min at 27,000 g. The supernatant was boiled for 5 min, cooled, and clarified for 15 min at 27,000 g. For yeast CaM, 5 mM CaCl2 and 1 mM DTT were added, and the protein bound to a Phenyl-Sepharose CL-4B column (GE Healthcare) was preequilibrated with 50 mM Tris-Cl, pH 7.5, 100 mM NaCl, 5 mM CaCl2, and 1 mM DTT. The column was washed with the same buffer containing 0.1 mM CaCl2 and then the same low-calcium buffer with 0.5 M NaCl. Bound protein was eluted with 50 mM Tris-Cl, pH 7.5, 1 mM DTT, and 1 mM EGTA. The protein was stored in 50% glycerol, 10 mM imidazole, pH 7.4, 0.2 M NaCl, 1 mM EGTA, and 1 mM DTT at −20°C. For Mlc1p, the clarified supernatant after boiling was precipitated with ammonium sulfate to 80% saturation, pelleted for 15 min at 27,000 g, and dialyzed against 20 mM imidazole, pH 7.4, 200 mM NaCl, and 1 mM EGTA. Mlc1p was stored in 50% glycerol, 10 mM imidazole, pH 7.4, 0.2 M NaCl, 1 mM EGTA, and 1 mM DTT at −20°C.
ASH1 mRNA reporter system
A two-plasmid system was used to follow ASH1 mRNA localization. The two plasmids pG14-MS2-GFP with LEU2 selection and YEplac195 lacZ-MS2-ASH1 with URA3 selection were gifts from R. Long (Medical College of Wisconsin, Milwaukee, WI; Bertrand et al., 1998). pG14-MS2-GFP expresses an MS2 coat protein–GFP under a constitutive promoter. The MS2-GFP chimera contains an NLS to confine unbound MS2-GFP to the nucleus. YEplac195 lacZ-MS2-ASH1 expresses the ASH1 mRNA along with six stem loop–binding motifs for the MS2 coat protein under control of a galactose-inducible promoter. The URA3 marker in YEplac195 lacZ-MS2-ASH1 was replaced with HIS3.
Visualization of transformed yeast
The mutant yeast strain K5477 (MATα ade2 his3 leu2 trp1 ura3 can1-100 she2::URA [W303a background]) was used. The yeast strain she2Δ was transformed with three plasmids. Two were required for the ASH1 reporter system. The third plasmid contained the various forms of She2p, including a WT used as a positive control and a plasmid without insert as the negative control. Transformants were grown on the appropriate complete synthetic medium lacking the nutrients needed to maintain the plasmids. To induce expression of the various versions of She2p along with the reporter mRNA, a single colony was grown overnight in the same synthetic medium containing 2% galactose as the sole carbon source. 4 h before visualization, the yeast cells were streaked in a fresh patch to ensure exponential growth when viewed and scored. The live cells were mounted on a slide in the appropriate liquid medium containing galactose and viewed with an inverted microscope (TE2000-E2; Nikon) using a Plan Apo 60× oil objective lens and differential interference contrast and FITC filters. A fluorescence illuminator (X-Cite 120; EXFO) and a 14-bit camera (CoolSNAP HQ2; Photometrics) were used. Images were processed using NIS Elements (Nikon) and ImageJ (National Institutes of Health) software. Actively budding yeast were scored according to the location and number of fluorescent particles seen. Actively budding yeast cells that contained one fluorescent mRNA particle were scored according to whether the particle was located in the bud tip (correct localization) or the mother (incorrect localization).
Analytical ultracentrifugation
An analytical ultracentrifuge (Optima XL-I; Beckman Coulter) was used to determine the sedimentation coefficients of the expressed constructs. Sedimentation velocity runs were performed in the An-60 Ti rotor at 40,000 rpm and 20°C in 10 mM Hepes, pH 7.0, 0.3 M NaCl, 1 mM DTT, 1 mM EGTA, and 1 mM NaN3. Sedimentation values were corrected for density and viscosity of the solvent. Sedimentation coefficients were determined by curve fitting to one or more species using the dc/dt program (Philo, 2000). Sedimentation equilibrium data were collected at 15,000 rpm and 4°C, 10 mM Hepes, pH 7.0, 0.3 M NaCl, 1 mM DTT, 1 mM EGTA, and 1 mM NaN3. The data were analyzed with Origin software provided with the Optima XL-I ultracentrifuge. For molecular mass calculations, the fit to the mean of three independent scans at equilibrium was determined for each protein sample, and the mean and SD of those values were calculated for each construct.
Actin-pelleting assay
0.3 µM Myo4p–She3p was mixed with various She2p constructs (∼1.2 µM monomer) and incubated for 10 min in 10 mM imidazole, pH 7.5, 1 mM EGTA, 1 mM DTT, and 0.2 M NaCl. Both proteins were preclarified before mixing together. 2.4 µM actin was then added. The mixtures were pelleted for 20 min at 400,000 g. The supernatant was removed, and the pellet was washed twice with buffer. Protein content of the supernatant and pellet was assessed by SDS-PAGE.
FLAG resin pull-down assay
A truncated Myo4p construct lacking the motor domain and neck was coexpressed with She3p and assessed for binding to various She2p constructs. Both proteins were clarified at 400,000 g for 15 min before mixing. Myo4p–She3p and She2p (each at 1 µM) were mixed together in 250 µl and loaded onto 0.3-ml FLAG columns (Sigma-Aldrich). The columns were washed with 10 mM imidazole, pH 7.5, 1 mM EGTA, and 0.2 M NaCl, and bound proteins were eluted with 0.1 mg/ml FLAG peptide in wash buffer. Fractions were analyzed on SDS gels.
EM
Myo4p–She3p and She2p-WT* were dialyzed into 10 mM MOPS, pH 7, 50 mM KCl, 1 mM MgCl2, 1 mM EGTA, and 1 mM DTT. Myo4p–She3p and She2p were mixed at a molar ratio of 6:1, incubated for 30 min in ice, and then diluted in the same buffer containing 50% glycerol to a final concentration of 60–120 µg/ml. 5–10 µl of each sample was sprayed on a freshly split mica surface, dried for 2 h under vacuum, rotary shadowed with platinum at an angle of 7°, and replicated with carbon in a Balzers 410 freeze-fracture machine. Replicas were photographed in an electron microscope (410; Philips) operating at 60 kV at a magnification of 75,300. The negatives were digitally scanned at 1,000 pixels per inch. Images were obtained from areas at the edge of each droplet that showed distinct molecules and a clear background.
TIRF microscopy
For experiments in which run lengths were measured, motion was followed from the YFP signal on She2p. She2p-YFP was clarified at 400,000 g for 20 min to remove aggregates. 0.2 µM Myo4p–She3p was mixed with 0.05 µM She2p-YFP (moles monomer) and was allowed to incubate on ice for at least 1 h. Flow cells were prepared by introducing the following solutions into the flow cell: 0.1 mg/ml N-ethylmaleimide–modified skeletal muscle myosin (5-min incubation), 5× rinse of 1 mg/ml BSA (2 min), Alexa Fluor 594 phalloidin yeast actin–Tpm1p filaments or chicken skeletal muscle actin filaments (2–5 min), 5× rinse with motility buffer, and, finally, Myo4p–She3p diluted in motility buffer with either 10 µM, 1 mM, or 2 mM MgATP. Motility buffer consists of 50 mM KCl, 25 mM imidazole, pH 7.4, 4 mM MgCl2, 1 mM EGTA, 50 mM DTT, 1 mg/ml BSA, 0.2 mg/ml yeast CaM, 0.2 mg/ml Mlc1p, an ATP-regenerating system (0.5 mM phosphoenolpyruvate and 100 U/ml pyruvate kinase), and an oxygen-scavenging system (3 mg/ml glucose, 0.1 mg/ml glucose oxidase, and 0.18 mg/ml catalase). The final concentrations were 20 nM Myo4p–She3p and 5 nM She2p-YFP. N-ethylmaleimide–modified skeletal muscle myosin forms a strong and ATP-insensitive bond with actin and was used to attach the actin filaments to the coverslip.
Data were collected on an inverted microscope (Eclipse Ti-U; Nikon) equipped with a 100× Plan Apo objective lens (1.49 NA) and auxiliary 1.5× magnification for through-the-objective TIRF microscopy. The YFP was excited with a 473-nm laser line. Images were obtained using a camera (XR/Turbo-Z; Stanford Photonics) running Piper Control software (v2.3.39). The pixel resolution was 95.4 nm. Data were collected at 10–20 frames per second. Movement of She2p-YFP was tracked manually using ImageJ.
For experiments in which the stepping pattern of the Myo4p–She3p-She2p complex was determined, motion was followed from a Qdot attached to the N terminus of Myo4p. She2p (without YFP) was clarified at 400,000 g for 20 min before use. N-terminally biotinated Myo4p-She3p was mixed with red (655-nm emission) streptavidin Qdots (Invitrogen) at a ratio of five Qdots per Myo4p monomer. A similar solution was prepared using green (565-nm emission) Qdots, and both solutions were incubated on ice for 20 min. They were then mixed in a 1:1 ratio, and She2p was added, resulting in final concentrations of 40 nM N-terminally biotinated Myo4p-She3p and 160 nM She2p (monomer concentration). Flow cells were prepared as described above. Alexa Fluor 594 phalloidin yeast actin–Tpm1p filaments were used. The Myo4p–She3p-She2p–Qdot mixture was diluted 200-fold in motility buffer containing 1 µM MgATP and 2 µM Tpm1p and added to the flow cell.
Data were collected on an inverted microscope (TE2000; Nikon) equipped with a 100× Plan Apo objective lens (1.49 NA) and auxiliary 1.5× magnification for through-the-objective TIRF microscopy. Qdots were excited with a 488-nm argon laser line. Images were obtained using a charge-coupled device camera (XR Mega-S30; Stanford Photonics) running Piper Control software (v2.3.14). Dual-color simultaneous imaging was achieved through a beam splitter (Dual-View; Optical Insights). The pixel resolution was 58.5 nm, but data were collected using 2 × 2–pixel binning, resulting in a final resolution of 117.6 nm. Data were collected at 15 frames per second.
Movement of Qdots on actin was tracked using ImageJ and the automated motion-tracking plugin SpotTracker (Sage et al., 2005). Data from the red and green channels were then color aligned by taking simultaneous red and green images of multicolored fluorescent microspheres at different locations within the field of view. Using a custom MATLAB program (MathWorks), a cross-correlation routine was used to determine the local displacements required to align the red image with the green one. The horizontal and vertical displacements required to correct each data point from the red channel were then calculated from nearby correlation values using 2D linear interpolation. Displacement versus time was calculated from the color-corrected data. Step size and dwell time were calculated in MATLAB using the Kerssemakers step-finding algorithm (Kerssemakers et al., 2006).
Online supplemental material
Fig. S1 shows gels of expressed Myo4p–She3p and a FLAG resin pull-down assay. Fig. S2 compares the fluorescence intensity of She2p-YFP and myosin Va–YFP. Fig. S3 shows the stepping pattern of Myo4p–She3p bound to She2p. Video 1 shows a Myo4p–She3p-She2p motor complex moving processively on an actin filament.
Acknowledgments
We thank Dave Warshaw for the use of the TIRF microscopes, Guy Kennedy for assistance with the TIRF microscopy, Art Michalek for the dual-color alignment, and Patricia Fagnant for the yeast actin expression.
This work was supported by National Institutes of Health grants AR35661 to H.L. Sweeney and GM078097 to K.M. Trybus.
References
Author notes
E.B. Krementsova and A.R. Hodges contributed equally to this paper.



![Figure 2. Ability of various She2p constructs to correctly localize ASH1 mRNA particles in budding yeast. (A) Representative images of correctly and incorrectly localized ASH1 mRNA. Correctly localized mRNA is found in the bud, whereas particles that are not transported remain in the mother cell. (left) Differential interference contrast [DIC]. (middle) Epifluorescence to visualize GFP-labeled ASH1 mRNA. (right) Merged images. (B) Percentage of correctly localized ASH1 mRNA particles for various She2p constructs. The indicated constructs were transformed into a she2Δ strain. Constructs lacking She2p do not localize ASH1 mRNA. WT* is the WT construct with four Cys residues (14, 68, 106, and 180) mutated to Ser. All other constructs are based on the WT* backbone. Addition of a fluorescent protein to either the N or C terminus of She2p had no effect on cellular function. Data are shown for a C-terminal fluorescent protein, but the data are identical for YFP at the N terminus. The negative control (neg) is a plasmid lacking an insert. The number of cells counted is indicated above each bar. Data were derived from at least two independent transformations per construct.](https://cdn.rupress.org/rup/content_public/journal/jcb/195/4/10.1083_jcb.201106146/5/m_jcb_201106146_rgb_fig2.jpeg?Expires=1781618535&Signature=Tyk6CEmHA9wXVFGKwiUrW0ZNYviP8YKtBg3ZHKXF8Ws4iT91fqIJP4rA-wrqxSNT7e9lrcm22NobADYMcdeTjGzwAmnocElZxkC0vfwK~6LVGg8fhU4hlxH4ifSgg8zxZEX4QCgoqFVZDyUIuFMOg-mVq7dhKVKargkB-CLUPgUctviC6IUCh7XgtvH8xhROpXRT68gO3DrJL9a3Wrklbgeght1G6gPJyA30HLREbda7KNkSZO3SBCJMlJyxPph~9-hawlsxeQCAQ1W3WKFK8XAswkKu33NzqqZ6BP2PZ8vcnSM6EKpk9D~B~0XklL0XWZ1h6d3IzapcGTwk61JqAg__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)