The serine/threonine kinase Par1b promotes cell–cell adhesion and determines the polarity of the luminal domain in epithelial cells. In this study, we demonstrate that Par1b also regulates cell–extracellular matrix (ECM) signaling in kidney-derived Madin–Darby canine kidney (MDCK) cells and identified the rho–guanosine triphosphatase adaptor and scaffolding protein IRSp53 as a Par1b substrate involved in this pathway. Par1b overexpression inhibits basal lamina formation, cell spreading, focal adhesion, stress fiber formation, and compaction, whereas Par1b depletion has the opposite effect. IRSp53 depletion mimics Par1b overexpression on cell–ECM signaling and lumen polarity but had no effect on adherens junction formation. Par1b directly phosphorylates IRSp53 on S366 in cell lysates and stimulates phosphorylation on S453/3/5 via an indirect mechanism. A Par1b phosphorylation–deficient IRSp53 mutant but not the wild-type protein efficiently rescues both the cell spreading and the lumen polarity defects in Par1b MDCK cells. Our data suggest a model in which Par1b phosphorylation prevents recruitment of IRSp53 effector proteins to its Src homology domain 3 by promoting 14-3-3 binding in the vicinity of that domain.
Introduction
A hallmark of the differentiation of transporting epithelia is the generation of a luminal or apical domain (Rodriguez-Boulan and Nelson, 1989; O’Brien et al., 2002). Nonstratified epithelia differ in the position of their luminal domain: columnar epithelia (e.g., kidney, intestine, or pancreas), display their lumina at the cell apex, separated by tight junctions from lateral domains enriched in cell–cell adhesion molecules, and opposite to basal domains enriched in receptors for the ECM. In contrast, hepatic epithelial cells establish their lumina, the bile canaliculi (BC), at sites of intercellular contact, separated by tight junctions from basolateral domains specialized for cell–cell interactions and exchange with the bloodstream (sinusoidal membranes).
In columnar epithelia, apical polarity determinants such as the Crumbs/Pals/Patj and the Cdc42–Par6–aPKC signaling complexes and Rab-GTPases, such as Rab8a and 11a that connect them to the polarized exocytic machinery, are essential for the formation of a luminal domain in vivo and in culture (Martin-Belmonte et al., 2007; Wang and Margolis, 2007; Bryant et al., 2010). Likewise, E-cadherin–mediated signaling provides a cue for lumen formation, per se (Vega-Salas et al., 1988). Cell–ECM signaling, on the other hand, appears to determine the orientation of polarity rather than cell polarity itself (Bryant and Mostov, 2008). Thus, when cultured in the absence of neighboring cells, kidney-derived MDCK cells restrict luminal markers to the free cell surface (Vega-Salas et al., 1987) When suspended in collagen I gels, MDCK cells proliferate to form hollow cysts. β1-Integrin orients the apical surface to face the cyst lumen by activating Rac1 and inhibiting rhoA and its targets ROCK and myosin II. Rac1 inside-out signaling in turn organizes laminin into a basal lamina that is essential for lumen orientation (O’Brien et al., 2001; Yu et al., 2005, 2008). Hepatocytes on the other hand are surrounded by ECM components, but unlike columnar epithelia, do not organize these components into a basal lamina. Indeed, basement membrane deposition, as it occurs in pathological situations such as cholestasis, leads to loss of hepatocyte polarity (Martinez-Hernandez and Amenta, 1993). In primary hepatocyte cultures, BC form only on moderately adhesive surfaces or in ECM sandwich cultures but not when monolayers are maintained on substrates that promote extensive cell spreading (Michalopoulos and Pitot, 1975; Dunn et al., 1991; Moghe et al., 1996; Cho et al., 2006). Thus, hepatocytes appear to engage different ECM-signaling mechanisms than do columnar epithelia to organize their intercellular luminal domains.
Work from our group has identified the serine/threonine kinase Par1b/EMK1/MARK2 as a regulator of epithelial lumina. Depletion of Par1b or inhibition of its function with a dominant-negative construct prevented lumen formation in MDCK cells in 3D collagen cultures and the formation of BC-like lumina in the hepatic cell line WIFB. However, Par1b overexpression promoted lateral lumina, resembling the organization of hepatic BC in MDCK cells, a model for columnar epithelia with an apical luminal domain (Cohen et al., 2004). Although Par1 substrates that mediate polarity in other systems such as during convergent extension, during neurite polarization and in the establishment of oocyte polarity are beginning to emerge (Matenia and Mandelkow, 2009), no Par1b substrates relevant for epithelial lumen polarity have been identified so far. It has been proposed, however, that Par1b regulates the organization of the epithelial lateral membrane because the active form of the kinase is enriched at sites of cell–cell contact (Suzuki et al., 2004) and inhibition of Par1b disrupted cell–cell adhesion (Böhm et al., 1997; Elbert et al., 2006).
In this study, we report that Par1b regulates cell spreading and the actin cytoskeleton at the basal domain of MDCK cells, and we identify IRSp53 (insulin receptor substrate p53/58) as a novel Par1b substrate involved in cell adhesion signaling. IRSp53 connects the rho-GTPases Cdc42 and Rac1 to their downstream effector molecules via a Src homology domain 3 (SH3), thus functioning as a rho-GTPase signaling platform (Scita et al., 2008). Overexpression of IRSp53 in most cell types induces extensive actin-containing cell protrusions dependent on Cdc42, Rac1, and their SH3 domain–containing effectors (Krugmann et al., 2001; Miki and Takenawa, 2002). The inverted BAR domain in IRSp53 has been proposed to mediate the membrane deformation that accompanies actin polymerization to allow for filopodia formation (Ahmed et al., 2010). In contrast, the functions of endogenous IRSp53 are less understood. An IRSp53 knockout mouse displayed defects in dendritic spine formation, a Rac1-dependent process (Sawallisch et al., 2009). IRSp53 knockdown (KD) inhibited cytokine-induced membrane ruffling in cultured macrophages (Abou-Kheir et al., 2008), and reduced cell spreading has been reported for IRSp53-depleted NIH3T3 cells (Roy et al., 2009). In polarized MDCK cells, IRSp53 is localized to the lateral domain and tight junction region by interacting with the PDZ domain proteins MALS and Lin7 via its C-terminal PDZ-binding domain (Hori et al., 2003). The role of IRSp53 in MDCK polarity has not been studied directly, although it was recently reported that a fusion protein between a PDZ mutant of Lin7 and IRSp53 could rescue the delay in tight junction formation observed in the Lin7 mutant lacking the PDZ domain (Massari et al., 2009). In this study, we report that Par1b overexpression and depletion of its substrate IRSp53 both inhibit basal lamina formation in 3D cultures of MDCK cells and reduced cell spreading and stress fiber formation in 2D cultures. The ECM-signaling phenotype was accompanied by a switch from apical to lateral lumen polarity, suggesting that cell adhesion signaling constitutes the underlying mechanism for the lumen polarity phenotype observed under both experimental conditions.
Results
Par1b inhibits cell spreading, laminin assembly, and stress fiber formation
MDCK cells overexpressing Par1b under a doxycycline (dox)-dependent promoter (Par1b-MDCK Tet-Off; Cohen et al., 2004) form tighter colonies and lateral lumina, whereas Par1b-depleted monolayers are less compact than controls (Fig. 1 A, left, “on plastic”). These observations prompted us to analyze whether Par1b levels affect the kinetics and extent of cell spreading. Dox-regulated expression of a Par1b shRNAmir construct (Par1b-KD-MDCK Tet-Off; see Par1b depletion in Fig. 2 D) promoted cell spreading between 30 and 120 min after plating (Fig. 1 B, left), whereas dox-dependent overexpression of the kinase inhibited spreading on collagen I (Fig. 1 B, right).
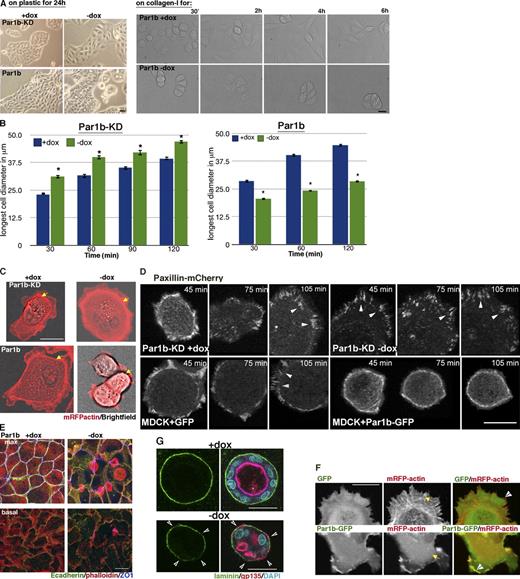
Par1b inhibits cell spreading and stress fiber formation and promotes cell compaction. (A) Phase-contrast images of cell islands cultured on tissue culture plastic for 24 h (left) or brightfield images of a time lapse (Video 6) after cell plating on collagen I (right). Translucent holes in the 24-h Par1b (−dox) culture representing lateral lumina. (B) Spreading on collagen I. *, P < 0.001 for differences between ±dox. Error bars indicate SEM. (C, D, and F) Still images from time-lapse videos 30 (C and F) or 45 min (D) after plating on collagen. (C) Brightfield and mRFP fluorescence (Videos 1 and 2) are shown. Arrows point to stress fibers. (D) Note the dox-dependent size difference in Par1b cells and Paxillin-mCherry (Video 3). Arrowheads point to FAs. (F) GFP or Par1b-GFP and mRFP actin (Video 5) are shown. Note both the difference in stress fibers (arrows) and that Par1b-GFP but not GFP localizes to the tips of actin bundles in lamella (arrowheads). (E) Polarized Par1b cultures 24 h after Ca2+ switch. Phalloidin labels stress fibers at the basal domain (bottom) and microvilli-rich lateral lumina and apical domains in the maximum (max) projections (top; Video 6). (G) Confocal sections through collagen cysts of Par1b-MDCK cells (±dox; see Video 4 for full stacks). Arrowheads point to gaps in laminin. Also note that Par1b cysts are smaller than WT cysts (mean: 23 vs. 30 cells/cyst; n = 13/17). Bars: (A–F) 10 µm; (G) 25 µm.
Par1b inhibits cell spreading and stress fiber formation and promotes cell compaction. (A) Phase-contrast images of cell islands cultured on tissue culture plastic for 24 h (left) or brightfield images of a time lapse (Video 6) after cell plating on collagen I (right). Translucent holes in the 24-h Par1b (−dox) culture representing lateral lumina. (B) Spreading on collagen I. *, P < 0.001 for differences between ±dox. Error bars indicate SEM. (C, D, and F) Still images from time-lapse videos 30 (C and F) or 45 min (D) after plating on collagen. (C) Brightfield and mRFP fluorescence (Videos 1 and 2) are shown. Arrows point to stress fibers. (D) Note the dox-dependent size difference in Par1b cells and Paxillin-mCherry (Video 3). Arrowheads point to FAs. (F) GFP or Par1b-GFP and mRFP actin (Video 5) are shown. Note both the difference in stress fibers (arrows) and that Par1b-GFP but not GFP localizes to the tips of actin bundles in lamella (arrowheads). (E) Polarized Par1b cultures 24 h after Ca2+ switch. Phalloidin labels stress fibers at the basal domain (bottom) and microvilli-rich lateral lumina and apical domains in the maximum (max) projections (top; Video 6). (G) Confocal sections through collagen cysts of Par1b-MDCK cells (±dox; see Video 4 for full stacks). Arrowheads point to gaps in laminin. Also note that Par1b cysts are smaller than WT cysts (mean: 23 vs. 30 cells/cyst; n = 13/17). Bars: (A–F) 10 µm; (G) 25 µm.
Par1b promotes IRSp53 phosphorylation and 14-3-3 binding. (A) IRSp53 in total lysates from Par1b and Par1b-KD cells (1/10 input) and from lysate fractions adsorbed on immobilized 14-3-3τ and competitively eluted. Arrows show a decrease in IRSp53 mobility in the Par1b lysates. (B) In vitro kinase assays of recombinant IRSp53 with Par1b-WT or Par1b-KA followed by λ-phosphatase (+) or mock (m) treatment and subsequent binding to immobilized 14-3-3τ. (top) 1/10 of IRSp53 input is shown. (middle) IB is shown, with IRSp53 competitively eluted from 14-3-3 resin. (bottom) Autorad is shown, with [32P]γATP-Par1b–phosphorylated IRSp53. (C) IRSp53-HA or IRSp53ΔSH3-HA cells transduced with control, Par1b-WT, or Par1b-KA adenovirus. IB of 1/10 Par1b, IB of HA input, and phospho-serine after HA IP are shown. (D) IRSp53-HA immunoprecipitated from [32P]orthophosphate-labeled (autorad) and unlabeled (IB, IRSp53-HA) Par1b-KD cells. Par1b IB shows dox-dependent depletion of the two Par1b splice variants.
Par1b promotes IRSp53 phosphorylation and 14-3-3 binding. (A) IRSp53 in total lysates from Par1b and Par1b-KD cells (1/10 input) and from lysate fractions adsorbed on immobilized 14-3-3τ and competitively eluted. Arrows show a decrease in IRSp53 mobility in the Par1b lysates. (B) In vitro kinase assays of recombinant IRSp53 with Par1b-WT or Par1b-KA followed by λ-phosphatase (+) or mock (m) treatment and subsequent binding to immobilized 14-3-3τ. (top) 1/10 of IRSp53 input is shown. (middle) IB is shown, with IRSp53 competitively eluted from 14-3-3 resin. (bottom) Autorad is shown, with [32P]γATP-Par1b–phosphorylated IRSp53. (C) IRSp53-HA or IRSp53ΔSH3-HA cells transduced with control, Par1b-WT, or Par1b-KA adenovirus. IB of 1/10 Par1b, IB of HA input, and phospho-serine after HA IP are shown. (D) IRSp53-HA immunoprecipitated from [32P]orthophosphate-labeled (autorad) and unlabeled (IB, IRSp53-HA) Par1b-KD cells. Par1b IB shows dox-dependent depletion of the two Par1b splice variants.
Live cell analysis of newly plated Par1b-MDCK or Par1b-KD-MDCK cells cotransfected with mRFP-actin revealed that Par1b overexpression disrupted the continuous lamellipodia skirt surrounding the cell periphery in controls and reduced the appearance of circumferential stress fibers and thin bundles of actin projecting radially away from the cell edge. Instead, individual, large and dynamic lamella emerged from the cell body in Par1b-MDCK cells (Fig. 1, C and F, Par1b; and Videos 1 and 5). Par1b-GFP colocalized with F-actin–rich regions at the tips of those dynamic cell protrusions and filopodia (Fig. 1 F and Video 5). We also performed live cell analysis of the focal adhesion (FA)–associated protein paxillin-mCherry (Schaller, 2001). Freshly plated GFP-expressing cells showed multiple FA sites, apparent as discrete streaks, in the cell periphery where lamellipodia formed (Fig. 1 D, arrowheads, MDCK+GFP; and Video 3, top). In contrast, in Par1b-GFP–expressing cells, discrete paxillin-mCherry–positive FAs were absent, and paxillin-mCherry only appeared as a continuous ring at the cell periphery (Fig. 1 D, MDCK+Par1b-GFP; and Video 3, top right). In contrast, in Par1b-KD MDCK cells, mRFP-actin exhibited increased stress fiber formation in the absence of dox (when Par1b is absent) compared with the control cells maintained in dox (Fig. 1 C, Par1b-KD; and Video 2) and more numerous and stronger paxillin-mCherry–positive FAs that reached further into the cell interior than in the controls (Fig. 1 D, top; and Video 3, bottom). Brightfield time-lapse imaging of Par1b-MDCK cell islands at 15-min intervals over a 6-h period showed contacting cells to spread and flatten at a slower rate compared with controls, but at later time points, they contracted again and balled up into tightly compacted cell clusters (Fig. 1 A, right, “on collagen”; and Video 6). This phenotype is consistent with a scenario in which the balance between cell–cell adhesion (which promotes compaction) and cell–matrix adhesion (which promotes cell spreading), is shifted toward cell–cell adhesion in Par1b-overexpressing cells.
Spreading and actin dynamics in Par1b-KD-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP fluorescence time-lapse series (1 frame/3 s) are shown. Left, +dox; right, −dox. Bar, 20 µm.
Spreading and actin dynamics in Par1b-KD-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP fluorescence time-lapse series (1 frame/3 s) are shown. Left, +dox; right, −dox. Bar, 20 µm.
Paxillin dynamics in T23 (Tet-Off)-MDCK cells expressing either GFP or Par1b-GFP and paxillin-mCherry. (top) mCherry and GFP fluorescence time-lapse series (1 frame/15 min) are shown. (bottom) Paxillin dynamics in Par1b-KD cells cultured in +dox or −dox and expressing paxillin-mCherry. mCherry fluorescence time-lapse series (1 frame/15 min) is shown. Arrowheads point to discrete, dynamic FAs that are apparent in the GFP but not the Par1b-GFP cell.
Paxillin dynamics in T23 (Tet-Off)-MDCK cells expressing either GFP or Par1b-GFP and paxillin-mCherry. (top) mCherry and GFP fluorescence time-lapse series (1 frame/15 min) are shown. (bottom) Paxillin dynamics in Par1b-KD cells cultured in +dox or −dox and expressing paxillin-mCherry. mCherry fluorescence time-lapse series (1 frame/15 min) is shown. Arrowheads point to discrete, dynamic FAs that are apparent in the GFP but not the Par1b-GFP cell.
Actin dynamics of T23 (control)-MDCK cells expressing either GFP or Par1b-GFP and mRFP actin. Top, GFP; bottom, Par1b-GFP. GFP and mRFP fluorescence time-lapse series (1 frame/3 min) are shown. Note that GFP and actin colors are inverted in the top panel for better visualization of the stress fibers. Bars, 10 µm.
Actin dynamics of T23 (control)-MDCK cells expressing either GFP or Par1b-GFP and mRFP actin. Top, GFP; bottom, Par1b-GFP. GFP and mRFP fluorescence time-lapse series (1 frame/3 min) are shown. Note that GFP and actin colors are inverted in the top panel for better visualization of the stress fibers. Bars, 10 µm.
Cell spreading and compaction in Par1b-MDCK monolayers cultured in ±dox. Brightfield time-lapse series (1 frame/15 min) is shown. Left, +dox; right, −dox. Bars, 40 µm.
Cell spreading and compaction in Par1b-MDCK monolayers cultured in ±dox. Brightfield time-lapse series (1 frame/15 min) is shown. Left, +dox; right, −dox. Bars, 40 µm.
Polarized Par1b-MDCK cells organize with a hepatic polarity phenotype in which the luminal domain interrupts the lateral domain of neighboring cells rather than being established at the apex (Fig. 1 A, Par1b, −dox, “on plastic”). We noticed that Par1b-MDCK with lateral lumina (Fig. 1 E, max projection −dox) lacked stress fibers at the basal domain (Fig. 1 E, −dox; and Video 7, top). Thus, defects in cell–matrix signaling were apparent not only in freshly plated cells but also in polarized Par1b monolayers.
Phalloidin staining in Par1b-MDCK cells ±dox and IRSp53KD cells ±dox. (top) Phalloidin staining in Par1b-MDCK cells ±dox. (bottom) Phalloidin staining in IRSp53KD cells ±dox. Confocal x-y stacks with 0.3-µm step size are shown.
Phalloidin staining in Par1b-MDCK cells ±dox and IRSp53KD cells ±dox. (top) Phalloidin staining in Par1b-MDCK cells ±dox. (bottom) Phalloidin staining in IRSp53KD cells ±dox. Confocal x-y stacks with 0.3-µm step size are shown.
When cultured in a 3D collagen I matrix, Par1b-MDCK cells not only formed multiple lumina rather then the central cavity observed in control cells but also exhibited disrupted laminin staining at the cyst circumference (Fig. 1 G; Fig. S1; and Video 4, top), suggesting defects in basal lamina formation. Cyst patches lacking laminin frequently showed inverted apical surface staining (Fig. 1 G; Fig. S1; and Video 4, top), a phenotype previously reported in MDCK cysts expressing dominant-negative Rac1 (O’Brien et al., 2001). We conclude that Par1b not only functions at the lateral domain to regulate cell–cell adhesion but is also involved in actin organization and FA assembly associated with cell spreading and regulates the assembly of the epithelial basal lamina.
Confocal x-y series through collagen I cysts of Par1b-MDCK cells and IRSp53-KD cells cultured in ±dox. (top) Confocal x-y series (0.76-µm steps) through collagen I cysts of Par1b-MDCK cells cultured in +dox and −dox. (bottom) Confocal x-y series (1-µm steps) through collagen I cysts of IRSp53KD-MDCK cells cultured in +dox and −dox. Red, gp135; green, laminin; blue, DAPI. Bars, 25 µm.
Confocal x-y series through collagen I cysts of Par1b-MDCK cells and IRSp53-KD cells cultured in ±dox. (top) Confocal x-y series (0.76-µm steps) through collagen I cysts of Par1b-MDCK cells cultured in +dox and −dox. (bottom) Confocal x-y series (1-µm steps) through collagen I cysts of IRSp53KD-MDCK cells cultured in +dox and −dox. Red, gp135; green, laminin; blue, DAPI. Bars, 25 µm.
Par1b phosphorylates IRSp53 in vitro and in vivo and promotes its 14-3-3 binding
Because none of the known Par1 substrates have been implicated in ECM signaling, we initiated an unbiased substrate screen based on the observation that Par1 phosphorylation promotes 14-3-3 binding in some of its substrates (Matenia and Mandelkow, 2009). We compared the 14-3-3 binding profile of membrane/cytoskeletal fractions prepared from MDCK cells overexpressing Par1b wild type (WT) to that of cells expressing a kinase-deficient mutant, Par1b-K49A (Elbert et al., 2006), referred to as Par1b-KA, and we identified IRSp53 as a phospho-protein enriched in the Par1b-WT sample (Fig. S2). Immunoblotting with IRSp53 antibodies confirmed increased IRSp53 binding from MDCK lysates to immobilized 14-3-3τ in cells that overexpressed Par1b in a dox-regulated manner (Fig. 2 A, control Par1b pair). On the other hand, dox-regulated Par1b KD (Fig. 2 A, control Par1b-KD pair) revealed reduced IRSp53 binding to 14-3-3 in Par1b-depleted MDCK cells. We also noticed a small but reproducible reduction in IRSp53 mobility upon Par1b overexpression, which is consistent with putative Par1b-dependent phosphorylation of IRSp53 (Fig. 2 A, arrow). Indeed, expression of Par1b-WT but not of Par1b-KA induced serine phosphorylation in recombinant IRSp53 and in a mutant of IRSp53 lacking the SH3 domain (Fig. 2 C). In contrast, Par1b depletion reduced the incorporation of [32P]orthophosphate into IRSp53 by ∼50% (by 70 and 40% in two separate experiments; Fig. 2 D). Importantly, recombinant IRSp53 was phosphorylated by Par1b in vitro (Fig. 2 B, [32P]ATP) and bound to 14-3-3 in a Par1b phosphorylation–dependent manner (Fig. 2 B, 14-3-3). 14-3-3 binding was abolished upon λ-phosphatase treatment.
Par1b mediates phosphorylation on the IRSp53 SH3 domain flanking serines S366 and S453/4/5
In vitro kinase assays using fragments of IRSp53 as substrates and subsequent in vitro mutagenesis of serines in candidate 14-3-3 binding motifs present in the Par1b-phosphorylated C-terminal IRSp53 fragment (i.e., S325, S365, S366, S453-5, S492, and S511; not depicted) revealed S366 as the main Par1b phosphorylation site in IRSp53 (Fig. 3, A [IRSp53 domain organization] and B). In contrast, the serine cluster S453,454,455 emerged as the major Par1b-dependent IRSp53 site when we probed IRSp53 mutants immunoprecipitated from cell lysates with a phospho-serine antibody (Fig. 3 C). Mutation of all four serines to alanines (IRSp53A4) completely abolished Par1b-mediated phosphorylation both in vitro and in vivo (Fig. 3, B and C). Thus, we conducted most of our subsequent functional studies comparing IRSp53-WT with the IRSp53-A4 mutant. The discrepancy between the Par1b-dependent in vitro and in vivo phosphorylation sites raised the question whether Par1b indeed phosphorylates IRSp53 in cells or, rather, activates an IRSp53 kinase in vivo. Although there are currently no biochemical approaches to directly measure phosphorylation events in intact cells, we adapted a previously developed strategy (Blethrow et al., 2008) that allowed us to assess whether Par1b phosphorylates IRSp53 after cells had been detergent permeabilized, a prerequisite for the entry of ATP analogues. We expressed in MDCK cells an engineered Par1b kinase, Par1b-ATP* with an enlarged ATP-binding pocket, to identify specific kinase substrates by offering the bulky ATPγS analogue N6-benzyl-ATP[35γS] that can only be used by the recombinant kinase but not by endogenous kinases also present in the lysate. To validate the approach, we first established that the “gatekeeper-modified” kinase Par1b-ATP* but not Par1b-WT transferred [35S]thio phosphate from N6-benzyl-ATP[35γS] onto recombinant full-length IRSp53 when the ATPγS analogue was used in in vitro kinase assays with recombinant proteins. As with the C-terminal fragment, the S366A but not the S453-5A substitution abolished the specific in vitro phosphorylation of full-length IRSp53 (Fig. 3 D). Importantly, Par1b-ATP* but not unmodified Par1b also transferred [35S]thio phosphate from N6-benzyl-ATP[35γS] onto full-length IRSp53 when the ATPγS analogue was added to whole cell lysates prepared from cells expressing Par1b-ATP* and the IRSp53 proteins, indicating that Par1b directly phosphorylates IRSp53 in this context (Fig. 3 E). As in assays with the isolated proteins, IRSp53S366A and IRSp53A4 were not phosphorylated by Par1b in whole cell lysates, although IRSp53S453-5A was, indicating that residue S366 is phosphorylated by Par1b, whereas the S453/4/5 phosphorylation occurs either via a Par1b-dependent kinase or is regulated by a Par1b-dependent serine phosphatase (Fig. 3 E). In addition, we ruled out that serines S117 and S148, present in two predicted low stringency 14-3-3 binding motifs in the IRSp53 N-terminal domain (Mackie and Aitken, 2005), are phosphorylated by Par1b (Fig. 3 E).
Par1 phosphorylation on S366 and Par1b-depedendent S453/4/5 phosphorylation promote 14-3-3 binding of IRSp53. (A) Diagram of Par1b-dependent phosphorylation sites in IRSp53. (B) In vitro kinase assays with WT and Par1b-KA using [32P]γATP and IRSp53-CT (WT and point mutants) as substrates. (C) IRSp53-HA–transfected cells (WT, A4, and S453-5A) were transduced with control (c) or Par1b virus, and the HA IPs were probed for P-serine and subsequently for HA. (D) In vitro kinase assays with WT and Par1b-ATP* using [35S]benzyl-ATPγS and full-length IRSp53 WT and mutants as substrates. Autoradiographs/Coomassie stain of IRSp53. (E) Triton X-100 lysates of cells coexpressing Par1b-WT or Par1b-ATP* and WT IRSp53-Myc or IRSp53 mutants were incubated with [35S]benzyl-ATPγS. Autoradiographs and IB for IRSp53-Myc and Par1b. (F) Recombinant WT IRSp53 and mutant proteins subjected to in vitro kinase assays with Par1b-WT or Par1b-KA were affinity isolated on 14-3-3 beads and competitively eluted (top). IRSp53 input (middle) and Par1b levels (bottom) are also shown. (G) 14-3-3 binding of IRSp53 proteins from cell lysates. Note the lower expression of the S117A sample accounts for the weaker signal in the bound fraction. The graph shows SEM from six experiments (three experiments for S117A/S148A). (H) Co-IP of 14-3-3-HA with IRSp53-Myc proteins from cell lysates.
Par1 phosphorylation on S366 and Par1b-depedendent S453/4/5 phosphorylation promote 14-3-3 binding of IRSp53. (A) Diagram of Par1b-dependent phosphorylation sites in IRSp53. (B) In vitro kinase assays with WT and Par1b-KA using [32P]γATP and IRSp53-CT (WT and point mutants) as substrates. (C) IRSp53-HA–transfected cells (WT, A4, and S453-5A) were transduced with control (c) or Par1b virus, and the HA IPs were probed for P-serine and subsequently for HA. (D) In vitro kinase assays with WT and Par1b-ATP* using [35S]benzyl-ATPγS and full-length IRSp53 WT and mutants as substrates. Autoradiographs/Coomassie stain of IRSp53. (E) Triton X-100 lysates of cells coexpressing Par1b-WT or Par1b-ATP* and WT IRSp53-Myc or IRSp53 mutants were incubated with [35S]benzyl-ATPγS. Autoradiographs and IB for IRSp53-Myc and Par1b. (F) Recombinant WT IRSp53 and mutant proteins subjected to in vitro kinase assays with Par1b-WT or Par1b-KA were affinity isolated on 14-3-3 beads and competitively eluted (top). IRSp53 input (middle) and Par1b levels (bottom) are also shown. (G) 14-3-3 binding of IRSp53 proteins from cell lysates. Note the lower expression of the S117A sample accounts for the weaker signal in the bound fraction. The graph shows SEM from six experiments (three experiments for S117A/S148A). (H) Co-IP of 14-3-3-HA with IRSp53-Myc proteins from cell lysates.
Thus, our combined approaches allowed us to identify S366 on IRSp53 as a direct Par1b substrate and the serine cluster S453-455 as a target of a Par1b-dependent kinase or phosphatase. IRSp53 and Par1b did not form stable complexes in cell lysates (unpublished data), suggesting that IRSp53 phosphorylation either does not take place within a Par1b-containing scaffolding complex but, rather, in a kiss and run fashion or occurs within the detergent insoluble cytoskeletal fraction that is unavailable for coimmunoprecipitation (co-IP) analysis.
We next determined whether the Par1b-dependent phosphorylation sites S366 and S453/454/545 are required for 14-3-3 binding of IRSp53. The 14-3-3 binding efficiency of the IRSp53 mutants S366A, S453-5A, and A4 corresponded to the extent of their in vitro Par1b phosphorylation: IRSp53-S366A showed reduced 14-3-3 binding compared with the WT protein, whereas the A4 mutant did not bind 14-3-3 at all in this direct binding assay (Fig. 3 F, top, lanes 1–4). Similar results were obtained when recombinant WT, S366A, and A4 IRSp53 proteins were expressed in MDCK cells and the lysates incubated with immobilized 14-3-3 (Fig. 3 G). The S453-5A mutation also reduced 14-3-3 binding of IRSp53 when isolated from MDCK lysates, whereas the S117A and S148A mutations were without effect (Fig. 3 G). Lastly, we coimmunoprecipitated recombinant IRSp53-WT but not IRSp53-A4 with recombinant 14-3-3τ from MDCK lysates (Fig. 3 H). A phosphomimetic aspartate mutant, IRSp53-D4, was also unable to interact with 14-3-3 in either assay (Fig. 3 H and not depicted). This is similar to other cases of 14-3-3 binding where phosphomimetics were ineffective (Johnson et al., 2010). Together, these data indicate that Par1b-mediated phosphorylation of IRSp53 is directly responsible for its 14-3-3 binding in MDCK cells.
While preparing this manuscript, Robens et al. (2010) reported residues T340 and T360 in IRSp53 as essential for IRSp53–14-3-3 interactions in Cos-7 cell lysates. However, when expressed in MDCK cells, a T360A IRSp53 mutant interacted with 14-3-3 as efficiently as it did the WT protein (Fig. 3 G), suggesting that cell type–specific kinase activities might generate several distinct 14-3-3 binding sites centered around the SH3 domain of IRSp53. Alternatively, the different sites might recruit different 14-3-3 isoforms. However, we found that Par1b-phosphorylated IRSp53 interacts with all three 14-3-3 isoforms we tested: τ, ζ, and σ (Fig. S3).
IRSp53 depletion in MDCK cells mimics aspects of the Par1b overexpression phenotype
To determine whether IRSp53 is a Par1b substrate involved in MDCK cell polarization and lumen organization, we first analyzed the phenotype of MDCK clones that feature dox-regulated depletion of IRSp53 by expressing an IRSp53-directed shRNAmir under an inducible promoter (Fig. 4 A). The target region was based on a previously published IRSp53-directed RNAi sequence (Shi et al., 2005). IRSp53 depletion yielded the hepatic polarity phenotype characteristic of Par1b-overexpressing MDCK monolayers (Fig. 4 C, top, “max”; compare with Fig. 1 E) although in fewer cells (38% of cells vs. 70% in Par1b clones 24 h after polarity was initiated) and not as robustly because the number of IRSp53-depleted cells with hepatic-type polarity decreased to 21% 48 h after the initiation of cell–cell adhesion (see Fig. 7). In 3D collagen cultures, IRSp53-KD caused multiple lumina as seen in Par1b-MDCK cells (Fig. 4 G; Fig. S1; and Video 4, bottom). These data suggest that, although not sufficient to account for the Par1b polarity phenotype, inhibition of IRSp53 signaling contributes to Par1b-induced hepatic-type lumen organization.
IRSp53 depletion mimics aspects of the Par1b overexpression phenotype. (A) Dox-inducible IRSp53 depletion is shown. (B) Brightfield time lapse of IRSp53-KD cell islands on collagen I (Video 10). (C) Polarized cultures 24 h after Ca2+ switch. Phalloidin labels stress fibers at the basal domain (bottom) and microvilli-rich lateral lumina and apical domains in the maximum (max) projections (top; see Video 7). (D and F) Still images from time-lapse videos taken 30 min after plating on collagen. (D) Brightfield and mRFP fluorescence (Video 8). (F) Paxillin-mCherry (Video 9, top) is shown. Arrowheads point to FAs. (E) Spreading of two IRSp53-KD clones on collagen I in DME and SMEM; all differences between ±dox are significant with P < 0.001. Error bars indicate SEM. (G) Confocal section through collagen cysts (see Video 4 for full stacks) annotated as in Fig. 1 G. IRSp53KD cysts are smaller than control cysts (14 vs. 25 cells/cyst; n = 31). Bars: (B–D and F) 10 µm; (G) 25 µm.
IRSp53 depletion mimics aspects of the Par1b overexpression phenotype. (A) Dox-inducible IRSp53 depletion is shown. (B) Brightfield time lapse of IRSp53-KD cell islands on collagen I (Video 10). (C) Polarized cultures 24 h after Ca2+ switch. Phalloidin labels stress fibers at the basal domain (bottom) and microvilli-rich lateral lumina and apical domains in the maximum (max) projections (top; see Video 7). (D and F) Still images from time-lapse videos taken 30 min after plating on collagen. (D) Brightfield and mRFP fluorescence (Video 8). (F) Paxillin-mCherry (Video 9, top) is shown. Arrowheads point to FAs. (E) Spreading of two IRSp53-KD clones on collagen I in DME and SMEM; all differences between ±dox are significant with P < 0.001. Error bars indicate SEM. (G) Confocal section through collagen cysts (see Video 4 for full stacks) annotated as in Fig. 1 G. IRSp53KD cysts are smaller than control cysts (14 vs. 25 cells/cyst; n = 31). Bars: (B–D and F) 10 µm; (G) 25 µm.
The localization of both Par1b and IRSp53 to the lateral membrane in polarized MDCK cells and to peripheral actin structures in freshly plated cells (Fig. 1 F and not depicted; Böhm et al., 1997; Hori et al., 2003) suggested that IRSp53 could either regulate cell–cell or cell–ECM signaling downstream of Par1b.
Because we had previously implicated cell–cell adhesion as a target of Par1b signaling (Cohen et al., 2007), we first tested whether IRSp53 depletion altered E-cadherin–mediated adhesion and tight junction formation in Ca2+ switch assays (Gonzalez-Mariscal et al., 1990). As shown in Fig. S4 (A and B), IRSp53 depletion did not alter the kinetics of E-cadherin recruitment or its incorporation into the cortical actin cytoskeleton. Although the formation of tight junctions, as indicated by ZO-1 recruitment to cell–cell contacts sites, appeared to be delayed 3 h after Ca switch in IRSp53 depleted cells, it was comparable with control conditions at the 8- and 24-h time points (Fig. S4 C).
However, we found that IRSp53 depletion reduced spreading of freshly plated MDCK cells on collagen I substrate and prevented the complete circumferential lamella skirt and circumferential stress fibers seen in control cells between 30 and 120 min after plating (Fig. 4 D and Video 7) similarly to our observations with Par1b overexpression (Fig. 1, B and C, Par1b). Likewise, paxillin-mCherry–positive FAs were largely absent in IRSp53-depleted cells (Fig. 4 F; and Video 9, top). A quantitative analysis presented in Fig. 4 E shows spreading defects in two inducible IRSp53-shRNAmir clones up to 120 min upon plating. Spreading defects were similar in cultures maintained in low-Ca2+ medium, indicating that they are independent of cell–cell adhesion (Fig. 4 E, Spinner Modified Eagle’s medium [SMEM]). Consistent with IRSp53 regulating cell spreading but not cell–cell adhesion, we also observed an extreme compaction of freshly plated IRSp53-depleted cell islands when imaged over a 6-h period (Fig. 4 B [4–6 h]; and Video 10). Again, this phenomenon mimicked the effect of Par1b overexpression (compare with Fig. 1 A, “on collagen”; Video 6). In addition, we found that, like Par1b-MDCK cells, IRSp53-depleted cells with lateral lumina lacked stress fibers at the basal domain (compare Fig. 5 C; and Video 7, bottom; with Fig. 1 E; and Video 7, top). Lastly, the IRSp53-KD cysts featured drastically reduced and fragmented laminin staining, indicating defects in basal lamina assembly (Fig. 4 G; Fig. S1; and Video 4, bottom) Collectively, our findings indicate that Par1b and IRSp53 have opposing roles in cell–matrix signaling that governs cell spreading, FA formation, and the organization of the actin cytoskeleton at the basal surface and basement membrane organization.
Paxillin dynamics in IRSp53-KD cells cultured in ±dox and expressing paxilin-mCherry. (top) mCherry fluorescence time-lapse series (1 frame/15 min) is shown. (bottom) Paxillin dynamics in IRSp53-WT clone #1 (left) or IRSp53-A4 clone #5 (right) expressing Par1b-GFP and paxillin-mCherry. mCherry and GFP fluorescence time-lapse series (1 frame/15 min) is shown.
Paxillin dynamics in IRSp53-KD cells cultured in ±dox and expressing paxilin-mCherry. (top) mCherry fluorescence time-lapse series (1 frame/15 min) is shown. (bottom) Paxillin dynamics in IRSp53-WT clone #1 (left) or IRSp53-A4 clone #5 (right) expressing Par1b-GFP and paxillin-mCherry. mCherry and GFP fluorescence time-lapse series (1 frame/15 min) is shown.
Cell spreading and compaction in IRSp53-KD-MDCK monolayers cultured in ±dox. Brightfield time-lapse series (1 frame/10 min). Left, +dox; right, −dox. Bars, 20 µm.
Cell spreading and compaction in IRSp53-KD-MDCK monolayers cultured in ±dox. Brightfield time-lapse series (1 frame/10 min). Left, +dox; right, −dox. Bars, 20 µm.
A nonphosphorylatable IRSp53 mutant rescues the Par1b phenotype. (A) Parental MDCK cells and five clones expressing RNAi-resistant IRSp53-A4 and four clones of IRSp53-WT cells were transiently transfected with mock or IRSp53shmir cDNA and analyzed for cell spreading 2 h after plating. (B) 2-h spreading in parental cells, eight IRSp53AA, and five IRSp53WT clones that were transduced with control or Par1b adenovirus. (C) Paxillin-mCherry in WT clone #1 and A4 clone #5 expressing Par1b-GFP 30 min after plating on collagen I. Arrowheads point to FAs (Video 9, bottom). (D) Par1b-MDCK cells transfected with mock or IRSp53 cDNAs (WT, A4, DD, and ΔSH3) were quantitatively analyzed for lumen polarity as described in Materials and methods. *, P < 0.0001 with difference to parental or mock-transfected cells. Error bars indicate SEM. Bars: (A and B) 30 µm; (C and D) 10 µm.
A nonphosphorylatable IRSp53 mutant rescues the Par1b phenotype. (A) Parental MDCK cells and five clones expressing RNAi-resistant IRSp53-A4 and four clones of IRSp53-WT cells were transiently transfected with mock or IRSp53shmir cDNA and analyzed for cell spreading 2 h after plating. (B) 2-h spreading in parental cells, eight IRSp53AA, and five IRSp53WT clones that were transduced with control or Par1b adenovirus. (C) Paxillin-mCherry in WT clone #1 and A4 clone #5 expressing Par1b-GFP 30 min after plating on collagen I. Arrowheads point to FAs (Video 9, bottom). (D) Par1b-MDCK cells transfected with mock or IRSp53 cDNAs (WT, A4, DD, and ΔSH3) were quantitatively analyzed for lumen polarity as described in Materials and methods. *, P < 0.0001 with difference to parental or mock-transfected cells. Error bars indicate SEM. Bars: (A and B) 30 µm; (C and D) 10 µm.
IRSp53-A4 rescues the Par1b overexpression phenotypes
Thus far, our data suggest that Par1b-mediated phosphorylation of IRSp53 inhibits its function and is responsible for some of the Par1b-induced morphological and polarity phenotypes in MDCK cells. To test this hypothesis, we asked whether expression of IRSp53-WT or its phosphorylation-deficient mutant IRSp53-A4 could alleviate Par1b-induced spreading defects and antagonize lateral lumen polarity.
We first determined whether KD-resistant IRSp53-WT and IRSp53-A4 were able to rescue the IRSp53-KD–induced spreading defect. We generated stable cell lines expressing near-endogenous levels of recombinant IRSp53 constructs under the weak thymidine kinase promoter (Fig. S5 C). Into those clones, we transiently transfected the shIRSp53mir-cDNA to reduce endogenous IRSp53 levels. Real-time PCR with canine-specific primers indicated a KD efficiency between 43 and 65% for different clones (Fig. S5 B), resulting in a more modest decrease of spreading compared with the stable IRSp53-KD clones (Fig. 4 A, immunoblot [IB]; compare parental mock with IRSp53-KD mRNA in Fig. S5 B). All five IRSp53-A4 clones and all four IRSp53-WT clones rescued the spreading defects and, to varying degrees, showed increased spreading compared with the parental cells (Fig. 5 A). None of the clones displayed a lateral polarity phenotype (unpublished data). These findings verify that the IRSp53 shRNAmir phenotype is not caused by an off-target effect.
Next, we transduced the IRSp53-WT and IRSp53-A4 clones with recombinant adenoviruses expressing either l-Cat (controls) or Par1b. The Par1b-induced spreading defect was rescued in six of the eight tested IRSp53-A4 clones, i.e., the mean cell diameter of the Par1b-transduced clones was with statistical significance different from that of parental Par1b-transduced cells but not from parental cells transduced with the control virus. In contrast, only one of the five IRSp53-WT clones showed a statistically significant increase in mean cell diameter over the parental cells when transduced with Par1b virus (Fig. 5 B). Likewise, when we cotransfected two IRSp53-WT and two IRSp53-A4 clones with Par1b-GFP and paxillin-mCherry, we observed extensive FAs only in the IRSp53-A4 but not the WT clones (Fig. 5 C and Video 9, bottom, WT clone #1 and A4 clone #5). Thus, we conclude that IRSp53-A4 is more efficient than IRSp53-WT in antagonizing the Par1b-induced spreading defect in MDCK cells.
When we tested the effect of IRSp53-WT and IRSp53-A4 on Par1b-induced lateral lumen formation, we observed similarly that the A4 mutant was far more effective than IRSp53-WT in reverting the lateral lumen polarity phenotype, yielding apical luminal domains (Fig. 5 D). Importantly, an SH3-deficient IRSp53 mutant was without any effect, indicating that the IRSp53 effector domain is important for the rescue. In these experiments, we used our Par1b cell line and transiently transfected the cells with recombinant IRSp53 constructs. Using Par1b virus transduction in the aforementioned IRSp53 clones yielded similar results (unpublished data). Furthermore, we determined that reducing IRSp53 levels and expressing low levels of recombinant Par1b promoted lateral lumina in a synergistic fashion (Fig. 6). A “leaky” IRSp53KD clone showed an ∼50% reduction of IRSp53 in +dox (Fig. 6 A) and exhibited few lateral lumina when transduced with a control virus (Fig. 6 B, IRSp53KD+dox/control virus). Par1b adenovirus transduction at levels that caused few lateral lumina in parental MDCK cells (Fig. 6 B, parental+dox/Par1b virus), however, featured close to 45% lateral lumen polarity in IRSp53KD+dox monolayers (Fig. 6 B, IRSp53KD+dox/Par1b virus).
Decreased IRSp53 and increased Par1b levels have a synergistic effect on lateral lumen formation. (A) IRSp53 expression in parental cells and an IRSp53KD clone in ±dox and after adenovirus transduction as indicated. (B) Lateral lumen polarity analyzed as described in Materials and methods. Error bars indicate SEM.
Decreased IRSp53 and increased Par1b levels have a synergistic effect on lateral lumen formation. (A) IRSp53 expression in parental cells and an IRSp53KD clone in ±dox and after adenovirus transduction as indicated. (B) Lateral lumen polarity analyzed as described in Materials and methods. Error bars indicate SEM.
Collectively, the Par1b phosphorylation-deficient IRSp53 mutant A4 was able to antagonize the effect of Par1b overexpression in cell spreading and lumen formation in MDCK cells while IRSp53-WT was, in each case, either less effective or ineffective in overcoming the Par1b phenotype. We conclude that Par1b phosphorylation inhibits IRSp53 to yield cytoskeletal defects that underlie the Par1b phenotype.
Par1b-induced inhibition of IRSp53 and myosin promote hepatic-type lumen polarity in a synergistic fashion
We had previously reported that Par1b-mediated inhibition of myosin II contributes to the hepatic polarity phenotype in Par1b-MDCK cells (Cohen et al., 2007). The myosin II–specific inhibitor blebbistatin (BB) promoted lateral lumina during polarization, but only transiently. To determine whether myosin II and IRSp53 function synergistically to yield the Par1b polarity phenotype, we compared the effect of IRSp53 depletion on lumen organization 24 and 48 h after the initiation of cell–cell contacts either in the presence or absence of BB. Although neither BB nor IRSp53 depletion alone matched the extent of lateral lumen formation found in Par1b-MDCK cells, in combination, they caused 66% of the cells to adopt hepatic-type polarity at the 24-h time point, although significantly fewer lumina were observed at 48 h (Fig. 7). Thus, Par1b signaling presumably involves still additional factors that account for the robustness of the lumen phenotype once polarity is fully established. We conclude that myosin II and IRSp53 inhibition by Par1b contribute to lumen organization in a synergistic fashion, most likely operating in different signaling pathways.
IRSp53 and myosin II function synergistically to regulate epithelial lumen polarity. IRSp53KD (−dox) and control (+dox) subjected to Ca2+ switch assays for 24 and 48 h in the presence or absence of 50 µM BB. Lumen polarity was quantified as described in Materials and methods. Statistically significant difference is shown with P < 0.0001 (*). Images represent the 24-h time point. Error bars indicate SEM. Bars, 10 µm.
IRSp53 and myosin II function synergistically to regulate epithelial lumen polarity. IRSp53KD (−dox) and control (+dox) subjected to Ca2+ switch assays for 24 and 48 h in the presence or absence of 50 µM BB. Lumen polarity was quantified as described in Materials and methods. Statistically significant difference is shown with P < 0.0001 (*). Images represent the 24-h time point. Error bars indicate SEM. Bars, 10 µm.
Par1b-mediated 14-3-3 binding inhibits protein recruitment to the IRSp53 C-terminal domain
Our observation that IRSp53’s role in lumen polarity downstream of Par1b is dependent on its SH3 domain (Fig. 5 D) prompted us to test whether Par1b-induced 14-3-3 binding interferes with the recruitment of IRSp53 effector proteins to its SH3 domain. Indeed, recombinant IRSp53-A4 coimmunoprecipitated slightly more recombinant WAVE2 than IRSp53-WT. This difference was even more pronounced in the presence of recombinant Par1b (Fig. 8 A), suggesting that Par1b phosphorylation interferes with WAVE–IRSp53 binding. In contrast, no difference was apparent in the interaction of WT and A4-IRSp53 with activated Cdc42 (Q61L; Fig. 8 B). We performed these experiments in 293 cells because coexpression of WAVE2 and IRSp53 caused WAVE2 aggregates in MDCK cells.
Par1b phosphorylation inhibits the recruitment of WAVE2 and other IRSp53-binding proteins to its C-terminal, SH3-containing domain. (A and B) WAVE2 co-IPs with IRSp53 in 293 cells expressing WAVE2-GFP, Myc-tagged IRSp53 constructs (WT, A4, or IRSp53 Δ317–521; ΔCT), and Par1b where indicated. 1/10 of the lysate was analyzed for WAVE and Par1b expression. (B) Data are from three experiments with and without recombinant Par1b and three experiments without recombinant Par1b only. Error bars indicate SEM. (C) Cdc42Q61L co-IPs with IRSp53-Myc constructs from 293 cells. Arrows indicate Cdc42, and asterisks indicate IgG H and L chains. Data are representative of three experiments. (D–F) GST-IRSp53CT was subjected to Par1b phosphorylation, immobilized on glutathione and incubated with 14-3-3τ as indicated. (D) Experimental schematic is shown. (E) Binding of proteins from [35S]Met/Cys-labeled 293 lysates to immobilized IRSp53CT was analyzed by Coomassie and autoradiography. Arrows indicate specific bands that are reduced in the Par1b sample. Asterisk indicates cellular 14-3-3, with one out of two identical results represented. (F) 14-3-3 binding to IRSp53CT was analyzed by IB, and IRSp53-CT levels by Ponceau red stain are shown.
Par1b phosphorylation inhibits the recruitment of WAVE2 and other IRSp53-binding proteins to its C-terminal, SH3-containing domain. (A and B) WAVE2 co-IPs with IRSp53 in 293 cells expressing WAVE2-GFP, Myc-tagged IRSp53 constructs (WT, A4, or IRSp53 Δ317–521; ΔCT), and Par1b where indicated. 1/10 of the lysate was analyzed for WAVE and Par1b expression. (B) Data are from three experiments with and without recombinant Par1b and three experiments without recombinant Par1b only. Error bars indicate SEM. (C) Cdc42Q61L co-IPs with IRSp53-Myc constructs from 293 cells. Arrows indicate Cdc42, and asterisks indicate IgG H and L chains. Data are representative of three experiments. (D–F) GST-IRSp53CT was subjected to Par1b phosphorylation, immobilized on glutathione and incubated with 14-3-3τ as indicated. (D) Experimental schematic is shown. (E) Binding of proteins from [35S]Met/Cys-labeled 293 lysates to immobilized IRSp53CT was analyzed by Coomassie and autoradiography. Arrows indicate specific bands that are reduced in the Par1b sample. Asterisk indicates cellular 14-3-3, with one out of two identical results represented. (F) 14-3-3 binding to IRSp53CT was analyzed by IB, and IRSp53-CT levels by Ponceau red stain are shown.
We next assessed the binding of proteins from lysates of [35S]Met metabolically labeled cells to immobilized GST-IRSp53-CT, the IRSp53 fragment that includes the SH3 domain and the Par1b phosphorylation sites (Fig. 2 A). When in vitro phosphorylated by Par1b and subsequently incubated with recombinant 14-3-3, GST-IRSp53-CT recruited at least six of the specifically binding proteins to a lesser extent than GST-IRSp53-CT that had been mock phosphorylated and that did not bind 14-3-3 (Fig. 8, C and D). Although the differences were modest (between 25 and 30%), they were expected to be so because only a fraction of IRSp53-CT becomes phosphorylated and binds 14-3-3 in vitro (Fig. 8 D).
Together, these data indicate that Par1b phosphorylation inhibits the recruitment of IRSp53-binding proteins to the SH3 domain and potentially other domains within the IRSp53 C terminus. The inhibition of multiple protein interactions is consistent with a steric hindrance caused by 14-3-3 binding rather than the regulation of a specific protein–protein interaction by phosphorylation.
Discussion
The role of Par1b in epithelial cells has been associated with its localization and activity at the lateral membrane and, specifically, with E-cadherin function. In this study, we show that Par1b also regulates cell spreading, FA formation, the organization of the actin cytoskeleton at the basal surface, and laminin deposition, consistent with a role in cell–matrix signaling. We identified IRSp53 as a Par1b substrate mediating its role in basal actin organization in epithelial cells. This conclusion is based on the following findings: (a) Par1b directly phosphorylates IRSp53 in vitro and in cell lysates at S366 and, in addition, stimulates IRSp53 phosphorylation at S453/4/5 indirectly in vivo, whereas Par1b depletion reduces IRSp53 phosphorylation in intact cells. (b) IRSp53 KD mimics all of the observed effects of Par1b overexpression on cell substrate signaling. (c) Importantly, a Par1b phosphorylation–deficient IRSp53 mutant rescued the spreading defect of Par1b-overexpressing cells, whereas expression of WT-IRSp53 was ineffective. These data are consistent with a scenario in which Par1b phosphorylation inhibits IRSp53 function. Fig. 9 presents our model for the mechanism of this inhibitory effect: (a) Par1b phosphorylation creates 14-3-3 binding sites proximal and distal to the IRSp53 SH3 effector domain. Both Par1b phosphorylation sites contributed to effective binding of IRSp53 from MDCK lysates to 14-3-3 and are located within sequences that conform with low stringency to a class I 14-3-3 binding consensus motif according to Scan Site analysis (http://scansite.mit.edu/motifscan_seq.phtml). Cooperative binding to multiple low-stringency sites has been shown for some 14-3-3 interactions (Dougherty and Morrison, 2004). It is noteworthy in this respect that the N-terminal helical domain (aa 1–260) of IRSp53 mediates its dimerization and might thereby contribute to 14-3-3 binding as previously suggested (Mackie and Aitken, 2005) by increasing the concentration of low-affinity binding sites. 14-3-3 molecules are obligate dimers that interact with two binding sites in either the same protein or by scaffolding two molecules (Johnson et al., 2010). (b) Recruitment of 14-3-3 masks the SH3 domain for binding of effector proteins that mediate actin polymerization downstream of rho-GTPases. WT-IRSp53, but not the phosphorylation-resistant mutant A4, showed reduced interaction with WAVE2 when Par1b was coexpressed. The inhibitory effect is not limited to WAVE2, as six unidentified proteins showed reduced in vitro binding to the IRSp53 C-terminal domain when preceded by Par1b-dependent 14-3-3 recruitment. Our model is similar to that recently proposed by Robens et al. (2010), who identified T340/T360 close to the Par1b site at S366, as important for 14-3-3 binding and inhibitory for effector recruitment to IRSp53 in Cos7 cells. Although the authors did not identify the responsible kinase, it is conceivable that in different cell types, distinct kinases regulate 14-3-3-binding in the vicinity of the SH3 domain to inhibit IRSp53’s scaffolding function.
Model for Par1b regulation of IRSp53 in epithelial lumen formation. Activated Rac1 or Cdc42 recruit Rho-GTPase effector proteins such as Mena, WAVE, mDia, and Eps8 involved in microfilament organization to the IRSp53 SH3 domain. Phosphorylation of S366 by Par1b and Par1b stimulation of S434-5 phosphorylation by an unidentified kinase recruits 14-3-3 proteins that interfere with WAVE2 binding and the recruitment of other proteins that interact with the IRSp53 C-terminal domain. Par1b overexpression inhibits IRSp53-dependent actin dynamics that regulate cell–ECM signaling and result in altered lumen polarity in MDCK cells. In other cell types, such as Cos7, unidentified kinases inhibit IRSp53 function by creating 14-3-3 binding sites on T340/T360.
Model for Par1b regulation of IRSp53 in epithelial lumen formation. Activated Rac1 or Cdc42 recruit Rho-GTPase effector proteins such as Mena, WAVE, mDia, and Eps8 involved in microfilament organization to the IRSp53 SH3 domain. Phosphorylation of S366 by Par1b and Par1b stimulation of S434-5 phosphorylation by an unidentified kinase recruits 14-3-3 proteins that interfere with WAVE2 binding and the recruitment of other proteins that interact with the IRSp53 C-terminal domain. Par1b overexpression inhibits IRSp53-dependent actin dynamics that regulate cell–ECM signaling and result in altered lumen polarity in MDCK cells. In other cell types, such as Cos7, unidentified kinases inhibit IRSp53 function by creating 14-3-3 binding sites on T340/T360.
IRSp53 regulates outside-in and inside-out cell–matrix interactions in MDCK cells. On one hand, IRSp53 directs cytoskeletal dynamics and FA formation in response to integrin engagement on collagen substrates; on the other hand, it participates in the formation of the epithelial basal lamina. In the early cell spreading phase, integrin engagement activates the FAK–Src kinase complex that in turn stimulates Cdc42 and Rac1 activity while rhoA is inhibited. Several known IRSp53-interacting proteins have been identified in this pathway, including the Cdc42 effectors Mena and N-WASP, the Rac1 effector WAVE, racGEF Tiam1, and mDia1. During the late spreading phase, a poorly understood switch from Rac1 to rhoA activity occurs that might be triggered by an increase in membrane tension as cells extend their surface area (Huveneers and Danen, 2009). Aberrant lamella in spreading of IRSp53-depleted cells suggest a role for IRSp53 in Rac1/Cdc42 signaling; whether lack of radial stress fibers is simply the consequence of early spreading defects or whether IRSp53 also regulates rhoA directly cannot be resolved at the moment, although the reduced diametrical stress fibers might indicate a direct effect of IRSp53 on rhoA activity in polarized cells. Actin dynamics at the leading edge of lamellae also govern the clustering and positioning of activated β1-integrins (Galbraith et al., 2007) that in turn could account for the role of IRSp53 in FA formation. As in early spreading, activation of Rac1 and concomitant inhibition of rhoA underlie the assembly of laminin into a basal lamina in MDCK cysts (Yu et al., 2005, 2008), and thus, similar IRSp53-signaling platforms could operate to regulate spreading and basal lamina assembly.
Although prominently associated with the region of cell–cell contact, no defects in adherens junction formation were apparent in IRSp53-depleted MDCK cells. This was somewhat surprising, given that depletion of WAVE2 or the WAVE-stabilizing proteins Abi1 and HSP300, which can form a complex with IRSp53 to mediate Arp2/3-dependent actin polymerization downstream of Rac1, caused severe defects in E-cadherin recruitment to cell–cell contact sites in polarizing cells (Yamazaki et al., 2007). The scenario is reminiscent of that described for Tiam1, which despite being recruited to nascent E-cadherin junctions, was dispensable for Rac1 activation and heterotypic E-cadherin engagement and clustering (Kraemer et al., 2007). Thus, IRSp53-independent mechanisms of Rac1–WAVE activation operate, or can at least compensate, for IRSp53 depletion at the lateral membrane.
In summary, we identified IRSp53 as the first Par1b substrate that contributes to its role in lumen polarity. The fact that IRSp53 works downstream of Par1b in cell spreading and lateral lumen polarity, whereas only Par1b but not IRSp53 regulates cell–cell adhesion, suggests that Par1b function in lumen formation involves cell–ECM signaling.
Materials and methods
Cell lines and culture
For cell spreading analysis, cells were trypsinized and plated at 25,000 cells/cm2 on 10 µg/cm2 rat tail collagen I (BD)–coated coverslips or MatTek chambers (for live cell analysis) for 30, 60, 90, and 120 min or for up to 6 h. For synchronized cell polarization analysis in Figs. 1 E, 4 C, 5 D, 6, 7, and S4, Ca2+ switch assays were performed as described previously (Cohen et al., 2004). Transduction with recombinant Par1b adenovirus (Figs. 2 A, 3 C, and 6 B; Cohen et al., 2004) was in serum-free SMEM for 1 h with 1 plaque-forming unit/cell. mRFP-actin, Par1b-GFP, paxillin-mCherry, and tagged IRSp53 cDNAs were transfected by nucleofection (Lonza) as previously described (Cohen et al., 2004). All experiments in which the effect of Par1b was assessed (except for Par1b adenovirus–transduced and Par1b-GFP–transfected cells) were conducted in a Par1b-MDCK cell line expressing C-terminal Myc-tagged canine EMK1/Par1b under a dox-regulated promoter 10-fold over endogenous levels as previously described (Cohen et al., 2004). For control conditions, expression of the recombinant protein was repressed by 1 µg/ml dox. IRSp53 shRNAmir and Par1b shRNAmir inducible cell lines were generated from the MDCK Tet-Off cell line T23 (provided by K. Mostov, University of California, San Francisco, San Francisco, CA). Induction by dox withdrawal occurred for 4 d. IRSp53-shRNAmir cells were induced in the presence of 50 µM enoxacin to enhance the RNAi response (Shan et al., 2008) for 4 d. Controls were maintained in the presence of dox but also enoxacin treated. Dox-inducible HA-IRSp53 and HA-IRSp53ΔSH3 MDCK Tet-Off cell lines were provided by J. Casanova (University of Virginia, Charlottesville, VA; Shi et al., 2005) and induced for 24 h by dox removal. Stable MDCK cell lines constitutively expressing IRSp53-HA3 and IRSp53A4-HA3 were generated from T23-MDCK cells. For Figs. 2 C, 5 B, and 6, adenoviruses encoding l-Cat (chloramphenicol acetyl transferase), Par1b, and kinase-dead Par1b-K49A (Par1b-KA) were transduced as previously described (Cohen et al., 2007). For 3D culture, Par1b and IRSp53KD cells were cultured in ±dox for 2 d before seeding in 3D collagen I cultures and grown for 6 d as previously described (Cohen and Müsch, 2003).
cDNA constructs and recombinant proteins
Par1b-shmir and IRSp53-shmir oligos were generated by PCR amplification as previously described (Sun et al., 2006) and cloned into the pTRE2 vector. The following target sequences for both mRNAs were previously described and validated (Cohen et al., 2004; Shi et al., 2005): Par1b, 5′-AAGAGGTAGCTGTGAAGATCAT-3′; IRSp53, 5′-CGACACTTGTATGAAGATGTAA-3′. mRFP-actin cDNA was provided by A. Barth (Stanford University, Palo Alto, CA), and Par1b-GFP was generated from the long splice variant of canine Par1b (Cohen et al., 2004) cloned into pEGFP-N1. The paxillin-mCherry plasmid was provided by S. Hanks (Vanderbilt University, Nashville, TN). cDNA encoding Cdc42Q61L-HA3 was provided by R. Cerione (Cornell University, Ithaca, NY). WAVE-GFP cDNA was provided by D. Cox (Albert-Einstein College of Medicine, The Bronx, NY). cDNA of human Myc-tagged IRSp53 (in pRK5; Krugmann et al., 2001) was provided by A. Hall (Memorial Sloan-Kettering, New York, NY), and HA-IRSp53 and ΔSH3-HA-IRSp53 (in pTRE2; Shi et al., 2005) were provided by J. Casanova. Both cDNAs were used as templates for site-directed mutagenesis to introduce the Ser/Thr-Ala mutations using the QuickChange Mutagenesis kit (Agilent Technologies). All mutant constructs were verified by sequencing. The human IRSp53-WT and IRSp53-A4 cDNAs were also cloned into pKH3 (Mattingly et al., 1994) for triple-HA tagging and subsequently exchanged for the luciferase cDNA in the pRL-TK vector (Promega) to yield IRSp53-HA3 under the thymidine kinase promoter. cDNAs encoding GST-tagged IRSp53 aa 1–178, aa 1–317, aa 180–317, aa 180–521, and aa 319–521 (Krugmann et al., 2001) were provided by A. Hall, and GST-tagged proteins were prepared from BL21 cells according to standard procedures. Full-length IRSp53 was cloned into pT7TEV-HMBP3 (provided by E. Yakubovskaya, Stony Brook University, Stony Brook, NY; Yakubovskaya et al., 2010), purified as His-tagged MBP fusion protein from bacteria by affinity chromatography on Ni-agarose, followed by TEV protease cleavage to release the untagged IRSp53 moiety. His-tagged recombinant TEV was readsorbed on Ni beads. 14-3-3τ cDNA (I.M.A.G.E. clone 5100237) was cloned into pGEX4T-1, purified from bacteria according to standard procedures, and covalently coupled to Affi15 gel (Bio-Rad Laboratories) according to the manufacturer’s instructions at a concentration of 20 mg/ml gel. Recombinant 14-3-3σ and His-14-3-3ζ were purchased from ProSpec.
Real-time PCR
Primer pairs 5′-CCTGCCCAGACGGCCGGTAC-3′/5′-CTGCTCACAGCCCGTGCTCCG-3′ and 5′-GGGAAATCGTGCGTGACATTAAG-3′/5′-TGTGTTGGCGTACAGGTCTTTG-3′ specific for canine IRSp53 mRNA and actin, respectively, were used to amplify ∼100-bp fragments from total reverse-transcribed MDCK cDNA using Sypro Green Mix (Thermo Fisher Scientific) in a realplex Mastercycler (Eppendorf).
Antibodies
The following antibodies were used in this study: gp135 (clone 3F21D8; provided by G. Ojakian, Downstate Medical Center, State University of New York, Brooklyn, NY), rat monoclonal ZO-1 (Millipore), E-cadherin monoclonal clone rr1 (provided by the Developmental Studies Hybridoma Bank Iowa as hybridoma and ascites fluid; Gumbiner and Simons, 1986), HA clone 16B12 (Covance), Myc clone 9E10 (EMD), IRSp53 goat polyclonal (ab15697; Abcam), and mouse monoclonal (#612674; BD), anti–phospho-serine (#600-401-261; Rockland), and anti-laminin (L9393; Sigma-Aldrich).
Immunolabeling techniques and IF analysis
For immunofluorescence analysis, cells were fixed in 2% paraformaldehyde at 37°C, permeabilized with 0.2% Triton X-100, and blocked with 1% BSA. Antibody incubation was performed in PBS/1% BSA. Secondary antibodies used were coupled to Alexa Fluor 488, Cy3, or Alexa Fluor 647. In addition, rhodamin-phalloidin was used. Coverslips were mounted in nonhardening medium containing glycerol/2.5% DABCO. 3D cysts were incubated with a laminin antibody for 4 h at 37°C before fixation and labeled with primary and secondary antibodies at 4°C overnight. Confocal microscopy was performed with a microscope (TCS SP5; Leica) using either an HCX Plan Apo CS 63.0× 1.40 NA oil or 40.0× 1.25 NA oil objective. Individual confocal x-y or x-z sections, or where indicated, x-y-z or x-z-y projections generated with LAS AF software (Leica) are presented. The phase-contrast images in Fig. 1 A were acquired on a DMIL microscope (Leica) with a 20× NA 0.3 objective equipped with a digital camera (Leica). All image figures were composed with Photoshop (Adobe). Brightness was adjusted using the “Brightness” function on the entire image. For quantitative IB analysis, we probed all blots either with 125I–protein A (Fig. 2, A and C; and Fig. 3 C) or DyLight 680/800–coupled secondary antibodies (all other IBs) and analyzed them in a laser scanner (Typhoon [Amersham] or FLA-9000 [Fujifilm]). Cell lysis in the experiments for Figs. 2 and 3 was performed in RIPA buffer (20 mM Tris-/HCl, pH 7.5, 150 mM NaCl, 5 mM EDTA, 1% NP-40, 0.25% deoxycholate, and protease and phosphatase inhibitors) as previously described (Elbert et al., 2006). For Fig. 2 D, cells were labeled with 0.2 mCi/ml [32P]Na phosphate for 90 min before cell lysis and IP with HA antibodies. For the experiments in Fig. 8, cells were lysed in a Triton X-100 buffer (20 mM Tris-HCl, pH 7.5, 150 mM NaCl, 5 mM EDTA, 1% Triton X-100, 10% glycerol, and protease and phosphatase inhibitors). GST-IRSp53CT and 14-3-3 binding to immobilized IRSp53-CT occurred in the same buffer. Metabolic labeling occurred with 0.1 mCi/ml [35S] translabel (PerkinElmer) in Met/Cys-free DME overnight.
Quantification of lumen polarity and cell spreading
Quantification of cells exhibiting lateral lumina and of cell spreading was obtained from 40× confocal images of fixed cells, which were labeled for gp135 and ZO-1 or rhodamine-phalloidin, respectively. Lateral lumina were defined by gp135 accumulation between neighboring cells flanked by a circumferential ring of ZO-1. Data in Fig. 5 D are from two experiments with five images (∼150 cells/image), and data in Figs. 6 and 7 are from three experiments with five images (∼150 cells/image) per time point. For cell spreading, the longest cell diameter, including lamellipodia extending beyond the cell body, was measured using ImageJ (National Institutes of Health). Data are from three experiments with five images (∼50 cells per image). Standard errors are presented, and significance was determined in two-tailed Student’s t tests.
Live cell microscopy
Live cell imaging was performed starting 30–60 min after plating on collagen I with a microscope (TCS SP5) equipped with a motorized x-y stage for multiple position finding (Leica) and with an 8,000-Hz resonant scanner (lenses: HCX Plan Apo CS 63.0× NA 1.40 oil or 40.0× NA 1.25 oil). Cells were imaged on collagen I–coated MaTek chambers at 37°C in a CO2-enriched atmosphere in recoding medium (DME without Phenol red, 10% FCS, 3.7 mg/ml NaHCO3, and 25 mM Hepes, pH 7.4; line mean, 32–128; frame mean, 1). Single confocal planes (pinhole setting 1 or 2 AU) and a 488-nm line of an Argon laser and 594-nm laser for GFP, mRFP, and brightfield were used. Image sequences were processed with ImageJ and QuickTime.
14-3-3 affinity chromatography
About 360 million TAP-tagged Par1b or Par1b-K49A–expressing cells were homogenized by nitrogen cavitation and 8 ml postnuclear supernatant separated into cytosolic and membrane/cytoskeletal fraction by centrifugation at 150,000 g for 90 min. The pellet was solubilized in RIPA buffer containing 1% NP-40 and 0.5% deoxycholate (protease inhibitors: 2 mM AEBSF and 10 µg/ml each leupeptin, pepstatin, and antipain; phosphatase inhibitors: 2 mM orthovanadate, 10 mM NaF, 10 mM glycerophosphate, and 5 mM Na-pyophosphate) and by passing three times through a 30-G needle. The solubilized fraction (cleared by a 45-min spin at 17,000 g) was incubated overnight with 20 µl immobilized 14-3-3τ and eluted with 2× 100 µl 0.5 mM R18 peptide (Wang et al., 1999). Protein bands excised from colloidal Coomassie-stained gels were identified by nano liquid chromatography/tandem mass spectrometry peptide sequencing by ProtTech, Inc. IRSp53 was identified on the basis of four peptides: (ALAGVTYAAK, YSDKELQYIDAISNK, SFHNELLTQLEQK, and EGDLITLLVPEAR). 14-3-3-binding of recombinant IRSp53 from unfractionated RIPA lysates occurred in a similar manner. For recombinant binding assays, 5 µg recombinant IRSp53 was subjected to kinase assays with Par1b or Par1b-KA immunoisolated from 1 mg cell lysate in the presence of 1 mM ATP and subsequently incubated with 5 µl immobilized 14-3-3τ.
Par1b kinase assays
In vitro kinase assays with immunoprecipitated Par1b and recombinant IRSp53 substrate were conducted as previously described for Dvl (Elbert et al., 2006), with the difference that TAP-tagged Par1b and Par1b-KA and TAP-tagged Par1b with an enlarged ATP-binding pocket (Par1b *ATP) were affinity isolated on IgG Sepharose and that the ATP analogue N6-benzyl-ATP[35γS] rather than [32P]γATP was used for the assay in Fig. 3 D. The details of this approach were based on previous experiments (Blethrow et al., 2004, 2008). For the kinase assays in cell lysates, TAP-Par1b and TAP-Par1b*ATP cell lines were transiently transfected with Myc-IRSp53 constructs, and a Triton X-100 lysate (concentration of 5–10 mg/ml) was incubated with 1 mM GTP and 100,000 cpm N6-benzyl-ATP[35γS] for 1 h at 30°C.
Online supplemental material
Fig. S1 shows that 3D collagen cysts of Par1b-overexpressing and IRSp53-depleted cells have multiple lumen, inverted apical surfaces, and disrupted laminin at the basal domain. Fig. S2 shows that IRSp53 becomes enriched in the 14-3-3 binding fraction in Par1b-overexpressing cells. Fig. S3 shows that Par1b phosphorylation stimulates IRSp53 binding to 13-4-4τ, -ζ, and -σ Fig. S4 shows that IRSp53 depletion does not compromise E-cadherin recruitment to cell–cell adhesion sites or its association with microfilaments in Ca2+ switch assays Fig. S5 shows expression of shRNAmir-resistant IRSp53 in IRSp53-depleted cells. Video 1 shows spreading and actin dynamics in Par1b-MDCK cells cultured in either +dox or −dox and expressing mRFP-actin. Video 2 shows spreading and actin dynamics in Par1b-KD-MDCK cells. Video 3 shows paxillin dynamics in Par1b and Par1b-KD MDCK cells expressing paxillin-mCherry. Video 4 shows confocal x-y series through collagen I cysts of Par1b-MDCK cells and IRSp53-KD cells cultured in +dox and −dox. Video 5 shows actin dynamics of T23 (control)-MDCK cells expressing either GFP or Par1b-GFP and mRFP actin. Video 6 shows cell spreading and compaction in Par1b-MDCK monolayers cultured in +dox or −dox. Video 7 shows phalloidin staining in Par1b-MDCK cells ±dox and in IRSp53KD cells ±dox. Video 8 shows spreading and actin dynamics in IRSp53KD-MDCK cells cultured in +dox or −dox and expressing mRFP-actin. Video 9 shows paxillin dynamics in IRSp53-KD cells cultured in +dox or −dox and expressing paxillin-mCherry and rescue with IRSp53 WT and IRSp53-4A. Video 10 shows cell spreading and compaction in IRSp53-KD-MDCK monolayers cultured in +dox or −dox.
Spreading and actin dynamics in Par1b-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP fluorescence time-lapse series (1 frame/1 min) are shown. Left, +dox; right, −dox. Bar, 40 µm.
Spreading and actin dynamics in Par1b-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP fluorescence time-lapse series (1 frame/1 min) are shown. Left, +dox; right, −dox. Bar, 40 µm.
Spreading and actin dynamics in IRSp53KD-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP-fluorescence time-lapse series (1 frame/3 s) are shown. Left, +dox; right, −dox. Bars, 10 µm.
Spreading and actin dynamics in IRSp53KD-MDCK cells cultured in ±dox and expressing mRFP-actin. Brightfield and mRFP-fluorescence time-lapse series (1 frame/3 s) are shown. Left, +dox; right, −dox. Bars, 10 µm.
Acknowledgments
We are grateful to Drs. Alan Hall, James Casanova, Steve Hanks, Irina Kaverina, Elena Yakubovskaya, Angela Barth, and Dianne Cox for kindly providing cDNAs used in this study. We thank our colleagues Dianne Cox and members of the Müsch laboratory, Jenny Nachbar and Alex Treyer, for discussions during the course of study and for critically reading the manuscript.
A. Müsch is supported by the National Institutes of Health (grant RO1 DK064842) and the Albert Einstein College of Medicine Start-Up Funds.


![Figure 2. Par1b promotes IRSp53 phosphorylation and 14-3-3 binding. (A) IRSp53 in total lysates from Par1b and Par1b-KD cells (1/10 input) and from lysate fractions adsorbed on immobilized 14-3-3τ and competitively eluted. Arrows show a decrease in IRSp53 mobility in the Par1b lysates. (B) In vitro kinase assays of recombinant IRSp53 with Par1b-WT or Par1b-KA followed by λ-phosphatase (+) or mock (m) treatment and subsequent binding to immobilized 14-3-3τ. (top) 1/10 of IRSp53 input is shown. (middle) IB is shown, with IRSp53 competitively eluted from 14-3-3 resin. (bottom) Autorad is shown, with [32P]γATP-Par1b–phosphorylated IRSp53. (C) IRSp53-HA or IRSp53ΔSH3-HA cells transduced with control, Par1b-WT, or Par1b-KA adenovirus. IB of 1/10 Par1b, IB of HA input, and phospho-serine after HA IP are shown. (D) IRSp53-HA immunoprecipitated from [32P]orthophosphate-labeled (autorad) and unlabeled (IB, IRSp53-HA) Par1b-KD cells. Par1b IB shows dox-dependent depletion of the two Par1b splice variants.](https://cdn.rupress.org/rup/content_public/journal/jcb/192/3/10.1083_jcb.201007002/5/m_jcb_201007002_gs_fig2.jpeg?Expires=1781624585&Signature=UlfTJAnLJOqwPkoF~OnzBnJ6GGwo4OmNuep8dMCP40kZy3bVJrRc2U0Vfaj6-35dIiyLBaAj81iPHrQq~D3iN2FkrzXSXhRNraoKmVwr~jw8TZkNZEYRGTScEiOK1z4aVCJpN6CtYoH3g0zSeOp~e11QqDVlJ-5SH1OsX5J3v~a95JD~lG7k3O2N0sF16svcYQTbickQdjVND1V8cipPDp-OC2DbLHLhCTS9Q4ia00MoV9Xp0ug3gDIyrxtN8Joa9OspzdJrc9kr1tqDuYnj74eEFe8tfjg23eR8s0E3g0KvxkC4HP5~2AH1b0i53xTlxPGA~IC~T0pSJYY6tuAihw__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
![Figure 3. Par1 phosphorylation on S366 and Par1b-depedendent S453/4/5 phosphorylation promote 14-3-3 binding of IRSp53. (A) Diagram of Par1b-dependent phosphorylation sites in IRSp53. (B) In vitro kinase assays with WT and Par1b-KA using [32P]γATP and IRSp53-CT (WT and point mutants) as substrates. (C) IRSp53-HA–transfected cells (WT, A4, and S453-5A) were transduced with control (c) or Par1b virus, and the HA IPs were probed for P-serine and subsequently for HA. (D) In vitro kinase assays with WT and Par1b-ATP* using [35S]benzyl-ATPγS and full-length IRSp53 WT and mutants as substrates. Autoradiographs/Coomassie stain of IRSp53. (E) Triton X-100 lysates of cells coexpressing Par1b-WT or Par1b-ATP* and WT IRSp53-Myc or IRSp53 mutants were incubated with [35S]benzyl-ATPγS. Autoradiographs and IB for IRSp53-Myc and Par1b. (F) Recombinant WT IRSp53 and mutant proteins subjected to in vitro kinase assays with Par1b-WT or Par1b-KA were affinity isolated on 14-3-3 beads and competitively eluted (top). IRSp53 input (middle) and Par1b levels (bottom) are also shown. (G) 14-3-3 binding of IRSp53 proteins from cell lysates. Note the lower expression of the S117A sample accounts for the weaker signal in the bound fraction. The graph shows SEM from six experiments (three experiments for S117A/S148A). (H) Co-IP of 14-3-3-HA with IRSp53-Myc proteins from cell lysates.](https://cdn.rupress.org/rup/content_public/journal/jcb/192/3/10.1083_jcb.201007002/5/m_jcb_201007002_rgb_fig3.jpeg?Expires=1781624585&Signature=ArLVSTJuSooSTcQAzuo8MxGzCWWV1sASY7-2n-PkxsqkHI7Xwwo84bbeTfggn9aIZwy9~4-Cn-jdzNa2e47gbJNrLdju7PPc1R6atuCcGhG6Mw7qab8F8EEQUxmxGNTWtKPUVEtH~vq6PsJuCWZwjbnlOPd8oP30NoJuofAIhOOg9rTvfjAOhzKt-TuKKTd8VB5xOVQJAI4awubI1wch688ynpUI4hycavik2k18YMiSJ~2niC684q6OJBmImCVRQEFaXRvVpmPRX5N8DSzq3RehrQ9nS0M9brA23J8pTTDPo6v1gLTa7vrTDOvl8d3DkMqbqBbBD5FoAHsb3BYzFg__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)




![Figure 8. Par1b phosphorylation inhibits the recruitment of WAVE2 and other IRSp53-binding proteins to its C-terminal, SH3-containing domain. (A and B) WAVE2 co-IPs with IRSp53 in 293 cells expressing WAVE2-GFP, Myc-tagged IRSp53 constructs (WT, A4, or IRSp53 Δ317–521; ΔCT), and Par1b where indicated. 1/10 of the lysate was analyzed for WAVE and Par1b expression. (B) Data are from three experiments with and without recombinant Par1b and three experiments without recombinant Par1b only. Error bars indicate SEM. (C) Cdc42Q61L co-IPs with IRSp53-Myc constructs from 293 cells. Arrows indicate Cdc42, and asterisks indicate IgG H and L chains. Data are representative of three experiments. (D–F) GST-IRSp53CT was subjected to Par1b phosphorylation, immobilized on glutathione and incubated with 14-3-3τ as indicated. (D) Experimental schematic is shown. (E) Binding of proteins from [35S]Met/Cys-labeled 293 lysates to immobilized IRSp53CT was analyzed by Coomassie and autoradiography. Arrows indicate specific bands that are reduced in the Par1b sample. Asterisk indicates cellular 14-3-3, with one out of two identical results represented. (F) 14-3-3 binding to IRSp53CT was analyzed by IB, and IRSp53-CT levels by Ponceau red stain are shown.](https://cdn.rupress.org/rup/content_public/journal/jcb/192/3/10.1083_jcb.201007002/5/m_jcb_201007002_rgb_fig8.jpeg?Expires=1781624585&Signature=J1nfqcbgXSk9uiSdQj6KRkrHKkY7xLkAaP~QuDMoN9aCZ51Si63pajS9Vzqcr9NDUTVjeEwZSdLcKfXA6zOWTbKgkayzoUo12OCHN-GHRmrL7PPrHE79Xd7jATl1nlU5mE0QyuOGwv9AnbTBM0ke7V~ef26ANME~SvdzkbFWhIFT4prFHQoCcJvPQolDJ1918hFueiRe-paex5vHnmhOEdBhLTKZFRMAjeC2~xGXaqBByENKV--10USImAO31uys6i80zdGPDnjOGRqq4NPi3-i7WWjyvLQdNI6hmyjyYSwYLWl76PCST99JhN9ztaJ4EiJGeSFXrqzLy5NmRDm~Uw__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)



