Ataxia telangiectasia and Rad3-related (ATR) kinase and its interacting protein ATRIP orchestrate the replication stress response. Homozygous splice variants in the ATRIP gene, resulting in ATRIP deficiency, were identified in two patients of independent ancestry with microcephaly, primordial dwarfism, and recurrent infections. The c.829+5G>T patient exhibited lymphopenia, poor vaccine responses, autoimmune features with hemolytic anemia, and neutropenia. Immunophenotyping revealed reduced CD16+/CD56dim NK cells and absent naïve T cells, MAIT cells, and iNKT cells. Lymphocytic defects were characterized by TCR oligoclonality, abnormal class switch recombination, and impaired T cell proliferation. ATRIP deficiency resulted in low-grade ATR activation but impaired CHK1 phosphorylation under genotoxic stress. ATRIP-deficient cells inadequately regulated DNA replication, leading to chromosomal instability, compromised cell cycle control, and impaired cell viability. CRISPR-SelectTIME confirmed reduced cell fitness for both variants. This study establishes ATRIP deficiency as a monogenic cause of microcephalic primordial dwarfism, highlights ATRIP’s critical role in protecting immune cells from replication stress, and offers new insights into its canonical functions.
Introduction
Maintaining genomic integrity requires cells to deploy defense mechanisms to counteract various DNA-damaging assaults. The phosphatidylinositol 3-kinase-related kinases (PIKKs) ATR, ATM, and DNA-PKcs act as key mediators of the DNA damage response (DDR), initiating signaling cascades that coordinate cell cycle progression, checkpoint activation, and concurrent DNA repair (Blackford and Jackson, 2017; Ciccia and Elledge, 2010). While ATM and DNA-PKcs respond to DNA double-strand breaks (DSBs), ATR primarily safeguards DNA synthesis during the S phase (Ciccia and Elledge, 2010; Cimprich and Cortez, 2008). Replication protein A (RPA)-coated single-stranded DNA (ssDNA) arises after the processing of damaged DNA, including stalled replication forks, resected DSBs, and UV-induced bulky adducts, and serves as a platform for ATR activation. By phosphorylating substrates such as CHK1, ATR orchestrates the replication stress response, triggering intra-S and G2/M checkpoints, suppressing origin firing, stabilizing replication forks, and promoting replication fork restart (Matsuoka et al., 2007; Jazayeri et al., 2006; Saldivar et al., 2017). ATR is essential for mammalian DNA replication, and complete loss of ATR was shown to result in early embryonic lethality in mice (de Klein et al., 2000; Brown and Baltimore, 2000; Nam et al., 2011a; Lam, 2022; Cortez et al., 2001). A homozygous hypomorphic leaky splice variant in ATR (NM_001184.4 (ATR): c.2022A>G, p.[Gly674=]) was first reported by O’Driscoll et al. (2003) in two families with Seckel syndrome (Goodship et al., 2000) (OMIM 210600), a form of microcephalic primordial dwarfism (MPD) associated with intellectual disability (ID) and distinct craniofacial features. To date, four additional ATR patients have been described, all harboring biallelic hypomorphic variants, leading to highly diminished but not abolished ATR protein levels (Ogi et al., 2012; Mokrani-Benhelli et al., 2013; Llorens-Agost et al., 2018).
The ubiquitously expressed ATR-interacting protein (ATRIP) is an essential binding partner of ATR, illustrated by the co-dependency of ATRIP and ATR for protein stability (Cortez et al., 2001). On a functional level, ATRIP recognizes and binds RPA-ssDNA nucleoprotein filaments, allowing ATR recruitment and subsequent activation by the kinases TOPBP1 and the recently identified ETAA1 (Bass et al., 2016). The structural basis for this signaling nexus has recently been resolved: the N-terminal RPA-binding motif of ATRIP directly interacts with RPA while its coiled-coil domain is required for the simultaneous homodimerization of two ATRIP molecules. Through its C-terminal HEAT motifs, ATRIP associates with the N-terminal region of ATR, and these ATR–ATRIP complexes form stable dimers of heterodimers upon recruitment to RPA-ssDNA (Wang et al., 2017; Deshpande et al., 2017; Rao et al., 2018). Mutational analyses revealed that ATRIP–TOPBP1 interactions and downstream ATR activation depend on a region adjacent to the coiled-coil domain in ATRIP (Rao et al., 2018). Unlike TOPBP1, which relies on both ATRIP and the RAD9–RAD1–HUS1 (9-1-1) clamp for recruitment to the ATRIP–ATR complex, ETAA1 directly interacts with RPA-ssDNA and ATR (Delacroix et al., 2007; Haahr et al., 2016; Bass et al., 2016). Defective ATRIP function is associated with human disease as reduced ATRIP expression was found in a patient with MPD (Ogi et al., 2012) and heterozygous ATRIP loss-of-function (LOF) variants have been associated with breast cancer susceptibility (Cybulski et al., 2023). Moreover, conditional ATRIP loss in the central nervous system in mice resulted in microcephaly, providing further evidence for a causal link between ATRIP deficiency and MPD (Matos-Rodrigues et al., 2020).
MPDs are a genetically heterogeneous group of overlapping disorders, defined by severe microcephaly and intrauterine and postnatal growth restriction. Pathogenic variants associated with MPDs have been identified in genes involved in seemingly distinct cellular processes such as DNA replication initiation, DNA repair, cell cycle progression, and centrosome function (Klingseisen and Jackson, 2011; Reynolds et al., 2017; Harley et al., 2016; Khetarpal et al., 2016). Disruption of the encoded proteins consistently restricted cell proliferation dynamics, thus exposing the common biological basis of MPDs. In addition, several of the genes involved in DNA replication initiation (MCM4, MCM10, GINS1, GINS4, POLD1-2, and POLE1-2), when disrupted, gave rise to a syndromic combined immunodeficiency (CID). Although clinically variable, these immunodeficiencies have in common that they are characterized by susceptibility to severe infections by herpes viruses and defects in the natural killer (NK) cell compartment (Conde et al., 2019; Conte et al., 2022; Cottineau et al., 2017; Cui et al., 2020; Frugoni et al., 2016; Gineau et al., 2012; Hughes et al., 2012; Nichols-Vinueza et al., 2021; Pachlopnik Schmid et al., 2012).
Here, we report that ATRIP deficiency, caused by homozygous splice variants in ATRIP, manifests with features of MPD and immunodeficiency. The LOF ATRIP variants allow residual ATR activity, although profoundly impaired. We performed a comprehensive analysis of the functional impact of ATRIP loss on ATR signaling activity and delineated the accompanying downstream cellular outcomes. In addition to genomic instability, patient-derived cells display an inadequate ATR response to genotoxic lesions, as evidenced by dysregulated cell cycle progression, increased chromosomal sensitivity, impaired proliferation, and pronounced decline in cell viability. Modeling of the ATRIP variants by CRISPR-SelectTIME revealed reduced cell fitness (Niu et al., 2022). Our observations highlight a more nuanced role of ATRIP within the ATR signaling pathway, clarify the biological basis of ATRIP-mediated human disease, and expand the disease phenotype associated with ATRIP deficiency.
Results
Homozygous intronic ATRIP variants in patients with MPD and immunodeficiency
Patient F1Pt was born at term, small for gestational age, and presented with severe microcephaly, developmental delay, and dysmorphic features (Fig. S1, a and b). Clinical investigations revealed failure to thrive, growth retardation, grossly normal skeletal development, and a mild ID. At age 2, she developed severe varicella and recurring respiratory tract infections (RTIs). Laboratory investigations showed lymphopenia and intermittent neutropenia (Table 1). Whole exome sequencing (WES) revealed a homozygous splice variant in intron 5 of the ATRIP gene (NM_130384.3): c.829+5G>T (Chr3(GRCh38):g.48457421G>T), which was confirmed by Sanger sequencing (Fig. 1 a and Fig. S1 c). Importantly, WES analysis of a large cohort of patients with microcephaly identified a male patient of North Indian ancestry (patient F46.1) with another homozygous splice variant in intron 5 of ATRIP (c.829+2T>G) (Masih et al., 2022). He presented with similar phenotypic features of facial dysmorphism, short stature, microcephaly, moderate ID, and recurrent RTIs (Fig. S1 a). Finally, microcephaly and short stature were previously reported in a patient with reduced ATRIP expression due to defective splicing (patient CV1720) (Ogi et al., 2012). Reappraisal of patient CV1720’s clinical status revealed a similar phenotype of late-onset lymphopenia and intermittent neutropenia (Table 1). More detailed clinical information can be found in the case descriptions (see Materials and methods and Table S1).
Expanded genetic and molecular analysis confirms complete loss-of-function ATRIP variant in patient F1Pt. (a) Photographs of patients F1Pt and F46.1 demonstrating facial similarities, including sloping forehead and beak-like nose. (b) Weight (Wgt), height (Hgt), and head circumference (occipital frontal circumference; OFC) at birth plotted as z-scores (SD from population mean for age and sex). Dashed line at −2 SD indicates cut-off for normal population distribution. ATRIP patients are denoted by red dots; previous reported ATR patients are denoted by blue dots. (c) Electropherograms of genomic DNA extracted from blood for F1Pt and a HC. Nucleotide numbering is in accordance with ENST00000320211.1. Images represent results from five independent experiments. (d) Population genetics: Highest SpliceAI Delta Score against gnomAD v4.0.0. allele frequency (AF) for splice region variants in ATRIP (ENST00000320211.1). Splice region variants are defined as nucleotide changes within the ±20 base pairs (bp) flanking the exon. Black dots and blue cross signs represent heterozygous and homozygous variants, respectively. More details regarding homozygous splice variants can be found in Table S4. Red shaded dots represent ATRIP variants of interest (c.829+5G>T and c.829+2T>G). (e) Fragment analysis and size profiles of PCR-amplified cDNA extracted from fibroblasts for F1Pt and a HC. Arrows indicate the position of forward and reverse primers used for PCR amplification. Percentages represent relative quantification of the 538 bp wild type (WT) and 380 bp mutant (r.7671_829del) fragment. Data are reflective of two independent experiments. (f) Electropherograms of cDNA extracted from PHA blasts for F1Fa (father), F1Mo (mother), and a HC. Nucleotide numbering is in accordance with ENST00000320211.1. Data are reflective of five independent experiments. (g) Sashimi plot of targeted RNA-seq data generated in Integrative Genomics Viewer. Input RNA was extracted from PHA blasts of F1Pt and a HC. Exon numbering is in accordance with ENST00000320211.1. (h) Fragment analysis of PCR-amplified cDNA using two primer pairs (S1: E3–E5; S2: E5–E7, indicated by arrows) on fibroblasts from F1Pt and a HC. Data are reflective of two independent experiments. (i) Reverse transcription quantitative PCR (RT-qPCR) analysis on fibroblasts of F1Pt and HCs (n = 3) of amplicon in exon 3–4 and exon 5–6. The relative expression to β-actin in a logarithmic scale is shown. Data from two independent experiments are shown. (j) Endogenous protein expression of ATRIP and interaction partners in EBV-LCLs from F1Pt, F1Fa, and HCs (n = 3). β-Tubulin was used as loading control. Western blot image is reflective of two independent experiments. (k) Quantification in arbitrary units of digitized chemiluminescent signals from Fig. 1 e normalized to β-tubulin signal from the same lane. Graph depicts fold change of normalized protein levels over the mean of HCs (n = 3) of at least four immunoblots. Source data are available for this figure: SourceDataFS1.
Expanded genetic and molecular analysis confirms complete loss-of-function ATRIP variant in patient F1Pt. (a) Photographs of patients F1Pt and F46.1 demonstrating facial similarities, including sloping forehead and beak-like nose. (b) Weight (Wgt), height (Hgt), and head circumference (occipital frontal circumference; OFC) at birth plotted as z-scores (SD from population mean for age and sex). Dashed line at −2 SD indicates cut-off for normal population distribution. ATRIP patients are denoted by red dots; previous reported ATR patients are denoted by blue dots. (c) Electropherograms of genomic DNA extracted from blood for F1Pt and a HC. Nucleotide numbering is in accordance with ENST00000320211.1. Images represent results from five independent experiments. (d) Population genetics: Highest SpliceAI Delta Score against gnomAD v4.0.0. allele frequency (AF) for splice region variants in ATRIP (ENST00000320211.1). Splice region variants are defined as nucleotide changes within the ±20 base pairs (bp) flanking the exon. Black dots and blue cross signs represent heterozygous and homozygous variants, respectively. More details regarding homozygous splice variants can be found in Table S4. Red shaded dots represent ATRIP variants of interest (c.829+5G>T and c.829+2T>G). (e) Fragment analysis and size profiles of PCR-amplified cDNA extracted from fibroblasts for F1Pt and a HC. Arrows indicate the position of forward and reverse primers used for PCR amplification. Percentages represent relative quantification of the 538 bp wild type (WT) and 380 bp mutant (r.7671_829del) fragment. Data are reflective of two independent experiments. (f) Electropherograms of cDNA extracted from PHA blasts for F1Fa (father), F1Mo (mother), and a HC. Nucleotide numbering is in accordance with ENST00000320211.1. Data are reflective of five independent experiments. (g) Sashimi plot of targeted RNA-seq data generated in Integrative Genomics Viewer. Input RNA was extracted from PHA blasts of F1Pt and a HC. Exon numbering is in accordance with ENST00000320211.1. (h) Fragment analysis of PCR-amplified cDNA using two primer pairs (S1: E3–E5; S2: E5–E7, indicated by arrows) on fibroblasts from F1Pt and a HC. Data are reflective of two independent experiments. (i) Reverse transcription quantitative PCR (RT-qPCR) analysis on fibroblasts of F1Pt and HCs (n = 3) of amplicon in exon 3–4 and exon 5–6. The relative expression to β-actin in a logarithmic scale is shown. Data from two independent experiments are shown. (j) Endogenous protein expression of ATRIP and interaction partners in EBV-LCLs from F1Pt, F1Fa, and HCs (n = 3). β-Tubulin was used as loading control. Western blot image is reflective of two independent experiments. (k) Quantification in arbitrary units of digitized chemiluminescent signals from Fig. 1 e normalized to β-tubulin signal from the same lane. Graph depicts fold change of normalized protein levels over the mean of HCs (n = 3) of at least four immunoblots. Source data are available for this figure: SourceDataFS1.
Full blood count and basic immunology screening of ATRIP patients (F1Pt and CV1720)
| F1Pt | CV1720 | ||||||
|---|---|---|---|---|---|---|---|
| Immunological features | 4 years, 5 mo | 17 years, 6 mo | 19 years, 8 mo | 7 years, 11 mo | 12 years, 5 mo | 13 years, 8 mo | 14 years, 4 mo |
| Peripheral blood count | |||||||
| White blood cell counts (/µl) | 10,830 (6,000–15,000) | 1,850 (4,300–9,640) | 3,060 (4,300–9,640) | 5,100 (5,000–15,500) | 3,200 (4,500–13,000) | 3,500 (4,500–13,000) | 4,500 (4,500–13,000) |
| Platelets (×103/µl) | 311 (229–533) | 87 (175–343) | 194 (175–343) | 279 (140–400) | 218 (140–400) | 199 (140–400) | 190 (140–400) |
| Hemoglobin (g/dl) | 12.1 (11.7–15.1) | 7.4 (11.7–15.1) | 13.4 (11.7–15.1) | 13.9 (11.5–14.5) | 14.0 (13.0–18.0) | 16.0 (13.0–18.0) | 15.0 (13.0–18.0) |
| Lymphocytes (/µl) | 1,191 (4,000–10,000) | 1,308 (1,230–3,420) | 1,050 (1,230–3,420) | 1,870 (1,500–6,500) | 1,430 (1,500–6,000) | 1,190 (1,500–6,000) | 1,830 (1,500–6,000) |
| Neutrophils (/µl) | 7,256 (1,500–7,500) | 250 (1,930–5,780) | 1,280 (1,930–5,780) | 2,760 (1,500–8,000) | 1,410 (1,800–8,000) | 1,820 (1,800–8,000) | 2,090 (1,800–8,000) |
| Monocytes (/µl) | 866 (400–1,200) | 117 (260–780) | 410 (260–780) | 350 (200–800) | 230 (200–800) | 420 (200–800) | 330 (200–800) |
| Eosinophils (/µl) | 106 (200–600) | 161 (30–370) | 260 (30–370) | 100 (40–400) | 70 (40–400) | 70 (40–400) | 180 (40–400) |
| Basophils (/µl) | 217 (10–100) | 0 (20–80) | 40 (20–80) | 40 (20–100) | 30 (20–100) | 0 (20–100) | 20 (20–100) |
| Lymphocyte subsets | |||||||
| T cells (CD3+) (/µl) | 834 (900–4,500) | 1,070 (700–2,100) | 1,090 (700–2,100) | N/A | N/A | N/A | 1,060 (1,000–2,200) |
| Helper T cells (CD4+) (/µl) | 262 (500–2,400) | 551 (300–1,400) | 413 (300–1,400) | N/A | N/A | N/A | 710 (530–1,300) |
| Cytotoxic T cells (CD8+) (/µl) (CD8+) | 524 (300–1,600) | 492 (200–1,200) | 639 (200–1,200) | N/A | N/A | N/A | 277 (330–920) |
| B cells (CD19+) (/µl) | 155 (200–2,100) | 139 (100–500) | 16 (100–500) | N/A | N/A | N/A | 480 (110–570) |
| NK cells (CD56+/CD16+) (/µl) | N/A | 58.9 (90–600) | 53.5 (90–600) | N/A | N/A | N/A | 150 (70–480) |
| Immunoglobulin levels | |||||||
| IgM (g/liter) | 2.2 (0.27–0.63) | 1.51 (0.4–2.48) | 0.31 (0.4–2.48)a | N/A | N/A | N/A | N/A |
| IgA (g/liter) | 1.4 (0.5–1.24) | 1.43 (0.71–3.65) | 0.46 (0.71–3.65)a | N/A | N/A | N/A | N/A |
| Total IgG (g/liter) | 21.4 (4.7–9.3) | 15.0 (7.0–16.0) | 11.3 (7.0–16.0) | N/A | N/A | N/A | N/A |
| IgG2 (g/liter) | 0.28 (0.72–3.4) | 1.79 (1.06–6.1)b | 2.47 (1.5–6.4)b | N/A | N/A | N/A | N/A |
| F1Pt | CV1720 | ||||||
|---|---|---|---|---|---|---|---|
| Immunological features | 4 years, 5 mo | 17 years, 6 mo | 19 years, 8 mo | 7 years, 11 mo | 12 years, 5 mo | 13 years, 8 mo | 14 years, 4 mo |
| Peripheral blood count | |||||||
| White blood cell counts (/µl) | 10,830 (6,000–15,000) | 1,850 (4,300–9,640) | 3,060 (4,300–9,640) | 5,100 (5,000–15,500) | 3,200 (4,500–13,000) | 3,500 (4,500–13,000) | 4,500 (4,500–13,000) |
| Platelets (×103/µl) | 311 (229–533) | 87 (175–343) | 194 (175–343) | 279 (140–400) | 218 (140–400) | 199 (140–400) | 190 (140–400) |
| Hemoglobin (g/dl) | 12.1 (11.7–15.1) | 7.4 (11.7–15.1) | 13.4 (11.7–15.1) | 13.9 (11.5–14.5) | 14.0 (13.0–18.0) | 16.0 (13.0–18.0) | 15.0 (13.0–18.0) |
| Lymphocytes (/µl) | 1,191 (4,000–10,000) | 1,308 (1,230–3,420) | 1,050 (1,230–3,420) | 1,870 (1,500–6,500) | 1,430 (1,500–6,000) | 1,190 (1,500–6,000) | 1,830 (1,500–6,000) |
| Neutrophils (/µl) | 7,256 (1,500–7,500) | 250 (1,930–5,780) | 1,280 (1,930–5,780) | 2,760 (1,500–8,000) | 1,410 (1,800–8,000) | 1,820 (1,800–8,000) | 2,090 (1,800–8,000) |
| Monocytes (/µl) | 866 (400–1,200) | 117 (260–780) | 410 (260–780) | 350 (200–800) | 230 (200–800) | 420 (200–800) | 330 (200–800) |
| Eosinophils (/µl) | 106 (200–600) | 161 (30–370) | 260 (30–370) | 100 (40–400) | 70 (40–400) | 70 (40–400) | 180 (40–400) |
| Basophils (/µl) | 217 (10–100) | 0 (20–80) | 40 (20–80) | 40 (20–100) | 30 (20–100) | 0 (20–100) | 20 (20–100) |
| Lymphocyte subsets | |||||||
| T cells (CD3+) (/µl) | 834 (900–4,500) | 1,070 (700–2,100) | 1,090 (700–2,100) | N/A | N/A | N/A | 1,060 (1,000–2,200) |
| Helper T cells (CD4+) (/µl) | 262 (500–2,400) | 551 (300–1,400) | 413 (300–1,400) | N/A | N/A | N/A | 710 (530–1,300) |
| Cytotoxic T cells (CD8+) (/µl) (CD8+) | 524 (300–1,600) | 492 (200–1,200) | 639 (200–1,200) | N/A | N/A | N/A | 277 (330–920) |
| B cells (CD19+) (/µl) | 155 (200–2,100) | 139 (100–500) | 16 (100–500) | N/A | N/A | N/A | 480 (110–570) |
| NK cells (CD56+/CD16+) (/µl) | N/A | 58.9 (90–600) | 53.5 (90–600) | N/A | N/A | N/A | 150 (70–480) |
| Immunoglobulin levels | |||||||
| IgM (g/liter) | 2.2 (0.27–0.63) | 1.51 (0.4–2.48) | 0.31 (0.4–2.48) | N/A | N/A | N/A | N/A |
| IgA (g/liter) | 1.4 (0.5–1.24) | 1.43 (0.71–3.65) | 0.46 (0.71–3.65) | N/A | N/A | N/A | N/A |
| Total IgG (g/liter) | 21.4 (4.7–9.3) | 15.0 (7.0–16.0) | 11.3 (7.0–16.0) | N/A | N/A | N/A | N/A |
| IgG2 (g/liter) | 0.28 (0.72–3.4) | 1.79 (1.06–6.1) | 2.47 (1.5–6.4) | N/A | N/A | N/A | N/A |
In case of F1Pt, the time points represent data prior to immunoglobulin treatment (age 4 years, 5 mo), before anti-CD20 mAb treatment (age 17 years, 6 mo), and after anti-CD20 mAb treatment (age 19 years, 8 mo). Bold and italic text signify reduced and elevated values, respectively, in comparison to age-matched reference values enclosed within brackets.
After anti-CD20 mAb treatment.
Under immunoglobulin replacement therapy.
Identification of homozygous complete LOFATRIP variants in patients with MPD. (a) Family pedigree with allele segregation of ATRIP splice variant. Index patient (F1Pt) is marked with an arrow, and clinical phenotype and genotype are indicated in the legend. A detailed phenotypical description can be found in Materials and methods. (b) Fragment analysis and size profiles of PCR-amplified cDNA extracted from PHA blasts for F1Pt, parents (F1Fa and F1Mo), and HC. Arrows indicate the position of forward and reverse primers used for PCR amplification. Percentages represent relative quantification of the 538 bp wild type (WT) and 380 bp mutant (r.671_829del) fragment. Data are representative of five independent experiments. (c) Electropherograms of cDNA extracted from PHA blasts of F1Pt and a control. Nucleotide numbering is in accordance with ENST00000320211.1. Depicted profiles are reflective of five independent experiments. (d) Schematic illustration of biallelic ATRIP variant effect at both transcript and amino acid level. (e) Endogenous protein expression of ATRIP and complex partners in primary fibroblasts from F1Pt, F02-98 (ATR patient), and controls (HCs). β-Tubulin was used as the loading control. Western blot image is reflective of four independent experiments. (f) Peak size profile in base pairs (bp) of PCR-amplified cDNA extracted from EBV-LCLs generated from the father (F1Fa) treated with and without puromycin. Percentages represent the relative ratio of WT (835 bp) and mutant (380 bp) fragments. Data are reflective of two independent experiments. (g) Schematic overview of the effect of the biallelic ATRIP variant at the protein level and an overview of the ATR-ATRIP complex with downstream effector CHK1. (h) HEK293T cells were transiently co-transfected with 3xFLAG-tagged ATRIP (3xFLAG-wtATRIP or 3xFLAG-mutATRIP), RPA70-Myc, and ATR-V5. After immunoprecipitation with anti-FLAG or control IgG, the interaction between ATRIP (FLAG) and ATR (V5), RPA70 (Myc), and endogenous TOPBP1 was examined by western blot analysis. β-Tubulin serves as a loading control. Results are reflective of three independent experiments. Source data are available for this figure: SourceData F1.
Identification of homozygous complete LOFATRIP variants in patients with MPD. (a) Family pedigree with allele segregation of ATRIP splice variant. Index patient (F1Pt) is marked with an arrow, and clinical phenotype and genotype are indicated in the legend. A detailed phenotypical description can be found in Materials and methods. (b) Fragment analysis and size profiles of PCR-amplified cDNA extracted from PHA blasts for F1Pt, parents (F1Fa and F1Mo), and HC. Arrows indicate the position of forward and reverse primers used for PCR amplification. Percentages represent relative quantification of the 538 bp wild type (WT) and 380 bp mutant (r.671_829del) fragment. Data are representative of five independent experiments. (c) Electropherograms of cDNA extracted from PHA blasts of F1Pt and a control. Nucleotide numbering is in accordance with ENST00000320211.1. Depicted profiles are reflective of five independent experiments. (d) Schematic illustration of biallelic ATRIP variant effect at both transcript and amino acid level. (e) Endogenous protein expression of ATRIP and complex partners in primary fibroblasts from F1Pt, F02-98 (ATR patient), and controls (HCs). β-Tubulin was used as the loading control. Western blot image is reflective of four independent experiments. (f) Peak size profile in base pairs (bp) of PCR-amplified cDNA extracted from EBV-LCLs generated from the father (F1Fa) treated with and without puromycin. Percentages represent the relative ratio of WT (835 bp) and mutant (380 bp) fragments. Data are reflective of two independent experiments. (g) Schematic overview of the effect of the biallelic ATRIP variant at the protein level and an overview of the ATR-ATRIP complex with downstream effector CHK1. (h) HEK293T cells were transiently co-transfected with 3xFLAG-tagged ATRIP (3xFLAG-wtATRIP or 3xFLAG-mutATRIP), RPA70-Myc, and ATR-V5. After immunoprecipitation with anti-FLAG or control IgG, the interaction between ATRIP (FLAG) and ATR (V5), RPA70 (Myc), and endogenous TOPBP1 was examined by western blot analysis. β-Tubulin serves as a loading control. Results are reflective of three independent experiments. Source data are available for this figure: SourceData F1.
In silico tools predicted inactivation of the wild type (WT) splice donor site near exon 5 for both novel variants (Table S2). Analysis of the population database gnomAD v4.0.0 revealed absence of the c.829+2T>G variant and presence of only five alleles of the c.829+5G>T variant, exclusively in a heterozygous state in the European (non-Finnish) subpopulation (MAF: 0.0004369%). Eleven homozygous variants in ATRIP splice boundaries, identified in gnomAD v4.0.0 (Table S3), were present with high allele frequencies (0.0001368–56.0807635%) and showed exclusively low recall cutoff SpliceAI scores (recall cutoff < 0.2; 0.00–0.18) (Fig. S1 d). Eight of the eleven homozygous variants were classified as benign (6/8) or likely benign (2/8) in ClinVar (Table S3). Based on the clinical and genetic evidence presented above, ATRIP was considered a plausible candidate gene for MPD and immunodeficiency.
ATRIP c.829+5G>T results in skipping of exon 5
To investigate the effect at the transcriptional level, reverse transcription PCR (RT-PCR) and Sanger sequencing were performed in phytohemagglutinin (PHA) blasts (Fig. 1, b and c) and fibroblasts (Fig. S1 e) from F1Pt. This failed to reveal a full-length transcript, but instead, showed the presence of a transcript lacking exon 5 (∆ex5, r.671_829del; p.(Ser224ArgfsTer2)) (Fig. 1 d). In PHA blasts from her parents (F1Mo and F1Fa), both the full-length and the shorter transcript were observed, in compliance with their zygosity (Fig. 1 b and Fig. S1 f). The ∆ex5 transcript was also detected in healthy control (HC) PHA blasts (3%) and fibroblasts (6%), indicating the presence of a naturally occurring low-abundance isoform (Fig. 1 b and Fig. S1 e), consistent with the transcripts documented in the Ensembl genome browser (ENST00000635082.1). We considered the possibility that c.829+5G>T represents a leaky splice variant; however, based on targeted RNA sequencing (RNA-seq) (Fig. S1 g), RT-PCR (Fig. S1 h), and reverse transcription quantitative PCR (RT-qPCR) (Fig. S1 i), we found no evidence supporting this. Although aberrant splicing arising from c.829+2T>G was not investigated, the consequences of c.829+2T>G are expected to coincide with c.829+5G>T as it inactivates the consensus splice donor site in intron 5, demonstrated by the in silico splice prediction tools (Table S2).
Reduced ATR protein in absence of full-length ATRIP
Alternative splicing of ATRIP pre-mRNAs harboring c.829+5G>T or c.829+2T>G variants results in ∆ex5 and a subsequent shift of the reading frame (r.671_829del). At the amino acid level, this translates into a serine to arginine substitution at position 224, immediately followed by the premature stop codon TAG (p.(Ser224ArgfsTer2)) (Fig. 1 d). Using western blot, we confirmed the absence of full-length ATRIP in both fibroblasts and Epstein–Barr virus immortalized lymphoblastoid cell lines (EBV-LCLs) of F1Pt (Fig. 1 e and Fig. S1, j and k). In the absence of full-length ATRIP, ATR expression was strongly reduced in both cell types, while parental LCLs from F1Fa showed an ∼50% reduction in both ATR and ATRIP protein levels. In line with the presumed co-dependency of ATRIP and ATR, fibroblasts from an ATR patient (F02-98) (O’Driscoll et al., 2003) exhibited strongly reduced ATRIP levels (Fig. 1 e and Fig. S1 k). In contrast, expression levels of interaction partners RPA1 and TOPBP1 were undisturbed by the absence of ATRIP (Fig. 1 e and Fig. S1, j and k).
Mutant ATRIP is LOF
The premature translation stop as a consequence of ∆ex5 is anticipated to trigger nonsense-mediated mRNA decay (NMD). When abolishing NMD using puromycin, equal quantities of both ∆ex5 (±380 bp, 51%) and full-length (±538 bp, 49%) amplicons were observed in cDNA from EBV-LCLs of F1Fa (Fig. 1 f). However, in the absence of puromycin, half of the ∆ex5 transcript persisted (±380 bp, 36% versus ±538 bp, 64%), indicating that the ∆ex5 transcript resulting from c.829+5G>T might translate into a truncated protein. Due to the unavailability of high-quality antibodies targeting the N-terminal region of ATRIP, we confirmed the presence of a truncated protein by overexpressing 3xFLAG-tagged wild type (wt) or mutant (mut) ATRIP, which resulted in a stable 82 and 37 kDa protein, respectively (Fig. 1 h). According to the remaining amino acid sequence (AA1–224), this 37 kDa ATRIP protein contains both the coiled-coil and RPA binding motif but lacks the TOPBP1 and ATR binding domains (Fig. 1 g) (Mordes et al., 2008). To assess this, HEK293T cells were co-transfected with ATR-V5, RPA1-Myc, and the 3xFLAG-wtATRIP or 3xFLAG-mutATRIP constructs. Upon pulldown with an anti-FLAG antibody, immunoblotting confirmed that the 37 kDa mutant was able to interact with RPA1, in accordance with earlier mutational studies (Ball et al., 2005; Namiki and Zou, 2006; Itakura et al., 2004). No ATR protein could be visualized, indicating an abolished interaction. Faint TOPBP1 was detected, but at levels comparable with those of the IgG controls, suggestive of a defective interaction with TOPBP1 (Fig. 1 h).
Presence of CD4+ T, B, NK lymphopenia, and autoimmune features with hemolytic anemia and neutropenia
Given the features of immunodeficiency in patients with ATRIP deficiency, we set out to characterize the immunophenotype of F1Pt. In line with the RTIs and a severe Varicella zoster infection, F1Pt presented with reduced numbers of B and NK cells (Fig. 2 a and Table 1). Low numbers of CD4+ T cells and an inverted CD4+/CD8+ ratio were noted (Fig. 2 a and Table 1). These features were unique to F1Pt, as both the heterozygous parents and the WT sister had immune profiles similar to HCs (Fig. 2 b). Immunoglobulin substitution therapy related to IgG2 subclass deficiency and a specific pneumococcal antibody deficiency, along with azithromycin maintenance, effectively controlled the RTIs (Fig. 2 c and Table 1). Over time, F1Pt developed a progressive neutropenia in the presence of antineutrophil cytoplasmic antibodies, which was responsive to emergency granulopoiesis during acute infections (Fig. 2 d). At 18 years of age, F1Pt presented with recurrent autoimmune hemolytic anemia (AIHA) that subsided upon treatment with corticosteroids and anti-CD20 monoclonal antibody (mAb) treatment (Fig. 2 e).
Loss of ATRIP is associated with an immune deficiency, characterized by CD4 + T cell lymphopenia and reduced B and CD16 + /CD56 dim NK lymphocytes. (a) Total numbers (No) of CD4+ T, CD8+ T, NK, and B cells in the peripheral blood from the ATRIP patient (F1Pt) over time. Shading indicates the age-based reference range. (b) FCM immunophenotyping of F1Pt, family members, and age-matched HCs. Percentages of CD4+ T, CD8+ T, NK, and B cells in PBMCs from F1Pt, sister (F1Si), mother (F1Mo), and father (F1Fa). Data represent one experiment, with each data point representing one biological replicate. Mean and SEM are shown. (c) IgG2 concentration (Conc.) in the peripheral blood from F1Pt over time. Shading indicates the age-based reference range. Immunoglobulin substation therapy around age 4 is indicated. (d) Total numbers (No) of neutrophils in the peripheral blood from F1Pt, demonstrating intermittent neutropenia. Shading indicates the age-based reference range. (e) Hemoglobulin concentration (Conc.) in the peripheral blood from F1Pt over time. Shading indicates an age-based reference range. Corticosteroid and anti-CD20 mAb treatment (aCD20) is indicated. (f) UMAP plot depicting cluster annotation of 25 unique T and NK subsets (left). Analysis was performed using concatenated 25-parameter FCM data of PBMCs obtained from HCs (n = 6) and F1Pt. Contour plots of HCs (middle, top) and F1Pt (middle, bottom). Bar graph showing the relative proportion of HCs and F1Pt within each T and NK subset cluster (right). (g) UMAP plot demonstrating cluster annotation of 18 unique B and innate subsets (left). Analysis was performed using concatenated 25-parameter FCM data of PBMCs obtained from HCs (n = 6) and F1Pt. Contour plots of HCs (middle, top) and F1Pt (middle, bottom). Bar graph depicting the relative contribution of HCs and F1Pt within each B and innate subset cluster (right). Data are representative of one experiment (f and g).
Loss of ATRIP is associated with an immune deficiency, characterized by CD4 + T cell lymphopenia and reduced B and CD16 + /CD56 dim NK lymphocytes. (a) Total numbers (No) of CD4+ T, CD8+ T, NK, and B cells in the peripheral blood from the ATRIP patient (F1Pt) over time. Shading indicates the age-based reference range. (b) FCM immunophenotyping of F1Pt, family members, and age-matched HCs. Percentages of CD4+ T, CD8+ T, NK, and B cells in PBMCs from F1Pt, sister (F1Si), mother (F1Mo), and father (F1Fa). Data represent one experiment, with each data point representing one biological replicate. Mean and SEM are shown. (c) IgG2 concentration (Conc.) in the peripheral blood from F1Pt over time. Shading indicates the age-based reference range. Immunoglobulin substation therapy around age 4 is indicated. (d) Total numbers (No) of neutrophils in the peripheral blood from F1Pt, demonstrating intermittent neutropenia. Shading indicates the age-based reference range. (e) Hemoglobulin concentration (Conc.) in the peripheral blood from F1Pt over time. Shading indicates an age-based reference range. Corticosteroid and anti-CD20 mAb treatment (aCD20) is indicated. (f) UMAP plot depicting cluster annotation of 25 unique T and NK subsets (left). Analysis was performed using concatenated 25-parameter FCM data of PBMCs obtained from HCs (n = 6) and F1Pt. Contour plots of HCs (middle, top) and F1Pt (middle, bottom). Bar graph showing the relative proportion of HCs and F1Pt within each T and NK subset cluster (right). (g) UMAP plot demonstrating cluster annotation of 18 unique B and innate subsets (left). Analysis was performed using concatenated 25-parameter FCM data of PBMCs obtained from HCs (n = 6) and F1Pt. Contour plots of HCs (middle, top) and F1Pt (middle, bottom). Bar graph depicting the relative contribution of HCs and F1Pt within each B and innate subset cluster (right). Data are representative of one experiment (f and g).
Expansion of T effector cells, low levels of CD16+/CD56dim NK, and absent MAIT and iNKT lineages
To study the impact of ATRIP deficiency on the lymphocyte compartment in more detail, we performed high-parametric flow cytometry (FCM) (Fig. 2, f and g) on peripheral blood mononuclear cells (PBMCs) of F1Pt and age-matched HCs. Among the CD4+ and CD8+ T cells in F1Pt, we noted the expansion of effector cells at the expense of naïve T cells (Fig. 2 f). Especially in the CD8+ T cell compartment, increased percentages of HLA-DR+ and CD27−CD28− T cells were noted, indicating ongoing activation and exhaustion. Similarly, CD95+ double-negative (DN) T cells were expanded. In addition to diminished naïve T cell levels, we noted the absence of both mucosal-associated invariant T (MAIT) and invariant NK T (iNKT) cells. The observed NK lymphopenia was characterized by a selective reduction of CD16+/CD56dim NK cells (Fig. 2 f). NK cytotoxic potential was assessed and revealed normal CD107a upregulation upon stimulation with K652. In line with the lower proportion of CD16+/CD56dim NK cells in F1Pt, perforin expression was reduced (data not shown). In a similar approach, the B cell and innate compartment were investigated in detail (Fig. 2 g). CD21lo B cells were expanded in F1Pt at the expense of CD21hi memory B cells and plasmablasts. In the innate compartment, we noted lower percentages of type 1 dendritic cells (DC1) and plasmacytoid (p)DCs. Among monocyte subsets, both inflammatory and CD169+ expressing monocyte subsets were increased. Manual gating of the FCM data confirmed the main findings of this analysis (Fig. S2, a–c).
Comprehensive immunological phenotyping of patient F1Pt reveals significant disruptions across T, B, and NK cell subsets . (a) Pie charts displaying the distribution of naïve T (CCR7+CD45RA+), TEM (CCR7−CD45RA−), T central memory (CCR7+CD45RA−), and terminally differentiated T effector (CCR7−CD45RA+) cells in CD8+ and CD4+ T cells of ATRIP patient (F1Pt) and HCs (n = 6). (b) Percentages of T and NK subsets in PBMCs of F1Pt and HCs (n = 6), based on manual gating of 25-parameter FCM data. Mean and SEM are shown. (c) Percentages of B and innate subsets in PBMCs of F1Pt and HCs (n = 6), based on manual gating of 25-parameter FCM data. Mean and SEM are shown. (d) Dotplot depicting signature genes defining the UMAP clusters shown in Fig. 3 a. (e) Circos plots showing the TRBV and TRAV pairing pattern of T cells of ATRIP patient (F1Pt) and HCs. (f) Frequency of unique T cell clones in F1Pt and HCs. (g) Distribution of the CDR3 region lengths of TCR-α and TCR-β clones of F1Pt and HCs T cells. (h) Circos plots demonstrating the IGH, IGK, and IGK pairing pattern of F1Pt and HCs B cells. (i) Frequency of unique B cell clones in F1Pt and HCs. (j) Distribution of the CDR3 region lengths of heavy and light chain of F1Pt and HCs B cells.
Comprehensive immunological phenotyping of patient F1Pt reveals significant disruptions across T, B, and NK cell subsets . (a) Pie charts displaying the distribution of naïve T (CCR7+CD45RA+), TEM (CCR7−CD45RA−), T central memory (CCR7+CD45RA−), and terminally differentiated T effector (CCR7−CD45RA+) cells in CD8+ and CD4+ T cells of ATRIP patient (F1Pt) and HCs (n = 6). (b) Percentages of T and NK subsets in PBMCs of F1Pt and HCs (n = 6), based on manual gating of 25-parameter FCM data. Mean and SEM are shown. (c) Percentages of B and innate subsets in PBMCs of F1Pt and HCs (n = 6), based on manual gating of 25-parameter FCM data. Mean and SEM are shown. (d) Dotplot depicting signature genes defining the UMAP clusters shown in Fig. 3 a. (e) Circos plots showing the TRBV and TRAV pairing pattern of T cells of ATRIP patient (F1Pt) and HCs. (f) Frequency of unique T cell clones in F1Pt and HCs. (g) Distribution of the CDR3 region lengths of TCR-α and TCR-β clones of F1Pt and HCs T cells. (h) Circos plots demonstrating the IGH, IGK, and IGK pairing pattern of F1Pt and HCs B cells. (i) Frequency of unique B cell clones in F1Pt and HCs. (j) Distribution of the CDR3 region lengths of heavy and light chain of F1Pt and HCs B cells.
TCR oligoclonality and altered antibody class switch recombination (CSR) in the absence of ATRIP
Given the diminished levels of naïve T cells and the absence of MAIT cells in F1Pt, we looked for potential disparities in the recombinational processes of antigen receptors. To study this in more detail, cellular indexing of transcriptomes and epitopes by sequencing (CITE-seq) combined with TCR and B cell receptor (BCR) sequencing of PBMCs, collected prior to anti-CD20 mAb treatment, was performed. CITE-seq profiling confirmed the observed immune defects (Fig. 3 a and Fig. S2 d). By visualizing both the usage and pairing of unique TRAV and TRBV genes, we observed a decreased diversity in the TCR repertoire in F1Pt compared with HCs (Fig. 3, b and c; and Fig. S2, e and f). Oligoclonality was most evident in CD8+ effector memory (TEM) cells, suggesting clonal expansion (Fig. 3, b and c). Using a similar analysis based on IGH transcripts, we observed no restrictions in BCR repertoire diversity (Fig. S2, h and i). Changes in complementarity-determining region 3 (CDR3) length and composition have been described in immunodeficiencies (Lee et al., 2016; Foth et al., 2021). Comparing the CDR3 length profiles of TRAV, TRBV, and IGH transcripts, we observed no consistent differences between F1Pt and HCs (Fig. 3 d and Fig. S2, g and j). These findings are indicative of competent V(D)J recombination, suggesting adequate repair by non-homologous end-joining (NHEJ) in the absence of ATRIP.
TCR oligoclonality in CD8 + T EM cells and altered CSR in absence of ATRIP. (a) CITE-seq profiling of PBMCs from HCs (n = 3) and ATRIP patient (F1Pt), identifying 25 immune subsets. UMAP visualization of pooled CITE-seq data of HCs and F1Pt, displaying the identified subsets (left). UMAP plot of HCs (middle, top) and F1Pt (middle, bottom). Ratio of the 25 subsets in HCs and F1Pt, ranked based on the prevalence in F1Pt (right). (b) Circos plots showing the TRBV and TRAV pairing pattern of CD8+ TEM cells of HCs and ATRIP patient (F1Pt). (c) Frequency of unique CD8+ TEM cell clones in HCs and F1Pt. (d) Distribution of the CDR3 region lengths of TCR-α and TCR-β clones from HCs and F1Pt CD8+ TEM cells. (e) Frequency of CSR junctions per S region in HCs (n = 3) (junctions; n = 8,305) and F1Pt (junctions; n = 2,758). Mean and SD are shown. (f) Proportion of non-productive junctions (inversional recombination) per S region in HCs (n = 3) (junctions; n = 652) and F1Pt (junctions; n = 225). Mean and SD are shown. (g) Pie charts demonstrating the microhomology usage at Sμ–Sα junctions in HCs (n = 3) and F1Pt. Data represent one experiment (a–g).
TCR oligoclonality in CD8 + T EM cells and altered CSR in absence of ATRIP. (a) CITE-seq profiling of PBMCs from HCs (n = 3) and ATRIP patient (F1Pt), identifying 25 immune subsets. UMAP visualization of pooled CITE-seq data of HCs and F1Pt, displaying the identified subsets (left). UMAP plot of HCs (middle, top) and F1Pt (middle, bottom). Ratio of the 25 subsets in HCs and F1Pt, ranked based on the prevalence in F1Pt (right). (b) Circos plots showing the TRBV and TRAV pairing pattern of CD8+ TEM cells of HCs and ATRIP patient (F1Pt). (c) Frequency of unique CD8+ TEM cell clones in HCs and F1Pt. (d) Distribution of the CDR3 region lengths of TCR-α and TCR-β clones from HCs and F1Pt CD8+ TEM cells. (e) Frequency of CSR junctions per S region in HCs (n = 3) (junctions; n = 8,305) and F1Pt (junctions; n = 2,758). Mean and SD are shown. (f) Proportion of non-productive junctions (inversional recombination) per S region in HCs (n = 3) (junctions; n = 652) and F1Pt (junctions; n = 225). Mean and SD are shown. (g) Pie charts demonstrating the microhomology usage at Sμ–Sα junctions in HCs (n = 3) and F1Pt. Data represent one experiment (a–g).
Affinity maturation and isotype switching of antibodies require efficient DNA recombination processes called somatic hypermutation and CSR, respectively. A potential role for ATR signaling in CSR has been suggested in previous studies, owing to its regulatory function in DNA replication and proliferation (Pan-Hammarström et al., 2006; Mokrani-Benhelli et al., 2013; Sun et al., 2021). Using a modified version of linear amplification–mediated high-throughput genome-wide translocation sequencing (LAM-HTGTS), we assessed CSR patterns in ATRIP-deficient PBMCs (Hu et al., 2016; Takada et al., 2024). Our results revealed an altered isotype switching profile in F1Pt, with decreased proportions of recombination between Sµ and Sγ1, Sγ2, and Sγ4 (Fig. 3 e). A marked increase in inversional Sµ–Sγ1 and Sµ–Sγ4 junctions was noted, indicating reduced efficiency in IgG subclass switching (Fig. 3 f). We observed increased usage of longer microhomologies (≥10 bp) at the Sµ–Sα switch junctions in F1Pt as compared with HCs (Fig. 3 g), indicating increased repair via alternative end joining. Given that alternative end joining depends on DNA end resection during the early S phase, these results are suggestive of disruptions in cell cycle dynamics (Saha et al., 2017).
ATRIP c.829+5G>T presents with a DNA repair signature and IFN-driven immune dysregulation
We performed gene set enrichment analysis (GSEA) to examine the differentially expressed genes in PBMCs of F1Pt compared with HCs. Among the most altered biological processes (adj P < 0.05; Log2: ±1), we noted a distinct DNA repair signature in PBMCs from F1Pt (Fig. 4, a and b; and Fig. S3, a and b), which was also evident in both T effector and NK subsets (Fig. S3, c and d). Likely related to F1Pt’s autoimmune phenotype, an upregulated IFN (both type I and II IFN) response was observed. In line with this, an elevated six-gene IFN-stimulated gene (ISG) score was observed, confirming dysregulated type I IFN signaling (Fig. S3 e) (Reynolds et al., 2019; Rice et al., 2013). Analysis of cytokine expression in serum samples collected during an episode of AIHA and after successful treatment with anti-CD20 mAb validated increased IFN signaling (IFNγ and IP10) and indicated both innate and T lymphocyte subset activation (IL-10, IL-18, IL-27, IL-31) (Fig. 4 c). While treatment of AIHA with corticosteroids and anti-CD20 mAb resulted in partial reduction of all cytokines, the Th2-signature cytokines IL–10 and IL–31 persisted. FCM analysis confirmed the response to anti-CD20 mAb therapy, with partial normalization of IFN-regulated proteins CD64 and CD169 on classical monocytes (Fig. 4 d). While effectively targeting B cells, anti-CD20 mAb treatment did not normalize NK and T cell subset percentages, nor did it normalize T cell maturation (Fig. S3, f–h). Additionally, treatment had minimal effect on T cell activation, as indicated by increased levels of ICOS, OX40, and PD1 (Fig. S3 i).
ATRIP deficiency presents with a DNA repair signature, IFN -driven immune dysregulation, and impaired T cell proliferation. (a) Heatmap representing the top 10 enriched hallmark gene sets (MSigDB) in PBMCs of F1Pt compared to HCs (n = 3). (b) Volcano plot showing the differentially expressed genes in PBMCs of F1Pt compared to HCs (n = 3). (c) Heatmap displaying the serum concentration of the cytokines IL–10, IL–21, IL–18, IP–10, IL–6, IL–17A, IL–31, IFNγ, IL–27, MIP31, ICAM–1, IL-1RA, Eotaxin-3, IL–22, and IL–1β in HCs (n = 13) and F1Pt before and after treatment with anti-CD20 mAb. (d) CD169 and CD64 expression on CD14+ monocytes of HCs (n = 6) and F1Pt before and after treatment with anti-CD20 mAb (aCD20). Histograms of a representative HC and F1Pt are shown. Bar plots depict median fluorescence (MFI). Mean and SEM are shown. (e) LTT demonstrates the proliferative reponse of F1Pt whole blood to various stimuli. Data shows at least two independent analyses. (f) Proliferation analysis of CD8+ and CD4+ PHA blasts from HCs (n = 9) and F1Pt. PBMCs were labeled with CTV and stimulated with PHA for 96 h (dark grey or orange) or not stimulated (light grey, control). PF and PI of three independent experiments are depicted, with each data point representing one biological replicate. Mean and SD are shown. ns: not significant, *P < 0.05, ***P < 0.001, ****P < 0.0001 (multiple unpaired t tests).
ATRIP deficiency presents with a DNA repair signature, IFN -driven immune dysregulation, and impaired T cell proliferation. (a) Heatmap representing the top 10 enriched hallmark gene sets (MSigDB) in PBMCs of F1Pt compared to HCs (n = 3). (b) Volcano plot showing the differentially expressed genes in PBMCs of F1Pt compared to HCs (n = 3). (c) Heatmap displaying the serum concentration of the cytokines IL–10, IL–21, IL–18, IP–10, IL–6, IL–17A, IL–31, IFNγ, IL–27, MIP31, ICAM–1, IL-1RA, Eotaxin-3, IL–22, and IL–1β in HCs (n = 13) and F1Pt before and after treatment with anti-CD20 mAb. (d) CD169 and CD64 expression on CD14+ monocytes of HCs (n = 6) and F1Pt before and after treatment with anti-CD20 mAb (aCD20). Histograms of a representative HC and F1Pt are shown. Bar plots depict median fluorescence (MFI). Mean and SEM are shown. (e) LTT demonstrates the proliferative reponse of F1Pt whole blood to various stimuli. Data shows at least two independent analyses. (f) Proliferation analysis of CD8+ and CD4+ PHA blasts from HCs (n = 9) and F1Pt. PBMCs were labeled with CTV and stimulated with PHA for 96 h (dark grey or orange) or not stimulated (light grey, control). PF and PI of three independent experiments are depicted, with each data point representing one biological replicate. Mean and SD are shown. ns: not significant, *P < 0.05, ***P < 0.001, ****P < 0.0001 (multiple unpaired t tests).
Transcriptomic and immune profiling indicates a DNA repair signature and interferon-driven immune activation in ATRIP deficient cells . (a) MSigDB hallmark gene sets differentially expressed in PBMCs from F1Pt compared to HCs (n = 3). Normalized enrichment score (NES) values of the gene sets are depicted. (b) Enrichment plots for two MSigDB hallmark gene sets differentially expressed in PBMCs from F1Pt compared to HCs (n = 3). The profile of the running enrichment score (ES) is depicted for both hallmark gene sets. (c) Heatmap displaying the top 20 enriched hallmark gene sets (MSigDB) in T effector cells of F1Pt compared to HCs (n = 3). (d) Heatmap showing the top 20 enriched altered hallmark gene sets (MSigDB) in NK cells of F1Pt compared to HCs (n = 3). (e) Relative expression of six ISGs in whole blood of patient F1Pt compared to HCs (n = 6) (left). For each sample, an ISG score was calculated from the median fold induction of the 6 ISGs (right). The mean ISG score of 6 HCs + 2 SD of the mean was calculated and ISG scores above this threshold (>3.793; indicated by the dotted line) was labeled as positive. (f) Contour plot showing CD4+ T cell maturation in HC and F1Pt. F1Pt (−) represents pre-treatment with anti-CD20 mAb (aCD20), F1Pt (+) represents post-treatment with anti-CD20. (g) Contour plot displaying CD8+ T cell maturation in HC and F1Pt. F1Pt (−) represents pre-treatment with anti-CD20, F1Pt (+) represents post-treatment with anti-CD20. (h) Frequencies of CD4+ T, CD8+ T, NK, and B cells in PBMCs from HCs (n = 18) and F1Pt before and after treatment with anti-CD20. Data represent one experiment, with each data point representing one biological replicate. Mean and SEM are shown. (i) ICOS, OX40, PD1, and CTLA4 expression on CD4+ T cells of HCs (n = 6) and F1Pt before and after treatment with anti-CD20 mAb. Bar plots display median fluorescence (MFI). Mean and SEM are shown.
Transcriptomic and immune profiling indicates a DNA repair signature and interferon-driven immune activation in ATRIP deficient cells . (a) MSigDB hallmark gene sets differentially expressed in PBMCs from F1Pt compared to HCs (n = 3). Normalized enrichment score (NES) values of the gene sets are depicted. (b) Enrichment plots for two MSigDB hallmark gene sets differentially expressed in PBMCs from F1Pt compared to HCs (n = 3). The profile of the running enrichment score (ES) is depicted for both hallmark gene sets. (c) Heatmap displaying the top 20 enriched hallmark gene sets (MSigDB) in T effector cells of F1Pt compared to HCs (n = 3). (d) Heatmap showing the top 20 enriched altered hallmark gene sets (MSigDB) in NK cells of F1Pt compared to HCs (n = 3). (e) Relative expression of six ISGs in whole blood of patient F1Pt compared to HCs (n = 6) (left). For each sample, an ISG score was calculated from the median fold induction of the 6 ISGs (right). The mean ISG score of 6 HCs + 2 SD of the mean was calculated and ISG scores above this threshold (>3.793; indicated by the dotted line) was labeled as positive. (f) Contour plot showing CD4+ T cell maturation in HC and F1Pt. F1Pt (−) represents pre-treatment with anti-CD20 mAb (aCD20), F1Pt (+) represents post-treatment with anti-CD20. (g) Contour plot displaying CD8+ T cell maturation in HC and F1Pt. F1Pt (−) represents pre-treatment with anti-CD20, F1Pt (+) represents post-treatment with anti-CD20. (h) Frequencies of CD4+ T, CD8+ T, NK, and B cells in PBMCs from HCs (n = 18) and F1Pt before and after treatment with anti-CD20. Data represent one experiment, with each data point representing one biological replicate. Mean and SEM are shown. (i) ICOS, OX40, PD1, and CTLA4 expression on CD4+ T cells of HCs (n = 6) and F1Pt before and after treatment with anti-CD20 mAb. Bar plots display median fluorescence (MFI). Mean and SEM are shown.
Impaired T cell proliferation in the absence of ATRIP
Considering that GSEA and CSR analyses indicated dysregulated DNA repair and cell cycle dynamics in F1Pt cells, we proceeded to investigate the proliferative capacity of T cells in more detail. The lymphocyte transformation test (LTT) performed on whole blood of F1Pt revealed reduced or absent T cell proliferation upon stimulation with both mitogens and specific antigens (candida, tetanus), respectively (Fig. 4 e). Using a more comprehensive study determining both the precursor frequency (PF) and proliferation index (PI) of CellTrace Violet (CTV)–labeled T cells, we noted a substantial reduction in the proliferation capacity of both CD4+ and CD8+ T cells of F1Pt upon stimulation with the mitogen PHA (Fig. 4 f). In conclusion, the immunophenotype of ATRIP deficiency is characterized by defects in all three lymphocytic lineages, an IFN-driven immune dysregulation, and a profound T cell proliferative defect.
Loss of ATRIP does not impair ATR recruitment and autophosphorylation
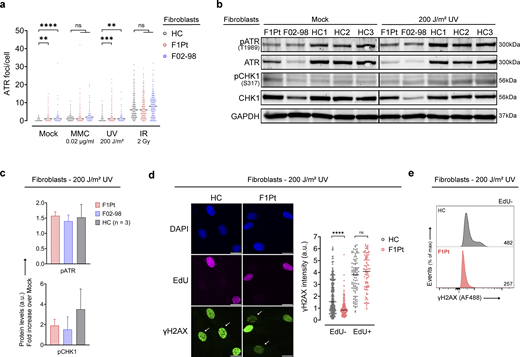
We set out to characterize the molecular processes driving disease in the absence of ATRIP. As localization of ATR to DNA damage sites is a prerequisite for ATR activation, we assessed nuclear ATR foci by immunofluorescence. Unexpectedly, this revealed that F1Pt fibroblasts were competent in forming ATR foci in response to the known ATR-activating DNA damage inducers mitomycin C (MMC), a DNA interstrand crosslinker (ICL), UV, and ionizing radiation (IR). In addition, F02-98 fibroblasts (ATR patient) showed normal recruitment of ATR (Fig. 5 a and Fig. S4 a), despite substantially reduced total ATR protein levels (11% of HC levels; Fig. S1 g). No significant differences in foci number per cell were observed between HC and F1Pt fibroblasts following MMC and IR treatment, whereas UV exposure resulted in a slightly higher amount of ATR foci in F1Pt and F02-98 fibroblasts. Notably, a higher number of spontaneous foci in mock conditions was observed, suggesting persistent DNA damage (Fig. 5 a and Fig. S4 a).
Absence of ATRIP does not abolish recruitment of ATR and its ability to phosphorylate substrates but reveals an insufficient ATR signaling response. (a) Fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98) were left untreated or exposed to 0.02 µg/ml MMC, 200 J/m2 UV, or 2 Gy IR. ATR was stained by immunofluorescence after 24 h (MMC) or 3 h (UV and IR) after exposure and ATR nuclear foci were quantified. Dot plot represents pooled data from three independent experiments; at least 150 cells were analyzed for each condition. The median number of foci is depicted. ns: not significant, **P < 0.01, ***P < 0.001, ****P < 0.0001 (Kruskall–Wallis test and Dunn’s multiple comparisons test). (b) Protein expression of phosphorylation events (T1989-pATR, S317-pCHK1) and total protein (ATR, CHK1) 3 h after 200 J/m2 UV radiation. Immunoblotting was performed on fibroblasts from F1Pt, F02-98, and HCs (n = 3). Western blot is representative of three independent experiments. GAPDH serves as a loading control. (c) pATR and pCHK1 levels shown in Fig. 2 b were quantified. Bar graph depicts pATR and pCHK1 levels post UV treatment, expressed as a fold increase over the levels observed in the Mock condition. Mean and SD are shown. (d) Quantification of γH2AX nuclear fluorescence in HC and F1Pt fibroblasts 3 h after 200 J/m2 UV exposure and concomitant EdU pulse-labeling. The mean γH2AX intensity per nucleus is shown for EdU− and EdU+ fibroblasts. Dot plot represents pooled data from three independent experiments. At least 200 (EdU−) or 90 (EdU+) cells were analyzed per condition. The median value is depicted. ns: not significant, ****P < 0.0001 (multiple Mann–Whitney tests and Bonferroni–Dunn multiple comparisons test). Representative immunofluorescence images with DAPI, EdU, and γH2AX staining are shown (left). White arrows indicate EdU− cells. Scale bars are 20 µm. (e) γH2AX expression was determined by flow cytometric analysis 3 h following 200 J/m2 UV exposure in EdU− (G0/G1 and G2/M phase) fibroblasts of HC and F1Pt. Median fluorescence intensity (MFI) of γH2AX (AF488) is annotated on the histogram. Data are reflective of one experiment. Source data are available for this figure: SourceData F5.
Absence of ATRIP does not abolish recruitment of ATR and its ability to phosphorylate substrates but reveals an insufficient ATR signaling response. (a) Fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98) were left untreated or exposed to 0.02 µg/ml MMC, 200 J/m2 UV, or 2 Gy IR. ATR was stained by immunofluorescence after 24 h (MMC) or 3 h (UV and IR) after exposure and ATR nuclear foci were quantified. Dot plot represents pooled data from three independent experiments; at least 150 cells were analyzed for each condition. The median number of foci is depicted. ns: not significant, **P < 0.01, ***P < 0.001, ****P < 0.0001 (Kruskall–Wallis test and Dunn’s multiple comparisons test). (b) Protein expression of phosphorylation events (T1989-pATR, S317-pCHK1) and total protein (ATR, CHK1) 3 h after 200 J/m2 UV radiation. Immunoblotting was performed on fibroblasts from F1Pt, F02-98, and HCs (n = 3). Western blot is representative of three independent experiments. GAPDH serves as a loading control. (c) pATR and pCHK1 levels shown in Fig. 2 b were quantified. Bar graph depicts pATR and pCHK1 levels post UV treatment, expressed as a fold increase over the levels observed in the Mock condition. Mean and SD are shown. (d) Quantification of γH2AX nuclear fluorescence in HC and F1Pt fibroblasts 3 h after 200 J/m2 UV exposure and concomitant EdU pulse-labeling. The mean γH2AX intensity per nucleus is shown for EdU− and EdU+ fibroblasts. Dot plot represents pooled data from three independent experiments. At least 200 (EdU−) or 90 (EdU+) cells were analyzed per condition. The median value is depicted. ns: not significant, ****P < 0.0001 (multiple Mann–Whitney tests and Bonferroni–Dunn multiple comparisons test). Representative immunofluorescence images with DAPI, EdU, and γH2AX staining are shown (left). White arrows indicate EdU− cells. Scale bars are 20 µm. (e) γH2AX expression was determined by flow cytometric analysis 3 h following 200 J/m2 UV exposure in EdU− (G0/G1 and G2/M phase) fibroblasts of HC and F1Pt. Median fluorescence intensity (MFI) of γH2AX (AF488) is annotated on the histogram. Data are reflective of one experiment. Source data are available for this figure: SourceData F5.
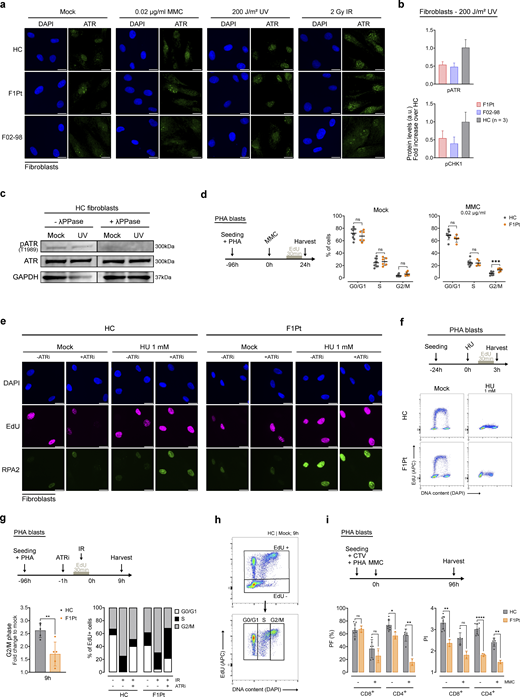
ATRIP deficiency disrupts ATR signaling, DNA replication, cell cycle progression, and T cell proliferation. (a) Representative immunofluorescence images of cells shown in Fig. 5 a with DAPI and ATR staining. Fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98) were left untreated or exposed to 0.02 µg/ml MMC, 200 J/m2 UV, or 2 Gy IR. ATR was stained by immunofluorescence following 24 h (MMC) or 3 h (UV and IR) of treatment. Images are representative of three independent experiments. Scale bars are 20 µm. (b) pATR and pCHK1 levels shown in Fig. 5 b were quantified and represent three independent experiments. Bar graph depicts pATR and pCHK1 levels post 200 J/m2 UV treatment, expressed as a fold increase over the mean levels observed in three HCs. Mean and SD are depicted. (c) Immunoblotting of T1989-pATR and total ATR with and without lambda phosphatase (λPPase) treatment on HC fibroblasts, untreated or 3 h after 200 J/m2 UV exposure. GAPDH serves as a loading control. (d) Cell cycle distributions of PHA blasts from HCs and ATRIP patient (F1Pt). Cells were either untreated or treated with 0.02 µg/ml MMC and subsequently harvested at the timepoints indicated on the schematics. Scatter dot plot depicts data from at least five independent experiments. Mean and SD are shown. ns: not significant, ***P < 0.001 (multiple paired t tests). (e) Representative immunofluorescence images with DAPI, EdU, and RPA staining of data shown in Fig. 6 d. HC and F1Pt fibroblasts were untreated or exposed to 1 mM HU for 3 h. Images are representative of three independent experiments. Scale bars are 20 µm. (f) Flow cytometric (FCM) EdU pulse-labeling profiles of HC and F1Pt fibroblasts demonstrating the inhibiting effect of HU treatment on replication progression. Cells were exposed to 1 mM HU for 3 h and subsequently harvested. Data are representative of two experiments. (g) EdU pulse-chase analysis after exposure to 4 Gy IR in the absence or presence of 20 nM ATRi. HC and F1Pt PHA blasts were harvested after 9 h. Bar plot (left) shows EdU+ cells present in G2/M phase after IR exposure, depicted as a fold change over the percentage observed in the mock condition. Data represent five independent experiments. Mean and SD are shown. **P < 0.01 (two-tailed paired t test). Bar plot (right) shows percentages of EdU+ cells present in G0/G1, S, and G2/M phase. Data from one experiment are shown. (h) FCM gating strategy of EdU pulse-chase kinetics presented in Fig. 6 g and Fig. S4 g. (i) PF and PI of CD8+ and CD4+ PHA blasts from HC and F1Pt. Cells were labeled with CTV and subsequently cultured for 96 h in the presence or absence of 0.02 µg/ml MMC. Data of at least two independent experiments are shown, with each data point representing one biological replicate. Mean and SD are shown. ns: not significant, *P < 0.05, **P < 0.01, ****P < 0.0001 (multiple paired t tests). Source data are available for this figure: SourceDataFS4.
ATRIP deficiency disrupts ATR signaling, DNA replication, cell cycle progression, and T cell proliferation. (a) Representative immunofluorescence images of cells shown in Fig. 5 a with DAPI and ATR staining. Fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98) were left untreated or exposed to 0.02 µg/ml MMC, 200 J/m2 UV, or 2 Gy IR. ATR was stained by immunofluorescence following 24 h (MMC) or 3 h (UV and IR) of treatment. Images are representative of three independent experiments. Scale bars are 20 µm. (b) pATR and pCHK1 levels shown in Fig. 5 b were quantified and represent three independent experiments. Bar graph depicts pATR and pCHK1 levels post 200 J/m2 UV treatment, expressed as a fold increase over the mean levels observed in three HCs. Mean and SD are depicted. (c) Immunoblotting of T1989-pATR and total ATR with and without lambda phosphatase (λPPase) treatment on HC fibroblasts, untreated or 3 h after 200 J/m2 UV exposure. GAPDH serves as a loading control. (d) Cell cycle distributions of PHA blasts from HCs and ATRIP patient (F1Pt). Cells were either untreated or treated with 0.02 µg/ml MMC and subsequently harvested at the timepoints indicated on the schematics. Scatter dot plot depicts data from at least five independent experiments. Mean and SD are shown. ns: not significant, ***P < 0.001 (multiple paired t tests). (e) Representative immunofluorescence images with DAPI, EdU, and RPA staining of data shown in Fig. 6 d. HC and F1Pt fibroblasts were untreated or exposed to 1 mM HU for 3 h. Images are representative of three independent experiments. Scale bars are 20 µm. (f) Flow cytometric (FCM) EdU pulse-labeling profiles of HC and F1Pt fibroblasts demonstrating the inhibiting effect of HU treatment on replication progression. Cells were exposed to 1 mM HU for 3 h and subsequently harvested. Data are representative of two experiments. (g) EdU pulse-chase analysis after exposure to 4 Gy IR in the absence or presence of 20 nM ATRi. HC and F1Pt PHA blasts were harvested after 9 h. Bar plot (left) shows EdU+ cells present in G2/M phase after IR exposure, depicted as a fold change over the percentage observed in the mock condition. Data represent five independent experiments. Mean and SD are shown. **P < 0.01 (two-tailed paired t test). Bar plot (right) shows percentages of EdU+ cells present in G0/G1, S, and G2/M phase. Data from one experiment are shown. (h) FCM gating strategy of EdU pulse-chase kinetics presented in Fig. 6 g and Fig. S4 g. (i) PF and PI of CD8+ and CD4+ PHA blasts from HC and F1Pt. Cells were labeled with CTV and subsequently cultured for 96 h in the presence or absence of 0.02 µg/ml MMC. Data of at least two independent experiments are shown, with each data point representing one biological replicate. Mean and SD are shown. ns: not significant, *P < 0.05, **P < 0.01, ****P < 0.0001 (multiple paired t tests). Source data are available for this figure: SourceDataFS4.
Next, ATR-dependent substrate phosphorylation was investigated as a measure of ATR activation. Autophosphorylation of ATR on residue T1989 has been identified as a hallmark of its active state and is regulated by DNA damage induction (Liu et al., 2011; Nam et al., 2011b). T1989 phosphorylation (hereafter referred to as pATR) was readily detected in untreated F1Pt and F02-98 cells and further increased upon UV treatment, although absolute pATR levels were reduced compared with control fibroblasts (Fig. 5 b and Fig. S4 b). Interestingly, when expressing UV-induced pATR levels as a fold increase over basal levels, a similar 1.5-fold induction was observed for F1Pt, F02-98, and control cells (Fig. 5 c). Of note, the specificity for T1989 phosphorylation was confirmed using lambda phosphatase (λPPase) treatment (Fig. S4 c). Full activation of CHK1, the major downstream effector of the ATR signaling pathway, requires phosphorylation by ATR (Saldivar et al., 2017). Following UV exposure, CHK1 phosphorylation on residue S317 (indicated as pCHK1) was reduced in both F1Pt and F02-98 fibroblasts compared with HCs (Fig. 5 b and Fig. S4 b). This reduction was also evident when the fold induction over basal levels was quantified (Fig. 5 c). Of note, total CHK1 levels were reduced in F02-98, but not in F1Pt fibroblasts (Fig. 5 b), a possible consequence of the high passage number and senescence of the F02-98 cells (Lossaint et al., 2022).
DNA repair intermediates formed after UV irradiation trigger H2AX phosphorylation at S139 (known as γH2AX), a process mainly mediated by ATR and found to be impaired in ATR patient cells (O’Driscoll et al., 2003; Stiff et al., 2006; Hanasoge and Ljungman, 2007; Mokrani-Benhelli et al., 2013). Following UV exposure, a strong pan-nuclear γH2AX positivity was readily noted via immunofluorescence in control fibroblasts (Fig. 5 d). In contrast, γH2AX was almost entirely absent in EdU− nuclei of F1Pt fibroblasts, while it was retained in EdU+ nuclei. Consistent with this, FCM analysis of γH2AX expression in EdU− cells showed decreased MFI values in F1Pt (Fig. 5 e). In conclusion, while the exact mechanism of ATR recruitment and activation remains unclear, our observations indicate that ATRIP is dispensable for the initial recruitment and subsequent autophosphorylation of ATR. Downstream ATR substrate phosphorylation was impaired but not abolished, providing evidence for an inadequate ATR signaling response in the absence of ATRIP.
Compromised replication stress response and cell cycle progression in the absence of ATRIP
Replication stress induces stalling of replication forks and exposes ssDNA that is rapidly coated with RPA. Subsequent ATR activation safeguards genomic stability and prevents further fork stalling and collapse by restraining fork progression and suppressing origin firing. In addition, the ATR kinase is responsible for stabilizing stalled forks, thereby facilitating fork restart upon resolved stress. We aimed to thoroughly define the in vitro phenotype associated with ATRIP deficiency by investigating the cellular consequences of genotoxic treatments.
Previous studies noted comparable cell cycle distributions in untreated cells of ATR patients and HCs (Ogi et al., 2012; Mokrani-Benhelli et al., 2013). Consistent with this, we observed no differences in cell cycle ratios between F1Pt and control fibroblasts or PHA blasts (Fig. 6, a and b; and Fig. S4 d). To investigate the replication stress response, we treated F1Pt fibroblasts with MMC and assessed the replicative response using FCM (Fig. 6 a). Whereas HC cells efficiently suppressed origin firing and fork progression upon MMC treatment (indicated by reduced EdU intensity), F1Pt cells showed no substantial response (Fig. 6 c). To verify ATR involvement, we pretreated cells with an ATR inhibitor (ATRi). Upon ATRi treatment, the EdU intensity of HC cells mimicked that of ATRIP-deficient cells, confirming impaired ATR signaling in F1Pt cells. Additionally, DAPI histograms of S phase cells showed accumulation of HC cells in the late S phase following MMC, in contrast to F1Pt cells that retained a diffuse DAPI profile similar to non-treated cells.
Loss of ATRIP results in compromised DNA replication and impaired cell cycle progression following replication stress. (a) Schematic outlining the treatment protocol of the FCM assay used in b and c. (b) Cell cycle distributions of untreated fibroblasts from a HC and ATRIP patient (F1Pt). Representative FCM plots are shown (left). Scatter dot plot depicts data from eight independent experiments (right). Mean and SD are shown. ns: not significant (multiple paired t tests). (c) FCM EdU pulse-labeling profiles of MMC treated HC and F1Pt fibroblasts (left). Cells were exposed to 0.02 µg/ml MMC for 24 h in the absence or presence of an ATR kinase inhibitor (ATRi, 20 nM). Histograms of EdU and DAPI intensity in S phase cells are shown (right). The median fluorescence intensity (MFI) is annotated on the histogram. Data are representative of three independent experiments. (d) Quantification of RPA nuclear fluorescence in HC and F1Pt fibroblasts 3 h after 1 mM HU exposure and concomitant EdU pulse-labeling. The mean RPA intensity per nucleus is shown for EdU+ cells. Dot plot represents data from pooled data from three experiments. The median value is depicted. ns: not significant; ****P < 0.0001 (multiple Mann–Whitney tests and Bonferroni–Dunn multiple comparisons test). (e) FCM EdU pulse-labeling profiles of HC and F1Pt fibroblasts after the release from HU treatment (left). Cells were exposed to 1 mM HU for 3 h, released for 3 h, and subsequently harvested. Histograms of EdU intensity in S phase cells are shown (right). Data are representative of two experiments. (f) Cell cycle profiles of HC and F1Pt fibroblasts following 72 h of 0.02 µg/ml MMC treatment, with and without 20 nM ATRi. The percentage of cells in G2/M phase is indicated. Data are representative of three independent experiments. (g) EdU pulse-chase kinetics of HC and F1Pt PHA blasts. Cells were untreated or treated with genotoxic inducers (0.02 µg/ml MMC or 200 J/m2 UV), pulse-labeled with EdU, and harvested at indicated time points. Kinetic plots show percentages of EdU+ cells present in S phase and are representative of three independent experiments. (h) CTV profiles of CD8+ and CD4+ PHA blasts from HC and F1Pt after 96 h of culture in the presence or absence of 0.02 µg/ml MMC. Data are representative of two experiments.
Loss of ATRIP results in compromised DNA replication and impaired cell cycle progression following replication stress. (a) Schematic outlining the treatment protocol of the FCM assay used in b and c. (b) Cell cycle distributions of untreated fibroblasts from a HC and ATRIP patient (F1Pt). Representative FCM plots are shown (left). Scatter dot plot depicts data from eight independent experiments (right). Mean and SD are shown. ns: not significant (multiple paired t tests). (c) FCM EdU pulse-labeling profiles of MMC treated HC and F1Pt fibroblasts (left). Cells were exposed to 0.02 µg/ml MMC for 24 h in the absence or presence of an ATR kinase inhibitor (ATRi, 20 nM). Histograms of EdU and DAPI intensity in S phase cells are shown (right). The median fluorescence intensity (MFI) is annotated on the histogram. Data are representative of three independent experiments. (d) Quantification of RPA nuclear fluorescence in HC and F1Pt fibroblasts 3 h after 1 mM HU exposure and concomitant EdU pulse-labeling. The mean RPA intensity per nucleus is shown for EdU+ cells. Dot plot represents data from pooled data from three experiments. The median value is depicted. ns: not significant; ****P < 0.0001 (multiple Mann–Whitney tests and Bonferroni–Dunn multiple comparisons test). (e) FCM EdU pulse-labeling profiles of HC and F1Pt fibroblasts after the release from HU treatment (left). Cells were exposed to 1 mM HU for 3 h, released for 3 h, and subsequently harvested. Histograms of EdU intensity in S phase cells are shown (right). Data are representative of two experiments. (f) Cell cycle profiles of HC and F1Pt fibroblasts following 72 h of 0.02 µg/ml MMC treatment, with and without 20 nM ATRi. The percentage of cells in G2/M phase is indicated. Data are representative of three independent experiments. (g) EdU pulse-chase kinetics of HC and F1Pt PHA blasts. Cells were untreated or treated with genotoxic inducers (0.02 µg/ml MMC or 200 J/m2 UV), pulse-labeled with EdU, and harvested at indicated time points. Kinetic plots show percentages of EdU+ cells present in S phase and are representative of three independent experiments. (h) CTV profiles of CD8+ and CD4+ PHA blasts from HC and F1Pt after 96 h of culture in the presence or absence of 0.02 µg/ml MMC. Data are representative of two experiments.
In the absence of timely ATR activation, excessive origin firing results in the accumulation of RPA-ssDNA nucleoprotein filaments, progressively depleting the nuclear pool of RPA and preceding compromised fork integrity (Toledo et al., 2013). Using immunofluorescence, we observed significantly higher RPA2 intensity in S phase F1Pt fibroblasts compared with HCs, both in mock and upon hydroxyurea (HU) treatment (Fig. 6 d and Fig. S4 e). HU induces widespread fork stalling through dNTP depletion, as evidenced by the complete loss of EdU incorporation after 3 h of HU treatment (Fig. S4 f) (Eklund et al., 2001). Additional treatment with ATRi further increased RPA2 intensity in both F1Pt and HCs, although the RPA2 intensity levels in HCs did not reach those observed in F1Pt.
Next, we determined whether patient fibroblasts retained their ability to recover from acute HU-induced replication stress. While HC cells readily resumed DNA synthesis, F1Pt fibroblasts demonstrated reduced levels of fork recovery following replication fork stalling, indicated by decreased EdU incorporation following HU release. Combined HU and ATRi treatment impeded replication recovery in HC cells and further intensified the observed impairment in F1Pt cells (Fig. 6 e).
Regulation of cell cycle progression is crucial to allow the repair of damaged DNA and the completion of DNA replication prior to mitosis. Pronounced G2/M arrest has been described as a hallmark of ATR deficiency, reflecting the direct consequence of acquired DNA damage during the S phase (Toledo et al., 2013). Accordingly, F1Pt fibroblasts and PHA blasts accumulated in the G2/M phase after MMC exposure (Fig. 6 f and Fig. S4 d). ATRi treatment in HC cells resulted in similar checkpoint activation, confirming that the observed G2/M arrest is a direct result of defective ATR signaling in the S phase (Fig. 6 f). An additional established function of ATR is the G2/M checkpoint activation in response to IR, preventing premature mitotic entry in the presence of severely damaged DNA (Fig. S4 g) (Cliby et al., 1998; Liu et al., 2000). In ATRIP-deficient cells, the ATR signaling pathway retained sufficient activity to initiate G2/M checkpoint arrest upon irradiation, albeit at a significantly lower level compared with HC cells (Fig. S4 g).
As our data did not indicate significant deviations in the cell cycle distribution of unperturbed patient cells, we extended our investigations to assess the S phase kinetics using an EdU pulse-chase assay (Fig. 6 g and Fig. S4 h) (Duthoo et al., 2022). We found that unperturbed F1Pt PHA blasts displayed a prolonged S phase compared with HC cells. Exposure to MMC and UV increased the S phase–delay in both F1Pt and HC cells, with a more pronounced effect in F1Pt cells. In light of these findings, we speculated that proficient ATR signaling is essential for facilitating T cell proliferation under conditions of excessive replicative stress. Indeed, upon treatment with MMC, we observed a strongly diminished proliferative response and subsequent division rate of CTV-labeled T cells of F1Pt (PF and PI, respectively), particularly in CD4+ T cells (Fig. 6 h and Fig. S4 i).
Taken together, defective ATR signaling in ATRIP-deficient cells results in an impaired replication stress response, consequently compromising cell cycle progression and proliferation.
ATRIP c.829+5G>T cells display DNA damage, chromosomal sensitivity, and impeded survival
The consequences of compromised replication and subsequent accumulation of DNA damage typically manifest after mitosis, resulting in chromosomal breakage and cell death. To assess spontaneously occurring DNA damage, we examined the formation of γH2AX nuclear foci, a well-established marker of DNA DSBs. Quantification of γH2AX in EdU− cells revealed a significantly higher number of foci in F1Pt and F02-98 fibroblasts (Fig. 7 a and Fig. S5 a). To evaluate DSB repair fidelity, we analyzed micronucleus (MN) formation using three different assays: the G0 MN assay, the MMC MN test, and a newly developed S MN assay (Fig. 7 b). F1Pt PHA blasts showed increased spontaneous MN levels with the S MN assay, as opposed to the normal levels observed with the G0 MN test (Fig. 7 c). This discrepancy suggests that DNA damage and subsequent chromosomal breakage only become apparent after multiple cell cycles have been completed. Correspondingly, immunofluorescence preparations of proliferating fibroblasts showed higher MN frequencies in F1Pt and F02-98 compared with HCs (data not shown).
ATRIP deficiency leads to increased spontaneous DNA damage, chromosomal sensitivity, and impaired survival upon exposure to genotoxic inducers requiring ATR pathway activation. (a) Immunofluorescence analysis of γH2AX foci in untreated EdU− fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98). γH2AX nuclear foci were quantified and are shown in a dot plot representing pooled data from three independent experiments. At least 150 cells were analyzed for each condition. The median number of foci is depicted. ****P < 0.0001 (Kruskall–Wallis test and Dunn’s multiple comparisons test). (b) Schematic of principle and treatment protocols of the MN assays depicted in c and d. Cytokinesis is blocked with cytochalasin B (CytoB), allowing scoring of MN in BN cells. (c and d) Using the G0 and S MN assay, micronuclei were scored in untreated PHA blasts (c) and in PHA blasts exposed to 1 Gy of IR (d). The MMC MN assay was used to quantify micronuclei in PHA blasts treated with 0.02 µg/ml MMC (d). Mean MN values of a HC group are indicated by dashed lines. Dotted lines correspond to the mean + 3 SD and serve as a cut-off for sensitivity. Data of two independent experiments are shown for F1Pt, in each experiment an internal control (IC) was included. One experiment is performed for the family members (F1Si, F1Fa, F1Mo). (e) Bar graph depicts the total percentage of cell death after 120 h of culture for untreated fibroblasts of HC, F1Pt, and F02-98. Mean and SD of at least three independent experiments is shown. (f) Schematic outlining of the treatment protocol of the cytotoxicity assay used in g and h. (g and h) Fibroblasts from HC, F1Pt, and F02-98 were untreated or exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) without (g) and with (h) kinase inhibitors (20 nM ATRi or 10 µM ATMi) as indicated. Cell death was monitored by live imaging up to 120 h by quantifying the number of cells stained with SYTOX Green. Cell death kinetics are reflective of at least two independent experiments. (i) Bar graph indicating % cell death after 120 h of treatment for the experiments shown in g and h.
ATRIP deficiency leads to increased spontaneous DNA damage, chromosomal sensitivity, and impaired survival upon exposure to genotoxic inducers requiring ATR pathway activation. (a) Immunofluorescence analysis of γH2AX foci in untreated EdU− fibroblasts from a HC, ATRIP patient (F1Pt), and ATR patient (F02-98). γH2AX nuclear foci were quantified and are shown in a dot plot representing pooled data from three independent experiments. At least 150 cells were analyzed for each condition. The median number of foci is depicted. ****P < 0.0001 (Kruskall–Wallis test and Dunn’s multiple comparisons test). (b) Schematic of principle and treatment protocols of the MN assays depicted in c and d. Cytokinesis is blocked with cytochalasin B (CytoB), allowing scoring of MN in BN cells. (c and d) Using the G0 and S MN assay, micronuclei were scored in untreated PHA blasts (c) and in PHA blasts exposed to 1 Gy of IR (d). The MMC MN assay was used to quantify micronuclei in PHA blasts treated with 0.02 µg/ml MMC (d). Mean MN values of a HC group are indicated by dashed lines. Dotted lines correspond to the mean + 3 SD and serve as a cut-off for sensitivity. Data of two independent experiments are shown for F1Pt, in each experiment an internal control (IC) was included. One experiment is performed for the family members (F1Si, F1Fa, F1Mo). (e) Bar graph depicts the total percentage of cell death after 120 h of culture for untreated fibroblasts of HC, F1Pt, and F02-98. Mean and SD of at least three independent experiments is shown. (f) Schematic outlining of the treatment protocol of the cytotoxicity assay used in g and h. (g and h) Fibroblasts from HC, F1Pt, and F02-98 were untreated or exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) without (g) and with (h) kinase inhibitors (20 nM ATRi or 10 µM ATMi) as indicated. Cell death was monitored by live imaging up to 120 h by quantifying the number of cells stained with SYTOX Green. Cell death kinetics are reflective of at least two independent experiments. (i) Bar graph indicating % cell death after 120 h of treatment for the experiments shown in g and h.
Loss of ATRIP increases DNA damage, chromosomal instability, and reduced cellular fitness. (a) Representative immunofluorescence images with DAPI and γH2AX staining of data shown in Fig. 7 a. γH2AX foci are shown in untreated fibroblasts from a control (HC), ATRIP patient (F1Pt), and ATR patient (F02-98). Images are representative of three independent experiments. Scale bars are 20 µm. (b) Using the G0 and S MN assay, micronuclei were scored in PHA blasts exposed to 0.5 Gy IR. Mean MN values of a reference HC group are indicated by dashed lines. Dotted lines correspond to the mean of HCs + 3 SD and serve as a cut-off for sensitivity to IR. Data of two independent experiments are shown for F1Pt, in each experiment an internal control (IC) was included. One experiment was performed for the family members (F1Si, F1Fa, F1Mo). (c and d) Fibroblasts from HC, F1Pt, and F02-98 were exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) in combination with a specific kinase inhibitor (2 µM DNA-PKi) (c) or only treated with 20 nM ATRi, 10 µM ATMi, or 2 µM DNA-PKi (d). Cell death was monitored by live imaging up to 120 h by quantifying the number of cells stained with SYTOX Green. Represented cell death kinetic plots are reflective of at least two independent experiments. (e and f) Corresponding proliferation curves of the data shown in Fig. 7, g and h. Fibroblasts from HC, F1Pt, and F02-98 were untreated or exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) without (e) or with (f) specific kinase inhibitors (20 nM ATRi or 10 µM ATMi) as indicated. Percentage of confluency was monitored by live imaging up to 120 h. Data are representative of at least two independent experiments. (g) Schematics of genome editing of the ATR and ATRIP allele (top). Blue lines represent the location of crRNAs, orange arrows depict the joint primers annealing to sequence outside the crRNA cutting regions, and red arrows represent the primers annealing to depletion region, used for wild type (WT) allele detection. The western blot verification of mono-allelic cells is shown (bottom). Cells were seeded in 6 cm dish, and 1 μg/ml doxycycline was added in the culture medium 24 h before cell lysis to induce the expression of cas9 protein. (h) Protein expression of ATR, ATRIP, RPA1, RPA2, and TOPBP1 in the novel generated MCF10A mono-allelic cells. β-Actin was used as loading control. (i) Cell fitness of MCF10A mono-allelic cells with frameshift mutations serving as internal control. Each variant has three independent biological replicates. Mean and SEM are shown. *P < 0.05, **P < 0.01, ***<0.001 (two-tailed paired t tests). Source data are available for this figure: SourceDataFS5.
Loss of ATRIP increases DNA damage, chromosomal instability, and reduced cellular fitness. (a) Representative immunofluorescence images with DAPI and γH2AX staining of data shown in Fig. 7 a. γH2AX foci are shown in untreated fibroblasts from a control (HC), ATRIP patient (F1Pt), and ATR patient (F02-98). Images are representative of three independent experiments. Scale bars are 20 µm. (b) Using the G0 and S MN assay, micronuclei were scored in PHA blasts exposed to 0.5 Gy IR. Mean MN values of a reference HC group are indicated by dashed lines. Dotted lines correspond to the mean of HCs + 3 SD and serve as a cut-off for sensitivity to IR. Data of two independent experiments are shown for F1Pt, in each experiment an internal control (IC) was included. One experiment was performed for the family members (F1Si, F1Fa, F1Mo). (c and d) Fibroblasts from HC, F1Pt, and F02-98 were exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) in combination with a specific kinase inhibitor (2 µM DNA-PKi) (c) or only treated with 20 nM ATRi, 10 µM ATMi, or 2 µM DNA-PKi (d). Cell death was monitored by live imaging up to 120 h by quantifying the number of cells stained with SYTOX Green. Represented cell death kinetic plots are reflective of at least two independent experiments. (e and f) Corresponding proliferation curves of the data shown in Fig. 7, g and h. Fibroblasts from HC, F1Pt, and F02-98 were untreated or exposed to genotoxic inducers (0.2 µg/ml MMC, 20 J/m2 UV, or 10 Gy IR) without (e) or with (f) specific kinase inhibitors (20 nM ATRi or 10 µM ATMi) as indicated. Percentage of confluency was monitored by live imaging up to 120 h. Data are representative of at least two independent experiments. (g) Schematics of genome editing of the ATR and ATRIP allele (top). Blue lines represent the location of crRNAs, orange arrows depict the joint primers annealing to sequence outside the crRNA cutting regions, and red arrows represent the primers annealing to depletion region, used for wild type (WT) allele detection. The western blot verification of mono-allelic cells is shown (bottom). Cells were seeded in 6 cm dish, and 1 μg/ml doxycycline was added in the culture medium 24 h before cell lysis to induce the expression of cas9 protein. (h) Protein expression of ATR, ATRIP, RPA1, RPA2, and TOPBP1 in the novel generated MCF10A mono-allelic cells. β-Actin was used as loading control. (i) Cell fitness of MCF10A mono-allelic cells with frameshift mutations serving as internal control. Each variant has three independent biological replicates. Mean and SEM are shown. *P < 0.05, **P < 0.01, ***<0.001 (two-tailed paired t tests). Source data are available for this figure: SourceDataFS5.
Next, we assessed chromosomal sensitivity to IR and MMC. No increase in radiation-induced MN yields was observed in F1Pt PHA blasts with the G0 MN assay, indicating a proficient NHEJ DSB repair pathway. Irradiation during the S phase led to a higher yield of MN (Fig. 7 d and Fig. S5 b), highlighting a specific S phase sensitivity to radiation in the absence of ATRIP, a phenotype not earlier associated with cells deficient in ATR signaling (O’Driscoll et al., 2003; Murga et al., 2009). As anticipated, F1Pt cells also showed impaired ICL repair, reflected by the increase in MN formation in the presence of MMC (Fig. 7 d). No increase in MN levels was detected in PHA blasts of the parents using both the MMC and S MN assay (Fig. 7, c and d; and Fig. S5 b), indicating that the heterozygous presence of the c.829+5G>T variant does not compromise ATR signaling.
Notably, persistent DNA damage accumulated during unperturbed replication in F1Pt and F02-98 fibroblasts negatively affected cell survival (Fig. 7, e and g). In addition, we meticulously quantified cell death and cell proliferation in the absence of functional ATR signaling following DNA damage induction by MMC, UV, or IR (Fig. 7, f–i; and Fig. S5, e and f). Treatment with 0.2 µg/ml MMC and 20 J/m2 UV revealed a profound sensitivity of F1Pt and F02-98 fibroblasts, whereas cell survival of control fibroblasts was slightly impaired by MMC and unaffected by UV (Fig. 7 g). These results confirm the previously described MMC and UV hypersensitivity in the absence of functional ATR signaling (O’Driscoll et al., 2003; Mokrani-Benhelli et al., 2013; Murga et al., 2009). In line with the G0 and S MN assay results of PHA blasts, exposure to 10 Gy IR only elicited a slight increase in cell death in F1Pt and F02-98 fibroblasts (Fig. 7 g). Combined treatment of ATRi and a DNA damage inducer progressively compromised cell survival. Although minor in the case of F1Pt and F02-98 fibroblasts, this increase was consistently observed for all three genotoxic treatment conditions, suggesting residual ATR kinase activity (Fig. 7 h).
In contrast to F1Pt and F02-98 fibroblasts, the combined treatment of ATRi with genotoxic stressors profoundly impaired cell survival in HC fibroblasts (Fig. 7 h). To assess redundancy with the other DDR kinases, we quantified cell death rates in the presence of specific inhibitors targeting ATM and DNA-PKcs. ATM inhibition (ATMi) resulted in particularly high lethality in combination with ATRIP deficiency (F1Pt) or hypomorphic ATR variants (F02-98), contrasting with the response observed in HC fibroblasts (Fig. 7 h and Fig. S5 d). This was most evident upon IR-induced genotoxic stress, implying a substantial redundancy between the ATR and ATM kinase. Concomitant DNA-PK inhibition with genotoxic treatment did not evoke additional sensitivities in F1Pt, F02-98, or HC fibroblasts (Fig. S5, c and d). In conclusion, these findings demonstrate that loss of ATRIP results in chromosomal sensitivity and compromised cell viability, although some residual ATR signaling was retained. Moreover, concurrent ATM kinase activity is needed to mitigate the most detrimental effects of genotoxic stressors, as previously suggested for ATR-deficient cells (Mokrani-Benhelli et al., 2013; Murga et al., 2009).
Modeling of ATRIP variants by CRISPR-SelectTIME exposes reduced cell fitness
Finally, to confirm the impact of the c.829+5G>T and c.829+2T>G ATRIP variants in an independent cellular model, cell fitness was assessed using the CRISPR-SelectTIME methodology (Niu et al., 2022). To accurately study the effects of these variants, an MCF10A cell line harboring only one functional ATRIP or ATR allele was generated (Fig. 8 a; and Fig. S5, g and h). In addition to the variants identified in this study, a hypomorphic leaky splice variant in ATR (c.2022A>G, homozygous in F02-98 [O’Driscoll et al., 2003]), the proposed causal ATRIP variants identified in patient CV1720 (Ogi et al., 2012) (c.2278C>T and c.248-14A<G), and a homozygous benign ATRIP splice variant present in gnomAD v4.0.0 (c.2056-6_2056-3del) were modeled. As expected, the presence of the c.2022A>G ATR variant resulted in strongly reduced cell fitness (Fig. 8 b). CRISPR-SelectTIME revealed similarly large effect sizes for both ATRIP splice variants, c.829+5G>T (homozygous in F1Pt) and c.829+2T>G (homozygous in F46.1). Moreover, we confirmed the anticipated neutrality of the homozygous ATRIP variant reported in gnomAD v4.0.0 (c.2056-6_2056-3del). Surprisingly, for the reported ATRIP variants of patient CV1720, reduced fitness was only evident for the c.2278C>T variant. Although abnormal splicing was observed in the presence of the second ATRIP allele of patient CV1720, we were unable to validate the causality of the candidate variant c.248-14A>G using CRISPR-SelectTIME (Fig. 8 c and Fig. S5 i), which might reflect the originally reported hypomorphic nature of this variant (Ogi et al., 2012). Given the role of both ATR and ATRIP in regulating DNA replication and proliferation dynamics, application of the CRISPR-SelectTIME methodology in a monoallelic ATRIP cell line appeared particularly suitable to establish the pathogenic role of the identified ATRIP variants c.829+5G>T and c.829+2T>G, next to the known c.2278C>T variant. Moreover, frequencies of these variants and their corresponding internal controls (frameshifts) were strikingly comparable, indicating similar selection against both editing outcomes (Fig. 8 c and Fig. S5 i).
Quantitative functional assay identifies fitness defect in genome-engineered ATRIP c.829+5G>T and ATRIP c.829+2T>G cells. (a) Schematic representation of CRISPR-SelectTIME process for cell fitness testing. The depletion region of the ATR and ATRIP genes are indicated. The CRISPR-Select cassettes for all the variants were transfected on day 2. Yellow cross indicates the various editing outcomes on target gene. The cell samples from day 2 (D2) and day 12 (D12) were collected and lysed for DNA extraction. PCR amplified the target edited sites of each sample and products were analyzed by NGS sequencing. (b) Schematic illustration of the modeled compound heterozygous (hez) and homozygous (hoz) ATRIP variants at the transcript level. (c) Cell fitness of MCF10A mono-allelic cells with the ATR and ATRIP variants. Each symbol within the bar represents an independent experiment. Each variant has three independent biological replicates. Mean and SEM are shown. Data of D12 is normalized to D2. **P < 0.01, ***P < 0.001, ns: non-significant (two-tailed paired t tests).
Quantitative functional assay identifies fitness defect in genome-engineered ATRIP c.829+5G>T and ATRIP c.829+2T>G cells. (a) Schematic representation of CRISPR-SelectTIME process for cell fitness testing. The depletion region of the ATR and ATRIP genes are indicated. The CRISPR-Select cassettes for all the variants were transfected on day 2. Yellow cross indicates the various editing outcomes on target gene. The cell samples from day 2 (D2) and day 12 (D12) were collected and lysed for DNA extraction. PCR amplified the target edited sites of each sample and products were analyzed by NGS sequencing. (b) Schematic illustration of the modeled compound heterozygous (hez) and homozygous (hoz) ATRIP variants at the transcript level. (c) Cell fitness of MCF10A mono-allelic cells with the ATR and ATRIP variants. Each symbol within the bar represents an independent experiment. Each variant has three independent biological replicates. Mean and SEM are shown. Data of D12 is normalized to D2. **P < 0.01, ***P < 0.001, ns: non-significant (two-tailed paired t tests).
Discussion
In this study, we demonstrated that biallelic LOF ATRIP variants result in human disease, characterized by short stature, microcephaly, ID, and immunodeficiency. The causal role of the pathogenic ATRIP variants is supported by multiple lines of evidence: (1) the presence of homozygous novel or rare variants with a similar LOF consequence on protein level in two patients of independent ancestry, (2) the complete absence of homozygous LOF ATRIP variants in the non-affected population (cf. gnomAD), (3) the finding of an impaired replication stress response associated with disturbed proliferation and survival in primary patient cells, and (4) reduced cell fitness when introducing these variants in an independent cellular model.
Ablation of ATR, TOPBP1, or CHK1 in murine models has consistently resulted in embryonic lethality, emphasizing the essential nature of the ATR signaling pathway (de Klein et al., 2000; Brown and Baltimore, 2000; Jeon et al., 2011; Zhou et al., 2013; Liu et al., 2000). This is further illustrated by the identification of exclusively hypomorphic ATR mutations in patients with MPD (Llorens-Agost et al., 2018; Mokrani-Benhelli et al., 2013; Ogi et al., 2012; O’Driscoll et al., 2003). In a similar vein, it has been suggested that ATRIP deficiency would be incompatible with life. However, our study demonstrates that LOF ATRIP variants are tolerated to some extent in human individuals, albeit resulting in a severe MPD phenotype. In line with our findings, knock-out atrip zebrafish were also shown to be viable but displayed a severely compromised body length and lifespan, suggesting a crucial role of atrip in growth and development during the juvenile stage (Shin et al., 2021).
ATRIP plays a crucial role in facilitating the stabilization, recruitment, and activation of the ATR kinase (Cortez et al., 2001; Zou and Elledge, 2003; Mordes et al., 2008). Studies using both cellular models and patient cells have shown that partial or complete ATRIP loss results in a significant reduction in ATR protein levels (Ball et al., 2005; Cortez et al., 2001; Ogi et al., 2012). This co-dependency between ATR and ATRIP for protein stability is further evidenced by our data, showing markedly diminished ATR levels in the complete absence of ATRIP. Several lines of evidence have demonstrated that ATRIP acts as a sensor of RPA-ssDNA nucleoprotein filaments, promoting ATR recruitment and subsequent activation of downstream substrates (Zou and Elledge, 2003; Bass et al., 2016). Interestingly, an early mutational study detected efficient CHK1 phosphorylation in the presence of mutated ATRIP devoid of its RPA-binding domain (Ball et al., 2005). Correspondingly, our findings revealed that the remaining pool of ATR was able to form foci and autophosphorylate (T1989 phosphorylation) upon exposure to genotoxic stressors in the absence of ATRIP, although at lower levels. These results suggest that ATRIP is essential for ATR stability but not for ATR recruitment and activation. While both TOPBP1 and ETAA1 are able to activate ATR, only the ATR-TOPBP1 axis requires ATRIP for recruitment to RPA-ssDNA filaments (Bass et al., 2016; Mordes et al., 2008; Haahr et al., 2016). It remains to be clarified whether binding and activation of ATR by ETAA1 is retained upon ATRIP loss. We speculate that interactions with ETAA1 and potentially other unanticipated complex partners may underlie the ability of ATR to recognize DNA damage and display kinase activity in an ATRIP-independent manner.
Studies using murine models have provided compelling evidence implicating ATR signaling in the development, maturation, and maintenance of the immune system (Murga et al., 2009; Ruzankina et al., 2007; Schuler et al., 2017; Schoeler et al., 2019; Menolfi et al., 2018), an association which had not been clearly described in humans to date. These mice consistently presented with developmental abnormalities analogous to MPD and features of premature aging, demonstrating that unresolved replicative stress, proliferative failure, and cellular senescence likely underlie both the developmental and immunological phenotype. The association of MPD with immunodeficiency is further supported by an emerging class of inborn errors of immunity (IEI) linked to monogenic defects in DNA replication factors (Conde et al., 2019; Cottineau et al., 2017; Gineau et al., 2012; Frugoni et al., 2016; Cui et al., 2020; Willemsen et al., 2023). These disorders typically present a combination of immunodeficiency with perturbed growth, microcephaly, progeroid features, and developmental abnormalities. All three ATRIP patients had clinical features suggestive of immunodeficiency, and the immunophenotype—characterized by lymphopenia, impaired proliferation, and autoimmunity with dysregulated IFN signaling—draws intriguing parallels to observations in other DNA replication–associated disorders. Our data revealed that ATRIP loss compromises the replication stress response, leading to a DNA repair signature in immune cells in vivo and impaired lymphocyte proliferation in vitro. Despite our observations in ATRIP deficient patients, no definite immunological phenotype has been observed in patients with biallelic ATR mutations (based on personal correspondence with respective clinicians). Although Mokrani-Benhelli et al. (2013) noted minor immune irregularities in a 9-year-old patient with biallelic ATR mutations, drawing parallels to our patient F1Pt, the phenotype was relatively mild and no up-to-date immunological data could be obtained. Several explanations for these disparate observations are possible: (1) the hypomorphic nature of the ATR mutations may mitigate the impact of ATR deficiency on the human immune system, (2) ATR signaling in immune cells such as lymphocytes is critically dependent on ATRIP, and (3) ATRIP has ATR-independent functions in the human immune system. In line with this, considerable heterogeneity in immunological features is also observed in both DNA repair and DNA replication-associated IEI, highlighting that mutations within the same cellular processes can lead to diverse phenotypes (Cottineau et al., 2017; Cui et al., 2020; Guo et al., 2015; Willemsen et al., 2023). Finally, several older studies reported immunological and hematological abnormalities, such as pancytopenia, in patients with a phenotypical diagnosis of Seckel syndrome (Butler et al., 1987; Woods et al., 1995; Arnold et al., 1999; Hayani et al., 1994). It is worth noting, however, that the molecular causes of Seckel syndrome had not yet been identified at that time, potentially leading to misdiagnoses due to overlapping features with other MPD-like disorders (Majewski et al., 1982; Thompson and Pembrey, 1985). More studies, comprehensively describing the immunological phenotype in molecularly characterized disorders of DNA replication and DNA repair, will be required to deepen our understanding of the role of ATR and ATRIP in the biology of immune cells.
In summary, our study provides evidence supporting ATRIP as a disease-associated gene in MPD and IEI and offers valuable insights into the cellular and immunological characteristics. Our findings challenge the conventional paradigm of the ATR–ATRIP interaction, underscoring the need for further investigation into the molecular mechanisms governing ATR activation upon DNA damage. Although our data suggests an essential role for ATRIP in safeguarding immune cells against the consequences of unresolved replicative stress, it remains unclear whether this reflects an unidentified noncanonical function of ATRIP or whether this is mediated by the ATR signaling pathway. Careful clinical monitoring and additional research are needed to further delineate the phenotypic spectrum associated with both biallelic ATR and ATRIP mutations and to better understand the cellular mechanisms driving the immunological phenotype.
Materials and methods
Case descriptions
Patient F1Pt, a girl of Belgian ancestry, was born at 39 wk of gestation with low birth weight (1,450 g), small stature, and reduced head circumference (40 and 27.5 cm, respectively) (Fig. S1 b). In the 15th wk of gestation, intrauterine growth retardation and oligohydramnios were observed. Amniocentesis revealed a normal karyotype. The pregnancy was complicated by maternal hypothyroidism due to Hashimoto’s disease, for which levothyroxine supplementation was initiated. Additionally, antibiotics were administered for perinatal chorioamnionitis. At birth, ophthalmological examinations and brain MRI did not show significant abnormalities. Skeletal radiography revealed osteoporotic changes in the bones, delayed bone age, and the presence of 11 pairs of ribs, without other unusual skeletal developmental issues. Ultrasound imaging of the kidneys demonstrated poor corticomedullary differentiation. Shortly after birth, the patient experienced feeding difficulties necessitating enteral nutrition until the age of 2. Tooth eruption was delayed, with the first tooth appearing after the age of 1. Postnatal growth retardation was significant and a trial of growth hormone treatment only marginally adjusted growth velocity. During early development, speech acquisition was delayed and comprehensive neurodevelopmental testing at a later stage revealed a mild ID with a Terman IQ score of 59. At 2 years of age, she developed a severe Varicella zoster infection, necessitating hospitalization and intravenous acyclovir treatment. At the age of 3, the patient began experiencing recurrent and persistent infections, including multiple episodes of bronchopneumonia, bilateral otitis media, gram-negative sepsis, campylobacter enteritis, and prolonged fever, accompanied by hemorrhagic stomatitis. These infections necessitated hospital admission and continuous antibiotic therapy. By the age of 4, prophylactic antibiotics were insufficient in managing the patient’s symptoms, and immunoglobulin substitution therapy was initiated. Computed tomography of the lungs showed mild bilateral bronchiectasis and fibrosis of the lung apices. The addition of azithromycin had a beneficial effect on lung inflammation. Immunological laboratory results indicated a mild leukopenia with significant lymphopenia, and a combined immune deficiency was diagnosed, characterized by decreased CD4+ T lymphocytes and reduced NK cells, an inverted CD4+/CD8+ T lymphocyte ratio, IgG2 deficiency, progressive neutropenia, and a deficiency in pneumococcal vaccine response (23 serotypes). Prophylaxis with sulfamethoxazole/trimethoprim was commenced. From age 14 onwards, a persistent EBV infection with a viral load of ∼5,000 copies per μg of DNA and hepatosplenomegaly was noted. At the age of 15, she developed severe normocytic anemia (7.3 g/dl). Bone marrow aspirate and trephine biopsies were performed. The erythroid cell displayed clear megaloblastoid changes and abnormal nuclear changes. In the case of the leukocytes, investigations revealed both hypoplasia and hypercellularity on different occasions. The examined leukocytes displayed a leftward shift. There were no increased blasts. The megakaryocytic lineage displayed aplasia in several aspirates. Trephine biopsies did not reveal signs of fibrosis (normal reticulin stain). AIHA based on cold agglutinins was diagnosed and initially treated with pulses of corticosteroids. The autoimmune anemia relapsed twice, first at 17 and then at 18 years of age, and concomitant thrombocytopenia and neutropenia were observed, prompting B cell eradication with rituximab (anti-CD20 mAb therapy), effectively restoring levels of all lineages. To date, at the age of 22, no malignant manifestations have been identified in the patient. No other family members present with a similar short stature, and, to our knowledge, there is no increased incidence of immunodeficiency or malignancy in the family. In adulthood, the patient presents as a delicate individual with proportionate short stature persistently below the third percentile throughout development. She reached a final height of 133 cm and a weight of 24.8 kg. Severe progressive microcephaly is evident, characterized by a mild sloping forehead, microphthalmia with convergent strabismus, and horizontal palpebral fissures. Additionally, she exhibits a prominent nose with a high nasal bridge, a short philtrum, and micrognathia. Her palate is normally developed, although dental crowding is observed. The shape of her ears is normal with attached earlobes but positioned rather low. Subtle hypopigmentation, a small single café-au-lait spot, and reduced subcutaneous fat content are also noted. Extremities appear normal without brachydactyly or abnormal palmar creases, although clinodactyly of the fifth digit is present bilaterally. She has a high-pitched voice and a friendly, outgoing personality.
Of note, several differential diagnoses for F1Pt have been excluded over the years: Bloom syndrome (based on absence of sister chromatid exchanges and screening of the BLM gene), Fanconi anemia (based on a historically inconclusive MMC test but normal exome sequencing), Nijmegen Breakage syndrome (based on absence of increased chromosome breaks in non-cycling cells after irradiation), and LIG4 syndrome (based on gene sequencing). Furthermore, microcephalic primary dwarfism syndromes like Seckel syndrome, microcephalic/Majewski osteodysplastic primordial dwarfism type II (MOPD II) (PCNT gene) were suggested, as well as cystic fibrosis. Exome-wide de novo filtering on trio WES data (F1Pt with parents) did not reveal other candidate genes (Table S4). Cytogenetically, she has a normal female karyotype (46, XX), and microarray-based comparative genomic hybridization (array-CGH) did not reveal any aberrations.
Patient F46-1, a boy of North Indian ancestry, was recently identified in a cohort of patients with microcephaly (Masih et al., 2022). He presented with severe microcephaly (OFC: −9.4 SD) and a short stature. He had a developmental delay and a moderate ID. He presented with a sloping forehead, a pronounced beak-like nose, micrognathia, and low-set ears. He was known with recurrent RTIs (around four per year) with at least one documented case of otitis media and gastroenteritis at the age of two, requiring hospital admission and intravenous fluids (information acquired through personal communication with authors). Unfortunately, the patient succumbed to his illness at 6 years of age, and samples for further investigation were unavailable.
Patient CV1720, a boy of Gujarati-Indian origin, was reported in 2012 as the first identified ATRIP-deficient MPD patient (Ogi et al., 2012). He was born at term via normal vaginal delivery with a reduced head circumference (27.1 cm) and low birth weight (2,060 g) (Fig. S1 b). Early developmental assessments showed delays in growth and development, with clinical features including severe microcephaly, small earlobes, micrognathia, and dental crowding. He was evaluated by an ophthalmologist and prescribed glasses. He was referred to a rheumatologist for possible Raynaud’s phenomenon, but no features of juvenile idiopathic arthritis were noted. Between the ages of 10 and 14, he presented with late-onset lymphopenia and intermittent neutropenia, with mild CD8+ T cell lymphopenia at the age of 13. His hemoglobin and platelet counts remained normal. He has no history of early childhood infections and has never been hospitalized due to infections. He continues to be routinely monitored, with no current need for hematological intervention.
Study approval
Ethical approval for this study was granted by the ethics committee of Ghent University Hospital in Ghent, Belgium (2012/593 and 2019/1565). Clinical data and samples were collected with informed consent from the participants of the study, in accordance with the 1975 Helsinki Declaration.
Cell culture
LCLs were derived from blood following EBV transformation. LCLs were maintained in Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with GlutaMAX, 10% FBS (Bodinco and Tico Europe), 1% penicillin–streptomycin (10,000 U/ml; Gibco), 1 mM sodium pyruvate (Gibco), and 50 μM 2-mercaptoethanol (Gibco). PHA blasts were generated by culturing cryopreserved isolated PBMCs in the LCL medium supplemented with 2% PHA-M (Gibco). HEK293T cells (The American Type Culture Collection [ATCC]) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS, 1% penicillin–streptomycin (10,000 U/ml), and 2 mM GlutaMAX (Gibco). Primary human dermal fibroblast cells were cultured in DMEM supplemented with 10% FBS, 1% penicillin–streptomycin (10,000 U/ml), 2 mM GlutaMAX, 0.1 mM sodium pyruvate, and 50 µM 2-mercaptoethanol. MCF10A cells (CRL-10317; ATCC) were cultured in Dulbecco’s Modified Eagle Medium:Nutrient Mixture F-12 (DMEM/F-12) (31330038; Thermo Fisher Scientific), supplemented with 5% horse serum (16050122; Thermo Fisher Scientific), 1% penicillin–streptomycin (15070063; Thermo Fisher Scientific), 10 μg/ml insulin (I1882; Sigma-Aldrich), 0.5 μg/ml hydrocortisone (H0888-1G; Sigma-Aldrich), 20 ng/ml EGF (AF-100-15; Peprotech), and 100 ng/ml cholera toxin (C8052-5MG; Sigma-Aldrich). ATR-deficient fibroblasts from a previously described ATR patient (F02-98) were purchased from Coriell Institute (GM18366) (O’Driscoll et al., 2003). All cell lines were incubated at 37°C with 5% CO2.
Sequencing analysis
Trio WES on the genomic DNA of the patient and unaffected parents was performed using KAPA HyperExome probes (Roche). Paired-end massive parallel sequencing was performed on a NovaSeq 6000 Instrument (Illumina). Data analysis was performed on an in-house developed platform, in accordance with the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG-AMP) guidelines (Nykamp et al., 2017). Variant confirmation and segregation analysis was performed by Sanger Sequencing on an ABI 3730 platform (Applied Biosystems).
RNA for evaluation of aberrant splicing was obtained using the Maxwell RSC simplyRNA Tissue Kit on a Maxwell RSC Instrument (Promega Corporation) from PHA blasts stimulated with 1 μl/ml PHA-M and 50 μl/ml interleukin-2 (IL-2) (Sigma-Aldrich), LCLs, and patient-derived dermal fibroblasts. cDNA was synthesized using Superscript III Reverse Transcriptase (Invitrogen) according to the manufacturer’s recommendations. Amplicon length analysis was performed using a TapeStation (Agilent Technologies) or an ABI 3730 platform (Applied Biosystems).
Targeted RNA-seq was performed with the SureSelectXT HS2 RNA System kit (Agilent Technologies) using SureSelectXT Human All Exon V7 probes (Agilent Technologies) for target enrichment and sequencing was performed on a NovaSeq 6000 Instrument (Illumina).
All primers were purchased from Integrated DNA Technologies and are listed in Table S5. Nucleotide and protein numbering is in accordance with transcript ENST00000320211.1 and NM_130384.3, respectively.
DNA damage inducers and inhibitors
Cells were irradiated with 254 nm ultraviolet radiation (UV-C) in the absence of culture medium using the UVP Crosslinker (Analytikjena) or with X-rays (220 kV, 13 mA, 0.15 mm copper filter, 3 Gy/min) using the Small Animal Radiation Research Platform (Xstrahl) at indicated doses. The following genotoxic inducers and chemical inhibitors were used: MMC (Sigma-Aldrich), HU (Sigma-Aldrich), ATR kinase inhibitor (ATRi) (20 nM; BAY-1895344, also known as Elimusertib), ATM kinase inhibitor (ATMi) (10 µM; KU-55933), and DNA-PK inhibitor (DNA-PKi) (2 µM; KU-57788, also known as NU7441). Inhibitors were added to the medium 1 h prior to treatment with genotoxic stressors.
Plasmids and cloning
Plasmid DNA constructs containing the wild type (WT) cDNA sequence for ATRIP, ATR, and RPA1 inserted into a modified pcDNA3.1(+) expression vector encoding a N-terminal 3xFlag-tag, a C-terminal V5-tag, or a C-terminal c-Myc-tag, respectively, were purchased from GenScript. Mutant ATRIP plasmid constructs containing the cDNA sequence for WT ATRIP lacking exon 5 inserted into a pcDNA3.1(+) vector encoding an N-terminal 3xFlag-tag were generated by GenScript. All plasmids were transformed in DH10B or DH5α Escherichia coli. DNA was isolated using NucleoBond Xtra Midi kit (Macherey-Nagel). HEK293T cells were transiently co-transfected with ATR, RPA1, and either WT ATRIP or mutant ATRIP plasmid DNA. Transfections were performed with branched 25 kDa polethylenimine.
RT-qPCR
ATRIP transcript levels
Fibroblasts from HCs and patient F1Pt were lysed in RLT buffer (Qiagen) and stored at −80°C until further processing. RNA was purified using the RNeasy Mini kit (Qiagen) according to the manufacturer’s protocol. Total RNA was reverse transcribed into cDNA using the SensiFast cDNA Synthesis Kit (Bioline) and RT-qPCR was performed using the LightCycler 480 (Roche). Gene transcript levels were normalized against an endogenous β-actin control. Samples were run in duplicates in two independent experiments. Results are presented as relative expression compared to the HCs, determined using the 2−ΔΔCt method. To determine mRNA expression levels of a shorter mutant transcript and a transcript containing exon 5 in F1Pt fibroblasts, primers against an amplicon in exon 3–4 and exon 5–6 of ATRIP were used, respectively. Predesigned qPCR primer pairs were ordered from Integrated DNA Technologies (primers are listed in Table S5).
ISG analysis
Whole blood was collected into PAXgene tubes (PreAnalytix) and stored at −80°C until processing. Total RNA was extracted using the PAXgene (PreAnalytix) RNA isolation kit and RNA was reverse transcribed into cDNA using the SensiFast cDNA Synthesis Kit (Bioline). RT-qPCR was performed using the LightCycler 480, analyzing the expression levels of six ISGs and two housekeeping genes using predesigned TaqMan Gene Expression Assays. Data were normalized to the housekeeping genes and results are presented as relative expression compared to the HCs. For each sample, an ISG score was calculated from the median fold induction of the 6 ISGs. A positive ISG score threshold was set based on the mean ISG score of six HCs + 2 SD from the mean (>3.793).
scRNA-, TCR-, BCR-, and CITE-seq
After thawing, up to 2 × 106 PBMCs of each individual were counted, isolated, and spun down. The cell pellet was resuspended and incubated for 30 min on ice with a staining mix in PBS containing 0.04% BSA, CD19-PE-Cy7, eFluor506 Fixable Viability dye, Human TruStain FcX (422302; BioLegend), and TotalSeq-C hashing antibodies (1:500; BioLegend). Two patient samples were multiplexed per lane using TotalSeq-C Cell Hashing Antibodies. Both total live PBMCs and B cells were sorted and loaded on different lanes. Sorted single-cell suspensions were resuspended at an estimated final concentration of 2,000 cells/µl and loaded on a Chromium GemCode Single Cell Instrument (10x Genomics) to generate single-cell gel beads in emulsion. The scRNA/Feature Barcoding/TCR libraries were prepared using the GemCode Single Cell 5′ Gel Bead and Library kit, version 1.1 (1000165; 10x Genomics), according to the manufacturer’s instructions. The cDNA content of prefragmentation and post-sample index PCR samples was analyzed using the 2100 BioAnalyzer (Agilent). Sequencing libraries were loaded on an Illumina NovaSeq flow cell at Vlaams Instituut voor Biotechnologie (VIB) Nucleomics core with sequencing settings according to the recommendations of 10x Genomics, pooled in a 75:20:5 ratio for the gene expression, TCR, and antibody-derived libraries, respectively.
LAM-HTGTS
The LAM-HTGTS method, modified for targeted amplification of CSR junctions, was performed on PBMCs of patient F1P and age-matched HCs (n = 3). The method and subsequent CSR junction analysis were performed as outlined by Du et al. (2024).
Immunoblotting
Cells were lysed in radioimmunoprecipitation assay buffer (150 mM NaCl, 1.0% IGEPAL CA-630, 0.5% sodium deoxycholate, 0.1% SDS, and 50 mM Tris, pH 8.0) (Roche) complemented with protease inhibitors (cOmplete ULTRA; Roche) and phosphatase inhibitors (PhosSTOP; Roche). The soluble fraction was normalized prior to protein separation by SDS-PAGE using a 4–15% Criterion TGX Stain-Free Protein Gel (Bio-Rad) followed by semi-dry transfer to a nitrocellulose membrane (Bio-Rad). Membranes were blocked with 5% nonfat dry milk (Cell Signaling Technology [CST]) in phosphate-buffered saline with 0.1% Tween 20 (PBS-T) or 5% bovine serum albumin (BSA) (Roche) in PBS-T for 1 h at room temperature, incubated with primary antibodies overnight at 4°C or for 2 h at room temperature, followed by incubation with secondary antibodies conjugated to horseradish peroxidase (HRP). Membranes were developed using enhanced chemiluminescence reagents (SuperSignal West Dura or Femto; Thermo Fisher Scientific) on a Chemidoc imaging system (Bio-Rad). All immunoblotting experiments were performed at least twice, and representative data are shown. Protein levels were quantified using the Image Lab software (Bio-Rad). All uncropped images can be retrieved in the Source File. The following primary antibodies were probed: anti-Flag (1:1,000; F3165; Sigma-Aldrich), anti-V5 (1:1,000; R960-25; Invitrogen), anti-c-Myc (1:2,000; ab9106; Abcam), anti-ATRIP (1:1,000; #97687; Cell Signaling Technology [CST]), anti-TOPBP1 (1:2,000; A300-111A; Bethyl Laboratories), anti-RPA70/RPA1 (1:2,000; ab176467; Abcam), anti-ATR (1:1,000; #2790; CST), anti-CHK1 (1:1,000; sc-8408; Santa Cruz), anti-phospho-ATR (T1989) (1:1,000; PA5-77873; Invitrogen), anti-phospho-CHK1 (S317) (1:1,000; #8191; CST), anti-β-tubulin-HRP (1:5,000; ab21058; Abcam), and anti-GAPDH-HRP (1:2,000; #8884; CST). The secondary antibodies goat anti-mouse IgG (H+L)-HRP (1:10,000; G-21040; Invitrogen) and goat anti-rabbit IgG-HRP (1:2,000; #7074; CST) were used. To confirm the specificity of the anti-phospho-ATR antibody, the membrane was dephosphorylated using Lambda Protein Phosphatase (P0753S; New England Biolabs). Briefly, the membrane was incubated with a blocking buffer containing 400 U/ml Lambda Protein Phosphatase and 1 mM MnCl2 overnight at 4°C prior to incubation with the primary antibody.
Co-immunoprecipitation
WT or mutant ATRIP protein was immunoprecipitated by incubating 250 µg of whole cell lysate with 4 µg of anti-FLAG antibody (F3165; Sigma-Aldrich) or 4 µg of isotype-matched IgG antibody (12-371; Sigma-Aldrich) overnight at 4°C, followed by incubation with Dynabeads Protein G (Invitrogen) for 1 h at 4°C. Beads with protein complexes were washed six times with IP High buffer without extra salt and detergents (54001; Nuclear complex Co-IP Kit; Active Motif). Precipitates were eluted by boiling in Laemmli Sample buffer and analyzed by immunoblotting.
Immunofluorescence
Primary fibroblasts were grown on 22-mm glass coverslips (Menzel Gläser) 24 h before starting the experiment. Cells were pulse-labeled with 10 µM EdU (Click-iT Plus EdU Alexa Fluor 647 Imaging Kit; C10640; Invitrogen) for 30 min and concomitantly treated with a DNA damage inducer. For RPA staining, cells were treated with pre-extraction buffer (0.5% Triton X-100, 20 mM HEPES, pH 7.4, 100 mM NaCl, 3 mM MgCl2, and 300 mM sucrose in distilled water) prior to fixation. At the indicated time after treatment, cells were fixed with 3% paraformaldehyde (PFA) in PBS for 20 min, followed by simultaneous permeabilization and blocking in 1% BSA (Sigma-Aldrich) and 0.2% Triton X-100 in PBS for 30 min. Primary antibody incubations were performed for 1 h at room temperature with anti-γH2AX S139 (1:500; 613402; BioLegend) and anti-RPA32/RPA2 (1:500; MABE285; Merck) or overnight at 4°C with anti-ATR (1:250; #13934; CST). After washing, coverslips were incubated with secondary antibodies for 1 h at room temperature (goat anti-rabbit Alexa Fluor 488 [1:1,000; A-11008; Invitrogen] or goat anti-mouse Alexa Fluor 488 [1:1,000; A-11001; Invitrogen]). EdU staining was performed by incubation with Click-iT reaction cocktail according to the manufacturer’s protocol. Coverslips were mounted in Fluoromount (Sigma-Aldrich) containing DAPI nuclear stain. Images were acquired with a Leica DM6 B microscope (Leica Microsystems) using a HCX PL APO 40×/1.3 oil objective, equipped with Las X software (Leica Microsystems). Identical microscope settings and image processing steps were used for all samples of the same experiment. Quantifications of DNA damage foci and analysis of signal intensities were performed using Fiji software (v2.9.0). Pooled data from three independent experiments are shown. A minimum of 150 cells were analyzed per experimental condition.
FCM
Cell cycle analysis
Fibroblasts were seeded at a density of 7,000 cells/cm2 and cultured for 24 h. The cells were exposed to a DNA damage inducer (0.02 µg/ml MMC, 200 J/m2 UV-C, or 1 mM HU) and subsequently cultured for the indicated time. The cells were pulse-labeled with 10 µM EdU for 30 min before harvesting. The cells were fixed and permeabilized using the Foxp3/Transcription Factor Staining Buffer Set (00-5523-00; Thermo Fisher Scientific) following the manufacturer’s protocol. EdU staining was performed by incubating the cells with the Click-iT reaction cocktail, prepared as per the manufacturer’s instructions. The cells were stained with permeabilization buffer containing anti-γH2AX-AF488 (S139) (2F3; 613406; BioLegend) for 30 min on ice when indicated. DNA content was stained using a permeabilization buffer containing DAPI. Acquisition of the stained cells was performed using a BD LSRFortessa or BD FACSymphony A3 (BD Biosciences). Subsequent data analysis was performed with the FlowJo v10.10.0 software (BD Life Sciences).
EdU pulse-chase assay
The EdU pulse-chase assay was performed as previously described by Duthoo et al. (2022) with minor adjustments. Briefly, PHA blasts were cultured for 96 h in a 48-well plate (250,000 cells in 500 µl) in the presence of 2% PHA-M. Next, the cells were pulse-labeled with 10 µM EdU for 30 min and concurrently exposed to a DNA damage inducer (200 J/m2 UV-C or 4 Gy IR). MMC (0.02 µg/ml) was added to the culture 24 h prior to EdU labeling. Subsequent harvesting of the cells was performed at various time points, ranging from 0 to 15 h, with intervals of 3 h. The cells were fixed and permeabilized using the Foxp3/Transcription Factor Staining Buffer Set. Subsequent EdU staining was performed using the Click-iT reaction cocktail and DNA was stained with permeabilization buffer containing DAPI. The acquisition was performed using a BD LSRFortessa (BD Biosciences), and data were analyzed with the FlowJo software.
Proliferation assay
Cryopreserved PBMCs were resuspended in PBS at a density of 1 × 106 cells per ml. Cells were labeled with CTV (1 µM; C34557; Thermo Fisher Scientific) and incubated for 10 min at 37°C. Unbound CTV was quenched by washing the cells with PBS supplemented with 1% FCS. The cells were seeded in a 48-well plate (250,000 cells in 500 µl), treated with 0.02 µg/ml MMC, and subsequently stimulated with 2% PHA-M. After 96 h of culture, PHA blasts were harvested and stained with anti-CD3-AF700 (SK7; 344822; BioLegend), anti-CD4-FITC (RPA-T4; 561005; BD Bioscience), anti-CD8-PE-Cy7 (RPA-T8; 557746; BD Bioscience), and FcR block (422302; BioLegend) in PBS. After 30 min of staining, cells were washed, stained with propidium iodide (PI), and acquired using a BD LSRFortessa (BD Biosciences). Subsequent data analysis was performed with the FlowJo software.
Immunophenotyping
Cryopreserved PBMCs were thawed in 37°C preheated complete medium (RPMI-1640 medium supplemented with GlutaMAX, 10% FCS, 1% penicillin–streptomycin [10,000 U/ml; 15140122; Gibco], 1 mM sodium pyruvate [11360070; Gibco], 1% non-essential amino acids [11140035; Gibco], and 50 μM 2-mercaptoethanol [31350010; Gibco]). Cells were left to recuperate for 30 min at 37°C and 5% CO2 after removal of DMSO. Cells were counted and 2,000,000 cells were plated for each panel. Cells were first stained with FcR block (422302; BioLegend) together with biotin-conjugated antibodies and Zombie UV Fixable Viability dye (423107; BioLegend) in PBS. In the second step, a first set of surface markers was stained with a mixture of antibodies in FACS buffer (DPBS pH7.4, 1% BSA, 0.05% NaN3, and 1 mM EDTA) and Brilliant Stain buffer (BD Biosciences). After 30 min of staining, cells were washed and stained overnight with a second set of antibodies. Cells were fixed, permeabilized, and intracellularly stained with antibodies using the Foxp3/Transcription Factor Staining Buffer Set according to the manufacturer’s protocol. Acquisition and analysis of labeled cell suspensions were performed with a FACSymphony flow cytometer (BD Biosciences) with subsequent analysis of data with the FlowJo software. Antibodies used to define PBMC populations can be found in Table S6.
MN assays
The G0 and MMC MN assay
The G0 and MMC MN assay were performed, as described previously by Beyls et al. (2021) and Francies et al. (2018), respectively. Briefly, fresh whole blood cultures were prepared by adding 0.5 ml blood to 4.5 ml RPMI. The culture medium was supplemented with 10% FBS and 1% penicillin–streptomycin. The cultures were irradiated with 0.5 or 1 Gy (G0 MN assay) or treated with 0.02 µg/ml MMC (MMC MN assay) and subsequently stimulated with 2% PHA-M. Cytochalasin B (6 µg/ml; Sigma-Aldrich) was added to block cytokinesis 23 h after stimulation. After 70 h culture time, cells were harvested and exposed to a cold hypotonic shock with 0.075 M KCl and fixed in an ice-cold methanol/acetic acid solution. Acridine orange (10 µg/ml; Sigma-Aldrich) stained slides were scored according to the criteria described by Fenech (2007). MN was scored manually on a fluorescence microscope in 1,000 binucleate (BN) cells per condition.
The S MN assay
Fresh whole blood cultures were set up as described above and cell proliferation was immediately stimulated with 2% PHA-M. Following 96 h of culture, the cells were pulse-labeled with 10 µM EdU for 30 min, irradiated with 0.5 or 1 Gy, and cytochalasin B (6 µg/ml; Sigma-Aldrich) was added. After an additional 24 h of culture, the cells were harvested and fixed according to the G0 MN protocol described previously. Slides were prepared and the cells were additionally fixed in 3% PFA for 20 min and permeabilized with ice-cold 0.2% Triton X-100 for 10 min. Subsequently, EdU staining was performed according to the manufacturer’s protocol and the slides were mounted in Fluoromount containing DAPI nuclear stain. MN was scored in 1,000 EdU-positive BN cells per condition.
Cytotoxicity assay
Fibroblasts were seeded into 24-well plates at a density of 6,000 cells/cm2 24 h before starting the experiment. Cells were exposed to a DNA damage inducer (0.2 µg/ml MMC, 20 J/m2 UV-C, or 10 Gy IR) and incubated with SYTOX Green nucleic acid stain (1:100,000; Invitrogen) to indicate dead cells during cell culture. Brightfield and fluorescence images were captured for 5 days with an 8-h interval with the CELLCYTE X (Cytena). Cell confluency and the number of dead cells were quantified using CELLCYTE X Analysis software.
CRISPR-SelectTIME in monoallelic cell system
Generation of monoallelic ATR+/− and ATRIP+/− cell lines
An iCas9-MCF10A-WT clonal cell line with stably integrated TRE3G Edit-R Inducible Lentiviral Cas9 construct (CAS11229; Horizon) was gifted by Roderick L. Beijersbergen (The Netherlands Cancer Institute, Amsterdam, Netherlands). To generate ATR+/− cells, iCas9-MCF10A-WT cells were transfected with dual gRNAs targeting intron 1 and intron 46 of the ATR gene. ATRIP+/− cells were generated by transfecting the iCas9-MCF10A-WT cells with dual gRNAs targeting intron 1 and intron 8 of the ATRIP gene. Two CRISPR RNAs (crRNAs) for Streptococcus pyogenes Cas9 were designed using the online software Crispor (http://crispor.tefor.net/crispor.py). Note, the crRNAs should be located in the non-coding region to avoid disturbing adjacent genes. The knock-out efficiency of the designed crRNAs was analyzed by the online tool ICE (https://ice.synthego.com/#/), and subsequently, the crRNAs with a high knock-out efficiency were selected to establish the mono-allelic cell line. The iCas9-MCF10A-WT cells were seeded with 60% confluency in a 6-cm culture dish. To induce the expression of the cas9 protein, 1 μg/ml doxycycline was added to the medium. After 24 h, 7.5 µl of each crRNA (2 nmol) was mixed with 15 µl of trans-activating crRNA (tracrRNA, 5 nmol) and was incubated at room temperature for 10 min to form crRNA:tracrRNA duplexes. Cells were subsequently transfected by 4 ml of culture medium containing 30 µl crRNA:tracrRNA duplexes, 15 µl lipofectamine RNAiMAX (13778; Thermo Fisher Scientific), and 500 µl Opti-MEM (31985062; Thermo Fisher Scientific). After 2 days of culture, single cells were sorted into 96-well plates with 100 µl fresh medium per well using a FACS Aria III instrument (BD Biosciences) and subsequently kept in culture for 2–3 wk until clonal cell lines were formed. Genomic PCR was performed to evaluate the monoallelic cell line. Primers targeting WT, ATR, or ATRIP deletion alleles were designed through Primer3 (https://primer3.ut.ee/). Primers annealing to sequences outside the depletion region were used to identify the knock-out alleles. Primers annealing to sequences inside the deleted regions were used to detect the WT allele. The cell colonies containing DNA that were successfully amplified by both two types of primers were monoallelic cell lines. DNA extraction was performed using the Quick-DNA Miniprep Plus Kit (D4069; Zymo Research). PCR was performed using 100 ng genomic DNA and 1% agarose gel electrophoreses. For further verification, the PCR products of target sites were purified using the QIAquick Gel Extraction Kit (28706; Qiagen) and sent to Eurofins for Sanger sequencing. The designed crRNAs and PCR primers are listed in Table S7. Additionally, the expression levels of ATRIP, ATR, TOPBP1, RPA1, and RPA2 in the generated monoallelic cell lines was confirmed by immunoblotting. The following primary antibodies were probed: anti-ATR (1:500; sc-1887; Santa Cruz), anti-ATRIP (1:500; #2737S; CST), anti-TOPBP1 (1:1,000; A300-111A; Bethyl Laboratories), anti-RPA70/RPA1 (1:500; NA13; Millipore), anti-RPA32/RPA2 (1:2,000; MABE285; Millipore), and anti-actin (1:50,000; MAB1501; Millipore).
In vitro CRISPR-SelectTIME assay
In a 6-well plate, 17,500 iCas9-MCF10A-ATR+/− or ATRIP+/− cells were seeded in 2 ml of culture medium per well. To induce the expression of Cas9 protein, 1 μg/ml doxycycline was added in each well 24 h before cell transfection. After 24 h, 7.5 µl of the designed crRNA (2 nmol) was incubated for 10 min at room temperature with 7.5 µl of tracrRNA (5 nmol). Next, a mixture of 15 µl crRNA:tracrRNA duplexes, 7.5 µl lipofectamine RNAiMAX, 250 µl Opti-MEM, and 2 µl mixture of designed variant (10 µM) and WT* single-stranded oligodeoxynucleotide repair templates was dripped onto the iCas9-MCF10A cells in fresh medium. The DNA repair template for each variant was designed in the online software Benchling (https://benchling.com). The design of the synonymous mutations referring to the codon usage information is available at https://www.kazusa.or.jp. After 2 days, 50% of the edited cells were collected and DNA extraction was performed using the Quick-DNA Miniprep Plus Kit. The remaining edited cells were kept in culture and passaged every 3 days. After an additional 10 days of culture, all cells were collected and lysed for DNA extraction using the Quick-DNA Miniprep Plus Kit. Using Benchling and Primer3, PCR primers for the amplification of the edited target sites in each variant were designed. Two rounds of PCR were performed to prepare the PCR products for subsequent amplicon NGS. First, primer pairs were used to amplify the specific edited target sites, containing overhangs with binding sites for the primers of the second round. Second, specific barcodes were added to each sample, including overhangs with barcodes or adaptors for NGS. After purifying the PCR products using the QIAquick Gel Extraction Kit, NGS was performed to sequence the PCR products of the DNA samples collected on day 2 and day 12. The ratio of read numbers of the knock-in of a variant relative to the WT* on day 2 and day 12 were compared to determine the functional characteristics of the different mutations. The designed crRNAs, PCR primers, and DNA repair templates are listed in Tables S8 and S9.
Statistics
GraphPad Prism software (v10) was used for all statistical analyses. As indicated in the figure legends, results were analyzed with a two-tailed Mann–Whitney test, multiple paired t tests, or Kruskal–Wallis test combined with Dunn’s multiple comparisons test. The threshold for statistical significance was set to P < 0.05. Representative immunofluorescence images and flow cytometric plots are shown. R (v4.3.2) and RStudio (v2023.09.1 + 494) were used for the analysis of scRNA-, TCR-, BCR-, and CITE-seq data. The analytical pipeline consisted of Cellranger (v6.1.1) for quality control. CITE-seq and scRNA-seq data were analyzed using the Seurat package (v5), with an embedded Presto package (v1.0.0) for differential gene expression analysis. Analysis of TCR-seq and BCR-seq data was conducted using the Diversity AnaLysis Interface (DALI, v2.0.0).
Online supplemental material
Fig. S1 provides additional information related to Fig. 1, including data on ATRIP deficient patients and population genetics, validation of exon 5 skipping caused by the variant, and confirmation of the absence of full-length ATRIP protein in patient-derived cells. Fig. S2 complements both Figs. 2 and 3, presenting additional data on the immunophenotype of patient F1Pt. Fig. S3 includes additional data relevant to Fig. 4, including additional scRNA-seq data demonstrating a DNA repair and IFN signature in ATRIP-deficient T effector and NK subsets, and FCM data detailing the effect of anti-CD20 mAb treatment in patient F1Pt. Fig. S4 provides supporting data for Figs. 5 and 6, including immunofluorescence images, western blot data, and FCM results that show the impact of ATRIP deficiency on ATR signaling, DNA replication, cell cycle progression, and proliferation. Fig. S5 provides supporting data for Figs. 7 and 8, detailing the cellular phenotype of ATRIP deficiency, including increased DNA damage, chromosomal sensitivity, and reduced cellular fitness. Table S1 provides information on the clinical features of patient F1Pt. Table S2 details the scores obtained from various in silico prediction tools and Table S3 lists information on the homozygous ATRIP splice variants found in gnomAD v4.0.0. Table S4 details the candidate genes that were screened to exclude differential diagnosis in patient F1Pt. Table S5 lists primers used for variant analysis, splice analysis, and RT-qPCR. Table S6 details all antibodies used for FCM. Table S7 lists crRNAs and PCR primers designed for generating the monoallelic cell lines. Table S8 includes crRNAs and PCR primers designed for variant analysis, and Table S9 lists all designed mutation and WT DNA repair templates for CRISPR-SelectTIME.
Data availability
The scRNA-seq dataset is publicly accessible on GEO (accession no: GSE284526) and the code used for scRNA-seq data analysis is available on GitHub at https://github.com/LeanderJonckheere/ATRIP_deficiency_JEM. The ATRIP variant has been submitted to ClinVar (VCV000982310.2, VCV003393375.1) and the Leiden Open Variation Database (https://databases.lovd.nl/shared/variants/0000959751#00003155). All other data are provided within the article and its supplementary materials or can be obtained from the corresponding authors upon reasonable request.
Materials and correspondence: Correspondence and requests for materials should be addressed to F. Haerynck ([email protected]) and K.B.M. Claes ([email protected]).
Acknowledgments
We gratefully thank the patients and family for consenting to this research. The valuable input from Prof. Penny Jeggo, Prof. Grant Stewart, Prof. Mark O’Driscoll, Dr. Angela Brady, Dr. Mohnish Suri, Dr. Pradeep Vasudevan, Dr. Emma Hobson, Dr. Kaljit Bhuller, and Dr. Mandal Kausik on the clinical phenotype of the ATRIP and ATR patients is warmly appreciated. We thank Karlien Claes for patient-centered support, Sylvie De Buck for administrative assistance, Lieselotte Vande Walle and Sarah Ghistelinck for experimental expertise, and Julie Smet for providing NK cytotoxicity data. We acknowledge the VIB Flow Core and VIB Single Cell Core of the VIB Center for inflammation research for their assistance and expertise in experimental design.
This work is supported (not financially) by the European Reference Network for Rare Immunodeficiency, Autoinflammatory, and Autoimmune Diseases Network (ERN-RITA). This work was funded by Research Foundation Flanders (FWO) (FWOTBM2018000102) and VIB Grand Challenge. S.J. Tavernier is a beneficiary of a senior postdoctoral FWO grant (1236923N). S. Rieman is funded by Ghent University (BOF23/DOC/013). Centre for Primary Immune Deficiency Ghent (CPIG) is recognized as a Jeffrey Modell Diagnostic and Research Center and funded by the Jeffrey Modell Foundation.
Author contributions: E. Duthoo: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Resources, Validation, Visualization, Writing - original draft, Writing - review & editing, E. Beyls: Conceptualization, Formal analysis, Investigation, Methodology, Project administration, Validation, Visualization, Writing - original draft, Writing - review & editing, L. Backers: Conceptualization, Data curation, Formal analysis, Investigation, Software, Validation, Visualization, Writing - original draft, T. Gudjónsson: Investigation, Methodology, Supervision, P. Huang: Data curation, Formal analysis, Investigation, Methodology, Writing - review & editing, L. Jonckheere: Data curation, Formal analysis, Resources, Software, Visualization, S. Riemann: Data curation, Formal analysis, B. Parton: Data curation, Investigation, Methodology, Resources, L. Du: Investigation, V. Debacker: Investigation, Resources, Writing - review & editing, M. De Bruyne: Formal analysis, Investigation, L. Hoste: Data curation, Investigation, Writing - review & editing, A. Baeyens: Methodology, Resources, Supervision, Writing - review & editing, A. Vral: Conceptualization, Funding acquisition, Methodology, Writing - review & editing, E. Van Braeckel: Funding acquisition, Supervision, Writing - review & editing, J. Staal: Methodology, Resources, G. Mortier: Resources, Validation, Writing - review & editing, T. Kerre: Data curation, Formal analysis, Writing - review & editing, Q. Pan-Hammarström: Supervision, C.S. Sørensen: Conceptualization, Funding acquisition, Methodology, Project administration, Supervision, Validation, Visualization, Writing - original draft, Writing - review & editing, F. Haerynck: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Writing - review & editing, K.B.M. Claes: Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing - original draft, Writing - review & editing, S.J. Tavernier: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Supervision, Validation, Writing - original draft, Writing - review & editing.
References
Author notes
E. Duthoo, E. Beyls, and L. Backers contributed equally to this paper.
F. Haerynck, K.B.M. Claes, and S.J. Tavernier share last authorship.
Disclosures: S. Riemann reported grants from Ghent University (predoctoral mandate BOF23/DOC/013), non-financial support from AstraZeneca, and non-financial support from GSK outside the submitted work. L. Jonckheere reported a grant from the Research Foundation—Flanders (FWO) (predoctoral mandate 11J2525N). C.S. Sørensen reported a patent to the European Patent Office, application no. 21816113.1 on CRISPR-Select pending. No other disclosures were reported.