NF-κB is an integral component of the immune response to Toxoplasma gondii. Although evidence exists that T. gondii can directly modulate the NF-κB pathway, the parasite-derived effectors involved are unknown. We determined that type II strains of T. gondii activate more NF-κB than type I or type III strains, and using forward genetics we found that this difference is a result of the polymorphic protein GRA15, a novel dense granule protein which T. gondii secretes into the host cell upon invasion. A GRA15-deficient type II strain has a severe defect in both NF-κB nuclear translocation and NF-κB–mediated transcription. Furthermore, human cells expressing type II GRA15 also activate NF-κB, demonstrating that GRA15 alone is sufficient for NF-κB activation. Along with the rhoptry protein ROP16, GRA15 is responsible for a large part of the strain differences in the induction of IL-12 secretion by infected mouse macrophages. In vivo bioluminescent imaging showed that a GRA15-deficient type II strain grows faster compared with wild-type, most likely through its reduced induction of IFN-γ. These results show for the first time that a dense granule protein can modulate host signaling pathways, and dense granule proteins can therefore join rhoptry proteins in T. gondii’s host cell–modifying arsenal.
Toxoplasma gondii is an obligate intracellular parasite capable of infecting a wide range of warm-blooded hosts, including humans. T. gondii establishes a lifelong chronic infection in the host by evading and subverting the immune system. T. gondii infection is usually asymptomatic in healthy humans but can lead to flu-like and neurological symptoms in immunosuppressed patients and the fetuses of pregnant women infected for the first time. The vast majority of T. gondii strains isolated from Europe and North America belong to three clonal lineages, types I, II, and III, which differ in many phenotypes, including virulence (Saeij et al., 2005). In mice, type I strains are categorically lethal, with an LD100 = 1, whereas type II or type III infections are not (LD50 ≈ 102 and ≈ 105, respectively; Sibley and Boothroyd, 1992; Saeij et al., 2006). Strain differences in the modulation of host immune signaling pathways are one way by which this diversity arises. For example, the strain-specific modulation of the STAT3/6 signaling pathway by the secreted kinase ROP16 accounts for some of the strain differences in virulence (Saeij et al., 2006). Evidence also exists for the strain-specific modulation of NF-κB (Robben et al., 2004), an important host signaling pathway in the regulation of inflammatory, immune, and antiapoptotic responses.
The NF-κB family of transcription factors is composed of five members: p50 (NF-κB1), p52 (NF-κB2), p65 (RelA), RelB, and c-Rel (Vallabhapurapu and Karin, 2009). In unstimulated cells, homo- or heterodimers of NF-κB are sequestered in the cytoplasm by members of the IκB (inhibitor of κB) family. Activation of NF-κB is initiated by the degradation of IκB proteins. This occurs via the activation of kinases called IκB kinases (IKKs), which phosphorylate two serine residues located in IκB regulatory domains, leading to their ubiquitination and subsequent proteasomal degradation. The NF-κB complex is then free to enter the nucleus where it can induce expression of specific genes that have NF-κB-binding sites in their promoters.
Many pathogens have developed strategies to modulate the host NF-κB pathway (Tato and Hunter, 2002). Several bacteria and viruses inhibit NF-κB activation and its resultant recruitment and activation of immune cells, resulting in enhanced survival of the pathogen. Other pathogens induce NF-κB activation, which inhibits apoptosis, an important defense against intracellular pathogens, and increases cell migration, thereby recruiting new cells to infect. Furthermore, NF-κB–mediated inflammation leads to tissue damage, allowing pathogens to cross tissue barriers. Thus, depending on the host, pathogen, and site of infection, an active NF-κB pathway can benefit either the host or the pathogen.
Mice deficient in some NF-κB family members have increased susceptibility to T. gondii, indicating the importance of this pathway in pathogen resistance (Mason et al., 2004). C-Rel−/− mice are highly susceptible to the acute stage of T. gondii i.p. infection, which can be rescued solely by treatment with IL-12, indicating that a major role of NF-κB in resistance to T. gondii is the induction of IL-12 secretion (Mason et al., 2004). IL-12 is a major mediator of the proinflammatory Th1 response development, and the major cause of chronic phase death in mice lacking RelB, p52, or the IκB protein Bcl-3 is also a deficient T cell response (Mason et al., 2004). Although it is clear that the NF-κB pathway is important for an adequate response to T. gondii infection, the mice used in these studies all lack a particular NF-κB subunit in every cell of their bodies, and it is currently unknown what the role of NF-κB is in specific cell types, such as those directly infected with T. gondii.
Evidence currently exists for both inhibition and activation of the NF-κB pathway in host cells by T. gondii. Less than 6 h after infection, a type I strain was shown to block the nuclear translocation of p65 and the in vitro binding of NF-κB subunits to DNA (Butcher et al., 2001; Shapira et al., 2002; Kim et al., 2004). Induction of IL-12 in response to TNF or LPS stimulation was also reduced (Butcher et al., 2001; Kim et al., 2004). This inhibition of the NF-κB pathway was dependent on active invasion by live parasites (Butcher and Denkers, 2002). After >6 h of infection, inhibition of p65 nuclear translocation and in vitro DNA binding was no longer observed (Kim et al., 2004; Leng et al., 2009). However, chromatin immunoprecipitation experiments showed that the in vivo binding of p65 to the TNF promoter was blocked even at late time points (Leng et al., 2009). Other groups, however, have shown that NF-κB is activated by a type I strain of T. gondii, and that this activation is necessary for the inhibition of apoptosis (Molestina et al., 2003; Payne et al., 2003; Molestina and Sinai, 2005b). A consistent observation has been the phosphorylation and ubiquitination of IκBα upon T. gondii infection, although it is unclear what effect this has on the nuclear translocation of NF-κB and transcription of downstream genes (Butcher et al., 2001; Molestina et al., 2003; Shapira et al., 2005). All of these studies used a type I strain of T. gondii, suggesting that different observations might be the result of different cell types and/or host species. Strain differences have been observed in the manipulation of the host NF-κB pathway by T. gondii. Type II strains were shown to cause the translocation of NF-κB to the nucleus of mouse splenocytes and mouse BM-derived macrophages (BMMs), whereas type I strains did not (Dobbin et al., 2002; Robben et al., 2004). This strain difference was also shown to have downstream effects, as infection of BMM with type II parasites resulted in high levels of IL-12 secretion compared with infection with type I parasites (Robben et al., 2004). At present, the T. gondii factors involved in the modulation of the NF-κB pathway are not known.
In our experiments, infection with type II strains induces a high level of NF-κB activation, whereas infection with type I or III strains does not. Using F1 progeny from a type II × type III cross, we identify a type II gene responsible for NF-κB activation, GRA15. The protein product of GRA15 is a novel dense granule protein that is necessary and sufficient for p65 nuclear translocation and NF-κB–mediated host cell transcription. We show that GRA15 activates the NF-κB pathway independent of MyD88 and TRIF but dependent on TRAF6 and the IKK complex. Although GRA15 does not affect overall virulence of parasites, it does have more subtle phenotypes in vivo, affecting both parasite growth and cytokine levels.
RESULTS
Host gene expression analysis shows strain-specific activation of the NF-κB pathway
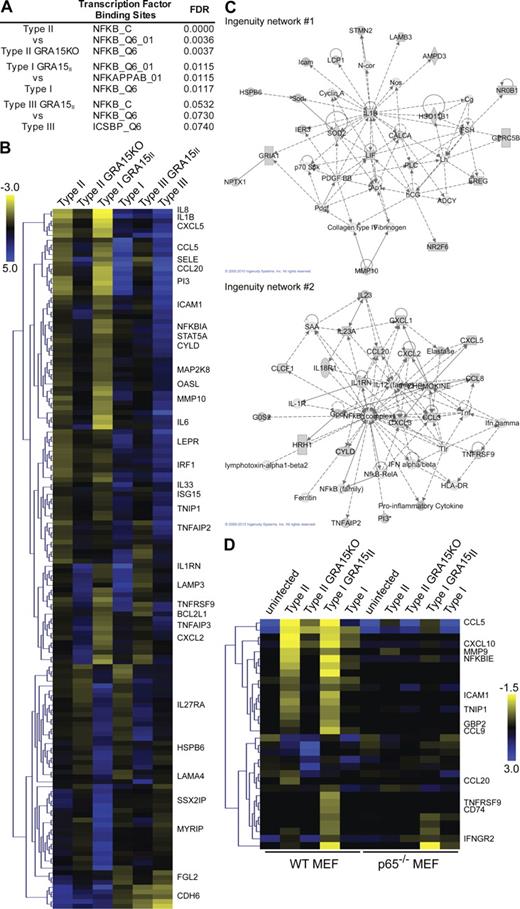
We previously generated a large gene expression dataset from human foreskin fibroblasts (HFFs) infected with type I, II, or III T. gondii strains (Saeij et al., 2007). In our dataset analysis, we focused on the human genes that were differentially regulated by type II strain infection because published data indicated that type II strains might induce more NF-κB nuclear translocation than type I strains (Dobbin et al., 2002; Robben et al., 2004). If T. gondii strains differ in the activation of the NF-κB pathway, this should lead to differences in expression of genes with NF-κB transcription factor binding sites (TFBSs) in their promoters. 105 genes were found to be more than twofold up-regulated in type II infections compared with type I and type III infections (Fig. S1 A). Analysis of TFBS in the regulatory elements of these genes revealed enrichment of NF-κB TFBS in their promoters (Fig. S1 B), and a network analysis of molecular relationships between the products of these 105 genes resulted in high scores for two networks whose central factors were the transcription factor NF-κB (network 1) and IL1β/PTGS2(COX-2) (network 2). IL1β and COX-2 are also regulated by NF-κB (Newton et al., 1997; Vallabhapurapu and Karin, 2009). These data suggest that there is at least one polymorphic locus between type II and type I/III strains that modulates the NF-κB pathway.
We also looked for polymorphic loci between type II and type III strains that modulate host gene expression using quantitative trait locus analysis of human gene expression levels of cells infected with 19 different F1 progeny from II × III crosses. We identified 3,188 human cDNAs that were regulated by a specific T. gondii genomic locus (Saeij et al., 2007). 1,176 of these human cDNAs were regulated by a locus on chromosome VIIb. The T. gondii polymorphic ROP16 kinase resides on this chromosome and, via its strain-specific activation of STAT3/6, is responsible for the differential expression of many of the genes that are regulated by a chromosome VIIb locus (Saeij et al., 2007). Loci on chromosome X also influenced the expression of 563 human cDNAs. To discover if these cDNAs are regulated by a common transcription factor, we determined if any TFBSs were enriched in the promoters of genes that are differentially modulated by F1 progeny with a type II allele versus a type III allele at each chromosome X marker. At many markers, the expression of genes with NF-κB TFBS in their promoters was enriched in F1 progeny with a type II genotype, suggesting that a T. gondii factor responsible for strain differences in NF-κB activation resides on chromosome X (Fig. S1 C). Additionally, network analysis of molecular relationships between the 563 genes that were significantly influenced by a chromosome X locus and their gene products resulted in high scores for two networks, one of which had the transcription factor NF-κB as its central factor (Fig. S1 C). We therefore hypothesized that a polymorphic T. gondii locus on chromosome X contributes to differential regulation of the host NF-κB pathway by type II and type I/III strains.
Toxoplasma type II parasites activate NF-κB
To investigate modulation of the NF-κB pathway by T. gondii, we infected HFFs with type I, II, or III T. gondii strains for 1–24 h and measured nuclear translocation of the NF-κB p65 subunit by immunofluorescence (IF). Starting after 4 h of infection and continuing until at least 24 h of infection, many cells infected with a type II strain contained high levels of p65 in their nucleus, whereas a type I or a type III strain did not induce translocation of high levels of p65 to the nucleus (Fig. 1 A and B; and not depicted). We have observed the translocation of p65 by infection with various type II strains, including ME49, Pru, DAG, or Beverley, and the absence of high levels of p65 translocation after infection with both RH or GT1 type I strains and CEP or VEG type III strains. Uninfected HFFs surrounding infected HFFs did not contain increased p65 in the nucleus, indicating that activation is not caused by a secreted host factor or a contaminant in the medium. Activation of p65 translocation after infection with type II strains was not inhibited by previous infection with type I or type III strains, demonstrating that this translocation is a result of specific activation by type II parasites rather than inhibition by type I/III parasites (Fig. 1, C and D; and not depicted). We have observed the activation of NF-κB p65 nuclear translocation by type II parasites in 293T cells, HeLa cells, mouse BMM, RAW264.7 murine macrophages, mouse embryonic fibroblasts (MEFs), and rat embryonic fibroblasts (see Fig. 3; and not depicted).
T. gondii strains differ in the activation of NF-κB. HFFs were infected with T. gondii strains for 18–24 h, fixed, and stained with α–NF-κB p65 (red) and Hoechst dye (blue). Shown are infection with a type II (GFP) strain (A) or a type III (GFP) strain (B), and HFFs coinfected with a type I (non-GFP, arrow) and a type II (GFP) strain of T. gondii (C, brightfield; D, IF). Cells infected with only type I parasites (arrowhead) do not contain nuclear NF-κB. This experiment has been repeated >10× with similar results. Bars, 10 µm.
T. gondii strains differ in the activation of NF-κB. HFFs were infected with T. gondii strains for 18–24 h, fixed, and stained with α–NF-κB p65 (red) and Hoechst dye (blue). Shown are infection with a type II (GFP) strain (A) or a type III (GFP) strain (B), and HFFs coinfected with a type I (non-GFP, arrow) and a type II (GFP) strain of T. gondii (C, brightfield; D, IF). Cells infected with only type I parasites (arrowhead) do not contain nuclear NF-κB. This experiment has been repeated >10× with similar results. Bars, 10 µm.
Type I parasites do not inhibit NF-κB activation
Although we observed that a type I strain does not inhibit p65 nuclear translocation in a co-infection with a type II strain, previous studies have shown that in mouse macrophages, infection with type I parasites can inhibit the activation of NF-κB in response to LPS or TNF (Butcher et al., 2001; Shapira et al., 2002; Kim et al., 2004). To further investigate if type I parasites can inhibit NF-κB translocation, we infected BMM with type I parasites for 1 or 18 h, stimulated the cells with LPS or TNF, and measured the translocation of p65 to the nucleus by IF (Fig. 2 A). In uninfected cells, both LPS and TNF stimulation induced the translocation of p65 subunits to the nucleus. Prestimulation with LPS inhibited later LPS-induced translocation, as the activation of Toll-like receptor pathways induces negative-feedback mechanisms to inhibit further signaling (Lang and Mansell, 2007; Wang et al., 2009). However, preinfection with type I parasites did not inhibit LPS- or TNF-stimulated translocation at early or late time points after infection. In fact, preinfection with T. gondii led to higher levels of p65 translocation after LPS stimulation, perhaps as a result of increased TLR4 expression (Kim et al., 2004).
Type I parasites do not inhibit NF-κB activation. (A) C57BL/6 BMMs were untreated, stimulated with 100 ng/ml LPS or 20 ng/ml mouse TNF, or infected with type I parasites (MOI = 2) for 1 h (top) or 18 h (bottom). Cells were then restimulated for 30 min with LPS or 45 min with TNF, fixed, and stained with α–NF-κB p65 and Hoechst dye. The intensity of nuclear NF-κB p65 was quantitated in at least 10 cells per treatment. Values represent the fold induction of nuclear p65 levels over uninfected unstimulated cells. This experiment was done once in BMM. A second experiment of HFFs infected with type I parasites and subsequently stimulated with TNF yielded similar results. Error bars represent standard deviation. (B) A 293 NF-κB GFP reporter cell line was plated on coverslips and infected with type I parasites (MOI = 1–2) for 45 min and then stimulated with 100 ng/ml human TNF for 4 h, fixed, and stained with α-SAG1 (red). Bars, 10 µm. The GFP intensity was quantitated for at least 30 infected cells and 30 uninfected cells (bottom; n.s. = not significant, two sample Student’s t tests). Horizontal bars represent the mean GFP intensity of cells. This experiment was performed twice, with the same qualitative results. Similar results were also obtained with a 293T NF-κB luciferase reporter cell line. (C) Microarray analysis was done on HFFs preinfected with a type I strain (MOI = 7.5) for 18 h, or left uninfected, and subsequently stimulated with 20 ng/ml human TNF for 6 h. Genes were preranked for both samples by the difference in expression, as compared with uninfected untreated HFFs, and GSEA was used to determine whether genes with NF-κB TFBS in their promoters or belonging to NF-κB–related canonical pathways were enriched in either or both samples. False discovery rate q-values <0.25 were considered significant. One array per strain and treatment was done.
Type I parasites do not inhibit NF-κB activation. (A) C57BL/6 BMMs were untreated, stimulated with 100 ng/ml LPS or 20 ng/ml mouse TNF, or infected with type I parasites (MOI = 2) for 1 h (top) or 18 h (bottom). Cells were then restimulated for 30 min with LPS or 45 min with TNF, fixed, and stained with α–NF-κB p65 and Hoechst dye. The intensity of nuclear NF-κB p65 was quantitated in at least 10 cells per treatment. Values represent the fold induction of nuclear p65 levels over uninfected unstimulated cells. This experiment was done once in BMM. A second experiment of HFFs infected with type I parasites and subsequently stimulated with TNF yielded similar results. Error bars represent standard deviation. (B) A 293 NF-κB GFP reporter cell line was plated on coverslips and infected with type I parasites (MOI = 1–2) for 45 min and then stimulated with 100 ng/ml human TNF for 4 h, fixed, and stained with α-SAG1 (red). Bars, 10 µm. The GFP intensity was quantitated for at least 30 infected cells and 30 uninfected cells (bottom; n.s. = not significant, two sample Student’s t tests). Horizontal bars represent the mean GFP intensity of cells. This experiment was performed twice, with the same qualitative results. Similar results were also obtained with a 293T NF-κB luciferase reporter cell line. (C) Microarray analysis was done on HFFs preinfected with a type I strain (MOI = 7.5) for 18 h, or left uninfected, and subsequently stimulated with 20 ng/ml human TNF for 6 h. Genes were preranked for both samples by the difference in expression, as compared with uninfected untreated HFFs, and GSEA was used to determine whether genes with NF-κB TFBS in their promoters or belonging to NF-κB–related canonical pathways were enriched in either or both samples. False discovery rate q-values <0.25 were considered significant. One array per strain and treatment was done.
To test whether TNF-stimulated NF-κB–mediated transcription can be inhibited by type I parasites, we used a 293 NF-κB reporter cell line that expresses GFP upon NF-κB activation. Infection of this reporter cell line with type II parasites results in high levels of GFP in infected cells (Fig. S9 A). We added type I parasites to these cells for 45 min, stimulated the cells with TNF, and measured GFP levels of infected and uninfected cells by microscopy (Fig. 2 B). After 4 h of stimulation, both infected and uninfected cells had varying levels of GFP, with some cells containing almost no GFP and <50% of cells having a high level of GFP. However, the distribution of GFP intensity in the populations of infected and uninfected cells was not significantly different. Unstimulated cells, either infected or uninfected, had negligible levels of GFP (unpublished data). Type I parasites also did not inhibit NF-κB–mediated transcription of luciferase in a 293 NF-κB luciferase reporter cell line (unpublished data).
To investigate if type I preinfection might inhibit transcription of specific subsets of NF-κB–regulated genes, we infected HFFs with a type I strain for 18 h or left cells uninfected and subsequently stimulated the cells with TNF for 6 h. We then performed gene expression analysis using Affymetrix microarrays. Comparing the expression data to uninfected HFF expression data, the genes regulated by TNF stimulation alone and type I preinfection followed by TNF stimulation were not identical; however, genes with NF-κB TFBS in their promoters or belonging to an NF-κB–related pathway were equally enriched in those two samples (Fig. 2 C). We therefore conclude that type I parasites do not inhibit TNF- or LPS-stimulated NF-κB p65 translocation or TNF-stimulated NF-κB–mediated transcription, but they may be able to modulate other pathways or host cell transcription factors which are important for expression of a small subset of TNF stimulated genes.
A T. gondii genomic locus on chromosome X mediates strain-specific activation of NF-κB
To find the T. gondii genomic regions mediating the type II versus type I/III strain-specific difference in activation of NF-κB, we infected HFFs with 27 F1 progeny derived from crosses between type II and type III strains and measured NF-κB activation by IF. Only HFFs infected with F1 progeny having type II alleles for the genetic markers ROP2 and GRA6 at the right end of T. gondii chromosome X contained nuclear NF-κB p65 (Fig. S2). Thus, the genomic region in the vicinity of genetic markers ROP2-GRA6 harbors one or more genes involved in the activation of NF-κB. We developed new RFLP markers to more accurately define the place of recombination in progeny that are recombinant for chromosome X around ROP2 and GRA6 and therefore limit the genomic region involved in the activation of NF-κB and the number of possible candidate genes. With these new markers, genotyping of STE7 refined the 3′ boundary of the region and genotyping of S26 refined the 5′ boundary of the region (Fig. S2). The refined region between the markers SAG2E and RC4 contains 45 predicted type II genes (ToxoDB.org, v6.0).
To identify the T. gondii gene responsible for NF-κB activation, we used a candidate gene approach. From the 45 predicted type II genes, the SAG2CDXY (SRS49a/b/c/d) locus was excluded, as a type II strain with this locus deleted still activates NF-κB (Saeij et al., 2008; unpublished data). Our first criterion for a protein able to interface with the host cell and modulate host cell signaling was the presence of a signal sequence. Of the 41 remaining genes, 17 are predicted to have a signal sequence. Of these 17 genes, four (ROP8, ROP2A, GRA6, and 63.m00001) were consistently expressed in tachyzoites in infected macrophage cells, as determined by T. gondii microarrays (unpublished data). Our top candidate genes (ROP2A, GRA6, and 63.m00001) were then tested by adding a type II C-terminal HA-tagged copy of the candidate gene, including at least 1,500 bp of the putative endogenous promoter, into a type I and/or III strain and assaying whether these transgenic type I/III strains activate NF-κB. Type I and III strains stably expressing a copy of 63.m00001 activated NF-κB in HFFs, whereas strains expressing a copy of GRA6 or ROP2 did not, indicating that 63.m00001, hereafter referred to as GRA15, is the locus on chromosome X which mediates the strain-specific activation of NF-κB (Fig. 3 and Fig. S3).
GRA15 mediates NF-κB p65 translocation. (A) HFFs were infected with T. gondii strains for 18 h, fixed, and stained with α–NF-κB p65 (red), α-SAG1 (green), and Hoechst dye (blue). Bars, 10 µm. (B) The amount of p65 in the nucleus was quantitated in at least 15 HFF cells for each strain. Asterisks indicate significantly higher levels of nuclear p65 compared with uninfected cells (*, P < 0.001, two sample Student’s t tests). (C) Mouse BMDMs (BALB/c) were infected with T. gondii strains for 24 h, fixed, and stained with α–NF-κB p65 (red), α-SAG1 (green), and Hoechst dye (blue). Bars, 10 µm. (D) The level of nuclear p65 in infected cells was quantitated in at least 12 infected cells per strain. Asterisks indicate significantly higher levels of nuclear p65 compared with uninfected cells (*, P < 0.005, two sample Student’s t tests). One replicate experiment was done in both HFFs and mouse BMDM with similar qualitative results. Horizontal bars represent the mean nuclear p65 intensity over all cells.
GRA15 mediates NF-κB p65 translocation. (A) HFFs were infected with T. gondii strains for 18 h, fixed, and stained with α–NF-κB p65 (red), α-SAG1 (green), and Hoechst dye (blue). Bars, 10 µm. (B) The amount of p65 in the nucleus was quantitated in at least 15 HFF cells for each strain. Asterisks indicate significantly higher levels of nuclear p65 compared with uninfected cells (*, P < 0.001, two sample Student’s t tests). (C) Mouse BMDMs (BALB/c) were infected with T. gondii strains for 24 h, fixed, and stained with α–NF-κB p65 (red), α-SAG1 (green), and Hoechst dye (blue). Bars, 10 µm. (D) The level of nuclear p65 in infected cells was quantitated in at least 12 infected cells per strain. Asterisks indicate significantly higher levels of nuclear p65 compared with uninfected cells (*, P < 0.005, two sample Student’s t tests). One replicate experiment was done in both HFFs and mouse BMDM with similar qualitative results. Horizontal bars represent the mean nuclear p65 intensity over all cells.
GRA15 mediates NF-κB p65 translocation
We quantified the nuclear p65 IF signal of cells infected with type I, II, or III strains, confirming that a subset of type II–infected cells had a high level of nuclear p65 (intensity >800), resulting in a significant difference in the average nuclear p65 of type II–infected cells compared with type I or type III–infected cells (Fig. 3, A and B). Type I (but not type III)–infected cells did have significantly more nuclear NF-κB p65 than uninfected cells. Type I and type III strains engineered to stably express a type II HA-tagged copy of GRA15 (type I GRA15II and type III GRA15II) activated NF-κB (Fig. 3, A and B). As with type II–infected cells, a subset of type I GRA15II–infected cells had very high levels of nuclear NF-κB, and the average nuclear p65 was significantly higher in these cells compared with type I–infected cells. A time-course experiment showed that this activation occurred starting 4 h after infection (Fig. S4).
To determine if type II GRA15 was necessary for NF-κB p65 translocation in host cells, we generated type II GRA15KO strains (Fig. S5). Removal of the GRA15 locus abolished p65 translocation by type II parasites in host cells, eliminating the subset of infected cells with a high level of nuclear p65 (Fig. 3, A and B). The level of nuclear p65 in cells infected with type II GRA15KO parasites was not significantly different from the level in uninfected cells (Fig. 3 B). To confirm that GRA15 is responsible for the NF-κB activation phenotype, we transfected GRA15II back into a type II GRA15KO strain. NF-κB p65 nuclear translocation was rescued in this type II GRA15KO GRA15II strain (Fig. 3, A and B). Additionally, a type I GRA15KO GRA15II strain activated translocation of p65, and infection with this strain or a type I GRA15II strain did not activate significantly different levels of nuclear p65, confirming that the type II copy of GRA15 alone was sufficient for the nuclear translocation of p65 by type I strains of T. gondii. Similarly, in mouse BMM, infection with type II strains, but not type I/III strains, activated a high level of p65 nuclear translocation. This activation was also a result of the type II GRA15 gene (Fig. 3, C and D).
To examine if p65 was the only NF-κB family subunit activated by GRA15, we infected HFFs with type I GRA15II parasites for 24 h and examined p50, p52, RelB, and c-Rel nuclear localization by IF (Fig. S6). Only p50 was specifically and significantly translocated to the nucleus by type I GRA15II strains (Fig. S6 A). However, there was a significant increase in levels of nuclear c-Rel upon infection with both type I and type I GRA15II strains (see Fig. S6 D).
GRA15 activates NF-κB–mediated transcription
To determine if the nuclear NF-κB p65 triggered by GRA15II is transcriptionally active, we infected HFFs with type II, type II GRA15KO, type I, type I GRA15II, type III, or type III GRA15II T. gondii and hybridized RNA from the infections to Affymetrix human genome arrays to determine host cell gene expression. The expression of genes with NF-κB TFBS in their promoters and gene products belonging to an NF-κB–related pathway was enriched in type II versus type II GRA15KO infections and type I/III GRA15II versus type I/III infections (Fig. 4 A and Supplemental data 1). 146 transcripts were strongly regulated by GRA15, regardless of the T. gondii strain genetic background, and we defined these genes as core GRA15-regulated genes (Fig. 4 B). Network analysis of molecular relationships between these 146 genes and their gene products also demonstrated an enrichment of genes involved in cytokine and NF-κB signaling (Fig. 4 C).
GRA15 activates NF-κB–mediated transcription. HFFs or MEFs were infected for 18–24 h with type II, type II GRA15KO, type I, type III, type I GRA15II, or type III GRA15II T. gondii strains, and host cell gene expression was analyzed by Affymetrix microarrays. At least two arrays were done per strain in HFFs and one array was done per strain in MEFs. (A) The top three enriched known TFBSs from GSEAs comparing type II versus type II GRA15KO and type I/III GRA15II versus type I/III infections are shown. (B) For the 146 genes that are defined as core GRA15-regulated genes, mean log2 gene expression values were median centered, genes were clustered by hierarchical clustering, and a heat map is presented. The complete set of genes is listed in Supplemental data 2. (C) Ingenuity pathway analysis was done for these 146 genes. The top two scoring networks are shown. (D) In MEF arrays, 32 genes have the same expression level in uninfected unstimulated WT and p65−/− cells and are regulated by GRA15. For these genes, log2 expression values were median centered, genes were clustered by hierarchical clustering, and a heat map is shown. The complete set of genes is listed in Supplemental data 2.
GRA15 activates NF-κB–mediated transcription. HFFs or MEFs were infected for 18–24 h with type II, type II GRA15KO, type I, type III, type I GRA15II, or type III GRA15II T. gondii strains, and host cell gene expression was analyzed by Affymetrix microarrays. At least two arrays were done per strain in HFFs and one array was done per strain in MEFs. (A) The top three enriched known TFBSs from GSEAs comparing type II versus type II GRA15KO and type I/III GRA15II versus type I/III infections are shown. (B) For the 146 genes that are defined as core GRA15-regulated genes, mean log2 gene expression values were median centered, genes were clustered by hierarchical clustering, and a heat map is presented. The complete set of genes is listed in Supplemental data 2. (C) Ingenuity pathway analysis was done for these 146 genes. The top two scoring networks are shown. (D) In MEF arrays, 32 genes have the same expression level in uninfected unstimulated WT and p65−/− cells and are regulated by GRA15. For these genes, log2 expression values were median centered, genes were clustered by hierarchical clustering, and a heat map is shown. The complete set of genes is listed in Supplemental data 2.
Type I T. gondii parasites also activate NF-κB p65 translocation in HFFs, although at a much lower level than type II strains (Fig. 3). Microarray analysis confirmed that a type I strain can cause the activation of NF-κB–mediated transcription. The expression of genes with NF-κB TFBS in their promoters and gene products belonging to an NF-κB–related pathway was enriched in type I–infected HFFs over uninfected HFFs (Supplemental data 1). This activation is not dependent on GRA15; the expression of NF-κB regulated genes was enriched in a type I GRA15KO infection compared with uninfected cells, and not in a type I infection compared with a type I GRA15KO infection (unpublished data).
We also infected WT and p65−/− MEFs with type II, type II GRA15KO, type I, or type I GRA15II T. gondii strains and analyzed host cell gene expression by microarray (Fig. 4 D). In WT MEFs, we found 32 genes to be core GRA15-regulated genes (more than twofold different in type II vs. type II GRA15KO or type I GRA15II vs. type I infections and similar expression level in uninfected unstimulated WT and p65−/− MEFs). Of these 32 genes, only three are also GRA15 regulated in p65−/− host cells. This data indicates that the majority of host cell transcription induced by GRA15 was activated via the canonical p65/p50 NF-κB heterodimer; however, it is possible that a small subset of genes was activated by other NF-κB subunits, such as c-Rel/p50 dimers, or other transcription factors.
GRA15 is a polymorphic secreted dense granule protein
The GRA15 coding region is predicted to be 1,908 bp in type I and III strains, but only 1,653 bp in type II strains, as a result of either an insertion or deletion (indel; ToxoDB.org, v6.0). An intron is predicted in the type I and type III copies, very close to the indel. We sequenced the region around this indel of both type I and type II cDNA. Neither strain was found to have an intron in this region, and the indel was confirmed. To determine the full transcript of GRA15II, we performed 5′ and 3′ rapid amplification of cDNA ends (RACE). Two GRA15II transcription start sites were found, one at −508 to −504 bp upstream of ATG and one at −277 bp upstream of ATG. Three polyadenylation sites were found, at +930, +992, and +1,144 bp downstream of the stop codon.
ToxoDB.org currently contains sequences from one strain for each of the three North American/European clonal lineages of Toxoplasma: GT1 (type I), ME49 (type II), and VEG (type III). The genomic sequences of strains within the same lineage are thought to be very similar; however, we sequenced the GRA15 locus from RH (type I), Pru (type II), and CEP (type III) strains as well, three other strains which we have used in our experiments. Although Pru and CEP have an identical sequence to ME49 and VEG, respectively, the RH sequence contains a frameshift mutation at base 872. This frameshift leads to the mutation of a stretch of 22 amino acids (2 remain conserved), followed by a premature stop codon, truncating the protein to 312 amino acids, instead of 635 amino acids. The type I/III and II protein sequences differ most strikingly at the 84–amino acid indel near the C terminus of the protein. Besides this indel, five other amino acids are polymorphic between types I/III and II and one other single amino acid is inserted or deleted (Fig. S7).
We next looked by IF at the localization of GRA15 in the parasite and infected host cells. To determine if GRA15 was secreted into the host cell, we performed evacuole staining on a 5-min type I GRA15II-HA infection of HFF cells using an antibody against the HA tag (Fig. 5 A). HA staining is clearly present in evacuoles, partially colocalizing with evacuoles containing rhoptry proteins, indicating that GRA15 is a secreted protein. Evacuoles containing GRA15 can also be seen after attachment of cytochalasin d–treated parasites (unpublished data). In parasitophorous vacuoles (PVs), costaining of the GRA15II-HA protein with either a rhoptry marker, ROP1, or a dense granule marker, GRA7, showed colocalization of GRA15II-HA with GRA7, with almost no overlap between the HA tag and ROP1 (Fig. 5 C). GRA15 staining overlaps GRA7 staining in the dense granules and within the PV. GRA15 also localizes to the outside of the PV membrane (PVM), and in a co-infection of type I (non-GFP) and type I GRA15II-HA (GFP) parasites, HA staining can be seen localized in the PVM and on the outside of the PVM of a parasite expressing GRA15II-HA (GFP), as well as on the outside of the PVM of a parasite not expressing GRA15II-HA (Fig. 5 B), which is consistent with dense granule localization. We therefore conclude that GRA15 is a dense granule protein.
GRA15 is a secreted dense granule protein. (A) Parasites expressing GFP and an HA-tagged copy of GRA15II were added to HFFs for 5 min to allow attachment and evacuole formation. Cells were then fixed and stained with α-HA (red), α-ROP1 (green), and Hoechst dye (blue). The HA tag is present with ROP1 in evacuoles, indicating that GRA15 can be secreted into the host cell. This experiment was repeated once with cytochalasin d–treated parasites with the same results. (B) Co-infection of type I (non-GFP) and type I GRA15II-HA (GFP) parasites, done once. Arrows indicate HA staining on the PVM of a non-GFP vacuole. (C) Co-staining of GRA15II-HA with a dense granule marker, GRA7, shows colocalization of HA staining and GRA7 in both the dense granules and the PV. Conversely, costaining of GRA15II-HA with a rhoptry marker, ROP1, shows almost no overlap between the HA tag and the rhoptries. Co-staining was done once, but the same GRA15II-HA staining pattern has been observed in more than five independent experiments. Bars, 5 µm.
GRA15 is a secreted dense granule protein. (A) Parasites expressing GFP and an HA-tagged copy of GRA15II were added to HFFs for 5 min to allow attachment and evacuole formation. Cells were then fixed and stained with α-HA (red), α-ROP1 (green), and Hoechst dye (blue). The HA tag is present with ROP1 in evacuoles, indicating that GRA15 can be secreted into the host cell. This experiment was repeated once with cytochalasin d–treated parasites with the same results. (B) Co-infection of type I (non-GFP) and type I GRA15II-HA (GFP) parasites, done once. Arrows indicate HA staining on the PVM of a non-GFP vacuole. (C) Co-staining of GRA15II-HA with a dense granule marker, GRA7, shows colocalization of HA staining and GRA7 in both the dense granules and the PV. Conversely, costaining of GRA15II-HA with a rhoptry marker, ROP1, shows almost no overlap between the HA tag and the rhoptries. Co-staining was done once, but the same GRA15II-HA staining pattern has been observed in more than five independent experiments. Bars, 5 µm.
GRA15 affects total levels of phospho-IκBα but does not affect PVM-associated phospho-IκBα
The nuclear translocation of NF-κB transcription factor subunits is dependent on the phosphorylation and degradation of an inhibitory protein, IκBα. We determined if GRA15 affected the overall levels of phospho-IκBα in infected cells by Western blotting and quantified the fraction of phosphorylated IκBα compared with the total level of IκBα (Fig. 6 A). Infection with a strain of T. gondii expressing GRA15II led to an increase in the fraction of total IκBα that was phosphorylated, although not to the extent of TNF-induced levels. This indicates that GRA15 activates NF-κB through the phosphorylation of IκBα.
GRA15 activity affects total levels of phospho-IκBα but not PVM-associated phospho-IκBα. (A) HFFs were infected with T. gondii strains for 24 h or stimulated with TNF for indicated times, and cell lysates were collected, run on an SDS-PAGE gel, and Western blotted for phospho-IκBα, total IκBα, SAG1 (parasite loading control), and GAPDH (host cell loading control). From the total IκBα blot, the fraction of phosphorylated IκBα was determined by comparing the intensity of the upper band (phosphorylated form) to the total intensity of the lower and upper band. This experiment was repeated once with type I and type I GRA15II strains only with similar results. (B) HFFs were coinfected with type I (non-GFP) and type II (GFP) parasites, fixed, and stained with α–phospho-IκBα (red) and Hoechst dye (blue). A type I PVM (non-GFP, arrowhead) and a type II PVM (GFP, arrow) are indicated. Bar, 5 µm. (C) HFFs were infected with T. gondii strains expressing GFP for 24 h, fixed, and stained with α–phospho-IκBα (red) and Hoechst dye (blue). This experiment has been done three times with similar results. Bars, 1 µm.
GRA15 activity affects total levels of phospho-IκBα but not PVM-associated phospho-IκBα. (A) HFFs were infected with T. gondii strains for 24 h or stimulated with TNF for indicated times, and cell lysates were collected, run on an SDS-PAGE gel, and Western blotted for phospho-IκBα, total IκBα, SAG1 (parasite loading control), and GAPDH (host cell loading control). From the total IκBα blot, the fraction of phosphorylated IκBα was determined by comparing the intensity of the upper band (phosphorylated form) to the total intensity of the lower and upper band. This experiment was repeated once with type I and type I GRA15II strains only with similar results. (B) HFFs were coinfected with type I (non-GFP) and type II (GFP) parasites, fixed, and stained with α–phospho-IκBα (red) and Hoechst dye (blue). A type I PVM (non-GFP, arrowhead) and a type II PVM (GFP, arrow) are indicated. Bar, 5 µm. (C) HFFs were infected with T. gondii strains expressing GFP for 24 h, fixed, and stained with α–phospho-IκBα (red) and Hoechst dye (blue). This experiment has been done three times with similar results. Bars, 1 µm.
Previously, a type I T. gondii protein extract was found to have IκBα-phosphorylating activity in vitro (Molestina et al., 2003). There is also evidence that this kinase activity can occur in vivo, as PVM-associated phospho-IκBα can still be observed in infected IKK-α/β double knockout MEFs (Molestina and Sinai, 2005a). However, in these IKK-α/β double knockout cells, after type I T. gondii infection NF-κB does not translocate to the nucleus or bind to DNA in vitro, and NF-κB–mediated gene expression is severely decreased (Molestina and Sinai, 2005b). To determine if the accumulation of phospho-IκBα on the PVM correlated with NF-κB activation, we infected HFFs with T. gondii strains and stained infected cells with a phospho-IκBα antibody. Although infection with type II parasites activated NF-κB to a much greater extent and led to higher levels of total phospho-IκBα compared with type I parasite infection, in a mixed infection of type I (non-GFP) and type II (GFP) T. gondii, phospho-IκBα accumulated almost exclusively on type I vacuoles (Fig. 6 B). We also found that the accumulation of phospho-IκBα at the PVM was independent of GRA15, as both type I and type I GRA15II PVMs accumulated phospho-IκBα, and neither type II nor type II GRA15KO PVMs accumulated visible levels of phospho-IκBα (Fig. 6 C). Phospho-IκBα also did not accumulate on type III PVMs (Fig. 6 C). We conclude that the accumulation of PVM-associated phospho-IκBα is specific to type I parasites and is not correlated with the overall level of NF-κB activation in the host cell.
GRA15 activation of NF-κB is dependent on the IKK complex and TRAF6 but independent of MyD88 and TRIF
BLAST and Pfam searches for proteins with similar amino acid sequences or domains to GRA15II returned no significant results, providing no clues to the mechanism of GRA15 NF-κB activation (Altschul et al., 1990; Finn et al., 2008). To start to answer this question, we determined which components of the NF-κB signaling pathway were necessary for GRA15 activity. When NF-κB is activated, IκB proteins are phosphorylated and then degraded by the proteasome. We previously determined that GRA15 leads to the phosphorylation of IκBα (Fig. 6), and to determine if GRA15 activity is dependent on the proteasome, we pretreated cells with MG132, a proteasomal inhibitor. Our results show that activation of NF-κB by GRA15 required functional proteasomal degradation (Fig. 7 A). IκB proteins are normally phosphorylated by the IKK complex, consisting of IKK-α, IKK-β, and IKK-γ (NEMO). IKK-γ is a regulatory subunit in the complex, whereas the β and α subunits are active kinases. IKK-β has greater kinase activity than IKK-α and is the principal kinase responsible for the phosphorylation of IκBα (Ghosh and Karin, 2002; Li and Verma, 2002). In WT MEFs, a type I GRA15II strain induces a 3.4-fold increase in nuclear p65 compared with uninfected cells. However, in IKK-β−/− MEFs, this increase is only 1.9-fold (Fig. 7 B). Many pathogens activate NF-κB via TLR agonists, and TLR signaling is mediated by the MyD88 and TRIF adaptor proteins. However, GRA15 is able to activate p65 nuclear translocation in MyD88/TRIF double knockout cells, indicating that it is not just a TLR ligand (Fig. 7 D). GRA15-mediated p65 activation is also not dependent on RIP1 but is dependent on TRAF6 (not depicted and Fig. 7 C). Thus GRA15 appears to modulate NF-κB at a specific step in the pathway downstream of MyD88 and TRIF but upstream of, or in a complex with, TRAF6 and IKK proteins.
GRA15 activity is dependent on IKK-β and TRAF6 and independent of MyD88 and TRIF. Cells were infected with type I GRA15II or type I parasites for 4 h, stimulated with 20 ng/ml TNF for 1 h, or left unstimulated (US) and uninfected (UI). Cells were fixed and probed with an α–NF-κB p65 antibody and mean nuclear staining was measured. (A) HFF cells were preincubated with media containing 20 ng/ml MG-132 proteasome inhibitor before infection and TNF stimulation. (B–D) The activity of GRA15II in the absence of different components of the NF-κB pathway was assayed. (B) IKK-β−/− MEFs. (C) TRAF6−/− MEFs. (D) MyD88−/−/TRIF−/− BMMs. These experiments were repeated at least two times and quantification was performed on a representative experiment for each factor assayed. Asterisks (*) indicate data are significantly different (P-value < 0.05, Student’s t test), and n.s. indicates data are not significantly different. Horizontal bars represent the mean nuclear p65 intensity over all cells.
GRA15 activity is dependent on IKK-β and TRAF6 and independent of MyD88 and TRIF. Cells were infected with type I GRA15II or type I parasites for 4 h, stimulated with 20 ng/ml TNF for 1 h, or left unstimulated (US) and uninfected (UI). Cells were fixed and probed with an α–NF-κB p65 antibody and mean nuclear staining was measured. (A) HFF cells were preincubated with media containing 20 ng/ml MG-132 proteasome inhibitor before infection and TNF stimulation. (B–D) The activity of GRA15II in the absence of different components of the NF-κB pathway was assayed. (B) IKK-β−/− MEFs. (C) TRAF6−/− MEFs. (D) MyD88−/−/TRIF−/− BMMs. These experiments were repeated at least two times and quantification was performed on a representative experiment for each factor assayed. Asterisks (*) indicate data are significantly different (P-value < 0.05, Student’s t test), and n.s. indicates data are not significantly different. Horizontal bars represent the mean nuclear p65 intensity over all cells.
GRA15 expressed in Hela cells is sufficient to activate NF-κB
We wanted to determine whether GRA15II alone is sufficient to activate p65 nuclear translocation or if other T. gondii secreted factors that are common to all type I, II, and III strains are also needed for this process. When we transiently transfected HeLa cells with a vector expressing the type II copy of GRA15 N-terminally fused with GFP, the nuclei of transfected GFP-positive cells contained p65, whereas the nuclei of nontransfected GFP-negative cells in the same culture did not (Fig. 8 A). The level of this nuclear localization is equivalent with activation by intracellular type I GRA15II parasites (Fig. 8 B). Expression of 55.m04955, an unrelated T. gondii protein, or Mob1A, a human protein present in the original vector, did not induce p65 translocation, indicating that this NF-κB activation is not the result of cell stress from protein overexpression (Fig. 8 A). GRA15II expression alone is therefore sufficient to recapitulate the induction of p65 nuclear translocation.
GRA15 expression alone is sufficient to activate NF-κB in HeLa cells. (A) HeLa cells were transfected with GRA15II, an unrelated T. gondii gene (55.m04955), or an unrelated human gene (Mob1A) fused to GFP. Cells were then fixed and stained with α–NF-κB p65 (red) and Hoechst dye (blue). All cells expressing GRA15II-GFP contain activated NF-κB p65, whereas cells expressing 55.m04955-GFP or Mob1A-GFP do not. Nontransfected non-GFP cells in the same culture also have no nuclear NF-κB p65. This experiment was repeated two more times with the same results. (B) HeLa cells were infected with type I GRA15II or type I parasites for 24 h, fixed, and stained with α–NF-κB p65 (red) and Hoechst dye (blue). Cells infected with a type I GRA15II strain contain comparable amounts of nuclear NF-κB p65 to transfected cells. This experiment was repeated a second time with similar results. Bars, 10 µm.
GRA15 expression alone is sufficient to activate NF-κB in HeLa cells. (A) HeLa cells were transfected with GRA15II, an unrelated T. gondii gene (55.m04955), or an unrelated human gene (Mob1A) fused to GFP. Cells were then fixed and stained with α–NF-κB p65 (red) and Hoechst dye (blue). All cells expressing GRA15II-GFP contain activated NF-κB p65, whereas cells expressing 55.m04955-GFP or Mob1A-GFP do not. Nontransfected non-GFP cells in the same culture also have no nuclear NF-κB p65. This experiment was repeated two more times with the same results. (B) HeLa cells were infected with type I GRA15II or type I parasites for 24 h, fixed, and stained with α–NF-κB p65 (red) and Hoechst dye (blue). Cells infected with a type I GRA15II strain contain comparable amounts of nuclear NF-κB p65 to transfected cells. This experiment was repeated a second time with similar results. Bars, 10 µm.
GRA15 affects in vitro parasite growth
To assay the effect of GRA15 on in vitro parasite growth, we infected monolayers of fibroblasts with type I, type I GRA15II, type II, or type II GRA15KO parasites, allowed the parasites to grow for 4–7 d, and then quantified the area of plaques formed on the monolayers. In HFF host cells, a type II GRA15KO strain formed significantly larger plaques than a type II strain (P = 0.002, Student’s t test), and a type I strain formed significantly larger plaques than a type I GRA15II strain (P = 0.024, Student’s t test; Fig. S8 A). However, in MEF host cells, type II and type II GRA15KO strains did not make significantly different sized plaques (P = 0.841, Student’s t test), and the same was true of type I and type I GRA15II strains (P = 0.371, Student’s t test; Fig. S8 A). This data indicates that GRA15 inhibits in vitro parasite growth in human fibroblasts but not mouse fibroblasts.
GRA15 affects IL-12 production in vitro
In vitro infection of macrophages with different strains of T. gondii results in different levels of IL-12p40 secretion, with type II strains inducing much higher levels of IL-12p40, and some of this variation has been suggested to be a result of strain differences in NF-κB activation (Robben et al., 2004; Kim et al., 2006; Saeij et al., 2007). To examine the role of GRA15-mediated NF-κB activation in the induction of IL-12p40 secretion, mouse BMMs were infected with type I, type I GRA15II, type II, or type II GRA15KO T. gondii, and levels of IL-12p40 in the supernatant were determined by cytokine ELISA (Fig. 9 A). As expected, type I–infected BMMs secrete a low level of IL-12p40 that is not significantly higher than the level secreted by uninfected cells, whereas type II induces a high level of IL-12p40 secretion. When GRA15 is removed from the type II strain, IL-12p40 secretion decreases more than sixfold, implying a considerable role for this gene product in modulating host cell IL-12 signaling (P = 0.001, Student’s t test). Similarly, the introduction of GRA15II in a type I strain leads to a significant increase in IL-12p40 secretion by BMM (P = 0.008, Student’s t test). IL-12p70 secretion was also higher after type II infection, compared with a type I infection, and partially dependent on the presence of a type II copy of GRA15 (unpublished data).
GRA15II promotes IL-12 secretion in vitro and affects parasite growth and host cytokine production in vivo. (A) BALB/c BMMs were infected with T. gondii strains for 24 h, supernatants were collected, and IL-12p40 levels were determined by cytokine ELISA. These experiments were performed at least three times in BMM using triplicate samples, as well as in RAW264.7 macrophages, all with similar results. (B–D) Mice were infected i.p. with tachyzoites of either a type II or a type II GRA15KO strain. (B) C57BL/6 or BALB/c mice were infected with 5,000 tachyzoites and survival of mice was monitored. In one experiment, five BALB/c mice were infected per strain, and in three separate experiments, a total of 25 C57BL/6 mice were infected per strain. (C) BALB/c mice were infected with parasites that express the enzyme luciferase. 5 d after infection, mice were i.p. injected with luciferin, anesthetized, and the flux (photons/sec/cm2/sr) was determined as a measure of parasite burden. Mice infected with a type II GRA15KO strain had significantly greater total flux (p/s) and, therefore, significantly greater parasite burden than mice infected with a type II strain. This burden difference 5 d after infection was observed in three independent experiments. (D) 1 or 2 d after infection, infected BALB/c mice were euthanized and an i.p. cavity wash was collected for IFN-γ, IL-12p70, and IL-12p40 cytokine ELISA. On day 2 after infection, mice infected with a type II GRA15KO strain had significantly lower levels of IFN-γ in the i.p. cavity than mice infected with a type II strain (P = 0.05, two-sample Student’s t test). Five mice were infected per strain per day. Day 2 cytokine levels were measured in a separate experiment with similar results. Error bars represent standard deviation from one experiment.
GRA15II promotes IL-12 secretion in vitro and affects parasite growth and host cytokine production in vivo. (A) BALB/c BMMs were infected with T. gondii strains for 24 h, supernatants were collected, and IL-12p40 levels were determined by cytokine ELISA. These experiments were performed at least three times in BMM using triplicate samples, as well as in RAW264.7 macrophages, all with similar results. (B–D) Mice were infected i.p. with tachyzoites of either a type II or a type II GRA15KO strain. (B) C57BL/6 or BALB/c mice were infected with 5,000 tachyzoites and survival of mice was monitored. In one experiment, five BALB/c mice were infected per strain, and in three separate experiments, a total of 25 C57BL/6 mice were infected per strain. (C) BALB/c mice were infected with parasites that express the enzyme luciferase. 5 d after infection, mice were i.p. injected with luciferin, anesthetized, and the flux (photons/sec/cm2/sr) was determined as a measure of parasite burden. Mice infected with a type II GRA15KO strain had significantly greater total flux (p/s) and, therefore, significantly greater parasite burden than mice infected with a type II strain. This burden difference 5 d after infection was observed in three independent experiments. (D) 1 or 2 d after infection, infected BALB/c mice were euthanized and an i.p. cavity wash was collected for IFN-γ, IL-12p70, and IL-12p40 cytokine ELISA. On day 2 after infection, mice infected with a type II GRA15KO strain had significantly lower levels of IFN-γ in the i.p. cavity than mice infected with a type II strain (P = 0.05, two-sample Student’s t test). Five mice were infected per strain per day. Day 2 cytokine levels were measured in a separate experiment with similar results. Error bars represent standard deviation from one experiment.
The T. gondii polymorphic rhoptry kinase ROP16 also affects IL-12 secretion, and previous microarray analyses determined that the expression of many GRA15-regulated genes are also strongly affected by a locus on chromosome VIIb, where ROP16 resides (Saeij et al., 2007). In these analyses, some genes, such as SOCS-2 and SOCS-3, were regulated by loci on both chromosome VIIb and X (Fig. S9 A). We wondered if the effect of ROP16 on these genes was through modulation of the NF-κB pathway. We infected an NF-κB GFP reporter cell line with a type II strain or a type II transgenic strain expressing a type I copy of ROP16. A significantly greater percentage of type II–infected cells than type II ROP16I–infected cells are GFP positive (P = 1.5 × 10−7, χ2 test), indicating that in a type II background, a type I copy of ROP16 significantly inhibits NF-κB activation (Fig. S9 B). In an NF-κB luciferase reporter cell line, infection with a type II ROP16I strain also induced significantly less luciferase activity than infection with a type II strain (unpublished data). Thus, both GRA15 and ROP16 affect NF-κB activity.
GRA15 affects in vivo parasite growth and cytokine production
To assay the effect of GRA15 on parasite virulence, we infected C57BL/6 or BALB/c mice by i.p. injection with 5,000 tachyzoites of a type II or a type II GRA15KO strain and monitored mouse survival during the acute phase of infection (days 0–20; Fig. 9 B). C57BL/6 mice infected with either strain succumbed to infection at the same time, between days 5 and 18 after injection. Additionally, the same percentage of mice in each group survived, ∼40%. BALB/c mice infected with either a type II or a type II GRA15KO strain did not die after infection.
To determine the effect of GRA15 on in vivo parasite burden, BALB/c mice infected i.p. with tachyzoites of either a type II or a type II GRA15KO strain that express the enzyme luciferase were imaged throughout infection. At day 5 after infection, mice infected with a type II GRA15KO strain had a significantly higher parasite burden than mice infected with a type II strain (Fig. 9 C; P = 0.01, Student’s t test). Similarly, expression of GRA15II in a type I strain inhibited in vivo parasite growth (Fig. S8 B). Together, these experiments indicate that GRA15 inhibits in vivo parasite growth in both a type I and a type II background.
GRA15 affects IL-12 secretion by BMM in vitro, and we also assessed whether it affects cytokine secretion in vivo at the site of infection. BALB/c mice were infected i.p. with tachyzoites of either a type II or a type II GRA15KO strain. 1 or 2 d after infection, mice were sacrificed, the i.p. cavity was washed, and cytokine levels in the wash were determined by ELISA. On day 2 after infection, mice infected with a type II GRA15KO strain had significantly less IFN-γ in their i.p. cavities than mice infected with a type II strain (Fig. 9 D; P = 0.05, Student’s t test). Although differences in IL-12p40 or p70 were not significant on either day 1 or day 2 because of large variations between mice, the average cytokine levels in type II GRA15KO–infected mice were consistently lower than cytokine levels in type II–infected mice. At these early time points, there was not a significant difference in parasite load between mice infected with either strain, as determined by imaging (unpublished data).
DISCUSSION
The modulation of the NF-κB pathway by T. gondii has long been an area of debate, with some studies stating that T. gondii activates NF-κB and others that T. gondii inhibits NF-κB activation. In this study, we have conclusively shown that the three North American clonal lineages of T. gondii differ in their activation of the host NF-κB pathway; type II strains activate a high level of NF-κB p65 translocation, whereas type I and III strains do not (Fig. 3). Using F1 progeny from a type II × type III cross, we found that a locus on chromosome X is responsible for this polymorphic phenotype, and we identified the novel T. gondii factor GRA15 at this locus (Fig. S2). The type II copy of GRA15 (63.m00001) is necessary in type II strains and sufficient in type I and III strains for NF-κB nuclear translocation and transcriptional activity (Figs. 3 and 4; and Supplemental data 1). We have observed this activation in a variety of human, mouse, and rat cell types, indicating that GRA15’s activity is independent of host cell type and species. Additionally, when GRA15II is expressed in HeLa cells, it is sufficient to activate NF-κB (Fig. 8 A).
It had been previously reported that infection with type I strains of T. gondii activates NF-κB in MEF and Henle 407 intestinal epithelial host cells (Molestina et al., 2003; Ju et al., 2009). Careful quantification of IF experiments showed that type I strains do slightly activate NF-κB p65 translocation, but the level of nuclear p65 is much higher in type II–infected cells than type I–infected cells (Fig. 3). It is possible that other groups have concluded that type I strains do not activate NF-κB because this activation is so low compared with that of LPS or TNF stimulation (Butcher et al., 2001; Shapira et al., 2002, 2005). Our IF experiments also showed that a type I strain activates c-Rel nuclear translocation, which might also affect host cell transcription (Fig. S6). By microarray analysis, some NF-κB–regulated genes are induced by type I infection, but a much greater number are induced by type II infection, and infection with a type I strain does not activate detectable GFP or luciferase transcription from an NF-κB reporter cell line (unpublished data), which is in concordance with another published study (Shapira et al., 2005). This low-level activation is also not dependent on GRA15, as there is no enrichment in NF-κB activation in a type I infection over a type I GRA15KO infection. Type III strains do not activate any p65 nuclear translocation or NF-κB–mediated transcription.
Our results agree with previous observations that phosphorylated IκBα accumulates on the PVM upon T. gondii type I infection (Butcher et al., 2001; Molestina et al., 2003; Shapira et al., 2005). However, levels of PVM-associated phospho-IκBα are not correlated with total levels of phospho-IκBα or the level of NF-κB transcriptional activity in the host cell. Phospho-IκBα is not observed on type II PVMs, and, although infection with T. gondii strains expressing GRA15II leads to higher total levels of phospho-IκBα, the presence of GRA15 does not affect PVM-associated phospho-IκBα (Fig. 6). A type I T. gondii protein capable of phosphorylating IκBα may play a role in low-level activation of NF-κB in type I strains (Molestina and Sinai, 2005a). However, it is clear that the GRA15 protein in type II strains activates NF-κB to a much greater extent.
In contrast, some groups have reported that infection with type I T. gondii strains inhibits NF-κB activation after stimulation with the cytokine TNF or the TLR ligand LPS (Butcher et al., 2001; Shapira et al., 2002; Kim et al., 2004). This inhibition has been observed after <6 h of infection in HFFs (Shapira et al., 2005), mouse BMM (Shapira et al., 2002; Kim et al., 2004), and thioglycolate-elicited cells (Butcher et al., 2001) and was not observed after 6 or 12 h of infection in mouse BMM (Kim et al., 2004; Leng et al., 2009). Our experiments confirm that type I T. gondii cannot inhibit LPS-stimulated NF-κB nuclear translocation or TNF-stimulated NF-κB transcriptional activity at a late time point in infection (Fig. 2, A and C). But, after a short infection (<5 h), we also do not observe inhibition of NF-κB p65 nuclear translocation or NF-κB–mediated transcription, contradicting previous studies (Fig. 2, A and B). It is true that some infected cells do not respond to LPS or TNF stimulation, and this observation may have lead to the conclusion that type I strains can inhibit NF-κB signaling. However, quantification of NF-κB p65 translocation and NF-κB reporter transcription in many uninfected and infected cells shows that preinfection with T. gondii does not alter the response of populations of cells to these stimuli.
The GRA15 gene product is a novel T. gondii dense granule protein that is secreted into the host cell upon invasion (Fig. 5), representing the first identified Apicomplexan dense granule protein that can modulate host cell signaling pathways. Rhoptry proteins have already been identified as parasite factors that can alter host cell behavior, but our findings indicate that dense granule proteins should be viewed as candidate factors as well. It is still unclear why only the type II copy of GRA15 activates NF-κB. Amino acid polymorphisms between the type II and type I/III copies of GRA15, including an insertion/deletion and several single amino acid changes, or the expression level of GRA15 may be responsible for this polymorphic phenotype.
Although the GRA15 protein is secreted into the host cell upon parasite invasion, cells infected with type I GRA15II parasites did not have substantial p65 in their nuclei until ∼4 h after infection (Fig. S4), and in our experiments NF-κB activation was usually assayed 18–25 h after infection. These slow kinetics are not unprecedented; Rac GTPase has been shown to initiate NF-κB nuclear translocation with a time course similar to that of GRA15 (Boyer et al., 2004). Additionally, the amount of GRA15 interacting with host cell proteins likely increases after PV formation as dense granule proteins are made and continuously secreted from intracellular parasites, and higher levels of GRA15 may be necessary to initiate NF-κB activation.
The precise mechanism by which GRA15 activates NF-κB has yet to be discovered. Our data suggests that GRA15 initiates canonical NF-κB activation, which preferentially induces the p65/p50 heterodimer (Hayden and Ghosh, 2004). A simple hypothesis is that GRA15 acts as a TLR ligand to activate the canonical NF-κB pathway; however, the activation of p65 translocation by GRA15 is not dependent on either MyD88 or TRIF, two proteins which are essential for TLR signaling. We did find that the activity of GRA15 is dependent on both IKK-β and TRAF6, suggesting that GRA15 acts either upstream of or in a complex with these proteins. A type I GRA15II strain activated more nuclear p65 translocation in WT MEFs than in IKK-β−/− or TRAF6−/− MEFs, although in IKK-β−/− cells this difference was not significant (P = 0.12). This is probably because IKK-α also has phosphorylating activity. Our microarray data further defines the placement of GRA15 in the NF-κB signaling pathway. GRA15 is able to constitutively activate NF-κB, but this activation leads to the expression of negative feedback regulators, such as the deubiquitinating enzymes A20 (TNFAIP3) and CYLD, which normally act to quickly down-regulate NF-κB signaling. Because TRAF6 and the IKK complex are both targets of these deubiquitinating enzymes, it is likely that GRA15 acts in concert with these proteins rather than upstream (Sun, 2008). We are currently looking for direct binding partners of GRA15 by coimmunoprecipitation.
We found that a type I copy of ROP16 can inhibit NF-κB activation in a type II strain (Fig. S9 B). Why this inhibition occurs in a type II strain but not a type I strain is unknown, but the genetic backgrounds of type I and type II strains are very different and other polymorphic factors likely exist that affect STAT and/or NF-κB signaling pathways. How ROP16 inhibits NF-κB activation is also unclear, but it is likely to be through its activation of STAT6 and/or STAT3 (Ohmori and Hamilton, 2000; Nelson et al., 2003; Butcher et al., 2005; Hoentjen et al., 2005). This inhibition has significant consequences, for example, GRA15 and ROP16 have opposing effects on the expression levels of many genes, including IL-12, a particularly important cytokine in T. gondii infection (Fig. 9 A; Gazzinelli et al., 1994; Saeij et al., 2007). In fact, the single amino acid difference in ROP16 that causes it to be less active in type II strains (Yamamoto et al., 2009) may have been selected for in type II strains specifically to increase NF-κB activation by GRA15II. However, GRA15 and ROP16 are expected to have additive or synergistic effects on the expression of other genes, such as the SOCS genes (Fig. S9 A). In any case, the modulation of host cell gene expression will depend upon the exact allelic combination of a variety of factors that T. gondii possesses, including GRA15 and ROP16.
Early in infection (days 1–5), GRA15 affected both cytokine production and parasite growth in vivo. When a host is first infected by live parasites, type II strains expressing GRA15 activate NF-κB in host cells and induce IL-12 secretion, whereas infection with type II GRA15KO parasites or type I/III strains does not cause this early activation. IL-12 stimulates NK cells and T cells to secrete IFN-γ, and the observed effect of GRA15 on IFN-γ levels was likely via an effect on IL-12. Although levels of IL-12 in the i.p. cavity were not significantly different between mice infected with a type II or type II GRA15KO strain, the levels were consistently lower in type II GRA15KO–infected mice, and this difference may have been enough to lead to a significant difference in IFN-γ levels (Fig. 9 D). IFN-γ is the main mediator of host resistance to T. gondii, and differences in IFN-γ levels probably also explain the growth difference that we observed in vivo between a type II and type II GRA15KO strain at a slightly later time point 5 d after infection (Fig. 9 C). However, as the infection progresses and parasites lyse out of host cells, pathogen-associated molecular pattern proteins within the PV, such as profilin and cyclophilin, are released, and NF-κB will be activated via TLR signaling and CCR5 signaling by all strains. At this stage, IL-12 production, IFN-γ production, and parasite growth are then independent of the GRA15 locus, which might explain why a type II strain does not differ in overall virulence from a type II GRA15KO strain (Fig. 9 B). The location of GRA15 on chromosome X does indicate that it could represent a previously identified chromosome X virulence locus (Saeij et al., 2006).
Plaque assays showed that GRA15 also affects parasite growth in vitro (Fig. S8 A). However, this in vitro effect occurred specifically in human cells and not in mouse cells. The cause of this difference is currently unknown, but one possibility is that genes affecting amino acid levels, lipid levels, or levels of other nutrients are partially NF-κB regulated in human cells but not in mouse cells.
GRA15II may also have other effects in vivo that remain untested. As an intracellular pathogen, T. gondii must use host cells to traffic through the body of the host animal. NF-κB activation by GRA15II increases expression of many chemokines and adhesion molecules (our microarray data), and strain differences in NF-κB activation may therefore lead to differences in the ability to induce migration of host cells, as previously reported (Lambert et al., 2006, 2009). Infection of hosts with other pathogens is also highly relevant to disease outcome, and the activation of NF-κB by T. gondii might increase transcription of HIV retroviral sequences with NF-κB binding sites in their promoters (Gazzinelli et al., 1996). Lastly, NF-κB activation leads to a proinflammatory Th1-type immune response that may promote inflammatory disease manifestations such as encephalitis and colitis, both of which have been observed mainly after type II strain infections (Hunter and Remington, 1994; Liesenfeld, 1999).
MATERIALS AND METHODS
Parasites and cells.
Parasites were maintained in vitro by serial passage on monolayers of HFFs at 37°C in 5% CO2. RH or GT1 were used as representative type I strains, ME49 or Pru as representative type II strains, and CEP or VEG as representative type III strains. A Pru strain engineered to express firefly luciferase and GFP (Pru ΔHXGPRT A7; Kim et al., 2007), and CEP and RH strains engineered to express click beetle luciferase and GFP (CEP HXGPRT- C22 and RH 1–1; Boyle et al., 2007), have been described previously. Pru and RH strains expressing HXGPRT and generated from unsuccessful knockout transfections were used in assays comparing parasite growth. F1 progeny from type II × type III crosses were described previously (Sibley et al., 1992; Khan et al., 2005). A Pru strain expressing a type I copy of ROP16 was also described previously (Saeij et al., 2007). HFFs were grown in DME (Invitrogen) supplemented with 10% heat inactivated FBS (PAA), 2 mM l-glutamine, 50 µg/ml each of penicillin and streptomycin, and 20 µg/ml gentamycin. BMMs were obtained from female BALB/c, C57BL/6, or MyD88/TRIF double knockout (a gift from H. Ploegh, Whitehead Institute for Biomedical Research and Massachussetts Institute of Technology, Cambridge, MA) mice. BM was isolated by flushing hind tibias and femurs using a 27-gauge needle and/or by crushing the bones using a mortar and pestle, followed by passage over a cell strainer. Cells were suspended in DME supplemented with 20% L929 cell-conditioned medium, 10% heat-inactivated FBS, 2 mM l-glutamine, 1 mM sodium pyruvate, 1× MEM nonessential amino acids, and 50 µg/ml each of penicillin and streptomycin. 3–6 × 106 cells were plated in 10-cm nontissue culture–treated dishes (VWR) and incubated at 37°C, with 5% CO2 in humidified air. After 6–7 d, cells were washed with PBS to remove nonadherent cells, harvested by dislodging with a cell scraper in PBS, and replated for the assay. 293 stable cell lines with four copies of the NF-κB consensus transcriptional response element driving the expression of GFP or GFP and luciferase (System Biosciences) were grown in DME supplemented with 10% heat-inactivated FBS, 2 mM l-glutamine, 1 mM sodium pyruvate, 1× MEM nonessential amino acids, 10 mM HEPES, 50 µg/ml each of penicillin and streptomycin, and 20 µg/ml gentamycin. 293 cells were passed every 2–4 d using 0.05% trypsin-EDTA. WT MEFs were gifts from M. Karin (University of California, San Diego, School of Medicine, La Jolla, CA) and A. Sinai (University of Kentucky College of Medicine, Lexington, KY), IKK-β−/− MEFs were a gift from M. Karin (Li et al., 1999), NF-κBp65−/− MEFs were a gift from A. Sinai, and TRAF6−/− MEFs were provided by K. Fitzgerald (University of Massachussetts Medical School, Worcester, MA). MEFs were grown in DME supplemented with 10% nonheat inactivated FBS (PAA), 2 mM l-glutamine, 1 mM sodium pyruvate, 1× MEM nonessential amino acids, 10 mM Hepes, and 50 µg/ml each of penicillin and streptomycin. MEFs were passed using 0.05% trypsin-EDTA. All parasite strains and cell lines were routinely checked for Mycoplasma contamination and it was never detected.
Reagents.
Antibodies against HA (3F10; Roche), mouse NF-κB p65 (sc-8008), human NF-κB p65 (sc-109), NF-κB p50 (sc-8414), NF-κB p52 (sc-7386), NF-κB RelB (sc-28689), NF-κB c-Rel (sc-71), T. gondii surface antigen (SAG)-1 (DG52; Burg et al., 1988), T. gondii dense granule protein GRA7 (Dunn et al., 2008), T. gondii rhoptry protein ROP1 (Tg49; Ossorio et al., 1992), phospho-IκBα (sc-8404), total IκBα (sc-847), and GAPDH (sc-32233) were used in the IF assay or in Western blotting. IF secondary antibodies were coupled with Alexa Fluor 488 or Alexa Fluor 594 (Invitrogen). Secondary antibodies used in Western blotting were conjugated to peroxidase (Kirkegaard & Perry Laboratories). Purified LPS (EMD), recombinant mouse TNF (AbD Serotec), and recombinant human TNF (Invitrogen) were used to stimulate cells. MG-132 (EMD) was used in proteasomal inhibition.
IF.
Parasites were allowed to invade cells on coverslips and incubated for different time points. The cells were then fixed with 3% (vol/vol) formaldehyde in PBS for 20 min at room temperature, permeabilized with 100% ethanol and/or 0.2% (vol/vol) Triton X-100, and blocked in PBS with 3% (wt/vol) BSA and 5% (vol/vol) goat serum. Coverslips were incubated with primary antibody for 1 h at room temperature or overnight at 4°C, and fluorescent secondary antibodies and Hoechst dye were used for antigen and DNA visualization, respectively. Coverslips were mounted on a glass slide with Vectashield (Vector Laboratories), and photographs were taken using NIS-Elements software (Nikon) and a digital camera (CoolSNAP EZ; Roper Industries) connected to an inverted fluorescence microscope (model eclipse Ti-S; Nikon). Quantification of nuclear signal was performed by randomly selecting at least 10 infected cells per T. gondii strain and measuring the mean signal intensity per nucleus using the NIS-Elements software and Hoechst dye to define nuclei. For evacuole staining, this standard IF protocol was modified slightly. Parasites were added to HFFs on coverslips, spun down to bring them into contact with host cells, and allowed to attach to and invade host cells for 5 min at 37°C. Unattached parasites were washed off with PBS, and cells were fixed 3% (vol/vol) formaldehyde in PBS for 20 min at room temperature, blocked in PBS with 5% (vol/vol) fetal bovine serum and 5% (vol/vol) normal goat serum for 1–2 h at room temperature, and permeabilized by incubation in PBS with 0.2% (wt/vol) saponin at 37°C for 20 min. For proteasomal inhibition HFF monolayers were pretreated with 20 ng/ml of MG-132 for 1 h at 37°C. Cells were infected with parasites and spun down. 4 h after MG-132 addition, cells were washed with PBS and fresh media containing no inhibitor was added. The monolayer was incubated for one additional hour before fixation. To synchronize infection during time course assays, HFF monolayers were incubated on ice with cold media for 10 min before infection. For infection, supernatant from fully lysed parasite flasks was pelleted and washed three times with PBS and resuspended in cold media. After infection, monolayers were incubated on ice for 30 min, and then unattached parasites were washed off with cold PBS. Fresh prewarmed media was added and cells were incubated at 37°C to allow invasion and infection for the determined length of time.
Generation of transgenic parasites.
The GRA15 coding region and putative promoter (1,940 bp upstream of the start codon) was amplified from type II T. gondii genomic DNA by PCR (forward, 5′-CCCAAGCTTGACTGCCACGTGTAGTATCC-3′; reverse, 5′-TTACGCGTAGTCCGGGACGTCGTACGGGTATGGAGTTACCGCTGATTGTGT-3′). Sequence coding for an HA tag was included in the reverse primer (denoted with italics) to C-terminally tag the protein. GRA15IIHA was then inserted into pCR8/GW (Invitrogen) by TOPO-TA cloning. Gateway cassette A was ligated into pTKO (gift from G. Zeiner, Stanford University School of Medicine, Stanford, California) at the EcoRV site, creating a Gateway destination vector (Invitrogen), pTKO-att (Fig. S10), and GRA15IIHA was cloned into pTKO-att by LR recombination (Invitrogen). The pTKO-att-GRA15IIHA vector was then linearized by digestion with XhoI (NEB). XhoI cuts off 244 bp of the putative promoter, leaving 1,696 bp intact upstream of the start codon. Linearized vector was transfected into RHΔHXGPRT and CEP HXGPRT− C22 parasites by electroporation. Electroporation was done in a 2-mm cuvette (Bio-Rad Laboratories) with 2 mM ATP (MP Biomedicals) and 5 mM GSH (EMD) in a Gene Pulser Xcell (Bio-Rad Laboratories), with the following settings: 25 μFD, 1.25 kV, ∞ Ω. Stable integrants were selected in media with 50 µg/ml mycophenolic acid (Axxora) and 50 µg/ml xanthine (Alfa Aesar) and cloned by limiting dilution. Expression of GRA15II was confirmed by IF for HA staining. Parasite strains already containing the HXGPRT gene (RH GRA15KO and Pru A7 GRA15KO) were cotransfected with 35 µg pTKO-att-GRA15II and 1 µg pTUB5-BLE (Soldati et al., 1995), containing the ble selectable marker. Stable integrants were selected extracellularly with 50 µg/ml Phleomycin (InvivoGen), and HA staining was confirmed by IF.
The ROP2 and GRA6 coding regions and putative promoters (at least 1,500 bp upstream of the start codon) were amplified from type II T. gondii genomic DNA by PCR (ROP2 forward, 5′-CACCGAGGTTGGAACTGTG-3′; ROP2 reverse, 5′-CTTACGCGTAGTCCGGGACGTCGTACGGGTAGATTGCCGTAACCGCCT-3′; GRA6 forward, 5′-CCCAAGCTTGAAGGACTGCGTTGAGTGTTTT-3′; and GRA6 reverse, 5′-GGAATTCTTACGCGTAGTCCGGGACGTCGTACGGGTAAAAATCAAACTCATTCACACTTC-3′). Sequence coding for an HA tag was included in the reverse primers (denoted with italics) to C-terminally tag the proteins. ROP2IIHA was then inserted into pENTR/D (Invitrogen) and GRA6IIHA was inserted into pCR8/GW (Invitrogen) by TOPO cloning, and then cloned into pTKO-att by LR recombination (Invitrogen). The vectors were then linearized by digestion with NotI (NEB) and transfected into parasites by electroporation. Electroporation and selection was done as in the previous paragraph, and HA staining was confirmed by IF.
Generation of GRA15 knockout.
A targeting construct (Fig. S5 A) was engineered using a modified pTKO-att vector, pTKO2, and Multisite Gateway Pro 3-Fragment Recombination (Invitrogen). The hypoxanthine-xanthine-guanine ribosyl transferase (HXGPRT) selectable marker was removed from pTKO-att by Cre recombinase (NEB) to form pTKO2 (Fig. S10). 5′ and 3′ flanking regions of GRA15 were cloned from type I and type II genomic DNA. Primers contained att recombination sites (denoted in primer sequence with italics) and amplified 2,083 bp, 100 bp upstream of the GRA15 start codon and 2,071 bp, 34 bp downstream of the GRA15 stop codon (5′ forward, 5′-GGGGACAAGTTTGTACAAAAAAGCAGGCTTAAGGGTCTGAACGTGTGCA-3′; 5′ reverse, 5′-GGGGACAACTTTGTATAGAAAAGTTGGGTGACCCGGCTTAAGTTGGTG-3′; 3′ forward, 5′-GGGGACAACTTTGTATAATAAAGTTGCATGACCAAAAACCGATAA-3′; and 3′ reverse, 5′-GGGGACCACTTTGTACAAGAAAGCTGGGTACAAGTCGGCACATGCTTAGA-3′). These flanking regions were then cloned around the HXGPRT selectable marker flanked by 5′ and 3′ UTRs from DHFR, amplified from pTKO with primers containing att recombination sites (denoted in primer sequence with italics; DHFR::HPT forward, 5′-GGGGACAACTTTTCTATACAAAGTTGCTCAGCACGAAACCTTGCAT-3′; and DHFR::HPT reverse, 5′-GGGGACAACTTTATTATACAAAGTTGTGTGTCACTGTAGCCTGCC-3′). Before transfection, the knockout vector was linearized with the restriction enzyme NotI (NEB). RHΔHXGPRT, PruΔHXGPRT, and PruΔHXGPRT A7 parasites were transfected with the knockout construct by electroporation, as described in the previous section. Stable integrants were selected as in the previous section and cloned by limiting dilution. PCR with a forward primer upstream of the 5′ flanking region (P1, 5′-CATGGATGCTAATCGGCTTT-3′) and a reverse primer within the HXGPRT cassette (P2, 5′-GATCCAGACGTCTTCAATGC-3′; and P3, 5′-GGGGACAACTTTATTATACAAAGTTGTGTGTCACTGTAGCCTGCC-3′) confirmed a disruption in the GRA15 locus. Additionally, PCR was done to confirm the inability to amplify GRA15 (P4, 5′-GATGATGGATCCATAATTCGGTGGCTTGGG-3′; and P5, 5′-GGGGACCACTTTGTACAAGAAAGCTGGGTATCGGCACATGCTTAGAAG-3′; Fig. S5 B).
Microarray.
For human arrays, HFFs were grown in a T25 to confluency. Parasite strains were syringe lysed and washed once with PBS. HFFs were infected with Pru ΔHXGPRT A7, Pru ΔHXGPRT, Pru A7 GRA15KO, Pru GRA15KO, RH 1–1, RH ΔHXGPRT, RH GRA15II (a transgenic RH strain expressing a type II copy of GRA15), RH GRA15KO, CEP HXGPRT− C22, or CEP C22 GRA15II (a transgenic CEP strain expressing a type II copy of GRA15) at varying MOIs. For mouse arrays, WT or p65−/− MEFs were grown in a 12-well plate to confluency. Parasite strains were syringe lysed and washed twice with PBS. MEFs were infected with Pru ΔHXGPRT A7, Pru A7 GRA15KO, RH 1–1, or RH GRA15II at varying MOIs.
Plaque assays were done to assess viability of parasites and infections with similar MOIs were chosen. Some samples were also stimulated with TNF. At least two biological replicates were done for every sample, except RH GRA15KO infection, TNF stimulation, RH ΔHXGPRT preinfection followed by TNF stimulation, and all MEF samples. 18–24 h after infection or 6 h after stimulation, total RNA was isolated using TRIzol reagent (Invitrogen) according to the manufacturer’s protocol and cleaned up using RNeasy Mini or MinElute kit (QIAGEN). RNA was labeled and hybridized to a human or mouse Affymetrix array (Human U133A 2.0 or Mouse 430A 2.0) according to the manufacturer’s protocol. Probe intensities were measured with the Affymetrix GeneChip Scanner 3000 7G and were processed into image analysis (.CEL) files with either GeneChip Operating Software or Expression Console Software (Affymetrix). Intensity values were normalized using the MAS5 algorithm such that the median intensity on the array was 500 using Expression Console software. The MAS5 algorithm gives a signal intensity value for every probe as well as a present, marginal, or absent call, based on mismatch probes. For all probes called present, signal intensity values <50 were increased to a minimum value of 50. For all probes called marginal or absent, the signal intensity value was set to 50. Expression data were clustered using MultiExperiment Viewer (Saeed et al., 2003, 2006). Microarray data has been uploaded to GEO Datasets under accession number GSE 25476.
Gene set enrichment analysis (GSEA) was used to find candidate transcription factors and canonical pathways that are modulated differently between T. gondii infections (Mootha et al., 2003; Subramanian et al., 2005). This program uses a priori–defined sets of genes and determines whether the members of these sets of genes are randomly distributed throughout a ranked list or primarily found at the top or bottom. As GSEA is generally used to generate hypotheses, gene sets enriched with a false discovery rate <0.25 were considered significant. Both transcription factor and canonical pathway gene sets from the Molecular Signatures Database were evaluated for enrichment (c2.cp.v2.5.symbols, c3.tft.v2.5.symbols.gmt; Subramanian et al., 2005). The gsea2 java release was run using all default settings. For analyses on RH GRA15KO, TNF, and RH ΔHXGPRT + TNF, and all MEF infections, for which we only did one array, MOI-matched arrays and the GSEA preranked function were used.
Distant regulatory elements of coregulated genes (DiRE; Gotea and Ovcharenko, 2008) and ingenuity pathway analysis (IPA) were also used. For every gene in a list, DiRE detects regulatory elements throughout the entire gene locus and looks for enrichment of TFBSs. IPA takes a list of genes and overlays them onto a global molecular network developed from information contained in the Ingenuity Pathways Knowledge Base. Networks of these genes are then algorithmically generated based on their connectivity. A network is a graphical representation of the molecular relationships between genes/gene products. Genes or gene products are represented as nodes, and the biological relationship between two nodes is represented as an edge. All edges are supported by at least one reference from the literature, from a textbook, or from canonical information stored in the Ingenuity Pathways Knowledge Base. The two networks with the highest score are indicated.
5′ and 3′ RACE.
Total RNA was isolated from HFFs infected with Pru parasites using TRIzol reagent (Invitrogen) and cleaned up using the RNeasy Mini kit (QIAGEN). RACE-ready cDNA was synthesized using a GeneRacer kit with SuperScript III RT (Invitrogen). Nested PCR was done on the RACE-ready cDNA to determine both 5′ and 3′ transcript ends using gene-specific primers (5′ GRA15reverse, 5′-AGTCCTCCCCGTTTTCGGTCTGTT-3′; 5′ GRA15 nested reverse, 5′-GACTCTGAACGGGGACGGGTAGTC-3′; 3′ GRA15 forward, 5′-CTGTCCACTCAATAGACCCCGTTGT-3′; and 3′ GRA15 nested forward, 5′-AAGATGCCGTGCAAAGCCAACTTC-3′), provided GeneRacer 5′ and 3′ primers, and Phusion enzyme (Finnzymes). PCR products were cloned into the pCR4Blunt-TOPO vector (Invitrogen) and sequenced. Sequences were analyzed with Sequencher software (Gene Codes).
Characterization of GRA15 sequence.
The coding sequence for GRA15 from types I (GT1), II (ME49), and III (VEG) was predicted from ToxoDB genomic sequence using ORF Finder (National Center for Biotechnology Information). GRA15 genomic DNA from additional strains (RH, Pru, CEP) was amplified by PCR and sequenced (forward, 5′-TCCGACTCAGTGCGGGAAA-3′; and reverse, 5′-ATCCAGGTCCCCAAAGG-3′). To check for the presence of a predicted intron in the type I/III ORF, type I cDNA was amplified by PCR and sequenced (forward, 5′-CACGTACACAACCCATCTCG-3′; and reverse, 5′-CGAATTCTCATGGAGTTACCGCTGATT-3′). 5′ and 3′ UTRs were determined by RACE, as described in the previous section. Amino acid alignments were done with ClustalW2 (EMBL-EBI). Similarity to known sequences was queried using BLAST (National Center for Biotechnology Information; Altschul et al., 1990) and PfamA (Finn et al., 2008).
Western blot.
HFFs in a 6-well plate were infected with parasites (MOI = 5) for 24 h. Infected cells were washed with ice-cold PBS, lysed by addition of lysis buffer, boiled for 5 min, and subjected to 10% SDS-PAGE. Proteins were transferred to a polyvinylidene difluoride membrane, which was blocked in PBS/0.1% Tween-20/5% nonfat dry milk and incubated with primary and secondary antibodies. The blot was incubated with a luminal-based substrate (Immun-Star WesternC; Bio-Rad Laboratories) and chemiluminescence was detected using a charge-coupled device camera (Chemidoc XRS; Bio-Rad Laboratories). The bands were visualized using Quantity One 1-D analysis software and analyzed using ImageJ (National Institutes of Health).
GRA15 expression in HeLa cells.
A type II copy of GRA15 or a type II copy of 55.m04955 was inserted into pIC242 (gift from I. Cheeseman, Whitehead Institute for Biomedical Research and Massachusetts Institute of Technology, Cambridge, MA), a Moloney Murine Leukemia Virus retroviral vector containing an N-terminal GFP protein fusion, by restriction/ligation. Amino acids 51–551 of GRA15II were included, as amino acids 1–50 were predicted to be a signal sequence by the signal peptide cleavage prediction server SignalP (Nielsen and Krogh, 1998; Bendtsen et al., 2004). These insertions replaced the original gene insert, Mob1A. Expression of GFP fusion proteins was promoted by the endogenous retroviral long terminal repeats. HeLa cells were then transiently transfected with expression vectors by lipofection using Fugene 6 Plus Transfection Reagent (Roche). Confluent cell cultures were split 1:10 into a 24-well plate (BD) containing glass coverslips. The cells were allowed to incubate at 37°C and 5% CO2 for 4 h. After incubation, the medium was replaced with 1 ml of fresh supplemented DME, and liposomes were added dropwise to the cells. Liposomes were generated according to manufacturer protocol. In brief, 3 µl Fugene reagent was mixed into 20 µl of unsupplemented DME and allowed to stand at room temperature for 5 min. Next, 0.5 µg of appropriate plasmid DNA for each transfection was added, mixed, and incorporated into liposomes for 20 min before addition to cells. Cells were left in contact with liposome for 24 h until the cells were fixed and stained for NF-κB p65. This experiment was performed two times.
In vitro cytokine ELISA.
BALB/c BMDMs were seeded (105 per well) in 96-well plates and left to adhere overnight at 37°C in 5% CO2. Cells were infected with freshly lysed T. gondii tachyzoites at MOI = 20, 10, and 5, and supernatants (200 µl) were collected 24–48 h after infection and stored at −20°C if necessary. IL-12p40 levels were determined, for the cells infected with equal numbers of viable parasites as determined by plaque assay, using a commercially available ELISA kit (OptEIA Mouse IL-12 (p40) ELISA Set; BD) according to the manufacturer’s instructions.
Infection of mice.
Female BALB/c or C57BL/6 mice that were 6–10 wk old (The Jackson Laboratory) were used in all experiments. For i.p. infection, tachyzoites were grown in vitro and extracted from host cells by passage through a 27-gauge needle, washed three times in PBS, and quantified with a hemocytometer. Parasites were diluted in PBS, and mice were inoculated i.p. with tachyzoites of each strain (in 300 µl) using a 28-gauge needle. To image mice infected with a parasite strain that expressed the enzyme luciferase, mice were injected i.p. with 3 mg firefly d-luciferin dissolved in PBS, anesthetized with isoflurane, and imaged with an IVIS Spectrum-bioluminescent and fluorescent imaging system (Xenogen Corporation). Images were processed and analyzed with Living Image software. The MIT Committee on Animal Care approved all protocols. All mice were maintained in specific pathogen-free conditions, in accordance with institutional and federal regulations.
i.p. wash and in vivo cytokine ELISA.
1 or 2 d after i.p. infection, mice were sacrificed and the i.p. cavity was washed with 5 ml PBS. The i.p. wash was spun at 450 g for 5 min to pellet cells. Supernatant was collected and stored at −80°C if necessary. IFN-γ, IL-12p40, and IL-12p70 levels were determined using commercially available ELISA kits (ELISA Ready-SET-Go!; eBioscience) according to the manufacturer’s instructions.
Plaque assay.
For all assays comparing the effect of T. gondii on the host cell, cells were infected with different MOIs and a plaque assay was done to determine the viability of each strain. The infections with the closest MOIs were then used. For the plaque assay, 100 parasites per well were added to confluent HFFs in a 24-well plate and were incubated for 5–7 d at 37°C. The number of plaques was counted using a microscope. Plaque assays were also performed to assess the viability of parasites used to infect mice. To assay in vitro parasite growth, plaque size was measured using NIS-Elements software (Nikon) and a digital camera (CoolSNAP EZ; Roper Scientific) connected to an inverted fluorescence microscope (eclipse Ti-S; Nikon).
Online supplemental material.
Fig. S1 shows that a type II locus on chromosome X strain-specifically induces expression of NF-κB–regulated genes. Fig. S2 shows mapping the strain-specific activation of NF-κB. Fig. S3 shows that type III strains complemented with GRA6 or ROP2 do not activate NF-κB. Fig. S4 shows time course of NF-κB activation by GRA15II. Fig. S5 shows generation and confirmation of GRA15KO. Fig. S6 shows GRA15 activation of NF-κB family subunits. Fig. S7 shows GRA15 amino acid alignment. Fig. S8 shows that GRA15 affects in vitro parasite growth and inhibits in vivo parasite growth in a type I background. Fig. S9 shows that ROP16 can affect the expression of GRA15-regulated genes. Fig. S10 shows pTKO, pTKO-att, and pTKO2 vectors. Supplemental data 1 shows GSEA gene sets. Supplemental data 2 shows complete heat map gene lists.
Acknowledgments
We would like to thank Gus Zeiner for the pTKO plasmid, Hidde Ploegh for MyD88−/−/TRIF−/− mice, Michael Karin for WT and IKKβ−/− MEFs, Anthony Sinai for WT and NF-κBp65−/− MEFs, and Katherine Fitzgerald for TRAF6−/− MEFs. We would also like to thank the MIT BioMicro center for technical assistance and members of the Saeij laboratory for useful comments.
J. Saeij was supported by a Scientist Development Grant from the American Heart Association (0835099N), by a Massachusetts Life Sciences Center New Investigator Award, by the Singapore-MIT Alliance for Research and Technology (SMART), and by National Institutes of Health (AI080621). E. Rosowski and D. Lu were supported by a Pre-Doctoral Grant in the Biological Sciences (5-T32-GM007287-33). E. Rosowski was also supported by the Cleo and Paul Schimmel Fund. K. Jensen was supported by a postdoctoral fellowship from the Cancer Research Institute. L. Rodda was supported by the MIT UROP office and the John Reed Fund.
The authors have no competing financial interests.
References
- BMM
BM-derived macrophage
- DiRE
distant regulatory elements of coregulated genes
- GSEA
gene set enrichment analysis
- HFF
human foreskin fibroblast
- IF
immunofluorescence
- IKK
IκB kinase
- MEF
mouse embryonic fibroblast
- PV
parasitophorous vacuole
- PVM
PV membrane
- RACE
rapid amplification of cDNA ends
- TFBS
transcription factor binding site
Author notes
E.E. Rosowski and D. Lu contributed equally to this paper.