To allow chromosome segregation, topoisomerase II (topo II) must resolve sister chromatid intertwines (SCI) formed during deoxynucleic acid (DNA) replication. How this process extends to the full genome is not well understood. In budding yeast, the unique structure of the ribosomal DNA (rDNA) array is thought to cause late SCI resolution of this genomic region during anaphase. In this paper, we show that chromosome length, and not the presence of rDNA repeats, is the critical feature determining the time of topo II–dependent segregation. Segregation of chromosomes lacking rDNA also requires the function of topo II in anaphase, and increasing chromosome length aggravates missegregation in topo II mutant cells. Furthermore, anaphase Stu2-dependent microtubule dynamics are critical for separation of long chromosomes. Finally, defects caused by topo II or Stu2 impairment depend on attachment of telomeres to the nuclear envelope. We propose that topological constraints imposed by chromosome length and perinuclear attachment determine the amount of SCI that topo II and dynamic microtubules resolve during anaphase.
Introduction
Genetic information is maintained during eukaryotic cell proliferation through replication of chromosomes in S phase, and their subsequent segregation to daughter cells during mitosis. Partitioning of these replicated chromosomes requires dissolution of cohesin linkages at the metaphase-to-anaphase transition (Nasmyth, 2002) as well as resolution of sister chromatid intertwines (SCI). Intertwines arise during replication termination events (DiNardo et al., 1984) and by rotation of the replication fork during DNA strand elongation, which converts superhelical stress ahead of the fork into SCI behind it (Postow et al., 2001). SCI are resolved through the action of type II topoisomerase (topo II), which introduces double-strand DNA breaks to achieve the passage of one double helix through another (Wang, 2002). Consequently, topo II–type enzymes are essential for chromosome segregation from bacteria to human cells (Holm et al., 1985; Uemura et al., 1987; Kato et al., 1990; Ishida et al., 1994).
Although topo II is active throughout the whole eukaryotic cell cycle, the time of complete SCI resolution is unclear. In the budding yeast Saccharomyces cerevisiae, analysis of the topological state of circular plasmids shows that small (<15 kb) replicated dimers are fully resolved into decatenated monomers before anaphase (Koshland and Hartwell, 1987; Baxter et al., 2011), whereas larger 60-kb minichromosomes are still present as catenated dimers in metaphase (Farcas et al., 2011; Charbin et al., 2014). At which point during the cell cycle is SCI resolution of endogenous yeast chromosomes completed remains a key unresolved issue. This was directly addressed in the case of the ribosomal DNA (rDNA) locus. Budding yeast rDNA has been extensively used as a model for chromosome condensation and segregation (Guacci et al., 1994; Bhalla et al., 2002; Lavoie et al., 2004; Machín et al., 2005; D’Ambrosio et al., 2008). The rDNA locus consists of 100–200 tandem repeats of a 9.1-kb element in chromosome XII, making it the longest budding yeast chromosome. Topo II is required during anaphase for segregation of the rDNA, suggesting that SCI are retained in this region until midanaphase (D’Ambrosio et al., 2008). Whether decatenation of the rDNA is representative of the rest of the genome was not known, but it was hypothesized to relate to specific features of this repetitive genomic region. Such features include a high density of replication origins, elevated transcription levels, and/or tightly regulated recombination (Ivessa and Zakian, 2002; Kobayashi and Ganley, 2005; Clemente-Blanco et al., 2009). Importantly, a possible role of chromosome length in rDNA decatenation dynamics remained unexplored.
In animal cells, chromosomes are typically longer than in budding yeast, and inactivation of topo II as late as anaphase leads to chromosome missegregation, indicating that SCI persist until late cell cycle stages (Ishida et al., 1994; Oliveira et al., 2010). Consistently, anaphase DNA bridges that are resolved in a topo II–dependent manner have been observed in human cells (Baumann et al., 2007; Chan et al., 2007; Wang et al., 2008). Thus, whether there are differences in the time of SCI resolution between budding yeast and animal cells and, if so, what the underlying reasons are remain unclear. Understanding how progression through the cell cycle affects SCI resolution in budding yeast, one of the main model systems to study chromosome segregation, is therefore of critical importance.
To directly address the role of chromosome arm length in decatenation, we established a system to study segregation of budding yeast chromosomes of a length comparable to or greater than chromosome XII but lacking rDNA sequences. We took this approach because reducing the number of rDNA repeats would also drastically shorten chromosome XII and possibly affect other cellular processes. Chromosome lengthening was achieved by end-to-end fusions to generate rDNA-free long compound (LC) chromosomes. This method was previously used to generate a stable fusion of chromosomes IV and XII. Aurora B–dependent hypercondensation promotes the segregation of this LC during anaphase and ensures that its length scales appropriately with that of the anaphase spindle (Neurohr et al., 2011). Whether LC hypercondensation is influenced by the presence of the rDNA array was not known. Here, we report that extra-long chromosomes lacking rDNA sequences also undergo anaphase hypercondensation. More importantly, we find that segregation of rDNA-free long chromosomes requires topo II activity throughout anaphase. Microtubule dynamics dependent on the XMAP215/Dis1 family member Stu2 contribute importantly to this process. Finally, we establish that attachment of telomeres to the nuclear envelope (NE) contributes to SCI accumulation in endogenous yeast chromosomes. These processes are not specific to the rDNA locus. Our findings raise the possibility that in both yeast and animal cells, accumulation of SCI is influenced by chromosome length and perinuclear attachment, whereas anaphase microtubule dynamics promote topo II–dependent SCI resolution during anaphase.
Results
Segregation dynamics of rDNA-free long chromosomes
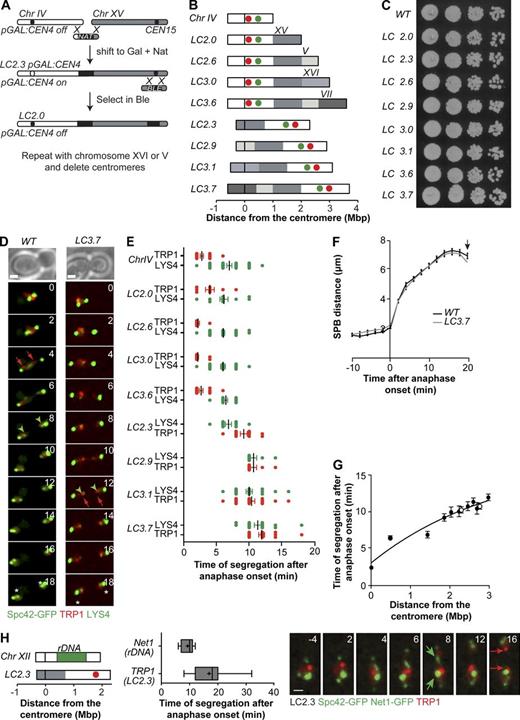
To investigate the relationship between yeast chromosome length and segregation dynamics independent of the presence of the rDNA array, we generated compound chromosomes as long as or longer than chromosome XII but lacking rDNA sequences. Chromosome IV was fused to up to three other chromosomes in haploid cells, by repeated cycles of homologous recombination and simultaneous inactivation of supernumerary centromeres (Figs. 1 A and S1 A; Materials and methods). These genome rearrangements resulted in overall changes in chromosome length but not in genomic content. Correct fusions were confirmed by PCR, pulse-field gel electrophoresis (Fig. S1, B and C), and segregation timing analysis (see two following paragraphs and Fig. 1 E). In this study, long chromosomes are termed LC(N), with N indicating the length in megabases of the long arm (Fig. 1 B, scheme). This approach yielded rDNA-free LC chromosomes with long arms ranging from 2.0 to 3.7 Mb in length (Table S1). As a reference, the long arm of chromosome XII is ∼2–3 Mb in length, assuming 100–200 rDNA copies.
Segregation of extra-long chromosomes lacking rDNA. (A) LC chromosomes were obtained by homologous recombination between subtelomeric regions of chromosomes IV and XV. Centromere 4 was inactivated by transcription from the GAL1,10 promoter. The process was then repeated with additional chromosomes, and supernumerary centromeres were deleted to enable growth in glucose media. Resulting fusions are shown in B. Gal, galactose; Nat, nourseothricin; Ble, bleomycin. (B) Long chromosomes used in this study, named accordingly to their longest chromosome arm. The black bar represents the centromere, and chromosomes within the fusion are color coded; the length of the arms is shown in megabases. The red and green circles indicate the position of TetO (in TRP1) and LacO (in LYS4), respectively. See also Fig. S1 and Table S1. (C) Wild-type and LC cultures were grown to mid–log phase in YPD, and serial dilutions were plated in YPD plates and incubated at 30°C for 2 d. (D) Segregation of wild-type chromosome IV and LC3.7 imaged at 30°C. Spc42-GFP marks the SPBs (bright green dots). The segregation of LYS4 labeled with LacI-GFP (faint green dots) is marked with green arrowheads, and that of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows. Asterisks indicate the estimated time of spindle breakdown, defined when SPB distance decreased by >10% relative to its maximum value. Numbers indicate the time in minutes. Time 0 is anaphase onset (distance between SPBs >3 µm). (E) The time of segregation of TRP1 (red dots) and LYS4 (green dots; scored when sister spots are separated by >2 µm) was determined for the indicated chromosomes. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 25 cells). (F) The distance between SPBs (Spc42-GFP) in wild-type and LC3.7 cells was determined during spindle elongation (n > 30). The arrow marks the mean time of spindle breakdown for both strains. (G) Time of segregation of TRP1 and LYS4 in LC chromosomes as a function of genomic distance to the centromere (n > 25 cells). White circles correspond to LYS4 and TRP1 in the rDNA-containing LC(IV:XII)cen4Δ. Data in F and G are shown as means and SEM. The line is a polynomial fit to the data points. (H) Time of segregation of rDNA and TRP1 in LC2.3 in cells imaged at 30°C. The segregation of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows, and Net1-GFP (nucleolar marker) is marked with green arrows. Numbers indicate the time in minutes. Time 0 is the anaphase onset (distance between SPBs >3 µm). Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). Chr, chromosome; WT, wild type. Bars, 1 µm.
Segregation of extra-long chromosomes lacking rDNA. (A) LC chromosomes were obtained by homologous recombination between subtelomeric regions of chromosomes IV and XV. Centromere 4 was inactivated by transcription from the GAL1,10 promoter. The process was then repeated with additional chromosomes, and supernumerary centromeres were deleted to enable growth in glucose media. Resulting fusions are shown in B. Gal, galactose; Nat, nourseothricin; Ble, bleomycin. (B) Long chromosomes used in this study, named accordingly to their longest chromosome arm. The black bar represents the centromere, and chromosomes within the fusion are color coded; the length of the arms is shown in megabases. The red and green circles indicate the position of TetO (in TRP1) and LacO (in LYS4), respectively. See also Fig. S1 and Table S1. (C) Wild-type and LC cultures were grown to mid–log phase in YPD, and serial dilutions were plated in YPD plates and incubated at 30°C for 2 d. (D) Segregation of wild-type chromosome IV and LC3.7 imaged at 30°C. Spc42-GFP marks the SPBs (bright green dots). The segregation of LYS4 labeled with LacI-GFP (faint green dots) is marked with green arrowheads, and that of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows. Asterisks indicate the estimated time of spindle breakdown, defined when SPB distance decreased by >10% relative to its maximum value. Numbers indicate the time in minutes. Time 0 is anaphase onset (distance between SPBs >3 µm). (E) The time of segregation of TRP1 (red dots) and LYS4 (green dots; scored when sister spots are separated by >2 µm) was determined for the indicated chromosomes. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 25 cells). (F) The distance between SPBs (Spc42-GFP) in wild-type and LC3.7 cells was determined during spindle elongation (n > 30). The arrow marks the mean time of spindle breakdown for both strains. (G) Time of segregation of TRP1 and LYS4 in LC chromosomes as a function of genomic distance to the centromere (n > 25 cells). White circles correspond to LYS4 and TRP1 in the rDNA-containing LC(IV:XII)cen4Δ. Data in F and G are shown as means and SEM. The line is a polynomial fit to the data points. (H) Time of segregation of rDNA and TRP1 in LC2.3 in cells imaged at 30°C. The segregation of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows, and Net1-GFP (nucleolar marker) is marked with green arrows. Numbers indicate the time in minutes. Time 0 is the anaphase onset (distance between SPBs >3 µm). Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). Chr, chromosome; WT, wild type. Bars, 1 µm.
First, we assessed whether these manipulations altered cellular viability. A fusion of chromosomes IV and XII, termed LC(IV:XII), does not impair cell growth or chromosome segregation (Neurohr et al., 2011). Similarly, the presence of rDNA-free LC chromosomes did not affect cell growth in rich media (Fig. 1 C) or worsen growth under conditions of replicative stress (Fig. S1 D). We next tested whether there might be more subtle impairment of mitotic progression as a result of chromosome lengthening. Anaphase dynamics of wild-type chromosome IV and LCs were determined by live-cell imaging. Two different loci in the same chromosome arm were visualized through TetR-mRFP and LacI-GFP reporters, in cells bearing tetracycline and lactose operator arrays. These were inserted 10 kb from CEN4 in wild-type chromosome IV (TRP1 locus) and in the middle of chromosome IV right arm, 470 kb away from TRP1 (LYS4 locus; Fig. 1 B, scheme). Spindle elongation was visualized in the same cells via the spindle pole body (SPB) component Spc42, fused to GFP (Spc42-GFP). Time-lapse imaging showed that spindle elongation dynamics and anaphase duration were indistinguishable between wild-type and LC cells (Fig. 1 F for a comparison between wild type and LC3.7). In addition, the times of TRP1 and LYS4 segregation (scored when sister loci separated by >2 µm) were similar to wild type in LCs in which CEN4 was the active centromere, indicating that segregation of centromere-proximal regions is not affected by changes in chromosome length. Thus, our results confirm and extend previous findings suggesting that chromosome replication and segregation are remarkably robust with respect to changes in chromosome length.
The segregation timing of TRP1 and LYS4 was proportional to their distance from the centromere, as evident from analysis of LCs in which these loci are located at increasing distance from the active centromere (Fig. 1, D and E). Notably, the relationship between centromere distance and segregation time was similar in all LCs, including the rDNA-containing LC(IV:XII)cen12Δ and LC(IV:XII)cen4Δ (Fig. 1 G). Thus, the presence of rDNA sequences did not have a major influence in the segregation time of long chromosome arms. Finally, the nucleolar marker Net1 was fused to GFP in LC2.3 cells to determine the time of rDNA segregation in chromosome XII, relative to that of TRP1 placed in the telomere-proximal region of an rDNA-free lengthened chromosome. Although chromosome segregation was slightly delayed in Net1-GFP cells relative to cells with untagged Net1, separation of the rDNA masses in chromosome XII (extending from 0.45 to ∼1.5 Mb away from the centromere, assuming 120 rDNA copies in our genetic background; Neurohr et al., 2011) always preceded segregation of TRP1 in LC2.3 (2 Mb from the centromere; Fig. 1 H). Thus, chromosome arm length, and not a specific property of the rDNA region, is the major determinant of segregation timing in yeast chromosomes.
Anaphase hypercondensation occurs independently of rDNA sequences
The oversized chromosome arm of LC(IV:XII) undergoes hypercondensation in anaphase, promoting its timely segregation. Hypercondensation requires Aurora B and the condensin complex. Accordingly, LC(IV:XII) cells carrying a conditional condensin mutation show reduced viability at semipermissive temperature and condensin- and Aurora B–dependent shortening of TRP1-LYS4 distances in anaphase (Neurohr et al., 2011).
How general is hypercondensation, and does it depend on rDNA? To address this, the ycg1-2 condensin mutation was introduced into cells with rDNA-free LCs. These cells showed increased temperature sensitivity and chromosome segregation defects compared with condensin mutants of wild-type karyotype, suggesting that hypercondensation is not restricted to rDNA-containing LCs (Fig. S2). To establish whether LCs lacking rDNA also undergo anaphase hypercondensation, the distance between TRP1 and LYS4 was measured in cells carrying either wild-type chromosome IV or LC3.6 in which CEN4 is the active centromere. Distances were measured in metaphase (10 min before spindle elongation) and after segregation of LYS4 (Fig. 2 A, gray area). To test the role of Aurora B/Ipl1 in anaphase compaction, wild type and cells carrying the temperature-sensitive ipl1-321 allele were arrested in G1 with mating pheromone, released into a new cycle at 25°C, and shifted to 35°C after 1 h to inactivate Aurora B during anaphase without affecting its function in chromosome biorientation (Tanaka et al., 2002). In wild-type cells, mean TRP1-LYS4 distances in daughter cells after chromosome segregation were reduced to 85% of their metaphase value (P < 0.01; Fig. 2 B, compare the metaphase and anaphase bud of chromosome IV), as described previously (Neurohr et al., 2011). TRP1-LYS4 metaphase distances were not significantly altered in LC3.6 relative to chromosome IV, but anaphase compaction was further reduced in both mother and bud compartments in LC3.6 cells (Fig. 2 B, anaphase), as reported for LC(IV:XII) (Neurohr et al., 2011). Furthermore, anaphase hypercompaction of the TRP1-LYS4 region in LC3.6 was defective after inactivation of Ipl1 in anaphase (Fig. 2 C). This demonstrates that Aurora B promotes anaphase hypercondensation in extra-long chromosomes independently of the presence of rDNA repeats.
Hypercondensation of an rDNA-free LC depends on Aurora B/Ipl1. (A) Distances between TRP1 and LYS4 measured in time series in wild-type and LC3.6 daughter cells. The shaded area (anaphase, after segregation) corresponds to the time points used to determine distances in B. (B) TRP1-LYS4 distances during metaphase (10 min before anaphase) and anaphase in wild-type and LC3.6 cells. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). ***, P < 0.01. (C) Wild type and cells carrying the temperature-sensitive ipl1-321 allele were arrested in G1 with mating pheromone, released into a new cycle at 25°C, and shifted to 35°C after 1 h, to inactivate Aurora B during anaphase without affecting its function in chromosome biorientation. Only cells that initiated anaphase within the first 20 min of imaging were considered for analysis. Error bars are SEMs, calculated from TRP1-LYS4 distances as in A by error propagation. ***, P < 0.001. Chr, chromosome; WT, wild type.
Hypercondensation of an rDNA-free LC depends on Aurora B/Ipl1. (A) Distances between TRP1 and LYS4 measured in time series in wild-type and LC3.6 daughter cells. The shaded area (anaphase, after segregation) corresponds to the time points used to determine distances in B. (B) TRP1-LYS4 distances during metaphase (10 min before anaphase) and anaphase in wild-type and LC3.6 cells. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). ***, P < 0.01. (C) Wild type and cells carrying the temperature-sensitive ipl1-321 allele were arrested in G1 with mating pheromone, released into a new cycle at 25°C, and shifted to 35°C after 1 h, to inactivate Aurora B during anaphase without affecting its function in chromosome biorientation. Only cells that initiated anaphase within the first 20 min of imaging were considered for analysis. Error bars are SEMs, calculated from TRP1-LYS4 distances as in A by error propagation. ***, P < 0.001. Chr, chromosome; WT, wild type.
Segregation of long chromosome arms requires topo II activity in anaphase
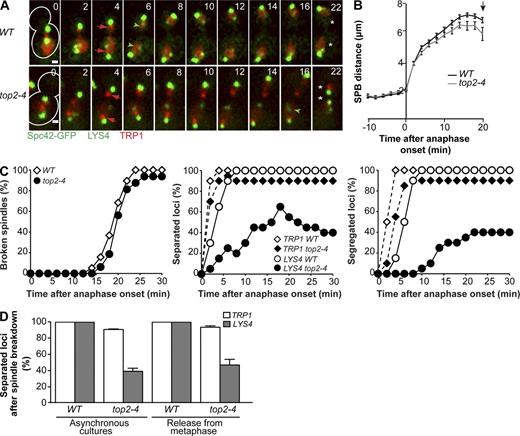
Previous work suggested that persistent catenation during anaphase determined the late segregation of rDNA repeats (D’Ambrosio et al., 2008). Because rDNA-free long chromosome arms segregate during late anaphase, we asked whether their separation also depended on topo II activity during anaphase. We thus studied the dynamics of TRP1 and LYS4 loci in the temperature-sensitive topo II mutant top2-4 by live-cell imaging. Wild-type and top2-4 cultures were grown to log phase at 25°C and shifted to the restrictive temperature (37°C) 15 min before imaging. Analysis was restricted to cells that started anaphase within the first 20 min of imaging and therefore had completed S phase before the temperature shift. Spindle elongation dynamics and the time of spindle breakdown, inferred by the sudden decrease in inter-SPB distance at the end of anaphase (Fig. 3 A, example cells) and by visualization of Tub1-GFP were similar in wild type and top2-4 mutants (Figs. 3, B and C; and S3). In wild-type cells, separation of TRP1 and LYS4 loci (separation, when two dots could be distinguished) was observed in early anaphase, and their segregation to opposite spindle poles (segregation, scored when the distance between sister loci was >2 µm) followed within 2 min (Fig. 3, A–C, WT). In top2-4 mutants, segregation of centromere-proximal TRP1 occurred with dynamics similar to those of wild-type cells, but separation of mid-arm–associated LYS4 was slower and less efficient in top2-4 cells. LYS4 separation was observed in ≤60% of top2-4 cells in early anaphase, but these loci rejoined in late anaphase in a fraction of the cells. Therefore, only 40% of top2-4 cells achieved proper LYS4 segregation after spindle breakdown (Fig. 3, A–C, top2-4). Analysis of top2-4 cells blocked in metaphase at 25°C by depletion of the anaphase-promoting complex activator Cdc20 and released into anaphase at 37°C by Cdc20 expression driven from the MET3 promoter confirmed the late mitotic function of topo II in chromosome IV segregation. Inactivation of topo II after 3 h of Cdc20 depletion at 25°C was sufficient to induce LYS4 separation defects in 60% of cells (Fig. 3 D, release from metaphase), as observed in the previous analysis (Fig. 3 D, asynchronous cultures). Comparable defects in chromosome IV segregation have been observed after inactivation of topo II during the entire cell cycle (Bhalla et al., 2002), suggesting that Top2 continues decatenating chromosome IV throughout anaphase and that one of its major contributions may occur then.
Topo II is required during anaphase for the segregation of chromosome IV. (A) Wild-type and top2-4 cells were grown to log phase at 25°C and imaged at 37°C. Imaging started 15 min after the temperature shift; only cells that initiated anaphase within the first 20 min of imaging were analyzed. Spc42-GFP marks the SPBs (bright green dots). The segregation of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows, whereas LYS4 loci labeled with LacI-GFP (faint green dots) are marked with green arrowheads. Note defective segregation of LYS4 in the top2-4 cell. Asterisks indicate spindle breakdown, defined when SPB distance decreased by >10% relative to its maximum value. Numbers indicate the time in minutes. Time 0 is anaphase onset. Cell outlines are depicted with white lines. Bars, 1 µm. (B) The distance between SPBs (Spc42-GFP) in wild-type and top2-4 cells treated as in A was determined during spindle elongation (n > 30 cells). The arrow marks the mean time of estimated spindle breakdown for both strains. Means and SEM are shown. See also Fig. S3. (C) Kinetics of spindle breakdown, separation, and segregation of centromeres (TRP1) and chromosome arm regions (LYS4) in cells treated as in A. Spindle breakdown was inferred by the sudden decrease in inter-SPB distance at the end of anaphase. TRP1 and LYS4 loci are scored as separated when two dots were distinguishable; segregation of the TRP1 and LYS4 is scored when sister spots are separated by >2 µm. n = 20 cells per strain. (D) The separation of TRP1 and LYS4 (scored when two sister spots can be visualized after spindle breakdown) determined by live-cell imaging in three independent experiments for cells treated as in A (asynchronous cultures). In a separate experiment, cells arrested in metaphase using pMET::CDC20 were released into anaphase at 37°C (release from metaphase). n > 20 cells each in three independent experiments. Means and SD are shown. WT, wild type.
Topo II is required during anaphase for the segregation of chromosome IV. (A) Wild-type and top2-4 cells were grown to log phase at 25°C and imaged at 37°C. Imaging started 15 min after the temperature shift; only cells that initiated anaphase within the first 20 min of imaging were analyzed. Spc42-GFP marks the SPBs (bright green dots). The segregation of TRP1 (red dots visualized with TetR-mCherry) is marked with red arrows, whereas LYS4 loci labeled with LacI-GFP (faint green dots) are marked with green arrowheads. Note defective segregation of LYS4 in the top2-4 cell. Asterisks indicate spindle breakdown, defined when SPB distance decreased by >10% relative to its maximum value. Numbers indicate the time in minutes. Time 0 is anaphase onset. Cell outlines are depicted with white lines. Bars, 1 µm. (B) The distance between SPBs (Spc42-GFP) in wild-type and top2-4 cells treated as in A was determined during spindle elongation (n > 30 cells). The arrow marks the mean time of estimated spindle breakdown for both strains. Means and SEM are shown. See also Fig. S3. (C) Kinetics of spindle breakdown, separation, and segregation of centromeres (TRP1) and chromosome arm regions (LYS4) in cells treated as in A. Spindle breakdown was inferred by the sudden decrease in inter-SPB distance at the end of anaphase. TRP1 and LYS4 loci are scored as separated when two dots were distinguishable; segregation of the TRP1 and LYS4 is scored when sister spots are separated by >2 µm. n = 20 cells per strain. (D) The separation of TRP1 and LYS4 (scored when two sister spots can be visualized after spindle breakdown) determined by live-cell imaging in three independent experiments for cells treated as in A (asynchronous cultures). In a separate experiment, cells arrested in metaphase using pMET::CDC20 were released into anaphase at 37°C (release from metaphase). n > 20 cells each in three independent experiments. Means and SD are shown. WT, wild type.
To gain a more mechanistic understanding of the link between chromosome length and topo II–dependent segregation dynamics, we studied LC top2-4 cells at a semipermissive temperature. The single top2-4 mutant grew well at 30°C, but growth of LC top2-4 cells was severely compromised at the same temperature (Fig. 4 A), indicating that higher levels of topo II activity are required for the survival of cells bearing longer chromosomes. We thus investigated how anaphase-specific inactivation of topo II affected the separation of sister loci in chromosomes of various lengths. LYS4 separation dynamics were determined in cells from asynchronous cultures after temperature shift from 25 to 30°C, using the same experimental setup described in the previous section to assess anaphase-specific topo II function. Separation and segregation were efficient in wild-type cells, top2-4 mutants with normal chromosomes, and LC2.9 cells in which LYS4 is located 2 Mb away from the active centromere (Fig. 4 B). In contrast, LYS4 separation was impaired in LC2.9 top2-4 under the same conditions. In these cells, transient separation of sister loci in early anaphase was often followed by their reassociation in late anaphase and resulting missegregation, indicating incomplete removal of intertwines (Fig. 4 B). In top2 mutants, the LYS4 separation defect was proportional to its distance from the centromere. Indeed, separation was defective in 60% of LC2.9 top2-4 and 90% of LC3.7 top2-4 mutants, in which the labeled locus is, respectively, placed at 1.8 and 2.6 Mb from to the centromere. In contrast, no defects were observed in LC3.6 top2-4 cells, in which LYS4 is in its normal position 450 kb away from CEN4 (Fig. 4 C). Together, these results show that not only is topo II activity during anaphase critical for chromosome segregation but also that its requirement is proportional to the length of chromosome arms.
Topo II is required during anaphase for the segregation of distal regions of long chromosomes. (A) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD. Serial dilutions plated at the indicated temperature were incubated for 2 d. (B) Kinetics of spindle breakdown (Spc42-GFP) and of separation and segregation of chromosome IV arm (LYS4) in cells of the indicated genotypes. Cultures were grown to log phase at 25°C and imaged at 30°C. Imaging started 15 min after the temperature shift, and only cells that initiated anaphase within the first 20 min of imaging were analyzed. n = 20 cells per strain. (C) Spindle elongation and chromosome segregation in representative LC top2-4 cells shifted from 25 to 30°C, 15 min before imaging start. Sister TRP1 and LYS4 spots segregate before spindle breakdown when located closer to the centromere (red arrows and green arrowheads in LC3.6), whereas they fail to separate when placed in a more distal chromosome region (single red arrow and green arrowhead in LC3.7). Numbers indicate the time in minutes. Note the transient separation of loci in LC3.7 top2-4 (12–14 min) followed by their reassociation until spindle disassembly (16–20 min). Asterisks show the time of spindle breakdown. Cell outlines are depicted with white lines. Bars, 1 µm. The graph (bottom left) shows the fraction of cells with unseparated LYS4 loci after spindle disassembly. Means and SD are shown. n > 20 cells in each of three independent experiments. The scheme (bottom right) represents the different chromosomes as in Fig. 1 B: black bars represent centromeres, and chromosomes within the fusion are color coded; the length of the arms is shown in megabases. WT, wild type.
Topo II is required during anaphase for the segregation of distal regions of long chromosomes. (A) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD. Serial dilutions plated at the indicated temperature were incubated for 2 d. (B) Kinetics of spindle breakdown (Spc42-GFP) and of separation and segregation of chromosome IV arm (LYS4) in cells of the indicated genotypes. Cultures were grown to log phase at 25°C and imaged at 30°C. Imaging started 15 min after the temperature shift, and only cells that initiated anaphase within the first 20 min of imaging were analyzed. n = 20 cells per strain. (C) Spindle elongation and chromosome segregation in representative LC top2-4 cells shifted from 25 to 30°C, 15 min before imaging start. Sister TRP1 and LYS4 spots segregate before spindle breakdown when located closer to the centromere (red arrows and green arrowheads in LC3.6), whereas they fail to separate when placed in a more distal chromosome region (single red arrow and green arrowhead in LC3.7). Numbers indicate the time in minutes. Note the transient separation of loci in LC3.7 top2-4 (12–14 min) followed by their reassociation until spindle disassembly (16–20 min). Asterisks show the time of spindle breakdown. Cell outlines are depicted with white lines. Bars, 1 µm. The graph (bottom left) shows the fraction of cells with unseparated LYS4 loci after spindle disassembly. Means and SD are shown. n > 20 cells in each of three independent experiments. The scheme (bottom right) represents the different chromosomes as in Fig. 1 B: black bars represent centromeres, and chromosomes within the fusion are color coded; the length of the arms is shown in megabases. WT, wild type.
Telomere detachment from the NE alleviates top2-associated defects
Topo II might be required during anaphase for the removal of intertwines in the rDNA as a result of this region’s repetitive structure. However, our results suggest a more general hypothesis that topo II is required during anaphase for the removal of persistent intertwines in any long chromosome arms. We wondered whether in this case, anchoring of yeast telomeres to the NE (Taddei et al., 2010) might interfere with chromosome rotation required for SCI removal. We thus tested whether loss of telomere anchoring might alleviate chromosome segregation defects of top2 mutants.
Two partially redundant pathways mediate tethering of telomeres to the NE during interphase in budding yeast, which depend on the Ku (Yku70/80) protein complex and on Sir4 and Esc1 proteins (Hediger et al., 2002; Taddei et al., 2004). We confirmed that deletion of YKU70 or ESC1 altered the subnuclear localization of the telomere in the long arm of chromosome IV, by determining the distance between mCherry-labeled TEL4R (YDR539W) and the nuclear periphery marked with Nup49-GFP, as reported (Hediger et al., 2002). Cells in S/G2 phases were identified by their bud size. TEL4R was tightly associated with the nuclear periphery in wild-type cells but was displaced away from the NE in yku70Δ and esc1Δ mutants (Fig. 5 A). Thus, ESC1 and YKU70 are important for association of TEL4R with the nuclear periphery.
Chromosome detachment from the NE reduces segregation defects in top2-4 mutants. (A) Localization of mCherry-labeled TEL4R relative to the nuclear periphery (Nup49-GFP) determined in S/G2 cells at 30°C. To quantify TEL4R localization, its position in confocal optical slices was mapped to one of three concentric nuclear zones of equal surface. n = 20 cells (wild type [WT] and yku70Δ) or 14 cells (esc1Δ) imaged during 6–8 min every 2–4 min. Bars, 1 µm. (B) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD at 25°C. Serial dilutions were incubated for 2 (25°C) and 6 d (32°C). (C) Kinetics of spindle breakdown (Spc42-GFP) and of separation and segregation of chromosome IV arm (LYS4) in cells of the indicated genotypes. Cultures were grown at 25°C and shifted to the semirestrictive temperature (33°C) 15 min before imaging. n = 20 cells per strain. (D) Chromosome segregation defects determined by live-cell imaging at 33°C, in TEL4R:lexA cells of the indicated genotype in the presence of LexA or LexA-Yif1. n > 20 cells in three independent experiments. ***, P < 0.001. Means and SD are shown. (E) TRP1-LYS4 distances measured in metaphase cells (10 min before anaphase onset) in the indicated strains. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). ***, P < 0.01.
Chromosome detachment from the NE reduces segregation defects in top2-4 mutants. (A) Localization of mCherry-labeled TEL4R relative to the nuclear periphery (Nup49-GFP) determined in S/G2 cells at 30°C. To quantify TEL4R localization, its position in confocal optical slices was mapped to one of three concentric nuclear zones of equal surface. n = 20 cells (wild type [WT] and yku70Δ) or 14 cells (esc1Δ) imaged during 6–8 min every 2–4 min. Bars, 1 µm. (B) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD at 25°C. Serial dilutions were incubated for 2 (25°C) and 6 d (32°C). (C) Kinetics of spindle breakdown (Spc42-GFP) and of separation and segregation of chromosome IV arm (LYS4) in cells of the indicated genotypes. Cultures were grown at 25°C and shifted to the semirestrictive temperature (33°C) 15 min before imaging. n = 20 cells per strain. (D) Chromosome segregation defects determined by live-cell imaging at 33°C, in TEL4R:lexA cells of the indicated genotype in the presence of LexA or LexA-Yif1. n > 20 cells in three independent experiments. ***, P < 0.001. Means and SD are shown. (E) TRP1-LYS4 distances measured in metaphase cells (10 min before anaphase onset) in the indicated strains. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). ***, P < 0.01.
Deletion of ESC1 and YKU70 slightly improved the growth of top2-4 mutants at 32°C (Fig. 5 B) but not at 37°C (not depicted). We then determined the rate of chromosome missegregation in top2-4 esc1Δ and top2-4 yku70Δ in the first division after shifting to this semipermissive temperature, as described previously. Under these conditions, the LYS4 separation and segregation defects observed in top2-4 mutants were clearly alleviated by deletion of ESC1 or YKU70 (Fig. 5 C). Thus, loss of Esc1 or Yku70 facilitates segregation of chromosome arms specifically when the anaphase function of topo II is compromised.
To assess whether rescue of top2-associated defects by deletion of ESC1 and YKU70 was caused by their roles in telomere perinuclear attachment, we asked whether artificially restoring telomere anchoring in top2 esc1 and top2 yku70 mutants would impose segregation defects to similar levels as in top2 single mutants. Expression of LexA fused to the integral membrane protein Yif1 targets telomeres containing LexA repeats to the NE (Schober et al., 2009). We thus inserted LexA recognition sequences 3 kb away from TEL4R in wild type, top2-4, top2-4 esc1, and top2-4 yku70 and quantified LYS4 segregation in the presence of either LexA or LexA-Yif1. Expression of LexA-Yif1 did not cause missegregation in otherwise wild-type cells nor did it aggravate the defect of top2-4 mutants. However, LexA-Yif1 specifically restored chromosome IV segregation defects in top2 esc1 and top2 yku70 double mutants to levels similar to those of the top2-4 single mutant (Fig. 5 D). Thus, our data discover an interplay between chromosome perinuclear attachment and chromosome segregation defects caused by topo II impairment.
We observed that in metaphase LC chromosomes in which labeled loci were located distal to the active centromere, topo II inactivation caused a shortening of TRP1-LYS4 distances (Fig. 5 E). This is likely not caused by hypercondensation, which is an anaphase phenomenon. Instead, it is consistent with an accumulation of intertwines between these loci because an increase in torsional stress shortens the length of braided DNA molecules (Charvin et al., 2003). Notably, shortening of TRP1-LYS4 distances was abolished in top2 esc1 and top2 yku70 mutants (Fig. 5 E), consistent with a reduction in intertwines after telomere detachment from the NE. We note that shortening of the TRP1-LYS4 region after topo II inactivation was not detected in a wild-type chromosome IV (unpublished data), perhaps caused by reduced catenation levels in regions close to the centromere. Collectively, these results strongly suggest that telomere anchoring mediated by ESC1 and YKU70 favors the accumulation of intertwines between replicated chromosomes.
Dynamic microtubules are required for anaphase progression in cells with extra-long chromosomes
These results support a model in which intertwines between sister chromatids remain after metaphase and therefore must be resolved in anaphase to allow faithful chromosome segregation. What factors contribute to anaphase SCI resolution? To identify mechanisms important for the segregation of long chromosomes, we searched for mutations that affect anaphase processes (such as spindle elongation) and specifically impair the growth of LC cells. Anaphase spindle elongation relies on sliding of interpolar microtubules by the kinesin-5 motors Cin8 and Kip1 and stabilization of the spindle midzone by the microtubule bundler Ase1 (Winey and Bloom, 2012). However, deletion of CIN8, KIP1, or ASE1 did not impair growth of LC3.1 cells to a greater extent than in cells with normal chromosomes. Likewise, no synthetic growth defects were observed in LC3.1 cells lacking the microtubule-associated proteins Bim1, Cik1, and Irc15 or in whi3Δ mutants with reduced cell size (Nash et al., 2001) and spindle length (Fig. S4; Neurohr et al., 2011).
In contrast, partial inactivation of the XMAP215/Dis1-like microtubule polymerase Stu2, which controls microtubule dynamics and anaphase spindle elongation (Kosco et al., 2001; Severin et al., 2001), impaired growth of long chromosome cells more severely than that of cells with normally sized chromosomes. LC3.1 stu2-13 mutants showed slight growth defects at 25°C and strongly reduced viability at 30°C, relative to stu2-13 mutants with a normal karyotype (Fig. 6 A). The anaphase activity of Stu2 is counteracted by that of the depolymerizing kinesin-8 Kip3 (Severin et al., 2001; Pearson et al., 2003). Interestingly, Kip3 overexpression from the strong GAL1,10 promoter also impaired growth of LC3.1 cells but not of cells with normal karyotype (Fig. 6 B).
Stu2 is required for LC chromosome arm segregation. (A and B) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD. Serial dilutions were incubated for 2 d at the indicated temperature to inactivate Stu2 (A) or in galactose media to overexpress Kip3 (B). (C) Spindle elongation dynamics in cells representative of the indicated strains expressing Tub1-GFP and Spc42-GFP. Cultures were grown at 25°C to mid–log phase and shifted to 30°C, and imaging was started within 15 min. Time series of anaphase spindle elongation are shown for cells of the indicated strains (2-min image intervals); the frames of spindle disassembly are marked with asterisks. The graphs show the spindle lengths of 20 individual cells of the indicated strains from 10 min before anaphase onset until the time of spindle disassembly. Bars, 1 µm. (D–F) Segregation dynamics of arm loci (ARM; LYS4 in chromosome IV and LC3.1) and pericentromere loci (CEN; TRP1 in LC3.0) in cells of the indicated strains, treated as in C. Spindle breakdown was inferred by Spc42-GFP tracking. (n > 20 cells analyzed for each strain.) WT, wild type.
Stu2 is required for LC chromosome arm segregation. (A and B) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD. Serial dilutions were incubated for 2 d at the indicated temperature to inactivate Stu2 (A) or in galactose media to overexpress Kip3 (B). (C) Spindle elongation dynamics in cells representative of the indicated strains expressing Tub1-GFP and Spc42-GFP. Cultures were grown at 25°C to mid–log phase and shifted to 30°C, and imaging was started within 15 min. Time series of anaphase spindle elongation are shown for cells of the indicated strains (2-min image intervals); the frames of spindle disassembly are marked with asterisks. The graphs show the spindle lengths of 20 individual cells of the indicated strains from 10 min before anaphase onset until the time of spindle disassembly. Bars, 1 µm. (D–F) Segregation dynamics of arm loci (ARM; LYS4 in chromosome IV and LC3.1) and pericentromere loci (CEN; TRP1 in LC3.0) in cells of the indicated strains, treated as in C. Spindle breakdown was inferred by Spc42-GFP tracking. (n > 20 cells analyzed for each strain.) WT, wild type.
These findings suggest that segregation of long chromosomes is particularly sensitive to defects in Stu2-dependent microtubule dynamics. To characterize the specific defects of stu2 mutants, we imaged anaphase progression in stu2-13 and LC stu2-13 cells grown to mid–log phase at 25°C and shifted to 30°C for 15 min before the start of anaphase imaging. In cells with normal chromosomes, partial Stu2 inactivation caused a mild reduction in the final length of anaphase spindles visualized with Tub1-GFP relative to wild-type cells, consistent with the role of Stu2 in spindle elongation (Fig. 6 C; Severin et al., 2001). In contrast, spindles in LC stu2 double mutant cells frequently stalled in midanaphase at an intermediate length of 4–5 µm for >30 min (Fig. 6 C). Imaging of chromosome segregation in wild-type and stu2 cells showed that centromere-proximal TRP1 segregated timely in both strains (wild type: mean of 2.8 min and SD of 1.2 min, n = 23 cells; stu2-13: mean of 2.7 min and SD of 1.2 min, n = 20), whereas LYS4 segregation was delayed by 5 min in stu2 mutants relative to wild-type cells (Fig. 6 D). Furthermore, LC stu2 mutants displayed severe defects in separation and segregation of chromosome arms: LYS4 separation was observed in the majority of midanaphase cells, but sister loci eventually collapsed into a single dot and failed to segregate in 60% of anaphases (Figs. 6 E and S5 A, representative cells). In contrast, separation of centromere-associated TRP1 was normal in LC stu2 mutants (Fig. 6 F). Defects in spindle and chromosome segregation were not caused by light-induced damage during imaging, as consistent phenotypes were also observed in LC stu2-13 cells synchronized in G1 at 25°C, released into a new cycle at 30°C, and stained with DAPI at time intervals (Fig. S5 B).
Partial reestablishment of microtubule dynamics by deletion of Kip3, which restores spindle elongation in stu2 single mutants (Severin et al., 2001), improved anaphase progression in LC stu2 cells. Although LC3.1 stu2-13 kip3Δ cells grew poorly at 25°C and showed an increased frequency of metaphase-like short spindles (unpublished data), the delay in spindle disassembly and the chromosome separation defects of LC3.1 stu2-13 were rescued in the triple mutant (Figs. 7 A and S5 A). Together, these results indicate that defects in the balance of the microtubule dynamicity factor Stu2 and its counteracting kinesin Kip3 induce anaphase spindle elongation and chromosome separation defects specifically in the presence of extra-long chromosomes.
Anaphase defects in LC stu2-13 are alleviated upon detachment of telomeres from the NE. (A) Dynamics of spindle breakdown (Spc42-GFP) and chromosome arm segregation (LYS4-GFP/TRP1-mCherry) in cells of the indicated strains, treated as in Fig. 6 C. In LC3.1 stu2-13 kip3Δ, the LYS4-associated LacI-GFP signal was not visible in all time points, and TRP1 dots (which segregate together with LYS4 in LC3.1; Fig. 1 E) were tracked instead. n > 20 cells were analyzed for each strain. (B) Spindle elongation dynamics (Tub1-GFP and Spc42-GFP) in LC3.1 stu2-13 cells as in Fig. 6 C but with yku70Δ or esc1Δ. Data from LC3.1 stu2-13 cells from Fig. 6 C are included in A and B for comparison. (C) Spindle elongation and chromosome arm (LYS4) segregation/separation in the indicated strains, as in A.
Anaphase defects in LC stu2-13 are alleviated upon detachment of telomeres from the NE. (A) Dynamics of spindle breakdown (Spc42-GFP) and chromosome arm segregation (LYS4-GFP/TRP1-mCherry) in cells of the indicated strains, treated as in Fig. 6 C. In LC3.1 stu2-13 kip3Δ, the LYS4-associated LacI-GFP signal was not visible in all time points, and TRP1 dots (which segregate together with LYS4 in LC3.1; Fig. 1 E) were tracked instead. n > 20 cells were analyzed for each strain. (B) Spindle elongation dynamics (Tub1-GFP and Spc42-GFP) in LC3.1 stu2-13 cells as in Fig. 6 C but with yku70Δ or esc1Δ. Data from LC3.1 stu2-13 cells from Fig. 6 C are included in A and B for comparison. (C) Spindle elongation and chromosome arm (LYS4) segregation/separation in the indicated strains, as in A.
Deletion of ESC1 or YKU70, which partially restored chromosome segregation in top2-4 mutants, also reduced the anaphase defects of LC stu2 mutant cells. LC3.1 stu2-13 esc1Δ and LC3.1 stu2-13 yku70Δ cells showed a reduction in the anaphase delay at the semipermissive temperature to levels closer to those of stu2-13 mutants with normal chromosomes (Fig. 7, B [for spindles visualized with GFP-Tub1] and C [for spindle disassembly inferred from Spc42-GFP tracking]). Deletion of ESC1 and YKU70 not only reduced the duration of anaphase in LC stu2 but also improved chromosome separation efficiency, reducing the frequency of LYS4 missegregation events from >50% in LC stu2 cells to ∼10% in LC stu2 esc1Δ and LC stu2 yku70Δ (Fig. 7 C). Therefore, LC stu2 defects are not simply caused by the increased mass of the LC chromosome and can be alleviated by removal of YKU70- and ESC1-dependent telomere attachment to the NE. In summary, these results show that detaching chromosomes from the NE, which alleviates the defects of top2 mutants, also reduces the chromosome segregation defects caused by Stu2 malfunction. Thus, the anaphase function of Stu2 is likely involved in the resolution of intertwines retained in long chromosome arms by telomere–NE interactions.
Discussion
Topo II activity during anaphase is required for the segregation of long chromosome arms
In this study, we investigated how sister chromatid separation in budding yeast is influenced by the length and perinuclear attachment of chromosomes. Our results suggest that chromosome length and attachment to the NE promote the retention of SCI during S phase, whereas microtubule dynamics and topo II are specifically required during anaphase for their resolution. Topo II–dependent resolution of the rDNA array in anaphase was previously reported (D’Ambrosio et al., 2008), as was the need for anaphase activity of fission yeast topo II for segregation of rDNA regions (Nakazawa et al., 2011). Whether late decatenation was unique to the rDNA or whether it extended to other chromosome regions remained unknown. Our findings establish that topo II is required in anaphase not only for rDNA segregation but also for the segregation of other S. cerevisiae chromosomes, at least as long as chromosome IV. As the requirement for anaphase topo II activity increases in longer chromosomes, late decatenation of the rDNA might simply be a consequence of its length. Intertwines between non-rDNA–bearing sister chromatids may contribute to cohesin-independent cohesion during anaphase, observed previously in the rDNA region (D’Amours et al., 2004; Sullivan et al., 2004). Our data suggest that this may be a general property of linear chromosomes and may depend on their length.
Impact of chromosome length and perinuclear attachment in SCI levels
Segregation defects caused by topo II inactivation during anaphase were proportional to chromosome length and partially dependent on telomere–NE interactions during interphase. These results provide evidence that length and perinuclear attachment are important sources of SCI in linear chromosomes. These factors could hinder chromosome rotation and prevent supercoils ahead of the replication fork, catenations behind it, or a combination of both from falling off chromosome ends before mitosis. Consistent with this possibility, replication-dependent topological stress increases with chromosome length (Kegel et al., 2011), and topo II inactivation only leads to breakage of chromosomes longer than 400 kb (Spell and Holm, 1994).
Yeast telomeric sequences constrain the topology of DNA molecules (Mirabella and Gartenberg, 1997) and limit the movement of chromosome ends, which are anchored to the NE with intervals of attachment separated by periods of rapid diffusion (Hediger et al., 2002). Consistent with the notion of dynamic anchorage, supercoiling in top1 top2 mutants inhibits genes far from chromosome ends, whereas telomere-proximal genes remain active, presumably because of the release of supercoils off chromosome ends (Joshi et al., 2010). Detachment of telomeres from the nuclear periphery might further increase chromosome mobility to dissipate torsional stress. Alternatively, telomere tethering might specifically prevent SCI (but not supercoils) from sliding off the ends of long chromosomes. In any event, our findings suggest that topological constraints caused by chromosome bulkiness and anchoring at the NE prevent the dissipation of torsional stress accumulated during replication.
Animal cell chromosomes are organized in topological domains with different levels of torsional stress (Kouzine et al., 2013) and associate with the nuclear lamina at discrete sites (Guelen et al., 2008). Whether this topological organization contributes to SCI formation and/or stability in animal cells remains to be elucidated.
Anaphase microtubule dynamics promote SCI resolution
In contrast to linear chromosomes, decatenation of circular plasmids is largely achieved in metaphase-arrested yeast cells (Koshland and Hartwell, 1987; Baxter et al., 2011), although decatenation kinetics seem to be size dependent even in these cases (Farcas et al., 2011; Charbin et al., 2014). Microtubule depolymerization after treatment with nocodazole impairs decatenation of these plasmids in metaphase (Baxter et al., 2011; Farcas et al., 2011; Charbin et al., 2014) but which aspect of microtubule function was required for this process and its role in SCI resolution of endogenous chromosomes remained unclear. Our results indicate that the microtubule dynamicity factor Stu2 is required during anaphase for the separation of replicated loci specifically in long chromosome arms. Defects caused by Stu2 impairment were rescued by detachment of telomeres from the NE during interphase, suggesting an involvement of Stu2 in resolution of anaphase SCI.
Stu2 controls not only microtubule and hence spindle length but also the frequency of microtubule growth/shrinkage transitions (Kosco et al., 2001; Pearson et al., 2003). We do not think spindle length plays a direct role in SCI resolution because diminishing the extent of anaphase pole-to-pole separation by inactivation of microtubule motors such as Cin8 (Straight et al., 1998), destabilization of the spindle midzone (in ase1Δ cells; Schuyler et al., 2003), or reduction of cell and spindle size (whi3Δ; Neurohr et al., 2011) does not impair viability of cells with lengthened chromosomes. Instead, we favor the view that microtubule dynamics regulated by Stu2 promote the resolution of SCI by topo II. We envision that before anaphase, interpolar and/or kinetochore dynamic microtubules promote SCI resolution in chromosome regions near their attachment sites, comprising the totality of small centromeric plasmids and centromere-proximal regions of large chromosomes. In contrast, topo II fails to completely resolve intertwines in the chromosome arm at this stage. Upon anaphase onset, dynamic microtubules spread the appropriate tension through the arms of long chromosomes. We note that impairment of topo II and Stu2 have different consequences for anaphase progression: top2 mutants elongate and subsequently disassemble their spindles with wild-type kinetics, whereas LC stu2 cells undergo a pause in midanaphase, which is largely dependent on telomere constraints mediated by Yku70/Esc1. Although we don’t fully understand the precise role of Stu2 in SCI resolution, we speculate that microtubule dynamics promote cycles of tension and relaxation on the DNA to promote topo II resolution of SCI. This could prevent the formation of aberrant chromatin structures in regions of high SCI density capable of triggering a midanaphase delay.
In conclusion, our results support a model in which chromosome length and telomere attachment to the NE determine the level of SCI accumulated during replication of yeast linear chromosomes. These intertwines are resolved specifically during anaphase by the combined action of topo II and microtubule dynamics (Fig. 8). As anaphase condensation increases after chromosome lengthening (this study; Neurohr et al., 2011), and condensin promotes topo II–dependent decatenation (Baxter et al., 2011; Charbin et al., 2014), hypercondensation could further contribute to SCI resolution in highly catenated chromosome regions. These processes might allow robust decatenation of naturally long chromosomes after an increase in length-dependent torsional stress, as can occur upon translocations or size variation of rDNA (Kobayashi et al., 1998).
Microtubule dynamics and topo II solve SCI during anaphase. Intertwines accumulate between replicated chromatids (red and black lines) partially as a result of their attachment to the NE in interphase (blue square). Topo II and dynamic spindle microtubules (green lines) remove SCI from centromere-proximal regions in metaphase and from chromosome arm regions after cleavage of cohesin (orange) in anaphase, during spindle elongation. cat., catenated.
Microtubule dynamics and topo II solve SCI during anaphase. Intertwines accumulate between replicated chromatids (red and black lines) partially as a result of their attachment to the NE in interphase (blue square). Topo II and dynamic spindle microtubules (green lines) remove SCI from centromere-proximal regions in metaphase and from chromosome arm regions after cleavage of cohesin (orange) in anaphase, during spindle elongation. cat., catenated.
In animal cells, in which the level of mitotic chromosome condensation is higher than in budding yeast, chromosome arms appear morphologically disjoined during prophase; however, linkages between individualized chromosome arms have been identified in metaphase (Paliulis and Nicklas, 2004). Thus, resolution of SCI during anaphase is likely conserved throughout eukaryotes, as further indicated by the presence in human cells of thin chromatin bridges that are resolved in anaphase in a topo II–dependent manner (Baumann et al., 2007; Chan et al., 2007; Wang et al., 2008). It will be of interest to investigate whether microtubule dynamics are involved in the resolution of these catenated bridges in human cells. Finally, our results show that systematic investigation of the genetic interactions of yeast cells carrying lengthened chromosomes is a powerful tool for the characterization of segregation mechanisms of yeast chromosomes and of the much larger animal cell chromosomes.
Materials and methods
Strains
S. cerevisiae strains are derivatives of S288c. Gene deletions and insertions were generated by PCR-based methods. TetO/LacO cells and chromosome fusion protocols were previously described (Neurohr et al., 2011). In brief, haploid cells were transformed with a PCR fragment encoding a selection cassette flanked by sequences with homology to subtelomeric regions. A step-by-step description of the construction of all LCs can be found in Fig. S1, and Table S2 lists the oligonucleotides used in their construction. Standard PCR-based techniques were used to introduce the temperature-sensitive alleles stu2-13 (mutation in STU2 ORF, location unknown; Kosco et al., 2001) and top2-4 (C2459A; Holm et al., 1985) by allele replacement at the endogenous locus in wild-type and LC cells. YIp22 (pMET3-HA3-CDC20, TRP1/MscI) provided by F. Uhlmann (London Research Institute, London, England, UK; Uhlmann et al., 1999) was used in a one-step gene replacement at the CDC20 locus. 8×LexA operators and the URA3 auxotrophic marker were amplified by PCR from pSH18-34 (Invitrogen) and inserted in chromosome IV between nucleotide positions 1,521,063 and 1,521,613. pRS425-based plasmids encoding LexA or LexA fused to full-length Yif1 under the control of the ADH1 promoter were provided by S. Gasser (Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland; Schober et al., 2009). TEL4R strains are derivatives of yPT239-1 provided by E. Fabre (Hôpital Saint Louis, Paris, France; Therizols et al., 2010), containing TetR-mRFP integrated in YGL119W and 112 repeats of the tetracycline operator in YDR539W.
Cell growth and microscopy
Time-lapse analysis of chromosome segregation, except when indicated, was performed on cells synchronized with 10 mg/ml α factor (Sigma-Aldrich) for 2 h, released in fresh YPD (yeast extract, peptone, and dextrose) medium for 1 h at 25°C, and placed in a preequilibrated temperature-controlled microscope chamber 15 min before imaging. For metaphase arrest, pMET3-CDC20 cells grown in minimal synthetic media without methionine were incubated in YPD supplemented with 2 mg/ml methionine for 3 h, and released into −Met media. For live microscopy, cells were plated in minimal synthetic medium on concanavalin A–coated (Sigma-Aldrich) Lab-Tek chambers (Thermo Fisher Scientific), and imaging was performed using a spinning-disk confocal microscope (Revolution XD; Andor Technology) with a Plan Apochromat 100×, 1.45 NA objective equipped with a dual-mode electron-modifying charge-coupled device camera (iXon 897 E; Andor Technology). Time-lapse series of 4-µm stacks spaced 0.5 µm were acquired every 2 min. iQ Live Cell Imaging software (Andor Technology) was used for image acquisition. Images were analyzed on 2D maximum projections and denoised with the Despeckle function in ImageJ 1.46b (National Institutes of Health). Distances between signals were measured between local maxima. Graphs and statistical analysis (t test allowing for unequal variance) were performed with Prism (GraphPad Software) and Excel (Microsoft).
Pulse-field gel electrophoresis
Pulse-field gel electrophoresis was performed essentially as previously described (Herschleb et al., 2007). In brief, 6 × 106 cells were washed in ice-cold 0.05 M EDTA, pH 7.5, resuspended in 30 µl of 0.125-M EDTA, pH 7.5, and kept at 37°C before mixing with 40 µl of 1.2% low melting point agarose in 0.125 M EDTA preequilibrated at 42–50°C, pipetted into a casting mold, and kept at 4°C overnight. Agarose bricks were incubated for ≥12 h in 1 ml of 0.5-M EDTA and 7.5% β-mercaptoethanol at 37°C. The buffer was drained, replaced with NDSK buffer (0.5 M EDTA, 1% [wt/vol] N-laurylsarcosine, and 1 mg/ml proteinase K), incubated overnight at 50°C, washed with Tris/Borate/EDTA, molded into a 0.7% agarose gel, and run with the standard protocol for >4 Mb DNA (to separate long chromosomes) in the CHEF Mapper XA System (Bio-Rad Laboratories).
Online supplemental material
Fig. S1 describes the generation and characterization of extra-long chromosomes. Fig. S2 shows cell viability and chromosome segregation in LC ycg1-2 mutant cells at 30°C. Fig. S3 characterizes spindle dynamics of wild-type and top2-4 cells. Fig. S4 demonstrates lack of genetic interactions between LC and genes affecting spindle function and size. Fig. S5 documents that perturbation of microtubule dynamics impairs anaphase progression in LC cells. Table S1 summarizes the structural organization of extra-long chromosomes, and Table S2 lists oligonucleotides used.
Acknowledgments
We thank Pedro Carvalho, Snezhana Oliferenko, and Steven Gross for critical reading of the manuscript; Francesc Posas, Fèlix Campelo, Jordi Torres, Yves Barral, and Susan Gasser for helpful suggestions; Trinidad Sanmartín for technical support; the Centre for Genomic Regulation Genomics, Bioinformatics, and Advanced Light Microscopy Units; and the Institut de Recerca Biomèdica Advanced Digital Microscopy Facility.
This research is supported by grants from the European Research Council (ERC Starting Grant 260965) and the Spanish Ministry of Science (BFU09-08213) to M. Mendoza.
The authors declare no competing financial interests.
References
Author notes
I. Titos’s present address is Dept. of Neurobiology, Harvard Medical School, Boston, MA 02115.




![Figure 5. Chromosome detachment from the NE reduces segregation defects in top2-4 mutants. (A) Localization of mCherry-labeled TEL4R relative to the nuclear periphery (Nup49-GFP) determined in S/G2 cells at 30°C. To quantify TEL4R localization, its position in confocal optical slices was mapped to one of three concentric nuclear zones of equal surface. n = 20 cells (wild type [WT] and yku70Δ) or 14 cells (esc1Δ) imaged during 6–8 min every 2–4 min. Bars, 1 µm. (B) Cultures of wild-type and mutant strains were grown to mid–log phase in YPD at 25°C. Serial dilutions were incubated for 2 (25°C) and 6 d (32°C). (C) Kinetics of spindle breakdown (Spc42-GFP) and of separation and segregation of chromosome IV arm (LYS4) in cells of the indicated genotypes. Cultures were grown at 25°C and shifted to the semirestrictive temperature (33°C) 15 min before imaging. n = 20 cells per strain. (D) Chromosome segregation defects determined by live-cell imaging at 33°C, in TEL4R:lexA cells of the indicated genotype in the presence of LexA or LexA-Yif1. n > 20 cells in three independent experiments. ***, P < 0.001. Means and SD are shown. (E) TRP1-LYS4 distances measured in metaphase cells (10 min before anaphase onset) in the indicated strains. Boxes include 50% of data points, and whiskers are 90%. Median (lines) and mean (crosses) are shown (n > 20 cells). ***, P < 0.01.](https://cdn.rupress.org/rup/content_public/journal/jcb/206/6/10.1083_jcb.201404039/6/m_jcb_201404039_fig5.jpeg?Expires=1779194424&Signature=OZkNyiVdAbytJeNd~scT3GCPp~BeZNGI7bk~NpGrW1fUG-FEPgkf0AeVzWcAuwrEmi2LW4DXJX2gN04OO4ET~~L9MLGUqdPxcy75HBDP50ntV4e3VoMO8-rf4ZuHFJvHq1M-mfysa6n9KwDt43jedj7oWc05VyLGfL0jjdkyx-9YWKqdW1AyR07rdYPn1J8XpeSvqtLzWBY-LdCBuXhXkIvLIzOEq3PS9htkZzILeSmrA2USvgq~c3PMUL5q9AlaoUn~DblN-gE5SiKFedY-E4W~WicclT7yp8Q4WI4MG1BXZV2TOyMPU4hUw3~SQRj-2trwczjloLJFh1b5PEkMkg__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)





