Glycoproteins misfolded in the endoplasmic reticulum (ER) are subjected to ER-associated glycoprotein degradation (gpERAD) in which Htm1-mediated mannose trimming from the oligosaccharide Man8GlcNAc2 to Man7GlcNAc2 is the rate-limiting step in yeast. In contrast, the roles of the three Htm1 homologues (EDEM1/2/3) in mammalian gpERAD have remained elusive, with a key controversy being whether EDEMs function as mannosidases or as lectins. We therefore conducted transcription activator-like effector nuclease–mediated gene knockout analysis in human cell line and found that all endogenous EDEMs possess mannosidase activity. Mannose trimming from Man8GlcNAc2 to Man7GlcNAc2 is performed mainly by EDEM3 and to a lesser extent by EDEM1. Most surprisingly, the upstream mannose trimming from Man9GlcNAc2 to Man8GlcNAc2 is conducted mainly by EDEM2, which was previously considered to lack enzymatic activity. Based on the presence of two rate-limiting steps in mammalian gpERAD, we propose that mammalian cells double check gpERAD substrates before destruction by evolving EDEM2, a novel-type Htm1 homologue that catalyzes the first mannose trimming step from Man9GlcNAc2.
Introduction
Proteins misfolded in the ER are degraded by the proteasome via a series of events collectively termed ER-associated degradation (Xie and Ng, 2010; Smith et al., 2011; Brodsky, 2012). Among the various pathways used, the best characterized, particularly in yeast, is ER-associated glycoprotein degradation (gpERAD) in which two-step mannose trimming from high-mannose-type oligosaccharides plays crucial roles (Molinari, 2007; Hosokawa et al., 2010a; Kamiya et al., 2012). α1,2-mannosidase Mns1 catalyzes the first step, conversion of Man9GlcNAc2 (M9) to Man8GlcNAc2 isomer B (M8B), and α1,2-mannosidase Htm1 catalyzes the second step, conversion of M8B to oligosaccharides with the α1,6-mannose exposed (Mα1,6E; Fig. 1, C and E; and see Fig. 5 A). These products are then recognized by lectin Yos9 for subsequent disposal (Quan et al., 2008).
Characterization of DT40 and HCT116 cell lines in regard to gpERAD. (A) Schematic structures of yeast Mns1 and Htm1 and their homologues in chickens (g) and humans (h). Sequence identities are shown as percentages. (B) Phylogenic tree calculated by the neighbor-joining method. (C) Schematic structure of Glc3Man9GlcNAc2. (D) Elution profiles of N-glycans prepared from total cellular glycoproteins of WT DT40 and HCT116 cells. This experiment was completed once. (E) Isomer composition of N-glycans obtained from D. nd, not detected. (F) Endogenous mRNA levels encoding chicken and human EDEM1, EDEM2, EDEM3, SEL1L, and ERmanI relative to chicken and human GAPDH mRNA level, respectively, expressed in WT DT40 or HCT116 cells (n = 3).
Characterization of DT40 and HCT116 cell lines in regard to gpERAD. (A) Schematic structures of yeast Mns1 and Htm1 and their homologues in chickens (g) and humans (h). Sequence identities are shown as percentages. (B) Phylogenic tree calculated by the neighbor-joining method. (C) Schematic structure of Glc3Man9GlcNAc2. (D) Elution profiles of N-glycans prepared from total cellular glycoproteins of WT DT40 and HCT116 cells. This experiment was completed once. (E) Isomer composition of N-glycans obtained from D. nd, not detected. (F) Endogenous mRNA levels encoding chicken and human EDEM1, EDEM2, EDEM3, SEL1L, and ERmanI relative to chicken and human GAPDH mRNA level, respectively, expressed in WT DT40 or HCT116 cells (n = 3).
The mammalian ER expresses ER mannosidase I (ERmanI) as the sole homologue of Mns1, but expresses multiple homologues of Htm1, namely, EDEM1, EDEM2, and EDEM3 (Fig. 1, A and B). The exact roles of all these proteins in mammalian gpERAD have remained elusive. Overexpression and biochemical experiments indicated that ERmanI converted M9 to M8B (Gonzalez et al., 1999; Hosokawa et al., 2003). Overexpression of EDEM1 or EDEM3 but not EDEM2 promoted mannose trimming at various steps, including the second step (Hosokawa et al., 2003, 2010b; Mast et al., 2005; Hirao et al., 2006; Olivari et al., 2006). These results pointed to ERmanI as the first-step enzyme and to EDEM1 and EDEM3 as the second-step enzymes, and suggested that EDEM2 lacks α-mannosidase activity. However, this was puzzling to us because it had originally been proposed that EDEM1 has no α1,2-mannosidase activity (Hosokawa et al., 2001) and because it was also suggested that ERmanI is involved in the formation of Man7-5GlcNAc2 with Mα1,6E, based on the results of overexpression (Hosokawa et al., 2003), knockdown (Avezov et al., 2008), and biochemistry (Aikawa et al., 2012). Moreover, the finding that EDEM1 recognized not only misfolded glycoproteins but also misfolded nonglycoproteins and delivered them to the ER membrane for destruction by binding to the carbohydrate moiety of its downstream component SEL1L (Cormier et al., 2009) generated controversy as to whether EDEMs function as α1,2-mannosidases for mannose trimming or as lectins for substrate delivery (Tamura et al., 2010). We have therefore conducted gene knockout (KO) analyses in chicken and human cell lines to resolve this controversy and to determine which proteins catalyze the two key steps of mannose trimming in mammalian gpERAD.
Results and discussion
We started by determining the N-glycan profiles of total cellular glycoproteins prepared from wild-type (WT) DT40 cells derived from chicken B lymphocytes and HCT116 cells derived from human colonic carcinoma, both of which express Mns1 and Htm1 homologue mRNAs at comparable levels (Fig. 1 F). We focused on high-mannose-type oligosaccharides with the elution positions shown in Fig. 1 D. For comparison, Fig. 1 E shows the amount of each high-mannose-type oligosaccharide relative to the total amount of high-mannose-type oligosaccharides. M9 and M8B levels in the two cell types were comparable, consistent with the results obtained with HepG2 cells (Hosokawa et al., 2010b). In connection with this, our literature search unraveled an interesting difference between yeast and mammalian gpERAD. Namely, total N-glycans obtained after 10–20-min pulse labeling of yeast cells consisted almost exclusively of M8 (Clerc et al., 2009), indicating that conversion of M9 to M8B by Mns1 is highly efficient and occurs regardless of protein folding/misfolding status. Therefore, conversion of M8B to Mα1,6E by Htm1 is the initiating and rate-limiting step in yeast gpERAD (Gauss et al., 2011). In contrast, total N-glycans obtained after 30-min pulse labeling of HEK293 cells consisted predominantly of M9 (Hirao et al., 2006), indicating that conversion of M9 to M8B is inefficient in human cells.
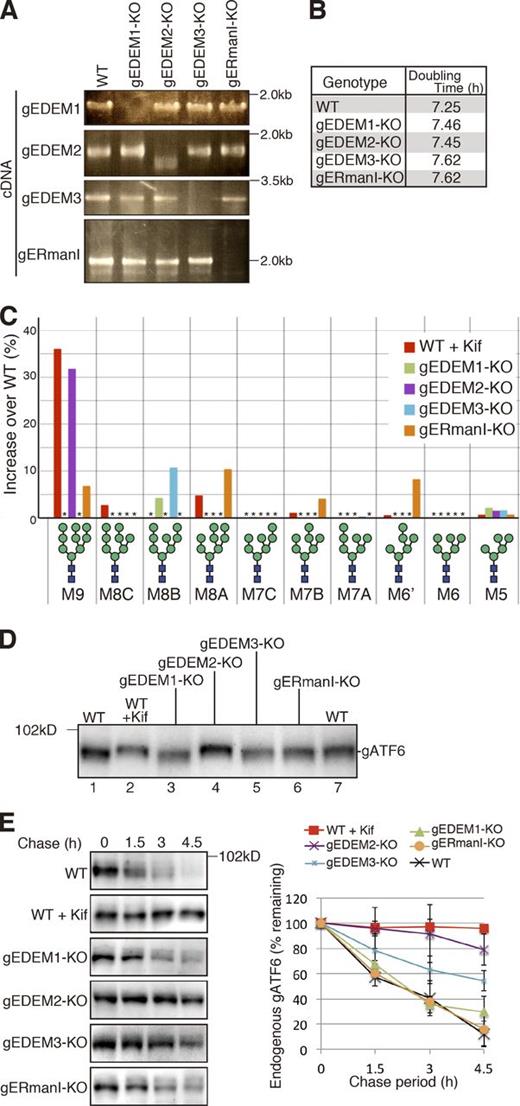
We knocked out MnsI and Htm1 homologues separately in DT40 cells using homologous recombination, which is exceptionally efficient in this cell line (Fig. S1). RT-PCR showed no expression of gEDEM1, gEDEM3, or gERmanI mRNA in the respective KO cells (Fig. 2 A). Although gEDEM2-KO cells expressed some short gEDEM2 mRNAs, these mRNAs had internal deletions and therefore failed to produce any proteins when translated in vitro (Fig. S1 K). All of these KO cells grew nearly as fast as WT cells (Fig. 2 B). We determined their N-glycan profiles in the same way as for WT cells (Fig. S2, A and B) and identified the oligosaccharides whose contents in KO cells exceeded those in untreated WT cells. The results are shown as the increase over WT (percentage of untreated WT cells subtracted from percentage of each type of KO cell) in Fig. 2 C. When WT cells were treated with kifunensine, an inhibitor of ERmanI (Tremblay and Herscovics, 1999), M9 became predominant, as expected, but surprisingly M9 increased only slightly in gERmanI-KO cells (Fig. 2 C). This finding indicates that the Mns1 homologue plays only a minor role in the conversion of M9 to M8B in DT40 cells, but is consistent with the finding that purified recombinant human ERmanI exhibited much weaker α1,2-mannosidase activity toward M9 attached to a native protein than to M9 attached to pyridylamine (Aikawa et al., 2012) and may be correlated with the unexpected localization of endogenous ERmanI at the Golgi apparatus (Pan et al., 2011). The fact that the levels of M8A, M7B, and M6′ also increased in gERmanI-KO cells (Fig. 2 C) indicates that gERmanI appears to trim the outermost mannose of the B branch randomly rather than to convert M9 to M8B specifically.
Effect of gene disruption on chicken gpERAD. (A) RT-PCR to amplify cDNA corresponding to the four α1,2-mannosidase mRNA expressed in DT40 cells of various genotypes. (B) Doubling time of WT and four α1,2-mannosidase KO cells (n = 3). (C) Display of oligosaccharides whose contents in kifunensine (Kif; 10 µg/ml, 6 h)-treated WT DT40 cells and four α1,2-mannosidase KO cells exceeded those in kifunensine-untreated WT cells with an increase over WT (%), as determined from isomer composition of their N-glycans (Fig. S2, A and B), which was completed once. Asterisks denote oligosaccharides whose contents did not exceed those in kifunensine-untreated WT cells. (D) Immunoblotting of cell lysates prepared from kifunensine-untreated or -treated WT DT40 cells and from the four α1,2-mannosidase KO cells using anti-chicken ATF6 (gATF6) antibody. (E) Cycloheximide chase to determine the degradation rate of endogenous gATF6 in kifunensine-untreated or -treated WT DT40 cells and in the four α1,2-mannosidase KO cells (n = 3).
Effect of gene disruption on chicken gpERAD. (A) RT-PCR to amplify cDNA corresponding to the four α1,2-mannosidase mRNA expressed in DT40 cells of various genotypes. (B) Doubling time of WT and four α1,2-mannosidase KO cells (n = 3). (C) Display of oligosaccharides whose contents in kifunensine (Kif; 10 µg/ml, 6 h)-treated WT DT40 cells and four α1,2-mannosidase KO cells exceeded those in kifunensine-untreated WT cells with an increase over WT (%), as determined from isomer composition of their N-glycans (Fig. S2, A and B), which was completed once. Asterisks denote oligosaccharides whose contents did not exceed those in kifunensine-untreated WT cells. (D) Immunoblotting of cell lysates prepared from kifunensine-untreated or -treated WT DT40 cells and from the four α1,2-mannosidase KO cells using anti-chicken ATF6 (gATF6) antibody. (E) Cycloheximide chase to determine the degradation rate of endogenous gATF6 in kifunensine-untreated or -treated WT DT40 cells and in the four α1,2-mannosidase KO cells (n = 3).
Contrary to our strong expectations from previous results (Mast et al., 2005), we were surprised to find that conversion of M9 to M8B was blocked as effectively in gEDEM2-KO cells as in WT cells treated with kifunensine (Fig. 2 C), indicating that the first-step mannose trimming in DT40 cells is mainly caused by gEDEM2 and that kifunensine inhibits both gERmanI and gEDEM2. In contrast, the level of M8B increased in gEDEM1-KO and gEDEM3-KO cells (Fig. 2 C), indicating that EDEM1 and EDEM3 are the second-step enzymes. These differences in N-glycan profiles were well reflected by the mobility in SDS-PAGE of ATF6, a type II transmembrane glycoprotein in the ER that functions as an unfolded protein response transducer (Haze et al., 1999): endogenous ATF6 migrated more slowly in kifunensine-treated WT and gEDEM2-KO cells than in other cell types (Fig. 2 D). We recently showed that ATF6 undergoes gpERAD in DT40 cells (Horimoto et al., 2013). Because its degradation by the proteasome is completely blocked by kifunensine treatment, it is a highly suitable substrate to determine the effect on gpERAD. Cycloheximide chase clearly showed that degradation of endogenous ATF6 occurred normally in gERmanI-KO cells, whereas it was most effectively blocked in kifunensine-treated WT and gEDEM2-KO cells (Fig. 2 E), as expected from the N-glycan profiles.
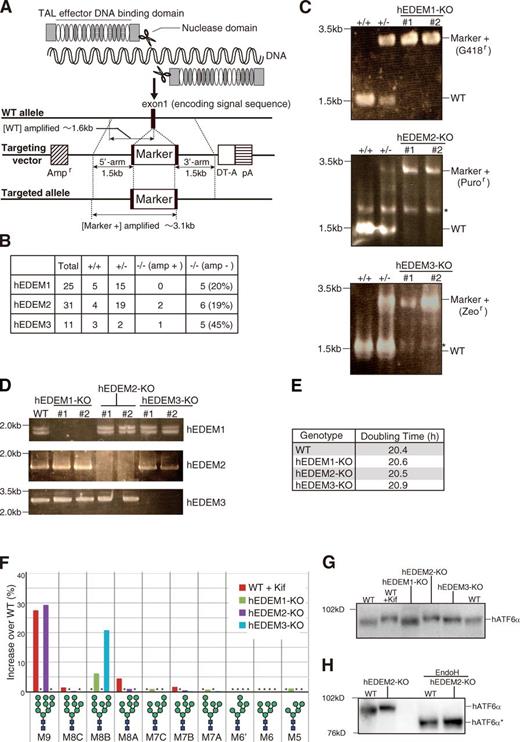
We then knocked out three EDEMs in the human HCT116 diploid cell line (Ochiai et al., 2014) using the transcription activator-like effector nuclease (TALEN) method (Joung and Sander, 2013). Specifically, we used the recently developed Platinum TALEN technology, which appears to have higher activity than previously reported methods (Sakuma et al., 2013). Single introduction of Platinum TALEN designed to cleave within exon 1 encoding the signal sequence of hEDEM1, hEDEM2, or hEDEM3, together with the respective targeting vector containing positive and negative selection markers, produced homo-KOs with efficiency of 20%, 19%, or 45%, respectively, and the ampicillin-resistance gene for Escherichia coli selection and the diphtheria toxin-A fragment gene were not incorporated into the genome when correctly targeted (Fig. 3, A and B). Genomic PCR (Fig. 3 C) and Southern blot hybridization (Fig. S3, A–D) showed incorporation of the drug-resistance gene at the expected locus in each case (four probes we tested did not work for hEDEM1 Southern blot hybridization), and RT-PCR revealed no expression of hEDEM1, hEDEM2, or hEDEM3 mRNA in the respective KO cells (Fig. 3 D). These KO cells grew as fast as WT cells (Fig. 3 E). Immunoblotting showed that ER stress marker proteins BiP, ATF4, and XBP1(S) were not induced in these KO cells (Fig. S3 E). The N-glycan profiles (Fig. S3, F and G) revealed that conversion of M9 to M8B was blocked in hEDEM2-KO cells as effectively as in kifunensine-treated WT HCT116 cells and that the level of M8B increased in hEDEM1-KO and hEDEM3-KO cells (Fig. 3 F). Thus, all endogenous hEDEMs function as α1,2-mannosidases. Endogenous human ATF6α migrated more slowly in kifunensine-treated WT and hEDEM2-KO cells than in untreated WT cells (Fig. 3 G), and this difference was lost after treatment with endoglycosidase H, as expected (Fig. 3 H). This difference in migration was also observed in different KO strains (Fig. S3 H). ATF6α in hEDEM3-KO cells appeared to migrate with a speed between that in untreated WT and hEDEM2-KO cells (Fig. 3 G), reflecting the marked increase in M8B level (Fig. 3 F). Degradation of endogenous ATF6α was blocked in WT HCT116 cells treated with MG132 or kifunensine, as expected, and was most effectively blocked in hEDEM2-KO cells (Fig. 4 A).
Effect of gene disruption on human gpERAD. (A) Strategy for TALEN-mediated gene disruption. Cleavage of the exon 1 encoding the signal sequence of human EDEM1, EDEM2, or EDEM3 with the designed TALEN facilitates subsequent homologous recombination with the respective targeting vector containing both positive and negative selection markers. (B) Results of screening. The presence (amp +) or absence (amp −) of the ampicillin-resistance gene in targeted alleles was checked by genomic PCR. (C) Genomic PCR to confirm homologous recombination in HCT116 cells of various genotypes. Asterisks denote nonspecific bands. (D) RT-PCR to amplify cDNA corresponding to hEDEM1, hEDEM2, or hEDEM3 mRNA expressed in HCT116 cells of various genotypes. The full-length cDNA and cDNA lacking the exon 6 were amplified as hEDEM1. (E) Doubling time of WT and three hEDEM-KO cells (n = 4). (F) Display, similarly to Fig. 2 C, of oligosaccharides whose contents in kifunensine (Kif; 10 µg/ml, 12 h)-treated WT HCT116 and three hEDEM-KO cells exceeded those in kifunensine-untreated WT cells with an increase over WT (%), as determined from isomer composition of their N-glycans (Fig. S3, F and G), which was completed once. Asterisks denote oligosaccharides whose contents did not exceed those in kifunensine-untreated WT cells. (G) Immunoblotting of cell lysates prepared from kifunensine-untreated or -treated WT HCT116 cells and from three hEDEM-KO cells using anti-human ATF6α (hATF6α) antibody. (H) Immunoblotting of cell lysates prepared from WT and hEDEM2-KO HCT116 cells with or without endoglycosidase H treatment. hATF6α* denotes the nonglycosylated hATF6α.
Effect of gene disruption on human gpERAD. (A) Strategy for TALEN-mediated gene disruption. Cleavage of the exon 1 encoding the signal sequence of human EDEM1, EDEM2, or EDEM3 with the designed TALEN facilitates subsequent homologous recombination with the respective targeting vector containing both positive and negative selection markers. (B) Results of screening. The presence (amp +) or absence (amp −) of the ampicillin-resistance gene in targeted alleles was checked by genomic PCR. (C) Genomic PCR to confirm homologous recombination in HCT116 cells of various genotypes. Asterisks denote nonspecific bands. (D) RT-PCR to amplify cDNA corresponding to hEDEM1, hEDEM2, or hEDEM3 mRNA expressed in HCT116 cells of various genotypes. The full-length cDNA and cDNA lacking the exon 6 were amplified as hEDEM1. (E) Doubling time of WT and three hEDEM-KO cells (n = 4). (F) Display, similarly to Fig. 2 C, of oligosaccharides whose contents in kifunensine (Kif; 10 µg/ml, 12 h)-treated WT HCT116 and three hEDEM-KO cells exceeded those in kifunensine-untreated WT cells with an increase over WT (%), as determined from isomer composition of their N-glycans (Fig. S3, F and G), which was completed once. Asterisks denote oligosaccharides whose contents did not exceed those in kifunensine-untreated WT cells. (G) Immunoblotting of cell lysates prepared from kifunensine-untreated or -treated WT HCT116 cells and from three hEDEM-KO cells using anti-human ATF6α (hATF6α) antibody. (H) Immunoblotting of cell lysates prepared from WT and hEDEM2-KO HCT116 cells with or without endoglycosidase H treatment. hATF6α* denotes the nonglycosylated hATF6α.
Requirement for α1,2-mannosidase activity of hEDEM2 in human gpERAD. (A) Pulse chase to determine the degradation rate of endogenous hATF6α in WT HCT116 cells with or without kifunensine or MG132 treatment and in three hEDEM-KO cells (n = 3). (B) Alignment of amino acid sequences around the EF hand motif, which is essential for the α1,2-mannosidase activity of Mns1, with those of Htm1 and their homologues in chicken and human. Identical amino acids are highlighted. (C) Schematic structure of hATF6α and hATF6α(C)-BACE(TM)-TAP. Potential N-glycosylation sites are shown schematically. (D) Immunoblotting of cell lysates prepared from WT and hEDEM2-KO HCT116 cells in which Myc-hATF6α (top) or hATF6α(C)-BACE(TM)-TAP (bottom) was expressed by transfection together with hEDEM2-Flag or hEDEM2-E117Q-Flag, using anti–c-myc and anti-Flag antibodies. (E) Immunoblottig of cell lysates prepared from WT HCT116, hEDEM2-KO, and hEDEM2-KO cells stably expressing hEDEM2-Flag or hEDEM2-E117Q-Flag using anti-human ATF6α and anti-Flag antibodies. (F) Pulse chase to determine the degradation rate of endogenous hATF6α in HCT116 hEDEM2-KO and hEDEM2-KO cells stably expressing hEDEM2-Flag or hEDEM2-E117Q-Flag (n = 3).
Requirement for α1,2-mannosidase activity of hEDEM2 in human gpERAD. (A) Pulse chase to determine the degradation rate of endogenous hATF6α in WT HCT116 cells with or without kifunensine or MG132 treatment and in three hEDEM-KO cells (n = 3). (B) Alignment of amino acid sequences around the EF hand motif, which is essential for the α1,2-mannosidase activity of Mns1, with those of Htm1 and their homologues in chicken and human. Identical amino acids are highlighted. (C) Schematic structure of hATF6α and hATF6α(C)-BACE(TM)-TAP. Potential N-glycosylation sites are shown schematically. (D) Immunoblotting of cell lysates prepared from WT and hEDEM2-KO HCT116 cells in which Myc-hATF6α (top) or hATF6α(C)-BACE(TM)-TAP (bottom) was expressed by transfection together with hEDEM2-Flag or hEDEM2-E117Q-Flag, using anti–c-myc and anti-Flag antibodies. (E) Immunoblottig of cell lysates prepared from WT HCT116, hEDEM2-KO, and hEDEM2-KO cells stably expressing hEDEM2-Flag or hEDEM2-E117Q-Flag using anti-human ATF6α and anti-Flag antibodies. (F) Pulse chase to determine the degradation rate of endogenous hATF6α in HCT116 hEDEM2-KO and hEDEM2-KO cells stably expressing hEDEM2-Flag or hEDEM2-E117Q-Flag (n = 3).
The enzymatic activity of Mns1 is strongly dependent on the EF hand motif in the mannosidase homology domain, and mutation of the glutamic acid within it (Fig. 4 B, red circle) is known to cause inactivation (Lipari and Herscovics, 1999). Importantly, when myc-tagged hATF6α (full length; Fig. 4 C, top) or hATF6α(C)-BACE(TM)-TAP (Horimoto et al., 2013; a shorter, artificial yet functional transmembrane protein containing the luminal region of hATF6α and the transmembrane region of BACE501, which is subjected to gpERAD; Fig. 4 C, bottom) was cotransfected with Flag-tagged hEDEM2 in hEDEM2-KO cells, these proteins migrated in SDS-PAGE at the same positions as in WT cells (Fig. 4 D). In contrast, no such rescue effect was observed with Flag-tagged mutant hEDEM2-E117Q (Fig. 4 D). These differences in migration were also observed for endogenous hATF6α in WT cells and hEDEM2-KO cells stably expressing Flag-tagged hEDEM2 or hEDEM2-E117Q (Fig. 4 E). Most importantly, stable introduction of Flag-tagged hEDEM2, but not Flag-tagged hEDEM2-E117Q, into hEDEM2-KO cells restored degradation of endogenous hATF6α (Fig. 4 F).
The mechanism of mammalian gpERAD was originally proposed based on the results of yeast genetics and biochemistry and overexpression in mammalian cells. However, here our pioneering application of TALEN to dissect and identify the functions of multigene family members (EDEMs) in human cells revealed that the roles of Mns1 and Htm1 homologues in mammalian gpERAD are completely discordant with those in that earlier proposed mechanism. The Mns1 homologue ERmanI plays a minor role in the conversion of M9 to M8B, the first step of gpERAD, unlike in yeast, whereas the previously passed-over Htm1 homologue EDEM2 most surprisingly plays a major role (Fig. 5 B).
Models of yeast and mammalian gpERAD. (A) In yeast, high-mannose-type oligosaccharide attached to asparagine (Glc3Man9GlcNAc2, G3M9) is first trimmed to M9 by glucosidases Gls1 and Gls2. M9 is trimmed to M8B by Mns1 and M8B is trimmed to M7A by Htm1. G1M9 is recognized by lectin chaperone Cne1 for folding, whereas M7A exposing α1,6-mannose is recognized by lectin Yos9 for subsequent degradation. The rate-limiting step in yeast gpERAD is the trimming from M8B to M7A. Refer to reviews by Molinari (2007) and Hosokawa et al. (2010a). (B) In mammals, G3M9 is trimmed to M9 by glucosidase I (GI) and glucosidase II (GII), homologues of yeast Gls1 and Gls2, respectively. G1M9 is recognized by lectin chaperones calnexin (CNX) and calreticulin (CRT), homologues of yeast Cne1, for folding. As our current results unambiguously show that M9 is mainly trimmed to M8B by EDEM2 and that M8B is trimmed by EDEM1 and EDEM3 to Man7-5GlcNAc2, which are recognized by lectin OS-9, a homologue of yeast Yos9, for degradation, mammalian gpERAD has two rate-limiting steps.
Models of yeast and mammalian gpERAD. (A) In yeast, high-mannose-type oligosaccharide attached to asparagine (Glc3Man9GlcNAc2, G3M9) is first trimmed to M9 by glucosidases Gls1 and Gls2. M9 is trimmed to M8B by Mns1 and M8B is trimmed to M7A by Htm1. G1M9 is recognized by lectin chaperone Cne1 for folding, whereas M7A exposing α1,6-mannose is recognized by lectin Yos9 for subsequent degradation. The rate-limiting step in yeast gpERAD is the trimming from M8B to M7A. Refer to reviews by Molinari (2007) and Hosokawa et al. (2010a). (B) In mammals, G3M9 is trimmed to M9 by glucosidase I (GI) and glucosidase II (GII), homologues of yeast Gls1 and Gls2, respectively. G1M9 is recognized by lectin chaperones calnexin (CNX) and calreticulin (CRT), homologues of yeast Cne1, for folding. As our current results unambiguously show that M9 is mainly trimmed to M8B by EDEM2 and that M8B is trimmed by EDEM1 and EDEM3 to Man7-5GlcNAc2, which are recognized by lectin OS-9, a homologue of yeast Yos9, for degradation, mammalian gpERAD has two rate-limiting steps.
We found that although hEDEM1 appeared to possess the weakest α1,2-mannosidase activity among the three hEDEMs (Fig. 3 F), deletion of hEDEM1 significantly delayed the degradation of hATF6α (Fig. 4 A). These observations are consistent with previous studies on EDEM1 (Hosokawa et al., 2010b; Ron et al., 2011; Shenkman et al., 2013). We also found that overexpression of both WT hEDEM1 and enzymatically inactive mutant hEDEM1-E225Q enhanced degradation of hATF6α(C)-BACE(TM)-TAP (Fig. S3 I). This suggests that hEDEM1 can function in gpERAD independently of its α1,2-mannosidase activity, when overexpressed or up-regulated by the unfolded protein response during ER stress (Yoshida et al., 2003), likely as a lectin for substrate delivery as proposed previously (Cormier et al., 2009).
The results obtained with TALEN-based KO in human cells are consistent with those we obtained here with conventional KO in chicken cells, demonstrating the reliability of TALEN technology. Particularly in human cells, the summation of the increase of M8B levels in hEDEM1-KO and hEDEM3-KO cells was nearly identical to the increase in M9 level in hEDEM2-KO cells (Fig. 3 F), indicating that conversion of M8B to Mα1,6E, the second step, is performed mainly by hEDEM3 and to a lesser extent by hEDEM1, not by hEDEM2. In connection with this subject, it was recently shown that SEL1L, a partner protein of the E3 ligase HRD1 spanning the ER membrane, binds to EDEM1 and EDEM3 but not to EDEM2 (Saeed et al., 2011; confirmation in Fig. S3 J). These results are consistent with our placement of EDEM1 and EDEM3 downstream of EDEM2 (Fig. 5 B).
Nonetheless, conversion of M9 to M8B by hEDEM2 is much slower than that by yeast Mns1 because M9 and M8B exist at comparable levels in human cells (Fig. 1 E). In other words, mammalian gpERAD has two rate-limiting steps in mannose trimming. As overexpression of hEDEM2 or hEDEM3 did not affect the level of hATF6α(C)-BACE(TM)-TAP (Fig. S3 I), the expression levels of EDEM2 or EDEM3 are not the primary determinants of the speed of mannose trimming. Interestingly, it was recently shown that EDEM1 and EDEM2 bind to ricin A chain, which is retrotranslocated from the ER to the cytosol similarly to ERAD substrates, but they bind much less efficiently to its P250A mutant, which is less hydrophobic than the WT (Sokołowska et al., 2011; Słomińska-Wojewódzka et al., 2014), suggesting that they recognize not only sugar structures but also protein structures.
Taking the aforementioned findings all together we propose that mammalian cells double check gpERAD substrates by evolving a novel-type Htm1 homologue that catalyzes the first mannose trimming step from M9 and thereby expressing three EDEMs with different specificity for glycoprotein folding/misfolding, first by EDEM2 (for conversion of M9 to M8B) and then by EDEM1/3 (for conversion of M8B to Mα1,6E), before recognition by lectin OS-9 (Fig. 5 B). This is more sophisticated than single check by Htm1 in yeast cells, as it avoids the unnecessary destruction of proteins, whose synthesis is energy expensive, and allows the escape of folded proteins from ERAD.
Materials and methods
Construction of plasmids
Recombinant DNA techniques were performed according to standard procedures (Sambrook et al., 1989) and the integrity of all constructed plasmids was confirmed by extensive sequencing analyses. Site-directed mutagenesis was performed using DpnI. p3*flag-CMVTM-14 expression vector (Sigma-Aldrich) was used to express a protein tagged with Flag at the C terminus. Platinum TALEN plasmids were constructed as described previously (Sakuma et al., 2013). In brief, each DNA-binding module was assembled into ptCMV-136/63-VR vectors using the two-step Golden Gate cloning method. Assembled sequences were 5′-TGGCGAGCGCTCGTCCTGgggctggtgctcctcCGGCTTGGCCTCCATGGA-3′ for hEDEM1, 5′-TCCGGCTGCTCATCCCGctcggcctcctgtgcGCGCTGCTGCCTCAGCA-3′ for hEDEM2, and 5′-TGTGGGTCCCCGGTTCCccagcgagcgcgatgGAGACTAGTGGCGGCGA-3′ for hEDEM3, where uppercase and lowercase letters indicate TALEN target sequences and spacer sequences, respectively.
Cell culture and transfection
DT40 cells were cultured and transfected as described previously (Ninagawa et al., 2011). HCT116 cells (CCL-247; ATCC) were cultured in Dulbecco’s modified Eagle’s medium (4.5 g/l glucose) supplemented with 10% fetal bovine serum, 2 mM glutamine, and antibiotics (100 U/ml penicillin and 100 µg/ml streptomycin) at 37°C in a humidified 5% CO2/95% air atmosphere. Transfection was performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. HCT116 cells stably expressing hEDEM2-Flag or hEDEM2-E117Q-Flag from p3*Flag-CMVTM14 expression vector were selected after culture for 20 d in the presence of 0.6 mg/ml G418.
Analysis of N-glycans of total cellular glycoproteins
Pyridylamination and structural identification of N-glycans of total cellular glycoproteins were performed as described previously (Hosokawa et al., 2010b). In brief, N-glycans released from delipidated cells by hydrazinolysis were reN-acetylated using acetic anhydrides, and then labeled with fluorescent 2-aminopyridine (Wako Pure Chemical Industries). Pyridylaminated (PA) oligosaccharides were fractionated by HPLC first on a TSK gel Amide-80 (amide-silica) column (Tosoh) and second on a Shim-pack HRC-octadecyl silica column (Shimadzu). Elution times of individual peaks from the amide-silica and octadecyl silica columns were normalized with respect to the degree of polymerization of PA-isomalto-oligosaccharide and are represented in units of glucose. N-Glycan structures were identified based on their elution times from these columns in comparison with those of PA-glycans in the GALAXY database (Takahashi and Kato, 2003) and confirmed by cochromatography with standard PA-high mannose type oligosaccharides (Tomiya et al., 1991; Kamiya et al., 2008).
Reagents
Puromycin (0.5 µg/ml), histidinol (1 mg/ml), mycophenolic acid (15 µg/ml), blasticidin S (25 µg/ml), G418 (2 mg/ml for DT40 and 0.6 mg/ml for HCT116), and zeocin (1 mg/ml) were used for the selection and maintenance of drug-resistant clones. Kifunensine was purchased from Cayman Chemical Company; endoglycosidase H from EMD Millipore; cycloheximide from Sigma-Aldrich; MG132 from Peptide Institute; and Z-VAD-fmk from Promega.
Immunological techniques
Immunoblotting analysis was performed according to the standard procedure (Sambrook et al., 1989) as described previously (Ninagawa et al., 2011). Chemiluminescence obtained using Western blotting Luminol Reagent (Santa Cruz Biotechnology, Inc.) was detected using an LAS-3000mini LuminoImage analyzer (Fujifilm). Rabbit anti–chicken ATF6 (Horimoto et al., 2013) and rabbit anti–human ATF6α (Haze et al., 1999) antibodies were raised previously. Mouse anti–c-myc antibody (9E10) was obtained from Wako Pure Chemical Industries; mouse anti-Flag antibody from Sigma-Aldrich; mouse anti-KDEL antibody from Medical and Biological Laboratories; rabbit anti–human ATF4 and rabbit anti–mouse XBP1 antibodies from Santa Cruz Biotechnology, Inc.; and rabbit anti–human GAPDH from Trevigen. Rabbit anti–human SEL1L antibody was a gift from N. Hosokawa (Kyoto University, Kyoto, Japan).
Pulse-chase experiments and subsequent immunoprecipitation were performed according to procedures described previously (Ninagawa et al., 2011). After incubation for 30 min in methionine- and cysteine-free Dulbecco’s modified Eagle’s medium (Invitrogen) supplemented with 2 mM glutamine and 10% dialyzed fetal bovine serum, cells were pulse labeled with 2.0 Mbq/dish EXPRE35S35S protein labeling mixture (PerkinElmer) and then chased in fresh complete medium. Cells were lysed in buffer A (50 mM Tris/Cl, pH 8.0, containing 1% NP-40, 150 mM NaCl, protease inhibitor cocktail (Nacalai), 20 µM MG132, and 2 µM Z-VAD-fmk). Immunoprecipitation was performed using anti-hATF6α antibody and protein A–coupled Sepharose beads (GE Healthcare). Beads were washed with high salt buffer (50 mM Tris/Cl, pH 8.0, containing 1% NP-40 and 150 mM NaCl) twice, washed with PBS, and heated at 65°C in Laemmli’s sample buffer. Immunoprecipitates were subjected to SDS-PAGE, and radioactive bands were analyzed using a FLA-3000G FluoroImage analyzer (Fujifilm).
Quantitative RT-PCR
Quantitative RT-PCR was performed using the SYBR Green method (Applied Biosystems) using a pair of primers, namely, 5′-CTGCACATGGCTCATGACCTA-3′ and 5′-ACCCGAGGATACGGAATTCC-3′ for gEDEM1, 5′-CTGAAGGCACTGTGGAAAAACC-3′ and 5′-GCCACCCATTTGGCAGTTAC-3′ for gEDEM2, 5′-GCGCAGCAGCAATTTGGT-3′ and 5′-CCGCTCCAACTCCACTATCTTT-3′ for gEDEM3, 5′-TTTGGTGTGCAACAACCTATG-3′ and 5′-CCTCTTATTGGACTGCTCTTCA-3′ for gSEL1L, 5′-AGTGCCAAGATGGATCACTTG-3′ and 5′-TGATCAGCAGTCAATCCATTG-3′ for gERmanI, 5′-CTGATGCCCCCATGTTTGTG-3′ and 5′-GCACGATGCATTGCTGACA-3′ for gGAPDH, 5′-AGTTCCTCCTGACACCAATA-3′ and 5′-GTCGACTCAGAATCCCAAAT-3′ for hEDEM1, 5′-CGCTTCGATGACTGGTACCT-3′ and 5′-TAGGCCTCCAAGGACTGGAA-3′ for hEDEM2, 5′-ATTAGCCAGCCACCTCTTCT-3′ and 5′-AGCAAAGCATCCATCCAAGT-3′ for hEDEM3, 5′-GGTTTGGCACCGATGTAGAT-3′ and 5′-AGCTTGTGCACTGTGTTGCT-3′ for hSEL1L, 5′-CCCGAGCCTAGGGACAAGAT-3′ and 5′-CCCGAGCCTAGGGACAAGAT-3′ for hERmanI, or 5′-GACCCCTTCATTGACCTCAA-3′ and 5′-TTGACGGTGCCATGGAATT-3′ for hGAPDH. 2,000, 8,000, 20,000, 80,000, and 200,000 molecules of plasmid carrying the respective gene were used as standards.
Southern blot hybridization
Southern blot hybridization was performed according to standard procedures (Sambrook et al., 1989) as described previously (Ninagawa et al., 2011). Specific probes were amplified by PCR from DT40 cell genomic DNA using the primers 5′-TGAAACTTGCACTGCAGGAG-3′ and 5′-ACCCCCTTCGTAAGTGGTTC-3′ for gEDEM1, 5′-GAAGGCTCTGAAGGCACTGT-3′ and 5′-TGTCGGATCCCTTTCAACTC-3′ for gEDEM2, 5′-AGCCGATTCACAGGAACATC-3′ and 5′-AAGAAGGGCAGTGCACAAAT-3′ for gEDEM3, or 5′-AAATTATTCTGATGCACATG-3′ and 5′-CTCTTGTATGCATCGATATGf-3′ for gERmanI, or from HCT116 cell genomic DNA using the primers 5′-CCACTGACCCTAACAATTGC-3′ and 5′-AAATGAGGCAAAGTAATGCC-3′ for hEDEM2 and 5′-GGTCACTTGATGGAAAAAAGC-3′ and 5′-ATTTGGTTGAACTTTTCCAG-3′ for hEDEM3. These probes were labeled with digoxigenin. Subsequent reaction with anti-digoxigenin antibody (Roche) and treatment with the chemiluminescent detection reagent CDP-star (GE Healthcare) were performed according to the manufacturer’s specifications. Chemiluminescence was visualized using an LAS-3000mini LuminoImage analyzer.
Genomic PCR
Homologous recombination in HCT116 cells was confirmed by genomic PCR using a pair of primers: 5′-CTATGTGCCAGCTACCATGTG-3′ and 5′-TACTCCATGGAGGCCAAGCC-3′ for hEDEM1, 5′-CCCCAGGGTAGTCATTTGTC-3′ and 5′-ATCTGGCGCGGAGCCGTCGG-3′ for hEDEM2, or 5′-GAGTACAGAGAGAAAAAGGAC-3′ and 5′-GCCACTAGTCTCCATCGCGC-3′ for hEDEM3. The presence of the ampicillin-resistance gene in targeted alleles was checked by genomic PCR using a pair of primers: 5′-GAGCACTTTTAAAGTTCTGC-3′ and 5′-TTACCAATGCTTAATCAGTG-3′.
RT-PCR
Total RNA prepared from WT DT40 cells or various KOs (∼5 × 106 cells) and from WT HCT116 cells or various KOs (∼3 × 106 cells) by the acid guanidinium/phenol/chloroform method using ISOGEN (Nippon Gene) was converted to cDNA using Moloney murine leukemia virus reverse transcription (Invitrogen) and random primers. Full-length open reading frame of gEDEM1, gEDEM2, gEDEM3, gERmanI, hEDEM1, hEDEM2, or hEDEM3 was amplified using PrimeSTAR HS DNA polymerase (Takara Bio Inc.) and a pair of primers, namely, 5′-GGGTACCATGCAATGGCGCTCGCTGGT-3′ and 5′-CTCTAGAGATCAAGCCCACCATCCGAT-3′ for gEDEM1, 5′-GGAATTCACCATGGCGCTGCTGCGCTCGCT-3′ and 5′-GCAGATCTGGAGTGTTGTCCAGGAAGACCT-3′ for gEDEM2, 5′-GGAATTCACCATGGGCGGAGCTGCGGGCTG-3′ and 5′-CTATCGATGGTAGTTCATCCTTTTCCATCA-3′ for gEDEM3, 5′-GAAGATCTCGCCATGTACGCTGCCGCCGCC-3′ and 5′-GCTCTAGATGCTGGCACCCAGATGGGGAG-3′ for gERmanI, 5′-GAAGATCTGACCATGCAATGGCGAGCGCTC-3′ and 5′-CGGGATCCAATCAAACCAACCATCTGGTC-3′ for hEDEM1, 5′-CCCAAGCTTGCTCTATGCCTTTCCGGCTGC-3′ and 5′-GCTCTAGATGAGGAGTCTAGGAAAACCTG-3′ for hEDEM2, or 5′-GAAGATCTGGCCATGAGCGAAGCCGGCG-3′ and 5′-CGGGATCCTAGCTCATCCTTCTCCATCA-3′ for hEDEM3.
In vitro translation
In vitro translation was performed using the TNT Quick Coupled Transcription/Translation Systems (Promega) and EASY TAG EXPRESS Protein labeling mix [35S] (PerkinElmer) according to the manufacturer’s instructions. Translated proteins were subjected to SDS-PAGE (12% gel) and radiolabeled proteins were visualized using an FLA-3000G FluoroImage analyzer.
Construction of gEDEM1 targeting vector
The 3.6-kb fragment of the gEDEM1 gene used for the 5′ arm was amplified by PCR from DT40 cell genomic DNA using the primers 5′-CCTCGAGTGGAGCTGTACCTGCTGTTG-3′ and 5′-CGGATCCGTAGCTGTCGTAGCCGAAGG-3′, and then subcloned between the XhoI and BamHI sites of pBluescript II KS (+) vector to create the pBluescript-5′ arm (gEDEM1). The 2.3-kb fragment of the gEDEM1 gene used for the 3′ arm was amplified similarly using the primers 5′-CCGGATCCGGGAAATTCTTCCGAGTTCC-3′ and 5′-CCCTCGAGGGAACTCCCTTCTTCAGGTTG-3′, and then subcloned between the BamHI and NotI sites of the pBluescript-5′ arm (gEDEM1) to create the pBluescript-5′-3′ arms (gEDEM1). The puromycin-resistance gene flanked by loxP sites designated Puro-loxP or histidinol-resistance (histidinol dehydrogenase [HisD]) gene was subcloned into the BamHI site of the pBluescript-5′-3′ arms (gEDEM1) to create pKO-gEDEM1-puromycin or pKO-gEDEM1-histidinol, respectively (Fig. S1 A). These constructs were transfected into DT40 cells by electroporation after linearization using SacI.
Construction of gEDEM2 targeting vector
The 3.2-kb fragment of the gEDEM2 gene used for the 5′ arm was amplified by PCR from DT40 cell genomic DNA using the primers 5′-GGACTAGTTAGCAGCAGGAGACCCTTGT-3′ and 5′-CGGGATCCGCTCGTAGGCGTGGTAGAAC-3′, and then subcloned between the SpeI and BamHI sites of pBluescript II KS (+) vector to create the pBluescript-5′ arm (gEDEM2). The 2.4-kb fragment of the gEDEM2 gene used for the 3′ arm was amplified similarly using the primers 5′-CGGGATCCGCAATGTCACCGAGTTCCAG-3′ and 5′-CCCTCGAGACCAATCCCAGCAGTACAGG-3′, and then subcloned between the BamHI and XhoI sites of the pBluescript-5′ arm (gEDEM2) to create the pBluescript-5′-3′ arms (gEDEM2). The neomycin-resistance gene flanked by loxP sites designated Neo-loxP or blasticidin S-resistance gene flanked by loxP sites designated Bsr-loxP was subcloned into the BamHI site of the pBluescript-5′-3′ arms (gEDEM2) to create pKO-gEDEM2-neomycin or pKO-gEDEM2-blasticidin S, respectively. HisD was also subcloned into the BamHI site of the pBluescript-5′-3′ arms (gEDEM2) to create pKO-gEDEM2-histidinol (Fig. S1 D). These constructs were transfected into DT40 cells by electroporation after linearization using SpeI or ScaI.
Construction of gEDEM3 targeting vector
The 2.2-kb fragment of the gEDEM3 gene used for the 5′ arm was amplified by PCR from DT40 cell genomic DNA using the primers 5′-CGAGCTCCCATGAGCAAGGAGGAGAAG-3′ and 5′-CGGGATCCTGGTCAAACATTTCCAGCAC-3′, and then subcloned between the SacI and BamHI sites of pBluescript II KS (+) vector to create the pBluescript-5′ arm (gEDEM3). The 3.4-kb fragment of the gEDEM3 gene used for the 3′ arm was amplified similarly using the primers 5′-CGGGATCCTTCTCCCTGACCTTGATTGA-3′ and 5′-CCCTCGAGAATCGGCTTAAAGCTGCAAA-3′, and then subcloned between the BamHI and XhoI sites of the pBluescript-5′ arm (gEDEM3) to create the pBluescript-5′-3′ arms (gEDEM3). The Neo-loxP was subcloned into the BamHI site of the pBluescript-5′-3′ arms (gEDEM3) to create pKO-gEDEM3-neomycin. The HisD or mycophenolic acid–resistance gene, designated E. coli xanthine-guanine phosphoribosyltransferase (EcoGPT), was also subcloned into the BamHI site of the pBluescript-5′-3′ arms (gEDEM3) to create pKO-gEDEM3-histidinol or pKO-gEDEM3-mycophenolic acid, respectively (Fig. S1 G). These constructs were transfected into DT40 cells by electroporation after linearization using PvuI or SacI.
Construction of gERmanI targeting vector
The 3.6-kb fragment of the gERmanI gene used for the 5′ arm was amplified by PCR from DT40 cell genomic DNA using the primers 5′-ATAAGAATGCGGCCGCACATGAACACTGAACTGCTG-3′ and 5′-CGGGATCCGCTTCCATTTCTTCCAGAGAG-3′, and then subcloned between the NotI and BamHI sites of pBluescript II KS (+) vector to create the pBluescript-5′ arm (gERmanI). The 2.8-kb fragment of the gERmanI gene used for the 3′ arm was amplified similarly using the primers 5′-CCATCGATGACTACCCAGAGCTTTCACC-3′ and 5′-CCGCTCGAGCAACTCAATCAAAATTCTG-3′, and then subcloned between the ClaI and XhoI sites of the pBluescript-5′ arm (gERmanI) to create the pBluescript-5′-3′ arms (gERmanI). Neo-loxP or Puro-loxP was subcloned into the BamHI site of the pBluescript-5′-3′ arms (gERmanI) to create pKO-gERmanI-neomycin or pKO-gERmanI-puromycin, respectively (Fig. S1 L). These constructs were transfected into DT40 cells by electroporation after linearization using NotI.
Construction of gEDEM1-, gEDEM2-, gEDEM3-, and gERmanI-KO cells
Exons encoding the N-terminal region of the mannosidase homology domain of gEDEM1, gEDEM2, or gEDEM3 were replaced with one of the five drug-resistance genes (Fig. S1, A, D, and G). Homologous recombination was checked by Southern blotting (Fig. S1, B, E, and H). We obtained two independent clones (−/− #1 and #2), in which the allelic gEDEM1 genes were disrupted by HisD and Puro-loxP (Fig. S1, B and C). As RT-PCR analysis showed that gEDEM1 mRNA was not expressed in these clones (Fig. S1 J), we used −/− #1 as gEDEM1-KO (Fig. 2 A). The gEDEM2 gene is encoded in chromosome 2, which is in trisomy in DT40 cells. We thereby obtained two independent clones (−/−/− #1 and #2) in which all three alleles of the gEDEM2 gene were disrupted by Neo-loxP, HisD, and Bsr-loxP (Fig. S1, E and F). Neo-loxP and Bsr-loxP were then eliminated from their genome using the Cre-loxP system to obtain −/−/− (loxP−) #1 and #2 for further disruption (Fig. S1, E and F). RT-PCR analysis showed that these loxP− clones expressed some shorter gEDEM2 mRNAs (Fig. S1 J). Thus, gEDEM2 cDNA was amplified by RT-PCR from WT cells and gEDEM2 −/−/− (loxP−) #1 and #2 using a pair of primers (5′-CGCAAGCTTGCGGCCATGGCGCTGCTGCG-3′ and 5′-CGCTCTAGATCAAGTGTTGTCCAGGAAGAC-3′), and then subcloned between the HindIII and XbaI sites of pcDNA3.1(+) (Invitrogen). Sequencing of the resulting plasmids revealed that gEDEM2 cDNA obtained from gEDEM2 −/−/− (loxP−) #1 and #2 was a mixture of two cDNAs designated a and b and that both contained one or two deletions, which created a stop codon in exon 5 (Fig. S1 K). Indeed, in vitro translation of the fragment did not produce any protein, whereas that of the WT fragment produced a protein of 75 kD, as expected (Fig. S1 K). We thus used −/−/− (loxP−) #1 as gEDEM2-KO (Fig. 2 A). Two independent clones (−/− #1 and #2) in which both alleles of the gEDEM3 gene were disrupted by EcoGPT and HisD or Neo-loxP were obtained (Fig. S1, H and I). As RT-PCR analysis showed that gEDEM3 mRNA was not expressed in these clones (Fig. S1 J), we used −/− #1 as gEDEM3-KO (Fig. 2 A).
Exons encoding the transmembrane domain of gERmanI were replaced with drug-resistance genes (Fig. S1 L). Two independent clones (−/− #1 and #2) in which both alleles of the gERmanI gene were disrupted by Neo-loxP and Puro-loxP were obtained (Fig. S1, M and N). As RT-PCR analysis showed that gERmanI mRNA was not expressed in these clones (Fig. S1 O), we used −/− #1 as gERmanI-KO (Fig. 2 A).
Construction of human targeting vectors
The DT-A-pA/loxP/PGK-Neo-pA/loxP vector was provided by Laboratory for Animal Resources and Genetic Engineering, Center for Developmental Biology, Institute of Physical and Chemical Research, Kobe, Japan. The puromycin- and zeocin-resistance genes were amplified from pPur (Takara Bio Inc.) and pVgRXR (Invitrogen) vectors, respectively, by PCR using the primers 5′-CGACCTGCAGCCAATATGACCGAGTACAAGCCCACGG-3′ and 5′-TTACAGCGGATCCCCTCAGGCACCGGGCTTGC-3′ and 5′-CGACCTGCAGCCAATATGGCCAAGTTGACCAGTGCC-3′ and 5′-TTACAGCGGATCCCCTCAGTCCTGCTCCTCGGCC-3′, respectively, and then inserted using the In-Fusion cloning method (Takara Bio Inc.) into the inverse PCR product of DT-A-pA/loxP/PGK-Neo-pA/loxP amplified with the primers 5′-GGGGATCCGCTGTAAGTCTGC-3′ and 5′-ATTGGCTGCAGGTCGAAAGGC-3′, resulting in obtaining DT-A-pA/loxP/PGK-Puro-pA/loxP and DT-A-pA/loxP/PGK-Zeo-pA/loxP, respectively, in which the neomycin-resistance gene was replaced with the puromycin- and zeocin-resistance genes, respectively.
Construction of hEDEM1 targeting vector
The 1.5-kb fragment of the hEDEM1 gene used for the 5′ arm was amplified by PCR from HCT116 cell genomic DNA using the primers 5′-ATAAGAATGCGGCCGCGTGCTCCTCCGGCTTGGCCTC-3′ and 5′-CCCAAGCTTCCTTATCAGAACCTAAG-3′, and then subcloned between the NotI and HindIII sites of the DT-A-pA/loxP/PGK-Neo-pA/loxP vector to create the DT-A-pA/loxP/PGK-Neo-pA/loxP-5′ arm (hEDEM1). The 1.5-kb fragment of the hEDEM1 gene used for the 3′ arm was amplified similarly using the primers 5′-GGGGTACCAAGTGTTTTCTTAGTTCCC-3′ and 5′-CCGCTCGAGCAGCCCCAGGACGAGCGCTC-3′, and then subcloned between the KpnI and XhoI sites of the DT-A-pA/loxP/PGK-Neo-pA/loxP-5′ arm (hEDEM1) to create pKO-hEDEM1-Neo, which was transfected into HCT116 cells. Approximately 100 colonies were obtained 20 d later.
Construction of hEDEM2 targeting vector
The 1.5-kb fragment of the hEDEM2 gene used for the 5′ arm was amplified by PCR from HCT116 cell genomic DNA using the primers 5′-GGGGTACCGCGCGATCCTCGCTCACTGC-3′ and 5′-CCGCTCGAGCCGGAAAGGCATAGAGCTCG-3′, and then subcloned between the KpnI and XhoI sites of the DT-A-pA/loxP/PGK-Puro-pA/loxP vector to create the DT-A-pA/loxP/PGK-Puro-pA/loxP-5′ arm (hEDEM2). The 1.5-kb fragment of the hEDEM2 gene used for the 3′ arm was amplified similarly using the primers 5′-ATAAGAATGCGGCCGCTGTGCGCGCTGCTGCCTCAG-3′ and 5′-CCCAAGCTTGGTCACTTCAAACGAGTGTAG-3′, and then subcloned between the NotI and HindIII sites of the DT-A-pA/loxP/PGK-Puro-pA/loxP-5′ arm (hEDEM2) to create pKO-hEDEM2-Puromycin, which was transfected into HCT116 cells. Approximately 100 colonies were obtained 17 d later.
Construction of hEDEM3 targeting vector
The 1.5-kb fragment of the hEDEM3 gene used for the 5′ arm was amplified by PCR from HCT116 cell genomic DNA using the primers 5′-GGGGTACCGAAAACCTTAAGGGAACACC-3′ and 5′-CCGCTCGAGCGCTGGGGAACCGGGGACCC3′, and then subcloned between the KpnI and XhoI sites of the DT-A-pA/loxP/PGK-Zeo-pA/loxP vector to create the DT-A-pA/loxP/PGK-Zeo-pA/loxP-5′ arm (hEDEM3). The 1.5-kb fragment of the hEDEM3 gene used for the 3′ arm was amplified similarly using the primers 5′-ATAAGAATGCGGCCGCAGCGCGATGGAGACTAGTGG-3′ and 5′-ATAAGAATGCGGCCGCTTTGCAATGATTTAAAGTAC-3′, and then subcloned into the NotI site of the DT-A-pA/loxP/PGK-Zeo-pA/loxP-5′ arm (hEDEM3) to create pKO-hEDEM3-Zeocin, which was transfected into HCT116 cells. Approximately 20 colonies were obtained 23 d later.
Online supplemental material
Fig. S1 shows how DT40 cells deficient in gEDEM1, gEDEM2, gEDEM3, or gERmanI are generated by homologous recombination. Fig. S2 shows characterization of high-mannose-type oligosaccharides of total glycoproteins prepared from DT40 cells of various genotypes. Fig. S3 shows characterization of HCT116 cells deficient in hEDEM1, hEDEM2, or hEDEM3 including Southern blot hybridization for confirmation of correct targeting and characterization of high-mannose-type oligosaccharides of total glycoproteins prepared from HCT116 cells of various genotypes.
Acknowledgments
We thank Ms. Kaoru Miyagawa for her technical and secretarial assistance, Ms. Yukiko Isono for her help in N-glycan analysis, and Dr. Elizabeth Nakajima for careful reading of this manuscript. We are grateful to Dr. Nobuko Hosokawa for providing anti-SEL1L antibody and to the Laboratory for Animal Resources and Genetic Engineering, Center for Developmental Biology, Institute of Physical and Chemical Research, Kobe, for providing the DT-A-pA/loxP/PGK-Neo-pA/loxP vector.
This work was financially supported in part by grants from the Ministry of Education, Culture, Sports, Science and Technology, Japan (19058009 and 26291040 to K. Mori, 26840065 to S. Ninagawa, 23221005 to T. Okada, 22020039 and 26102518 to Y. Kamiya, and 24249002 and 25102008 to K. Kato).
The authors declare no competing financial interests.
References
Author notes
Y. Kamiya’s present address is Graduate School of Engineering and EcoTopia Science Institute, Nagoya University, Nagoya 464-8603, Japan.