The promyelocytic leukemia (PML) protein organizes PML nuclear bodies (NBs), which are stress-responsive domains where many partner proteins accumulate. Here, we clarify the basis for NB formation and identify stress-induced partner sumoylation as the primary NB function. NB nucleation does not rely primarily on intermolecular interactions between the PML SUMO-interacting motif (SIM) and SUMO, but instead results from oxidation-mediated PML multimerization. Oxidized PML spherical meshes recruit UBC9, which enhances PML sumoylation, allow partner recruitment through SIM interactions, and ultimately enhance partner sumoylation. Intermolecular SUMO–SIM interactions then enforce partner sequestration within the NB inner core. Accordingly, oxidative stress enhances NB formation and global sumoylation in vivo. Some NB-associated sumoylated partners also become polyubiquitinated by RNF4, precipitating their proteasomal degradation. As several partners are protein-modifying enzymes, NBs could act as sensors that facilitate and confer oxidative stress sensitivity not only to sumoylation but also to other post-translational modifications, thereby explaining alterations of stress response upon PML or NB loss.
Introduction
The eukaryotic nucleus contains domains organized by master proteins, such as promyelocytic leukemia (PML), which drives the formation of PML nuclear bodies (NBs; Lallemand-Breitenbach and de Thé, 2010). PML NBs are stress-regulated, dynamic structures that concentrate hundreds of proteins and finely tune multiple pathways including senescence, stemness, stress response, and defense against viruses (Koken et al., 1995; Dellaire and Bazett-Jones, 2004; Bernardi and Pandolfi, 2007; Ching et al., 2013). Functionally, NB disruption through expression of the PML/RARA oncogene has been implicated in acute promyelocytic leukemia (APL) pathogenesis. As2O3 (arsenic), an effective APL therapy, restores NBs through PML and PML/RARA oxidation, disulfide-mediated multimerization, or direct binding to PML, both followed by PML/RARA sumoylation and degradation (Jeanne et al., 2010; Zhang et al., 2010; de Thé et al., 2012). Arsenic similarly enhances NB biogenesis and nuclear matrix association in non-APL cells (Zhu et al., 1997). Yet, PML NB assembly and function remain imperfectly understood (Lallemand-Breitenbach and de Thé, 2010).
PML is sumoylated on three target lysines and displays a SUMO-interacting motif (SIM; Hecker et al., 2006; Kamitani et al., 1998). Accordingly, it has been proposed that NB nucleation depends on intermolecular interactions between a sumoylated PML and a SIM on PML C terminus (Fig. 1 A; Müller et al., 1998; Shen et al., 2006). In that regard, inactivation of sumoylation impairs NB formation (Nacerddine et al., 2005). Many PML partner proteins are also modified by SUMOs, suggesting that PML SIM may tether sumoylated partners onto NBs (Matunis et al., 2006; Shen et al., 2006). At variance with this model, we and others have shown that K160, a major SUMO acceptor site on PML, is not required for NB formation, but is critical for the recruitment of partners (Ishov et al., 1999; Zhong et al., 2000; Zhu et al., 2005; Lallemand-Breitenbach et al., 2001, 2008). Furthermore, an isoform that lacks the SIM yields nuclear bodies (Weidtkamp-Peters et al., 2008). Thus, the respective roles of PML or partner sumoylation in NB assembly remain disputed.
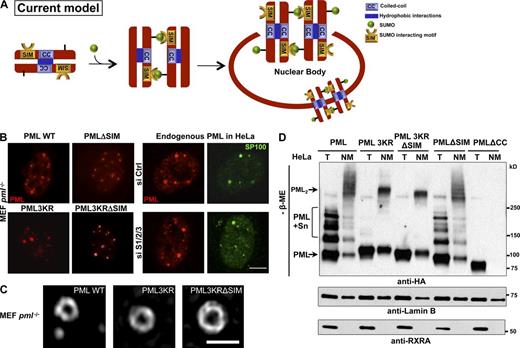
PML NB nucleation does not depend on SUMO interactions with SIM. (A) Schematic representation of the current model, in which NB biogenesis relies on noncovalent intermolecular interactions between PML-attached SUMO and the PML SIM. (B) PML and SP100 immunolocalization showing morphologically normal NBs in pml−/− immortalized MEFs stably expressing PML or the indicated mutants (left), and in HeLa cells in which SUMO1 and SUMO2/3 were inactivated (right). Bar, 5 µm. Quantitative data are indicated in Fig. S1 A. (C) Super-resolution microscopy analysis showing NBs formed by PML WT, PML3KR, or PML3KRΔSIM in pml−/− MEFs. Bar, 1 µm. (D) Western blot analysis under nonreducing conditions of pml−/− MEFs stably expressing the indicated HA-tagged PML mutants. Lysates of total cells (T) or nuclear matrix (NM) fractions. Lamin B and RXRA are fractionation controls. The top arrow (PML2) points to covalently linked PML multimers; sumoylated PML (PML + Sn) and unmodified PML are indicated.
PML NB nucleation does not depend on SUMO interactions with SIM. (A) Schematic representation of the current model, in which NB biogenesis relies on noncovalent intermolecular interactions between PML-attached SUMO and the PML SIM. (B) PML and SP100 immunolocalization showing morphologically normal NBs in pml−/− immortalized MEFs stably expressing PML or the indicated mutants (left), and in HeLa cells in which SUMO1 and SUMO2/3 were inactivated (right). Bar, 5 µm. Quantitative data are indicated in Fig. S1 A. (C) Super-resolution microscopy analysis showing NBs formed by PML WT, PML3KR, or PML3KRΔSIM in pml−/− MEFs. Bar, 1 µm. (D) Western blot analysis under nonreducing conditions of pml−/− MEFs stably expressing the indicated HA-tagged PML mutants. Lysates of total cells (T) or nuclear matrix (NM) fractions. Lamin B and RXRA are fractionation controls. The top arrow (PML2) points to covalently linked PML multimers; sumoylated PML (PML + Sn) and unmodified PML are indicated.
Sumoylation regulates multiple biological processes (Hay, 2005; Nacerddine et al., 2005; Cubeñas-Potts and Matunis, 2013). The SUMO E2–conjugating enzyme UBC9 is essential and may be sufficient for target sumoylation, but E3 enzymes may also bridge UBC9 to specific substrates, facilitating their sumoylation. That sumoylation can occur in the absence of specific E3s raises the issues of how its specificity and spatiotemporal regulation are achieved (Gareau and Lima, 2010). Cellular stress modulates global sumoylation through multiple mechanisms (Bossis and Melchior, 2006; Xu et al., 2009), and some key enzymes in the SUMO conjugation/deconjugation pathways are oxidative stress sensitive, including SUMO proteases and the SAE2-activating enzyme (Bossis and Melchior, 2006; Xu et al., 2009; Yeh, 2009). Among SUMO-regulated processes, poly- or multi-sumoylation may initiate polyubiquitination by the SUMO-targeted ubiquitin ligase (STUbL) Ring-finger protein 4 (RNF4) and proteasome-mediated degradation (Lallemand-Breitenbach et al., 2008; Tatham et al., 2008). Although this pathway was initially described for arsenic-induced PML or PML/RARA degradation, other NB-associated proteins may also be subjected to RNF4-mediated degradation.
The diversity of PML partner proteins recruited onto NBs has suggested a general function for NBs in their sequestration and/or activation (Eskiw et al., 2003; Lin et al., 2006; Bernardi and Pandolfi, 2007). In particular, overexpression of PML and/or of some specific partners modulates post-translational modifications of these partners (Bernardi and Pandolfi, 2007; Lallemand-Breitenbach and de Thé, 2010). Nevertheless, a global systematic analysis of PML NB function that establishes a role for NBs as global post-translational regulation sites is lacking.
Here, we dissect the mechanisms underlying NB biogenesis and show that NB formation regulates oxidative stress–responsive sumoylation through cooperation between PML, UBC9, and RNF4. Our results shed new light on how PML-dependent sumoylation could finely tune senescence or self-renewal.
Results
NB nucleation does not rely on PML SIM–SUMO interactions
The current model proposes that interaction of PML-conjugated SUMO with PML SIM nucleates NB biogenesis (Fig. 1 A). To test this, we stably expressed, in immortalized pml−/− mouse embryonic fibroblasts (MEFs), a PML mutant the SIM of which was excised (PMLΔSIM). However, this did not impede the formation of PML bodies (Fig. 1 B and Fig. S1 A). Similarly, a PML mutant devoid of its three sumoylation sites (PML3KR), and even PML3KRΔSIM, allowed the formation of the spherical PML lattice when stably expressed in pml−/− cells (Fig. 1, B and C). Interestingly, the latter did not exhibit a diffuse nuclear staining, possibly suggesting that SUMO and SIM allow interaction of PML with components of the chromatin. Although there were some changes in the mean number of bodies (Fig. S1 A), those directly reflected expression levels of the different PML mutants that have different stabilities or toxicities (Fig. S1, B and C; unpublished data). Importantly, super-resolution structured illumination microscopy (Gustafsson, 2000) revealed that these mutants form spherical structures exactly like wild-type PML (PML WT; Fig. 1 C). Finally, siRNA extinction of three SUMO paralogues did not prevent body formation (Fig. 1 B and Fig. S1, D and E), although it decreased their number and completely abrogated NB association of partners such as SP100 or DAXX (Fig. 1 B; unpublished data; Ishov et al., 1999; Lallemand-Breitenbach et al., 2001). The decrease in NB number may reflect the fact that the S/G2 phase of the cell cycle is associated with PML NB duplication and that SUMO extinction triggers a cell cycle arrest (Dellaire et al., 2006). Collectively, SUMO–SIM interactions are not required for the formation of the PML lattice, although they regulate subsequent partner protein recruitment into NBs to form mature NBs.
PML RBCC (RING-finger, B boxes, coiled-coil) domains are involved in noncovalent self-interactions and are required for NB formation (Kastner et al., 1992). NBs are components of the nuclear matrix (NM), and NM association largely relies on PML oxidation and intermolecular disulfide linkage (Jeanne et al., 2010). We thus compared the ability of PMLΔSIM, PML3KR, and PML3KRΔSIM to form NM-associated covalent multimers. In pml−/− MEFs stably expressing these PML mutants, total and NM extracts were analyzed by nonreducing Western blotting. The NM fraction was found to be dramatically enriched in covalent PML multimers—that disappeared upon reduction—in WT-, 3KR-, ΔSIM- and 3KRΔSIM-expressing cells, but, as expected, not in PMLΔCC-expressing cells (Fig. 1 D and Fig. S1 F). Thus, NB formation and NM association occur independently of PML sumoylation and SIM, but require noncovalent RBCC-mediated interactions for the assembly of oxidized PML multimers.
SIM and SUMOs are general NB-targeting signals
A literature search of NB resident proteins actually revealed that all those we examined both contain functional SIMs and may undergo sumoylation, suggesting that these motifs cooperate to enforce association to NBs (Fig. 2 A; and Fig. S2). To test this hypothesis, the SIMs of PML, HIPK2, or DAXX fused to the C terminus of GFP were stably expressed in CHO cells stably expressing PML (CHO-PML). Interestingly, GFP-SIM fusions, but not the GFP control, associated with NBs in an arsenic-enhanced manner (Fig. 2 B, top; quantification in Fig. 2 C). Similarly, by removing the di-glycine cleavage sites, we irreversibly fused SUMO paralogues to the N terminus of GFP. When expressed in CHO-PML cells, these SUMOΔGG-GFP fusions also localized to NBs, and were slightly sensitive to arsenic exposure (Fig. 2, B and C).
SIM and SUMOs are NB-targeting signals. (A) Prototypical PML partner with both a SIM and a SUMO conjugation site may interact with PML SUMO (1) and SIM (2) in an ordered manner. (B) Colocalization of GFP-SIM or SUMO1/2/3ΔGG-GFP fusions with NBs. (Top) As2O3 (arsenic)-enhanced NB recruitment of GFP-SIM fusions stably expressed in CHO-PML (arsenic: 10−6 M, 1 h). (Bottom) Recruitment to NBs of SUMO1, 2, or 3ΔGG-GFP fusions in CHO-PML but not in CHO cells. Insets show GFP labeling alone. (C) Quantification of GFP fusions’ recruitment on NBs: ratios of GFP intensities (in NBs versus in the nucleoplasm) were calculated cell by cell, and averaged from 20 cells. P-values are indicated. (D) FRAP analysis of NB-associated SUMOΔGG-GFPs or SIM-GFPs in CHO-PML cells (the graph represents means of five experiments); standard deviations are shown for GFP and S3ΔGG-GFPSIM. t1/2 recoveries after photo-bleaching are shown below (table). (E) Real-time diffusion analysis after Dendra-DAXX photoconversion performed on NBs or in the nucleoplasm of CHO-PML cells (means of five experiments are represented below). Error bars represent standard deviation. Half times of Dendra-DAXX diffusion after photoconversion are indicated below (t1/2). Insets show a representative green to red photoconverted ROI surrounding NB or in the nucleoplasm at t = 2.5 s, t = 9.5 s, and t = 45.3 s after switch. Bar, 0.5 µm. (F) Immunolocalization of PMLΔSIM or PMLK160R stably expressed in pml−/− MEFs or CHO cells, and of endogenous DAXX or transfected SP100, as indicated. (G) Immunolocalization of DAXX mutants stably expressed in CHO-PML cells. Bars, 5 µm.
SIM and SUMOs are NB-targeting signals. (A) Prototypical PML partner with both a SIM and a SUMO conjugation site may interact with PML SUMO (1) and SIM (2) in an ordered manner. (B) Colocalization of GFP-SIM or SUMO1/2/3ΔGG-GFP fusions with NBs. (Top) As2O3 (arsenic)-enhanced NB recruitment of GFP-SIM fusions stably expressed in CHO-PML (arsenic: 10−6 M, 1 h). (Bottom) Recruitment to NBs of SUMO1, 2, or 3ΔGG-GFP fusions in CHO-PML but not in CHO cells. Insets show GFP labeling alone. (C) Quantification of GFP fusions’ recruitment on NBs: ratios of GFP intensities (in NBs versus in the nucleoplasm) were calculated cell by cell, and averaged from 20 cells. P-values are indicated. (D) FRAP analysis of NB-associated SUMOΔGG-GFPs or SIM-GFPs in CHO-PML cells (the graph represents means of five experiments); standard deviations are shown for GFP and S3ΔGG-GFPSIM. t1/2 recoveries after photo-bleaching are shown below (table). (E) Real-time diffusion analysis after Dendra-DAXX photoconversion performed on NBs or in the nucleoplasm of CHO-PML cells (means of five experiments are represented below). Error bars represent standard deviation. Half times of Dendra-DAXX diffusion after photoconversion are indicated below (t1/2). Insets show a representative green to red photoconverted ROI surrounding NB or in the nucleoplasm at t = 2.5 s, t = 9.5 s, and t = 45.3 s after switch. Bar, 0.5 µm. (F) Immunolocalization of PMLΔSIM or PMLK160R stably expressed in pml−/− MEFs or CHO cells, and of endogenous DAXX or transfected SP100, as indicated. (G) Immunolocalization of DAXX mutants stably expressed in CHO-PML cells. Bars, 5 µm.
Partner recruitment/sequestration onto PML NBs are controlled by sequential and polarized SUMO–SIM interactions
Using FRAP analysis, we measured the exchange rate of the GFP fusions between NBs and the nucleoplasm. S3ΔGG-GFP was retained in NBs, and adding SIM to the S3ΔGG-GFP fusion enhanced its retention in NBs (Fig. 2 D). Similarly, a photo-switchable fusion between Dendra and DAXX (a well-known SUMO–SIM-containing NB partner) stably expressed in CHO-PML cells, diffused much faster in the nucleoplasm than in NBs, again evidencing a PML role in partner protein sequestration (Fig. 2 E). Our findings imply that SIM and SUMOs are NB-targeting/retention signals, explaining the NB association of SIM–SUMO-containing partners.
We then questioned how partner SIM or SUMO could interact with PML. Unexpectedly, NBs generated in CHO cells or pml−/− MEFs by PMLΔSIM stable expression efficiently recruited DAXX, RNF4, or SP100, whereas this was not observed with PMLK160R (Fig. 2 F; unpublished data; Ishov et al., 1999; Lallemand-Breitenbach et al., 2001). Note that PMLΔSIM was sumoylated and degraded upon arsenic exposure, corroborating the efficient recruitment of RNF4 (Fig. S1 C). Thus, PML SIM is unessential for partner recruitment.
Examining the partner side, multiple studies have demonstrated the requirement of SIM for the recruitment of some specific proteins in NBs and for their sumoylation (Lin et al., 2006). Indeed, deletion of DAXX C-terminal SIM abrogated its association to NBs (Fig. 2 G and Fig. S2; Lin et al., 2006). Although irreversible fusion of SUMO1 or -2 clearly enhanced DAXX association to NBs, this was insufficient to rescue DAXXΔSIM NB localization (Fig. 2 G). These data demonstrate that SIM and SUMO are not equivalent in promoting partner NB recruitment. Collectively, these data argue against the previously proposed role of PML SIM in either NB morphogenesis or partner recruitment. They strongly suggest that SUMO–SIM interactions are polarized and sequential. PML sumoylation on K160 recruits partners through their SIM. Partners undergo sumoylation and are then sequestered through interactions with partner, or PML, SIMs.
NBs consist of a matrix PML shell and a nonmatrix SUMO–SIM core
Our findings suggest that NBs could consist of two compartments: one, formed by a mesh of covalent PML multimers and associated to the NM; the other, formed by partner proteins, relying on multiple weak SUMO–SIM interactions. Yet, the latter are unlikely to secure partner association to the NM. To test this, in situ NM preparations were labeled with antibodies against PML or partners. Intense PML and SUMO2/3 staining was observed in NM preparations, whereas SUMO1, RNF4, and DAXX staining was dramatically diminished when compared with whole-cell preparations (Fig. 3 A).
Topologically and biochemically distinct compartments in NBs. (A) In situ NM preparations from MRC5 cells, showing the matrix association of endogenous PML/SUMO2/3, in contrast to endogenous DAXX, SUMO1, or transfected FLAG-RNF4. Control, total cell labeling; nuclear matrix, in situ NM preparation; double PML/partner protein (top) and partner only (bottom) labeling are shown. Bar, 5 µm. (B) Confocal (top bar, 5 µm) and immunoelectron microscopy analysis (bottom bar, 0.5 µm) of stably transfected CHO-SP100 or CHO-SP100/PML cells. In the dual PML/SP100 labeling (far right), PML is revealed by large and SP100 by smaller gold particles. (C) Deconvoluted confocal analysis of IFNα-treated MRC5 primary fibroblasts stained with anti-PML (red), anti-SUMO1, anti-SUMO-2/3, anti-SP100, or anti-DAXX (green) antibodies. Bar, 1 µm.
Topologically and biochemically distinct compartments in NBs. (A) In situ NM preparations from MRC5 cells, showing the matrix association of endogenous PML/SUMO2/3, in contrast to endogenous DAXX, SUMO1, or transfected FLAG-RNF4. Control, total cell labeling; nuclear matrix, in situ NM preparation; double PML/partner protein (top) and partner only (bottom) labeling are shown. Bar, 5 µm. (B) Confocal (top bar, 5 µm) and immunoelectron microscopy analysis (bottom bar, 0.5 µm) of stably transfected CHO-SP100 or CHO-SP100/PML cells. In the dual PML/SP100 labeling (far right), PML is revealed by large and SP100 by smaller gold particles. (C) Deconvoluted confocal analysis of IFNα-treated MRC5 primary fibroblasts stained with anti-PML (red), anti-SUMO1, anti-SUMO-2/3, anti-SP100, or anti-DAXX (green) antibodies. Bar, 1 µm.
To illustrate this biochemical difference at the level of localization, PML was stably expressed in CHO cells stably expressing SP100 (CHO-SP100). This led to the recruitment of diffuse SP100 into typical NBs (Fig. 3 B, top). A distinct PML-labeled shell was observed while SP100 staining concentrated within the central micro-granular core in electron microscopy (Fig. 3 B, bottom). Similarly, in transformed (SaOS) and primary (MRC5) cells, endogenous SP100, DAXX, SUMO1, and SUMO2/3, as well as ectopically expressed RNF4, were all found in this central core, and only SUMO2/3 was present together with PML on the outer shell (Fig. 3 C; unpublished data). Absence of SUMO1 on the external shell contrasts with a previous report and may be due to different cell types used (Lang et al., 2010). Collectively, in keeping with the super-resolution images, these data demonstrate that NBs consist of a matrix-associated peripheral shell, containing oxidized and SUMO-conjugated PML, and an inner nonmatrix core that accumulates partners through SUMO–SIM interactions.
PML recruits UBC9 into NBs favoring ROS-enhanced sumoylation
We found that UBC9 was concentrated into NBs (Fig. 4 A), consistent with reports showing that UBC9 may bind PML in vitro (Duprez et al., 1999). UBC9 NB association was observed in H1299 cells for endogenous proteins (Fig. 4 A, left), as well as in CHO cells stably expressing both UBC9-GFP and PML (Fig. 4 A, right). UBC9, which harbors a SIM and may be sumoylated (Knipscheer et al., 2008), was associated with the soluble inner fraction of the bodies (Fig. S3, A and B).
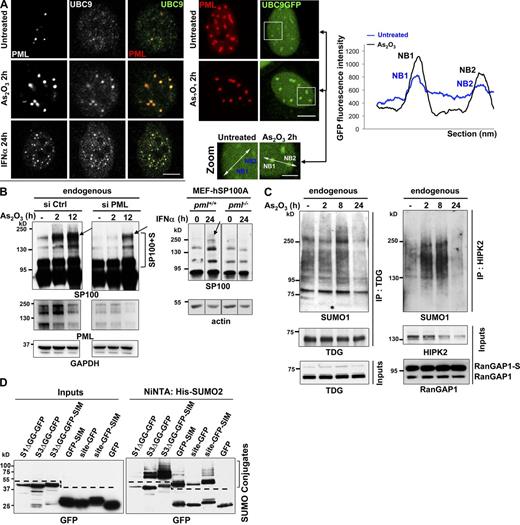
UBC9 recruitment into NBs favors hyper-sumoylation of partner proteins. (A, left) Confocal analysis of endogenous UBC9 localization in Triton X-100 pre-extracted H1299 cells, showing recruitment onto NBs after arsenic or IFNα exposure. Representative of two independent experiments, n ≥ 300 cells examined. Bar, 5 µm. (Right) Localization of UBC9-GFP stably expressed in CHO-PML cells before and after 2 h exposure to arsenic. Representative of three independent experiments, n ≥ 300 cells. Bar, 5 µm. Zooms show the regions (outlined above) used to quantify GFP fluorescence intensity (graph). Representative of three repeats. Bar, 2 µm. (B, left) Western blot analysis of endogenous SP100 profiles, performed on protein extracted 48 h after transfection with PML or control siRNAs. (Right) Sumoylation of hSP100A stably expressed in pml+/+ or pml−/− MEFs treated or not with IFNα for 24 h. Arrows and the bracket point to sumoylated species (see also Fig. S4, A, B, and D). (C) Western blot analysis of endogenous TDG (left) or HIPK2 (right) immunoprecipitated (IP) from H1299 cells treated as indicated. Inputs are shown. RanGAP1-S, sumoylated RanGAP1. (D) SUMO-GFP or SIM-GFP fusions are sumoylated. The indicated GFP fusions and (His)x6-SUMO2 were overexpressed in CHO-PML cells, purified over a Ni-NTA column and analyzed by Western blot using anti-GFP antibodies. Left, inputs; right, denaturing purification of sumoylated proteins. Dotted lines show sumoylated GFP fusions specifically purified from unspecific binding of GFP to the Ni-NTA column.
UBC9 recruitment into NBs favors hyper-sumoylation of partner proteins. (A, left) Confocal analysis of endogenous UBC9 localization in Triton X-100 pre-extracted H1299 cells, showing recruitment onto NBs after arsenic or IFNα exposure. Representative of two independent experiments, n ≥ 300 cells examined. Bar, 5 µm. (Right) Localization of UBC9-GFP stably expressed in CHO-PML cells before and after 2 h exposure to arsenic. Representative of three independent experiments, n ≥ 300 cells. Bar, 5 µm. Zooms show the regions (outlined above) used to quantify GFP fluorescence intensity (graph). Representative of three repeats. Bar, 2 µm. (B, left) Western blot analysis of endogenous SP100 profiles, performed on protein extracted 48 h after transfection with PML or control siRNAs. (Right) Sumoylation of hSP100A stably expressed in pml+/+ or pml−/− MEFs treated or not with IFNα for 24 h. Arrows and the bracket point to sumoylated species (see also Fig. S4, A, B, and D). (C) Western blot analysis of endogenous TDG (left) or HIPK2 (right) immunoprecipitated (IP) from H1299 cells treated as indicated. Inputs are shown. RanGAP1-S, sumoylated RanGAP1. (D) SUMO-GFP or SIM-GFP fusions are sumoylated. The indicated GFP fusions and (His)x6-SUMO2 were overexpressed in CHO-PML cells, purified over a Ni-NTA column and analyzed by Western blot using anti-GFP antibodies. Left, inputs; right, denaturing purification of sumoylated proteins. Dotted lines show sumoylated GFP fusions specifically purified from unspecific binding of GFP to the Ni-NTA column.
Arsenic, which induces oxidative stress, increased UBC9 recruitment into NBs (Fig. 4 A and Fig. S3 C), and actually depleted UBC9 from the nucleoplasm in both H1299 and CHO-PML cells (Fig. 4 A). Interferon (IFNα), used to boost endogenous PML expression and NB formation (Stadler et al., 1995), enhanced UBC9 targeting to NBs in the same manner as arsenic (Fig. 4 A, bottom). These findings imply that PML recruits UBC9 to NBs and may thus specifically favor sumoylation of PML partner proteins in situ, enhancing their sequestration by SIM–SUMO interactions.
SP100 is one of the most studied PML partners, whose basal sumoylation was proposed to be influenced by PML (Cuchet et al., 2011; Fig. S4 A, left). Arsenic elicited hyper-sumoylation of endogenous SP100 in HeLa and H1299 cells, which was sharply delayed and reduced by PML silencing (Fig. 4 B, left; and Fig. S4, A and B). Proximity ligation assays, which allow in situ detection of closely interacting individual proteins, confirmed that endogenous interactions between SP100 and SUMO1 occurred mainly in NBs and increased with exposure to arsenic or IFNα (Fig. S4 C). This strongly suggests that SP100 sumoylation occurs in NBs. Similarly, exposure to IFNα increased SP100 sumoylation in a PML-dependent manner, as assessed in pml−/− or pml+/+ MEFs stably expressing SP100A isoform (MEF-hSP100A; Fig. 4 B, right) or in HeLa cells expressing endogenous SP100 (Fig. S4 D). In the same manner as for SP100, sumoylation of endogenous thymine DNA glycosylase (TDG), a NB-resident protein (Takahashi et al., 2005), was transiently increased by arsenic (Fig. 4 C, left). This was also the case for NB-associated kinase HIPK2 and for DAXX (Ishov et al., 1999; D’Orazi et al., 2002; Fig. 4 C, right; unpublished data), but not for RanGAP1 (Fig. 4 C, right), a protein not associated with NBs.
We finally assessed modifications of SIM- and SUMOΔGG-GFP fusions that are artificially recruited into NBs by performing His-purification from cells coexpressing (His)x6-SUMO2 together with SUMO3ΔGG-GFP, GFP-SIM, or the TDG sumoylation site fused to either GFP or GFP-SIM (site-GFP and site-GFP-SIM, respectively). All the GFP fusions were found to be sumoylated, whereas GFP itself, which is not targeted to NBs, did not undergo significant modifications (Fig. 4 D).
Collectively, we formally establish that PML NBs enhance in situ the sumoylation of partner proteins. Most importantly, it confers oxidative stress and interferon sensitivity to partner sumoylation through concentration of UBC9 and its substrates within NB cores.
RNF4 recruitment into NBs induces loss of conjugated partners
Upon exposure to arsenic, RNF4 is recruited into NBs together with proteasome subunits (Lallemand-Breitenbach et al., 2008). Proximity ligation assays, performed to better assess PML–RNF4 interactions, revealed that the interaction signals predominantly took place in NBs and were sharply enhanced upon exposure to arsenic, implying that RNF4 primarily interacts with NB-associated PML (Fig. 5 A).
RNF4 recruitment into NBs results in sumoylation decay through partner ubiquitination. (A, top) PML–RNF4 interactions detected by PLA Duolink assay (red dots) and PML NB immunolocalization (green); Z-stack projections are shown. (Bottom) Quantification of the Duolink dots per cell and percentages of colocalization with NBs (means from 20 cells). (B) Western blot analysis of endogenous SP100 hyper-sumoylation (bracket) upon exposure to arsenic in H1299 transfected with the indicated siRNAs. (C) Western blot analysis of transduced hSP100A in MEFs treated as indicated, demonstrating arsenic-induced proteasomal degradation after 24 h. (D) SUMO and ubiquitin conjugation of SP100 upon exposure to arsenic. (Left) Endogenous SP100 immunoprecipitates from arsenic-treated HeLa cells probed with anti-SUMO1 and anti-SP100 antibodies. (Right) Nickel-purified His-ubiquitin conjugates from His-ubiquitin–overexpressing HeLa cells, probed with antibodies to SP100 and ubiquitin. (E) Polyubiquitination of indicated GFP fusions in CHO-PML cells overexpressing (His)x6-Ubiquitin, purified over nickel column and analyzed by Western blot using anti-GFP antibodies. Left, inputs; right, denaturing purification of ubiquitinated proteins.
RNF4 recruitment into NBs results in sumoylation decay through partner ubiquitination. (A, top) PML–RNF4 interactions detected by PLA Duolink assay (red dots) and PML NB immunolocalization (green); Z-stack projections are shown. (Bottom) Quantification of the Duolink dots per cell and percentages of colocalization with NBs (means from 20 cells). (B) Western blot analysis of endogenous SP100 hyper-sumoylation (bracket) upon exposure to arsenic in H1299 transfected with the indicated siRNAs. (C) Western blot analysis of transduced hSP100A in MEFs treated as indicated, demonstrating arsenic-induced proteasomal degradation after 24 h. (D) SUMO and ubiquitin conjugation of SP100 upon exposure to arsenic. (Left) Endogenous SP100 immunoprecipitates from arsenic-treated HeLa cells probed with anti-SUMO1 and anti-SP100 antibodies. (Right) Nickel-purified His-ubiquitin conjugates from His-ubiquitin–overexpressing HeLa cells, probed with antibodies to SP100 and ubiquitin. (E) Polyubiquitination of indicated GFP fusions in CHO-PML cells overexpressing (His)x6-Ubiquitin, purified over nickel column and analyzed by Western blot using anti-GFP antibodies. Left, inputs; right, denaturing purification of ubiquitinated proteins.
When examining arsenic-induced SP100, TDG, or HIPK2 hyper-sumoylation, we noted loss of sumoylated species after 24 h exposure (Fig. 5, B and C; and Fig. 4 C). This loss was reversed by the proteasome inhibitor MG132, and also by RNF4 siRNA silencing (Fig. 5 B, arrows; and Fig. 5 C; unpublished data), suggesting that with time some sumoylated species were ubiquitinated and degraded. Endogenous SP100 was therefore immunoprecipitated and this demonstrated that arsenic enhanced both endogenous SUMO1 and polyubiquitin conjugation (Fig. 5 D). A fraction of SP100 species was dually conjugated by both SUMO and ubiquitin, as demonstrated by two-step purification (Fig. S4 E).
We similarly assessed the ubiquitination status of site-GFP-SIM, GFP-SIM, and SUMOΔGG-GFP fusions that are recruited in NBs and sumoylated. His-purifications from cells coexpressing (His)x6-Ubiquitin and SUMO3ΔGG-GFP, GFP-SIM, or site-GFP-SIM (see previous section) indicated that these GFP fusions were also ubiquitinated, but not GFP itself (Fig. 5 E).
PML-dependent biphasic response of global sumoylation to oxidative stress
We then analyzed global sumoylation in response to ROS induced by arsenic (Fig. S3 C; Kawata et al., 2007). Arsenic elicited transient formation of high molecular weight (HMW) SUMO conjugates, while protein species in the 50–100 kD range were marginally affected (Fig. 6 A). After more than 12 h exposure to arsenic, both HMW SUMO1 and SUMO2/3 conjugates declined (Fig. 6 A). MG132 proteasome inhibitor dampened this late arsenic-triggered loss of sumoylated proteins, showing that sumoylation decrease reflects the degradation of some hyper-sumoylated proteins rather than de-sumoylation (Fig. 6 A). In that respect, SENP3 and SENP5, which were proposed to traffic between nucleoli and NBs (Gong and Yeh, 2006), were not recruited to NBs upon arsenic exposure (unpublished data).
PML and RNF4 cooperate to regulate sumoylation in response to oxidative stress. (A) Western blot analysis of SUMO1 and -2/3 conjugates from H1299 cells treated as indicated with arsenic, and for 8 h with MG132. (B) Western blot analysis of arsenic-induced transient hyper-sumoylation in H1299 cells transfected with the indicated siRNAs. SUMO1 and -2/3 conjugates are shown along with PML, RNF4, and actin controls. Star, sumoylation of RanGAP1, a non–NB-associated protein, does not respond to arsenic; PML-S, sumoylated PML. (C) Western blot analysis of immunoprecipitated polyubiquitin conjugates (IP: Ub) from pml+/+ or pml−/− MEFs treated or not with arsenic and IFNα for 48 h. (D) Western blot analysis of sumoylation after transduction of RNF4 (R) or catalytically inactive RNF4-DN mutant in IFNα-treated pml+/+ or pml−/− MEFs. Brackets, HMW SUMO1 conjugates; asterisk, sumoylated RanGAP1. (E) Western blot analysis of global sumoylation in immortalized pml+/+ or pml−/− MEFs.
PML and RNF4 cooperate to regulate sumoylation in response to oxidative stress. (A) Western blot analysis of SUMO1 and -2/3 conjugates from H1299 cells treated as indicated with arsenic, and for 8 h with MG132. (B) Western blot analysis of arsenic-induced transient hyper-sumoylation in H1299 cells transfected with the indicated siRNAs. SUMO1 and -2/3 conjugates are shown along with PML, RNF4, and actin controls. Star, sumoylation of RanGAP1, a non–NB-associated protein, does not respond to arsenic; PML-S, sumoylated PML. (C) Western blot analysis of immunoprecipitated polyubiquitin conjugates (IP: Ub) from pml+/+ or pml−/− MEFs treated or not with arsenic and IFNα for 48 h. (D) Western blot analysis of sumoylation after transduction of RNF4 (R) or catalytically inactive RNF4-DN mutant in IFNα-treated pml+/+ or pml−/− MEFs. Brackets, HMW SUMO1 conjugates; asterisk, sumoylated RanGAP1. (E) Western blot analysis of global sumoylation in immortalized pml+/+ or pml−/− MEFs.
Arsenic’s biphasic effect on global sumoylation again depended on both PML and RNF4 (Fig. 6 B). Indeed, PML silencing abrogated the initial arsenic-enhanced conjugation, whereas RNF4 extinction affected only the late degradation phase. HMW SUMO conjugates did not co-migrate with SUMO-modified PML and there was a delay between PML degradation and sumoylated protein decrease (Fig. 6 B; and not depicted), implying that arsenic-induced HMW species do not correspond solely to PML conjugates. Moreover, arsenic-induced degradation of sumoylated proteins was confirmed to be PML dependent in MEFs isolated from pml+/+ and pml−/− animals (Fig. S5 A).
MEFs were then treated with both arsenic and IFNα to enhance NB formation. Polyubiquitinated conjugates were immunoprecipitated and probed with antibodies to SUMO1, SUMO2/3, or PML. Formation of mixed ubiquitin–SUMO2/3 conjugates, a read-out for RNF4 activity, was strongly induced in pml+/+ but not in pml−/− MEFs (Fig. 6 C), suggesting that RNF4 required PML to efficiently ubiquitinate sumoylated proteins, at least in this setting of oxidative stress.
We then investigated whether overexpressing RNF4 could lead to the clearance of SUMO conjugates in a pml-dependent manner. RNF4 or a dominant-negative mutant (RNF4-DN) thereof was transduced in pml+/+ or pml−/− MEFs treated with IFNα to increase the differences in PML content. RNF4 overexpression markedly decreased the amounts of HMW SUMO1 or SUMO2/3 conjugates in pml+/+ MEFs only (Fig. 6 D). In contrast, these conjugates accumulated in RNF4-DN–transduced cells, also in a pml-dependent manner (Fig. 6 D). SUMO1 conjugation to RanGAP1 was unaffected (Fig. 6 D, star; also see star in Fig. 6 B). Similar effects, albeit of smaller magnitude, were noted in the absence of IFNα (unpublished data). Interestingly, in the absence of treatment, pml−/− MEFs reproducibly exhibited higher levels of HMW SUMO1 and 2/3 conjugates than pml+/+ MEFs (Fig. 6 E, also see untransfected cell lanes in Fig. 6 D). Accumulation of SUMO conjugates in the absence of PML was also noted in primary tissues such as bone marrow (Fig. S5 B). Collectively, these data demonstrate that upon oxidative stress PML NB formation enhances first the sumoylation of partner proteins and subsequently the degradation of some of these, revealing NBs as global sumoylation control machineries.
Oxidation triggers NB formation and sumoylation in vivo and in APL cells
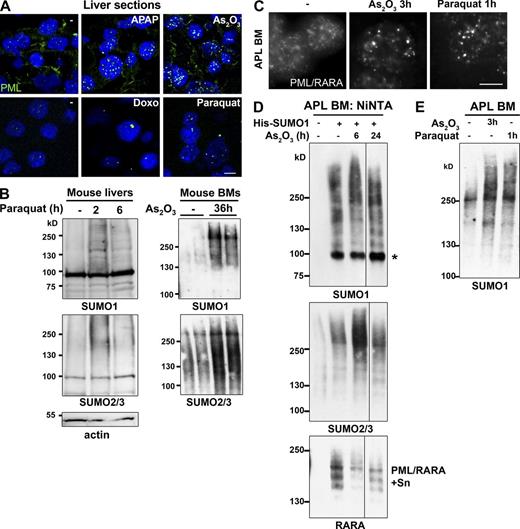
To address the role of oxidation in a physiological setting, we analyzed NB formation in vivo in conditions of oxidative stress. For this we examined livers from mice after a short (1–2 h) exposure to acetaminophen, doxorubicin, gamma irradiation, or paraquat (Kawata et al., 2007). Remarkably, the size and number of NBs were dramatically increased (Fig. 7 A and Fig. S5 C). Whereas doxorubicin and paraquat led to formation of much larger NBs, acetaminophen and irradiation led to a dramatic increase in the number of NBs (Fig. S5 C and not depicted). This was not associated with increased amounts of PML (unpublished data), except a twofold increase for doxorubicin, which activates pml expression through P53 (de Stanchina et al., 2004). Collectively, these data support the direct role of PML oxidation in NB nucleation and are consistent with previous pathology studies showing an increased amount of NBs in inflammatory tissues (Koken et al., 1995; Terris et al., 1995). Importantly, in mouse liver or bone marrow, in vivo treatments with arsenic or paraquat also led to a transient boost of sumoylation (Fig. 7 B).
Induction of NB formation and sumoylation in oxidant-treated mice. (A) In vivo oxidative stress effects on NB formation: Immunofluorescence analysis of PML NBs on liver sections from mice exposed to APAP (acetaminophen: N-acetyl-p-aminophenol), As2O3, doxorubicin (Doxo), or paraquat for 2 h. Quantifications are shown in Fig. S5 C. n ≥ 50 cells examined. (B) In vivo sumoylation after exposure to oxidative stress. (Left) Western blot analysis of liver cells from mice treated with paraquat. (Right) Western blot analysis of bone marrow (BM) cells from mice treated with arsenic; two mice are shown for each from two independent experiments. (C) Immunofluorescence analysis of APL cells obtained after in vivo administration of arsenic or paraquat for the indicated period of time. Bar, 5 µm. (D) His-SUMO1–transduced mouse APL bone marrow (BM) cells were isolated after in vivo arsenic administration. His-conjugates were purified and probed with antibodies to SUMO1, SUMO2/3, and RARA. PML/RARA-S, poly-sumoylated PML/RARA. Representative data from six mice, three independent experiments. (E) Western blot analysis of bone marrow cells of APL mice after in vivo treatment with arsenic or paraquat.
Induction of NB formation and sumoylation in oxidant-treated mice. (A) In vivo oxidative stress effects on NB formation: Immunofluorescence analysis of PML NBs on liver sections from mice exposed to APAP (acetaminophen: N-acetyl-p-aminophenol), As2O3, doxorubicin (Doxo), or paraquat for 2 h. Quantifications are shown in Fig. S5 C. n ≥ 50 cells examined. (B) In vivo sumoylation after exposure to oxidative stress. (Left) Western blot analysis of liver cells from mice treated with paraquat. (Right) Western blot analysis of bone marrow (BM) cells from mice treated with arsenic; two mice are shown for each from two independent experiments. (C) Immunofluorescence analysis of APL cells obtained after in vivo administration of arsenic or paraquat for the indicated period of time. Bar, 5 µm. (D) His-SUMO1–transduced mouse APL bone marrow (BM) cells were isolated after in vivo arsenic administration. His-conjugates were purified and probed with antibodies to SUMO1, SUMO2/3, and RARA. PML/RARA-S, poly-sumoylated PML/RARA. Representative data from six mice, three independent experiments. (E) Western blot analysis of bone marrow cells of APL mice after in vivo treatment with arsenic or paraquat.
In APL, PML NBs are disrupted in a treatment-reversible manner (de Thé and Chen, 2010; Lallemand-Breitenbach et al., 2012). We thus examined whether NB reformation by arsenic or paraquat would change the global sumoylation profile of APL cells (Fig. 7, C–E). Leukemic cells from APL mice were transduced or not with His-SUMO1 and transplanted into healthy mice, which were then treated after leukemia development. After short in vivo exposure to arsenic, APL cells displayed a transient increase in HMW SUMO1 and mixed SUMO1/2/3 conjugates (Fig. 7 D), concomitant with NB reformation (Fig. 7 C). Similarly, paraquat treatment in vivo induced both NB reformation and SUMO1 conjugation (Fig. 7, E and C). Rapid arsenic-induced degradation of sumoylated PML/RARA confirmed that the fusion did not account for the increase in global SUMO conjugation (Fig. 7 D). Thus, in vivo NB reformation is accompanied by enhanced global sumoylation, raising the issue that this may contribute to the efficacy of arsenic or retinoic acid therapy to cure APL.
Discussion
This study presents an integrated view of the sequential steps of NB biogenesis and is the first proposal for an implication of NBs in global oxidative stress–induced partner sumoylation and degradation.
Whereas previous studies had noted the requirement of the SIM for sumoylation (Takahashi et al., 2005; Knipscheer et al., 2008; Meulmeester et al., 2008) or for NB association of specific proteins (DAXX, HIKP2, Topors) (Weger et al., 2003; Lin et al., 2006; Sung et al., 2011), we provide evidence for a mechanism of NB biogenesis involving two distinct steps: an initial oxidation-sensitive PML multimerization and subsequently, a polarized SIM–SUMO-dependent recruitment of partner proteins. Indeed, analysis of PML3KRΔSIM demonstrates that SUMO–SIM interactions are not responsible, as previously thought (Müller et al., 1998; Shen et al., 2006), for the initial nucleation of the PML spherical mesh (Fig. 1). The latter is triggered by a complex polymerization scheme involving PML coiled-coil and disulfide bonds (Jeanne et al., 2010). Our in vivo experiments demonstrate that NB formation primarily mirrors cellular oxidative stress (Fig. 7), explaining NB prevalence in diseased, but not normal, tissues (Daniel et al., 1993; Koken et al., 1995; Gambacorta et al., 1996). UBC9 binds PML (Duprez et al., 1999), accumulates in NBs (Fig. 4), and enhances PML sumoylation in trans on the PML mesh (Jeanne et al., 2010). This sumoylated PML lattice then becomes a docking site for the dynamic association of partners through multiple labile SUMO–SIM interactions (Fig. 8), in line with the seeding model proposed for other nuclear domains (Rajendra et al., 2010). The initial SIM–SUMO interaction is polarized as the SIM anchors partners to the SUMO of PML K160, and in situ sumoylation then enforces partner retention within the NB core (Fig. 2). PML SIM is dispensable for partner recruitment and the fusion of SUMO to DAXXΔSIM fails to rescue NB association. This rules out the alternative model whereby partner SIM would recruit UBC9, triggering partner sumoylation in the nucleoplasm and subsequent recruitment onto NBs, partner SUMO binding PML SIM. Collectively, distribution of partner proteins within the nucleus will primarily depend on PML level as well as on its self-assembly and sumoylation, both reflecting redox status.
NBs are redox-regulated hubs of sumoylation and SUMO-initiated, RNF4-mediated ubiquitination. Nucleation of the NB mesh relies solely on ROS-induced PML oxidation, while intermolecular PML SUMO–SIM interactions are not involved in this initial nucleation step. Stress-induced UBC9 recruitment results in subsequent PML sumoylation, critical for partners’ recruitment through their SIM. In situ sumoylation of partners secures partner–PML, partner–partner, and partner–RNF4 interactions in NBs, promoting their ubiquitination or other post-translational modifications (PTMs) like acetylation or phosphorylation.
NBs are redox-regulated hubs of sumoylation and SUMO-initiated, RNF4-mediated ubiquitination. Nucleation of the NB mesh relies solely on ROS-induced PML oxidation, while intermolecular PML SUMO–SIM interactions are not involved in this initial nucleation step. Stress-induced UBC9 recruitment results in subsequent PML sumoylation, critical for partners’ recruitment through their SIM. In situ sumoylation of partners secures partner–PML, partner–partner, and partner–RNF4 interactions in NBs, promoting their ubiquitination or other post-translational modifications (PTMs) like acetylation or phosphorylation.
Consistent with the recruitment of UBC9 within NBs, we demonstrate that PML aggregation upon oxidative stress enhances global sumoylation (Figs. 6 and 7), most likely of NB partners. Overexpression of TRIM proteins was suggested to modulate sumoylation, notably in yeast (Quimby et al., 2006; Chu and Yang, 2011). Concentration of targets and enzymes or different components of protein–RNA complexes constitute a common feature of nuclear bodies (Cajal bodies, nuclear speckles…), which enables chemical reactions or complex formation between low-abundance nuclear species (Rajendra et al., 2010). That NBs constitute stress-responsive sumoylation factories may define a new class of E3 ligase acting through physical concentration, as suggested for the nuclear pore (Fig. 8; Zhang et al., 2002; Melchior et al., 2003; Nagai et al., 2011). It is possible that SUMO E3 ligases are also recruited into NBs (Rabellino et al., 2012), and that NB inner core provides a favorable redox environment for thiol enzymes involved in the sumoylation cascade (Bossis and Melchior, 2006). Recent studies have similarly demonstrated that concentration of viral genomes and transactivators into PML NBs contributes to viral fitness (Teng et al., 2012). The wave of sumoylation induced by PML NBs upon oxidative stress resembles the one occurring after DNA damage, in which SIMs orchestrate the coordinated sumoylation of DNA repair proteins (Psakhye and Jentsch, 2012). Interestingly, many of these proteins may be observed in PML NBs (Lallemand-Breitenbach and de Thé, 2010), and PML or RNF4 absence is associated with persistent DNA damage (Zhong et al., 1999; Galanty et al., 2012; Yin et al., 2012), both supporting a role of PML in sumoylation/degradation of DNA repair proteins.
Several NB partners are protein-modifying enzymes (RNF4, HIPK2, TDG, SIRT1, CBP, MDM2, HAUSP…), suggesting that partner NB association, through transient SIM–SUMO-mediated partner sequestration, could confer oxidative stress sensitivity to other post-translational modifications (Langley et al., 2002; Seet et al., 2006; Trotman et al., 2006; Kirkin and Dikic, 2007; Song et al., 2008; Regad et al., 2009; de la Vega et al., 2012; de Thé et al., 2012). In that respect, P53 is a downstream effector of PML-regulated senescence (Bischof et al., 2002; Chiantore et al., 2012), which together with many of its modifying enzymes, may be NB associated. This suggests that PML NB aggregation may confer redox sensitivity to P53 signaling. Supporting the hypothesis that NB aggregation facilitates other post-translational modifications through partner sumoylation, we demonstrate that PML is required for ubiquitination of many sumoylated proteins by the RNF4 ubiquitin ligase. Interestingly, UBC9 and RNF4 do not have the same requirements for NB association, as UBC9 directly binds to PML (Duprez et al., 1999), whereas RNF4 requires PML poly-sumoylation (Lallemand-Breitenbach et al., 2008). This likely contributes to slow kinetics of the degradation phase in arsenic-treated cells (Fig. 6 A), as opposed to immediate hyper-sumoylation upon NB formation. Degradation of PML upon longer arsenic exposure likely explains why new SUMO conjugates fail to replace those that are degraded. Increased SUMO levels in pml-deficient cells suggest that RNF4-mediated degradation quantitatively exceeds conjugation at a steady state (Fig. 6 E and Fig. S5 B). RNF4 recruitment onto a given target may be influenced by the type and number of SUMO paralogues, thus defining a “SUMO code” for subsequent degradation (Fig. 8).
Our studies not only clarify the sequential interactions involved in NB biogenesis, but also identify NBs as pivotal domains regulating stress-induced sumoylation, explaining defects in redox-sensitive pathways (NF-κB, PTEN, HIF1A…) in cells with disrupted NBs (Lallemand-Breitenbach and de Thé, 2010). These findings identify changes in nuclear architecture (i.e., NB biogenesis) as a key link between oxidative stress and a subsequent defined biochemical response. Our in vivo observations underscore the relevance of this response pathway in physiological or pathological conditions. Remarkably, two key PML-regulated pathways, senescence and defense against microbes, are also activated by hyper-sumoylation (Yates et al., 2008; Ribet et al., 2010; Everett et al., 2013), fully in line with our model. Future studies should thus assess the contribution of deregulated sumoylation to other PML-dependent phenotypes.
Materials and methods
Constructs and siRNAs
PMLΔSIM, PML3KRΔSIM mutants and GFP-SIMPML were constructed by deletion or insertion of the PML SIM coding sequence (aa 556–566) with the QuikChange II site-directed mutagenesis kit (QIAGEN) into MSCV-PML, MSCV-PML3KR, or pEGFP-N1. MSCV retroviral vectors for (His)x6-tagged PML and PML3KR mutant were described previously and used for stable expression (Lallemand-Breitenbach et al., 2008; Jeanne et al., 2010). HA-tagged versions of PML and mutants were used for Western blot analyses of NMs prepared from transduced pml−/− MEFs. HA tag was fused in frame with PML N terminus. PMLΔCC corresponds to the deletion of the coiled-coil motif (aa 216–333). Unless otherwise indicated, PML refers to the PML-III isoform. SUMO1, 2, and 3 cDNAs, deleted from the 5′ diglycine coding sequence, were amplified by PCR and cloned in the BglII–HindIII restriction sites of pEGFP-N1. Fusions of the DAXX or HIPK2 SIM (aa 730–741 and 857–871, respectively; Lin et al., 2006; Sung et al., 2011) to GFP (pEGFP-C1-SIMDaxx or HipK2) and deletion from the pSG5-DAXX were similarly constructed. TDG sumoylation site (aa 325–335; Takahashi et al., 2005) was inserted in pEGFP-SIM(PML) by mutagenesis. pDendra2 vector (Takara Bio Inc.) was used to clone DAXX coding sequence in-frame with the 5′ dendra gene. siRNAs against SUMOs (QIAGEN) were described previously: SUMO-1 (5′-GGACAGGAUAGCAGUGAGA-3′), SUMO-2 (5′-AGGGAUGAAUCUGUAACUUAA-3′), SUMO-3 (5′-GAGGCAUACACCACUUAGU-3′; Lallemand-Breitenbach et al., 2008). siRNAs against hRNF4 (5′-CCCUGUUUCCUAAGAACGAAA-3′) and siRNA against hPML (5′-AAGAGTCGGCCGACTTCTGGT-3′) transcripts were purchased from QIAGEN.
Antibodies, immunoblots, and immunofluorescence
Anti–human PML, anti–mouse PML, anti-SUMO1, SUMO2/3, anti-ubiquitin (FK2), and anti-RNF4 antibodies were reported previously (Lallemand-Breitenbach et al., 2008; Jeanne et al., 2010). Homemade rabbit or chicken polyclonal anti-SP100 antibodies were raised against full-length hSP100A and recognize all human isoforms (chicken anti-SP100 antibody used at 1:500 for immunofluorescence [IF] and rabbit anti-SP100 antibody at 1:2,000 for Western blot analysis). The polyclonal goat anti–human HIPK2 antibody (used at 1:1,000 for Western blot analysis) was from Santa Cruz Biotechnology, Inc., polyclonal rabbit anti–human UBC9 antibody (used at 1:500) was from Cell Signaling Technology; polyclonal rabbit anti–human TDG antibody (used at 1:1,000) was from Proteintech; polyclonal rabbit anti–human RXRA antibody (used at 1:1,000) was from Santa Cruz Biotechnology, Inc.; and anti-RanGAP1 was provided by F. Melchior (Universität Heidelberg, Heidelberg, Germany). Monoclonal mouse anti-HA antibody (Covance) was used at 1:2,000 for Western blot and 1:1,000 for IF. All antibodies were revealed by Alexa Fluor 488– or 594–labeled secondary antibodies (1:500) from Life Technologies or HRP-conjugated secondary antibodies (1:20,000) from The Jackson Laboratory. IF assays were performed as reported previously (Lallemand-Breitenbach et al., 2008; Jeanne et al., 2010). SUMO conjugates were separated on 4–12% gradient gels (Invitrogen) or homemade 7% or 10% SDS-PAGE. For analysis under nonreducing conditions, β-mercaptoethanol or DTT was omitted from the standard Laemmli buffer.
Microscopy image acquisition
Electron microscopy was performed as described previously (Lallemand-Breitenbach et al., 2001), with cells fixed in 1.6% glutaraldehyde or 4% paraformaldehyde for 30 min, which were then extensively washed in 0.1 M phosphate Sörensen buffer (pH 7.2–7.3). The fixed material was embedded in Lowicryl K4M. In situ NM preparations were prepared as described previously (Stuurman et al., 1990) and used for IF or resuspended by boiling in Laemmli buffer (NM fraction) for Western blot analysis. In brief, cells grown on coverslips or Lab-Tek II chambers (Thermo Fisher Scientific) were permeabilized in Kern matrix buffer (KMB) containing 1% NP-40. Nucleic acids were digested with 50 × 10−6 g/ml of RNase A and 0.2 × 10−6 g/ml of DNase I, or using 10−6 g/ml of micrococcal nuclease. A subsequent high-salt extraction was performed (2 M NaCl). After washes, the remaining insoluble proteins were fixed using 4% PFA.
Confocal analyses were performed on a confocal microscope (LSM510 Meta; Carl Zeiss) with a Plan-Apochromat 63×/1.4 NA oil-immersion objective at 20°C. Alexa Fluor 488, 594, 648 dye conjugates were used together with Vectashield mounting media containing DAPI (Vector Laboratories). Images were acquired with LSM510 software (Carl Zeiss). Increased resolution images were obtained with deconvolution software (AutoDeblur; ImageQuant) using standard blind iterative algorithms.
3D structured illumination microscope image acquisition
Super-resolution structured illumination microscopy was performed on a structured illumination microscope (ELYRA S.1; Carl Zeiss). A structured illumination pattern of the 561-nm diode laser was projected into the sample. The microscope was equipped with a Plan-Apochromat 100×/1.46 NA oil-immersion objective (Carl Zeiss), and we used immersion oil of refractive index 1.515. Emitted fluorescence light was directed through an appropriate dichroic beam splitter and emission filter onto an EMCCD camera (iXon 885; Andor Technology). Five translations and three rotations of the illumination pattern were recorded at each z-slice. 3D image stacks (120-nm increment along z axis) of typically 6-µm height were acquired and computationally reconstructed to generate super-resolution optical serial sections with twofold extended resolution in the three axes (lateral resolution 110 nm and axial resolution 280 nm). Image reconstruction and post-processing were performed with the ZEN imaging software package (Carl Zeiss) using the algorithm of Heintzmann and Cremer.
FRAP and photoconversion
FRAP was performed using a confocal microscope (LSM510 Meta; Carl Zeiss). For SUMOΔGG-GFP and GFP-SIM fusions, FRAP was performed on 2-µm regions of interest (ROIs) surrounding NBs using the 480-nm laser, at zoom 20, maximum speed scan (0.05 s between acquisitions). 10 images were acquired before bleach, 3 iterations were used to bleach, and time of recovery was between 15 and 30 s. Dendra-DAXX was photo-converted using a confocal microscope (LSM510 Meta; Carl Zeiss). Dendra-DAXX localization was followed using a 488-nm laser and ROI corresponding to Dendra-DAXX associated with NBs was then exposed to 405-nm laser for 2 s at zoom 20. After conversion, acquisitions were then performed at both 488 and 594 nm.
Cell culture, transfections, and treatments
Cells, cell cultures, treatments, and siRNA silencing have been described previously (Lallemand-Breitenbach et al., 2008; Jeanne et al., 2010). In brief, HeLa, H1299, MRC5, CHO, COS, SaOS cells, and pml−/− and pml+/+ MEFs were cultured in 10% FCS DMEM medium (Gibco). SV40 T-immortalized pml+/+ and pml−/− MEFs were obtained from pools of several embryos from two mice. As2O3 (Fluka) was used at 10−6 M for indicated times. Proteasome inhibitor MG132 (EMD Millipore) was used at 10−5 M, and IFNα (Roche) at 1,000 IU/ml.
siRNAs were transfected using HiPerFect reagent (QIAGEN) and samples were analyzed 48 or 72 h after transfection. Plasmid transfections were performed with the Effectene transfection reagent (QIAGEN). Stable CHO cell lines, expressing PMLIII isoform, was obtained by cotransfection of pSG5-PML with the Dsp-Hygro vector followed by selection for 2 wk in 800 µg/ml hygromycin-containing media. pml−/− MEFs were similarly transduced with MSCV-PML or PML mutants using viruses produced as described previously (Lallemand-Breitenbach et al., 2008). The cDNA encoding human SP100A isoform was also inserted in a MSCV retroviral vector, used to produce virus and to infect immortalized pml+/+ and pml−/− MEFs. Viruses encoding RNF4 or the RNF4-DN dominant-negative mutant have been described previously (Lallemand-Breitenbach et al., 2008). Cellular ROS levels were quantified by CellROX Deep Red reagent (Life Technologies). Probes were incubated for 20 min after arsenic exposure and quantified by FACS according to manufacturer’s guidelines.
Immunoprecipitations
Cells were washed in ice-cold PBS supplemented with 10 mM N-ethylmaleimide before lysis in 2% SDS and 50 mM Tris, pH 8. After brief sonication, cell lysates were diluted 10-fold in immunoprecipitation (IP) buffer containing 50 mM Tris, pH 8, 200 mM NaCl, 0.1 mM EDTA, 0.5% NP-40, 10% glycerol, and protease inhibitors. Lysates were incubated for 2 h at 4°C with the appropriate antibody, followed by incubation with protein A–agarose for 2 h. Beads were washed three times in the IP buffer before elution of immunoprecipitated proteins in sample buffer. For detection of His-tagged ubiquitin conjugates of endogenous SP100, His-tagged protein purification on Ni-NTA resin (QIAGEN or Invitrogen) was performed 24 h after transfection with His-ubiquitin or His-SUMO1– or 2–encoding vector as described previously (Lallemand-Breitenbach et al., 2008). Cells were lysed in denaturing buffer (6 M guanidium-HCl, 0.1 M NaH2PO4/Na2HPO4, and 10 mM imidazole, pH 8) and lysates were incubated with Ni-NTA resin (QIAGEN or Invitrogen) for 2 h. Three subsequent washes were performed with decreasing amounts of guanidium-HCl before elution in Laemmli buffer with 200 mM imidazole.
In vivo treatments
Experiments were performed in accordance with the French guidelines of institutional animal care committees, using protocols approved by the Comité Régional d’Ethique (protocol no. 4). The mouse APL model was described previously (Lallemand-Breitenbach et al., 1999). FVB-Nico mice were transplanted with leukemic blasts obtained from leukemic hMRP8-PML/RARA transgenic mice. Transplanted mice were treated with 150 mg/kg paraquat (paraquat dichloride, PESTANAL; Sigma-Aldrich), 5 mg/kg arsenic, or 300 mg/kg APAP (acetaminophen [N-acetyl-p-aminophenol]) through peritoneal injection, or with 20 mg/kg doxorubicin (Accord) through i.v. injections. Animals were sacrificed by cervical dislocation, organs were collected, and protein extracts analyzed by direct Western blot or after purification on Ni-NTA resin.
Proximity ligation assays
Duolink assays (Olink Bioscience) allowed the detection of individual proteins in close interaction (<40 nm), and were based on the in situ proximity ligation of DNA linked to secondary antibodies followed by PCR amplification. Assays were performed using anti-SP100 and -SUMO1, anti-SP100 and -SUMO2/3, anti-SP100 and -UBC9, anti-UBC9 and -PML, and anti-RNF4 and -PML primary antibodies.
Online supplemental material
Fig. S1 shows statistics on NB formation by PML mutants and upon SUMO siRNA knockdown, as well as efficient degradation of PML mutants upon arsenic exposure and controls for silencing efficiency. Fig. S2 shows SIMs and SUMO-conjugating sites on NB partners, as referenced in the supplemental references. Fig. S3 shows UBC9GFP localization in soluble NB inner core and arsenic-induced ROS production. Fig. S4 shows in situ NB-facilitated sumoylation and ubiquitination of SP100 induced by arsenic. Fig. S5 shows PML-dependent loss of high molecular weight SUMO conjugates ex vivo and in vivo, as well as quantification of NB formation in vivo in response to oxidative stress-inducing agents. Supplemental references list publications showing NB association of proteins with functional SIMs and SUMO-conjugating sites, as schematically represented in Fig. S2. Additional data are available in the JCB DataViewer at http://dx.doi.org/10.1083/jcb.201305148.dv.
Acknowledgments
We thank F. and E. Puvion for their help with electron microscopy. We thank P.P. Pandolfi for the gift of pml+/+ and pml−/− mice; M.L. Schmitz for HIPK2 expression vectors; F. Melchior for RanGAP1 antibodies; and J. Palvimo for RNF4 antibodies. We thank the core facility of the Institut Universitaire d’Hématologie for confocal microscopy analyses. The core facility is supported by grants from the Association Saint-Louis, Conseil Regional d’Ile-de-France, and the Ministère de la Recherche. We thank members of the laboratory, particularly J.C. Gluckman, J. Ablain, and A. Bazarbachi, as well as P. Cossart and C. Dargemont, for critical review of the manuscript.
The laboratory is supported by INSERM, CNRS, Université Paris-Diderot, Institut Universitaire de France, Ligue Contre le Cancer, Institut National du Cancer, Association pour la Recherche contre le Cancer (Griffuel Award to H. de Thé), Canceropôle Ile de France, the European Research Council (STEMAPL advanced grant to H. de Thé), PACRI and Saint Louis Institute, and the French National Research Agency (ANR) as part of the “Investissements d’Avenir” program (reference: ANR-11-PHUC-002). U. Sahin was supported by a fellowship of the Fondation pour la Recherche Medicale.
The authors declare no competing financial interests.
References
- APL
acute promyelocytic leukemia
- As2O3
arsenic trioxide
- DAXX
death domain–associated protein 6
- HIPK2
homeodomain-interacting protein kinase 2
- MEF
mouse embryonic fibroblast
- NB
nuclear body
- NM
nuclear matrix
- PML
promyelocytic leukemia
- RanGAP1
Ran GTPase-activating protein 1
- RARA
retinoic acid receptor A
- RNF4
Ring-finger protein 4
- ROS
reactive oxygen species
- SIM
SUMO-interacting motif
- SP100
nuclear autoantigen Sp-100
- SUMO
small ubiquitin-like modifier
- TDG
thymine-DNA glycosylase
- UBC9
ubiquitin-conjugating enzyme 9
Author notes
M. Jeanne’s present address is University of California, San Francisco, San Francisco, CA 94143.