Distributing junctional components around the cell periphery is key for epithelial tissue morphogenesis and homeostasis. We discovered that positioning of dynamic microtubules controls the asymmetric accumulation of E-cadherin. Microtubules are oriented preferentially along the dorso-ventral axis in Drosophila melanogaster embryonic epidermal cells, and thus more frequently contact E-cadherin at dorso-ventral cell–cell borders. This inhibits RhoGEF2, reducing membrane recruitment of Rho-kinase, and increasing a specific E-cadherin pool that is mobile when assayed by fluorescence recovery after photobleaching. This mobile E-cadherin is complexed with Bazooka/Par-3, which in turn is required for normal levels of mobile E-cadherin. Mobile E-cadherin–Bazooka prevents formation of multicellular rosette structures and cell motility across the segment border in Drosophila embryos. Altogether, the combined action of dynamic microtubules and Rho signaling determines the level and asymmetric distribution of a mobile E-cadherin–Bazooka complex, which regulates cell behavior during the generation of a patterned epithelium.
Introduction
Stable adhesion between cells is required to maintain the integrity of epithelial sheets during development and throughout life. It is crucial to balance the stability and the dynamics of cell adhesion so that cells can undergo morphogenetic changes, including convergent extension during development, and also respond to physical forces in mature epithelia. The major component of cell–cell adhesive contacts (adherens junctions) is E-cadherin (E-cad), a transmembrane protein that mediates homophilic adhesion (Zhang et al., 2009). The intracellular domain of E-cad recruits other proteins, including β-catenin, α-catenin, and p120catenin, to sites of adhesion, and couples adhesion to the actin cytoskeleton and signaling molecules (for reviews see Nelson, 2008; van Roy and Berx, 2008). Local E-cad concentration and dynamic behavior determines the strength of adhesion and E-cad signaling, which are the key factors for normal tissue morphogenesis and homeostasis (Niessen et al., 2011).
The distribution of E-cad junctions is tightly regulated, not only into a discrete band along the apical-basal axis, but also around the cell periphery. The even distribution of E-cad around the periphery requires Rap1, demonstrating that generating an even distribution requires an active mechanism (Knox and Brown, 2002). Microtubules (MTs) are known to regulate cortical dynamics and asymmetry, with MT plus ends being oriented preferentially toward the cell periphery. Dynamic instability of the plus ends allows MTs to grow outwards and explore peripheral structures, including sites of E-cad and integrin adhesion (e.g., Kaverina et al., 1999; Stehbens et al., 2006). Furthermore, MT plus ends generate cortical asymmetry to establish elongated cell shape in Schizosaccharomyces pombe (for review see Chang and Martin, 2009). Several +TIPs (MT plus end tracking proteins) transiently associate with MT plus ends and regulate their dynamics and interactions with other cell structures (e.g., for review see Akhmanova and Steinmetz, 2008). For example, +TIP End-Binding 1 (EB1) suppresses the transition from MT growth to shrinkage (catastrophes; e.g., Komarova et al., 2009). Furthermore, EB1 links MT plus ends to numerous other molecules, including regulators of MT dynamics and signaling proteins (for review see Akhmanova and Yap, 2008). Dynamic MTs are necessary for the local accumulation of E-cad in MCF-7 cells (Stehbens et al., 2006), which suggests that MT regulation of E-cad could also be important in regulating E-cad distribution and function in morphogenetic events.
Here, we examine a model system where the normal distribution of E-cad is uneven around the cell periphery, and find that this uneven distribution is important for regulating cell mixing within the epidermis of Drosophila melanogaster embryos. Pattern formation within the embryo requires combined mechanisms of cell fate determination and control of cell movement and mixing, as too much movement within cell layers may destroy the patterns laid down by patterning networks. The well-known cascade of pattern formation genes, from maternally localized axis-determining genes, to gap, pair-rule, and segment polarity genes, divides up the epidermis into segmental units, each further separated by a parasegment boundary into anterior and posterior compartments (e.g., for review see Sanson, 2001). The mechanisms that cause cells to respect segment and parasegment boundaries are still being elucidated. Recently, a transcellular acto-myosin cable was found to restrict cell movement across the parasegment boundary (Monier et al., 2010). However, cells within the engrailed-expressing posterior compartment can cross the segment borders, but then they shut off engrailed and change their fate (Vincent and O’Farrell, 1992). The mechanisms that control cell crossing at the segmental boundary are not known.
Here, we demonstrate that dynamic MTs regulate the asymmetric distribution of a specific mobile pool of E-cad. This mobile pool is not just a precursor to the immobile pool, but behaves as a distinct complex, containing the adaptor protein Bazooka/Par-3 (Baz), best known for its earlier function in setting up apical-basal cell polarity and positioning of E-cad junctions (for review see St Johnston and Ahringer, 2010). MTs elevate the mobile E-cad–Baz pool by inhibiting Rho signaling. Finally, we show that the elevated mobile E-cad at dorso-ventral (DV) borders reduces cell crossing of the segment boundary. Thus, we describe a pathway from dynamic MTs to the maintenance of segmental boundaries, via a dynamic E-cad–Baz complex.
Results
Identification of two pools of E-cad, one of which is polarized
To address mechanisms controlling the planar distribution of E-cad within adherens junctions, we focused on a subset of lateral epidermal cells in Drosophila embryos at the transition from stage 15 to stage 16 (further referred to as “stage 15,” see Material and methods for details). As described in the following paragraph, these cells have a natural asymmetry in E-cad distribution, and furthermore, in this model system we can inhibit protein function in stripes of cells and thereby produce perturbed and control cells side-by-side in each embryo (with engrailed::Gal4 [en::Gal4] driving upstream activation sequence [UAS] constructs). At this stage, these cells have completed all divisions, and the major morphogenetic rearrangements germ-band retraction and dorsal closure have been completed.
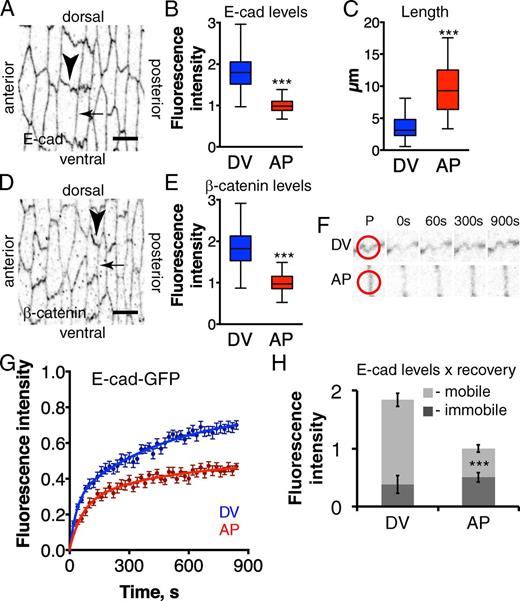
At stage 15, the embryonic epidermal cells have a rectangular shape, with short sides on DV cell borders and long sides on anterior–posterior (AP) borders; DV cell borders were 2.7-fold shorter than AP (3.6 ± 0.2 µm vs. 9.6 ± 0.4 µm; Fig. 1, A and C). The length difference is countered by differences in endogenous E-cad levels as detected by antibody staining: DV borders had 1.83 ± 0.07-fold more E-cad (fluorescent intensity per micrometer) than AP borders (Fig. 1, A and B; numerical values are given in Table S1; see also Price et al., 2006; Kaplan and Tolwinski, 2010). Detecting E-cad complexes with other reagents gave similar results: DV borders had 1.88 ± 0.05-, 1.83 ± 0.04-, and 1.84 ± 0.04-fold more ubiquitin promoter–driven E-cad–GFP, endogenous β-catenin, and β-catenin–YFP protein trap, respectively, than AP borders (Fig. 1, D–E; and not depicted). This difference in levels is not due to greater convolution at DV borders, as levels of other membrane-associated proteins, such as the cell adhesion protein Echinoid (Ed-YFP protein trap) and the Crumbs complex–associated protein Stardust, did not differ (Fig. 2, A and B; and not depicted). Thus, E-cad level differences indicate specific regulation around the cell periphery.
Mobile E-cad is asymmetrically distributed in epidermal cells. (A–E) There are two junction types in the lateral epidermis of stage 15 embryos: short DV borders (DV, arrowhead in A and D) and long AP borders (AP; arrow in A and D). DV borders have higher junctional E-cad (A and B, anti–E-cad) and β-catenin levels (D and E, anti–β-catenin), and are shorter (C) than AP. Bars, 5 µm. (F and G) There is greater exchange of E-cad–GFP at DV borders than AP, shown by greater FRAP; single FRAP examples (F), with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (mean ± SEM) plotted in G, with best fit curves as solid lines. (H) Combining the measurements of E-cad junctional levels and E-cad–GFP recovery provides estimates of mobile and immobile pools of E-cad at DV and AP borders (error bars indicate mean ± SEM), showing that DV borders have more mobile E-cad. Table S1 has detailed numbers for this and all other figures. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
Mobile E-cad is asymmetrically distributed in epidermal cells. (A–E) There are two junction types in the lateral epidermis of stage 15 embryos: short DV borders (DV, arrowhead in A and D) and long AP borders (AP; arrow in A and D). DV borders have higher junctional E-cad (A and B, anti–E-cad) and β-catenin levels (D and E, anti–β-catenin), and are shorter (C) than AP. Bars, 5 µm. (F and G) There is greater exchange of E-cad–GFP at DV borders than AP, shown by greater FRAP; single FRAP examples (F), with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (mean ± SEM) plotted in G, with best fit curves as solid lines. (H) Combining the measurements of E-cad junctional levels and E-cad–GFP recovery provides estimates of mobile and immobile pools of E-cad at DV and AP borders (error bars indicate mean ± SEM), showing that DV borders have more mobile E-cad. Table S1 has detailed numbers for this and all other figures. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
Mechanisms of E-cad recovery. (A and B) Localization of Echinoid-YFP (A; Ed, black/green in epidermal cells) and quantification (B) of Ed junctional levels in cells expressing EB1-DN (magenta in A) and adjacent wild-type cells. Bar, 5 µm. (C–H) E-cad–GFP FRAP recovery at DV borders: (C–D) comparing 1-µm and 2-µm bleach spots with the inset in D showing a closer view of the initial recovery to highlight the difference; (E and F) with ShiDN versus wild type; and (G and H) comparing the first and second bleach of the same spot. Examples of recovery are shown in C, E, and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in D, F, and H. The first frame after the first bleach event (B) and the prebleach frame before the second bleach event (S) are shown in G.
Mechanisms of E-cad recovery. (A and B) Localization of Echinoid-YFP (A; Ed, black/green in epidermal cells) and quantification (B) of Ed junctional levels in cells expressing EB1-DN (magenta in A) and adjacent wild-type cells. Bar, 5 µm. (C–H) E-cad–GFP FRAP recovery at DV borders: (C–D) comparing 1-µm and 2-µm bleach spots with the inset in D showing a closer view of the initial recovery to highlight the difference; (E and F) with ShiDN versus wild type; and (G and H) comparing the first and second bleach of the same spot. Examples of recovery are shown in C, E, and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in D, F, and H. The first frame after the first bleach event (B) and the prebleach frame before the second bleach event (S) are shown in G.
Examination of the dynamic behavior of E-cad by measuring FRAP of ubiquitin promoter–driven E-cad–GFP revealed two pools of E-cad within the adherens junctions: mobile and immobile. E-cad–GFP FRAP curves were best fit by a biexponential equation (Eq. 1; Fig. 1, F and G; best fit parameters are given in Table S1), resulting in fast (∼30 s) and slow (∼350 s) half times, indicative of two different recovery processes. The fast process is diffusion within the membrane because, as predicted (Sprague and McNally, 2005), the rate depends on the size of the bleach spot (Fig. 2, C and D), and is not affected by dominant-negative dynamin (Shibire-DN; Fig. 2, E and F). The slow process is endocytic trafficking, as it has the converse behavior (Fig. 2, C–F). The maximum recovery corresponds to the ratio of mobile and immobile protein at the cell–cell contact (Lippincott-Schwartz et al., 2003), and for E-cad–GFP, the ratio differs at DV and AP borders. We confirmed that the partial recovery of fluorescence is caused by a naturally occurring immobile fraction, rather than the immobility being induced by bleaching, by rebleaching a previously bleached spot (containing fluorescence only from recovered mobile E-cad) and obtaining full recovery (Fig. 2, G and H). We estimated the levels of mobile E-cad as the product of the total E-cad level times the fraction that recovered in FRAP at DV and AP borders; and immobile E-cad as the total minus mobile E-cad (Fig. 1 H and Table S1). This revealed that the asymmetry in E-cad distribution is caused by differences in mobile E-cad. The immobile E-cad was evenly distributed around the cell periphery, whereas mobile E-cad was elevated at DV borders.
Mobile E-cad contains and requires Bazooka/Par3
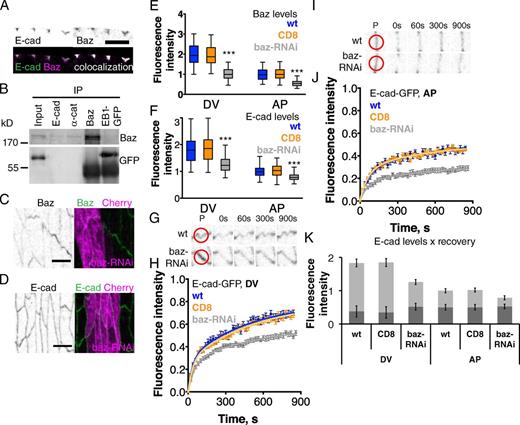
We examined whether there were differences in the intracellular proteins that associated with immobile and mobile E-cad. The three catenins, α, β, and p120, were associated with both fractions (Fig. S1 and not depicted). The elevated mobile E-cad on the DV borders was reminiscent of the polarized distribution of Baz earlier in embryonic development, where Baz is needed for elevated β-catenin levels at DV borders (Zallen and Wieschaus, 2004; Simões et al., 2010). At this later stage, we found that Baz was still elevated on DV borders (Fig. 3, C and E), where it colocalizes with E-cad (Fig. 3 A). At early developmental stages, Baz binds E-cad (Harris and Peifer, 2005; Wei et al., 2005), therefore we tested if this interaction occurs at stage 15. Endogenous E-cad and α-catenin, but not the control EB1-GFP, coimmunoprecipitated with endogenous Baz (Fig. 3 B). Additional lines of evidence show that mobile E-cad is bound to Baz and requires its function. Zygotic expression of UAS::baz-RNAi with en::Gal4 driver reduced cortical Baz levels at both AP and DV borders (Fig. 3, C and E), which reduced E-cad levels and asymmetry (Fig. 3, D and F) and the maximum recovery of E-cad–GFP at DV and AP borders (Fig. 3, G–J). As a control, expression of CD8 with the same Gal4 driver did not change Baz nor E-cad levels, nor E-cad–GFP recovery (Fig. 3). Estimation of mobile and immobile E-cad levels (Fig. 3 K) showed that immobile E-cad was not changed by Baz reduction, whereas mobile E-cad was reduced but maintained asymmetry. This contrasts with the consequences of reducing a protein associated with both E-cad pools, β-catenin (using arm-RNAi), which reduced both pools (Fig. S1).
Baz binds E-cad and is required for normal junctional levels of mobile E-cad. (A) E-cad and Baz colocalize at junctions as scored by antibody staining of endogenous proteins. (B) Endogenous E-cad and α-catenin (α-cat), but not EB1-GFP, coimmunoprecipitate with Baz. Input lane shows 5% of immunoprecipitation volume. (C–F) Baz (C; black/green, anti-Baz) and E-cad (D; black/green, anti–E-cad) in wild-type cells and adjacent cells expressing baz-RNAi (magenta) or CD8 as control (not depicted), and quantitation of levels (E and F). Bars, 5 µm. (G–J) E-cad–GFP FRAP in cells expressing baz-RNAi at DV (G and H) and AP borders (I and J). Examples of recovery are shown in G and I, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in H and J. See Fig. S1 for a similar experiment with β-catenin RNAi. (K) Combining the data permits an estimate of mobile and immobile E-cad pools (error bars indicate mean ± SEM) at DV and AP borders. ***, P < 0.0001.
Baz binds E-cad and is required for normal junctional levels of mobile E-cad. (A) E-cad and Baz colocalize at junctions as scored by antibody staining of endogenous proteins. (B) Endogenous E-cad and α-catenin (α-cat), but not EB1-GFP, coimmunoprecipitate with Baz. Input lane shows 5% of immunoprecipitation volume. (C–F) Baz (C; black/green, anti-Baz) and E-cad (D; black/green, anti–E-cad) in wild-type cells and adjacent cells expressing baz-RNAi (magenta) or CD8 as control (not depicted), and quantitation of levels (E and F). Bars, 5 µm. (G–J) E-cad–GFP FRAP in cells expressing baz-RNAi at DV (G and H) and AP borders (I and J). Examples of recovery are shown in G and I, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in H and J. See Fig. S1 for a similar experiment with β-catenin RNAi. (K) Combining the data permits an estimate of mobile and immobile E-cad pools (error bars indicate mean ± SEM) at DV and AP borders. ***, P < 0.0001.
As further confirmation that Baz is bound to mobile E-cad, we performed FRAP on Baz-GFP, expressed with UAS promoter and en::Gal4 (Fig. 4, A–D). Similar to endogenous Baz, Baz-GFP colocalized and coimmunoprecipitated with E-cad (Fig. 4 E and not depicted). Baz-GFP recovery was also best fit by a biexponential equation (Eq. 1) but fully recovered, showing the absence of an immobile fraction. Similar to E-cad, Baz-GFP recovers due to both diffusion (recovery depends on the size of bleach spot; Fig. 4, A and B), and dynamin-dependent endocytic trafficking (Fig. 4, C and D). The effect of Shi-DN on E-cad and Baz is consistent, despite the reduction of E-cad–GFP recovery and the apparent increase in Baz-GFP recovery (Figs. 2 F and 4 F). In each case, the recovery caused by endocytic trafficking is eliminated, leaving just recovery caused by diffusion; for Baz, all cell junctional protein is diffusive, whereas for E-cad there is also the immobile fraction. The half times of Baz-GFP and E-cad–GFP recovery were similar (Table S1), which is consistent with them being in a complex. Finally, we found that Baz and E-cad colocalized in the tubular structures formed when dynamin was inhibited (Fig. 4 E for Baz-GFP; and not depicted for endogenous Baz). Thus, putting all these results together, we have found that there are two pools of E-cad: an immobile fraction and a mobile fraction that we show here is specifically associated with Baz and at least in part dependent on Baz. These findings raised two main questions: (1) how does the mobile E-cad–Baz pool become especially elevated on DV borders, and (2) what is the function of this distinct cadherin subcomplex?
Mechanisms of Baz recovery. (A–D) Baz-GFP FRAP at DV borders. Baz-GFP recovery is via diffusion and trafficking, similar to E-cad–GFP: comparing 1-µm and 2-µm bleach spots with the inset in B showing a closer view of the initial recovery to highlight the difference (A and B), and in cells expressing ShiDN (C and D). Examples of recovery are shown in A and C, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in B and D. (E) E-cad and Baz-GFP colocalized in tubular structures (arrows) caused by expression of Shi-DN. Colocalization is shown in white in right image. Bar, 5 µm.
Mechanisms of Baz recovery. (A–D) Baz-GFP FRAP at DV borders. Baz-GFP recovery is via diffusion and trafficking, similar to E-cad–GFP: comparing 1-µm and 2-µm bleach spots with the inset in B showing a closer view of the initial recovery to highlight the difference (A and B), and in cells expressing ShiDN (C and D). Examples of recovery are shown in A and C, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) with best fit curves shown as solid lines in B and D. (E) E-cad and Baz-GFP colocalized in tubular structures (arrows) caused by expression of Shi-DN. Colocalization is shown in white in right image. Bar, 5 µm.
Polarized MTs increase the mobile pool of E-cad at DV cell–cell borders
As cadherins are regulated by dynamic MTs (Stehbens et al., 2006), we sought to test whether MTs regulate E-cad junctional levels. In our system, MTs form a dense apical network, similar to other Drosophila and mammalian cells (Gilbert et al., 1991; Rogers et al., 2008), with the apical MTs oriented predominantly along the DV axis (Fig. 5 A; Dickinson and Thatcher, 1997). To be able to visualize and manipulate dynamic MTs we generated two new tools.
EB1-DN and Spas change MT organization and dynamics. (A) Spas eliminated MTs in epidermal cells at the level of E-cad junctions (wild type [wt], Spas expressed in a stripe of two cells; α-tubulin, magenta; E-cad, green). (B) EB1 domains, showing MT-binding CH, coil-coiled (CC) dimerization, and acidic C terminus (Ac) are depicted, as well as new constructs with GFP and Cherry tags. (C) Sample kymographs showing growth of individual MTs in wild-type (wt) and EB1-DN cells. (D–G) EB1-DN reduced processivity of MT growth and the number of growing ends. Quantitation of individual growth event duration (D), MT growth speed (E), EB1-GFP fluorescence intensity at individual plus ends (F), and plus end density per square micrometer at the level of E-cad junctions (G). (H and I) EB1-DN changed oriented growth of MTs. (H) Projection of 20 images taken every 0.5 s. EB1-GFP, black/green; EB1-DN, magenta. (I) Quantitation of growth orientation of individual MTs in control and EB1-DN cells (Cherry-positive in H) relative to dorsal (D), ventral (V), anterior (A), and posterior (P) sides of the embryos. Red lines mark 45° segments. Bars, 5 µm. ***, P < 0.0001.
EB1-DN and Spas change MT organization and dynamics. (A) Spas eliminated MTs in epidermal cells at the level of E-cad junctions (wild type [wt], Spas expressed in a stripe of two cells; α-tubulin, magenta; E-cad, green). (B) EB1 domains, showing MT-binding CH, coil-coiled (CC) dimerization, and acidic C terminus (Ac) are depicted, as well as new constructs with GFP and Cherry tags. (C) Sample kymographs showing growth of individual MTs in wild-type (wt) and EB1-DN cells. (D–G) EB1-DN reduced processivity of MT growth and the number of growing ends. Quantitation of individual growth event duration (D), MT growth speed (E), EB1-GFP fluorescence intensity at individual plus ends (F), and plus end density per square micrometer at the level of E-cad junctions (G). (H and I) EB1-DN changed oriented growth of MTs. (H) Projection of 20 images taken every 0.5 s. EB1-GFP, black/green; EB1-DN, magenta. (I) Quantitation of growth orientation of individual MTs in control and EB1-DN cells (Cherry-positive in H) relative to dorsal (D), ventral (V), anterior (A), and posterior (P) sides of the embryos. Red lines mark 45° segments. Bars, 5 µm. ***, P < 0.0001.
To visualize dynamic MTs, we prepared a transgene encoding EB1, the MT plus end–binding protein, tagged with GFP and expressed under its own promoter (EB1-GFP), which is able to rescue Eb1 mutations (Fig. 5 B; see Materials and methods and Video 1). To inhibit MT dynamics, we made a dominant-negative version of Drosophila EB1 (EB1-DN; Fig. 5 B), using the same strategy as used for human EB1 (Komarova et al., 2009). By combining both of these reagents, we characterized how EB1-DN alters dynamic MTs. Expression of EB1-DN with a UAS promoter and en::Gal4 reduced the duration of MT growth without altering the mean growth speed (Fig. 5, C–E), which is equivalent to the EB1-DN effects in mammalian cells (Komarova et al., 2009) and EB1 depletion by RNAi in Drosophila cells (Rogers et al., 2002). In addition, we found that EB1-DN reduced EB1-GFP levels at individual plus ends, and the mean number of EB1-GFP–positive plus ends (Fig. 5, F and G). Importantly, EB1-DN perturbed the normal polarized orientation of MTs in the DV direction (Fig. 5, H and I), resulting in greater variation in direction of growth, although a bias toward the DV direction remained. This suggests that persistent growth is required for MTs to become oriented along the long axis of the apical surface. Hence, we can use EB1-DN to reduce the number and processivity of dynamic MTs. To disrupt MTs more generally, we overexpressed the MT-severing protein Spastin (Spas; Roll-Mecak and Vale, 2005). Spas overexpression almost completely eliminated MTs (Fig. 5 A) without causing substantial differences to the cell shape, making it suitable for our analysis.
Epidermis at stage 15 of embryo development expressing EB1-GFP (black) tracking MT plus ends in control (left) and cells expressing EB1-DN (right). Images were analyzed by time-lapse microscopy using spinning disc confocal microscope (Eclipse Ti-E inverted microscope [Nikon], equipped with a CFI Apochromat TIRF 100× 1.49 NA oil objective lens [Nikon] and a motorized CSU-X1-A1 confocal head [Yokogawa Corporation of America]). Frames were taken every 0.5 s for 50 s.
Epidermis at stage 15 of embryo development expressing EB1-GFP (black) tracking MT plus ends in control (left) and cells expressing EB1-DN (right). Images were analyzed by time-lapse microscopy using spinning disc confocal microscope (Eclipse Ti-E inverted microscope [Nikon], equipped with a CFI Apochromat TIRF 100× 1.49 NA oil objective lens [Nikon] and a motorized CSU-X1-A1 confocal head [Yokogawa Corporation of America]). Frames were taken every 0.5 s for 50 s.
We then tested how perturbing MTs altered E-cad distribution and dynamics. EB1-DN reduced endogenous E-cad at the DV borders, but not AP borders, whereas Spas reduced E-cad at both types of borders (Fig. 6, A and B). EB1-DN also reduced endogenous Baz, strongly at DV borders and more weakly at AP, whereas Spas strongly reduced Baz at both borders (Fig. 6, C and D). EB1-DN reduced the maximum of E-cad–GFP recovery in FRAP only at DV borders, whereas Spas reduced the maxima of recovery at both borders (Fig. 6, E–H). Combining the results showed that the changes to immobile E-cad were not significant and that immobile E-cad remained symmetrically distributed in both EB1-DN– and Spas-expressing cells (Fig. 6 I). EB1-DN resulted in a specific loss of mobile E-cad from DV borders. Embryos homozygous for an Eb1 mutant allele had the same changes to E-cad (Fig. S2), which confirms that the effects of EB1-DN are caused by reducing EB1 function. To test whether the effects of EB1-DN are general to transmembrane adhesion molecules, we examined Echinoid-YFP, but observed no change in distribution or dynamics (Fig. 2, A and B; Table S1; and data not depicted). Compared with EB1-DN, Spas reduced mobile E-cad more uniformly (Fig. 6 I). These results support a model where dynamic MT plus end proximity to cell borders increases mobile E-cad; EB1-DN causes a selective loss of plus ends at the DV borders, whereas Spas eliminates almost all plus ends. These data are consistent with loss of the asymmetrical distribution of E-cad, β-catenin, and Baz in fat and expanded mutant cells that also showed disrupted apical MT organization (Marcinkevicius and Zallen, 2013). Thus, we infer that MTs elevate mobile E-cad levels and that the polarization of MTs toward the DV borders leads to the higher levels of mobile E-cad at these borders. This does not seem to be caused by kinesin-driven delivery of E-cad to the cortex, as observed for N-cad (Mary et al., 2002), as elimination of Spas did not produce the effects predicted from loss of delivery: (1) accumulation of E-cad in intracellular vesicles as for N-cadherin (Mary et al., 2002) or when E-cad exocytosis is defective (Langevin et al., 2005), and (2) the depletion of mobile E-cad from the junctions, as in the impairment of endocytic trafficking with Shi-DN (Fig. 2 F).
EB1-DN and Spas reduce mobile E-cad–Baz and its asymmetry. (A–D) Endogenous E-cad (A) and Baz (C) localization in cells expressing EB1-DN or Spas (anti–E-cad, black/green in A; anti-Baz, black/green in C; Cherry-tagged products in experimental cells, magenta), and quantitation of E-cad (B) and Baz (D) levels at the junctions. Bars, 5 µm. (E–H) Recovery of E-cad–GFP in cells that express CD8, EB1-DN, or Spas at DV borders (E and F) and AP borders (G and H). Examples of recovery are shown in E and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) in F and H. wt, wild-type cells adjacent to experimental. (G) Combining this data provides an estimate of mobile and immobile E-cad pools (mean ± SEM); inhibition of MTs especially reduced the mobile E-cad at DV borders. Fig. S2 shows the same defect in Eb1 mutant embryos. **, P < 0.001; ***, P < 0.0001.
EB1-DN and Spas reduce mobile E-cad–Baz and its asymmetry. (A–D) Endogenous E-cad (A) and Baz (C) localization in cells expressing EB1-DN or Spas (anti–E-cad, black/green in A; anti-Baz, black/green in C; Cherry-tagged products in experimental cells, magenta), and quantitation of E-cad (B) and Baz (D) levels at the junctions. Bars, 5 µm. (E–H) Recovery of E-cad–GFP in cells that express CD8, EB1-DN, or Spas at DV borders (E and F) and AP borders (G and H). Examples of recovery are shown in E and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) in F and H. wt, wild-type cells adjacent to experimental. (G) Combining this data provides an estimate of mobile and immobile E-cad pools (mean ± SEM); inhibition of MTs especially reduced the mobile E-cad at DV borders. Fig. S2 shows the same defect in Eb1 mutant embryos. **, P < 0.001; ***, P < 0.0001.
Dynamic MT plus ends suppress Rho inhibition of mobile E-cad
In seeking to discover how dynamic MTs regulate mobile E-cad levels, we explored several proteins that become enriched at MT tips, and had success with RhoGEF2, which binds EB1 (Rogers et al., 2004), is enriched at AP borders, and is required for junctional asymmetry of E-cad in early Drosophila embryogenesis (Levayer et al., 2011; Warrington et al., 2013). The rapid exchange of proteins on the plus ends (Bieling et al., 2007; Dragestein et al., 2008) indicates that the association of RhoGEF2 will not lead to transport along the growing MT, but instead may concentrate RhoGEF2 with co-recruited positive or negative regulators, or alternatively may locally reduce the free RhoGEF2 concentration.
First, we tested if altering the RhoGEF2 concentration changes mobile E-cad–Baz levels. In contrast to when we perturbed MTs, changing RhoGEF2 levels affected both immobile and mobile E-cad, which suggests two Rho pathways, one of which regulates immobile E-cad independent of MTs. Overexpression of RhoGEF2 did not change total E-cad fluorescence at DV borders, and slightly increased it at AP borders (Fig. 7, A and B). At both borders it reduced the junctional levels of Baz (Fig. 7, C and D), and the mobile fraction of E-cad (Fig. 7, E–H). Combining these results (Fig. 7 I) shows that overexpressed RhoGEF2 decreased mobile E-cad–Baz while elevating immobile E-cad. This is consistent with a double-negative pathway where dynamic MTs inhibit RhoGEF2, which reduces mobile E-cad–Baz. In the reciprocal experiment reducing RhoGEF2, we only saw the expected elevation of mobile E-cad–Baz on AP borders, presumably because RhoGEF2 was already fully inhibited at DV borders, and this effect was obscured by the general reduction of E-cad levels. Thus, knocking down RhoGEF2 by RNAi had three effects: (1) reduced total endogenous E-cad levels at both borders (Fig. 7, A and B); (2) reduced junctional Baz levels at DV borders but not AP borders (Fig. 7, C and D), which represents an increase in Baz relative to E-cad total levels at AP borders; and (3), similarly, increased the fraction of mobile E-cad–GFP at AP borders (Fig. 7, G and H). Combining these results (Fig. 7 I) shows that, opposite to RhoGEF2 overexpression, RhoGEF2-RNAi elevated the levels of mobile E-cad at AP borders. At both borders it increased the fraction of E-cad that is mobile. In addition, it decreased immobile E-cad, significantly at AP borders, but not significantly at DV borders (Fig. 7, E–I; and Table S1). Thus, we conclude that RhoGEF2 negatively regulates the fraction of mobile E-cad, and that MTs may therefore act by inhibiting RhoGEF2. In addition, RhoGEF2 has an independent function in increasing levels of immobile/total E-cad.
RhoGEF2 negatively regulates mobile E-cad–Baz. (A–D) Endogenous E-cad (A; black/green, anti–E-cad) and Baz (C; black/green, anti-Baz) localization in wild type and upon RhoGEF2 overexpression or down-regulation with RhoGEF2-RNAi (A and C; magenta), and quantitation of E-cad (B) and Baz (D) junctional levels. Bars, 5 µm. (E–H) E-cad–GFP FRAP upon RhoGEF2 overexpression or down-regulation with RhoGEF2-RNAi at DV (E and F) and AP borders (G and H). Examples of recovery are shown in E and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) in F and H. (I) Combining this data provides an estimate of mobile and immobile E-cad pools (error bars indicate mean ± SEM) at DV and AP borders. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
RhoGEF2 negatively regulates mobile E-cad–Baz. (A–D) Endogenous E-cad (A; black/green, anti–E-cad) and Baz (C; black/green, anti-Baz) localization in wild type and upon RhoGEF2 overexpression or down-regulation with RhoGEF2-RNAi (A and C; magenta), and quantitation of E-cad (B) and Baz (D) junctional levels. Bars, 5 µm. (E–H) E-cad–GFP FRAP upon RhoGEF2 overexpression or down-regulation with RhoGEF2-RNAi at DV (E and F) and AP borders (G and H). Examples of recovery are shown in E and G, with red circles on the prebleached frame (P) showing the bleach spots, and averaged recovery curves (error bars indicate mean ± SEM) in F and H. (I) Combining this data provides an estimate of mobile and immobile E-cad pools (error bars indicate mean ± SEM) at DV and AP borders. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
We next sought evidence that Rho signaling levels are regulated by MTs differentially at DV and AP borders. We selected the Rho effector Rho-kinase (Rok) as a potential readout of Rho signaling, as it is elevated at AP borders at earlier developmental stages similar to RhoGEF2 (Simões et al., 2010; Levayer et al., 2011). We thought that Rok recruitment to junctions may reflect the local Rho signaling levels, and confirmed that its localization was affected by RhoGEF2. As predicted, the distribution of Rok-Venus, expressed from the spaghetti squash promoter, was complementary to mobile E-cad; i.e., higher on AP borders. Its junctional levels were increased by RhoGEF2 overexpression at AP and DV borders and reduced by RhoGEF2-RNAi at AP borders (Fig. 8), but not DV (which suggests that RhoGEF2 is already fully inactive on DV borders; see the previous paragraph). Thus, Rok-Venus membrane recruitment reflects RhoGEF2 activity.
Rok localization is regulated positively by RhoGEF2 and negatively by MTs. Rok-YFP localization (A; black/green) in wild-type (wt) and adjacent cells expressing RhoGEF2, RhoGEF2-RNAi, EB1-DN, or Spas marked with Cherry (magenta), and quantitation of Rok-YFP junctional levels (B). Bar, 5 µm. ***, P < 0.0001.
Rok localization is regulated positively by RhoGEF2 and negatively by MTs. Rok-YFP localization (A; black/green) in wild-type (wt) and adjacent cells expressing RhoGEF2, RhoGEF2-RNAi, EB1-DN, or Spas marked with Cherry (magenta), and quantitation of Rok-YFP junctional levels (B). Bar, 5 µm. ***, P < 0.0001.
We then tested if perturbing MTs changed the junctional distribution of Rok-Venus. As expected, if MTs inhibit RhoGEF2, EB1-DN increased Rok-Venus at DV borders, but not AP, whereas Spas up-regulated Rok-Venus uniformly, and the control CD8 had no effect (Fig. 8). Thus, MT inhibition increases Rok-Venus at the membrane, opposite to their effect on mobile E-cad–Baz (compare Fig. 8 and Fig. 6 I; for quantitative data see Table S1). Altogether, our results are consistent with dynamic MTs negatively regulating RhoGEF2, resulting in reduced recruitment of Rok-Venus and elevation of mobile E-cad. Although it seems likely that Rok inhibits mobile E-cad (see Discussion), we have not ruled out the reverse.
Mobile E-cad–Baz maintains epidermal cells within the posterior compartment
Finally, we addressed the function of this dynamic E-cad–Baz subcomplex. Our experimental tools reduced rather than eliminated mobile E-cad, so we expected modest changes in cell behavior. We discovered that mobile E-cad reduction hindered the ability of cells to avoid crossing the segment boundary. Engrailed-expressing cells in the posterior parasegment do occasionally cross the segment boundary, and then switch off engrailed (Vincent and O’Farrell, 1992). The segment boundary was identified by a straight alignment of cell–cell junctions, with cells on the posterior side having shorter AP borders (Fig. 9 A; compare AP borders in cells anterior and posterior to the segment boundary). Using en::Gal4 to drive stable fluorescent proteins increased the number of cells that we can score as having crossed over (Fig. 9 A; see Materials and methods). The proportion of segments showing cells that have crossed the segment boundary was increased by EB1-DN, Spas, or baz-RNAi, relative to wild type or CD8 (Fig. 9 B).
Mobile E-cad–Baz prevents cells from crossing the segment boundary. (A) Examples of stripes of en::Gal4 driving GFP (red), with cell outlines labeled with anti–E-cad (green). In most cases, cells do not cross the segment boundary (white line, left). When cells cross, one can see independent events with different levels of GFP (middle, arrowheads), a rare example of a cell pair in the process of crossing the boundary (right, arrow). Bars, 5 µm. (B) Quantitation of cell behavior in control cells and cells expressing CD8, EB1-DN, Spas, or baz-RNAi. (top) Percentage of stripes with cells expressing the engrailed-driven marker transgene on the other side of the segment boundary; (middle) relative rates of cell pair crossing, estimated using Poisson distribution; (bottom) percentage of rosettes, five- and six-cell contacts, between cells within the posterior compartment (the cells at the boundary and their anterior neighbors). In each case, the mean ± 95% CI is shown (error bars); for raw data and examples of fitting with Poisson distribution, see Fig. S3. (C) Examples of three-cell contact (3), four-cell contact (4), and five-cell contact rosette (R) between cells at the segment border and their anterior neighbors, indicated with asterisks in A. (D) Localization of MyoII-YFP (black/green) in wild-type cells and adjacent cells expressing CD8, EB1-DN, Spas, or baz-RNAi marked with Cherry (magenta). Bars, 5 µm. (E) Quantitation of cortical MyoII levels, see also Table S1. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
Mobile E-cad–Baz prevents cells from crossing the segment boundary. (A) Examples of stripes of en::Gal4 driving GFP (red), with cell outlines labeled with anti–E-cad (green). In most cases, cells do not cross the segment boundary (white line, left). When cells cross, one can see independent events with different levels of GFP (middle, arrowheads), a rare example of a cell pair in the process of crossing the boundary (right, arrow). Bars, 5 µm. (B) Quantitation of cell behavior in control cells and cells expressing CD8, EB1-DN, Spas, or baz-RNAi. (top) Percentage of stripes with cells expressing the engrailed-driven marker transgene on the other side of the segment boundary; (middle) relative rates of cell pair crossing, estimated using Poisson distribution; (bottom) percentage of rosettes, five- and six-cell contacts, between cells within the posterior compartment (the cells at the boundary and their anterior neighbors). In each case, the mean ± 95% CI is shown (error bars); for raw data and examples of fitting with Poisson distribution, see Fig. S3. (C) Examples of three-cell contact (3), four-cell contact (4), and five-cell contact rosette (R) between cells at the segment border and their anterior neighbors, indicated with asterisks in A. (D) Localization of MyoII-YFP (black/green) in wild-type cells and adjacent cells expressing CD8, EB1-DN, Spas, or baz-RNAi marked with Cherry (magenta). Bars, 5 µm. (E) Quantitation of cortical MyoII levels, see also Table S1. *, P < 0.01; **, P < 0.001; ***, P < 0.0001.
The distribution of the cells that have crossed the segment boundary gives some insight into the underlying process that results in crossing. Multiple crossing events can occur independently (Fig. 9 A), as indicated by examples having separate groups of cells across the boundary. Independent groups of crossed cells often had different degrees of reduction of en::Gal4–driven GFP (Fig. 9 A), which suggests that they crossed at different times. In addition, 90% of cell clusters across the boundary contained an even number of cells, suggesting that cells cross in pairs (Fig. S3). These findings suggest that cell pairs cross over an extended period, and we could estimate a relative rate of crossing, between wild-type and experimental perturbation, using Poisson distribution (Fig. S3). EB1-DN, Spas, and baz-RNAi caused elevations in the rate of 1.7 ± 0.2 (P < 0.001)-, 2.2 ± 0.3 (P < 0.0001)-, and 2.0 ± 0.2 (P < 0.0001)-fold, respectively (Fig. 9 B). Thus, the increased number of cases where cells have crossed the boundary is caused by an increase in the rate of pairs of cells crossing.
Crossing the segment boundary requires exchange of neighbors, and therefore remodeling of cell–cell junctions. Such remodeling can be documented by the formation of cell rosettes, as best characterized during convergent extension, earlier in embryonic development (Blankenship et al., 2006). We counted the types of multicellular contacts within the engrailed stripe, focusing on cells adjacent to the segment border (Fig. 9 C). The number of rosettes (contacts between five or more cells) was increased by 1.6 (P < 0.01)-, 2.1 (P < 0.0001)-, and 2.0 (P < 0.0001)-fold by EB1-DN, Spas, or baz-RNAi, respectively, in comparison to control (Fig. 9 B), whereas the number of four-cell contacts did not change (not depicted). The experimentally induced increases in the number of rosettes and crossing rate are similar, and suggest that rosette formation is the initial event that leads to segment boundary crossing. The mobile E-cad may stabilize this cell boundary, preventing its loss and rosette formation.
Previous work showed: (1) that cables of Myosin II (MyoII) prevent cell movement across the other (anterior) side of the stripe of engrailed-expressing cells in embryos and across DV and AP compartment boundaries in wing imaginal discs later in development (Major and Irvine, 2006; Landsberg et al., 2009; Monier et al., 2010), and (2) that, looking earlier in development, Baz reduces MyoII recruitment to adherens junctions, so that MyoII is lower at DV versus AP borders (Simões et al., 2010). MyoII, tagged with a YFP protein trap, was still asymmetrically distributed at stage 15, with higher levels at AP borders (Fig. 9, D and E), similar to Rok-Venus (Fig. 8 B and Table S1; Walters et al., 2006; Simone and DiNardo, 2010). It was therefore important to test whether increased crossing is caused by the reduction of dynamic E-cad–Baz, or the knock-on effect of elevating MyoII at DV borders, losing the normal distinction between high levels at AP and low at DV. Perturbing MTs did affect MyoII-YFP distribution, as EB1-DN increased MyoII-YFP at DV borders but not AP, and Spas increased MyoII-YFP at both borders (Fig. 9, D and E). However, when Baz was down-regulated using baz-RNAi, MyoII-YFP levels did not change (Fig. 9, D and E), even though loss of Baz affected MyoII distribution earlier in embryogenesis (Simões et al., 2010). As baz-RNAi increased rosette formation and the rate of border crossing to a similar level as Spas, but did not alter MyoII-YFP, we conclude that it is not perturbation of MyoII that reduces rosette formation to keep cells within the segment, but instead the loss of mobile E-cad.
Discussion
In this study we focused on the questions of how and why E-cad becomes asymmetrically distributed around the cell periphery in epithelial cells. We report three major findings. First, we demonstrate that there are two distinct E-cad pools at adherens junctions in Drosophila epidermal cells: a mobile pool that recovers after photobleaching and a stable pool that does not recover in the period of observation. Only the mobile pool of E-cad is associated with Baz, and elevated at DV borders, whereas the stable pool is uniformly distributed around the cell circumference. Second, we discovered the factors that are crucial for accumulation and polarized distribution of mobile E-cad–Baz: a polarized population of dynamic MTs locally reduces a Rho activity that antagonizes the mobile pool, leading to elevation of mobile E-cad at DV borders. Third, we identify a function for the mobile E-cad–Baz complex, which is to stabilize the short sides, reducing rearrangement of cell contacts into rosettes and crossing of the segment boundary.
Two pools of E-cad are present at adherens junctions
Since the initial discovery of E-cad as an adhesion molecule, numerous activities of E-cad have been identified (e.g., for review see Cavallaro and Dejana, 2011). Here we have described two pools of E-cad that can be distinguished by their different rates of recovery by FRAP, and differential association with Baz. As mobile E-cad–Baz levels can be altered independently of the immobile pool, the mobile pool does not appear to be a precursor of the immobile pool. Immobile and mobile E-cad fractions have also been characterized during convergent extension of the Drosophila embryo, earlier than the stage we have examined (Cavey et al., 2008). At this early stage the immobile pool is found in distinct clusters, whereas at the stage of our observations the two pools of E-cad are intermingled in the E-cad belt at the adherens junctions. Thus, with the right tools, an apparently homogenous adherens junction can be divided into subcomplexes, which suggests that these molecules have more than one function at this site. Similar cases of underlying diversity within apparently uniform domains of transmembrane proteins are been recently characterized. Thus, Crumbs within the stalk membrane of photoreceptor cells is in two complexes, distinguishable by the isoform of Stardust that is bound to Crumbs (Bulgakova et al., 2010). Within the ring of Echinoid, molecules bind either Baz or Canoe, as the binding is mutually exclusive, but at the resolution of light microscopy these two pools are intermingled (Wei et al., 2005). Thus, our findings add to a recent trend of discovery of underlying diversity within the adhesive structures formed by transmembrane receptors.
The association of Baz with E-cad is consistent with a variety of previous findings. Baz regulates the size and position of E-cad containing junctions along the lateral membrane (Harris and Peifer, 2004; Wang et al., 2012) and Baz can bind β-catenin (Wei et al., 2005). Here we have shown that E-cad–Baz turns over at adhesion sites faster than the E-cad not bound to Baz, and is regulated by dynamic MTs. Mutants that cause the mislocalization of Baz, e.g., crumbs, fail to maintain epithelial integrity during tissue remodeling (Campbell et al., 2009). This defect may be caused by the lack of E-cad–Baz turnover. In this study we were able to partially reduce mobile E-cad levels using baz-RNAi, or inhibition of dynamic MTs with EB1-DN or Spas, without affecting the immobile pool. This permitted the functional analysis of mobile E-cad in cell behavior (see the last portion of the Discussion).
Regulation of mobile E-cad–Baz by MTs and RhoGEF2
MTs are commonly oriented with their plus ends toward the cell cortex, and a subpopulation of MTs was shown to be targeted toward E-cad adhesion sites (Stehbens et al., 2006). In contrast to mammalian and Drosophila cells in culture, where the MTs extend radially (Stehbens et al., 2006; Rogers et al., 2008), the epidermal cells at the late stages of Drosophila development display apical acentrosomal MT arrays that are polarized along the DV axis of the embryo (this paper; Rogers et al., 2008; Marcinkevicius and Zallen, 2013), most likely downstream of the planar cell polarity–regulator Fat (Marcinkevicius and Zallen, 2013). Such organization of MTs results in a higher probability for the MTs to contact the DV borders than AP. Using EB1-DN to perturb MT dynamics resulted in a decrease in the mobile pool of E-cad at the DV borders, without any effect on the immobile pool, or E-cad at the AP borders. Therefore, we conclude that the polarized network of dynamic MTs regulate asymmetric distribution of the mobile pool of E-cad in the elongated epidermal cells within the Drosophila embryo.
Dynamic MTs have been suggested to function as “diffusional sinks”: the +TIPs transiently accumulate at plus ends, bind signaling molecules, and thus locally increase their concentration at the plus ends while depleting the amount at adjacent sites, such as the plasma membrane (Akhmanova et al., 2009). Our model of regulation of the mobile pool of E-cad by dynamic MTs fits with this mechanism (summarized in Figs. 10 A and S4). EB1 binds RhoGEF2, concentrating it at plus ends (Rogers et al., 2004). As DV borders are contacted by more MT plus ends, we predict that the free RhoGEF2 concentration near the membrane becomes more strongly reduced, lowering the activity of Rho as revealed by reduced Rok recruitment. The control of mobile E-cad–Baz levels by RhoGEF2 led us to infer that Rho signaling normally reduces mobile E-cad–Baz at adherens junctions. This inhibition could occur by the direct phosphorylation of Baz by Rok, as found in both Drosophila and mammalian cells (Nakayama et al., 2008; Simões et al., 2010), although we cannot yet rule out the possibility that the changes to mobile E-cad have an effect on Rok recruitment. Regulation by phosphorylation would predict that it is unphosphorylated Baz that is complexed with E-cad to ensure normal levels of junctional mobile E-cad. Baz binds β-catenin (Wei et al., 2005) and VE-cadherin (Iden et al., 2006), so the interaction of Baz with E-cad observed by coimmunoprecipitation could occur via either of these interactions, or indirectly (this paper; Harris and Peifer, 2005). Altogether, our speculative model proposes that MT plus ends sequester RhoGEF2 by EB1 to regulate spatial phosphorylation of Baz, which leads to an increase in the mobile E-cad–Baz complex (Fig. 10 A).
Models. (A) Model of regulation of mobile E-cad distribution by dynamic MTs. EB1 at MT plus ends sequesters RhoGEF2, suppressing Rok localization at DV borders. This releases inhibition of mobile E-cad–Baz, elevating it at DV borders. (B) Model of formation of a five-cell rosette (right, arrowhead) from four-cell contact (left) through either “unzipping” of the DV border (red, arrow) and reattachment with the AP border (red/black) or membrane internalization (red, arrows). (C) Model of cell crossing of the segment border through rosette resolution (blue asterisks). The collapsing junctions are in red, the newly formed junctions are in blue, and those changing from AP to DV are in yellow.
Models. (A) Model of regulation of mobile E-cad distribution by dynamic MTs. EB1 at MT plus ends sequesters RhoGEF2, suppressing Rok localization at DV borders. This releases inhibition of mobile E-cad–Baz, elevating it at DV borders. (B) Model of formation of a five-cell rosette (right, arrowhead) from four-cell contact (left) through either “unzipping” of the DV border (red, arrow) and reattachment with the AP border (red/black) or membrane internalization (red, arrows). (C) Model of cell crossing of the segment border through rosette resolution (blue asterisks). The collapsing junctions are in red, the newly formed junctions are in blue, and those changing from AP to DV are in yellow.
This model can explain what happens when MT dynamics are perturbed (Fig. S4). EB1-DN expression reduced the number of EB1-labeled growing plus ends and their preferential growth toward the DV borders. Therefore, the DV borders are approached by fewer EB1-labeled plus ends, resulting in less RhoGEF2 sequestering and greater cytoplasmic RhoGEF2 close to DV borders, and elevation of Rho activity, as reflected by the increase in Rok. At AP borders, the effect of reducing the number of EB1-labeled plus ends may be compensated by increased contact of the growing plus ends, explaining the lack of change in Rok and mobile E-cad (Fig. S4). Spas eliminates MTs and therefore there are no plus ends to concentrate RhoGEF2. This increases the RhoGEF2 concentration near both borders, elevating Rok and reducing mobile E-cad (Fig. S4). Thus, our results support the “diffusional sink” mechanism. To our knowledge, this work provides the first evidence that this mechanism operates to control cortical signaling.
Mobile E-cad reduces rosette formation and cell motility
Two phenotypes were observed when levels of mobile E-cad were reduced: increased formation of multicellular rosettes, and higher number of cells crossing the segment boundary. The changes in the rate of cells crossing the segment boundary and the number of rosettes caused by EB1-DN did not significantly differ from those caused by Spas. As the level of mobile E-cad at AP borders was not affected by EB1-DN, we conclude that it is largely the mobile E-cad at DV borders, but not AP borders, that prevents cells from crossing segment boundaries and rosette formation. How does mobile E-cad at DV borders normally reduce the formation of rosettes? Other recent examples support the idea that membrane domains are maintained by specific transmembrane proteins. The photoreceptor cells lacking Crumbs have a shorter apical membrane compartment, which is the normal site of Crumbs localization in a variety of model systems (for review see Gosens et al., 2008). Shortening of the lateral membrane during the follicular epithelial cell shape transition from cuboidal to squamous involves the removal of Fasciclin 2 from the membrane (Gomez et al., 2012). We suggest that, in a similar fashion, mobile E-cad–Baz protects the short cell–cell contacts from collapsing. These borders may require more mobile E-cad to maintain them because they are under lower tension than AP borders, as indicated by the differences in MyoII levels (Fig. 6 D; Monier et al., 2010). We envision two alternative models of such adhesion collapse (Fig. 10 B). In the first, cells exchange neighbors by detaching and reattaching their membranes. This implies that mobile E-cad is primarily involved in determining the strength of adhesion, as reduction in mobile E-cad–Baz increases membrane detachment, or “unzipping.” In the second, some borders between cells shrink by endocytosis of the membrane. This predicts that mobile E-cad blocks such membrane turnover. This could occur by inhibition of clathrin recruitment, as clathrin was found elevated at AP borders during germ band extension (Levayer et al., 2011). The recent finding that VE-Cad that cannot be internalized assembles correctly into adherens junctions, but prevents cell migration (Nanes et al., 2012), supports the turnover mechanism.
Formation of rosettes may be sufficient to cause cell crossing, as, earlier in development, multicellular rosettes drive directional cell intercalation during germ band extension (Blankenship et al., 2006). We suggest that pairs of cells are extruded across the segment border when rosettes are resolved along the AP axis of the embryo (Fig. 10 C). MyoII is required to prevent cells from crossing compartment boundaries during early embryogenesis and in the wing imaginal discs by forming an actomyosin barrier (Major and Irvine, 2006; Landsberg et al., 2009; Monier et al., 2010). We did not detect enrichment in MyoII accumulation at the segment border, in contrast to parasegment borders at earlier developmental stages or ventral epidermis (Monier et al., 2010). However, it is likely that elevated levels of MyoII at the AP borders support realignment of the segment borders after the cells cross it (Fig. 10 C). After the initial cell–pair extrusion, loss and creation of new cell borders will lead to neighbor exchange (Fig. 10 C).
It seems likely that the regulatory pathway from MTs to E-cad that we have identified also exists in mammalian cells. Disruption of MTs increases RhoA activity in mouse palate cells (Kitase and Shuler, 2012), whereas stabilization of MTs reduces recovery of E-cad in FRAP in mouse keratinocytes (Sumigray et al., 2012). In addition, Baz/Par-3 associates directly with VE-cad in CHO cells (Iden et al., 2006), and antagonizes MyoII in MDCK cells (Wan et al., 2012). However, this is not the only pathway for MTs to influence cadherin junctions. Recently, we demonstrated that dynamic MTs positively support Rho signaling at the epithelial zonula adherens through a mechanism that involved the MT-dependent localization of centralspindlin and Ect2 at the junctions (Ratheesh et al., 2012). This implies that a repertoire of mechanisms exists for MTs to regulate cell–cell junctions. Understanding how these mechanisms are deployed throughout development and across species, and how they may work through the two E-cad pools that we have discovered, is an important problem for future work.
Materials and methods
Fly stocks
The following transgenic fly stocks were used in this study: en::Gal4, UAS::CD8-mCherry, UAS::Shi-K44A, eb104524/CyO, UAS::RhoGEF2/CyO, and UAS::myr-GFP (all from Bloomington Drosophila Stock Center); Df(2R)ED1612 (DrosDel); ubi::E-cad–GFP (Oda and Tsukita, 2001); UAS::Spas/CyO (Sherwood et al., 2004); UAS::Baz-FL (Krahn et al., 2010); UAS::baz-RNAi, UAS::arm-RNAi, and UAS::RhoGEF2-RNAi (Vienna Drosophila RNAi Center stocks v2914, v107344, and v110577, respectively); sqh::RokK116A-Venus (Simões et al., 2010); and MyoII-YFP, armadillo-YFP, and Ed-YFP (all from Drosophila Genomics Resource Center). The flies and embryos were kept at 18°C.
Molecular cloning and transgenesis
The EB1-GFP construct contains an 8,322-bp fragment comprising from 3,876 bp 5′ to the ATG initiating codon to the last codon, a four-serine linker, the GFP coding sequence, and 1,151 bp 3′ of the stop codon. This DNA was cloned in the transposable P-element vector pWhiteRabbit for transgenesis. The dominant-negative EB1 (EB1-DN) construct contains a 609-bp fragment of Eb1 cDNA encoding aa 83–285, a four-serine linker, and the mCherry coding sequence. This region of eb1 is homologous to the sequence used to make a dominant-negative version of human EB1 (Komarova et al., 2009). This DNA was cloned in pUAST vector for transgenesis.
Data acquisition
All experiments were performed on lateral epidermal cells to exclude effects caused by denticle morphogenesis (Dickinson and Thatcher, 1997; Price et al., 2006) examining segments A1–A4. All experiments were done at the end of stage 15/beginning of stage 16 of embryonic development after completion of dorsal closure, which could be detected with all of the tagged proteins and antibody stainings. The image acquisition of junctional protein accumulation, FRAP, and segment boundary crossing was done using an upright confocal microscope (FV1000; Olympus) using 100×/1.40 NA oil UPlan-SApochromat objective lens. 16-bit depth images were taken at a magnification of 0.099 µm/pixel (FRAP), 0.062 µm/pixel (junctional protein accumulation), or 0.124 µm/pixel (segment boundary crossing). For analysis of MT dynamics and organization, and EB1 accumulation, images acquisition was made with an inverted microscope (Eclipse Ti-E; Nikon), equipped with a CFI Apochromat total internal reflection fluorescence (TIRF) 100× 1.49 NA oil objective lens (Nikon) and a motorized confocal head (CSU-X1-A1; Yokogawa). 16-bit images were projected onto the CCD chip at a magnification of 0.045 µm/pixel. Image acquisition was done with FV10-ASW software for the upright confocal microscope (FV1000; Olympus) or with MetaMorph software (http://www.moleculardevices.com/Products/Software/Meta-Imaging-Series/MetaMorph.html; Molecular Devices) for the inverted microscope (Eclipse Ti-E; Nikon).
For live imaging, we dechorionated embryos in 50% bleach, washed in water, and embedded in halocarbon oil 27 (Sigma-Aldrich). FRAP experiments were performed as described previously (Cavey et al., 2008). In each embryo, several circular regions of 1 µm or 0.5 µm radius were photobleached at junctions so that there was only one bleach event per cell. Photobleaching was performed with 12 scans at 8 µs/pixel at 100% 488 laser power, resulting in the reduction of E-cad–GFP signal by 60–80%. A stack of 6 z sections spaced by 0.38 µm was imaged just before photobleaching, and immediately after photobleaching, and then at 20-s intervals, generally for 15 min in total. For analysis of MT dynamics and EB1 accumulation, we used embryos carrying a maternally provided copy of the EB1-GFP transgene. Series of 100 images were taken with 0.1-s exposure at 0.5-s intervals.
Embryos were fixed in a 1:1 mixture of 4% formaldehyde in PBS/heptane for 20 min followed by removal of the vitelline envelope in 1:1 methanol/heptane. To stain MTs the embryos were fixed for 2 min in 1:1 4% EDTA in ice-cold methanol/heptane. The primary antibodies used were rat anti–E-cad (1:100, DCAD2; Developmental Studies Hybridoma Bank [DSHB]), rabbit anti-Baz (1:1,000; provided by A. Wodarz, University of Göttingen, Göttingen, Germany), and mouse anti–α-tubulin (1:500, 12G10; DSHB). Direct fluorescence of EB1-DN-mCherry, CD8-mCherry, myr-GFP, MyoII-YFP, and Rok-Venus proteins was used in fixed embryos. The secondary antibodies used were either Alexa Fluor 488 conjugated (Jackson ImmunoResearch Laboratories) or Cy3 conjugated (Invitrogen).
Data analysis
FRAP.
The recovery curves were obtained by manually measuring intensities of background, control region, and photobleached region using 2-µm or 1-µm diameter circular regions for each time point in Fiji software (http://fiji.sc/wiki/index.php/Fiji). For each genotype at least 15 individual recovery curves from at least five embryos were taken for each type of border (for exact numbers see Table S1). Each replicate value in each dataset was considered as an individual point for curve fitting. GraphPad Prism software (http://www.graphpad.com/) was used for nonlinear fitting and plotting on graphs. The recovery was fitted to a biexponential of the form
and to a single exponential of the form f(t) = 1 − Fim − A1 × e−t/Tfast, where Fim is a size of the immobile pool of the protein, Tfast and Tslow are the half times, and A1 and A2 are the amplitudes of the fast and slow components of the recovery. An F-test was used to choose the equation and compare datasets.
Mean fluorescence levels and MTs dynamics.
The junctional protein accumulation was obtained by manually measuring mean gray values at 20 junctions of each type in Fiji in five embryos using a circular region of 1-µm diameter spanning the cell–cell junction. A mean gray value of 10 measurements of the background done using a circular region of 1-µm diameter was subtracted from each measurement at the junction. Colocalization was tested with the colocalization plugin (http://rsbweb.nih.gov/ij/plugins/colocalization.html) in Fiji.
To measure fluorescence intensity and the amount of EB1-GFP, and to obtain dynamic properties of MTs plus ends, 10 time-lapse movies from five embryos were used: two movies from each embryo recording different regions with control and EB1-DN–expressing cells covered in each region. To measure fluorescence intensity and the amount of EB1-GFP, the first frame of each time series was used. A “3D object counter” plugin (http://rsbweb.nih.gov/ij/plugins/track/objects.html) was run on the same size regions expressing EB1-DN and not expressing it for each image. The number of detected objects divided by the size of the region was used as a measure of density. The mean intensity of each object was normalized to the mean intensity of the objects in the region that did not express EB1-DN for each image to exclude possible changes caused by photobleaching. To obtain dynamic properties of MTs, 10 kymographs with clearly distinguishable beginnings and ends of tracks were created using the Multiple Kymograph plugin (http://www.embl.de/eamnet/html/body_kymograph.html) and the “read velocity from tsp” macro (http://www.embl.de/eamnet/html/body_kymograph.html) in Fiji for each time-lapse movie. The nonparametric Mann-Whitney-Wilcoxon test was used in GraphPad Prism to compare the datasets.
MT organization.
To measure the polarization of MT, the angle of growth direction of each EB1-GFP–positive MT that was used to make a kymograph was measured relative to the embryo borders. The diagrams were created with our MatLab script. Statistical analysis was done in Excel (Microsoft) using an F-test to compare with a random distribution or Levene’s test to assess the equality of variances in different samples.
Rosette formation and border crossing.
The number of cells outside of the segment borders and types of contacts formed by the cells just anterior to the segment border were manually counted in Fiji in three stripes in each of 45 embryos per genotype (90 embryos for CD8-Cherry expression). The number of cells was counted after applying a threshold to the fluorescent signal of myristoylated GFP, which was the same for all embryos. The statistical analysis was done in GraphPad Prism using a χ2 test or nonlinear fitting with Poisson distribution. 95% confidence intervals (CI) for proportions were obtained using GraphPad QuickCalcs (http://www.graphpad.com/quickcalcs/ConfInterval1.cfm).
Immunoprecipitation and Western blotting
The EB1-GFP–expressing embryos were homogenized in lysis buffer (50 mM Tris-HCl, pH 8, 150 mM NaCl, 0.5% Triton X-100, 1 mM MgCl2, and complete proteinase inhibitor cocktail, EDTA-free [Roche]), cleared from cuticle debris by centrifugation for 3 min at 3,000 g, and loaded onto protein G Sepharose (GE Healthcare), which was preincubated for 2 h with one of the following antibodies: 3 µl of rabbit anti-GFP antibody (∼6 µg; Invitrogen), 10 µl of rat anti–E-cad (supernatant, DCAD2; DSHB), 10 µl of mouse anti–α-catenin (supernatant, DCAT-1; DSHB), or 2 µl of rabbit anti-Baz (provided by A. Wodarz). After overnight incubation at 4°C, Sepharose was washed three times in lysis buffer and then eluted with 15 µl of SDS sample buffer.
Online supplemental material
Fig. S1 shows that down-regulation of β-catenin (arm) reduces mobile and immobile pools of E-cad. Fig. S2 shows that Eb1 mutants and EB1-DN cause similar changes to E-cad distribution and dynamics. Fig. S3 shows that cells cross the segment boundary in pairs at a steady rate. Fig. S4 shows a model of regulation of mobile E-cad distribution by dynamic MTs with the changes caused by EB1-DN and Spas diagrammed. Video 1 shows epidermis at stage 15 of embryo development expressing EB1-GFP (black) tracking MT plus ends in control and cells expressing EB1-DN. Table S1 gives numerical values of protein accumulation at cell borders and best-fit parameters for FRAP experiments. Additional data are available in the JCB DataViewer at http://dx.doi.org/10.1083/jcb.201211159.dv.
Acknowledgments
We thank John Overton (for injection of constructs and maintenance of fly stocks) and the Gurdon Institute Imaging Facility.
This work was supported by grant RGP0002/2008-C from the Human Frontier Science Program and grant BB/K00056X/1 from the UK Biotechnology, Biological Sciences Research Council. Core funding was provided by the Wellcome Trust (092096) and Cancer Research UK (C6946/A14492). A.S. Yap is a Research Fellow of the National Health and Medical Research Council of Australia (631383).




![Figure 5. EB1-DN and Spas change MT organization and dynamics. (A) Spas eliminated MTs in epidermal cells at the level of E-cad junctions (wild type [wt], Spas expressed in a stripe of two cells; α-tubulin, magenta; E-cad, green). (B) EB1 domains, showing MT-binding CH, coil-coiled (CC) dimerization, and acidic C terminus (Ac) are depicted, as well as new constructs with GFP and Cherry tags. (C) Sample kymographs showing growth of individual MTs in wild-type (wt) and EB1-DN cells. (D–G) EB1-DN reduced processivity of MT growth and the number of growing ends. Quantitation of individual growth event duration (D), MT growth speed (E), EB1-GFP fluorescence intensity at individual plus ends (F), and plus end density per square micrometer at the level of E-cad junctions (G). (H and I) EB1-DN changed oriented growth of MTs. (H) Projection of 20 images taken every 0.5 s. EB1-GFP, black/green; EB1-DN, magenta. (I) Quantitation of growth orientation of individual MTs in control and EB1-DN cells (Cherry-positive in H) relative to dorsal (D), ventral (V), anterior (A), and posterior (P) sides of the embryos. Red lines mark 45° segments. Bars, 5 µm. ***, P < 0.0001.](https://cdn.rupress.org/rup/content_public/journal/jcb/201/6/10.1083_jcb.201211159/5/m_jcb_201211159_fig5.jpeg?Expires=1781714605&Signature=gIeUye9qp8g79tyL0MXuBoNV8Ps1xnOc6tdhAGQlH~qvGsg1RP0qVyY5ayOvUJsHRJKtEFXHBjM14fOy2184-P64fIcHzYzVljuKT-Af0wzYqDDEeCgZ7wbrJN2q-EwER65cEvYgn4g2rqszXhRHvndY6e2LvXW6YexNDBDGOcrk6yS1O9yd0ABoMvefB2pt21R1MucZR4jOdwSFeXjy0tLsgO-A1A~f6GFBGNZ94bTWRkag-sFlskUmKRPo8BXV0yrz0KfJ8omvFBNFOjndPxJXux1zVIi6AAnIDhaFwoLMcqUC-K95pvg-Znodq0Znx0CArh4FGDSgk09OkHlGLQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)







