Many mitotic factors were shown to be activated by Ran guanosine triphosphatase. Previous studies in Xenopus laevis egg extracts and in highly proliferative cells showed that mitotic chromosomes were surrounded by steep Ran guanosine triphosphate (GTP) concentration gradients, indicating that RanGTP-activated factors promote spindle assembly around chromosomes. However, the mitotic role of Ran in normal differentiated cells is not known. In this paper, we show that although the steep mitotic RanGTP gradients were present in rapidly growing cell lines and were required for chromosome congression in mitotic HeLa cells, the gradients were strongly reduced in slow-growing primary cells, such as HFF-1 fibroblasts. The overexpression of RCC1, the guanine nucleotide exchange factor for Ran, induced steeper mitotic RanGTP gradients in HFF-1 cells, showing the critical role of RCC1 levels in the regulation of mitosis by Ran. Remarkably, in vitro fusion of HFF-1 cells produced cells with steep mitotic RanGTP gradients comparable to HeLa cells, indicating that chromosomal gain can promote mitosis in aneuploid cancer cells via Ran.
Introduction
Mitotic entry is marked by a strong increase in the dynamic instability of microtubules (MTs; Zhai et al., 1996), leading to increased MT dependence on local regulation. During prometaphase (PM), chromosome-, kinetochore-, and centrosome-centered mechanisms direct the self-assembly of MTs into the mitotic spindle and facilitate correct MT connections to kinetochores on each chromosome (Walczak and Heald, 2008; Wadsworth et al., 2011). In one model explaining the rapid MT–kinetochore attachments, the growth of centrosomal MTs toward kinetochores is promoted by a chromosomal gradient of MT stabilization activity (Wollman et al., 2005). In another model, such chromosomal signals promote MT growth within the clusters of PM chromosomes, accelerating the initially lateral MT–kinetochore attachments in PM (Magidson et al., 2011). In both models, chromosomes could contribute to their mitotic segregation by activating spindle assembly factors (SAFs) through Ran GTPase (Clarke and Zhang, 2008; Kaláb and Heald, 2008).
The chromatin binding of RCC1, the guanine nucleotide exchange factor for Ran, and the cytoplasmic localization of RanGAP1 drive the rise of a concentration gradient of RanGTP surrounding the mitotic chromosomes. The binding of RanGTP diffusing from chromosomes to its ligands induces downstream gradients, including a gradient of SAFs activated by their RanGTP-induced release from importins (Kaláb and Heald, 2008). Although the RanGTP or RanGTP-regulated gradients were detected in meiotic Xenopus laevis egg extracts, maturing mouse oocytes, and tissue-culture cell lines (Kaláb et al., 2002, 2006; Caudron et al., 2005; Dumont et al., 2007), the mitotic role of Ran in normal somatic cells is not known.
Results and discussion
Cell type–specific diversity of the mitotic RanGTP and importin-β cargo gradients
To determine whether the RanGTP gradient supports mitosis in all human somatic cells or is an adaptation specific to certain kinds of cells, we measured RanGTP gradients in a panel of human cells, including primary cells, immortalized normal cells, cancer-derived cells, and tumorigenic cells (Fig. 1 and Table S1). These measurements were performed with fluorescence lifetime imaging microscopy (FLIM) using two previously developed Förster resonance energy transfer (FRET) sensors (Kaláb et al., 2002, 2006) with the donor–acceptor pairs replaced by mTFP-1 (Ai et al., 2008) and dsREACh (Materials and methods). For both sensors, we used live-cell FLIM measurements of their donor fluorescence lifetime (τdonor) to calculate FRET efficiency E using E = 1 − τdonor/τdonor REF (Sun et al., 2011), in which the τdonor REF = 2,519 ps is the mean τdonor of mTFP-1 expressed in cells in the absence of the acceptor (Fig. S1, E and F).
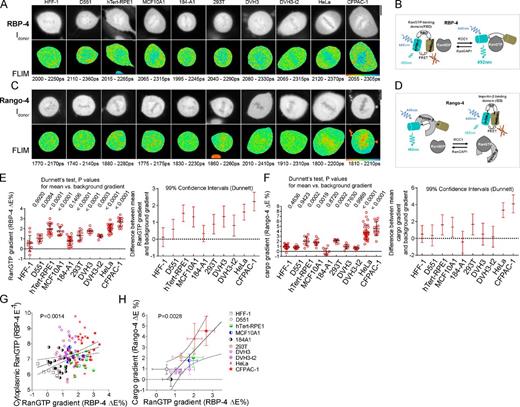
Cell-specific diversity of mitotic RanGTP and cargo gradients. (A and C) Mitotic RanGTP gradients detected with RBP-4 (A) and cargo gradients detected with Rango-4 (C) by FLIM in different cells. The top rows show the donor intensity Idonor, and bottom rows show the pseudocolor FLIM images. The range of the displayed τdonor values is indicated beneath the FLIM images. (B and D) Schematic of RBP-4 (B) and Rango-4 (D). (E and F, left) Scatter plots of the mitotic RanGTP gradients (E) and the cargo gradients (F) quantified as the difference between the cytoplasmic and chromatin E (ΔE; single-cell data, means ± SD). For each sensor and cell type, the gradients were compared by ANOVA/Dunnett with background gradient detected using an inactive FRET sensor (Fig. S1, E and F). Adjusted p-values for the difference between mean gradients and background gradient are shown above the scatter plots. (E and F, right) Dunnett’s test 99% confidence intervals for the difference between mean gradients and background gradient. (G and H) Regression analysis of the RanGTP gradient and cytoplasmic RanGTP levels (G) and of the RanGTP and cargo gradients (H; means ± SD). Dotted lines show linear regression slope ± 99% confidence band. Bars, 10 µm.
Cell-specific diversity of mitotic RanGTP and cargo gradients. (A and C) Mitotic RanGTP gradients detected with RBP-4 (A) and cargo gradients detected with Rango-4 (C) by FLIM in different cells. The top rows show the donor intensity Idonor, and bottom rows show the pseudocolor FLIM images. The range of the displayed τdonor values is indicated beneath the FLIM images. (B and D) Schematic of RBP-4 (B) and Rango-4 (D). (E and F, left) Scatter plots of the mitotic RanGTP gradients (E) and the cargo gradients (F) quantified as the difference between the cytoplasmic and chromatin E (ΔE; single-cell data, means ± SD). For each sensor and cell type, the gradients were compared by ANOVA/Dunnett with background gradient detected using an inactive FRET sensor (Fig. S1, E and F). Adjusted p-values for the difference between mean gradients and background gradient are shown above the scatter plots. (E and F, right) Dunnett’s test 99% confidence intervals for the difference between mean gradients and background gradient. (G and H) Regression analysis of the RanGTP gradient and cytoplasmic RanGTP levels (G) and of the RanGTP and cargo gradients (H; means ± SD). Dotted lines show linear regression slope ± 99% confidence band. Bars, 10 µm.
To measure free RanGTP, we used RBP-4 (RanGTP-binding probe-4, modified YFP–RanGTP-binding domain (RBD)–CFP; Kaláb et al., 2002), which indicates RanGTP binding by decreased E (Fig. 1, A and B). We quantified the mitotic RanGTP gradient by subtracting the mean chromatin RBP-4 E from the cytoplasmic E (RBP-4 ΔE; Fig. 1 E), and we used the inverse of cytoplasmic RBP-4 E (RBP-4 E−1) as a measure of cytoplasmic free RanGTP levels (Fig. 1 G). Separately, we used FLIM of Rango-4 (modified Rango; Kaláb et al., 2006) to measure the RanGTP-regulated gradient of free importin-β cargoes. Because Rango-4 signals its RanGTP-induced release from importin-β by increased E (Fig. 1, C and D), we quantified the free cargo gradient by subtracting mean cytoplasmic Rango-4 E from its E at the chromatin (Rango-4 ΔE; Fig. 1 F).
Our screen revealed a striking, cell-specific diversity of mitotic RanGTP and cargo gradients (Fig. 1). Rango-4 FLIM showed that mitotic HeLa cell chromosomes were surrounded by a free cargo gradient as previously observed (Kaláb et al., 2006; O’Connell et al., 2009; Soderholm et al., 2011), and RBP-4 FLIM detected the expected upstream RanGTP gradient (Kaláb et al., 2002). However, only in pancreatic adenocarcinoma-derived CFPAC-1 cells were both gradients as steep as in HeLa cells. The RanGTP gradient was at background level in HFF-1 fibroblasts and in 184-A1 breast epithelial cells and reduced in D551 (Detroit-551) fibroblasts (Fig. 1, A and E). A gradient steeper than that in D551 cells was present in tumorigenic DVH3 cells derived from a virally induced D551 cell fusion and in metastatic DVH3-t2 cells that were isolated from DVH3 cell–induced tumors (Duelli et al., 2007). A steep RanGTP gradient was detected in human telomerase reverse transcriptase (hTert)–RPE1 cells (normal retinal pigmented epithelial cells immortalized by telomerase), 293T cells (human embryonic kidney cells immortalized by adenoviral DNA), and immortalized breast epithelial cells MCF10A1 (Fig. 1, A and E). The amplitude of the RanGTP gradient correlated with increased cytoplasmic RanGTP levels (P = 0.0014; Fig. 1 G), showing that RanGTP could reach farther to the cytoplasm in cells with steep RanGTP gradients. Not surprisingly, given the absence of a RanGTP gradient, no mitotic cargo gradient was detected in HFF-1 and 184-A1 cells. However, the cargo gradient was also absent in D551, DVH3, and DVH3-t2 cells (Fig. 1 F), indicating their shared disconnect between the RanGTP and cargo gradients (Fig. 1, E, F, and H). The steep mitotic RanGTP gradient is therefore not universal in human somatic cells, and its coupling to the gradients of free importin-β cargoes is also cell type specific, raising questions about the causes and physiological role of such diversity.
Although FLIM provided sensitive and quantitative measurements, it required a long acquisition time (2 min) and complex data analysis. We therefore validated our results by using the more traditional emission intensity ratio method of FRET detection, after we replaced the nonfluorescent acceptor in our sensors with YPet (Nguyen and Daugherty, 2005). In mitotic cells expressing one of the sensors, we then measured FRET, first by the emission ratio technique and then by FLIM. FRET images obtained by both techniques showed the presence of RanGTP and cargo gradients in HeLa cells and no spatial gradients in HFF-1 cells (Fig. S1, A and B), validating our FLIM analyses.
High RCC1 levels and binding to chromatin drive steep mitotic RanGTP gradients
To identify the causes of the RanGTP gradient diversity, we first compared the levels and modifications of Ran and its regulators using immunoblots from lysates of nonsynchronized cells (Fig. 2 A). Interestingly, cells with small or nondetectable RanGTP gradients (HFF-1, D551, and 184-A1) shared low RCC1 levels and, consistent with their slower growth in culture, low levels of the mitotically phosphorylated Ser 10 on histone H3 and low levels of TPX2, the major RanGTP-regulated SAF, which is degraded at the exit from mitosis (Fig. 2 A; Stewart and Fang, 2005). To examine the role of such differences in RanGTP gradient formation, we focused further analyses on the comparison of HFF-1 and HeLa cells.
Differences in RCC1 levels and N-terminal methylation contribute to the cell-specific diversity of mitotic RanGTP gradients. (A and B) Immunoblots showing protein expression levels and posttranslational modifications in nonsynchronized cells (A) and in HFF-1 and HeLa cells synchronized by NZ shake-off in mitosis or interphase (B). As indicated, the position of methylated RCC1 (Me3-RCC1α) and phosphorylated RanGAP1 (p-RanGAP1-SUMO) are shown. Corresponding loading control data for each separate gel are shown. Asyn, asynchronized; Inter, interphase; Mito, mitosis. (C) IF of RCC1 in HeLa and HFF-1 cells. (D) Scatter plot of RCC1 chromatin/cytoplasmic ratio detected by IF in mitotic HeLa and HFF-1 cells. (E) Confocal fluorescence images of live cells expressing NTDα-mCherry. (F) Scatter plot of chromatin/cytoplasmic ratio of NTDα-mCherry in live cells. (G) FRAP measurements of wt and ASPK RCC1α-mCherry binding to chromatin in interphase (left) and mitotic cells (right). D, F, and G are single-cell data, means ± SD; t test in D; ANOVA/Bonferroni in F and G. In F, all pairwise comparisons are significant (P < 0.01), except for two pairs indicated (NS). In G, only significant pairwise comparisons are indicated. (H) mCherry fluorescence, donor intensity (Idonor), and FLIM images of mitotic HFF-1 cells coexpressing wt (top) or S2K RCC1α-mCherry and Rango-4. Bars, 10 µm.
Differences in RCC1 levels and N-terminal methylation contribute to the cell-specific diversity of mitotic RanGTP gradients. (A and B) Immunoblots showing protein expression levels and posttranslational modifications in nonsynchronized cells (A) and in HFF-1 and HeLa cells synchronized by NZ shake-off in mitosis or interphase (B). As indicated, the position of methylated RCC1 (Me3-RCC1α) and phosphorylated RanGAP1 (p-RanGAP1-SUMO) are shown. Corresponding loading control data for each separate gel are shown. Asyn, asynchronized; Inter, interphase; Mito, mitosis. (C) IF of RCC1 in HeLa and HFF-1 cells. (D) Scatter plot of RCC1 chromatin/cytoplasmic ratio detected by IF in mitotic HeLa and HFF-1 cells. (E) Confocal fluorescence images of live cells expressing NTDα-mCherry. (F) Scatter plot of chromatin/cytoplasmic ratio of NTDα-mCherry in live cells. (G) FRAP measurements of wt and ASPK RCC1α-mCherry binding to chromatin in interphase (left) and mitotic cells (right). D, F, and G are single-cell data, means ± SD; t test in D; ANOVA/Bonferroni in F and G. In F, all pairwise comparisons are significant (P < 0.01), except for two pairs indicated (NS). In G, only significant pairwise comparisons are indicated. (H) mCherry fluorescence, donor intensity (Idonor), and FLIM images of mitotic HFF-1 cells coexpressing wt (top) or S2K RCC1α-mCherry and Rango-4. Bars, 10 µm.
In asynchronous cell cultures, the levels of RanGAP1 were similar in HFF-1 and HeLa cells (the HFF-1/HeLa ratio of the RanGAP1 + RanGAP1–small ubiquitin-like modifier (SUMO) signal was 98 ± 5%; mean ± SD, n = 4; three repeats; Fig. 2 A). However, HFF-1 cells contained less Ran (71 ± 2% HFF-1/HeLa) and markedly lower levels of RCC1 (17 ± 3% HFF-1/HeLa) and TPX2 (10 ± 3% HFF-1/HeLa; mean ± SD, n ≥ 4 for all comparisons; three or four repeats; Fig. 2 A).
Immunoblots from cells synchronized by nocodazole (NZ) shake-off showed that interphase and mitotic HeLa cells had similar RCC1 levels (the ratio of mitotic/interphase RCC1 was 92 ± 20%; mean ± SD, n = 2; one repeat; Fig. 2 B), but in HFF-1 cells, there was more RCC1 in mitotically synchronized than in interphase cells (171 ± 1% mitotic/interphase HFF-1 cells; Fig. 2 B). It is therefore unlikely that the much lower RCC1 levels in asynchronous HFF-1 cells versus asynchronous HeLa cells (Fig. 2 A) resulted from a lower fraction of mitotic cells in the HFF-1 cell population.
Ran localization was similar between the two cells (Fig. S2 A). However, the immunofluorescence (IF) showed low RanGAP1 levels at the kinetochores (Fig. S2 B) and strongly reduced RCC1 staining on the chromosomes (Fig. 2, C and D) in HFF-1 cells compared with HeLa cells. Because RCC1-chromatin binding drives RanGTP gradient formation, we examined this further.
Both the C-terminal catalytic domain of RCC1 and its short flexible N-terminal domain (NTD) directly bind to DNA and histones in the nucleosome (Makde et al., 2010), where the chromatin binding of the C terminus is modulated by the loading of Ran on RCC1 (Li et al., 2003), and the chromatin binding of the NTD provides crucial support (Hitakomate et al., 2010). The binding of the RCC1 N terminus to DNA requires α-N-methylation of Ser 2, which is mediated by the N-terminal RCC1 methylase (NRMT; Chen et al., 2007; Schaner Tooley et al., 2010). In addition, three RCC1 splicing isoforms (RCC1α, β, and γ) expressed in human cells differ in their NTD composition and in how their binding to chromatin is affected by Ser 11 phosphorylation (Hood and Clarke, 2007).
Because quantitative PCR showed that the phosphorylation-sensitive RCC1γ isoform (Hood and Clarke, 2007) was the least abundant and that RCC1α was the prevailing isoform (Fig. S2 C), we focused on RCC1α and its methylation. Although immunoblotting with RCC1 α-N-trimethylation antibodies (Me3-SPK; Schaner Tooley et al., 2010) showed that, relative to RCC1 protein levels, the Me3-SPK on RCC1 was similar in HeLa and HFF-1 cells (41 ± 7% Me3-SPK HFF-1/HeLa), this comparison was limited by residual reactivity of the Me-3 antibody with nonmethylated RCC1 (Fig. 2 A and Fig. S2 E). We therefore analyzed the role of RCC1 N-terminal methylation using RCC1α-mCherry reporters expressed in HeLa and HFF-1 cells. As expected (Chen et al., 2007; Hitakomate et al., 2010; Schaner Tooley et al., 2010), the nonmethylatable ASPK-NTDα-mCherry (Schaner Tooley et al., 2010) did not bind to mitotic chromosomes (Fig. 2, E and F). However, the binding of wild-type (wt) NTDα-mCherry to mitotic chromosomes was weaker in HFF-1 cells, and the S2K NTDα-mCherry reporter carrying methyl-mimetic substitution showed stronger chromosome binding in both cells (Fig. 2, E and F).
We verified the role of N-terminal methylation in full-length RCC1-chromatin binding using FRAP (Fig. 2 G and Fig. S2, F and G). In cells expressing wt or ASPK RCC1α-mCherry, we photobleached an area of the nucleus or mitotic chromosomes and measured the FRAP t1/2 (Fig. 2 G and Fig. S2, F and G), which is inversely proportional to the chromatin binding of the fluorescent reporter (Hitakomate et al., 2010). In mitotic and interphase cells, the binding of wt RCC1α-mCherry to chromatin was stronger in HeLa than in HFF-1 cells. The ASPK mutation strongly reduced the RCC1-chromatin binding in HeLa but had almost no effect on the already weak binding in HFF-1 cells (Fig. 2 G). The lower α–N-terminal methylation therefore significantly contributed to reduced RCC1 binding to chromatin in HFF-1 cells throughout the cell cycle.
To examine the role of RCC1 levels in RanGTP gradient formation, we measured the mitotic cargo gradients in HFF-1 cells coexpressing Rango-4 and RCC1α-mCherry. We found that the expression of the wt and particularly of the S2K RCC1α-mCherry induced detectable mitotic cargo gradients in HFF-1 cells (Fig. 2 H), showing that increased RCC1 levels are sufficient to drive the formation of steeper mitotic RanGTP gradients.
However, the amplitude of the RanGTP gradient is expected to be sensitive to the balance of the cellular concentrations of Ran and its regulators (Caudron et al., 2005; Kaláb et al., 2006). For example, the smaller amplitudes of mitotic RanGTP gradients in DVH3 and DVH3-t2 cells than in HeLa cells (Fig. 1, A and E) then potentially could be explained by the combined effect of lower RCC1 levels and higher RanGAP1 levels, together with slightly lower Ran levels (Fig. 2 A). Specifically, relative to HeLa cells, the Ran levels were 81 ± 17% in DVH3 cells and 73 ± 12% in DVH3-t2 cells. At the same time, relative to HeLa cells, DVH3 cells contained 141 ± 32% total RanGAP1 (sum of RanGAP1 + RanGAP1-SUMO) and 68 ± 19% RCC1. Similarly, DVH3-t2 cells contained 126 ± 12% total RanGAP1 and only 69 ± 14% RCC1 compared with HeLa (all aforementioned relative expression values were derived from data in Fig. 2 A and its biological repeats; n = 4). Future studies will be needed to verify this model and what causes the absence of the cargo gradient in certain cells.
Steep mitotic RanGTP gradient promotes bipolar spindle assembly in PM
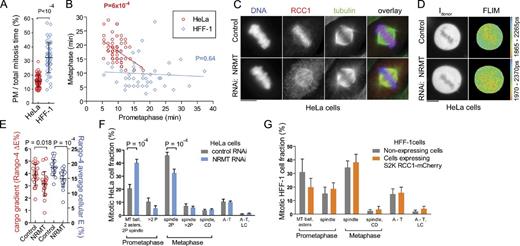
To examine how differences in RanGTP gradient steepness affect mitosis, we first compared mitotic progression in unperturbed HeLa and HFF-1 cells by live-cell microscopy (Fig. 3, A and B; and Fig. S3 A). Although the mitosis was shorter in HFF-1 cells (54 ± 12 vs. 61 ± 7 min in HeLa; P = 0.003), the PM time in HFF-1 cells was longer and more variable (17 ± 7 vs. 9 ± 2 min in HeLa; P < 10−4; Fig. 3 A and Fig. S3 A), indicating that the chromosome congression was slower and more stochastic in HFF-1 cells. Metaphase was longer in HeLa cells (18 ± 4 vs. 10 ± 5 min in HFF-1 cells; P < 10−4; Fig. S3 A), and unlike in HFF-1 cells, its duration in individual HeLa cells was inversely proportional to their PM time (Fig. 3 B), perhaps indicative of a longer time needed for sorting erroneous MT–kinetochore attachments formed during rapid spindle assembly.
Mitotic RanGTP gradient promotes rapid spindle assembly during PM. (A and B) Time-lapse imaging of untreated cells was used to measure the duration of mitotic phases (see also Fig. S3 A). (A) Scatter plot of PM/total mitosis time in HeLa and HFF-1 cells (single-cell data, means ± SD, t test). (B) Linear regression of the PM versus metaphase time in HeLa and HFF-1 cells. Best-fit regression lines and the corresponding p-values of the fits are indicated. (C) RCC1 IF in control or NRMT RNAi-treated HeLa cells. (D) Rango-4 donor intensity (Idonor) and FLIM images of control or NRMT RNAi-treated HeLa cells. (E) Scatter plot of mitotic cargo gradients and mean cellular Rango-4 E. Single-cell data, means ± SD, t test. (F and G) The fraction of mitotic cells in PM, metaphase, anaphase, and telophase (A-T) detected by IF in NRMT or control RNAi-treated HeLa cells (F) and in HFF-1 cells expressing or not expressing S2K RCC1α-mCherry (G). CD, chromosome congression defect; LC, lagging chromosomes; 2P, two poles or asters; >2P, multipolar spindles and asters. Means ± SD, ANOVA/Bonferroni test; n = 4 in F and n = 3 in G. Only significant changes are indicated. Bars, 10 µm.
Mitotic RanGTP gradient promotes rapid spindle assembly during PM. (A and B) Time-lapse imaging of untreated cells was used to measure the duration of mitotic phases (see also Fig. S3 A). (A) Scatter plot of PM/total mitosis time in HeLa and HFF-1 cells (single-cell data, means ± SD, t test). (B) Linear regression of the PM versus metaphase time in HeLa and HFF-1 cells. Best-fit regression lines and the corresponding p-values of the fits are indicated. (C) RCC1 IF in control or NRMT RNAi-treated HeLa cells. (D) Rango-4 donor intensity (Idonor) and FLIM images of control or NRMT RNAi-treated HeLa cells. (E) Scatter plot of mitotic cargo gradients and mean cellular Rango-4 E. Single-cell data, means ± SD, t test. (F and G) The fraction of mitotic cells in PM, metaphase, anaphase, and telophase (A-T) detected by IF in NRMT or control RNAi-treated HeLa cells (F) and in HFF-1 cells expressing or not expressing S2K RCC1α-mCherry (G). CD, chromosome congression defect; LC, lagging chromosomes; 2P, two poles or asters; >2P, multipolar spindles and asters. Means ± SD, ANOVA/Bonferroni test; n = 4 in F and n = 3 in G. Only significant changes are indicated. Bars, 10 µm.
Next, we examined whether the RanGTP gradient is responsible for such differences by inducing changes in the RanGTP gradient and following their effects on mitosis. We reduced the RanGTP gradient by decreasing RCC1-chromatin binding via NRMT RNAi in HeLa cells (Schaner Tooley et al., 2010). As previously reported (Schaner Tooley et al., 2010), the NRMT RNAi reduced the RCC1 methylation (Fig. S3 B) and disrupted RCC1-chromatin binding (Fig. 3 C). Although NRMT RNAi induced a subtle increase of RCC1 levels (to 151 ± 27%; Fig. S3 B), the RanGTP gradient was reduced by the treatment, as shown by the decreased mitotic cargo gradient and decreased mean free Rango-4 in mitotic cells (Fig. 3, D and E). The NRMT knockdown induced a significant increase in the PM fraction of mitotic HeLa cells (31.2 ± 2.7% control vs. 45.9 ± 4.8% NRMT RNAi; P < 10−3; Fig. 3 F and Fig. S3 C), indicating that the steep RanGTP gradient is required for the assembly of a fully functional bipolar spindle capable of driving the chromosome congression during PM. We confirmed these results by perturbing the RanGTP gradient via RanGAP1 RNAi in HeLa cells and via RCC1 inhibition in the temperature-sensitive tsBN2 cell line (Fig. S3, E–I; Li and Zheng, 2004). Consistent with previous studies (Joseph et al., 2002; Arnaoutov and Dasso, 2005), the depletion of RanGAP1 and RCC1 both induced strong defects in chromosome congression and segregation (Fig. S3, F–I). Both interventions also caused the accumulation of PM cells (Fig. S3, F–I), supporting the results of the NRMT knockdown.
Because of the low TPX2 levels in HFF-1 cells (Fig. 2, A and B), we expected that increasing the RanGTP gradient by RCC1 overexpression in HFF-1 cells (Fig. 2 H) might have only a subtle effect on spindle assembly and function. Indeed, S2K-RCC1α-mCherry overexpression caused only a statistically insignificant decrease in PM cells with astral MTs (19.9 ± 6.5 vs. 31.1 ± 9.6%; two-way analysis of variance [ANOVA], P > 0.05; Fig. 3 G). The activation of the RanGTP gradient alone therefore may not be sufficient to accelerate the assembly of a fully functional mitotic spindle.
Chromosomal gain drives steep mitotic RanGTP gradient
Because slower diffusion is expected to increase the steepness of the RanGTP gradient (Caudron et al., 2005), we wondered whether chromosomal crowding in cells with high chromosome numbers contributes to their steep mitotic RanGTP gradient. To test this, we measured mitotic RanGTP gradients in polyploid HFF-1 cells prepared in vitro (Jansen et al., 2007) by fusion between HFF-1 cells expressing RBP-4 and HFF-1 cells expressing MT plus end–binding protein EB3-mCherry. Remarkably, fused mitotic HFF-1 cells displayed steep RanGTP gradients comparable with mitotic HeLa cells (compare Fig. 4 B and Fig. 1 E), demonstrating that chromosomal gain is sufficient to drive the formation of a steep mitotic RanGTP gradient.
Chromosomal gain drives steep mitotic RanGTP gradient. (A) mCherry fluorescence, donor intensity (Idonor), and FLIM images of mitotic cells resulting from the fusion of HFF-1 cells expressing EB3-mCherry with HFF-1 cells expressing RBP-4 (bottom). In the control HFF-1 cells expressing RBP-4 (top), the PEG-induced cell fusion was omitted. The displayed τdonor range is shown on the right. (B) Scatter plot of the mitotic RanGTP gradients (RBP-4 ΔE) in control and fused HFF-1 cells (single-cell data, means ± SD, t test). Bar, 10 µm.
Chromosomal gain drives steep mitotic RanGTP gradient. (A) mCherry fluorescence, donor intensity (Idonor), and FLIM images of mitotic cells resulting from the fusion of HFF-1 cells expressing EB3-mCherry with HFF-1 cells expressing RBP-4 (bottom). In the control HFF-1 cells expressing RBP-4 (top), the PEG-induced cell fusion was omitted. The displayed τdonor range is shown on the right. (B) Scatter plot of the mitotic RanGTP gradients (RBP-4 ΔE) in control and fused HFF-1 cells (single-cell data, means ± SD, t test). Bar, 10 µm.
Virus-induced cell fusion (Duelli et al., 2007), mitotic slippage, or cytokinesis failure is thought to produce unstable tetraploid precursors of aneuploid cancer cells (Ganem et al., 2007; Vitale et al., 2011). The chromosomal gain-driven mitotic RanGTP gradient in such intermediates could selectively promote proliferation of intermediates expressing Ran-regulated mitotic factors. Intriguingly, the transcriptional analysis in human breast cancers and in mouse models identified a conserved TPX2-centered expression network that strongly predicted metastasis susceptibility in both species (Hu et al., 2012).
Our results indicate that the diversity of mitotic RanGTP gradients can be understood if Ran is considered as a component of a highly dynamic and spatially organized intracellular system. Although elevated RCC1 levels and increased chromosome numbers are important for the formation of the gradient, exceptions, such as in the case of DVH3 and DVH3-t2 cells (as discussed for data in Fig. 1, A and E; and Fig. 2 A), are to be expected from such a systems-level perspective. An important question left for the future is the physiological role of the regulation of mitotic RanGTP gradient. The steep RanGTP gradient could support cell cycle progression by counteracting mechanisms that sense prolonged PM and induce G1/S cell cycle arrest in the next cell cycle (Uetake and Sluder, 2010). Because the levels of RCC1 and Ran do not significantly change between interphase and mitosis (Fig. 2 B), cells with steep mitotic RanGTP gradients likely contain higher levels of nuclear RanGTP after the exit from mitosis, possibly facilitating the G1/S checkpoint transition via the interphase functions of Ran.
Materials and methods
Cell culture
tsBN2 cells were obtained from T. Nishimoto (Kyushu University, Fukuoka, Japan) and from M. Dasso (National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD) and were grown at 33°C in 5% CO2. All other cells were grown at 37°C in 5% CO2. Human adenocarcinoma epithelial cells (HeLa; CCL-2) were obtained from American Type Culture Collection and used to isolate the single cell–derived clone HeLa 6 (Kaláb et al., 2006; Kaláb and Soderholm, 2010), which was used throughout this study. Human ductal adenocarcinoma-derived CFPAC-1 cells (CRL-1918), human embryonic kidney 293T cell line (CRL-11268), normal newborn human male foreskin fibroblasts (HFF-1; SCRC-104), normal human skin fibroblast D551 (CCL-110), and telomerase-immortalized normal retinal pigmented epithelial cells hTert-RPE1 (CRL-4000) were purchased from American Type Culture Collection and cultured in 10% FBS/DMEM (Invitrogen). The MCF10A1 normal immortalized breast epithelial cells (Santner et al., 2001) were purchased from the Karmanos Cancer Institute and cultured in DMEM/F12 (1:1) with 5% horse serum, 0.029 mM Na bicarbonate, and 10 mM Hepes supplemented with 10 µg/ml insulin, 20 ng/ml EGF, 0.5 µg/ml hydrocortisone, and 100 ng/ml cholera toxin. The conditionally immortalized, nonmalignant breast epithelial cell line 184-A1 (a gift from M. Stampfer, University of California, Berkeley, Berkeley, CA; Garbe et al., 2009) was cultured in M87A + cholera toxin + oxytocin medium, which consists of 50% vol/vol mammary epithelial basal medium (Lonza), 50% vol/vol DMEM/F12, 7.5 µg/ml bovine insulin (Gemini Bio Products), 35 µg/ml bovine pituitary extract (Lonza), 0.3 µg/ml hydrocortisone (Sigma-Aldrich), 5 ng/ml EGF (Sigma-Aldrich), 2.5 µg/ml transferrin (Sigma-Aldrich), 5 µM isoproterenol (Sigma-Aldrich), 2 mM glutamine (Lonza), 5 nM tri-iodothyronine (Sigma-Aldrich), 0.5 nM β-estradiol (Sigma-Aldrich), 0.25% FBS (Invitrogen), 0.5 ng/ml cholera toxin (Sigma-Aldrich), 1 mg/ml AlbuMAX (Invitrogen), 0.75 mg/ml NaHCO3 (Sigma-Aldrich), and 0.1 nM oxytocin (Bachem). The human tumor-inducing DVH3 cell line and the mouse tumor-derived human tumorigenic cell line DVH3-t2 (Duelli et al., 2007) were gifts from D. Duelli (Rosalind Franklin Medical University, Chicago, IL) and were grown 10% FBS/DMEM.
IF
For IF, cells were grown on cleaned glass coverslips or on dishes (ibiTreat 60 µ-Dish; ibidi). After fixation with the antigen-specific method (see next paragraph), the cells were permeabilized for 2–5 min with 1% Triton X-100 and 2 µg/ml Hoechst 33342 (Sigma-Aldrich) in general tubulin buffer (GTB; 80 mM Pipes, pH 6.9, 1 mM MgCl2, and 1 mM EGTA) and blocked at 4°C for 1 h or overnight with IF buffer (IFB; 6% normal donkey serum [Jackson ImmunoResearch Laboratories, Inc.], 0.2% saponin [Sigma-Aldrich], and 0.02% sodium azide in GTB). Samples were incubated with primary antibodies diluted in IFB at room temperature for 1–2 h or overnight at 4°C followed by washes with 0.2% saponin/GTB and a 0.5–1-h incubation with secondary antibodies (dye-labeled purified donkey IgG; Jackson ImmunoResearch Laboratories, Inc.) diluted in IFB. After final washes in GTB containing 0.2% saponin and 0.1%Triton X-100, the samples were mounted using ProLong Gold Antifade (Invitrogen) and stored at −30°C or photographed immediately. RFP-Booster-Atto594 (ChromoTek) was used to detect mCherry-tagged RCC1 or mCherry in fixed cells for experiments in Fig. 3 G. For RCC1 and tubulin detection (Fig. 2 C and Fig. 3 C), cells were fixed for 2–5 min at −30°C in 5% acetic acid, 50% methanol, and 45% ethanol. For tubulin and DNA detection (Fig. S3, C, D, and G), the cells were fixed with freshly prepared 4% paraformaldehyde in GTB for 20 min at 37°C. For the detection of tubulin, RanGAP1, and DNA (Fig. S2 B), cells were fixed in 2% paraformaldehyde in GTB for 15 min at room temperature. The fixation and IF of tubulin and DNA in synchronized tsBN2 and BHK cells (Fig. S3 I) are described in the paragraph Analysis of mitosis progression in tsBN2 cells treated with nonpermissive/permissive temperature.
Antibodies
The following commercial antibodies were used for immunoblotting and IF assays: rabbit monoclonal to RCC1 (used for IF and immunoblotting; #5134; Epitomics), goat polyclonal to RCC1 (used for immunoblotting; sc-1162; Santa Cruz Biotechnology, Inc.), mouse monoclonal to Ran (#610340; BD), rabbit polyclonal to TPX2 (NB500-179; Novus Biologicals), mouse monoclonal E7 antibody to tubulin (Developmental Studies Hybridoma Bank), rabbit polyclonal to β-tubulin (#A01203; GenScript), human anticentromere antibody (HCT 0100; ImmunoVision), and rabbit monoclonal to phospho–Ser 10 in histone H3 (#E173; Epitomics). The rabbit polyclonal antibody to Me3-SPK (Chen et al., 2007; Schaner Tooley et al., 2010) was a gift from C. Tooley and I. Macara (University of Virginia, Charlottesville, VA), the affinity-purified rabbit polyclonal antibody to RanGAP1 was a gift from A. Arnaoutov and M. Dasso (National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD), and goat polyclonal antibody to RanGAP1 was a gift from F. Melchior (Heidelberg University, Heidelberg, Germany).
Cell transduction with lentiviruses and cell transfection
Lentiviral production was performed in house or contracted to the Protein Expression Laboratory, SAIC-Frederick, Inc., using protocols recommended by Invitrogen. Human embryonic kidney cells, 293FT (Invitrogen), were transfected with the expression vector and packaging mix (pLP1, pLP2, and pLP/vesicular stomatitis virus; ViraPower; Invitrogen) using Lipofectamine 2000. Lentivirus supernatant was harvested 3 d after transfection, snap frozen in liquid nitrogen, and stored at −80°C. On the day of transduction, an aliquot of the lentivirus stock was thawed on ice and added to growth medium containing Polybrene to obtain a final 6 µg Polybrene/milliliter of medium containing 50–200 µl of the viral stock/milliliter. For FRET sensor lentiviruses, the supernatant was concentrated 10 times using Lenti-X Concentrator (Takara Bio Inc.; Invitrogen), and the Polybrene concentration was lowered to 2 µg/ml to reduce cell stress. Cells were cultured in the virus-containing medium overnight, washed with Dulbecco’s PBS (DPBS), and returned to normal growth medium, and FLIM experiments were performed the next day. Transfection of HeLa cells with plasmids was performed using the transfection reagent (FuGENE 6; Roche) according to the instructions of the manufacturer.
Analysis of mitosis progression in tsBN2 cells treated with nonpermissive/permissive temperature
tsBN2 and BHK21 cells were grown in 75-cm2 flasks to ∼70% confluency at 33°C, rinsed in Opti-MEM with 4% FBS, and treated with 8 µM NZ for 6 h. Flasks were then vortexed (10 s), and media containing resuspended mitotic cells (>95% enrichment) were split into two equal aliquots for each sample. One half of tsBN2 and BHK21 cells, respectively, was incubated at 33°C, the other half was incubated at 40.5°C, and both were incubated for 2.5 h. The cells were then washed three times in 6 ml DPBS (Gibco) and once in Opti-MEM with 4% FBS at 22°C before being resuspended in 600 µl of 22°C Opti-MEM with 4% FBS. 100-µl aliquots of each cell suspension were then simultaneously transferred to a 30°C water bath and subsequently fixed by the addition of 500 µl of 4% paraformaldehyde, 0.02% glutaraldehyde, and 0.02% Triton X-100 in PHEM (60 mM Pipes, 25 mM Hepes, 10 mM EGTA, 4 mM MgSO4, pH 7.0 [buffered with KOH]) at 4–30-min intervals. The fixed suspensions were washed twice in PHEM before overnight sedimentation on poly-lysine (Sigma-Aldrich)–coated 12-mm-diam coverslips at 4°C. IF with tubulin antibodies and DNA stain was performed as described in the IF section, except that the slides were quenched 3× in a freshly prepared solution of BH4 (∼10 mg/50 ml GTB) before staining.
RanGAP1 RNAi
Suspensions of HeLa cells (approximately 60,000 cells in 500 µl/well) were seeded on 12-mm round glass coverslips in a 24-well plate (Falcon). The next day, the cells were transfected with 20 pmol/well with annealed and deprotected siRNA oligonucleotide (oligo) targeting a unique sequence in human RanGAP1 (5′-GAAACCGUCUGGAGAAUGAdTdT-3′; obtained from Thermo Fisher Scientific) using Lipofectamine 2000 and a protocol recommended by the manufacturer (Invitrogen). Control cells were treated with Lipofectamine 2000 only. After 5 d of treatment, cells were harvested for immunoblotting (using a protocol described for Fig. 2 A in Electrophoresis and quantitative immunoblotting) or fixed for IF with tubulin antibodies and DNA stain.
NRMT RNAi
Control siRNA oligo (5′-UGGUUUACAUGUCGACUAA-3′; ON-TARGETplus nontargeting siRNA #1; #D-001810-01-20) and NRMT siRNA oligo (5′-AGAGAAGCAATTCTATTCCAAG-3′; Schaner Tooley et al., 2010) were obtained from Thermo Fisher Scientific. HeLa and HFF-1 cells were grown in 6-well plates (35-mm-diam wells, containing two 12-mm round glass coverslips each) in antibiotic-free medium and transfected with a 100-nM final concentration siRNA using Oligofectamine (Invitrogen). After 120-h treatment, cells grown on coverslips were fixed by methanol/acetic acid/ethanol and processed for IF with RCC1 and tubulin antibodies and DNA stain as described in the IF section. The remainder of the cells in wells was harvested by trypsinization, washed in DPBS, and prepared for SDS-PAGE and immunoblotting as described in a subsequent section (Electrophoresis and quantitative immunoblotting). To perform FLIM in RNAi-treated cells, HeLa cells were treated with NRMT RNAi or control oligos as described in this paragraph. 3 d after the start of the treatment, control and NRMT RNAi-treated cells were transfected with a plasmid for the expression of Rango-4 (pK314 [pSG8 Rango-4]). FLIM analysis of both cell samples was performed 5 d after the start of treatment.
Scoring mitotic phenotypes in HFF-1 and HeLa cells
Mitotic phenotypes were manually scored by inspecting the fixed and immunostained (tubulin or TPX2; DNA) cells under 100×, 1.40 NA oil immersion objective on an inverted microscope (IX81; Olympus) equipped with a system controller (IX2-UCB; Olympus) and a shutter controller (Lambda 10-B; Sutter Instrument) and operated via MetaMorph software (Molecular Devices). Using the same lens, representative images (Fig. S3, C and D) were acquired using a charge-coupled device camera (C4742-98; Hamamatsu Photonics). During manual scoring, each mitotic cell entering the view was inspected in the DNA and tubulin or TPX2 channel and classified into categories including PM, metaphase, anaphase, and telophase. As the defining difference between PM and metaphase cells, we considered the state of chromosome congression rather than bipolar spindle assembly. Because the spindle axis was namely in HFF-1 cells tilted vertically, all cells were inspected by focusing along the z axis to discriminate between the spherical (PM) or disc-shaped (metaphase) arrangement of chromosomes. Cells with the most chromosomes dispersed and showing no sign of organization to the metaphase plate were scored as PM, and within that category, we scored the MT organization: MT ball, asters, and spindle. The MT ball corresponds to MTs showing no clear polar or radial structures, clearly radial MT arrays were scored as asters, and MT structures showing clear pole-to-pole connections were scored as spindles. Cells with spindles that were also showing clear evidence of chromosome congression to the disc-shaped metaphase plate were scored as metaphase, and within that category, we scored the number of spindle poles and number of lagging chromosomes. Similarly, we scored anaphase and telophase cells to categories with and without lagging chromosomes.
HFF-1 cell fusion
The fusion of HFF-1 cells was induced using a modification of the previously described protocol (Jansen et al., 2007). HFF-1 cells were seeded on 6-well plates and separately transduced with lentiviruses for the expression of RBP-4 FRET sensor or with lentiviruses for EB3-mCherry microtubule plus-end marker. 2 d later, the cells were trypsinized, and the cells expressing RBP-4 were mixed 1:1 with cells expressing EB3-mCherry and seeded on fibronectin-coated 35-mm ibidi dishes (2 × 105–2.5 × 105 cells/dish) in 2 ml of growth medium supplemented with 10 µm/ml Y-27632 Rho kinase inhibitor. After 1.5 h, the medium was aspirated, and cells were overlaid with 0.4 ml of 50% polyethylene glycol (PEG)-1500 in 75 mM Hepes, pH 8.0 (Roche), for 30 s at room temperature. Cells were then washed five times with DPBS and returned to culture at 37°C in 5% CO2 in the growth medium containing Y-27632. After an overnight recovery, the mitotic cells coexpressing mCherry and FRET sensor were analyzed by FLIM. Control nonfused cells expressing the RBP-4 sensor were treated as described for the RBP-4– and EB3-mCherry–expressing cells, except that the PEG treatment was omitted.
Synchronization of HeLa and HFF-1 cells (Fig. 2 B)
HeLa cells were seeded on plastic100-mm dishes (Falcon), and HFF-1 cells were seeded on acid-etched glass 100-mm dishes (Pyrex 3160; Pyrex) and kept in culture for 2–3 d to achieve optimal adherence. The cultures were then supplemented with 10 µM NZ added to their growth media and incubated for 6 h at 37°C in 5% CO2 to synchronize in mitosis. The dishes were rinsed with fresh media containing 10 µM NZ and vortexed in 3–4 ml of 10 µM NZ/media for 10 s at maximum speed (Vortex Genie 2; Daigger). The mitotic cells dissociated from the dishes were then washed and resuspended in warm DPBS and rapidly processed to prepare samples for SDS-PAGE and protein concentration measurements as described in Electrophoresis and quantitative immunoblotting. To collect interphase cells, the cells remaining on plates after shake-off were washed with warm DPBS and vortexed again to dissociate remaining mitotic cells, and the adherent cells were harvested by trypsinization, washed with warm FBS-containing media to neutralize trypsin, and processed as described for the mitotic cells. To prepare acid-etched tissue-culture glass plates, Petri dishes (Pyrex 3160) were incubated in 1 M HCl at 60–65°C overnight, washed extensively in distilled water, and autoclaved.
Confocal live-cell microscopy
Confocal images were sequentially acquired using ZEN 2009 software (Carl Zeiss) on a confocal system (LSM 710; Carl Zeiss) including a 34-channel spectral detector (Carl Zeiss) with an inverted microscope (Observer.Z1; Carl Zeiss) and an infrared laser (680–1,080 nm; Chameleon Ultra II; Coherent, Inc.). For most live-cell imaging experiments, cells were grown in a 2-well or 4-well chambered coverglass (Lab-Tek II; Thermo Fisher Scientific). For FRAP measurements of different RCC1-mCherry constructs, HeLa cells were grown in 8-well chambers (µ-Slide ibiTreat; ibidi). An incubator (Incubator XL S1; PeCon) mounted on the microscope stage and equipped with the Temperature Module S1, CO2 Module S1, and Heating Unit XL S1 was used to maintain the cells at 37°C in 5% CO2 and constant humidity. During all live-cell imaging experiments, cells were kept in the same growth media as for maintaining the cell culture.
Time-lapse bright-field imaging microscopy of cell division
Bright-field images were collected using the same microscope system as the confocal live-cell microscopy described in the previous section, except the cells were illuminated with a 5.0-mW HeNe laser (LASOS Lasertechnik GmbH) at 0.2% maximum power using the Plan Apochromat 20×, 0.8 NA lens, and the transmitted light was detected using the internal transmitted light photomultiplier tube transmitted light detector of the microscope. The pixel dwell time of the laser was fixed at 1.27 µs, and the detector gain was adjusted at each imaging field at the beginning of the time lapse to avoid under- or overexposure. Images were acquired as z stacks of five slices with 4-µm intervals every 80 s for ≤23 h and were stitched together sequentially with ZEN 2009 software to create time-lapse videos. ZEN autofocus macro was used between each time frame to prevent focus drifting. We acquired two videos for HeLa and seven videos for HFF-1 cells. Using the time stamp feature of the ZEN software, in all cells that underwent complete mitosis, we recorded the time of the (1) nuclear envelope breakdown, (2) metaphase plate formation, (3) metaphase plate disassembly, and (4) nuclear envelope reformation. The PM time corresponds to the interval between 1 and 2, the metaphase time corresponds to the interval between 2 and 3, and the interval between 3 and 4 is the anaphase + telophase time.
FLIM
Spatially resolved, time-correlated single-photon counting (TCSPC) datasets were acquired using a Plan Apochromat 63×,1.40 NA oil immersion lens on an inverted nonlinear optics (NLO) microscope (LSM 710) equipped with a TCSPC controller (SPC-830; Becker & Hickl). Samples were excited by one-photon 440-nm pulses generated by a frequency-doubling 80-MHz, 880-nm infrared laser using a harmonic generator (Mira Femto System Harmonic Generator 9300; Coherent, Inc.). The emission was collected from a custom side port, filtered through a 510 ± 45–nm band-pass filter (510/84; Semrock), and detected by a module (HPM-100-40; Becker & Hickl) containing a hybrid photomultiplier (R10467-40 GaAsP; Hamamatsu Photonics). Recording conditions were chosen to limit the emission to 0.2–1 × 106 counts per second. Images of 128 × 128 pixels (1,024 time bins/pixel) were averaged over 120 s. Before scanning each mitotic cell for FLIM, a bright-field image was acquired using the NLO microscope (LSM 710) as for FLIM, except that the cell was illuminated with a diode-pumped solid-state 561-nm, 10-mW laser 561-nm excitation line at 1% maximum power, and transmitted light was collected using the internal transmitted light photomultiplier tube transmitted light detector of the microscope. The pixel dwell time of the laser was fixed at 0.64 µs, and the detector gain was adjusted for each cell to obtain optimal exposure. Acquired bright-field images were 16 bit in depth and 512 × 512 pixels in dimensions.
FLIM data analysis
Out-of-cell fluorescence was omitted from the analysis by choosing a threshold at 1% of the mean photon count inside of the cell. Fluorescence lifetime images were produced and analyzed using SPCI (single photon counting image) software (Becker & Hickl). In all samples, the median donor fluorescence lifetime τdonor was calculated from datasets at bin = 1, assuming incomplete two-exponential decay and using trapezoid integration and no prefixed parameters. Instrumental response was determined using the SPCI automatic calculation routine. At least eight mitotic cells were analyzed per cell type and FRET sensor (range 8–34, 261 total; 132 with Rango-4, and 129 with RBP-4). Excluded from further analyses were FRET sensor-expressing cells displaying mean cellular τdonor > 2,300 ps, corresponding to three cells in total (one Rango-4 HFF-1 cell, one RBP-4 HFF-1 cell, and one RBP-4 hTert-RPE1 cell). Presumably, the excluded cells expressed sensors in which the dsREACh acceptors were only partially folded or translated. To prepare line scans of FLIM images, the τdonor values were exported from SPCI and imported to MetaMorph (version 7.5).
FLIM data pseudocolor display
The RGB pseudocolor images of the median τdonor values detected in cells were prepared using SPCI software, choosing an identical span of the displayed values for each FRET sensor: 250 ps for RBP-4, 400 ps for Rango-4, 150 ps for RBP-4Y, and 350 ps for Rango-4Y. We chose each of these display spans as ∼100 ps wider than the largest amplitude of the τdonor value detected among all mitotic cells. The FLIM display ranges were centered at the cell-specific mean τdonor rounded to 5 ps. For example, in a HeLa cell expressing Rango-4 (Fig. 1 C), in which the SPCI determined mean cellular τdonor = 1,997 ps and range of 1,842–2,145 ps, the FLIM display range was set to 1,800–2,200 ps.
FRET emission intensity ratio imaging
FRET emission intensity images were collected using a NLO microscope (LSM 710) with the same objective lens and light source used for the FLIM microscopy described in the FLIM section. The master gain was fixed at 900 and 1,150 for mTFP-1 and YPet channels, respectively, to directly compare the value of IFRET/Idonor between different cells. The digital gain was set at 15.0 for both channels. All acquired images were 16 bit and 512 × 512 pixels in dimension. The IFRET/Idonor ratiometric images were prepared using MetaMorph (version 7.5).
FRAP
FRAP measurements of mCherry-tagged RCC1 constructs expressed in live cells were performed on a confocal NLO microscope (LSM 710) using a Plan Apochromat 63×, 1.40 NA oil immersion lens (Carl Zeiss) and a diode-pumped solid-state 561-nm, 10-mW laser 561-nm excitation line. Five prebleach scans were followed by photobleaching with two passes of a 100% power 561-nm laser and 30 (interphase cells) or 55 (mitotic cells) postbleach scans. In the interphase cells, the photobleached area was a 1-µm circle inside the nucleus. To suppress the noise caused by chromosome motion during recovery, the photobleached area comprised approximately one half of the metaphase plate in the mitotic cells. Pre- and postbleach scans were acquired at 1 frame/s, and the excitation power was adjusted to assure no detectable bleaching in neighboring cells that were not subjected to FRAP (corresponding to 0.2–0.7% of maximum 561-nm laser power). ZEN 2009 software was used to calculate the fluorescence recovery t1/2.
Cloning general procedures
Gene construction kits (Textco BioSoftware) and Vector NTI software (Invitrogen) were used for the in silico assembly and for sequencing of all constructs. Site-directed mutagenesis was performed using the site-directed mutagenesis kit (QuikChange Lightning Multi; Agilent Technologies). PCRs were performed with high fidelity polymerases (Pfu Ultra or Pfx; Invitrogen). Construction of lentiviral expression vectors was performed with the pLenti6.3/V5-TOPO TA Cloning kit (Invitrogen), the ViraPower HiPerform T-REx Gateway Expression System (Invitrogen), or the pENTR/D-TOPO Cloning kit (Invitrogen). All constructs were verified by restriction digests and sequencing across fragments generated by PCR.
Cloning of RCC1α-mCherry constructs
PCR cloning was used to insert human full-length RCC1α or RCC1α (1–27 aa) between Xho1 and Age1 sites and mCherry between Age1 and Not1 sites in pEGFPN1, resulting in p-RCC1α-mCherry (pK260) and p-RCC1α NTD-mCherry (pK295) plasmids for transfection-mediated expression in tissue-culture cells. PGK (phosphoglycerate kinase) Histone H2B-mCherry (plasmid 21217; Addgene; Kita-Matsuo et al., 2009) was used as the PCR template for mCherry. The pK260 and pK295 plasmids were then used as templates for the PCR-mediated assembly of lentiviral expression plasmids in linearized topoisomerase-conjugated pLenti6.3/V5-TOPO.
Full-length wt RCC1α-mCherry was amplified from pK260 by PCR, using 5′ RCC1α and 3′ mCherry primers. After the addition of TA overhangs using Taq polymerase, the purified PCR product was inserted by TOPO cloning into pLenti6.3/V5-TOPO, resulting in pLenti6.3-RCC1α-mCherry (pK267). Using the same strategy and PCR with mutated 5′ RCC1α oligos, we constructed pLenti RCC1α-mCherry with S2K (pK286) and ASPK (pK285) mutations. Similarly, using the same primers, but with p-RCC1α NTD-mCherry (pK295) as a template, we prepared constructs for the lentiviral expression of wt and mutated mCherry-tagged NTD: pLenti wt-NTD-mCherry (pK296), pLenti ASPK-NTD-mCherry (pK297), and pLenti S2K-NTD-mCherry (pK306).
Cloning of FRET sensors
As in YFP-RBD-CFP (Kaláb et al., 2002), the sensory domain in RBP-4 and RBP-4Y FRET sensors is an RBD of Yrb1 (Saccharomyces cerevisiae homologue of human RanBP1). In Rango-4Y and Rango-4, the sensory domain is identical to the importin-β–binding domain (IBB) of human snurportin 1, which was used in previous versions of Rango (Kaláb et al., 2006; Kaláb and Soderholm, 2010). The FRET donor in RBP-4Y and Rango-4Y is the monomeric teal fluorescent protein mTFP-1 (purchased as pmTFP-1-ER plasmid from Allele Biotechnology; Ai et al., 2008), and the acceptor is YPet (Nguyen and Daugherty, 2005). The FRET donor in RBP-4 and Rango-4 is again mTFP-1, and the acceptor is a moderately dimerizing nonfluorescent acceptor, dsREACh, that we developed by removing the monomerization mutation R223F from sREACh (gift from R. Yasuda, Duke University Medical Center, Durham, NC; Murakoshi et al., 2008) and introducing a dimerization-promoting mutation, S208F. Plasmid for transfection-mediated expression of Rango-4 in HeLa (pK215 [pSG Rango-4]) was prepared by stepwise assembly of mTFP-1, snurportin IBB, and dsREACh sequences in a pSG8 vector using PCR and restriction–ligation cloning.
Using PCR and restriction–ligation cloning, the open reading frames for RBP-4Y, Rango-4Y, RBP-4, and Rango-4 were inserted into pENTR/D-TOPO-V5. In this manner, we prepared pK224 (pENTR RBP-4Y), pK225 (pENTR Rango-4Y), pK226 (pENTR RBP-4), and pK221 (pENTR Rango-4). LR Gateway recombination reactions (Invitrogen) of the aforementioned pENTR vectors with pLenti6.3/TO/V5-DEST were then used to prepare pK232 (pLenti RBP-4Y), pK231 (pLenti Rango-4Y), pK234 (pLenti RBP-4), and pLenti Rango-4 (pK233). Plasmid pK315 (pSG8 mTFP-1–IBB) for the expression of mTFP-1–IBB donor-only FLIM standard was constructed by ligating a PCR-amplified mTFP-1–IBB fragment from pK233 (pLenti Rango-4) into the pSG-8 vector. Plasmid for the detection of background chromatin cytoplasmic gradient measurements was prepared by replacing RBD in pK215 with a PCR-amplified fragment corresponding to amino acids 17–36 in human importin-α1, resulting in pK364 (pSG8 mTFP-1-linker-dsREACh). A summary of the FRET expression constructs is shown in Table S2.
Other expression constructs
Plasmid for the expression of mCherry-RanT24N (p-mCherry-RanT24N; pTK21; Kiyomitsu and Cheeseman, 2012) was a gift of T. Kyomitsu and I. Cheeseman (Massachusetts Institute of Technology, Cambridge, MA). The pLenti6.3 GFP-Ran (pK281) construct was created by first assembling the EGFP-Ran in pENTR/D-TOPO using PCR cloning followed by Gateway reaction between pENTR/D-TOPO-EGFP-Ran and pLenti6.3/TO/V5-DEST (Invitrogen).
RNA isolation and quantitative PCR
Tissue-culture cells were harvested by trypsinization and washed in DPBS, and total cellular RNA was isolated using the RNA isolation kit (PureLink Micro-to-Midi Total; Invitrogen). The RNA was then used as a template to prepare complementary DNA with the cDNA synthesis kit (iScript; Bio-Rad Laboratories). RT-PCR reactions were performed on the real-time PCR system (7900HT Fast; Applied Biosystems) using Power SYBR green PCR master mix (Applied Biosystems). RT-PCR primers were as described previously (Hood and Clarke, 2007): RCC1α forward, 5′-AAGAAGGTGAAGGTCTCACAC-3′; RCC1β forward, 5′-CTCCTGCCAAGGTGCCTG-3′; RCC1γ forward, 5′-CTCCTGCCAAGTCTCACAC-3′; and RCC1 common reverse, 5′-GCACAACATCCTCCGGAATG-3′. The results were quantified using 7900HT version 2.3 Sequence Detection System.
Electrophoresis and quantitative immunoblotting
To prepare SDS-PAGE samples for all experiments except NRMT RNAi (see next paragraph), cells were released from tissue-culture flasks by trypsinization, resuspended in 5 ml of warm FBS-containing media, pelleted (2 min at 125 g), and resuspended in warm DPBS. Each cell suspension was divided in an aliquot for protein concentration measurements (100 µl) and aliquots for the SDS-PAGE sample preparation (1 ml each). These aliquots were pelleted for 2 min at 120 g at room temperature. The aliquots for SDS-PAGE were immediately resuspended in SB (100 µl of 2× SDS-PAGE sample buffer with 5% β-mercaptoethanol), heated at 100°C for 5 min, and stored at −30°C before use. The cell pellets for protein concentration measurement were resuspended in 40 µl protein extraction reagent (M-PER; Thermo Fisher Scientific), incubated on ice for 10 min, sonicated (3 × 5 s) on ice, and clarified by centrifugation (10 min at 16,200 g at 4°C). The protein concentration was measured with the Protein Assay (Bio-Rad Laboratories) using BSA as a standard.
To prepare SDS-PAGE samples from NRMT or control RNAi-treated HeLa cells (Fig. S3 B), cells were harvested by trypsinization, resuspended in 5 ml FBS-containing media, washed with DPBS, and extracted in M-PER supplemented with protease inhibitor cocktail (Roche) and phosphatase inhibitor cocktail (PhosSTOP; Roche) for 10 min on ice. After the lysates were clarified by centrifugation (10 min for 16,200 g at 4°C), aliquots were taken for protein concentration measurement, and the rest of the lysates were mixed with SB, heated at 100°C for 5 min, and stored at −30°C before use.
Proteins were separated by SDS-PAGE using homemade gels containing a 100:1 wt/wt acrylamide/bis-acrylamide (Bio-Rad Laboratories) ratio and SDS L4509 (Sigma-Aldrich). Equal protein mass of cell lysates (20 µg/lane in all figures except Fig. 2 B in which 5 µg/lane was loaded) was separated on either 15% SDS-PAGE gels (for the detection of pS10-histone H3) or 10% SDS-PAGE gels (other antibodies). Gels were blotted to polyvinylidene fluoride transfer membrane (Immobilon-FL; Millipore) or nitrocellulose membrane (Odyssey; LI-COR Biosciences).
Immunoblots were blocked with 4% BSA (Sigma-Aldrich) made in TTBS (TBS with 0.05% Tween 20) or 2.5% skim milk in TTBS. After incubation with primary antibodies and washes in TTBS, the blots were incubated with IRDye 800– and/or IRDye 680–conjugated goat or donkey secondary antibodies (LI-COR Biosciences) and washed in TTBS. Then, the fluorescence emission spectra were acquired using the Odyssey Classical Infrared Imaging System (LI-COR Biosciences). Tubulin staining was used as an equal loading control for all samples, taking advantage of the two-color detection of the LI-COR Biosciences system and/or after separating the blots in horizontal stripes that were developed with different antibodies.
To calculate the relative protein expression or modification levels from data, Odyssey CLx software (LI-COR Biosciences) was first used to determine the background-subtracted emission intensity of protein bands. The relative expression values were calculated from such background-subtracted signals that were normalized to the levels of tubulin detected in the same lane on the same blot. The relative expression values for asynchronous cell cultures of HFF-1, HeLa, DVH3, and DVH3-t2 cells were derived from data shown in Fig. 2 A and from at least three additional biological repeats (new cell culture, SDS-PAGE, and immunoblot) of this experiment. The comparisons of RCC1 levels in cell cycle–synchronized HeLa and HFF-1 cells were obtained from data shown in Fig. 2 B and from one additional biological repeat of this experiment. The comparison of RCC1 levels in HeLa cells treated with control or NRMT RNAi was obtained from data in Fig. S3 B and three additional repeats of the NRMT RNAi experiment. Recombinant N-terminally biotinylated RCC1α (Halpin et al., 2011) in Fig. S2 was provided by D. Halpin and R. Heald (University of California, Berkeley, Berkeley, CA).
Image processing
Photoshop CS3 (Adobe) was used to crop and arrange all images, and contrast was adjusted separately for each RGB channel using the Auto Color Correction Options with no clipping. Figures were assembled and annotated using Illustrator CS3 (Adobe).
Statistical analyses
Statistical analyses were performed with Prism version 6 (GraphPad Software) and with Excel (Microsoft).
Online supplemental material
Fig. S1 shows validation of the RBP-4 and Rango-4 sensors and data on reference τdonor and on background τdonor gradient. Fig. S2 shows the localization of Ran and RanGAP1, quantitative RCC1 PCR, Me3-SPK antibody characterization, and RCC1-mCherry FRAP data. Fig. S3 shows data on mitotic phase duration in live cells, RNAi validation, IF detection of mitotic phases, RanGAP RNAi in HeLa cells, and RCC1 inhibition in tsBN2 cells. Table S1 shows characteristics of cells involved in this study. Table S2 shows a summary of DNA constructs for the expression of FRET sensors.
Acknowledgments
The authors wish to thank L. Samelson, C. Parent, J. Rubin, R. Heald, K. Weis, and T. Misteli for comments on the manuscript; C. Tooley, I. Macara, M. Stampfer, R. Yasuda, D. Duelli, M. Dasso, A. Arnaoutov, D. Halpin, R. Heald, I. Cheeseman, and T. Kiyomitsu for reagents and plasmids; and D. Duelli for data on cell characteristics. Imaging was performed at the Confocal Microscopy Core Facility of the Center for Cancer Research, National Cancer Institute, in Bethesda, MD. Some initial experiments were performed in the laboratories of K. Weis and R. Heald and at the Molecular Imaging Center at the University of California, Berkeley.
K. Hasegawa, S.J. Ryu, and P. Kaláb were supported by the Intramural Research Program of the Center for Cancer Research, National Cancer Institute, National Institutes of Health.
References
- ANOVA
analysis of variance
- DPBS
Dulbecco’s PBS
- FLIM
fluorescence lifetime imaging microscopy
- FRET
Förster resonance energy transfer
- GTB
general tubulin buffer
- hTert
human telomerase reverse transcriptase
- IBB
importin-β–binding domain
- IF
immunofluorescence
- IFB
IF buffer
- MT
microtubule
- NLO
nonlinear optics
- NRMT
N-terminal RCC1 methylase
- NTD
N-terminal domain
- NZ
nocodazole
- PEG
polyethylene glycol
- PM
prometaphase
- RBD
RanGTP-binding domain
- SAF
spindle assembly factor
- SUMO
small ubiquitin-like modifier
- TCSPC
time-correlated single-photon counting
- wt
wild type
Author notes
K. Hasegawa and S.J. Ryu contributed equally to this paper.