The formation of functional kinetochores requires the accurate assembly of a large number of protein complexes. The Hsp90–Sgt1 chaperone complex is important for this process; however, its targets are not conserved and its exact contribution to kinetochore assembly is unclear. Here, we show that human Hsp90–Sgt1 interacts with the Mis12 complex, a so-called keystone complex required to assemble a large fraction of the kinetochore. Inhibition of Hsp90 or Sgt1 destabilizes the Mis12 complex and delays proper chromosome alignment due to inefficient formation of microtubule-binding sites. Interestingly, coinhibition of Sgt1 and the SCF subunit, Skp1, increases Mis12 complexes at kinetochores and restores timely chromosome alignment but forms less-robust microtubule-binding sites. We propose that a balance of Mis12 complex assembly and turnover is required for the efficient and accurate assembly of kinetochore–microtubule binding sites. These findings support a novel role for Hsp90–Sgt1 chaperones in ensuring the fidelity of multiprotein complex assembly.
Introduction
Kinetochores are assembled from multiple distinct protein complexes to form a multivalent microtubule-binding site. Kinetochores contact the lateral faces of microtubules, and these initial low-affinity contacts are believed to mature to more stable end-on attachments (for a recent review see Cheeseman and Desai, 2008). A combination of microtubule- and motor-derived forces acting at the kinetochore and along chromosome arms contribute to chromosome alignment and establish tension between sister chromatids during metaphase (Hays and Salmon, 1990; Sharp et al., 2000). The identification of a large number of kinetochore proteins from multiple species has greatly advanced our understanding of kinetochore structure, yet the rules for assembling these complexes are still poorly understood.
In budding yeast, the centromere binding complex 3 (CBF3) is first assembled before it can bind to sequence-specific elements on centromeric DNA (CEN-DNA) and recruit additional complexes that form a single microtubule-binding site (Espelin et al., 1997; Kaplan et al., 1997). After CBF3 binds to CEN-DNA, the assembly of a functional kinetochore is thought to occur in a hierarchical manner, with complexes building outwards until a microtubule-binding site forms (Ortiz et al., 1999). However, it is interesting to note that relatively few mutants completely eliminate kinetochore–microtubule binding in yeast, which suggests that a strict hierarchy between centromeric DNA and microtubule binding may be an oversimplification. CBF3 is composed of three core subunits, and the Hsp90–Sgt1 chaperone complex balances its assembly with its turnover (Rodrigo-Brenni et al., 2004). Analysis of reconstituted Hsp90–Sgt1 complexes led to the proposed model that Sgt1 acts as an adapter, linking Hsp90 to a subset of its clients (Catlett and Kaplan, 2006). Sgt1 directly interacts with Hsp90 and with Skp1, a protein shared with both CBF3 and the SCF E3 ubiquitin ligase complex. The SCF-dependent turnover of CBF3 subunits suggests that Sgt1 connects Hsp90 clients to SCF-mediated degradation (Kaplan et al., 1997). Consistent with this idea, mutants that affect the affinity of Sgt1 for Skp1 favor either assembly or turnover of CBF3 (Rodrigo-Brenni et al., 2004).
In contrast to the relatively simple point kinetochores that form on sequence-determined centromeres in budding yeast, specification of mammalian kinetochores is largely epigenetic and is dependent on the proper loading of nucleosomes containing the histone H3 variant CENP-A (for a review see Gieni et al., 2008). The CENP-A nucleosome is required to recruit a kinetochore assembly platform in interphase that consists of the CENP-H, CENP-M, and CENP-O complexes, also called the constitutive centromere complex, or CCAN (Okada et al., 2006; Cheeseman and Desai, 2008). Outer kinetochore complexes are assembled at centromeres in the G2 stage of the cell cycle (Hori et al., 2003; Cheeseman et al., 2004; Yang et al., 2008); Knl1 and the Mis12 complex are believed to modulate the formation of the Ndc80 microtubule-binding complex (Cheeseman et al., 2006). Rather than forming individual microtubule-binding units, electron tomography experiments suggest that microtubule-binding sites form a network that allows both low- and high-affinity microtubule interactions (Dong et al., 2007). It has been suggested that this architecture allows for an initial low-affinity capture of microtubules by kinetochores followed by a maturation process that gives rise to high-affinity interactions. Implicit in this model is the idea that the proper assembly of a kinetochore network is necessary to ensure the efficient formation of robust microtubule attachment sites.
Experiments to test the interdependency of kinetochore proteins on assembly of a microtubule-binding site are also consistent with a network-like organization. For example, inhibition of the outer kinetochore protein Knl-1 reduces the formation of microtubule attachment sites that include Ndc80Hec1 as well as the “keystone” complex, Mis12 (Cheeseman et al., 2008). Similarly, the depletion of Mis12 subunits reduces the levels of both the outer kinetochore protein Ndc80Hec1 and the inner kinetochore proteins CENP-H and CENP-A (Kline et al., 2006), which suggests that kinetochore complexes assemble into an interdependent network.
Despite advances in understanding the dependencies of kinetochore protein assembly at centromeres, it is less clear how individual subunits correctly assemble into subcomplexes and then into higher-order structures. The fact that both simple and complex kinetochores require Hsp90–Sgt1 suggests that chaperones provide a conserved mechanism for assembling large protein networks. Although mammalian cells lack a homologue of CBF3, inhibition of either Hsp90 or Sgt1 leads to the reduction in a large number of kinetochore-associated proteins (Steensgaard et al., 2004; Niikura et al., 2006) and reduces the efficiency of microtubule attachments. The partial loss of microtubule attachment activity suggests that Hsp90–Sgt1 modulates the efficiency of the higher-order kinetochore assembly required for chromosome alignment. Thus, identification of the target of Hps90-Sgt1 will provide important insights into the rate-limiting steps required to ensure the efficient assembly of kinetochore–microtubule attachment sites.
Using a combination of genetic and biochemical approaches, we show that the Mis12 complex is the primary target of Hsp90–Sgt1 at the kinetochore. Inhibition of either Sgt1 or Hsp90 dramatically reduces the levels of Mis12 subunits and gives rise to kinetochore defects identical to loss of Mis12 subunits. Mis12 complexes interact with both Hsp90 and Sgt1, and Sgt1 stabilizes this complex, which is consistent with its proposed role as a client adaptor. The SCF component Skp1 opposes the turnover of Mis12 complexes and rescues the loss of high-affinity microtubule-binding sites after Sgt1 inhibition. Interestingly, inhibition of Skp1 reduces the robustness of microtubule attachment sites, arguing that both assembly and turnover of Mis12 are important for proper kinetochore function.
Results
Hsp90 is required for Dsn1 stability and Mis12 complex localization
Previous studies using Hsp90 inhibitors and Sgt1 siRNA suggest that the chaperone complex is required for recruitment of both constitutive and outer kinetochore complexes (Steensgaard et al., 2004; Niikura et al., 2006). However, because of the interdependence of kinetochore complexes during assembly, it is unclear which kinetochore complex or complexes might require chaperone function. To address this issue, we used the rapid kinetics of Hsp90 inhibition by 17-(allylamino)-17-demethoxygeldanamycin (17-AAG) to identify kinetochore complexes most immediately affected. After inhibition of Hsp90, clients often exhibit a decrease in protein levels, either because they degrade or become insoluble. After treatment of HeLa cells for 12–18 h with 100 nM 17-AAG, immunoblotting showed that both outer (e.g., Ndc80Hec1) and constitutive kinetochore complexes (e.g., CENP-K, -H, -U, and -N) remain at control levels. In contrast, the Dsn1 subunit of the Mis12 complex was reduced to 65% of control; the levels of other Mis12 subunits were unaffected by 17-AAG (Fig. 1 A). Treatment with 500 nM 17-AAG for 18 h further decreased the levels of Dsn1 to less than half (47% of control; Fig. 1 B). Consistent with the immunoblots, short-term treatment of cells with 500 nM 17-AAG resulted in a general decrease in Mis12 subunits at kinetochores (Fig. 1 C). It is interesting to note that not all Mis12 components behaved the same way after 17-AAG treatment; Dsn1 levels at kinetochores were decreased on average to 78% of control, whereas Nnf1 was more dramatically reduced (63% of control; Fig. 1 D). The Nsl1 signal was similarly decreased at kinetochores but was also found in large aggregates near chromosomes (Fig. 1 C, arrow). The observed heterogeneity suggests that Mis12 subunits have different fates in the absence of Hsp90. Overall, these observations support the idea that the Mis12 complex, and in particular Dsn1, is a target of the Hsp90 chaperone.
Hsp90 inhibition decreases Dsn1 levels and Mis12 complex recruitment to the kinetochore. (A) HeLa cells were treated with 100 nM 17-AAG for 18 h, and extracts were analyzed by immunoblotting for the indicated kinetochore antigens. Asterisks indicate nonspecific background on the immunoblot. The fluorescent intensities from the secondary antibody bound to the indicated Mis12 complex subunits were quantified (see Materials and methods) and are presented as a percentage signal from the control siRNA treatment. Differences in the amount of extract loaded were normalized using tubulin immunoblots. (B) HeLa cells were treated with the indicated concentrations of 17-AAG and analyzed as in A. (C) HeLa cells treated with the 500 nM 17-AAG or with carrier alone for 18 h were processed for immunofluorescence using antibodies against the indicated Mis12 subunit, DAPI, and ACA. Merged and individual fluorescent images are presented as indicated. The arrow indicates Nsl1 protein aggregates. Bar, 10 µM. (D) The mean maximal fluorescent intensities for control cells were set to 100%, and the fraction of signal remaining after 17-AAG treatment was calculated. Error bars represent the deviation from the mean observed in multiple kinetochores and cells. This trend was observed in multiple repetitions of the experiment.
Hsp90 inhibition decreases Dsn1 levels and Mis12 complex recruitment to the kinetochore. (A) HeLa cells were treated with 100 nM 17-AAG for 18 h, and extracts were analyzed by immunoblotting for the indicated kinetochore antigens. Asterisks indicate nonspecific background on the immunoblot. The fluorescent intensities from the secondary antibody bound to the indicated Mis12 complex subunits were quantified (see Materials and methods) and are presented as a percentage signal from the control siRNA treatment. Differences in the amount of extract loaded were normalized using tubulin immunoblots. (B) HeLa cells were treated with the indicated concentrations of 17-AAG and analyzed as in A. (C) HeLa cells treated with the 500 nM 17-AAG or with carrier alone for 18 h were processed for immunofluorescence using antibodies against the indicated Mis12 subunit, DAPI, and ACA. Merged and individual fluorescent images are presented as indicated. The arrow indicates Nsl1 protein aggregates. Bar, 10 µM. (D) The mean maximal fluorescent intensities for control cells were set to 100%, and the fraction of signal remaining after 17-AAG treatment was calculated. Error bars represent the deviation from the mean observed in multiple kinetochores and cells. This trend was observed in multiple repetitions of the experiment.
Sgt1 inhibition affects the stability of the Mis12 complex
We next asked if inhibiting Sgt1 in HeLa cells similarly affected the Mis12 complex. After depletion of Sgt1 using siRNA (∼70% reduction; see Fig. 3 C), we observed a similar reduction in the fluorescent intensities of Mis12 subunits at the kinetochore compared with 17-AAG treatment (compare Fig. 2 A and Fig. 1 C). However, the decrease in signals was far more dramatic with Sgt1 siRNA; Dsn1, Nnf1, and Nsl1 were a mean of 47%, 11%, and 9% of control signals, respectively (Fig. 2 B). The differences between Hsp90 and Sgt1 inhibition may reflect either the critical targeting role of Sgt1 or the fact that 17-AAG can trap Hsp90 client intermediates, resulting in partially formed complexes. Regardless of these differences, we conclude that the localization of Mis12 kinetochore complexes requires the function of both Hsp90 and Sgt1.
Sgt1 depletion prevents efficient Mis12 subunit recruitment to kinetochores. (A) After treatment of HeLa cells with control or Sgt1 siRNAs for 96 h, cells were prepared for immunofluorescence staining using antibodies against Dsn1, Nnf1, or Nsl1 as indicated, and chromosomes were stained with DAPI. Bar, 10 µM. (B) Maximum fluorescent signals at kinetochores were quantified using ImageJ (see Materials and methods) for each of the indicated antigens. A mean of 96 kinetochores from multiple cells was used to calculate the mean values and are plotted with error bars that represent the standard deviation.
Sgt1 depletion prevents efficient Mis12 subunit recruitment to kinetochores. (A) After treatment of HeLa cells with control or Sgt1 siRNAs for 96 h, cells were prepared for immunofluorescence staining using antibodies against Dsn1, Nnf1, or Nsl1 as indicated, and chromosomes were stained with DAPI. Bar, 10 µM. (B) Maximum fluorescent signals at kinetochores were quantified using ImageJ (see Materials and methods) for each of the indicated antigens. A mean of 96 kinetochores from multiple cells was used to calculate the mean values and are plotted with error bars that represent the standard deviation.
Sgt1 and Hsp90 coordinately stabilize Mis12 complex protein levels. (A) The levels of Mis12 proteins were measured by immunoblot after treatment with control or Sgt1 siRNAs for 96 h. Tubulin serves as a loading control and was used to quantify the fluorescent immunoblot signals (see Materials and methods). Changes in the levels of Mis12 subunits are expressed as a percentage of control extracts (set at 100%). (B) After treatment with the indicated siRNAs, Mis12 subunits were analyzed by immunoblotting. (C) siRNA knockdown efficiency was analyzed for Sgt1 and Hsp90 as in A. Error bars represent the standard deviation derived from multiple blots: Sgt1 siRNA (n = 4), Hsp90 RNAi (n = 2). (D) Dsn1 levels were measured by immunoblotting after treatment with Hsp90 or Hsp90 and Sgt1 RNAi for 48 h. Protein levels were analyzed as in A. (E) Immunoblots with antibodies to histone H3 phospho-serine 10 (H3pS10) and Aurora B after siRNA knockdown of Sgt1 or Hsp90 levels for 72 h. The percentage of mitotic cells observed at the time of harvest is indicated below the blot.
Sgt1 and Hsp90 coordinately stabilize Mis12 complex protein levels. (A) The levels of Mis12 proteins were measured by immunoblot after treatment with control or Sgt1 siRNAs for 96 h. Tubulin serves as a loading control and was used to quantify the fluorescent immunoblot signals (see Materials and methods). Changes in the levels of Mis12 subunits are expressed as a percentage of control extracts (set at 100%). (B) After treatment with the indicated siRNAs, Mis12 subunits were analyzed by immunoblotting. (C) siRNA knockdown efficiency was analyzed for Sgt1 and Hsp90 as in A. Error bars represent the standard deviation derived from multiple blots: Sgt1 siRNA (n = 4), Hsp90 RNAi (n = 2). (D) Dsn1 levels were measured by immunoblotting after treatment with Hsp90 or Hsp90 and Sgt1 RNAi for 48 h. Protein levels were analyzed as in A. (E) Immunoblots with antibodies to histone H3 phospho-serine 10 (H3pS10) and Aurora B after siRNA knockdown of Sgt1 or Hsp90 levels for 72 h. The percentage of mitotic cells observed at the time of harvest is indicated below the blot.
We reasoned that if Sgt1 targets Hsp90 to Mis12 complexes, then, like 17-AAG treatment, depletion of Sgt1 should also destabilize the Dsn1 subunit of the Mis12 complex. To test this possibility, we immunoblotted extracts after treating cells with Sgt1 siRNA. We observed a significant decrease in the levels of Dsn1 (50% of control; Fig. 3 A), which is comparable to 500 nM 17-AAG treatment (Fig. 1 B). Unlike 17-AAG treatments, we observed a significant decrease in the levels of the Nsl1 and Nnf1 subunits (59% and 57% of control, respectively) and a moderate reduction of Mis12 (87% of control; Fig. 3 A). Again, the more dramatic destabilization may arise from the ability of Sgt1 to target Mis12 subunits to Hsp90 and thus protect them from degradation. Interestingly, depletion of Dsn1, or any other Mis12 component, similarly destabilizes all Mis12 subunits (Fig. 3 B); we take this data as evidence that protein destabilization is linked to failures in complex assembly (Fig. 3 A).
If Sgt1 is required to target Mis12 subunits to Hsp90, we reasoned that partial depletions of Sgt1 and Hsp90 would exhibit an additive effect on Mis12 instability, or a synthetic interaction. We treated cells with siRNA against Sgt1 or Hsp90 for just 48 h and observed no decrease in Dsn1 levels when Sgt1 is inhibited (not depicted) and only a modest reduction in Dsn1 when Hsp90 is inhibited (72% of control; Fig. 3 D). However, codepleting Sgt1 and Hsp90 results in a 50% decrease of Dsn1 after 48 h. In addition, codepletion resulted in elevated cell death compared with the single depletion conditions (unpublished data). Collectively, the instability of Dsn1 after Hsp90 or Sgt1 depletion and the synthetic effect of codepletion argue that Hsp90 and Sgt1 function in the same pathway to assemble Mis12 complexes and target them to the kinetochore.
Sgt1 and Dsn1 similarly affect overall kinetochore assembly
It is possible that in addition to Mis12 complexes, Hsp90–Sgt1 targets other kinetochore complexes or regulators of kinetochore assembly. Aurora B is a known Hsp90 client (Miyata and Nishida, 2004) and has been found to regulate the localization of Dsn1 and Mis12 complexes during mitosis (Yang et al., 2008). To test whether Aurora B activity is affected by siRNA of Sgt1, we measured a known target of the kinase serine 10 on histone H3 (H3pS10). Immunoblots of extracts showed the expected enrichment of H3pS10 after nocodazole treatment. Interestingly, depletion of Sgt1 gave rise to an elevated H3pS10 signal detected in immunoblots (Fig. 3 E) and by immunofluorescence (not depicted), but with a similar mitotic index to nocodazole-treated cells (Fig. 3 E). In contrast, depletion of Hsp90 eliminated the H3pS10 signal (Fig. 3 E), a result that is consistent with published reports that Hsp90 is required for Aurora B activity (Lange et al., 2002; Terasawa and Minami, 2005). The fact that depletion of Sgt1 and Hsp90 had opposite effects on Aurora B highlights the distinct pathways regulated by Sgt1 and Hsp90. Importantly, these results also argue against Aurora B being the relevant Sgt1 target with respect to Mis12 complex assembly, as elevated Aurora B is predicted to enhance Mis12 complex recruitment to the kinetochore (see Discussion; Yang et al., 2008). Finally, we checked if the reported link between Sgt1 and the Drosophila Polo-like kinase 1 (Martins et al., 2009) could explain the Mis12 defect in Sgt1-inhibited cells. We found that inhibition of Polo by siRNA did not alter Dsn1 levels (not depicted), which led us to favor the idea that Sgt1 directly affects Mis12 complex assembly.
As a so-called keystone complex, Mis12 contributes to the localization of several other kinetochore complexes. If Sgt1 function is limited to Mis12 complex assembly, we predict that blocking Sgt1 function will phenocopy the depletion of individual Mis12 subunits. Indeed, inhibition of Sgt1 or Dsn1 reduced the kinetochore signals of Ndc80Hec1, CENP-K, CENP-N, and CENP-U but not CENP-A, -C, or -B (Fig. 4, A–D, respectively). In the case of CENP-K, CENP-N, and CENP-U, we analyzed signals in interphase cells. Interphase CCAN components are constitutively loaded and have low turnover rates; these characteristics have been noted previously (McClelland et al., 2007). As a result, staining was more robust than in mitotic cells, allowing more accurate measurements of fluorescent signals closely associated with anti-centromere antigen (ACA) staining (Fig. 4, A–D, insets). Importantly, siRNA against Sgt1 and Dsn1 reduced the kinetochore-associated signals of both inner and outer complexes to a similar extent (Fig. 4 E). In contrast, our findings and previously published work show that depletion of either the CENP-H complex component, CENP-K, or the microtubule-binding component Ndc80Hec1, does not affect Mis12 complex localization (unpublished data; Cheeseman et al., 2006; Kline et al., 2006). The shared set of localization defects observed after inhibition of Sgt1 or Dsn1 favor the idea that Sgt1 acts solely on the Mis12 complex.
Sgt1 and Dsn1 target a complementary set of kinetochore proteins. (A–D) HeLa cells were treated with control, Sgt1, or Dsn1 siRNAs for 72 h and prepared for immunofluorescence analysis using antibodies to Ndc80Hec1, CENP-K, CENP-N, CENP-U, or ACA as indicated. Chromosomes were stained with DAPI. Insets highlight the kinetochore foci indicated by the small arrows in the merged image and are enlarged 3.5-fold. Bars: (main panels) 10 µm; (insets) 1 µm. (E) Fluorescent signals were quantified as in Fig. 1 D. P-values were <0.05 for all treated samples with the exception of CENP-U, which exhibited a high degree of signal variation. Error bars represent the deviation from the mean observed in multiple kinetochores and cells. (F) Diagram of kinetochore–protein interactions based on our genetic analyses and published data (Foltz et al., 2006; Okada et al., 2006; Cheeseman et al., 2008). The Mis12 complex and arrows are highlighted in red to represent potential direct or indirect interaction pathways within the kinetochore network.
Sgt1 and Dsn1 target a complementary set of kinetochore proteins. (A–D) HeLa cells were treated with control, Sgt1, or Dsn1 siRNAs for 72 h and prepared for immunofluorescence analysis using antibodies to Ndc80Hec1, CENP-K, CENP-N, CENP-U, or ACA as indicated. Chromosomes were stained with DAPI. Insets highlight the kinetochore foci indicated by the small arrows in the merged image and are enlarged 3.5-fold. Bars: (main panels) 10 µm; (insets) 1 µm. (E) Fluorescent signals were quantified as in Fig. 1 D. P-values were <0.05 for all treated samples with the exception of CENP-U, which exhibited a high degree of signal variation. Error bars represent the deviation from the mean observed in multiple kinetochores and cells. (F) Diagram of kinetochore–protein interactions based on our genetic analyses and published data (Foltz et al., 2006; Okada et al., 2006; Cheeseman et al., 2008). The Mis12 complex and arrows are highlighted in red to represent potential direct or indirect interaction pathways within the kinetochore network.
The global defects in kinetochore assembly after inhibition of either Sgt1 or Dsn1 led us to ask if defects arise simply from protein mislocalization or if the link between misassembly and protein turnover extends to non-Mis12 kinetochore complexes. We found that siRNA directed against either Sgt1 or Dsn1 destabilized Ndc80Hec1, CENP-H, and CENP-K (Fig. 5 A). In contrast, CENP-N levels were only modestly affected, and CENP-U (CENP-O complex) levels did not change. The reported interaction between Mis12 and Ndc80Hec1 (Cheeseman et al., 2006), but not with CENP-N and CENP-U, is consistent with the idea that the assembly of other kinetochore complexes is linked to their stability. Importantly, siRNA directed against CENP-U or CENP-K (45% and 75% knockdown, respectively) did not significantly affect the levels of Mis12 subunits or Ndc80Hec1, making it unlikely that Sgt1 indirectly affects Mis12 complex stability (Fig. 5 B). We conclude that global kinetochore assembly defects after inhibition of Sgt1 or Mis12 components likely arise from the turnover of misassembled or unassembled complexes (see Discussion).
Inhibition of Sgt1 and Mis12 subunits gives rise to identical kinetochore defects. (A) HeLa cells were treated with control, Sgt1, or Dsn1 siRNAs for 72 h. Cell extracts were analyzed by immunoblotting to determine the levels of inner (CENP-K, -H, -U, and -N) and outer kinetochore proteins Ndc80Hec1. (B) HeLa cells were treated with control, CENP-U, and CENP-K siRNAs for 72 h, and extracts were analyzed by immunoblotting as in A using the indicated antibodies. For A and B, tubulin was used as a loading control, and brackets indicate CENP-U–specific bands. Asterisks represent nonspecific staining.
Inhibition of Sgt1 and Mis12 subunits gives rise to identical kinetochore defects. (A) HeLa cells were treated with control, Sgt1, or Dsn1 siRNAs for 72 h. Cell extracts were analyzed by immunoblotting to determine the levels of inner (CENP-K, -H, -U, and -N) and outer kinetochore proteins Ndc80Hec1. (B) HeLa cells were treated with control, CENP-U, and CENP-K siRNAs for 72 h, and extracts were analyzed by immunoblotting as in A using the indicated antibodies. For A and B, tubulin was used as a loading control, and brackets indicate CENP-U–specific bands. Asterisks represent nonspecific staining.
Sgt1 binds to and enhances Mis12 complex association with Hsp90
The genetic experiments presented strongly argue that the Mis12 complex interacts with Hsp90–Sgt1 as a client. To more directly test the chaperone–client relationship, we immunopurified Dsn1 from extracts of cells that had been arrested in mitosis with nocodazole and immunoblotted for Hsp90 or Sgt1; this interaction was observed in cycling cells but was less robust (unpublished data). We observed a modest but significant copurification of Hsp90 and Sgt1 with Dsn1, which is typical of levels observed with Hsp90 clients in cell extracts (Fig. 6 A; Hartson et al., 1999; Whitesell et al., 1998; Arlander et al., 2006). Consistent with its proposed role as a client adaptor (Catlett and Kaplan, 2006), 48 h of depletion of Sgt1 (to 36% of control) reduced the levels of Hsp90 that copurified with Dsn1 by >50% (Fig. 6 A). In addition, the fact that Hsp90 did not copurify with CENP-K or Ndc80Hec1 argues that its interaction with the Mis12 complex is specific and likely to occur before Mis12 incorporation into kinetochores (Fig. 6 B).
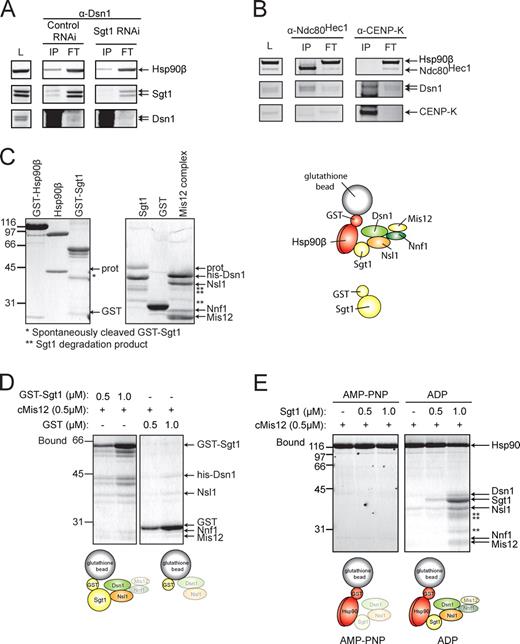
In vitro biochemical assays of the Sgt1–Hsp90 and Mis12 complex interactions. (A) HeLa cell extracts were subject to immunopurification (IP) after 48 h of treatment with control or Sgt1 siRNA and arrested for 16 h in nocodazole, using antibodies to Dsn1. The starting extract (L), the IP, and the post-IP supernatant (FT) were analyzed by immunoblotting using the indicated antibodies (labeled on the right side of the gel). (B) HeLa cell extracts were analyzed as in A, except that either Ndc80Hec1 or CENP-K was used for the immunoprecipitation. The immunoblot in the top panel was probed with antibodies against both Ndc80Hec1 and Hsp90. (C) The indicated proteins or Mis12 complex (referred to as Mis12) were purified, analyzed on SDS-PAGE, and stained with Coomassie dye. The migration position and cleavage products provide a reference for the same material used in the binding assays. The arrow labeled GST indicates the migration of cleaved GST; the arrow labeled prot indicates the migration position of the protease used to cleave GST from the fusion. *, the migration position of cleaved GST-Sgt1; **, the migration position of Sgt1 degradation. The diagram provides a key to complexes illustrated in the binding assays that follow. (D) The indicated µM concentration of GST-Sgt1 or GST were bound to glutathione beads and then used in the binding assay by adding Mis12 complex to 0.5 µM; complexes were identified that were associated with glutathione beads bound to the fusions. The diagram below the gel summarizes the binding observed, and the grayed subunits exhibit lower or nonspecific binding. Analysis of unbound fractions is included in Fig. S1. (E) GST-Hsp90 was preincubated with the indicated nucleotide (see Materials and methods) and then bound to the Mis12 complex in the presence or absence of Sgt1. Diagrams below the gels summarize the binding results. Numbers next to the gel blots indicate molecular mass in kD.
In vitro biochemical assays of the Sgt1–Hsp90 and Mis12 complex interactions. (A) HeLa cell extracts were subject to immunopurification (IP) after 48 h of treatment with control or Sgt1 siRNA and arrested for 16 h in nocodazole, using antibodies to Dsn1. The starting extract (L), the IP, and the post-IP supernatant (FT) were analyzed by immunoblotting using the indicated antibodies (labeled on the right side of the gel). (B) HeLa cell extracts were analyzed as in A, except that either Ndc80Hec1 or CENP-K was used for the immunoprecipitation. The immunoblot in the top panel was probed with antibodies against both Ndc80Hec1 and Hsp90. (C) The indicated proteins or Mis12 complex (referred to as Mis12) were purified, analyzed on SDS-PAGE, and stained with Coomassie dye. The migration position and cleavage products provide a reference for the same material used in the binding assays. The arrow labeled GST indicates the migration of cleaved GST; the arrow labeled prot indicates the migration position of the protease used to cleave GST from the fusion. *, the migration position of cleaved GST-Sgt1; **, the migration position of Sgt1 degradation. The diagram provides a key to complexes illustrated in the binding assays that follow. (D) The indicated µM concentration of GST-Sgt1 or GST were bound to glutathione beads and then used in the binding assay by adding Mis12 complex to 0.5 µM; complexes were identified that were associated with glutathione beads bound to the fusions. The diagram below the gel summarizes the binding observed, and the grayed subunits exhibit lower or nonspecific binding. Analysis of unbound fractions is included in Fig. S1. (E) GST-Hsp90 was preincubated with the indicated nucleotide (see Materials and methods) and then bound to the Mis12 complex in the presence or absence of Sgt1. Diagrams below the gels summarize the binding results. Numbers next to the gel blots indicate molecular mass in kD.
To study the Hsp90–Sgt1 interaction with the Mis12 complex in more detail, we separately purified recombinant Mis12 complex (His6-Dsn1, Nsl1, Nnf1, and Mis12), human GST-Sgt1 expressed in bacteria, and GST-Hsp90 expressed in Baculovirus-infected insect cells (Fig. 6 C and Materials and methods). Glycerol gradient sedimentation indicated that the bulk of Mis12 subunits migrate at the expected position for a properly assembled multiprotein complex (Fig. S1 A). We incubated purified Mis12 complex with two amounts of bead-bound GST-Sgt1 and visualized associated proteins. At low GST-Sgt1 levels (0.5 µM), we observed a specific and significant association with the Mis12 subunits, Dsn1 and Nsl1; at higher levels of GST-Sgt1 (1.0 µM), we observed enhanced Dsn1 and Nsl1 binding and now significant levels of the Mis12 and Nnf1 subunits (Fig. 6 D). We conclude that Sgt1 can directly interact with the Mis12 complex and that it may have a higher affinity for the Dsn1 and Nsl1 subunits.
We next examined the interaction between Mis12 complex and Hsp90. Mis12 complexes were incubated with GST-Hsp90 in the presence of ADP or the nonhydrolyzable ATP analogue, AMP-PNP. In the absence of Sgt1, we observed very weak binding of Nsl1 and Dsn1 independent of nucleotide (Fig. 6 E; −Sgt1 lanes). We observed similar weak binding of Nsl1 and Dsn1 in the presence of 0.5 µM or 1.0 µM Sgt1 and either AMP-PNP (Fig. 6 E) or 17-AAG (Fig. S2 C). Strikingly, binding of Mis12 complex to GST-Hsp90-ADP was dramatically enhanced after titrating in >1.0 µM Sgt1 (Figs. 6 E and S1 C). A similar result was obtained for Mis12 complexes isolated in glycerol gradient fractions (Fig. S1, A and B), which argues that under these conditions, GST-Hsp90 can interact with intact Mis12 complex and is not simply binding to misfolded proteins. These results are consistent with work on yeast complexes (Catlett and Kaplan, 2006) and demonstrate the conserved preference of the human Sgt1 for the ADP-bound form of human Hsp90. We also conclude that Sgt1 binds to Mis12 independently of Hsp90 and can mediate the formation of an Hsp90-ADP-Sgt1-client cocomplex, supporting a model whereby Sgt1 binds to Mis12 subunits and targets them to Hsp90 (see Discussion).
Skp1 balances Sgt1-dependent Mis12 complex stability with turnover
In budding yeast, turnover of the Hsp90–Sgt1 client complex CBF3 is linked to the SCF E3-ubiquitin ligase (Kaplan et al., 1997). In both yeast and human systems, Skp1 is an essential component of SCF and interacts with Sgt1 (Skowyra et al., 1997; Lyapina et al., 1998; Michel and Xiong, 1998; Yu et al., 1998; Zhang et al., 2008). To test if Skp1 and SCF are involved in the turnover of Mis12 complexes, we used double siRNA treatment. We reasoned that if Mis12 complex turnover requires SCF function, then Skp1 depletion should stabilize Mis12 subunits in the absence of Sgt1. Sgt1 siRNA alone results in reduced levels of Dsn1 and Ndc80Hec1 on mitotic kinetochores (Fig. 7, A and B) and increased frequency of misaligned chromosomes (Fig. 7 C). Strikingly, cotransfection of HeLa cells with Sgt1 and Skp1 siRNAs (see Fig. 7 D for Skp1 knockdown efficiency) suppressed the defect in Dsn1 and Ndc80Hec1 kinetochore localization (Fig. 6, A and B). The increase in kinetochore signals is accompanied by stabilization of protein levels (Fig. 8) and by formation of functional kinetochores, as indicated by a reduction in the frequency of cells with misaligned chromosomes (Fig. 7 C). Sgt1 levels remain low in the double siRNA-treated cells, which demonstrates that Skp1 does not rescue kinetochore defects by stabilizing Sgt1 (Fig. 8 E). Collectively these results suggest that Hsp90–Sgt1, together with Skp1, act in a conserved pathway that balances kinetochore complex stability with turnover.
Skp1 is required for Mis12 complex turnover after Sgt1 siRNA. (A) HeLa cells were treated with control, Sgt1, and Sgt1+Skp1 siRNAs for 72 h. Cells were processed for immunofluorescence with antibodies against tubulin, Dsn1, and ACA; chromosomes were stained with DAPI as indicated. Bar, 10 µM. (B) Fluorescent values from Dsn1 or Ndc80Hec1 signal were quantified for the indicated siRNA treatments. The mean intensities of control siRNA for multiple kinetochores, cells, and experiments were set at 100%. The error bars represent the deviation from the mean and are consistent with results from multiple experiments. (C) Metaphase-type cells with misaligned chromosomes (see the example in A; Sgt1 siRNA) were counted for the indicated siRNA treatments and divided by the total number of metaphase cells (n = 50 cells for each treatment group). (D) siRNA knockdown efficiency of Skp1 was analyzed by immunoblotting, quantified, and compared with control protein levels. Error bars represent the standard deviation over multiple blots (n = 6).
Skp1 is required for Mis12 complex turnover after Sgt1 siRNA. (A) HeLa cells were treated with control, Sgt1, and Sgt1+Skp1 siRNAs for 72 h. Cells were processed for immunofluorescence with antibodies against tubulin, Dsn1, and ACA; chromosomes were stained with DAPI as indicated. Bar, 10 µM. (B) Fluorescent values from Dsn1 or Ndc80Hec1 signal were quantified for the indicated siRNA treatments. The mean intensities of control siRNA for multiple kinetochores, cells, and experiments were set at 100%. The error bars represent the deviation from the mean and are consistent with results from multiple experiments. (C) Metaphase-type cells with misaligned chromosomes (see the example in A; Sgt1 siRNA) were counted for the indicated siRNA treatments and divided by the total number of metaphase cells (n = 50 cells for each treatment group). (D) siRNA knockdown efficiency of Skp1 was analyzed by immunoblotting, quantified, and compared with control protein levels. Error bars represent the standard deviation over multiple blots (n = 6).
The balance of assembly and turnover of Mis12 complexes contributes to the efficiency and distribution of microtubule attachment sites. (A) The fluorescent signal associated with Dsn1 at kinetochores was quantified in cells treated with control (blue) or Sgt1 siRNA (green). Signals for each kinetochore from multiple mitotic cells are represented in the scatter plot. The fluorescent cell images are examples of the data used for the plot. (B) The fluorescent signals from kinetochores only from cells treated with Sgt1 siRNA were divided into two populations: those from kinetochores of aligned or misaligned chromosomes. The signals from these two populations are represented in a scatter plot. The grayscale images on the right highlight the boxed regions as examples of Dsn1 fluorescence from aligned or misaligned kinetochores. After treatment with Sgt1 (C), control, or Ndc80Hec1 (D) siRNAs, cells were briefly cold-treated to destabilize nonkinetochore microtubules (see Materials and methods). Cells were processed for immunofluorescence and stained for tubulin, ACA, and chromosomes with DAPI. Multiple images for cells treated with Sgt1 siRNA exemplify the heterogeneity of kinetochore defects. Arrows indicate unattached kinetochores. (E) HeLa cells were treated with control, Sgt1, or Sgt1 and Skp1 siRNAs for 72 h, and extracts were subject to immunoblot analysis of the indicated proteins. The fluorescent signals associated with the indicated proteins (Dsn1, Nsl1, Mis12, and Nnf1) were quantified, and control treatment was set to 100%. Signals were normalized to tubulin, which serves as a loading control. (F) After treatment of HeLa cells with control or Skp1 siRNA and processing for immunofluorescence, the maximum fluorescent intensities of kinetochore-associated Dsn1, Nnf1, and Ndc80Hec1 from multiple mitotic cells were quantified. P-values were obtained from a two-tailed Student’s t test. (G) HeLa cells were treated with control or Skp1 siRNAs for 72 h and briefly cold-treated to eliminate non–kinetochore-bound microtubules. Cells were processed for immunofluorescence and stained with ACA and tubulin antibodies. Arrows indicate kinetochores with little or no associated microtubule signal. Bars: (A and B, left) 10 µm; (B, right) 1 µM; (C, D, and G) 10 µM.
The balance of assembly and turnover of Mis12 complexes contributes to the efficiency and distribution of microtubule attachment sites. (A) The fluorescent signal associated with Dsn1 at kinetochores was quantified in cells treated with control (blue) or Sgt1 siRNA (green). Signals for each kinetochore from multiple mitotic cells are represented in the scatter plot. The fluorescent cell images are examples of the data used for the plot. (B) The fluorescent signals from kinetochores only from cells treated with Sgt1 siRNA were divided into two populations: those from kinetochores of aligned or misaligned chromosomes. The signals from these two populations are represented in a scatter plot. The grayscale images on the right highlight the boxed regions as examples of Dsn1 fluorescence from aligned or misaligned kinetochores. After treatment with Sgt1 (C), control, or Ndc80Hec1 (D) siRNAs, cells were briefly cold-treated to destabilize nonkinetochore microtubules (see Materials and methods). Cells were processed for immunofluorescence and stained for tubulin, ACA, and chromosomes with DAPI. Multiple images for cells treated with Sgt1 siRNA exemplify the heterogeneity of kinetochore defects. Arrows indicate unattached kinetochores. (E) HeLa cells were treated with control, Sgt1, or Sgt1 and Skp1 siRNAs for 72 h, and extracts were subject to immunoblot analysis of the indicated proteins. The fluorescent signals associated with the indicated proteins (Dsn1, Nsl1, Mis12, and Nnf1) were quantified, and control treatment was set to 100%. Signals were normalized to tubulin, which serves as a loading control. (F) After treatment of HeLa cells with control or Skp1 siRNA and processing for immunofluorescence, the maximum fluorescent intensities of kinetochore-associated Dsn1, Nnf1, and Ndc80Hec1 from multiple mitotic cells were quantified. P-values were obtained from a two-tailed Student’s t test. (G) HeLa cells were treated with control or Skp1 siRNAs for 72 h and briefly cold-treated to eliminate non–kinetochore-bound microtubules. Cells were processed for immunofluorescence and stained with ACA and tubulin antibodies. Arrows indicate kinetochores with little or no associated microtubule signal. Bars: (A and B, left) 10 µm; (B, right) 1 µM; (C, D, and G) 10 µM.
Hsp90–Sgt1 and Skp1 ensure efficient formation of high-affinity kinetochore–microtubule attachment sites
Overall, our findings are consistent with Sgt1 and Hsp90 being important for the efficient formation of kinetochore–microtubule sites. Sgt1 or Dsn1 siRNA reduce but do not eliminate kinetochore-bound (Fig. 4, A and E) and soluble pools of Ndc80Hec1 (Fig. 5 A). Our observations as well as published data demonstrate only a partial chromosome alignment defect after Sgt1 or Dsn1 siRNA treatment. The partial chromosome alignment defects could arise because only some centromeres have formed fully functional microtubule attachment sites or because partially functional sites have formed on all centromeres. To begin to distinguish between these possibilities, we imaged various kinetochore markers after treatment of cells with Sgt1 siRNA. We found that all chromosomes contained reduced Dsn1 signal in the absence of Sgt1; quantification of individual kinetochore signals showed a very narrow distribution that clustered near the low end of signals found in control cells (Fig. 8 A). We then compared the intensities of Dsn1 on chromosomes that appeared aligned with those that had clearly failed to align at the metaphase plate (i.e., misaligned chromosomes); we observed that misaligned chromosomes have on average lower Dsn1 signals compared with signals on aligned chromosomes (Fig. 8 B). To determine if the assembly of Mis12 complexes is simply delayed (i.e., less efficient) in the absence of Sgt1, we analyzed post-metaphase cells after Sgt1 or Dsn1 depletion. Although depletion of Sgt1 results in enrichment in mitotic cells, a significant percentage of these cells proceed past metaphase (Fig. S3 C). Interestingly, in these early anaphase cells, Dsn1 levels return to near normal levels, arguing that assembly and recruitment of Mis12 complexes to the kinetochore is delayed in the absence of Sgt1. Although we cannot rule out that Sgt1 depletion is less efficient in those anaphase cells, the heterogeneity in Dsn1 signals contrasts with the situation in control cells and argues that these cells originated from metaphases with defective kinetochores. In addition, the behavior of anaphase cells after Sgt1 depletion contrasts with Dsn1 depletion, which have fewer cells that proceed past metaphase, maintain low levels of Dsn1, and exhibit lagging chromosomes in anaphase (Fig. S3, C and D). We take these data to indicate that Sgt1 is required for the timely assembly of Mis12 and kinetochore complexes.
The delayed assembly of Mis12 complexes is predicted to compromise the formation of kinetochore–microtubule binding sites. To test this possibility, we cold-treated control siRNA-treated cells so that only kinetochore microtubules that had made stable plus-end attachments were preserved (Salmon and Begg, 1980). In control cells, a robust spindle structure was preserved in metaphase cells, and bundles of microtubules were observed to terminate in kinetochores (Fig. 8 D). In contrast, Sgt1-depleted cells with partially aligned chromosomes exhibited disorganized spindle structures, with many kinetochores showing a complete absence of microtubule attachments (Fig. 8 C, arrowheads). We also observed a range of defects among treated mitotic cells, with some cells having nearly normal spindle structures (Fig. 8 C, top), a heterogeneity that we argue is consistent with the delayed formation of functional microtubule-binding sites. In contrast to these heterogeneous microtubule attachment defects, reduction in Ndc80Hec1 prevents nearly all microtubule bundles after cold treatment (Fig. 8 D, bottom). These results support the idea that Hsp90–Sgt1 is required, through stabilization of Mis12 complexes, to ensure the efficient formation of microtubule attachment sites.
We next considered the possibility that the turnover of Mis12 complexes may eliminate improperly assembled kinetochore complexes, thus ensuring the fidelity of microtubule binding site formation. Inhibition of Skp1 alone did not obviously affect chromosome alignment or increase the number of lagging chromosomes in anaphase (unpublished data). We did observe a modest increase in the number of cells with multipolar spindles, which is consistent with the reported role of Skp1 and SCF in centrosome integrity (unpublished data; Murphy, 2003). Analysis of protein levels by immunoblotting shows an elevation of Nsl1, Nnf1, and Mis12 but not Dsn1 in cells treated with Skp1 siRNA (Fig. 8 E). Although we found an increase in Mis12 complexes at the kinetochore (Fig. 8 F), there was no overall increase in the levels of Ndc80Hec1 (Fig. 8 F), which suggests that the additional Mis12 complexes are only partially functional. To evaluate whether partially functional Mis12 complexes might compromise microtubule attachments, we analyzed cold-stable microtubules. Surprisingly, we observed fewer cold-stable kinetochore–microtubule bundles after siRNA against Skp1 (Fig. 8 G, arrows; and Fig. S3). Even where kinetochores retained cold-stable microtubules, these were less robust than control cells. The decrease in the robustness of kinetochore–microtubule attachments but otherwise normal alignment suggests that high-affinity attachment sites form with normal kinetics, but that there are fewer per kinetochore (see Discussion). Together, these observations suggest that Skp1-mediated turnover of Mis12 complexes serves a nonessential role in ensuring optimal numbers of functional microtubule-binding sites.
Discussion
In this work, we sought to identify the kinetochore targets of mammalian Hsp90–Sgt1 to gain insight into the mechanism of chaperone-mediated kinetochore assembly. We found that Sgt1 binds to and targets the Mis12 complex to Hsp90 and is required for Mis12 stability and kinetochore association. Turnover of Mis12 complexes depends on the SCF ubiquitin ligase subunit, Skp1. Reminiscent of kinetochore complexes in budding yeast, we found that the balance of assembly and turnover directly contribute to kinetochore function in mammalian cells. Hsp90–Sgt1–mediated assembly of Mis12 complexes is necessary for the timely formation of high-affinity microtubule-binding sites, whereas the turnover of Mis12 complexes is required to ensure the proper number of microtubule attachments. We propose that the balance of complex assembly and turnover mediated by Hsp90–Sgt1 and SCF ensures the fidelity of the microtubule-binding site.
Hsp90–Sgt1 targets Mis12 complexes
The argument that the Mis12 complex is the main client of the Hsp90–Sgt1 chaperone complex is based on both genetic and biochemical evidence. The Mis12 subunit, Dsn1, is the first kinetochore protein to exhibit decreased levels after treatment of cells with the Hsp90 inhibitor, 17-AAG. There are numerous examples of degraded Hsp90 clients; the likely explanation is that the failure of Hsp90 to release or bind its client results in ubiquitin-mediated degradation. Similar to treatment with 17-AAG, siRNA of Sgt1 causes a dramatic decrease in Dsn1 but also results in the loss of Nsl1 and a significant decrease in the other Mis12 subunits, Nnf1 and Mis12. Our interpretation is that Sgt1 mediates the targeting of Dsn1 and Nsl1 to Hsp90; in the absence of targeting and presumably assembly, Nnf1 and Mis12 become susceptible to destruction machinery as well. Although we cannot rule out the possibility that Hsp90–Sgt1 has an additional client in the kinetochore, the fact that Dsn1 siRNA mimics the Sgt1 siRNA defects in kinetochore complex stability and recruitment strongly argues that the Mis12 complex is the main client responsible for kinetochore defects.
The biochemical reconstitution of human Hsp90, Sgt1, and Mis12 clearly shows that the Sgt1 and Mis12 complexes both interact with Hsp90-ADP, and this argues for the formation of a cocomplex between Hsp90-ADP and the Sgt1 and Mis12 subunits. The fact that Sgt1 can bind independently to Mis12 subunits and enhance the affinity of the Mis12 complex for Hsp90 is consistent with the client adaptor function previously proposed for yeast Sgt1 (Catlett and Kaplan, 2006). Our biochemical reconstitution took advantage of the stable and soluble behavior of an assembled Mis12 complex coexpressed in bacteria (unpublished data; Cheeseman et al., 2006; Kline et al., 2006). Despite starting with a fully assembled Mis12 complex, we observed an enrichment of the Dsn1-Nsl1 subunits associated with Hsp90–Sgt1. In the absence of Sgt1, Hsp90 binds low levels of Dsn1 and Nsl1 but not Nnf1 or Mis12. Interestingly, this occurs independently of nucleotide and may represent a weak initial interaction between Hsp90 and client that is stabilized by Sgt1 and ADP. Further examination of Hsp90–Sgt1 interactions with Mis12 subcomplexes and individual subunits will be important for ordering the steps of chaperone complex formation. Although we speculate that Hsp90–Sgt1 aids in Mis12 complex assembly, our data specifically shows that the two complexes interact and that inhibition of either Sgt1 or Hsp90 leads to protein turnover. Again, reconstitution of the complete client–chaperone complex with individual Mis12 subunits will be required to understand the precise role of Hsp90–Sgt1 in Mis12 complex assembly.
One of the more intriguing observations presented here is that siRNA of Skp1 suppresses the Mis12 complex instability induced by siRNA of Sgt1. We argue that Sgt1 is required for Mis12 complex stability, presumably by mediating complex assembly. Our results are also consistent with the idea that Skp1 and SCF are involved in the turnover of the Mis12 complex. Indeed, siRNA of Skp1 alone results in a modest but significant increase in the levels of Mis12 subunits in extracts as well as those recruited to the kinetochore. As SCF has a broad range of substrates, understanding its specific role at the kinetochore will require identifying the F-box subunit that targets Mis12 complexes. Regardless of which SCF complex is involved, it is striking that the same balance between stability and turnover that controls CBF3 levels and function in budding yeast also plays a role in human kinetochore function.
Balance of keystone Mis12 complex stability with turnover
The Mis12 complex has been described as a “keystone” kinetochore complex (Cheeseman and Desai, 2008), as it makes contacts with several other kinetochore complexes both proximal (e.g., CENP-H) and distal (e.g., Ndc80Hec1) to the centromeric DNA. In its role as a keystone complex, Mis12 complex is proposed to act as a node in the context of a network of protein complexes (Knl1, Ndc80, and motors) that form low- and high-affinity microtubule attachment sites. Although it is unclear what properties make the Mis12 complex uniquely require Hsp90 function, one interestingly possibility is that chaperone activity is required for appropriate protein contacts between Mis12 subunits and other kinetochore complexes. In this scenario, when Dsn1 or another Mis12 subunit binds to Hsp90–Sgt1, a stable interaction surface may be exposed that is required for the ordered assembly of other Mis12 proteins as well as interactions with other kinetochore complexes. In principle, mistakes in this process would result in incorrectly formed subcomplexes that either fail to get recruited to the kinetochore or fail to form robust microtubule-binding networks. In the case of treatment with Skp1 siRNA, we observed stabilized Mis12 subunits and an increase in fluorescent intensity at kinetochores but no increase in Ndc80Hec1 at kinetochores. Indeed, analysis of cold-stable kinetochore–microtubules argues that although kinetochore–microtubule attachments form efficiently when Skp1 is inhibited, they are less robust than controls. Our interpretation of this result is that improperly assembled complexes are recruited to kinetochores and form nonoptimal microtubule attachment sites. In this light, a more detailed characterization of Mis12 complexes that form in the absence of Skp1 or Sgt1 will be informative.
The characteristics of the Mis12 complex that support its role as a keystone kinetochore complex are shared with the CBF3 complex, the Hsp90–Sgt1 client in budding yeast. It is possible that the Mtw1 complex in yeast, which is homologous to the Mis12 complex in humans, is also an Hsp90–Sgt1 client. However, interactions between the Mtw1 complex and Hsp90–Sgt1 have not been reported, and sequence conservation is poor between the human and yeast subunits (15% or less for between subunits; unpublished data). The CBF3 complex may be more functionally similar to human Mis12 complex, as it also plays a “keystone” like role, recruiting multiple kinetochore complexes to centromeric DNA. Like Mis12 complexes, CBF3 has been shown to have multiple protein–protein contacts within the CBF3 complex and with other proteins, including but not limited to Cep3, Ctf13, Okp1, Bir1, and Smt3 (sumo) (Ortiz et al., 1999; Gillis et al., 2005; Montpetit et al., 2006). Indeed, mutants that affect CBF3 assembly can alter the stability of associated protein complexes (including Bir1 and Sli15; Thomas and Kaplan, 2007), similar to the behavior of the Mis12 complex observed in this work. We speculate that like the Mis12 complex in human cells, CBF3 also requires the Hsp90–Sgt1 chaperone complex to form appropriate protein–protein interactions, both at the intracomplex level and between other protein complexes. We argue that this property is critical for the ability of CBF3 to carry out multiple, distinct mitotic functions. It will be interesting to determine if Mis12 complexes carry out nonkinetochore functions. For example, inhibition of Mis12 components in fission yeast has been reported to alter chromatin organization during mitosis (Nakazawa et al., 2008). Further analysis of Mis12 complex siRNA phenotypes in mitosis as well as anaphase will be required to address this possibility.
Materials and methods
Cell culture, siRNAi transfection, and Hsp90-inhibitor treatments
Hela cell cultures were maintained in DME supplemented with glutamine, sodium pyruvate, penicillin-streptomycin (Invitrogen), and 10% FBS (Sigma-Aldrich).
Hsp90 inhibitor 17-AAG (Kosan Biosciences) solubilized in DMSO, or a DMSO vehicle control, were added to fresh media and then to cell cultures seeded at ∼50% confluency at dosages of either 100 nM or 500 nM, as indicated. Cell cultures treated with siRNA were transferred from DME to Opti-MEM, then supplemented with glutamine before siRNA transfection. siRNA reagents were obtained from Thermo Fisher Scientific and transfected at the indicated concentrations (Table S2) using Oligofectamine in Opti-MEM media (Invitrogen). FBS was added to 10% 4–6 h after transfection, and cultures were incubated as indicated.
Cloning and expression of recombinant proteins
Isolation of human Sgt1 and Hsp90β was performed by PCR amplification from a human leukocyte cDNA library (Takara Bio Inc.). Sgt1 and Hsp90β were amplified as described previously (Catlett and Kaplan, 2006) to introduce cloning sites allowing ligation into pGEX6-P1 (Invitrogen) or pMIT-77 (pFASTBAC-GST containing a PreScission protease cleavage site; GE Healthcare) expression vectors, respectively. Coding sequences were verified by DNA sequencing before protein expression. Human GST-Sgt1 and human Mis12 complex (from a polycistronic vector as described previously; Kline et al., 2006) were produced in BL21 DE3 cells using 0.1 mM IPTG to induce expression and grown for 5 h at 25°C. Human GST-Hsp90β was expressed in insect cells using the Bac-to-Bac expression system (Invitrogen).
Immunoblotting, in vitro binding assay, and immunoprecipitation
For immunoblot analysis, cells treated with siRNA or 17-AAG were rinsed in PBS (10 mM Na2HPO4, 2 mM KH2PO4, 137 mM NaCl2, and 3 mM KCl2, pH 7.4) and suspended in lysis buffer (10 mM Hepes, pH 8.0, 150 mM NaCl, 50 mM β-glycerolphosphate, 0.1 mM EDTA, 1% Triton X-100, 1 mM dithiothreitol, and 10% glycerol) plus protease inhibitors (1 mM phenylmethylsulfonyl fluoride, 1 mM N-tosyl-L-phenylalanine chloromethyl ketone, 10 µg/ml leupeptin, 10 µg/ml pepstatin, and 10 µg/ml chymostatin). Cells were incubated in lysis buffer for 15 min on ice and centrifuged at 14,000 rpm for 15 min at 4°C, then the soluble fraction was collected for analysis. Protein concentrations were measured by a Bradford assay (Bio-Rad Laboratories). For immunoblots, 40 µg of protein was loaded per lane on NuPAGE 4–12% 1.5-mm gradient gels in MES running buffer (Invitrogen) and transferred to 0.2 µM nitrocellulose (GE Healthcare) for 5 h in transfer buffer (20 mM tris base, 0.2 M glycine, and 20% methanol) at 4°C.
Immunoblots were blocked in 2% nonfat dry milk in PBS for 1 h and rinsed briefly in PBS, then primary antibodies were added (see Table S2 for working concentrations) in PBST-BSA (PBS, 0.01% Tween 20, and 2% BSA) for 1–3 h depending on the antigen. Membranes were rinsed three times for 5 min in PBST (PBS and 0.01% Tween 20) and secondary antibodies, goat anti–rabbit IgG 800 (LI-COR Biosciences), or goat anti–mouse IgG 680 (LI-COR Biosciences), then added 1/10,000 to PBST-BSA and incubated for 1 h. Membranes were rinsed two times for 5 min in PBST and once for 5 min in PBS before imaging. Immunoblots were scanned using the Odyssey system analyzed using Odyssey software version 3.0 (both from LI-COR Biosciences). This method increased blotting sensitivity that was not possible using chemiluminescent approaches.
Recombinant proteins were isolated using Ni-NTA agarose (QIAGEN) or glutathione affinity chromatography (GE Healthcare), incubated, and washed in 1% Triton X-100 lysis buffer to remove contaminating proteins and prepared for analysis by SDS-PAGE as described previously (Catlett and Kaplan, 2006). Binding assays were conducted as described previously in a total reaction volume of 200 µl (Catlett and Kaplan, 2006) with the following variations. Bead-bound GST-Hsp90β was incubated in binding buffer (25 mM Tris, pH 8.0, 150 mM KCl, 0.05% Triton X-100, 10% glycerol, 1 mM dithiothreitol, 10 mM MgCl2, and protease inhibitors as described above) plus 10 mM ATP for 1 h, rinsed twice in binding buffer, and then used in binding assays as previously described for 2 h at 37°C. Bead-bound fractions were suspended in Laemmli buffer, resolved on 12.5% SDS-PAGE, and stained in Gel-Code Blue (Thermo Fisher Scientific). The stained SDS-PAGE gels were scanned and quantified using the Odyssey system as recommended by the manufacturer.
Immunoprecipitation reactions were performed from Hela cell extracts prepared in lysis buffer as described above. Cells were treated with 100 ng/ml nocodazole for 16 h before extraction to enrich for mitotic cells. Protein concentrations of extracts were measured by a Bradford assay. For each reaction, 4 µg of primary antibody was incubated with 2.5 mg of Hela cell extract suspended in 200 µl of lysis buffer containing 10 mM ADP, 10 mM MgCl2, and protease inhibitors for 2 h at 4°C. 10 µl of protein A–sepharose CL-4B (Sigma-Aldrich) was added and incubated for 1 h at 4°C. Beads were rinsed four times in 500 µl of lysis buffer and resuspended in Laemmli buffer for SDS-PAGE and Western transfer.
Immunofluorescence and image acquisition
Cells were grown on coverslips under the culture conditions described above. Fixation and processing of coverslips for staining was performed in PHEM buffer (60 mM Pipes, 25 mM Hepes, 10 mM EGTA, and 2 mM MgCl2, pH 6.9) and 4% formaldehyde as described previously (Kline et al., 2006), with the exception of CENP-K and CENP-U analysis. For staining of CENP-U and CENP-K, coverslips were prepared by fixing cells in absolute methanol at −20°C for 20 min. Cells were rinsed 3× in PBS in between incubations. Primary and secondary incubations were conducted for 1 h at room temperature in PBS containing 0.1% Triton X-100 and 10% BSA. For cold-stable analysis, cells were incubated on ice for 15 min in Opti-MEM, then fixed for 10 min on ice followed by 10 min at room temperature using PHEM buffer containing 4% formaldehyde and 0.2% Triton X-100. Cells were then processed as above. Goat anti–mouse IgG Alexa Fluor 594 or 498 (Invitrogen)-, goat anti–rabbit IgG Alexa Fluor 594 (Invitrogen)-, or goat anti–human IgG Cy5 (Jackson ImmunoResearch Laboratories, Inc.)-conjugated secondary antibodies were used at 1:200. See Table S1 for primary antibody working dilutions.
Images were collected at room temperature using an inverted microscope (IX 70; Olympus) fitted to a Deltavision RT imaging system (Applied Precision). Images were recorded using a 60× NA 1.4 oil immersion lens and a CoolSnap HQ camera (Roper Industries). Deconvolution was performed using the SoftWorx v 3.5.1 software package (Applied Precision). Measurements of kinetochore intensities were conducted using maximum intensity projections of deconvolved images. Kinetochore exposure settings were held constant within each group of experiments, and kinetochore signals within each figure were scaled equally. Maximal kinetochore intensities and corresponding background levels were measured from radial 7 × 7 pixel areas for each kinetochore foci using ImageJ version 1.42 software (http://rsb.info.nih.gov/ij/) and exported to Excel software (Microsoft) for analysis. A minimum of 6 cells and 12 kinetochore foci per cell were measured for each condition (each experiment was replicated a minimum of three times). Kinetochore foci intensities were then pooled within a given experiment, and the mean kinetochore fluorescence intensity was calculated. Statistical significance was evaluated by a two-tailed Student’s t test; results with p-values <0.05 were considered statistically significant.
Online supplemental material
Fig. S1 is an analysis of recombinant Mis12 complex purity and composition by glycerol gradient followed by in vitro binding analysis to Hsp90–Sgt1 and an analysis of Sgt1-dependent Mis12 complex binding. Fig. S2 contains unbound fractions from binding assays depicted in Fig. 5. In addition, an Hsp90–Sgt1–Mis12 complex binding assay conducted in the presence of 17-AAG is provided. Fig. S3 contains single z projections of microtubule attachment site defects seen in Skp1-treated Hela cells. Control, Dsn1, and Sgt1 siRNA-treated anaphase cells are also provided to illustrate the increase in kinetochore-bound Dsn1 during anaphase in Sgt1 depleted cells. Table S1 lists antibodies used in this study. Table S2 lists siRNA reagents.
Acknowledgments
We are grateful to Arshad Desai for providing antibodies against the human Mis12 complex, Iain Cheeseman and Arshad Desai for providing the human Mis12 polycistronic expression vector, Patrick Meraldi for antibodies against CENP-N, Topher Carroll for antibodies against CENP-H, and Simon Chan and Daniel Starr for reading and providing feedback on the manuscript.
This work was funded by Research Scholar Grants from the American Cancer Society (RSG-02-035-01 and RSG-02-035-05-CCG) to K.B. Kaplan and funding from the National Center for Research Resources (NCRR; grant UL1 RR024146) to A.E. Davies.