Hematopoietic stem cells (HSCs) are maintained through the regulation of symmetric and asymmetric cell division. We report that conditional ablation of the RNA-binding protein Msi2 results in a failure of HSC maintenance and engraftment caused by a loss of quiescence and increased commitment divisions. Contrary to previous studies, we found that these phenotypes were independent of Numb. Global transcriptome profiling and RNA target analysis uncovered Msi2 interactions at multiple nodes within pathways that govern RNA translation, stem cell function, and TGF-β signaling. Msi2-null HSCs are insensitive to TGF-β–mediated expansion and have decreased signaling output, resulting in a loss of myeloid-restricted HSCs and myeloid reconstitution. Thus, Msi2 is an important regulator of the HSC translatome and balances HSC homeostasis and lineage bias.
Hematopoiesis is a tightly orchestrated process in which the hematopoietic stem cell (HSC) goes through symmetric and asymmetric divisions to self-renew and also to differentiate into progenitors that can give rise to different cell lineages (Brümmendorf et al., 1999; Beckmann et al., 2007; Wu et al., 2007). The balance between self-renewal and differentiation of the HSCs needs to be regulated for supporting a normal hematopoietic system. However, not much is known about the programs that regulate this balance.
The Musashi (Msi) family of RNA-binding proteins, including Msi1 and Msi2, contribute to the control of symmetric and asymmetric stem cell division, regulate stem cell function, and play a role in cell fate determination (Okano et al., 2005). In Drosophila melanogaster, Musashi was first identified to be important for sensory neuron development, where it is essential for asymmetric division of neuroblasts into daughter cells with neuronal and nonneuronal potential (Nakamura et al., 1994). Loss of Musashi function results in defective asymmetric division and the formation of two nonneuronal daughter cells at the expense of commitment to the neuronal lineage (Nakamura et al., 1994). In vertebrates, Msi1 and Msi2 are evolutionarily conserved, harboring two tandem RNA recognition motifs and a carboxyl-terminal poly-A–binding protein association domain (Okano et al., 2005; Kawahara et al., 2008).
Msi proteins are thought to function by binding to the 3′ untranslated regions (UTRs) of target mRNAs at a consensus sequence and then blocking translation by hindering access of the poly-A–binding protein to the elongation initiation complex (Kawahara et al., 2008). An inverse relationship between Msi and Numb expression as a result of Msi-mediated repression of Numb has been demonstrated in a variety of systems, including myeloid leukemia cells (Imai et al., 2001; Ito et al., 2010; Kharas et al., 2010). Msi2 is an important modulator of proliferation and differentiation in both normal HSCs and in myeloid malignancies (Hope et al., 2010; Ito et al., 2010; Kharas et al., 2010; de Andrés-Aguayo et al., 2011). Analysis of the hematopoietic compartment of Msi2 gene trap mice revealed a reduced number of short-term HSCs and lymphoid primed myeloid progenitor (LMPP) cells, but no significant defect was found in long-term HSCs (de Andrés-Aguayo et al., 2011). Although Msi2 is most highly expressed in the primitive hematopoietic compartment, and MSI2 overexpression drives quiescent HSCs out of G0 and into cycle (Kharas et al., 2010), it remains unclear whether and how Msi2 affects HSC self-renewal and commitment under homeostatic conditions. Furthermore, the critical RNA-binding targets of Msi2 in hematopoietic cells that regulate self-renewal and lineage commitment remain to be uncovered.
To determine the role of Msi2 in HSCs and avoid potentially confounding compensatory mechanisms arising from germline Msi2 loss, we generated Msi2 conditional knockout mice that allowed us to study Msi2 function in a cell-autonomous manner in adult tissues using spatiotemporally controlled deletion. Here, analysis of microarray data of Msi2 conditional knockout mice coupled with MSI2 HITS-CLIP (cross-linking and immunoprecipitation followed by high-throughput sequencing) profiling data allowed us to identify novel regulatory pathways downstream of Msi2 in HSCs (Chi et al., 2009).
RESULTS
Msi2 is required to maintain normal HSC numbers
To assess the role of Msi2 in the hematopoietic compartment, we developed a conditional knockout mouse model. We targeted the Msi2 locus in embryonic stem cells with a construct containing loxP sites flanking the first four Msi2 exons (Fig. 1 a). After removal of the neomycin resistance selection cassette, a Msi2flox/flox mouse colony was established and crossed with Mx1-Cre mice to generate an inducible Msi2 loss of function strain (Msi2flox/flox::Mx1-Cre). To delete the Msi2 gene in cells of the hematopoietic lineage, we induced the Cre transgene in Msi2flox/flox::Mx1-Cre mice by three polyinosinic:polycytidylic acid (pIpC) injections, which efficiently excised the Msi2 gene from the BM and spleen, as assessed by Southern blot and quantitative real-time PCR (qRT-PCR) analysis within the hematopoietic stem and progenitor cells (HSPCs; LSK, Lineageloc-kit+, Sca+; Fig. 1, b and c). Msi2-deleted mice will be referred to as Msi2Δ/Δ and control mice as either Msi2f/f or Msi2wt/Δ (heterozygous mice were phenotypically and functionally the same as Msi2f/f).
Msi2 conditional knockout mice have reduced HSC numbers. (a) Targeting scheme for Msi2 conditional knockout mice. (b) Southern blot of the indicated genotypes 4 wk after pIpC treatment in vivo after XbaI digestion of genomic DNA and hybridization with the probe depicted in panel a. (c) qRT-PCR of Msi2 normalized to Gapdh from LSK (lineagelo, Sca+Kit+)-sorted cells from mice 1 mo after pIpC injection (n = 3 per group). (d) Overall cell counts in mice as indicated after pIpC in the BM (left) and spleen (right; 3–6 wk, n = 4; 18–22 wk, n = 9, 10 from two independent experiments). (e) Representative flow cytometric analysis from mice 3–6 wk after pIpC (mean and SEM; n = 12; three independent experiments). (f and g) Absolute number of LSK (f) and LSK+CD150+CD48− cells (g) from the indicated mice after pIpC (3–6 wk: same mice as e; and18–22 wk: Msi2f/f, n = 12; Msi2Δ/Δ, n = 13 from four independent experiments). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
Msi2 conditional knockout mice have reduced HSC numbers. (a) Targeting scheme for Msi2 conditional knockout mice. (b) Southern blot of the indicated genotypes 4 wk after pIpC treatment in vivo after XbaI digestion of genomic DNA and hybridization with the probe depicted in panel a. (c) qRT-PCR of Msi2 normalized to Gapdh from LSK (lineagelo, Sca+Kit+)-sorted cells from mice 1 mo after pIpC injection (n = 3 per group). (d) Overall cell counts in mice as indicated after pIpC in the BM (left) and spleen (right; 3–6 wk, n = 4; 18–22 wk, n = 9, 10 from two independent experiments). (e) Representative flow cytometric analysis from mice 3–6 wk after pIpC (mean and SEM; n = 12; three independent experiments). (f and g) Absolute number of LSK (f) and LSK+CD150+CD48− cells (g) from the indicated mice after pIpC (3–6 wk: same mice as e; and18–22 wk: Msi2f/f, n = 12; Msi2Δ/Δ, n = 13 from four independent experiments). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
Msi2Δ/Δ mice had normal peripheral blood counts (not depicted) and BM and spleen cellularity at 3–6 wk after pIpC injections (Fig. 1 d). However, after 18 wk, the mice had reduced spleen weights (not depicted) and cellularity in the spleen and BM (Fig. 1 d). We previously observed alterations in myeloid differentiation upon MSI2 overexpression in vivo (Kharas et al., 2010). In contrast, we found no significant changes in the frequencies of mature myeloid cell types as well as B and T cells in the BM and spleen (not depicted). The decreased cellularity in both spleen and BM and phenotypes from a previous study on Msi2 suggested that there could be a defect in early stem and/or progenitor function (de Andrés-Aguayo et al., 2011). Thus, we examined the overall frequency and numbers of HSCs in these mice and found a reduction in the frequency and absolute number of LSKs as early as 4 wk and reduced overall numbers of HSCs at 18 wk (Fig. 1, e–g; and Fig. S1).
Msi2Δ/Δ HSCs are functionally defective in reconstitution
To test the function of Msi2Δ/Δ HSCs, we transplanted the BM of Msi2Δ/Δ or Msi2wt/Δ into congenic recipients. After a noncompetitive engraftment, we observed a reduction in spleen size and a dramatic reduction of donor chimerism at the level of phenotypic LSK+CD150+CD48− LSKs and hematopoietic progenitors after 6 mo (Fig. 2, a–c; and Fig. S2 a). To test whether Msi2 deletion was maintained in these long-term HSC transplants, we performed qRT-PCR for Msi2 in sorted CD45.2+LSK+ at various time points. Consistent with a role for Msi2 in maintaining the HSC compartment, we found that the residual chimerism in the LSKs (Fig. 2, b and c; and Fig. S2 a) represented cells that were unexcised for Msi2 and maintained comparable Msi2 expression with the control (Fig. 2 d). To further test the requirement for HSC engraftment, we repeated these experiments with sorted phenotypic HSC populations (LSK+CD150+CD48−CD34−) and support congenic BM (300k). We were unable to detect any chimerism of Msi2-deleted cells after 10 wk (Fig. 2 e). These data further suggest that Msi2 was required for the engraftment of HSCs.
Msi2 deficiency decreases repopulating activity. (a) 106 BM cells from mice treated with pIpC were transplanted, and these transplanted animals were sacrificed at 22–24 wk after pIpC with the indicated organ weights (n = 9 from two independent transplants). (b) Representative CD45.2 chimerism flow plot for the HSCs, LSK CD48−CD150+; mean and SEM shown next to gate and also in c. (c) CD45.2 chimerism analysis from HSCs, LSK+CD48−CD150+. See Materials and methods for the surface phenotype of the progenitors (n = 8, 9, same experiment as in panel a). (d) qRT-PCR expression of Msi2 normalized to Gapdh in sorted LSKs from the indicated primary mice or transplanted mice (c) and analyzed at various time points after pIpC (primary mice 4 wk [before transplant] n = 4 per genotype; 12 wk [before transplant], n = 3 per genotype; and 16–24 wk [transplanted mice], n = 9 or n = 8 for Msi2f/f or Msi2Δ/Δ, respectively). (e) BM cells were sorted for HSCs, LSK CD34−CD48−CD150+, and 3,000 HSCs were transplanted into lethally irradiated mice together with 300,000 BM support cells. Peripheral blood was then examined at 10 wk after engraftment (n = 4 mice per genotype). (f) Congenic mice (CD45.1) were noncompetitively transplanted with control or Msi2flox/flox marrow and were allowed to engraft normally (6 wk); peripheral blood chimerism was analyzed before injection of pIpC. (g) Chimerism analysis 18 wk after pIpC from mice in f (see Fig. S2 a [control, n = 10; or Msi2Δ/Δ, n = 11 from three independent transplants] or Fig. S2 b for gating; LT-HSC: LSK+CD34−FLK2/3−, ST-HSC: LSK+CD34+FLK2/3−, LMPP: LSK+CD34+FLK2/3+, representative of control, n = 3; or Msi2Δ/Δ, n = 5 mice). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
Msi2 deficiency decreases repopulating activity. (a) 106 BM cells from mice treated with pIpC were transplanted, and these transplanted animals were sacrificed at 22–24 wk after pIpC with the indicated organ weights (n = 9 from two independent transplants). (b) Representative CD45.2 chimerism flow plot for the HSCs, LSK CD48−CD150+; mean and SEM shown next to gate and also in c. (c) CD45.2 chimerism analysis from HSCs, LSK+CD48−CD150+. See Materials and methods for the surface phenotype of the progenitors (n = 8, 9, same experiment as in panel a). (d) qRT-PCR expression of Msi2 normalized to Gapdh in sorted LSKs from the indicated primary mice or transplanted mice (c) and analyzed at various time points after pIpC (primary mice 4 wk [before transplant] n = 4 per genotype; 12 wk [before transplant], n = 3 per genotype; and 16–24 wk [transplanted mice], n = 9 or n = 8 for Msi2f/f or Msi2Δ/Δ, respectively). (e) BM cells were sorted for HSCs, LSK CD34−CD48−CD150+, and 3,000 HSCs were transplanted into lethally irradiated mice together with 300,000 BM support cells. Peripheral blood was then examined at 10 wk after engraftment (n = 4 mice per genotype). (f) Congenic mice (CD45.1) were noncompetitively transplanted with control or Msi2flox/flox marrow and were allowed to engraft normally (6 wk); peripheral blood chimerism was analyzed before injection of pIpC. (g) Chimerism analysis 18 wk after pIpC from mice in f (see Fig. S2 a [control, n = 10; or Msi2Δ/Δ, n = 11 from three independent transplants] or Fig. S2 b for gating; LT-HSC: LSK+CD34−FLK2/3−, ST-HSC: LSK+CD34+FLK2/3−, LMPP: LSK+CD34+FLK2/3+, representative of control, n = 3; or Msi2Δ/Δ, n = 5 mice). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
To further delineate whether this phenotype is cell autonomous, we noncompetitively transplanted cells from both control and Msi2flox/flox::Mx1-Cre BM, which were then engrafted normally into congenic recipients (Fig. 2 f). After engraftment, Msi2 deletion was induced by pIpC injection, and the peripheral blood was assessed 6 wk later. We observed a reduction in the overall white blood cell count of mice engrafted with Msi2-deficient HSCs, including reductions in myeloid, lymphoid, and red blood cell numbers (not depicted). Similar to the transplants performed with Msi2Δ/Δ BM cells, the chimerism of the residual host BM outcompeted the donor HSCs and progenitors (Fig. 2 g and Fig. S2 b). Collectively, these results indicate that Msi2 is critical for HSC engraftment.
Consistent with a loss of stem cell homeostasis, we found that Msi2Δ/Δ mice had a reduction in the G0 quiescent population in both LSKs and HSCs (Fig. 3 a and Fig. S3 a). Quiescent cells within the HSC compartment have been shown to have higher engraftment potential in transplant assays (Passegué et al., 2005). To test whether the loss of Msi2 would sensitize the cells to proliferative stress, we challenged mice with 5-Fluorouracil (5-FU). These experiments were performed in a cell-autonomous context in which unexcised BM was engrafted, injected with pIpC, and then analyzed at various time points after 6 wk. We noted a dramatic reduction of chimerism in the peripheral blood in the myeloid compartment and recovery within the monocytes in Msi2Δ/Δ compared with controls (Fig. 3 b). This reduction in chimerism was also observed in the phenotypic HSCs (LSK+CD150+CD48−) in 5-FU–treated mice (Fig. 3 c and Fig. S3 b). Altogether, loss of Msi2 impairs HSCs during normal homeostasis, which is accentuated in response to perturbations including transplantation and replicative stress.
Msi2 deficiency reduces quiescence, and proliferative stress further depletes HSCs. (a) Representative flow cytometry cell cycle plots with statistical analysis (bottom; HSC: LSK CD34−, HSPC: LSK CD34+; control, n = 10; Msi2Δ/Δ, n = 7 from three independent experiments). See Fig. S3 a for gating. (b) The indicated mice were transplanted noncompetitively, injected with pIpC, and after 6 wk were injected with 5-FU. Peripheral blood analysis was performed with measurements of monocytes at the indicated time points and were normalized back to the same animal before 5-FU (n = 5, 8) and 6 wk after pIpC injections. (c) BM donor chimerism of HSCs (LSK CD48−CD150+) of mice (n = 5, 8) was assessed at 9 wk after pIpC and from b 40 d after 5-FU and non–5-FU mice (control, n = 8; and Msi2Δ/Δ, n = 10 from three independent experiments). See Fig. S3 b for gating. Mean and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
Msi2 deficiency reduces quiescence, and proliferative stress further depletes HSCs. (a) Representative flow cytometry cell cycle plots with statistical analysis (bottom; HSC: LSK CD34−, HSPC: LSK CD34+; control, n = 10; Msi2Δ/Δ, n = 7 from three independent experiments). See Fig. S3 a for gating. (b) The indicated mice were transplanted noncompetitively, injected with pIpC, and after 6 wk were injected with 5-FU. Peripheral blood analysis was performed with measurements of monocytes at the indicated time points and were normalized back to the same animal before 5-FU (n = 5, 8) and 6 wk after pIpC injections. (c) BM donor chimerism of HSCs (LSK CD48−CD150+) of mice (n = 5, 8) was assessed at 9 wk after pIpC and from b 40 d after 5-FU and non–5-FU mice (control, n = 8; and Msi2Δ/Δ, n = 10 from three independent experiments). See Fig. S3 b for gating. Mean and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
Msi2 determines myeloid lineage bias
To determine whether lineage fate decisions were also altered, we reexamined all of our transplant experiments for differential chimerism between myeloid and lymphoid populations. Interestingly, we observed that the defect was more severe in the myeloid lineage compared with the lymphoid lineage (Fig. 4, a–c). This prompted us to further define the HSC compartment and assess the heterogeneous phenotypic cell types that can display lineage bias toward the myeloid or lymphoid fate (Uchida et al., 2003; Dykstra et al., 2007; Chambers et al., 2008; Beerman et al., 2010; Pang et al., 2011). We did not observe a reduction in common lymphoid progenitors (CLPs; Fig. 4 d and Fig. S4 a). Although there was no difference in the frequency of HSCs among the LSKs (Fig. S1), we found a three- to fourfold reduction in the frequency of myeloid-biased HSCs (My-HSCs; LSK+CD34−CD48−CD150hi or LSK+CD34−CD48−CD150hiCD86lo; Fig. 4, e and f; and Fig. S4 b; Shimazu et al., 2012).
Msi2Δ/Δ cells have reduced myeloid lineage–biased HSCs and defective myeloid repopulating activity. (a) Chimerism analysis of the myeloid and lymphoid populations in the BM and spleen of mice 22–24 wk after transplant (same as in Fig. 2 [a and b]). (b) Chimerism of myeloid and lymphoid lineages in spleen of primary transplanted mice shown 12–18 wk after pIpC (n = 9, 10 from two independent experiments). (c) Chimerism analysis 9 wk after pIpC and 40 d after 5-FU injections (same as in Fig. 3 [b and c]). (d) Chimerism analysis at 18 wk after pIpC, and CLP (Linloc-kitmidSca+Flk2+IL7R+, CLP2: Linloc-kitloSca+Flk2+IL7R+) populations are based on gating in Fig. S4 a (control, n = 3; or Msi2Δ/Δ, n = 5 mice). (e) Representative flow analysis (left; with indicated staining from 18-wk-post-pIpC mice). Mean and SEM for the frequency of CD150hi My-HSCs gated from (left flow plot) 4 wk (n = 12 and n = 15 from five independent experiments) and 18 wk (n = 5 and Msi2Δ/Δ n = 8 from two independent experiments). (f) Flow cytometric analysis of My-HSCs: LinloCD45.2+LSK CD150hiCD86− (n = 6, n = 8 mice from two independent experiments). See gating strategy in Fig. S4 b. SEM is shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
Msi2Δ/Δ cells have reduced myeloid lineage–biased HSCs and defective myeloid repopulating activity. (a) Chimerism analysis of the myeloid and lymphoid populations in the BM and spleen of mice 22–24 wk after transplant (same as in Fig. 2 [a and b]). (b) Chimerism of myeloid and lymphoid lineages in spleen of primary transplanted mice shown 12–18 wk after pIpC (n = 9, 10 from two independent experiments). (c) Chimerism analysis 9 wk after pIpC and 40 d after 5-FU injections (same as in Fig. 3 [b and c]). (d) Chimerism analysis at 18 wk after pIpC, and CLP (Linloc-kitmidSca+Flk2+IL7R+, CLP2: Linloc-kitloSca+Flk2+IL7R+) populations are based on gating in Fig. S4 a (control, n = 3; or Msi2Δ/Δ, n = 5 mice). (e) Representative flow analysis (left; with indicated staining from 18-wk-post-pIpC mice). Mean and SEM for the frequency of CD150hi My-HSCs gated from (left flow plot) 4 wk (n = 12 and n = 15 from five independent experiments) and 18 wk (n = 5 and Msi2Δ/Δ n = 8 from two independent experiments). (f) Flow cytometric analysis of My-HSCs: LinloCD45.2+LSK CD150hiCD86− (n = 6, n = 8 mice from two independent experiments). See gating strategy in Fig. S4 b. SEM is shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.
Msi2Δ/Δ HSCs have increased commitment toward differentiation
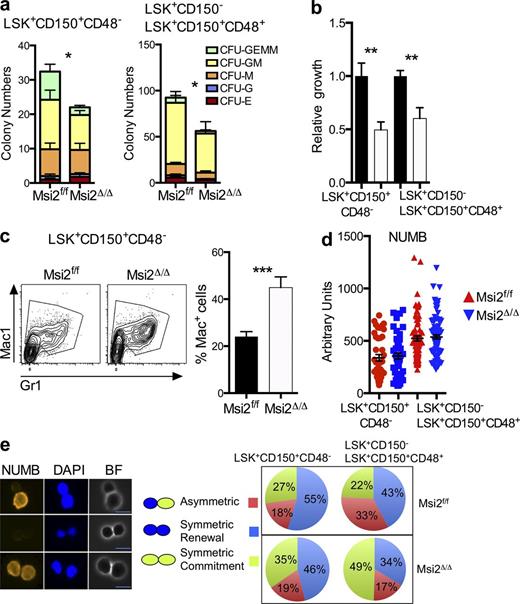
The reduced quiescence and increased sensitivity to stress prompted us to examine whether Msi2Δ/Δ HSCs were defective in their ability to undergo lineage commitment at the single cell level. Myeloid colony–forming assays with sorted phenotypic HSCs (LSK+CD150+CD48−) or HSPCs (LSK+CD150− and LSK+CD150+CD48+; Fig. S5) revealed decreased colony formation with significant reductions in multilineage myeloid differentiation (Fig. 5 a). Proliferation assays in the presence of myeloid cytokines on sorted HSCs and HSPCs resulted in reduced total cellular output (Fig. 5 b). Concurrently, we observed an increase in the frequency of cells committed to differentiation based on the expression of myeloid markers (Fig. 5 c). Collectively, Msi2 contributes to stem cell fate decisions based on the reduced cellular output, decreased colony formation, and increased cellular commitment.
HSCs and HSPCs from Msi2-deficient mice have reduced self-renewal divisions with differentiation independent of the Numb. (a) The indicated sorted populations were plated in myeloid methylcellulose colony assays (left, HSC: LSK+CD150+CD48−; and right, HSPC: LSK+CD150+CD48+ and LSK CD150−; means and SEM from n = 6 control and n = 5 Msi2Δ/Δ from two independent experiments; *, P < 0.05 for CFU-GM, CFU-G, and CFU-GEMM). G, granulocyte, GM, mixed granulocyte and monocyte colony; BFU-E, burst-forming units erythroid lineage; GEMM, mixed granulocyte, erythroid monocyte, and megakaryocyte; MEG, megakaryocyte. (b) Indicated sorted populations from Msi2Δ/Δ donors (6–9 wk; HSC mice: n = 14–15 per genotype from six independent experiments; HSPC mice: n = 5 from two independent experiments) were grown for 5 d, and cell numbers were divided by the number of cells in the control wells (see Fig. S5 for gating). (c) Representative flow for mature myeloid markers from HSCs grown same as in b and percentage of Mac-1–positive cells (histogram; control, n = 15; and Msi2Δ/Δ, n = 13 HSCs). (d) Sorted control and Msi2Δ/Δ HSCs and HSPCs from mice 4 wk after pIpC treatment cultured in vitro for 36 h and stained for expression of Numb and DAPI (four to five mice from two independent experiments; HSCs, n = 42 and 46 cells; HSPCs, n = 76 and 94). (e) Representative images for paired Numb daughter assay with DAPI (middle) and brightfield images (left) and experimental scheme (right) for studying asymmetric division in HSCs/HSPCs. Percentage of doublet cells in each type of cell division is shown (n = 4–5 individual mice from two independent experiments; total HSCs daughter pairs: 22 and 72, HSPC pairs: 84 and 107). Bars, 10 µm. Mean and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
HSCs and HSPCs from Msi2-deficient mice have reduced self-renewal divisions with differentiation independent of the Numb. (a) The indicated sorted populations were plated in myeloid methylcellulose colony assays (left, HSC: LSK+CD150+CD48−; and right, HSPC: LSK+CD150+CD48+ and LSK CD150−; means and SEM from n = 6 control and n = 5 Msi2Δ/Δ from two independent experiments; *, P < 0.05 for CFU-GM, CFU-G, and CFU-GEMM). G, granulocyte, GM, mixed granulocyte and monocyte colony; BFU-E, burst-forming units erythroid lineage; GEMM, mixed granulocyte, erythroid monocyte, and megakaryocyte; MEG, megakaryocyte. (b) Indicated sorted populations from Msi2Δ/Δ donors (6–9 wk; HSC mice: n = 14–15 per genotype from six independent experiments; HSPC mice: n = 5 from two independent experiments) were grown for 5 d, and cell numbers were divided by the number of cells in the control wells (see Fig. S5 for gating). (c) Representative flow for mature myeloid markers from HSCs grown same as in b and percentage of Mac-1–positive cells (histogram; control, n = 15; and Msi2Δ/Δ, n = 13 HSCs). (d) Sorted control and Msi2Δ/Δ HSCs and HSPCs from mice 4 wk after pIpC treatment cultured in vitro for 36 h and stained for expression of Numb and DAPI (four to five mice from two independent experiments; HSCs, n = 42 and 46 cells; HSPCs, n = 76 and 94). (e) Representative images for paired Numb daughter assay with DAPI (middle) and brightfield images (left) and experimental scheme (right) for studying asymmetric division in HSCs/HSPCs. Percentage of doublet cells in each type of cell division is shown (n = 4–5 individual mice from two independent experiments; total HSCs daughter pairs: 22 and 72, HSPC pairs: 84 and 107). Bars, 10 µm. Mean and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001).
To determine whether the observed defect in Msi2Δ/Δ HSCs could be caused by increased commitment divisions, we examined Numb protein levels and distribution in mitotic HSC daughter pairs. Numb can be used as a surrogate readout for asymmetric stem cell division and as a putative direct target of Msi2 mRNA binding in HSCs (Wu et al., 2007; Kharas et al., 2010). Surprisingly, we observed equivalent Numb protein levels in Msi2Δ/Δ HSCs or HSPCs compared with the control cells. This result indicates that Msi2 deficiency does not alter the global protein levels of Numb within an individual cell (Fig. 5 d). However, we did observe a decrease in the percentage of cells that underwent asymmetric Numb segregation in the HSPCs and an increase in the percentage of cells with symmetric commitment Numb staining in both the HSCs and HSPCs (Fig. 5 e). These results suggested that loss of Msi2 skewed stem cell fate decisions in favor of commitment divisions.
MSI2’s direct RNA binding targets control translation and the self-renewal program in HSCs
We decided to take a global approach to understanding how Msi2 loss alters the cellular program of stem cells. We first examined the pathways that were altered in the Msi2Δ/Δ LSK by performing transcriptome profiling at 4 wk after Msi2 deletion. We found 311 differentially expressed genes, of which 208 were down-regulated and 103 were up-regulated (fold change > 1.5; feature P < 0.05; Table S1). To easily visualize the top differentially expressed genes, we generated a heat map of the top 40 down-regulated and up-regulated genes (ranked by t-statistic; Fig. 6, a and b). We then functionally annotated our expression profiling and performed Gene Set Enrichment Analysis (GSEA; Subramanian et al., 2005) on all curated gene sets in the Molecular Signatures Database (MSigDB; 4,850 gene sets) combined with an additional set of relevant gene sets (90 gene sets from our experimentally derived or published hematopoietic self-renewal and differentiation signatures [Subramanian et al., 2005]; see Table S2). We found 13 gene sets (false discovery rate [FDR] < 0.01 and >20 genes) that were enriched in the control LSKs and 92 gene sets enriched in the Msi2Δ/Δ LSKs (Table S3). We found that the Msi2Δ/Δ LSKs have down-regulated genes that are normally expressed in the self-renewal signature of HSCs. Also, the Msi2Δ/Δ LSKs have up-regulated genes associated with differentiation to megakaryocyte erythroid progenitor (MEPs), common myeloid progenitors (CMPs), and granulocyte monocyte progenitors (GMPs; Fig. 6 c and Table S3). These data suggest that Msi2 maintains the stem cell gene expression program. We decided to examine the expression of a set of lineage transcription factors in a more phenotypically purified HSC population (LSK+CD150+CD48−CD34−). We found reductions in Egr1, Gfi1, and Junb expression with a modest, but significant increase in c-Myc transcript (Fig. 6 d). Consistent with the increase in the mRNA level, we also found increased c-MYC protein within the phenotypic HSCs (LSK+CD150+CD48−; Fig. 6 e). These data further supported the role for Msi2 in maintaining the self-renewal program and an increase in the commitment in the HSCs.
Loss of self-renewal gene expression program and increased differentiation in Msi2-deficient HSPCs. (a and b) Microarray heat maps of differentially expressed genes with the top 40 down-regulated (a) and top 40 up-regulated genes (b) shown and also ranked based on Student’s t test values (n = 4 per genotype; 4 wk after pIpC) of sorted LSKs. (c) Self-renewal and differentiation gene sets analyzed from the transcriptome analysis comparing control and Msi2Δ/Δ LSKs (same as in a and b). Enrichment score (ES) is shown. (d) qRT-PCR analysis of the indicated hematopoietic self-renewal and lineage transcription factors in sorted LSK+CD150+CD48−CD34− cells from Msi2Δ/Δ and Msi2f/f (n = 3 per genotype). (e) Sorted LSK+CD150+CD48− cells were stained with c-MYC–specific Ab (left) and DAPI (middle); a merged image is shown (right). Bars, 10 µm. Fluorescent intensity was quantified and normalized to the control (three independent experiments; 171 and 248 cells quantified per group in total). Means and SEM are shown (*, P < 0.05; ****, P = 3.56 × 10−7).
Loss of self-renewal gene expression program and increased differentiation in Msi2-deficient HSPCs. (a and b) Microarray heat maps of differentially expressed genes with the top 40 down-regulated (a) and top 40 up-regulated genes (b) shown and also ranked based on Student’s t test values (n = 4 per genotype; 4 wk after pIpC) of sorted LSKs. (c) Self-renewal and differentiation gene sets analyzed from the transcriptome analysis comparing control and Msi2Δ/Δ LSKs (same as in a and b). Enrichment score (ES) is shown. (d) qRT-PCR analysis of the indicated hematopoietic self-renewal and lineage transcription factors in sorted LSK+CD150+CD48−CD34− cells from Msi2Δ/Δ and Msi2f/f (n = 3 per genotype). (e) Sorted LSK+CD150+CD48− cells were stained with c-MYC–specific Ab (left) and DAPI (middle); a merged image is shown (right). Bars, 10 µm. Fluorescent intensity was quantified and normalized to the control (three independent experiments; 171 and 248 cells quantified per group in total). Means and SEM are shown (*, P < 0.05; ****, P = 3.56 × 10−7).
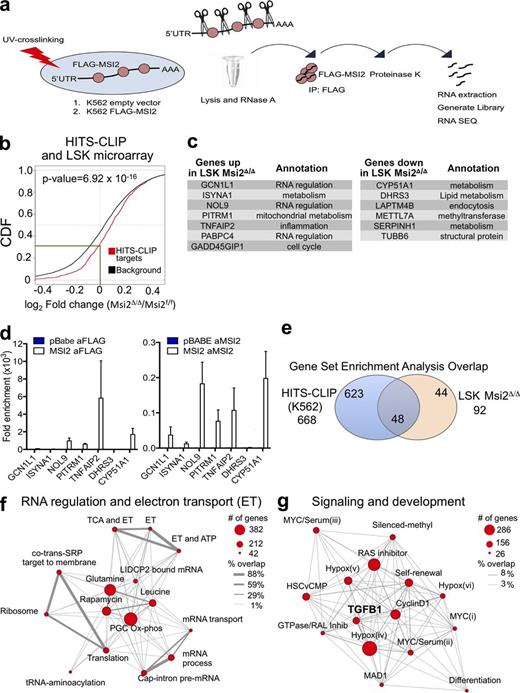
Although we observed significant alterations in multiple pathways in LSKs, it remained unclear whether this was a result of a direct mechanism of action. Therefore, we decided to globally assess direct MSI2-binding targets by UV cross-linking and immunoprecipitation of MSI2, followed by RNase digestion of unbound RNA and RNA sequencing of the resulting protected fragments (HITS-CLIP; Fig. 7 a). Because of the technical challenge of the HITS-CLIP protocol, which requires large amounts of starting material (i.e., 100 million cells) to capture specific interactions between endogenous proteins and target RNA, we overexpressed a FLAG-tagged version of MSI2 in parallel to a control vector lacking the MSI2 cDNA in K562 leukemia cells (Fig. 7 a). After performing HITS-CLIP, sequencing read alignment, and peak calling using our previously published algorithm (Loeb et al., 2012), we tested CLIP peaks for significant differential MSI2 binding over control and normalized peak expression by differential expression of all genes by RNA sequencing, using a joint statistical model of HITS-CLIP and RNA sequencing read count data. We found 1,097 unique targets that have at least one significant MSI2-binding site (FDR < 0.1 with a corrected CLIP log fold change of two or more). We observed binding that was distributed between the coding DNA sequence (CDS) and the UTRs (56% and 44%, respectively; Table S4). To determine whether gene expression changes in the LSK microarray after Msi2 deletion were enriched for direct Msi2 mRNA–binding HITS-CLIP targets, we performed a cumulative distribution function and found a statistically significant difference in HITS-CLIP targets compared with the background gene expression (genes expressed in LSKs but are not HITS-CLIP targets) that was expressed in the LSKs (right shift; p-value = 6.92 × 10−16; Fig. 7 b). Furthermore, ∼70% of the HITS-CLIP target genes were increased in expression after Msi2 deletion in the LSKs, and of these, ∼30% demonstrated a reduction of expression in the Msi2Δ/Δ LSKs compared with the control (Fig. 7 b). Moreover, we identified seven up-regulated genes and six down-regulated genes that were both differentially expressed in the LSKs and were HITS-CLIP targets (Fig. 7 c). These genes represented a wide variety of pathways, including RNA regulators, cell cycle, and various metabolic pathways. We then reexamined a subset of these genes in K562 cells by qRT-PCR. To validate our approach, we performed an additional RNA immunoprecipitation (RIP) experiment with antibodies (Abs) that target either endogenous MSI2 or the flag-tagged version of MSI2 (Flag-MSI2; Fig. 7 d). We were able to detect enrichment in the RIP experiments using the MSI2-targeting Abs compared with control cells that expressed the pBabe empty vector or control Ab (anti–rabbit IgG) in six out of the seven targets that were retested, suggesting that our HITS-CLIP and mRNA overlap was reproducible (Fig. 7 d).
MSI2’s direct RNA targets are involved in RNA metabolism and the self-renewal signaling program. (a) Schematic outline of the HITS-CLIP approach. (b) Target genes are mouse genes homologous to human MSI2 targets identified by HITS-CLIP in K562 (FDR < 0.1) and background genes are all the other genes expressed in mouse LSK cells. Log2 fold changes (logFCs) of gene expression between knockout and wild-type samples are fitted using limma package. When there are multiple microarray probes mapped to the same gene, logFC of that gene is represented by the median of all corresponding probes. P-value is obtained with one-sided K-S test on the logFC distributions of target and background genes (green line indicates the intersection of the HITS-CLIP targets with a zero logFC). CDF, cumulative distribution function. (c) Up- and down-regulated genes that are differentially expressed and overlap with the top HITS-CLIP targets. (d) Validation using qRT-PCR of a subset of the HITS-CLIP targets from c with RIP performed with anti-FLAG Ab and Ab specific for MSI2 in K562 cells (mean of two independent RIP experiments and fold change over control samples; pBabe cells were used as control for Flag Ab sample, and anti–rabbit Ab was used as control for Msi2 Ab). SEM is shown. (e) Venn diagram indicating the number of statistically significant gene sets from the HITS-CLIP or the transcriptome analysis comparing controls and Msi2Δ/Δ LSKs (FDR < 0.01). The number in the middle indicates the overlapping significant gene sets that are enriched in both the HITS-CLIP and the transcriptome analysis of LSKs. (f and g) Modules indicate the relationships between manually curated and selected gene sets (within the overlap in e and Table S6). “% overlap” indicates the number of genes that are common between the genes set.
MSI2’s direct RNA targets are involved in RNA metabolism and the self-renewal signaling program. (a) Schematic outline of the HITS-CLIP approach. (b) Target genes are mouse genes homologous to human MSI2 targets identified by HITS-CLIP in K562 (FDR < 0.1) and background genes are all the other genes expressed in mouse LSK cells. Log2 fold changes (logFCs) of gene expression between knockout and wild-type samples are fitted using limma package. When there are multiple microarray probes mapped to the same gene, logFC of that gene is represented by the median of all corresponding probes. P-value is obtained with one-sided K-S test on the logFC distributions of target and background genes (green line indicates the intersection of the HITS-CLIP targets with a zero logFC). CDF, cumulative distribution function. (c) Up- and down-regulated genes that are differentially expressed and overlap with the top HITS-CLIP targets. (d) Validation using qRT-PCR of a subset of the HITS-CLIP targets from c with RIP performed with anti-FLAG Ab and Ab specific for MSI2 in K562 cells (mean of two independent RIP experiments and fold change over control samples; pBabe cells were used as control for Flag Ab sample, and anti–rabbit Ab was used as control for Msi2 Ab). SEM is shown. (e) Venn diagram indicating the number of statistically significant gene sets from the HITS-CLIP or the transcriptome analysis comparing controls and Msi2Δ/Δ LSKs (FDR < 0.01). The number in the middle indicates the overlapping significant gene sets that are enriched in both the HITS-CLIP and the transcriptome analysis of LSKs. (f and g) Modules indicate the relationships between manually curated and selected gene sets (within the overlap in e and Table S6). “% overlap” indicates the number of genes that are common between the genes set.
We then again queried the MSigDB signatures with GSEA (Subramanian et al., 2005) using the full list of ranked (by fold change) RNA CLIP targets to understand the functional classification of MSI2’s direct RNA targets. MSI2 binding was positively enriched for 668 gene sets (FDR < 0.01; Table S5), whereas the nonspecific RNA interactions (low or negative fold change) failed to identify any significant pathways. These data suggest that MSI2 RNA binding generates a defined cellular program that may not be simply attributed to alterations of individual targets in a pathway. Similar to examining the individual genes in LSKs (Fig. 7 b), this large number of gene sets prompted us to further filter and overlap the molecular signatures with our data from the LSK microarray. Thus, we identified 52% (48 out of 92) gene sets enriched as MSI2-binding targets that were also enriched in the transcriptome data analysis from Msi2Δ/Δ LSKs (Fig. 7 e), providing a union between MSI2 RNA binding activity and subsequent transcriptional regulation of these pathways (Table S6). Interestingly, the gene sets were unidirectional, demonstrating enrichment only in the Msi2Δ/Δ LSKs and not in the control cells, implying that the majority of Msi2’s targets are being repressed. Of the 48 gene sets, we narrowed down the gene sets that were associated with hematopoietic biology. We then examined the genes sets and categorized them into two defined modules: “RNA regulation and electron transport” and “Signaling and development” gene sets (Fig. 7, f and g; and Table S6). We displayed the size of the gene sets (circle diameter) and the overlap of the individual genes between the identified gene sets (connecting line thickness) to visualize the relationship of these functional pathways. For example, within RNA regulation and electron transport, genes sets associated with genes that are normally down-regulated after mTOR inhibition (i.e., rapamycin, leucine, or glutamine deprivation) were both enriched for MSI2 binding and up-regulated in Msi2Δ/Δ LSKs. This central node connected three distinct peripheral nodes that included energy metabolism, mRNA processing, and translation (Fig. 7 f). In the signaling and development module, a variety of pathways including a developmental program of “HSC versus CMP” and “Self-renewal” and other signaling pathways including, Hypoxia, RAS, MYC, and CYCLIND1, among others, were detected, and the overlap among the genes within the cluster was 8% or less (Fig. 7 g). Interestingly, the Self-renewal and HSC versus CMP gene sets (already discussed in Fig. 6) further overlapped with HITS-CLIP targets. We also observed four Myc-related gene sets that overlapped with HITS-CLIP targets, which also contained 8% or fewer of the individual genes overlapping. Additionally, we observed that the TGFB1 pathway also overlapped with our HITS-CLIP targets (Fig. 7 g). Altogether, these modules indicate a complex role for MSI2 in regulating multiple critical cellular processes and pathways.
MSI2 modulates the TGF-β signaling pathway
Although the role for Msi2 in HSCs is multidimensional and complex, we decided to focus on the interaction between MSI2 and the TGF-β pathway for multiple reasons. Previous studies have demonstrated that TGF-β signaling is critical for HSC quiescence, cell cycle, and differentiation (Sitnicka et al., 1996; Kale and Vaidya, 2004; Kale, 2004; Yamazaki et al., 2011), and Msi2Δ/Δ LSKs had significantly down-regulated Tgfbr1 (Fig. 6 a). Moreover, the novel relationship between MSI2 and the TGF-β pathway is of interest as it came up in our HITS-CLIP and microarray data. Therefore, our combined genomic experiments coupled with our functional experiments prompted us to further examine the link between MSI2 and TGF-β signaling. First, we wanted to test whether Tgfbr1 expression levels were also down-regulated when we examined more phenotypically pure HSCs (LSK+CD150+CD48−CD34−). Consistent with our microarray results within the LSKs, we observed a reduction at the level of the mRNA expression in Msi2Δ/Δ HSCs (Fig. 8 a). However, when we further tested whether this resulted in the reduction in protein expression, we observed the opposite result with a significant increase in TGFBR1 in sorted LSK+CD150+CD48− cells (Fig. 8 b). This prompted us to further test the output of the TGF-β pathway in LSK+CD150+CD48− cells in the presence and absence of Msi2. An established readout for TGF-β signaling is the phosphorylation of downstream effectors, including Smad2/3. In contrast to the increase in receptor expression, we found reduced phosphorylation of Smad2/Smad3 in Msi2Δ/Δ HSCs (LSK+CD150+CD48−) directly isolated from BM or grown in vitro (Fig. 8, c and d). Another readout for TGF-β signaling in HSCs is the expression of one of its direct target genes, p57 (Scandura et al., 2004; Brenet et al., 2013). Recently, it has been reported that the loss of p57 in HSCs results in reduced quiescence and compensatory up-regulation of p27 (Cheng et al., 2001; Matsumoto et al., 2011; Zou et al., 2011). Consistent with reduced signaling output of TGF-β, we observed a significant decrease in p57 expression and up-regulation of p27 in Msi2Δ/Δ LSKs (Fig. 8 e).
Msi2 regulates TGF-β signaling. (a) mRNA expression of Tgfbr1 in LSK+CD150+CD48−CD34− (n = 3 per genotype). (b) Sorted LSK+CD150+CD48− cells from the indicated mice were stained with a TGFBR1-specific Ab (left) and DAPI (middle); a merged image is shown (right). Fluorescence was quantified and normalized to the control; the result is from three independent experiments; 292 and 175 cells were quantified per group in total. (c) Sorted LSK+CD150+CD48− cells were stained with pSmad2/3-specific Ab (representative image from freshly isolated cells, left) and DAPI (middle); a merged image is shown (right). Immunofluorescence was quantified and normalized to the control; results are from two independent experiments; 465 and 540 cells were quantified per group in total. (d) LSK+CD150+CD48− cells were grown in culture for 2 d. 38–40 cells were quantified per group in total from three independent experiments. (e) qRT-PCR of cell cycle regulators and p57 target gene of TGF-β signaling from control and Msi2Δ/Δ LSKs (n = 5, two independent experiments). (f) LSK+CD150+CD48− sorted populations from control and Msi2Δ/Δ donors (6–9 wk; HSC mice: n = 3–8 mice from the control for each condition and 3–10 for the Msi2Δ/Δ from up to six independent experiments) were grown (50 cells per well were plated initially for 7 d), and absolute cell numbers are shown. (g) qRT-PCR of Smad3 from LSK+CD150+CD48−CD34− cells (n = 3 per genotype). (h) BM cells were sorted for LSK+CD150+CD48− and stained with a specific Ab for SMAD3 (fluorescence was quantified and normalized to the control; four independent experiments; 531 control and 379 Msi2Δ/Δ cells quantified per group in total). (b, c, and h) Bars, 10 µm. (i) qRT-PCR of TGFBR1 and SMAD3 HITS-CLIP targets with RIP performed as in Fig. 7 d with FLAG and a rabbit Ab specific for MSI2 in K562 cells (mean of two independent RIP experiments and fold change over control samples and SEM). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ****, P = 1.17 × 10−41).
Msi2 regulates TGF-β signaling. (a) mRNA expression of Tgfbr1 in LSK+CD150+CD48−CD34− (n = 3 per genotype). (b) Sorted LSK+CD150+CD48− cells from the indicated mice were stained with a TGFBR1-specific Ab (left) and DAPI (middle); a merged image is shown (right). Fluorescence was quantified and normalized to the control; the result is from three independent experiments; 292 and 175 cells were quantified per group in total. (c) Sorted LSK+CD150+CD48− cells were stained with pSmad2/3-specific Ab (representative image from freshly isolated cells, left) and DAPI (middle); a merged image is shown (right). Immunofluorescence was quantified and normalized to the control; results are from two independent experiments; 465 and 540 cells were quantified per group in total. (d) LSK+CD150+CD48− cells were grown in culture for 2 d. 38–40 cells were quantified per group in total from three independent experiments. (e) qRT-PCR of cell cycle regulators and p57 target gene of TGF-β signaling from control and Msi2Δ/Δ LSKs (n = 5, two independent experiments). (f) LSK+CD150+CD48− sorted populations from control and Msi2Δ/Δ donors (6–9 wk; HSC mice: n = 3–8 mice from the control for each condition and 3–10 for the Msi2Δ/Δ from up to six independent experiments) were grown (50 cells per well were plated initially for 7 d), and absolute cell numbers are shown. (g) qRT-PCR of Smad3 from LSK+CD150+CD48−CD34− cells (n = 3 per genotype). (h) BM cells were sorted for LSK+CD150+CD48− and stained with a specific Ab for SMAD3 (fluorescence was quantified and normalized to the control; four independent experiments; 531 control and 379 Msi2Δ/Δ cells quantified per group in total). (b, c, and h) Bars, 10 µm. (i) qRT-PCR of TGFBR1 and SMAD3 HITS-CLIP targets with RIP performed as in Fig. 7 d with FLAG and a rabbit Ab specific for MSI2 in K562 cells (mean of two independent RIP experiments and fold change over control samples and SEM). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ****, P = 1.17 × 10−41).
These results further suggest a defect in the output of the TGF-β signaling pathway in vivo. The effect of TGF-β treatment is biphasic in HSCs, with high levels of TGF-β blocking proliferation and low levels of this pathway activation leading to increased proliferation (Kale and Vaidya, 2004; Kale, 2004). We therefore examined the functional response of Msi2Δ/Δ HSCs to various levels of TGF-β signaling in vitro. The reduced TGF-β signaling observed in steady-state Msi2Δ/Δ HSCs was overcome by the addition of high doses of exogenous TGF-β, resulting in potent growth suppression regardless of Msi2 status (Fig. 8 f). In contrast, when exposed to a dose (0.01 ng/ml) of TGF-β shown to expand normal HSCs, Msi2Δ/Δ HSCs failed to respond, consistent with diminished sensitivity to the proliferative effects of TGF-β1 (Fig. 8 f). However, this effect was lost at the lower doses of TGF-β, suggesting a narrow window for this effect. Interestingly, a recent study has also identified an increase in proliferation within the My-HSC compartment after stimulation with 0.01 ng/ml TGF-β1 (Challen et al., 2010). Furthermore, our results that indicate reduced output in signaling and proliferation match the marked reduction we observed specifically within the My-HSCs (Fig. 4, e and f).
To further understand how Msi2 loss could negatively control TGF-β output, we tested the mRNA expression of Smad3 and found equivalent levels in the LSK+CD150+CD48−CD34− (Fig. 8 g). In contrast to the TGFBR1 regulation and consistent with the phosphorylation of SMAD2/3, we found a reduction in the total SMAD3 abundance in the LSK+CD150+CD48− (Fig. 8 h). Although TGFBR1 and SMAD3 were not in the list of the top HITS-CLIP targets (FDR < 0.1), we did detect direct Msi2 binding in our HITS-CLIP (TGFBR1: fold change of 12.78 compared with control, p-value = 0.02, FDR = 0.12; and Smad3: fold change of 3.2, p-value = 0.02, FDR = 0.11; Table S6). We then validated this binding with an additional RIP for TGFBR1 and SMAD3 (Fig. 8 i). Altogether, these data suggested that Msi2 was directly modulating the TGF-β pathway at multiple nodes.
DISCUSSION
In this study, we examined the role of Msi2 in HSCs using a conditional knockout mouse model. Conditional ablation of Msi2 in the hematopoietic system results in dramatic reductions in steady-state HSC and HSPC numbers, with no changes in maturation of myeloid or lymphoid hematopoietic lineages in primary mice. We demonstrate that Msi2 is required for HSC quiescence and maintaining the balance of self-renewal and symmetric commitment that is required for normal hematopoiesis (Fig. 9). Applying global RNA target analysis of MSI2 binding in leukemic cells combined with transcriptional analysis in mouse HSCs, we identified common self-renewal and differentiation pathways. Additionally, our approach uncovered a novel role for Msi2 in the regulation of TGF-β signaling by demonstrating reduced TGF-β signaling output and impaired response to TGF-β stimulation in Msi2-deficient HSCs.
Model of Msi2 function in HSCs. (left) Myeloid- and lymphoid-biased HSCs undergo balanced symmetric and asymmetric divisions, giving rise to differentiated progeny. Normal My-HSCs proliferate in response to low levels of TGF-β signals. (right) Deletion of Msi2 in HSCs leads to loss of quiescence and increased symmetric commitment toward differentiated progeny. Additionally, Msi2 loss leads to a reduction in My-HSCs, which are unresponsive to TGF-β–induced proliferative signals.
Model of Msi2 function in HSCs. (left) Myeloid- and lymphoid-biased HSCs undergo balanced symmetric and asymmetric divisions, giving rise to differentiated progeny. Normal My-HSCs proliferate in response to low levels of TGF-β signals. (right) Deletion of Msi2 in HSCs leads to loss of quiescence and increased symmetric commitment toward differentiated progeny. Additionally, Msi2 loss leads to a reduction in My-HSCs, which are unresponsive to TGF-β–induced proliferative signals.
Previous studies from our group and others have identified a role for Msi2 in hematopoietic development, but it remained unclear how Msi2 functions and what pathways Msi2 regulates in hematopoietic cells (Hope et al., 2010; Ito et al., 2010; Kharas et al., 2010; de Andrés-Aguayo et al., 2011). Our study is consistent with experiments using shRNAs specific to Msi2 that revealed a block in the engraftment of Msi2-depleted cells (Hope et al., 2010; Kharas et al., 2010). However, our study differs with the analysis of hematopoietic cells from mice harboring an insertional gene trap at the Msi2 locus. Although these mice demonstrated a general reduction in repopulating activity in vivo, they attribute this defect to the LMPP and not the HSC (de Andrés-Aguayo et al., 2011). We find reductions in LSKs and LMPPs, but also find a significant defect in the HSCs after genetic ablation of Msi2. Consistent with the gene trap loss of function Msi2 mice, we also observed a reduction of circulating myeloid cells (de Andrés-Aguayo et al., 2011). We found that this defect initially occurred at the level of the My-HSC. Similarly, the same study suggested that there was a proliferative defect in the LMPP compartment. We were unable to detect reduced cycling cells, but found a failure to maintain quiescence and skewing away from symmetric self-renewal divisions toward differentiation in phenotypic HSCs. Transcriptome analysis in the gene trap mice suggested a loss of Myc target gene expression, whereas we observed the opposite: an increase of Myc target gene expression and increased levels of Myc in the HSC compartment at the mRNA and protein level. The apparent discrepancy in the defect within the HSCs between the two mouse models suggests that the loss of Msi2 may be compensated during development or could be caused by differences between gene trap–mediated insertional disruption and the conditional deletion approach in vivo.
Experiments on the Msi family have considered Numb to be a critical downstream target of Msi mRNA binding and translational suppression (Imai et al., 2001). In contrast to previously published studies connecting Musashi to Numb and to Notch signaling (Imai et al., 2001; Ito et al., 2010; Kharas et al., 2010), we were unable to uncover a role for these pathways in HSPCs. A study of normal CD34+ cells from healthy BM displayed no correlation between MSI2 and NUMB expression, whereas the same study found reduced expression of NUMB in MDS/AML samples with the highest level of MSI2 expression (Pereira et al., 2012). Finally, we are unable to detect binding of NUMB in our HITS-CLIP analysis or increased expression after MSI2-shRNA depletion in K562 cells (unpublished data).
To gain an unbiased understanding of the pathways downstream of Msi2 in maintaining proper HSC function, we used transcriptome-wide MSI2 RNA–binding target analysis. A previous global study examining Msi1 using native RIP (RIP-CHIP) identified targets known to contribute to oncogenic transformation (de Sousa Abreu et al., 2009). In our study, we used a recently developed approach for studying RNA-binding proteins (HITS-CLIP) that involves UV cross-linking to generate more specificity in binding targets through direct interactions between target proteins and RNA (Chi et al., 2009). This allowed us to define a global map of MSI2 direct binding RNA targets for the first time. Moreover, integration of our HITS-CLIP and transcriptome analysis performed in Msi2-deficient HSPCs identified a developmental program that is controlled through many direct RNA–protein interactions.
Based on the phenotypes we observed in self-renewal, quiescence, and our genomic data, we focused on the TGF-β pathway. A previous study in vitro with TGF ligands suggested that TGF-β signaling could block proliferation of HSCs and maintain quiescence (Sitnicka et al., 1996). However, TGF-β1 signaling–deficient mice revealed no detectable alterations in the HSC compartment in vivo, but demonstrated increased proliferation in vitro after SCF stimulation (Larsson et al., 2003, 2005). Additionally, the loss of Tgfbr1 had normal HSC function and self-renewal potential (Larsson et al., 2003, 2005). In contrast, loss of Tgfbr2 or Smad4 resulted in reduced chimerism in competitive transplants (Karlsson et al., 2007; Yamazaki et al., 2011). Other studies have demonstrated that My-HSCs and myeloid cell lines treated with low levels of TGF-β1 in vitro exhibit increased proliferation (Kale and Vaidya, 2004; Kale, 2004; Karlsson et al., 2007; Chambers et al., 2008; Challen et al., 2010).
Although we observed an increase in TGFBR1, this did not translate into increased signaling, as we found a reduction in SMAD2/3 phosphorylation, total levels of SMAD3, decreased p57 expression, and insensitivity to the growth stimulatory effects of TGF-β. Furthermore, the reduced TGF-β signaling ultimately resulted in a decrease in HSC self-renewal and a dramatic reduction in My-HSCs. This observation provides a potential explanation for the reduction in the myeloid lineage compared with the lymphoid lineage.
Msi2 is well known for its function as a translation inhibitor, but previous studies have shown that Msi2 can also act as a translation activator depending on the cell cycle status (Charlesworth et al., 2006; Arumugam et al., 2010, 2012). This can explain why Msi2 can bind to both TGFBR1 and Smad3 and yet affect their translation differently. It will be interesting to understand how Msi2 can act either as a translation inhibitor or activator.
Our study characterizes Msi2 deletion in the hematopoietic system and uses global RNA binding analysis to identify a novel link between Msi2 and TGF-β signaling in regulating quiescence and myeloid/lymphoid lineage determination of HSCs. It will be interesting to test whether this pathway is retained and functionally important for leukemic stem cells or whether Msi2 modulates other critical pathways in these cells. Collectively, these results have important implications for understanding how the activity of RNA-binding proteins contribute to normal tissue homeostasis and stem cell function and how MSI2 and its targets contribute to oncogenic transformation.
MATERIALS AND METHODS
qRT-PCR.
mRNA was isolated from FACS-purified populations using TRIzol (Life Technologies) and the RNeasy RNA extraction kit (QIAGEN) and reverse transcribed into cDNA with iSCRIPT (Bio-Rad Laboratories). qRT-PCR was performed as previously described with the indicated primers (supplemental dataset) for p21, p27, p57, Actin, JunB, Gfi1, Id2, Gata2, Evi1, c-myc, and Egr1. qRT-PCR primers designed for 3′ UTRs for RIP samples are listed in the supplemental dataset.
Msi2 conditional knockout mice.
We cloned one loxP site 3′ of the transcriptional start site and a floxed PGK-neomycin-resistance cassette 3′ of the fifth intron using a Kpn1 fragment encompassing the 3′ region of the Msi2 locus cloned into the pBS backbone. (Fig. 1 a). The targeting vector was electroporated into V6.5 embryonic stem cells, and G418-resistant colonies were subcloned. Homologous recombinants were detected by Southern blotting using 5′ and 3′ external probes, and correctly targeted embryonic stem clones were transiently treated with Cre to remove the Neo resistance cassette (verified by Southern blotting), before blastocyst injection of conditional 2-lox clones. Resulting chimeras were backcrossed to C57BL/6 mice and the F1 generation was bred to Mx1 Cre mice on a C57BL/6 background. Deletion was initiated using pIpC (GE Healthcare) by intraperitoneal injections every 3–4 d as indicated (150 mg/kg) per mouse three times. Cre recombination of the conditional 2-lox (floxed) locus to the inactive, 1-lox (null) locus was verified by Southern blotting after cutting genomic DNA with XbaI and hybridization with a 5′ external probe. All of the animal experiments were approved under the Institutional Animal Care and Use Committee.
Noncompetitive transplants.
Noncompetitive transplants were performed with 106 BM cells from 6–10-wk-old Msi2flox/flox or littermate control mice, transplanted into lethally irradiated B6SJL congenic CD45.1 recipients. In the primary transplants with Msi2Δ/Δ BM, transplants were performed 12–16 wk after Msi2 deletion. In the cell-autonomous transplants, transplanted marrow was allowed to engraft for 4–6 wk before pIpC was administered. Peripheral blood chimerism was assessed after engraftment before deletion to verify successful engraftment.
Peripheral blood analysis.
Peripheral blood was collected from the retroorbital cavity using a heparinized glass capillary tube. Complete peripheral blood count analysis including a differential blood count was obtained by using Hemavet (Drew Scientific).
Flow cytometry.
BM cells or splenocytes were harvested and subjected to red blood cell lysis. Fresh or frozen cells were stained with the following Abs: CD45.2-FITC and CD45.1-APC, Mac1-PE, Gr1-APC, c-Kit–APC, CD71-PE, Ter119-APC, B220-PE, and CD3-APC (BD) and analyzed on the BD FACSCalibur instrument. Staining for multiparameter flow cytometry was performed after a c-kit enrichment using 10 µl MACS beads (CD117) per mouse and then run on an AutoMACS (Miltenyi Biotec) according to the manufacturer’s instructions. The cells were then stained with the following cocktail: (Lineage; CD3, CD4, CD8, Gr1, B220, CD19, and TER119 all conjugated with PeCy5), Sca-Pac Blue, CD34-FITC or CD45.2-FITC, SLAM-APC, CD48-PE, c-KIT–Alexa Fluor 780, and FcgRIIb-PeCy7 (Fig. 2 d); HPCs (Linloc-Kit+Sca1−), GMPs (LK, FcγRIIbhiCD34+), CMPs (LK, FcγRIIbmid CD34+), MEPs (LK, FcγRIIbloCD34−), B cells (B220+), and T cells (CD3+) from the spleen were also sorted. For analysis of LMPPs and CLPs, the following cocktail was used: Lineage marker mix–PeCy5, Sca-Pecy7 IL-7Ra–Pac Blue, Flk2-PE, CD34-FITC, and Kit-APC (Martin et al., 2003; Adolfsson et al., 2005; Karsunky et al., 2008).
Colony-forming assays.
200 sorted HSCs or 500 sorted HSPCs were plated in in M3434 methylcellulose media (STEMCELL Technologies), and colonies were scored after 10 d and 7 d, respectively.
Microarray analysis of Msi2 expression in conditional knockout mice.
HSPCs (LineageloScaI+c-Kit+) were sorted 4 wk after pIpC injection. RNA was extracted using TRIzol and the RNeasy RNA extraction kit. RNA was then amplified using the NUGEN Pico amplification kit, fragmented, and hybridized on Mouse Expression Array 430 2.0 (Affymetrix) at the Microarray Core Sub-Facility (Molecular Genetics Core, Division of Genetics, Intellectual and Developmental Disabilities Research Center, Children’s Hospital Boston; CLS 16). Signal normalization was performed by robust multi-array average (RMA) method. Data were analyzed using GSEA across the complete list of genes ranked by signal to noise ratio (microarray data, n = 4 Msi2flox/flox mice and n = 4 Msi2Δ/Δ mice). The microarray data can be found in the GEO database under accession no. GSE53385.
Immunofluorescence.
HSCs or HSPCs were sorted from primary mice 6 wk after pIpC and cultured with STIF media (Stemspan media [STEMCELL Technologies] containing 10 ng/ml heparin, 10 ng/ml SCF, 20 ng/ml TPO, 20 ng/ml IGFII, and 10 ng/ml FGF) in 96 round-bottom wells for 16 h and then treated with 10 nM Nocodazole for 24 h. Cells that were fixed with 1.5% paraformaldehyde and permeabilized with cold methanol were cytospun onto glass slides and were then stained on slides with anti-Numb (Abcam) and secondary Ab (donkey anti–goat Alexa flour 546; Molecular Probes) with DAPI counterstaining (Kharas et al., 2010). Symmetric versus asymmetric percentages were assessed based on quantification of the signal intensity of each cell (divided by surface area) normalized for background staining using an Axio Imager M2 microscope (Carl Zeiss), followed by a confirmatory visual inspection that determined symmetric commitment versus symmetric renewal. Daughter cells with elevated equivalent staining of Numb were counted as symmetric commitment, whereas if one daughter cell contained more staining than the other cell, then the division was considered an asymmetric division. If there was low or no staining in the daughter pairs, this was scored as a symmetric renewal division. Cells were also stained for phospho-Smad2/3 (EMD Millipore), and staining was quantified as above in all cells including nondividing cells. Phospho-Smad2/3 was also performed on HSCs directly isolated from animals with similar results. Freshly isolated HSCs that were fixed and permeabilized as mentioned above were stained on glass slides for c-Myc (Epitomics), TGFBR1 (Abcam), and pan-Smad3 (Cell Signaling Technology) using secondary Ab (donkey anti–rabbit Alexa Fluor 488; Molecular Probes). Quantification of the signal intensity of each cell (divided by surface area) normalized for background staining was measured using AxioVision software (Carl Zeiss).
In vitro HSC and HSPC proliferation assay.
We sorted 100 HSCs (LSK CD48−CD150+) and 1,000 HSPCs (LSK+CD150− and LSK+CD150+CD48+) into Stemspan media containing 10 ng/ml IL-3, 10 ng/ml IL-6, 50 ng/ml SCF, 10 ng/ml TPO, and 20 ng/ml Flt3L in a round-bottom plate and counted at days 7 and 5, respectively. Then cells were analyzed using counting beads and indicated markers for differentiation by flow cytometry. For TGF-stimulated proliferation, 50 HSCs were sorted into a well of a 96-well plate containing media as above with different doses of TGF-b1 (R&D Systems). Cells were then counted 7 d later using counting beads (Life Technologies) with flow cytometry.
Statistical analyses.
For bar graphs, the unpaired two-tailed Student’s t test was used to compute p-values, except where stated otherwise. Error bars reflect the SEM, except where stated otherwise. All statistical analyses were performed using Prism 4.0 (GraphPad Software) and the R statistical environment.
RIP.
20 million K562 control vector cells and cells overexpressing Flag-MSI2 were used for RIP using the Magna RIP RNA binding protein immunoprecipitation kit (EMD Millipore). In brief, cells were washed with cold PBS and lysed with RIP lysis buffer provided from the kit. 5 µg anti–Flag M2 Ab (Sigma-Aldrich), anti–rabbit Ab, or anti-Msi2 Ab (EMD Millipore), which were incubated with magnetic beads, were used to immunoprecipitate Flag–MSI2–RNA complexes. Immunoprecipitated complexes were washed and treated with proteinase K. RNA was extracted using the phenol/chloroform method, and the resulting RNA was converted to cDNA using the Verso cDNA kit (Thermo Fisher Scientific). cDNA was then used for qRT-PCR validating candidates from the HITS-CLIP.
UV cross-linking, immunoprecipitation, and high-throughput RNA sequencing (HITS-CLIP).
The protocol used is mainly based on UV cross-linking and immunoprecipitation with high-throughput RNA sequencing (HITS-CLIP) published by Chi et al. (2009). In brief, 100 million empty vector control– or Flag-MSI2–overexpressing K562 cells were UV cross-linked with 4,000 J at 265 nm on ice. Cells were washed three times with cold PBS containing protease inhibitors, and the cell pellets were resuspended in a low salt buffer that is optimized for reducing nonspecific RNA interactions (100 mM NaCl, 0.1% SDS, 0.5% sodium deoxycholate, 0.5% Nonidet P-40) with protease inhibitors. Cells were sonicated with a Branson probe sonicator for three cycles of 10 s (0.7 s on/1.3 s off) at 20% power on ice. The lysates were then rested on ice for 20 min. RQ1 DNase (Promega) was added to the lysates and incubated at 37°C for 5 min. RNase A (USB Products) was added at a concentration of 1:1,000 and incubated at 37°C for 10 min. Lysates were spun and supernatants were incubated with protein G Dynabeads (Invitrogen) that were preincubated with anti–Flag M2 Ab in low salt buffer (Sigma-Aldrich). Beads were washed with low salt wash buffer three times followed by two washes with high salt buffer (500 mM NaCl, 0.1% SDS, 0.5% sodium deoxycholate, 0.5% Nonidet P-40) to reduce nonspecific protein interactions. 5 µg proteinase K (Invitrogen) was added to the washed beads and incubated at 70°C for 2 h. RNA was extracted using standard phenol/chloroform extraction methods, the RNA was precipitated at 80°C overnight, and the resulting RNA pellets were washed with 80% ethanol to retain short RNA fragments. RNA sizes were assessed on a 2100 Bioanalyzer (pico chip), and the sizes of extracted RNA ranged between 40 and 200 bases in length. RNA libraries were prepared and barcoded at the Broad Institute Sequencing Core facility and then sent to the core facility at the Sloan-Kettering Institute for single-end Illumina sequencing. A first round of sequencing for one control and two replicates produced ∼20 million 36-bp alignable reads for each library. The second replicate was sequenced for an additional 60 million reads to enhance coverage.
Statistical analysis and plots of HITS-CLIP data.
To identify potential binding sites, we used a custom R package to call peaks in read coverage in the Msi2 HITS-CLIP experiments (Loeb et al., 2012), and we tested for differential binding between Msi2-overexpressing cells and control while correcting for differential expression using a joint generalized linear model of HITS-CLIP and RNA sequencing read counts in the candidate peak regions. Peak windows that were differentially bound at an FDR threshold of 0.1 were reported unless otherwise noted.
Processing CLIP reads.
Using the FASTX-Toolkit from the Hannon Laboratory (Cold Spring Harbor Laboratory, Cold Spring Harbor, NY), 3′ ends of reads were trimmed until the base quality score was not <30 and retained if the remaining read length was at least 20 nt. After trimming, we mapped CLIP reads to the hg19 human genome with Bowtie2 (Langmead and Salzberg, 2012).
Identification and annotation of Msi2-binding sites from HITS-CLIP data.
To identify possible Msi2-binding sites in HITS-CLIP data, we reimplemented our previous CLIP-seq peak-calling pipeline as an R package to enable efficient and transcriptome-wide identification and annotation of peaks mapped by various sequencing technologies (Loeb et al., 2012). In brief, our approach was inspired by edge detection in computer vision, where sharp changes in brightness in an image are detected as edges of an object by computing the rate of change of the intensity gradient.
To identify peaks, we first combined the reads from all of the HITS-CLIP experiments together. Reads previously trimmed for low base quality were extended to a minimum length of 30 nt. We then constructed a 1D signal profile of read coverage, K[x], which contains cumulative read counts for each position x from all HITS-CLIP libraries. To simultaneously smooth and identify edges in the signal, this profile was convolved with a kernel derived from the second derivative of a Gaussian (gD″), with a mean of 0, standard deviation of 1, and bandwidth set to 15 nt. The edges in the original signal are located at the zero-crossings of the convolved signal.
The zero-crossings of the second derivative that switch from positive to negative indicate the edges that start a peak, and the points that switch from negative to positive identify the ends of each peak. Once the peaks were identified, we then quantified peaks in each experiment by counting the number of reads from each experiment that overlapped with each peak. Reads that overlapped more than one peak were assigned to the peak with which it had maximal overlap.
After peak identification, each CLIP peak was annotated according to the RefSeq gene annotation for hg19 (downloaded from UCSC on April 4, 2012). Genes with multiple isoforms were reduced to a unified gene model, which is the union of all annotated exons. To compensate for possible errors in annotation, for each gene we extended the first exon 1 kb upstream and the last exon 5 kb downstream. Using these models, HITS-CLIP peaks were then annotated with the specific genomic region (CDS, intron, 5′ UTR, and 3′ UTR) that the peaks overlap. If a genomic region was assigned to multiple categories, we assigned it to one of them with the following order: 3′ UTR, CDS, 5′ UTR, and intron. Peaks that mapped to multiple genomic regions were assigned to the region with maximum overlap.
Processing RNA-Seq reads.
Paired-end RNA-Seq reads were aligned to human genome hg19 with Tophat (Trapnell et al., 2010). Then we counted the number of RNA-Seq reads within each previously defined HITS-CLIP binding site using the HTSeq package from the Huber Laboratory (European Molecular Biology Laboratory, Heidelberg, Germany).
Identification of significant HITS-CLIP peaks by integrating gene expression.
As flag-tagged Msi2 protein was not expressed in control cells, HITS-CLIP reads from the control sample were generated by nonspecific binding of the Ab and other sources of background noise. A fraction of reads in samples with overexpressed Msi2 also came from these noise sources (Chi et al., 2009). Therefore, we were interested in identifying HITS-CLIP peaks with significantly higher read counts in Msi2-overexpressing cells relative to control, as they are likely to be the real Msi2-binding sites. However, as mRNA expression levels may change between different conditions, differential read counts at a site can be caused either by a change in transcript abundance or by differential Msi2 binding. To identify real differential binding events, it is necessary to integrate gene expression data into the analysis.
Specifically, we jointly modeled read count data from HITS-CLIP and RNA-Seq with a novel generalized linear model approach. We represented the read count from a window containing peak i in sample j as Kij, which was seen as realization of random variable Kij; here, the read count represents either HITS-CLIP reads or RNA-Seq reads in the window, depending on sample j. It is assumed that Kij follows a negative binomial distribution, which has been widely used in modeling read count data (Anders et al., 2012; McCarthy et al., 2012).
For each peak i, the expected value of Kij, denoted by μij, is fit via a logarithmic link by the following model:
Here represented scaled library size of sample j, which was the total read count in sample j scaled by the weighted trimmed mean of log expression ratios (Robinson and Oshlack, 2010). It was included as a normalization factor. After normalization, the logarithm of this variable is decomposed into four factors, where the regression coefficients have the following interpretation: represents the baseline log expression level measured by the window at peak i; represents the baseline log read count ratio of CLIP reads to RNA-Seq reads at peak i; represents the effect of Msi2 overexpression on read counts caused by mRNA expression changes. Finally, the interaction term represents differential Msi2 binding caused by overexpression; this coefficient will be nonzero if there is differential binding even after controlling for differential mRNA expression. Factors and equal 1 or 0, depending on the condition and library type of sample j. To test whether the interaction term is 0, we fit data to both the full model and a reduced model without the interaction term. Then the deviances of two models were used to conduct a likelihood ratio test. If there is no true differential binding effect, the difference in deviances between the nested models should be small compared with a χ2 distribution with one degree of freedom. In this way, we were able to characterize the significance of differential binding with a p-value. Finally, we defined Msi2-binding sites as sites with Benjamini-Hochberg–adjusted p-value <0.1 and > 0. The above analysis was conducted using the edgeR package (McCarthy et al., 2012).
Visualization of GSEA results.
GSEA (Subramanian et al., 2005) was conducted independently on expression profiles of Msi2Δ/Δ LSKs and HITS-CLIP targets in K562. The overlapping gene sets were significantly enriched (FDR < 0.01) in both analyses. The relationships between those common gene sets were visualized using HTSanalyzeR (Wang et al., 2011) and igraph packages in R. In Fig. 7 (f and g), each node represents a gene set, and its size represents the number of genes in the gene set. An edge between two nodes indicates that two gene sets share part of the genes, and the width of each edge shows the percentage of overlap between two gene sets.
Online supplemental material.
Fig. S1 shows the gating strategy for Fig. 1 g. Fig. S2 shows the gating strategy for Fig. 2. Fig. S3 shows the gating strategy for Fig. 3. Fig. S4 shows the gating strategy for Fig. 4. Fig. S5 shows the gating strategy for Fig. 5 (a and e). Table S1, included as a separate Excel file, shows microarray analysis of the differentially expressed genes in Msi2-deficient LSK cells. Table S2, included as a separate Excel file, lists the additional curated gene sets utilized in GSEA analyses. Table S3, included as a separate Excel file, lists the statistically significant gene sets from Msi2-deficient LSK microarray expression analysis. Table S4, included as a separate Excel file, shows the location and number of the significant HITS-CLIP targets. Table S5, included as a separate Excel file, lists the statistically significant gene sets from the HITS-CLIP ranked list in K562 cells. Table S6, included as a separate Excel file, lists the significant and overlapping gene sets from the HITS-CLIP and microarray analyses. The supplemental dataset, included as a separate Excel file, shows primer sequences used for the qRT-PCR experiments.
Acknowledgments
We would like to acknowledge Tingting Zhang, Gary D. Gilliland, Stephen Nimer, and Ross Levine for all their critical advice and suggestions. We would also like to thank Varun Shenoy, Eileen Shiuan, Aly Azeem Khan, Anisha Ganguly (Microarray Core Sub-Facility, Molecular Genetics Core, Division of Genetics, Intellectual and Developmental Disabilities Research Center, Children’s Hospital Boston; CLS 16), Agnes Viale, and the Memorial Sloan-Kettering Cancer Center Sequencing and Genomics Core for all their support.
M.G. Kharas was supported by the US National Institutes of Health (NIH) National Institute of Diabetes and Digestive and Kidney Diseases Career Development Award. The Tri-Institutional Starr Stem Cell Fellowship supports S.-M. Park. This work was also supported by grants from the NIH (M.G. Kharas), the Louis V. Gerstner Young Investigator Award, American Society of Hematology Junior Scholar Award, the Tri-Institutional Starr Foundation, and the Howard Hughes Medical Institute (to G.Q. Daley). C.J. Lengner was supported by an R01 from the National Cancer Institute (NIH) and a fellowship from the W.W. Smith Charitable Trust.
The authors have no conflicting financial interests.
References
- 5-FU
5-Fluorouracil
- Ab
antibody
- CDS
coding DNA sequence
- CLP
common lymphoid progenitor
- CMP
common myeloid progenitor
- FDR
false discovery rate
- HITS-CLIP
cross-linking and immunoprecipitation followed by high-throughput sequencing
- HSC
hematopoietic stem cell
- HSPC
hematopoietic stem and progenitor cell
- LMPP
lymphoid primed myeloid progenitor
- My-HSC
myeloid-biased HSC
- pIpC
polyinosinic:polycytidylic acid
- qRT-PCR
quantitative real-time PCR
- RIP
RNA immunoprecipitation
- UTR
untranslated region



![Figure 2. Msi2 deficiency decreases repopulating activity. (a) 106 BM cells from mice treated with pIpC were transplanted, and these transplanted animals were sacrificed at 22–24 wk after pIpC with the indicated organ weights (n = 9 from two independent transplants). (b) Representative CD45.2 chimerism flow plot for the HSCs, LSK CD48−CD150+; mean and SEM shown next to gate and also in c. (c) CD45.2 chimerism analysis from HSCs, LSK+CD48−CD150+. See Materials and methods for the surface phenotype of the progenitors (n = 8, 9, same experiment as in panel a). (d) qRT-PCR expression of Msi2 normalized to Gapdh in sorted LSKs from the indicated primary mice or transplanted mice (c) and analyzed at various time points after pIpC (primary mice 4 wk [before transplant] n = 4 per genotype; 12 wk [before transplant], n = 3 per genotype; and 16–24 wk [transplanted mice], n = 9 or n = 8 for Msi2f/f or Msi2Δ/Δ, respectively). (e) BM cells were sorted for HSCs, LSK CD34−CD48−CD150+, and 3,000 HSCs were transplanted into lethally irradiated mice together with 300,000 BM support cells. Peripheral blood was then examined at 10 wk after engraftment (n = 4 mice per genotype). (f) Congenic mice (CD45.1) were noncompetitively transplanted with control or Msi2flox/flox marrow and were allowed to engraft normally (6 wk); peripheral blood chimerism was analyzed before injection of pIpC. (g) Chimerism analysis 18 wk after pIpC from mice in f (see Fig. S2 a [control, n = 10; or Msi2Δ/Δ, n = 11 from three independent transplants] or Fig. S2 b for gating; LT-HSC: LSK+CD34−FLK2/3−, ST-HSC: LSK+CD34+FLK2/3−, LMPP: LSK+CD34+FLK2/3+, representative of control, n = 3; or Msi2Δ/Δ, n = 5 mice). Means and SEM are shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.](https://cdn.rupress.org/rup/content_public/journal/jem/211/1/10.1084_jem.20130736/4/m_jem_20130736r_fig2.jpeg?Expires=1781615175&Signature=a~Tw~pJgIoV-SCHH0PnHidKi0pkrD7ERuts~Wumf-e868sl1l-XPkjvnqQeQYKaVGOFRUn7kvte8u-38dr4cerQhyZCk0o3aReqQ1OMWT9K1S4gwUlJ2IQufeM~MrC0mWbHvlMc11nbuoEAjMZTd03cp9s74TcgCOKhjyd-d02B9obnPkl-XLP5HpKfslBSD1ndIu1oOrQpzedf4DcS3lapTJNqmmKAsYZe3oMCWvruU9Rj6Uq~DM3r8X~MhAhZiruKDV1gwA3q9Khm2JyOvNk9cMf7JlTDydsJ3EmKu5Yf6vcELXVwbrF6ugxs1G4tzD6LTIvnvZbBSw99G0WA28A__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
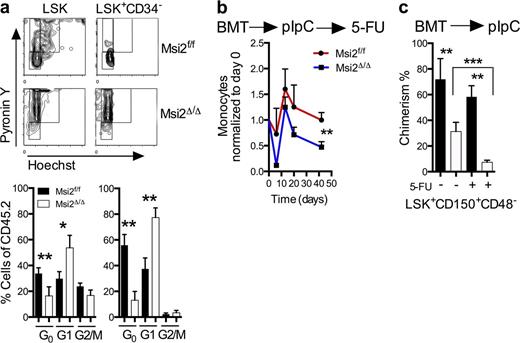
![Figure 4. Msi2Δ/Δ cells have reduced myeloid lineage–biased HSCs and defective myeloid repopulating activity. (a) Chimerism analysis of the myeloid and lymphoid populations in the BM and spleen of mice 22–24 wk after transplant (same as in Fig. 2 [a and b]). (b) Chimerism of myeloid and lymphoid lineages in spleen of primary transplanted mice shown 12–18 wk after pIpC (n = 9, 10 from two independent experiments). (c) Chimerism analysis 9 wk after pIpC and 40 d after 5-FU injections (same as in Fig. 3 [b and c]). (d) Chimerism analysis at 18 wk after pIpC, and CLP (Linloc-kitmidSca+Flk2+IL7R+, CLP2: Linloc-kitloSca+Flk2+IL7R+) populations are based on gating in Fig. S4 a (control, n = 3; or Msi2Δ/Δ, n = 5 mice). (e) Representative flow analysis (left; with indicated staining from 18-wk-post-pIpC mice). Mean and SEM for the frequency of CD150hi My-HSCs gated from (left flow plot) 4 wk (n = 12 and n = 15 from five independent experiments) and 18 wk (n = 5 and Msi2Δ/Δ n = 8 from two independent experiments). (f) Flow cytometric analysis of My-HSCs: LinloCD45.2+LSK CD150hiCD86− (n = 6, n = 8 mice from two independent experiments). See gating strategy in Fig. S4 b. SEM is shown (*, P < 0.05; **, P < 0.01; ***, P < 0.001). BMT, BM transplant.](https://cdn.rupress.org/rup/content_public/journal/jem/211/1/10.1084_jem.20130736/4/m_jem_20130736_fig4.jpeg?Expires=1781615175&Signature=x-sFm7xwg1Xu41IVWsD8LUK4~IYPhEck0oQoUuLb2bG1k9pjK5qqvtQdtTmoNqwQQusHu3yW1KmELMUznnMHQFg9mMi5sqi8o-6HC54Fax-U8DOjw95Ua-MZFcWLYYt5zyOrzHjf4GobaihsEB403k1V0kxV5kGl8JiVsmnMsM~z~5mT9klUh4D1w~o~AkRqbS46SlX19JGCfD7bA0q4LB~pueEv~K1IEAmxzGTukCKvREBHmzY19873fQW0ADQVuqUc~le66iNg0cUqWvPTnPxKfHDfY1lmei0oP1pej4pHTEjh-NaF5ihUjIdRIXhBeD1MKWpdP9O7fqvGOfivlQ__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)