Class switch recombination (CSR) allows the humoral immune response to exploit different effector pathways through specific secondary antibody isotypes. However, the molecular mechanisms and factors that control immunoglobulin (Ig) isotype choice for CSR are unclear. We report that deficiency for the Ikaros transcription factor results in increased and ectopic CSR to IgG2b and IgG2a, and reduced CSR to all other isotypes, regardless of stimulation. Ikaros suppresses active chromatin marks, transcription, and activation-induced cytidine deaminase (AID) accessibility at the γ2b and γ2a genes to inhibit class switching to these isotypes. Further, Ikaros directly regulates isotype gene transcription as it directly binds the Igh 3′ enhancer and interacts with isotype gene promoters. Finally, Ikaros-mediated repression of γ2b and γ2a transcription promotes switching to other isotype genes by allowing them to compete for AID-mediated recombination at the single-cell level. Thus, our results reveal transcriptional competition between constant region genes in individual cells to be a critical and general mechanism for isotype specification during CSR. We show that Ikaros is a master regulator of this competition.
Class switch recombination (CSR) diversifies the humoral immune response by joining a single antibody variable region gene with different constant region (CH) genes responsible for unique effector functions (1). This is crucial for establishing immunity, as patients selectively deficient in CSR suffer from recurrent and severe infections (2). CSR occurs between repetitive but nonhomologous DNA sequences called switch (S) regions, which are located upstream of each CH gene (except δ). CSR requires the expression of activation-induced cytidine deaminase (AID) (3, 4), an enzyme that is thought to directly deaminate single-stranded DNA (5, 6), though this mechanism is still under debate (7). DNA lesions induced by AID are processed to generate double-stranded DNA breaks (DSBs), which activate DNA damage response proteins to promote efficient long-range recombination (8). DSBs in Sμ and downstream S regions are ultimately joined through end joining mechanisms, allowing the expression of a new antibody isotype (1).
CSR requires transcription and is targeted to individual constant region genes by the selective activation of isotype-specific intronic (I) promoters in response to antigen, cytokine, and co-stimulatory signals (9). This “germline transcription” begins at I exons and proceeds through adjacent S regions and CH genes, giving rise to noncoding germline transcripts (GLTs). Transcription is thought to initiate CSR by promoting S region accessibility and exposing single-stranded DNA to AID (1). Indeed, CSR is abrogated by I promoter deletions (10, 11) and is restored by their replacement with heterologous promoters (12, 13). These latter studies also demonstrated that constitutively transcribed S regions are ectopically targeted for CSR, highlighting the role of S region transcription in isotype selection. However, the mechanisms establishing this targeting are not completely understood, and it is unclear how individual cells select between simultaneously transcribed S regions for CSR.
Germline transcription is regulated by an enhancer at the 3′ end of the Igh locus and by chromatin modifications. The 30-kb Igh 3′ enhancer lies downstream of Cα and contains four DNase hypersensitive (HS) regions: HS3a, HS1,2, HS3b, and HS4. Disruption of the enhancer reduces transcription and CSR to all isotypes, with γ3, γ2b, and γ2a most drastically affected (14, 15). As the 3′ enhancer is distant from I promoters (up to 110 kb), transcriptional control is believed to occur through promoter–enhancer looping (16). In addition, histone modifications, such as histone H3 acetylation (AcH3) at I exons and S regions, are tightly correlated with GLT induction, indicating that they may regulate germline transcription (17, 18). Nonetheless, the molecular mechanisms and factors controlling S region transcription and isotype specification during CSR remain largely undefined.
The Ikaros zinc finger transcription factor plays important roles in B cells. Ikaros is required for B cell specification (19, 20) and differentiation (20–22), as well as allelic exclusion at the Igκ locus (23, 24). We have studied Ikaros function in the B cell lineage using mice bearing a hypomorphic mutation in the ikzf1 (Ikaros) locus (IkL/L). IkL/L mice contain a LacZ reporter knocked into ikzf1 exon 2, resulting in the production of low levels of functional, but truncated, Ikaros proteins (∼10% of WT) in hematopoietic cells (21). Unlike Ikaros-null mice (19), IkL/L mice develop relatively normal numbers of mature, polyclonal B cells (21). Interestingly, IkL/L mice exhibit abnormal serum antibody titers, characterized by striking >50% reductions in IgG3 and IgG1, and >50% increases in IgG2b and IgG2a (21). This intriguing observation led us to hypothesize that Ikaros plays a role in isotype selection. In this paper, we report that Ikaros is indeed a central regulator of Igh locus transcription and isotype specification during CSR.
RESULTS
Ikaros deficiency skews CSR to IgG2b and IgG2a
To determine if Ikaros regulates CSR, switching to all isotypes was assessed in purified WT and IkL/L splenic B220+ B cells using a battery of in vitro culture conditions. CSR was measured by flow cytometry (FACS) for surface Ig isotype expression after 3–4 d in culture. After LPS stimulation, WT cells switched only to IgG3 and IgG2b, as expected (Fig. 1 A and Fig. S1 A). In contrast, IgG2b+ cells were 3.6-fold more frequent in IkL/L cultures, and IgG2a+ cells were also detected, whereas IgG3+ cells were reduced (23% of WT; Fig. 1 A and Fig. S1 A). After LPS + IFN-γ stimulation, both genotypes switched to the same isotypes, but IkL/L cultures produced more IgG2b+ and IgG2a+ cells (up 2.3- and 2.7-fold, respectively, vs. WT) and fewer IgG3+ cells (24% of WT; Fig. 1 B and Fig. S1 B). After LPS + IL-4 stimulation, WT cells switched to IgG1 (Fig. 1 C and Fig. S1 C) and IgE (as determined by RT–quantitative PCR [qPCR] of ε postswitch transcripts; Fig. 2 A) (25), as expected. In contrast, IkL/L cells switched ectopically to IgG2b, IgG2a, and IgG3, produced fewer IgG1+ cells than WT (WT, 30.1 ± 1.63%; IkL/L, 26.2 ± 1.57%), and exhibited a 54% reduction in ε postswitch transcripts, indicating fewer IgE+ cells (Fig. 1 C, Fig. 2 A, and Fig. S1). After LPS + IL-5 + TGF-β stimulation, WT cells switched to IgG3, IgG2b, and IgA, whereas IkL/L cultures generated 2.9-fold more IgG2b+ cells, unexpected IgG2a+ cells, and fewer IgG3+ and IgA+ cells (32 and 52% of WT, respectively; Fig. 1 D and Fig. S1 D). In all of the these conditions, the fraction of switched cells (e.g., IgG+ and IgA+) in Ikaros-deficient cultures increased 20–120% versus WT, suggesting a general increase in switching capacity (Fig. 1). Finally, we confirmed these results by ELISA (Fig. S2), demonstrating that isotype expression is significantly skewed toward IgG2b and IgG2a in activated Ikaros-deficient B cells.
CSR is skewed toward IgG2b and IgG2a in IkL/L B cells. CFSE-labeled B220+ WT and IkL/L B cells were stimulated for 72 h with (A) LPS, (B) LPS + IFN-γ, or (C) LPS + IL-4. (D) 24 h with LPS was followed by 72 h with LPS + IL-5 + TGF-β. CSR was analyzed by FACS, and percentages of Ig+ cells are indicated. The data are representative of more than four independent experiments. Bar graphs represent mean percentages plus SD of switched cells in each condition for four (D) or five (A–C) experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02).
CSR is skewed toward IgG2b and IgG2a in IkL/L B cells. CFSE-labeled B220+ WT and IkL/L B cells were stimulated for 72 h with (A) LPS, (B) LPS + IFN-γ, or (C) LPS + IL-4. (D) 24 h with LPS was followed by 72 h with LPS + IL-5 + TGF-β. CSR was analyzed by FACS, and percentages of Ig+ cells are indicated. The data are representative of more than four independent experiments. Bar graphs represent mean percentages plus SD of switched cells in each condition for four (D) or five (A–C) experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02).
Reduced IgE switching in IkL/L B cells despite normal ε GLT expression. (A) Iμ-Cε transcript expression in B220+ WT and IkL/L B cells after 72 h of stimulation with LPS + IL-4 was analyzed by RT-qPCR. The Iμ promoter is equally active in WT and IkL/L cells (Fig. 4, A–D), indicating that Iμ-Cε transcripts can be faithfully compared between genotypes to quantify IgE-switched cells. (B) RT-qPCR for ε GLT expression after 48 h of LPS + IL-4 stimulation. Igβ was used as a loading control and all values were normalized to WT. Significance was determined by a two-tailed t test assuming unequal variance (**, P < 0.02). Graphs represent means plus SD of two (A) or three (B) independent experiments. Neither Iμ-Cε transcripts nor ε GLTs were detected in other conditions (not depicted).
Reduced IgE switching in IkL/L B cells despite normal ε GLT expression. (A) Iμ-Cε transcript expression in B220+ WT and IkL/L B cells after 72 h of stimulation with LPS + IL-4 was analyzed by RT-qPCR. The Iμ promoter is equally active in WT and IkL/L cells (Fig. 4, A–D), indicating that Iμ-Cε transcripts can be faithfully compared between genotypes to quantify IgE-switched cells. (B) RT-qPCR for ε GLT expression after 48 h of LPS + IL-4 stimulation. Igβ was used as a loading control and all values were normalized to WT. Significance was determined by a two-tailed t test assuming unequal variance (**, P < 0.02). Graphs represent means plus SD of two (A) or three (B) independent experiments. Neither Iμ-Cε transcripts nor ε GLTs were detected in other conditions (not depicted).
Because CSR requires proliferation (26) and IkL/L B cells hyperproliferate at low stimuli concentrations and cell densities (21), overall increases in switched cells and/or specific increases in IgG2b+ and IgG2a+ cells could be caused by increased cell divisions in IkL/L cultures. To evaluate this, CFSE dilution was used to track cell division, and the percentage of IgG/A+ WT and IkL/L cells in each generation was compared for the described conditions. Each generation of IkL/L cells exhibited increases in IgG2b+ and IgG2a+ cells, as well as increased overall switching after all stimulations when compared with WT; increases in IgG3+ cells were also detected after LPS + IL-4 stimulation (Fig. S3). Further, IkL/L cells exhibited reduced CSR to IgG3 and IgA at every division analyzed and to IgG1 after the third division in conditions that induce WT cells to switch to these isotypes (LPS, LPS + IFN-γ, and LPS + IL-5 + TGF-β for IgG3, LPS + IL-4 for IgG1, and LPS + IL-5 + TGF-β for IgA; Fig. S3). These results indicate that significant, proliferation-independent isotype expression abnormalities exist in IkL/L B cells after CSR-inducing stimulation.
It was also possible that the increased IgG2b+ and IgG2a+ populations in IkL/L cultures could be caused by the expansion of autoreactive cells that had previously switched in vivo, rather than specific CSR defects, considering the lower activation thresholds of IkL/L B cells (21) and the production of autoantibodies in transgenic mice expressing a B cell–restricted dominant-negative Ikaros isoform (27). Thus, isotype expression was analyzed using whole and bona fide unswitched B cells (B220+, and CD43−IgG/A− and CD43−IgM+, respectively) that were stimulated with LPS, LPS + IFN-γ, and LPS + IL-4. Regardless of purification strategy, IkL/L cultures exhibited consistent and significant increases in IgG2b+ and IgG2a+ (and IgG3+ with LPS + IL-4) cells compared with WT cultures (Fig. S4). Further, there were no significant differences in IgG2b or IgG2a expression between B220+, CD43−IgG/A−, and CD43−IgM+ IkL/L cultures (Fig. S4). Thus, the aberrant isotype expression by Ikaros-deficient B cells results from defects in CSR and not from the expansion of in vivo–switched cells.
In summary, IkL/L cells (a) are more likely to undergo CSR, (b) display enhanced and ectopic CSR to IgG2b and IgG2a regardless of stimulation, and (c) show reduced CSR to all other isotypes (except for ectopic CSR to IgG3 after LPS + IL-4). Therefore, Ikaros deficiency results in a B cell–intrinsic defect in isotype specification during CSR.
Ikaros function is required during CSR
Because Ikaros is expressed throughout B cell development and controls processes such as V(D)J recombination, which may affect the B cell repertoire (19, 20), we asked if Ikaros is required in mature B cells during CSR. WT B cells were stimulated for 24 h with LPS to induce CSR, and were then transduced with retroviruses encoding the dominant-negative Ikaros 6 (Ik6) isoform and a truncated human nerve growth factor receptor (NGFR) reporter. The Ik6 isoform lacks DNA binding domain zinc fingers and, upon dimerization, inhibits DNA binding by functional Ikaros isoforms (28). 72 h after infection, switching and NGFR expression were analyzed by FACS. WT cells transduced with control virus (NGFRhi) switched exclusively to IgG3 and IgG2b, as did uninfected cells (NGFR−) from WT cultures transduced with Ik6-NGFR (Fig. 3). Ik6-NGFRhi cells, however, exhibited a threefold increase in IgG2b+ cells, ectopic CSR to IgG2a, 30–50% fewer IgG3+ cells, and more switched cells (Fig. 3 and Fig. S5 A). Thus, inhibiting Ikaros function in mature WT B cells during CSR recapitulated the IkL/L switching phenotype. Similarly, transduction of IkL/L cells with the full-length Ik1 isoform rescued IkL/L switching defects (Fig. S5, B and C). Importantly, equally effective rescue was achieved with the truncated Ik1 isoform that lacks exon 2–derived protein sequences and is expressed at low levels in IkL/L cells, indicating that these truncated Ikaros proteins function like full-length Ikaros during CSR (Fig. S5, B and C) (21). Collectively these results show that Ikaros regulates CSR specificity and efficiency in mature B cells.
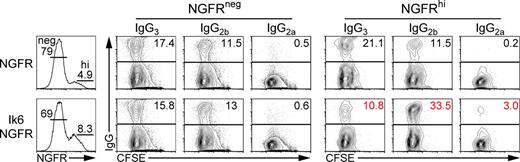
Ikaros regulates CSR in mature B cells. CFSE-stained B220+ WT B cells were activated for 24 h with LPS to induce CSR, infected with retroviruses encoding the NGFR reporter alone or with the dominant-negative Ik6 isoform (Ik6-NGFR), and stimulated for another 72 h with LPS. Ig and NGFR expression were analyzed by FACS. Numbers represent percentages of NGFR−, NGFR+, or IgG+ cells. The data are representative of four independent experiments (Fig. S5 A provides a statistical analysis).
Ikaros regulates CSR in mature B cells. CFSE-stained B220+ WT B cells were activated for 24 h with LPS to induce CSR, infected with retroviruses encoding the NGFR reporter alone or with the dominant-negative Ik6 isoform (Ik6-NGFR), and stimulated for another 72 h with LPS. Ig and NGFR expression were analyzed by FACS. Numbers represent percentages of NGFR−, NGFR+, or IgG+ cells. The data are representative of four independent experiments (Fig. S5 A provides a statistical analysis).
AID expression is normal in IkL/L B cells
AID protein levels correlate with aicda transcript levels and switching efficiency (29–31). Thus, if aicda/AID expression were increased in Ikaros-deficient B cells, it could account for increased switching. Aicda expression was examined by RT-qPCR in WT and IkL/L B cells, either freshly isolated or stimulated for 48 h with LPS, LPS + IFN-γ, or LPS + IL-4. Stimulation of WT and IkL/L B cells induced aicda expression >48-fold, and there was no significant difference in aicda levels between IkL/L and WT, though expression was slightly higher in mutant cells (Fig. S6). Further, multiplex single-cell (SC)–RT-PCR analysis revealed similar percentages of aicda+ cells in both genotypes after LPS (WT, 71%; IkL/L, 67%) and LPS + IL-4 (WT, 65%; IkL/L, 62%) stimulation, suggesting similar levels of aicda transcripts per cell (Table S1; see Fig. 9, A and B). Thus, higher CSR efficiency in IkL/L cells is not caused by increased aicda expression.
Ikaros deficiency results in deregulated S region transcription
To determine if Ikaros controls CSR specificity by regulating S region transcription, we measured GLT levels for μ, γ3, γ1, γ2b, γ2a, and ε in WT and IkL/L B cells by RT-qPCR. Freshly isolated IkL/L cells exhibited three- to sixfold increases in γ3, γ2b, and γ2a GLT expression at the population level (Fig. 4 A). Further, SC-RT-PCR revealed that ∼22 and ∼6% of IkL/L cells expressed γ3 and γ2b GLTs, respectively, whereas virtually no WT cells expressed either GLT, suggesting that Ikaros represses the transcription of a subset of S regions in the absence of ex vivo stimulation (Fig. S7 and Table S1). After 48 h with LPS, LPS + IFN-γ, or LPS + IL-4, WT B cells specifically induced GLTs for the isotypes to which they switched (Fig. 1, A–C; and Fig. 4, B–D). IkL/L cells, however, expressed markedly different patterns of GLTs, and only derepressed GLTs correlated well with CSR. Strikingly, γ2b and γ2a GLTs were either overexpressed (>3.5-fold) or expressed ectopically (γ2b with LPS + IL-4, and γ2a with LPS and LPS + IL-4) in IkL/L cells, and this expression correlated with increased or ectopic CSR to these isotypes (Fig. 1, A–C; Fig. 4, B–D; and Fig. S8). Similarly, γ3 GLTs were expressed ectopically with LPS + IL-4 stimulation, which correlated with ectopic IgG3 switching (Fig. 1 C and Fig. 4 D). In contrast, μ GLTs were expressed similarly in both genotypes, and surprisingly, despite expressing WT levels of γ3 (with LPS and LPS + IFN-γ), γ1 and ε GLTs, IkL/L cells switched less efficiently to these isotypes (Fig. 1, A–C; Fig. 2; and Fig. 4, B–D). Collectively, these results suggest that Ikaros represses the transcription of Sγ3, Sγ2b, and Sγ2a in resting and stimulated cells, and that increased germline transcription of Sγ2b and Sγ2a (and Sγ3 after LPS + IL-4) in Ikaros-deficient cells skews isotype selection during CSR.
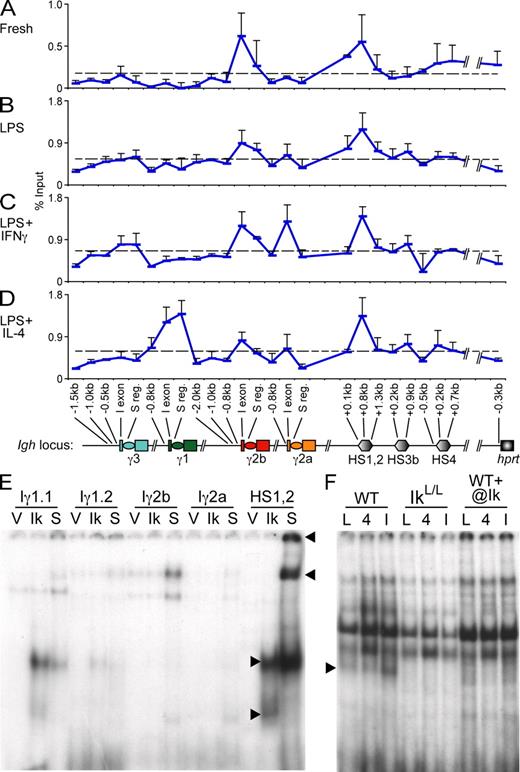
Deregulated expression of γ3, γ2b, and γ2a GLTs in IkL/L B cells. GLTs expressed by B220+ WT and IkL/L B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were analyzed by qPCR. GLTs were normalized to Igβ levels and all values are represented relative to those of unstimulated WT B cells. Graphs represent means plus SD of three independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05).
Deregulated expression of γ3, γ2b, and γ2a GLTs in IkL/L B cells. GLTs expressed by B220+ WT and IkL/L B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were analyzed by qPCR. GLTs were normalized to Igβ levels and all values are represented relative to those of unstimulated WT B cells. Graphs represent means plus SD of three independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05).
Derepression of S region transcription in IkL/L cells results in increased AID-dependent histone modifications
If enhanced Sγ2b and Sγ2a transcription skews switching in IkL/L cells, it would be reflected in increased AID accessibility and activity in these S regions, as AID-induced DSBs are necessary intermediates for CSR (32). To evaluate AID accessibility, we measured AcH4, which is induced in response to DSBs (33, 34) and is up-regulated in an AID-dependent manner at S regions during CSR (18). S region AcH4 levels were analyzed in both freshly isolated and stimulated (48 h) WT and IkL/L cells by chromatin immunoprecipitation (ChIP)–qPCR using anti-AcH4 antibodies. S region AcH4 levels were low in unstimulated B cells of both genotypes, as expected (Fig. 5 A). In WT cells, LPS and LPS + IFN-γ induced H4 acetylation at Sγ3 and Sγ2b, and LPS + IFN-γ also induced AcH4 at Sγ2a; LPS + IL-4 induced AcH4 exclusively at Sγ1 (Fig. 5, B–D). Thus, AcH4 levels mirrored GLT and isotype expression in WT cells, confirming that AID-dependent chromatin modifications correlate with transcription and switching (Figs. 1, 4, and 5) (18).
AcH4 correlates with transcriptional deregulation in IkL/L B cells. WT and IkL/L CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP with anti-AcH4 antibodies. Graphs represent the mean S region AcH4 enrichment indexes plus SD for three to four independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). AcH4 at Sγ2a was consistently higher in IkL/L versus WT samples after LPS + IFN-γ (Sγ2a IkL/L/WT ratio mean = 1.68; range = 1.21–2.13).
AcH4 correlates with transcriptional deregulation in IkL/L B cells. WT and IkL/L CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP with anti-AcH4 antibodies. Graphs represent the mean S region AcH4 enrichment indexes plus SD for three to four independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). AcH4 at Sγ2a was consistently higher in IkL/L versus WT samples after LPS + IFN-γ (Sγ2a IkL/L/WT ratio mean = 1.68; range = 1.21–2.13).
In IkL/L cells, however, histone H4 was hyperacetylated at ectopically and overtranscribed S regions. AcH4 levels were consistently higher at Sγ2b and Sγ2a in stimulated IkL/L cells compared with WT, correlating with increased transcription and switching to these isotypes (Fig. 1, A–C; Fig. 4, B–D; and Fig. 5, B–D). Similarly, ectopic AcH4 at Sγ3 in LPS + IL-4–stimulated IkL/L cells mirrored ectopic γ3 transcription and switching (Fig. 1 C, Fig. 4 D, and Fig. 5 D). In contrast, normal γ3 GLT expression in LPS- and LPS + IFN-γ–stimulated IkL/L cells induced 25–40% lower AcH4 levels at Sγ3 in comparison with WT, suggesting reduced AID-dependent activity (Fig. 4, B and C; and Fig. 5, B and C). Finally, similar γ1 GLT levels in LPS + IL-4–stimulated IkL/L and WT cells resulted in comparable induction of AcH4 at Sγ1 in both genotypes (Fig. 4 D and Fig. 5 D). Thus, increased transcription of Sγ2b and Sγ2a (and Sγ3 after LPS + IL-4) correlates with increased AID-dependent H4 acetylation, suggesting that abnormal germline transcription in IkL/L cells increases AID accessibility at Sγ2b and Sγ2a (and Sγ3) to skew CSR.
Ikaros does not repress germline transcription by regulating promoter–enhancer interactions in the Igh locus
Germline transcription of Sγ3, Sγ2b, and Sγ2a requires the Igh 3′ enhancer (14, 15), and Ikaros has been reported to regulate long-range promoter–enhancer looping at the β-globin locus (35). Therefore, we asked if Ikaros regulates germline transcription by controlling interactions between the 3′ enhancer and CH gene promoters. Igh locus promoter–enhancer interactions were assayed in WT and IkL/L B cells by chromosome conformation capture (3C) to measure interactions between distal chromosome regions (36). In these experiments, interacting chromatin was cross-linked with formaldehyde, digested with a restriction enzyme, and subjected to intramolecular DNA ligation and qPCR amplification of ligation products (Fig. S9 shows digestion and PCR controls). We examined interactions between a HindIII fragment containing the HS1,2 regulatory site and fragments covering the μ enhancer, downstream CH gene promoters, HS3a, and HS3b/4, as well as I regions not expected to participate in the regulation of germline transcription. Although HS1,2 was chosen as a reference point, patterns of promoter–enhancer cross-linking were similar, though weaker, using a 3′ enhancer fragment containing HS3b/4 (unpublished data). WT and IkL/L B cells were compared, either before or after 36 h of stimulation, when germline transcription was efficiently induced, but CSR was barely detectable by FACS (unpublished data). B cell–specific interactions were identified by comparison with PMA-stimulated T cells in which the Igh locus is silent.
In freshly isolated B cells, but not T cells, HS1,2 was frequently cross-linked to a fragment containing the μ enhancer, as previously reported (Fig. 6 A) (16). This interaction was reduced but still present in IkL/L B cells, suggesting that Ikaros contributes to μ–3′ enhancer interactions (Fig. 6 A). Cross-linking between HS1,2 and γ3 or γ2a in IkL/L B cells was similar to WT, despite ectopic γ3, γ2b, and γ2a transcription; only HS1,2–γ2b interactions were increased in mutant cells (Fig. 4 A and Fig. 6 A). Thus no clear correlation was detected between CH promoter–HS1,2 interactions and increased GLT expression in unstimulated IkL/L cells.
Ikaros does not regulate interactions between HS1,2 and CH gene promoters. CD43− WT and IkL/L B cells were analyzed by 3C, either (A) before or after 36 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4. CD4+ T cells stimulated for 36 h with PMA served as a negative control. The y axis represents the relative cross-linking frequency between a HindIII fragment covering HS1,2 and the rest of the locus. Points represent means plus SD of two independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). In A, HS1,2–HS3a and –HS3b/4 cross-linking frequencies in B cells, though similar between genotypes, were too high to be shown.
Ikaros does not regulate interactions between HS1,2 and CH gene promoters. CD43− WT and IkL/L B cells were analyzed by 3C, either (A) before or after 36 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4. CD4+ T cells stimulated for 36 h with PMA served as a negative control. The y axis represents the relative cross-linking frequency between a HindIII fragment covering HS1,2 and the rest of the locus. Points represent means plus SD of two independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). In A, HS1,2–HS3a and –HS3b/4 cross-linking frequencies in B cells, though similar between genotypes, were too high to be shown.
Next, we asked if Ikaros deficiency affects promoter–enhancer interactions after stimulation. In LPS-induced WT cells, there were B cell–specific HS1,2–γ3 and –γ2b cross-linking peaks, indicating HS1,2 interactions with these CH genes during transcription and switching (Fig. 1 A, Fig. 4 B, and Fig. 6 B). In IkL/L cells, HS1,2–γ2b and –γ2a cross-linking frequencies were similar to WT despite enhanced germline transcription of these genes (Fig. 4 B and Fig. 6 B); HS1,2–γ3 interactions were also similar between genotypes. After LPS + IFN-γ stimulation, WT cells exhibited significant B cell–specific cross-linking between HS1,2 and γ2a, as well as interaction peaks between HS1,2 and γ3 and γ2b, which correlated with transcription and switching for all three genes (Fig. 1 B, Fig. 4 C, and Fig. 6 C). HS1,2–γ2b and –γ2a interactions were nearly identical in IkL/L cells, despite sharply increased transcription and switching mainly to γ2a and, to a lesser extent, γ2b and γ3 (Fig. 1 B, Fig. 4 C, and Fig. 6 C). On the other hand, HS1,2–γ3 interactions were somewhat, but not significantly reduced in IkL/L cells (despite normal transcription), indicating a possible role for Ikaros in HS1,2–γ3 interactions after LPS + IFN-γ (Fig. 6 C). Finally, LPS + IL-4 induced consistently higher HS1,2–γ1 interactions in WT and IkL/L B cells versus WT T cells, which correlated with transcription and switching at γ1 in response to LPS + IL-4, as previously reported (Fig. 1 C, Fig. 4 D, and Fig. 6 D) (16). Importantly, ectopic transcription of γ3, γ2b, and γ2a in LPS + IL-4–stimulated IkL/L cells did not correlate with modified promoter–enhancer interactions, as there were no significant differences between WT and IkL/L cells, and only HS1,2–γ2a cross-linking was slightly increased in mutant populations (Fig. 4 D and Fig. 6 D). In summary, similar to previous reports, we found interactions between HS1,2 and CH genes that were transcribed and targeted for CSR in WT cells (16). Further, there was no clear correlation between CH promoter–HS1,2 interactions and defects in germline transcription. Therefore, deregulated S region transcription and aberrant switching in Ikaros-deficient B cells is not caused by altered interactions between the Igh 3′ enhancer and the promoters of specific CH genes.
Ikaros maintains repressive chromatin at the γ2b and γ2a genes
AcH3 is a hallmark of transcriptionally active chromatin (37) and correlates tightly with germline transcription (17, 18). Thus, we asked if Ikaros could regulate S region transcription by controlling H3 acetylation. AcH3 was examined at Cγ I exons and S regions in WT and IkL/L B cells by ChIP-qPCR, both before and after 48 h of stimulation. There were no significant differences in AcH3 levels between freshly isolated WT and IkL/L cells (Fig. 7 A). This suggests that AcH3 levels either do not contribute to GLT overexpression, or that our assay lacked the sensitivity required to detect differences in AcH3 associated with low levels of transcription (Fig. 4 A).
Increased AcH3 at γ3, γ2b, and γ2a in IkL/L B cells. WT and IkL/L CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP with anti-AcH3 antibodies. Graphs represent the mean I exon (I) and S region (S) AcH3 enrichment indexes plus SD for three independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). Sγ2b and Sγ2a AcH3 levels were consistently higher in IkL/L versus WT samples after LPS + IFN-γ (Sγ2b IkL/L/WT ratio mean = 2.68 [range = 1.82–3.47]; Sγ2a IkL/L/WT ratio mean = 2.41 [range = 1.45–3.87]).
Increased AcH3 at γ3, γ2b, and γ2a in IkL/L B cells. WT and IkL/L CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP with anti-AcH3 antibodies. Graphs represent the mean I exon (I) and S region (S) AcH3 enrichment indexes plus SD for three independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). Sγ2b and Sγ2a AcH3 levels were consistently higher in IkL/L versus WT samples after LPS + IFN-γ (Sγ2b IkL/L/WT ratio mean = 2.68 [range = 1.82–3.47]; Sγ2a IkL/L/WT ratio mean = 2.41 [range = 1.45–3.87]).
Next, we determined if AcH3 levels correlate with deregulated germline transcription in IkL/L B cells after stimulation. Stimulated WT cells induced H3 acetylation at the isotype genes they transcribed: I/Sγ3 and I/Sγ2b with LPS and LPS + IFN-γ, I/Sγ2a with LPS + IFN-γ, and I/Sγ1 with LPS + IL-4 (Fig. 4, B–D; and Fig. 7, B–D). Strikingly, in stimulated IkL/L cells, AcH3 levels at the γ2b and γ2a I exons and/or S regions were consistently higher than in WT cells, and correlated with increased and ectopic transcription (Fig. 4, B–D; and Fig. 7, B–D). Similarly, ectopic AcH3 at I/Sγ3 in LPS + IL-4–stimulated IkL/L cells correlated with ectopic transcription of that isotype (Fig. 4 D and Fig. 7 D). Normal levels of γ3 (LPS and LPS + IFN-γ) and γ1 (LPS + IL-4) transcription in IkL/L cells were associated with AcH3 levels at I/Sγ3 and I/Sγ1 that were equivalent to WT (Fig. 4, B–D; and Fig. 7, B–D). Importantly, AcH3 enrichment at the 3′ enhancer was also similar between WT and IkL/L cells (unpublished data). Thus, deregulated transcription and AcH3 are tightly correlated in IkL/L cells, indicating that Ikaros promotes a repressive chromatin state (e.g., histone H3 hypoacetylation) at CH promoters and genes during CSR.
Ikaros interacts directly with the Igh locus
We next asked if Ikaros could control chromatin remodeling and S region transcription through direct binding to Igh regulatory regions. We analyzed Ikaros binding to the Igh locus in freshly isolated and stimulated WT B cells by ChIP-qPCR using anti-Ikaros antibodies. We focused on regions in the Igh 3′ enhancer and Cγ gene promoters, I exons and S regions, that contained Ikaros consensus sites identified with the TFSEARCH algorithm (38).
Ikaros associated with the Igh locus in vivo in WT B cells (Fig. 8). In freshly isolated CD43− cells, Ikaros associated with Iγ2b and HS1,2 in the 3′ enhancer (Fig. 8 A). These associations were maintained after LPS stimulation (Fig. 8 B). When cells were stimulated with LPS + IFN-γ, Ikaros associated at low levels with Iγ3 and at higher levels with Iγ2b, Iγ2a, and HS1,2 (Fig. 8 C). Interestingly, this stimulation also induced transcription at Iγ3, Iγ2b, and Iγ2a, indicating that Ikaros associates strongly with some I exons upon their transcriptional activation (Fig. 8 C). Similarly, after LPS + IL-4 stimulation, Ikaros associated with the transcribed Iγ1 exon (Fig. 4 D and Fig. 8 D). In addition, there was moderate Ikaros association with Iγ2b, whereas the HS1,2 peak remained high (Fig. 8 D). Thus, Ikaros strongly associates with HS1,2 regardless of stimulation, with Iγ2b in all conditions but most strongly when Iγ2b is normally transcribed in WT, and with Iγ3, Iγ1, and Iγ2a when they are transcriptionally induced upon stimulation.
Ikaros associates with CH promoters and HS1,2 of the 3′ enhancer. WT CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP-qPCR with anti-Ikaros antibodies. Points represent mean Ikaros enrichment, measured as percentage input, plus SD from two to three independent experiments. Approximate amplicon positions relative to transcription start sites or the 5′ ends of HS sites are indicated. Dashed lines represent the mean percentage input for each condition and are defined as the threshold of specific Ikaros binding. (E and F) EMSAs were performed on putative Ikaros binding sites from sequences identified by ChIP (A–D). (E) Nuclear extracts from Cos cells transduced with an empty vector (V) or one encoding the Ik1 isoform (Ik) were incubated with the indicated probes (Table S2). Two putative Iγ1 Ikaros binding sites were tested. (F) The HS1,2 probe was incubated with nuclear extracts from WT or IkL/L B cells stimulated for 36 h with LPS (L), LPS + IL-4 (4), or LPS + IFN-γ (I). Specificity was verified by supershift with anti-Ikaros antibodies. Arrowheads indicate Ikaros complexes. Data in E and F are representative of two independent experiments.
Ikaros associates with CH promoters and HS1,2 of the 3′ enhancer. WT CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP-qPCR with anti-Ikaros antibodies. Points represent mean Ikaros enrichment, measured as percentage input, plus SD from two to three independent experiments. Approximate amplicon positions relative to transcription start sites or the 5′ ends of HS sites are indicated. Dashed lines represent the mean percentage input for each condition and are defined as the threshold of specific Ikaros binding. (E and F) EMSAs were performed on putative Ikaros binding sites from sequences identified by ChIP (A–D). (E) Nuclear extracts from Cos cells transduced with an empty vector (V) or one encoding the Ik1 isoform (Ik) were incubated with the indicated probes (Table S2). Two putative Iγ1 Ikaros binding sites were tested. (F) The HS1,2 probe was incubated with nuclear extracts from WT or IkL/L B cells stimulated for 36 h with LPS (L), LPS + IL-4 (4), or LPS + IFN-γ (I). Specificity was verified by supershift with anti-Ikaros antibodies. Arrowheads indicate Ikaros complexes. Data in E and F are representative of two independent experiments.
To determine if Ikaros binds directly to any of these sequences, we performed EMSA. Exogenous Ikaros, confirmed by supershift, bound strongly to a probe containing a putative Ikaros binding site from HS1,2 (Fig. 8 E). Interestingly, endogenous Ikaros complexes also bound this probe, indicating that Ikaros can directly bind HS1,2 (Fig. 8 F). In contrast, Ikaros did not bind to putative sites from Iγ2b or Iγ2a, and supershift with Ikaros-specific antibodies revealed only low affinity binding to sites in Iγ1, in keeping with a previous report (Fig. 8 E) (39). Collectively, these results indicate that Ikaros regulates germline transcription by binding directly to HS1,2, but suggest that its association with Iγ regions is indirect, possibly through enhancer–promoter interactions.
Ikaros controls transcriptional competition between S regions
The data described in the previous sections demonstrate that increased and ectopic transcription of γ2b and γ2a (and γ3 with LPS + IL-4) in IkL/L cells correlates with enhanced/ectopic switching to these isotypes. Paradoxically, CSR to IgG3, IgG1, and IgE is decreased in IkL/L cells despite normal GLT levels (Figs. 1, 2, and 4). We hypothesized that this discrepancy was caused by increased/ectopic transcription of Sγ2b/Sγ2a (and Sγ3) outcompeting transcription of other S regions for AID-mediated CSR. This model would require that (a) S regions are cotranscribed in individual cells and (b) transcription of Sγ2b/Sγ2a (and Sγ3 with LPS + IL-4) is selectively increased relative to that of other S regions. To test this hypothesis, we examined the expression of aicda, and γ3 and γ2b GLTs (and actb as a positive control) by SC-RT-PCR in LPS- and LPS + IL-4–stimulated (48 h) IgM+ cells (Fig. 9 and Table S1). Importantly, because interallelic CSR occurs efficiently in WT cells (40, 41), SC-RT-PCR was performed without regard for allele. In LPS-stimulated WT cells, 97.1% of switching competent cells (e.g., aicda+) expressed GLTs for γ2b (90.1%) and/or γ3 (78.5%), which correlated with switching to these two isotypes (Fig. 1 A; and Fig. 9, B and C). Interestingly, 91.7% of γ3 GLT+ cells also transcribed γ2b, whereas 15–20% of WT cells switched to IgG3 after 4 d, indicating that the majority of WT cells cotranscribe S regions and that cells cotranscribing γ2b and γ3 can still choose γ3 for CSR (Fig. 3 and Fig. 9 C). In LPS + IL-4–stimulated cells, the high efficiency of IgG1 switching suggests that γ1 GLTs are expressed in almost all cells (Fig. 1 C); γ1 PCRs were not performed because of the added complexity of performing a fifth PCR in multiplex. Interestingly, 53.9% of aicda+ WT cells expressed GLTs for γ3 (34.3%) and/or γ2b (29.8%), suggesting competition with Sγ1 in a subset of WT cells (Fig. 9, B and C). However, by combining SC-RT-PCR and RT-qPCR data (Fig. 4, B and D), we calculated that the levels of Sγ3 and Sγ2b transcripts per γ2b+/γ3+ cell were four times lower in LPS + IL-4–stimulated WT cells than in cells stimulated with LPS (Fig. 9 D). These low levels appear insufficient to induce CSR to γ2b and γ2a (Fig. 1 C). Thus, S regions are cotranscribed in WT cells and relative S region transcription rates per cell correlate with CSR, indicating that S region competition can occur under normal conditions and may regulate isotype choice.
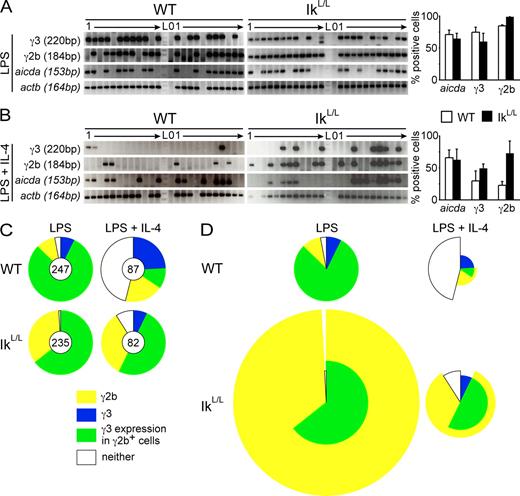
Increased S region competition at the SC level for CSR in IkL/L B cells. SC-RT-PCR was performed on IgM+ WT and IkL/L cells after 48 h of stimulation and two divisions (assessed by CFSE dilution). (A and B) Representative data from cohorts of (A) LPS- and (B) LPS + IL-4–stimulated cells are shown. Only actb+ wells were counted (positive control for sorting). Bar graphs represent mean percentages plus SD of cells positive for Aicda, or γ3 or γ2b GLTs, from two independent experiments (0, no cells; 1, one cell; raw data are shown in Table S1). (C) Pie charts represent mean percentages of aicda+ cells expressing GLTs for γ3, γ2b, both, or neither. The numbers of aicda+ cells analyzed over two experiments are indicated (chart centers). (D) Data from C was integrated with GLT expression levels measured by RT-qPCR (Fig. 4, B and D) to give a comprehensive view of γ2b and γ3 GLT expression in individual cells. Pie chart wedges represent the percentages of aicda+ B cells that express the indicated GLTs. Wedge radii are defined such that the wedge area corresponds to total GLT expression levels; γ3 and γ2b GLT levels were normalized to LPS-stimulated WT B cells, for which the radius was set to 1. Thus, the percentage of degrees out of 360 taken by each wedge corresponds to the percentage of expressing cells, and the wedge area corresponds to the integrated per cell expression level.
Increased S region competition at the SC level for CSR in IkL/L B cells. SC-RT-PCR was performed on IgM+ WT and IkL/L cells after 48 h of stimulation and two divisions (assessed by CFSE dilution). (A and B) Representative data from cohorts of (A) LPS- and (B) LPS + IL-4–stimulated cells are shown. Only actb+ wells were counted (positive control for sorting). Bar graphs represent mean percentages plus SD of cells positive for Aicda, or γ3 or γ2b GLTs, from two independent experiments (0, no cells; 1, one cell; raw data are shown in Table S1). (C) Pie charts represent mean percentages of aicda+ cells expressing GLTs for γ3, γ2b, both, or neither. The numbers of aicda+ cells analyzed over two experiments are indicated (chart centers). (D) Data from C was integrated with GLT expression levels measured by RT-qPCR (Fig. 4, B and D) to give a comprehensive view of γ2b and γ3 GLT expression in individual cells. Pie chart wedges represent the percentages of aicda+ B cells that express the indicated GLTs. Wedge radii are defined such that the wedge area corresponds to total GLT expression levels; γ3 and γ2b GLT levels were normalized to LPS-stimulated WT B cells, for which the radius was set to 1. Thus, the percentage of degrees out of 360 taken by each wedge corresponds to the percentage of expressing cells, and the wedge area corresponds to the integrated per cell expression level.
S region competition is increased in Ikaros-deficient B cells. After LPS stimulation, virtually every aicda+ IkL/L cell expressed γ2b GLTs (99.1% γ2b GLT+ cells), whereas the frequency of cells expressing γ3 GLTs was slightly lower than in WT (64.3% of IkL/L cells, down 14%; Fig. 9 A). Strikingly, every γ3 GLT+ IkL/L cell also expressed γ2b GLTs, putting Sγ3 in constant competition with Sγ2b (Fig. 9 C). Unlike in WT cells, however, Sγ3/Sγ2b cotranscription in IkL/L cells was associated with strongly skewed CSR to IgG2b (Fig. 1 A). To explain this difference, we examined per cell transcript levels and found that γ2b GLTs were expressed at 8.5 times higher levels per γ2b+ cell in IkL/L versus WT cultures, whereas γ3 GLT levels were similar on a per cell basis (up 1.6-fold in IkL/L vs. WT; Fig. 9 D). Thus, although both WT and IkL/L B cells cotranscribe Sγ3/Sγ2b, increased Sγ2b transcription in IkL/L cells coincides with IgG2b-skewed switching, indicating that higher Sγ2b transcription outcompetes Sγ3 transcription for AID-mediated CSR.
Similarly increased competition was found after LPS + IL-4 stimulation. 91.0% of aicda+ IkL/L cells expressed γ3 (57.2%, up 22.9%) and/or γ2b (83.8%, up 50.8%) GLTs, indicating that these S regions directly compete with Sγ1 for CSR in most cells (Fig. 9, B and C). In addition, LPS + IL-4–stimulated IkL/L cells expressed γ3 and γ2b GLTs at per cell levels that were three- to fourfold higher than those found in similarly stimulated WT cells, and nearly equivalent to those observed in LPS-stimulated WT cells (Fig. 9 D). This correlated with ectopic CSR to these isotypes and reduced CSR to IgG1 (Fig. 1 C). Thus, in LPS + IL-4–stimulated IkL/L cells, high levels of Sγ2b/Sγ3 transcription successfully compete with transcription of Sγ1 for CSR.
Collectively, these observations demonstrate that the majority of B cells cotranscribe S regions, and that per cell transcription rates correlate with isotype selection. Further, our results strongly support the hypothesis that increased transcription of Sγ2b and Sγ2a (and Sγ3 with LPS + IL-4) in IkL/L cells outcompetes transcription of other S regions for CSR and thus skews isotype selection.
DISCUSSION
In this study, we identify Ikaros as a critical regulator of Ig isotype selection in B cells during CSR. Ikaros deficiency results in increased and ectopic switching to IgG2b and IgG2a (and ectopic switching to IgG3 with LPS + IL-4) and enhanced overall switching, but reduced switching to IgG3 (with LPS and LPS + IFN-γ), IgG1, IgE, and IgA. This function is independent of Ikaros’s role in B cell development, as inhibition of Ikaros in mature WT cells skews isotype selection to IgG2b and IgG2a, whereas retroviral expression of Ikaros in IkL/L B cells rescues CSR. Further, in direct correlation with their CSR phenotype, IkL/L cells exhibit sharp increases in transcription and AcH4, an AID-dependent histone mark, at Sγ2b and Sγ2a (and Sγ3 with LPS + IL-4) but not other S regions. Ikaros directly binds the Igh locus in vivo and is required to prevent hyperacetylation of histone H3 at the γ2b and γ2a (and γ3 with LPS + IL-4) genes upon B cell activation. Thus, Ikaros maintains repressive chromatin at γ2b and γ2a (and γ3 with LPS + IL-4), and suppresses transcription, AID accessibility, and switching to these isotypes. These results identify Ikaros as the first factor that controls the range of isotypes targeted by CSR.
Our results also reveal that the choice to undergo CSR to specific CH genes is controlled by the balance of germline transcription across different S regions within individual cells. We have shown that the majority of aicda+ LPS-stimulated WT cells coexpress GLTs for γ2b and γ3, suggesting competition between these two isotypes for switching. Further, selective increases in γ2b GLTs in IkL/L cells, where γ3 is always cotranscribed with γ2b, correlate with increased CSR to IgG2b and decreased CSR to IgG3. Thus, repression of Sγ2b transcription by Ikaros is required to maintain a balance between Sγ2b and Sγ3 transcription, allowing for normal switching to both isotypes. Similarly, Ikaros fine tunes Sγ2b and Sγ2a transcription in response to LPS + IFN-γ, to allow switching to IgG3. Therefore, in addition to suppressing promiscuous switching to IgG3/2b/2a in response to LPS + IL-4, and IgG2a in response to LPS, Ikaros limits transcription across Sγ2b and Sγ2a so that germline transcription of other S regions (i.e., Sγ3 and Sα) can effectively compete for switching. These results indicate that the isotype fate of individual cells during CSR is determined by the relative levels of germline transcription across different S regions. Considering that single human B cells have also been reported to transcribe multiple S regions simultaneously (42), this is likely a general mechanism, across species, for isotype selection during CSR.
A question arising from our work is how Ikaros suppresses transcription at γ3, γ2b, and γ2a. Interestingly, we have found that histone H3 is hyperacetylated at these genes in IkL/L cells. Although it is possible that these increased AcH3 levels result from transcription-coupled processes, it is attractive to speculate that Ikaros regulates germline transcription by maintaining a repressive chromatin structure. In this respect, it is well documented that histone acetylation contributes to transcription by opening chromatin and/or providing binding platforms for regulatory factors (43, 44).
Considering that H3 is hyperacetylated in IkL/L cells and that Ikaros interacts with the Sin3 and NuRD histone deacetylase (HDAC) complexes (45–48), we expected Ikaros to recruit these complexes to the Igh locus to regulate germline transcription. However, we have been unable to find reduced Igh locus occupancy for Sin3a, Mi-2β (a core member of NuRD), HDAC1, or HDAC2 in IkL/L versus WT cells by ChIP-qPCR or double cross-linking (DC)–ChIP-qPCR (Fig. S10). Furthermore, shRNA knockdown of Mi-2β in WT B cells neither increased γ2b transcription nor altered switching specificity (Fig. S11). These data indicate that Ikaros does not suppress H3 acetylation and germline transcription by recruiting the NuRD and Sin3 HDAC complexes to the Igh locus.
Our results indicate that Ikaros may directly regulate germline transcription through the HS1,2 region of the Igh 3′ enhancer. Ikaros associates with HS1,2 in all conditions tested, and Ikaros proteins bind directly to a sequence in this region in vitro. In contrast, despite the association of Ikaros with the Iγ1, Iγ2b, and Iγ2a regions by ChIP, we were unable to identify sequences in these regions that could be strongly and directly bound by Ikaros. Although we cannot rule out binding to noncanonical sites, these data suggest that Ikaros binds directly to HS1,2 and interacts indirectly with Iγ1, Iγ2b, and Iγ2a. Our 3C data, as well as those from Wuerffel et al. (16), support this interpretation, as HS1,2 interacts with fragments containing the γ3, γ1, γ2b, and γ2a I promoters/exons. Interestingly, HS1,2–γ1 interactions increase when this gene is transcribed (e.g., LPS + IL-4; Figs. 4 and 6) (16), which correlates with the appearance of an Ikaros binding peak by ChIP. Similarly, Ikaros associates with Iγ2a only when cells are stimulated with LPS + IFN-γ, which induces both γ2a GLTs and increased HS1,2-Iγ2a interactions. Thus, we propose that Ikaros interacts indirectly with and regulates CH gene promoters through HS1,2.
The idea that Ikaros mediates repression through the HS1,2 regulatory region of the 3′ enhancer is bolstered by the observation that 3′ enhancer disruption most drastically affects transcription of and CSR to γ3, γ2b, and γ2a, the CH genes most strongly affected by Ikaros deficiency (14, 15). One issue with this model is that HS1,2 deletion does not affect CSR (14), suggesting that HS1,2 may be not be functionally important for CSR and/or that it functions independently of Ikaros. This explanation, however, is probably too simplistic. In another complex locus, the β-globin locus, deletions of individual regulatory elements result in little to no phenotype, whereas combined deletions can drastically inhibit globin gene expression, indicating that the loss of individual elements can be compensated for by redundant functions in other regions (49). Thus, compensation by other 3′ enhancer elements, which have many similar transcription factor motifs (50), could explain the lack of CSR phenotype in HS1,2−/− cells. In addition, HS1,2 can strongly synergize with other HS fragments to activate transcription (51), and the core region HS1,2 is highly conserved between mouse, rat, rabbit, and human (52), indicating that HS1,2 is likely to play important roles in CSR.
How then, can HS1,2−/− and IkL/L B cell CSR phenotypes be reconciled? An obvious possibility is that Ikaros could repress germline transcription and histone acetylation by antagonizing positive regulatory functions at this element. In this case, HS1,2 deletion would abolish both negative regulation by Ikaros and positive regulation by unknown factors, resulting in a neutral phenotype. In contrast, Ikaros deficiency would allow increased activity by positively acting factors, resulting in a significant phenotype. Interestingly, Ikaros has been shown to compete with positive regulators to modulate the transcription of target genes in other settings (22, 53, 54). In this respect, our EMSA assay indicates that multiple factors can bind at or near the HS1,2–Ikaros binding site (Fig. 8 F). Thus, it will be important to determine if Ikaros competes with other factors for binding to the HS1,2 regulatory element to fine tune germline transcription.
Finally, the efficiency of CSR is controlled by several mechanisms, including cell proliferation, AID expression, subcellular localization and posttranslational modifications, DSB formation, DNA repair, S region synapsis, and end joining (1). Our results suggest that S region transcription rates in WT cells also limit switching efficiency. This concept is supported by (a) the low rates of CSR compared with switching competency in WT cells (∼30% of cells switched after 4 d with LPS, whereas 70% were aicda+GLT+ after 48 h; Fig. 3, Fig. 9A, and Table S1), (b) the similar levels of CSR-competent cells in IkL/L and WT cultures (67% of IkL/L cells were aicda+GLT+ after 48 h; Fig. 9 A and Table S1), and (c) the direct correlation between increased per cell GLT levels, AID accessibility, and switching frequency in IkL/L cells (∼50% switched after 4 d; Fig. 5 B, Fig. 9 D, and Fig. S5 B). Why it would be advantageous to limit S region transcription rates and CSR efficiency under normal conditions is not clear. This mechanism could result in more flexible humoral immune responses in vivo, as slowing the rate of CSR may prevent B cells from switching en masse soon after activation, thus allowing isotype selection to be modified over time. In addition, reduced S region transcription, and therefore DSB formation, would decrease the probability of unrepaired DSBs, which can to lead to oncogenic translocations (55).
In summary, our results reveal transcriptional competition between constant region genes as a central and general mechanism for isotype specification during CSR. We have shown that Ikaros is a master regulator of this competition.
MATERIALS AND METHODS
Mice.
The IkL/L mouse line was previously described (21). Mice backcrossed >10 times onto the C57BL/6 genetic background were analyzed at 5–9 wk of age. All animal work was performed under protocols approved by the Direction des Services Vétérinaires du Bas-Rhin, France (authorization no. 67-343).
Cell culture.
Spleen cells were used for all experiments. B cells (B220+, IgM+CD43−, or CD43−IgG/A− cells; >98% purity) were sorted on a FACSVantage SE option DIVA (BD) or enriched by depletion of CD43+ cells with MACS beads (>90% purity), and CD4+ T cells were isolated with anti-CD4 MACS beads (>90% purity; Miltenyi Biotech). Cells were labeled with 5 µg/ml CFSE (10 min at 37°C; Sigma-Aldrich) and were maintained at 1.2 × 106 cells/ml in complete medium (RPMI 1640, 10% FCS, 25 mM Hepes, 1 mM sodium pyruvate, 2 mM l-glutamine, 1× nonessential amino acids, 5 × 10−5 M 2-mercaptoethanol, and 1% antibiotics) with 25 µg/ml LPS (serotype 0111:B4 from Escherichia coli; Sigma-Aldrich), 5 ng/ml IL-4 (Sigma-Aldrich), 100 ng/ml IFN-γ (PeproTech), 3 ng/ml TGF-β (R&D Systems), 5 ng/ml IL-5 (BD), and 0.5 ng/ml PMA (Sigma-Aldrich).
Flow cytometry.
Reagents included anti-B220–FITC (RA3-6B2), anti–mouse IgG2b–biotin (RMG2b-1; BioLegend), anti–mouse IgA–PE (SouthernBiotech), F(ab′)2 goat anti–mouse IgM–Cy5 and streptavidin-Cy5 or -PE (Jackson ImmunoResearch Laboratories), anti–mouse IgG3–biotin (R40-82), anti–mouse IgG1–biotin (A85-1), anti–mouse IgG2ab–biotin (Igh-b; 5.7) and anti-CD43–PE (BD), goat anti–mouse IgG (H+L)–biotin (Invitrogen), anti-NGFR–Cy5 (8737; provided by W. Pear, University of Pennsylvania, Philadelphia, PA), and 7-aminoactinomycin D (Sigma-Aldrich). Cells were analyzed with a FACSCalibur (BD) and FlowJo software (Tree Star, Inc.).
RT-qPCR.
RNA was isolated with RNeasy kits (QIAGEN). 350 ng RNA was reverse transcribed in 20 µl (50 mM Tris-HCl [pH 8.3], 75 mM KCl, 3 mM MgCl2, 10 mM dithiothreitol, 500 µM dNTP, 10 U recombinant RNasin ribonuclease inhibitor [Promega], 0.5 µM oligo d(T) [New England Biolabs, Inc.], and 40 U SuperScript II RT [Invitrogen]). SYBR Green JumpStart Taq ReadyMix (Sigma-Aldrich) was used for all qPCRs. Approximately 3 ng of cDNA was run (in triplicate) on a LightCycler 480 (96-well plate format) and analyzed with LightCycler 480 basic software (Roche). Transcript quantities for each gene were calculated relative to standard curves and normalized to Igβ transcripts. Gene of interest/Igβ ratios for each condition were averaged across experiments and normalized to the indicated WT condition, to give relative expression versus WT or versus unstimulated WT, as noted. Table S2 lists oligonucleotides for all experiments.
SC-RT-PCR.
Two rounds of PCR were performed with fully nested primers, as previously described (54). Only actb+ wells were analyzed (>90%).
Retrovirus production and transduction.
MigR1-NGFR (provided by W. Pear) is an murine stem cell virus–based retrovirus (56). Ik1 cDNA was amplified from mouse spleen cDNA by PCR (bp 271–1,818; available from GenBank/EMBL/DDBJ under accession no. NM_001025597). Ik1* cDNA was obtained by site-directed mutagenesis to delete exon 2–derived Ik1 sequences (bp 310–430). Ik6 cDNA was obtained by fusing PCR fragments for exons 1 and 2 (bp 271–429) and exon 7 (bp 1,117–1,818) of Ik1. cDNAs were cloned into the MigR1-NGFR vector and were designated Ik1-NGFR, Ik1*-NGFR, and Ik6-NGFR. pQsupR-Mi2 (Mi-2β shRNA; provided by S. Smale, University of California, Los Angeles, Los Angeles, CA) and pQsupR (mock) vectors were previously described (57). Vectors were transfected into Eco-Phoenix packaging cells (provided by G. Nolan, Stanford University, Stanford, CA) to produce high-titer retroviral supernatants. 3.5 × 105 CFSE-stained cells (1.2 × 106 cells/ml) were cultured in LPS for 24 h, transduced with retroviral supernatant (25% total vol; 4 µg/ml polybrene; 25 µg/ml LPS), and centrifuged for 90 min at 2,600 rpm. Medium was replaced after 12 h and cells were analyzed 60 h later.
ChIP.
The ChIP protocol was adapted from Wang et al. (18) and Millipore (http://www.millipore.com/techpublications/tech1/mcproto407). In brief, 1.8 × 107 B cells were cross-linked at 37°C for 10 min in 5 ml PBS/0.5% BSA/1% ultra-pure formaldehyde (Electron Microscopy Sciences). After quenching with 0.125 M glycine and a cold PBS wash (with 1× protease inhibitor cocktail [PIC]; Roche), cells were lysed in 5 ml of Triton X lysis buffer for 10 min on ice (1% Triton X-100, 50 mM MgCl2, 100 mM Tris-HCl [pH 7.1], 11% sucrose, 1× PIC). Nuclei were pelleted at 2,000 rpm for 10 min (4°C) and were lysed in 500 µl of SDS lysis buffer for 10 min on ice (1% SDS, 50 mM Tris-HCl, 10 mM EDTA, 1× PIC). Chromatin was sonicated to 500-1,000 bp using a Bioruptor 200 (Diagenode), and sonication efficiency was checked. After 2× dilution in ChIP buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCl [pH 8.1], 167 mM NaCl), chromatin was precleared by rotating for 2 h at 4°C with 80 µl 50% protein A slurry (0.2 mg/ml sheared salmon sperm DNA, 0.5 mg/ml BSA, 50% protein A; GE Healthcare). 1.3 × 106 cell equivalents were saved as input. 3.9 × 106 cell equivalents were incubated overnight with 2 µg anti-AcH3 (Millipore), 2 µg anti-AcH4 (Millipore), 2.5 µg anti-Ikaros (rabbit anti–mouse produced in house), anti–Mi-2β (5 µl of rabbit serum; gifts from P. Wade [National Institute of Environmental Health Sciences, Research Triangle Park, NC] and S. Smale), 5 µg anti-Sin3a (Santa Cruz Biotechnology, Inc.), 5 µg anti-HDAC1 (Abcam), 5 µg anti-HDAC2 (Abcam), and control antibodies (Bethyl Laboratories). ChIPs were recovered with 65 µl 50% protein A slurry for 5 h and processed according to the Millipore protocol. ChIP and input DNA was dissolved in 140 µl TE buffer (10 mM Tris-HCl, 1 mM EDTA [ph 8]), and inputs were diluted 10 times. 2-µl aliquots were analyzed (in duplicate) by qPCR. Calculations were as follows: percentage input (%I) = 100 × ([antibody bound] − [IgG bound])/(input*10); and histone acetylation index = (%IIgh sequence)/(%IHprt promoter). For DC-ChIP, cells were first fixed with disuccinimidyl glutarate (58).
3C.
3C was performed as previously described (59) with some modifications. In brief, 11 × 106 cells were fixed for 10 min in 10 ml 1× PBS/0.5% BSA/1.5% formaldehyde (Electron Microscopy Sciences) at room temperature with tumbling (10 rpm). Fixation was quenched with 0.125 M glycine. Cells were washed one time in cold PBS/1× PIC and lysed for 10 min on ice in 1 ml of lysis buffer (10 mM Tris [pH 8], 10 mM NaCl, 0.2% NP-40, 1× PIC). After centrifugation, cells were resuspended in 537 µl of digestion buffer (60 µl of 10× buffer R [Fermentas], 1× PIC). SDS was added to 0.3% and samples were rotated for 1 h at 37°C (99 rpm). Triton X was added to 1.8% to sequester SDS, bringing the total volume to 600 µl. Samples were rotated (99 rpm) for 1 h at 37°C before overnight digestion with 500 U HindIII (50 rpm; New England Biolabs, Inc.). HindIII was inactivated by adding SDS to 1.75% and incubating for 20 min at 65°C. 106 cell equivalents were removed, de–cross-linked, and analyzed by qPCR to monitor digestion efficiency (routinely 80–90%; Fig. S9 A). 107 cell equivalents were diluted to 8 ml in 1× ligation buffer (New England Biolabs), 1% Triton X, and 1× PIC, and SDS was sequestered by rocking for 1 h at 37°C. 4,000 NEB U of ligase (New England Biolabs, Inc.) were added and samples were rocked for 10 min at 4°C before incubation for 6 h at 16°C, followed by 30 min at room temperature. Samples were de–cross-linked overnight at 65°C with 300 µg protease K and were retreated with 300 µg protease K for 1 h at 45°C. DNA was extracted using ultra-pure phenol/chloroform/isoamyl alcohol (25:24:1; Invitrogen), washed two times with chloroform, ethanol precipitated with 2.5 M ammonium acetate, and washed four times with 75% ethanol. After suspension in TE, relative DNA concentrations were calculated by qPCR (ChIP primers “HS4 −0.5 kb”).
Ligation products were analyzed by qPCR in duplicate or triplicate using ∼200 ng DNA. Absolute quantities were calculated using a template consisting of equimolar ratios of all possible ligation products. The template was constructed as previously described (60) with PCR fragments from the Igh and Gapd loci and diluted in genomic DNA. qPCR specificity was confirmed with negative controls (without fixation, digestion, or ligation), melting point analysis, digestion of PCR products with HindIII, and migration of PCR and digestion products on 2% agarose gels (Fig. S9 B). Relative cross-linking frequency was calculated as follows: cross-linking frequency = ([HS1,2 − fragment X]/[HS4 −0.5 kb loading control])/([Gapd 3′ − Gapd 5′]/[HS4 −0.5 kb]).
EMSA.
Nuclear extracts (NEs) and EMSAs were prepared as previously described (61). 2.5 µg NE from Cos cells transfected with mock cDNA or cDNA coding for Ik1 proteins or 4 µg NE from CSR-induced B cells was used.
Online supplemental material.
Table S1 summarizes raw data from SC-RT-PCR experiments. Table S2 lists oligonucleotides. Fig. S1 shows statistical analysis of WT and IkL/L CSR frequencies after LPS, LPS + IL-4, LPS + IFN-γ, and LPS + IL-5 + TGF-β stimulation. Fig. S2 quantifies Ig isotype production by WT and IkL/L B cells using ELISA. Fig. S3 analyzes WT and IkL/L CSR frequency as a function of proliferation. Fig. S4 examines CSR in whole and bona fide unswitched WT and IkL/L B cells. Fig. S5 shows the effect of retroviral expression of dominant-negative Ikaros on CSR in WT cells, and full-length and exon 2–deleted Ikaros isoforms on CSR in IkL/L B cells. Fig. S6 shows RT-qPCR analysis of aicda expression in stimulated WT and IkL/L cells. Fig. S7 shows SC-RT-PCR analysis of aicda, γ3 GLT, and γ2b GLT expression in freshly isolated IgM+CD43− B cells and CD4+CD3+ T cells. Fig. S8 shows RT-qPCR analysis of μ, γ3, γ2b, and γ2a GLT expression in WT and IkL/L B cells after 48 h with LPS and one or three divisions. Fig. S9 shows digestion efficiency and PCR specificity controls for 3C experiments. Fig. S10 shows ChIP-qPCR and DC-ChIP-qPCR for Mi-2β, Sin3a, HDAC1, and HDAC2 at the Igh locus in WT and IkL/L cells. Fig. S11 shows the effect of Mi-2β shRNA knockdown on CSR in WT B cells.
Acknowledgments
We thank G. Nolan; W. Pear; S. Smale and P. Wade for critical reagents; J. Barths, C. Ebel, and P. Marchal for FACS; P. Marchal for technical assistance; and M. Gendron for animal husbandry.
M. Sellars received a predoctoral fellowship from La Fondation pour la Recherche Médicale. B. Reina-San-Martin is an AVENIR–Institut National de la Santé et de la Recherche Médicale (INSERM) young investigator. The work was supported by INSERM, the Centre National pour la Recherche Scientifique, and the Hôpital Universitaire de Strasbourg, as well as by a grant from L’Agence Nationale de la Recherche (to B. Reina-San-Martin, S. Chan, and P. Kastner).
The authors have no conflicting financial interests.
References
Abbreviations used: 3C, chromosome conformation capture; AcH3, histone H3 acetylation; AID, activation-induced cytidine deaminase; ChIP, chromatin immunoprecipitation; CSR, class switch recombination; DSB, double-stranded DNA break; GLT, germline transcript; HDAC, histone deacetylase; HS, hypersensitive; I, intronic; NGFR, nerve growth factor receptor; qPCR, quantitative PCR; S, switch; SC, single cell.
Author notes
M. Sellars’s present address is Molecular Pathogenesis Program, The Helen L. and Martin S. Kimmel Center for Biology and Medicine at the Skirball Institute for Biomolecular Medicine, New York University School of Medicine, New York, NY 10016.







![Figure 7. Increased AcH3 at γ3, γ2b, and γ2a in IkL/L B cells. WT and IkL/L CD43− B cells that were (A) freshly isolated or stimulated for 48 h with (B) LPS, (C) LPS + IFN-γ, or (D) LPS + IL-4 were subjected to ChIP with anti-AcH3 antibodies. Graphs represent the mean I exon (I) and S region (S) AcH3 enrichment indexes plus SD for three independent experiments. Significance was determined by a two-tailed t test assuming unequal variance (***, P < 0.005; **, P < 0.02; *, P < 0.05). Sγ2b and Sγ2a AcH3 levels were consistently higher in IkL/L versus WT samples after LPS + IFN-γ (Sγ2b IkL/L/WT ratio mean = 2.68 [range = 1.82–3.47]; Sγ2a IkL/L/WT ratio mean = 2.41 [range = 1.45–3.87]).](https://cdn.rupress.org/rup/content_public/journal/jem/206/5/10.1084_jem.20082311/6/m_jem_20082311_rgb_fig7.jpeg?Expires=1779236353&Signature=IGhCTosMsU0y3ynrR5V71nYAY3VgVklNjL3IlCq8O~~TANuuy3Lxaqu0MIxMOHEDkGj~T1Kr972sOkXjidpkq-abJXajtVQoIdnHFEbX6IZDeDF7idWaXDx53VrjZGBSL0VokURazUiflRbTDYzYgA8rcufA2qcHHQtjg~D8QcEpAydgP~aJmLrQ5Vm1hMDPyOA0BWd2aocf0~EH~V2cWUhC47E0RIxNdU8GR9H75sbBbDtkt7lP6pg1ClCL4DmYTv4WPRz~QE9jq66px~yyPadOcbbw~ttNQ9xiT-9UVNamqYhdNR20qSml6jk04jxFhHxThBlzFiPfoLwlErgw5g__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)