Legionella pneumophila is an intracellular bacterial pathogen that modulates membrane trafficking to survive and proliferate within host cells. After phagocytosis, the L. pneumophila–containing vacuole evades the endocytic pathway by excluding the host GTPase Rab5, a crucial regulator of phagosomal maturation. In this study, we show that the evolutionarily conserved lysine residue K134 of Rab5 undergoes ubiquitination during infection. This modification depends on Lpg2525, an F-box protein from L. pneumophila that acts as a component of the SKP–Cullin–F-box complex. We further demonstrate that Rab5 ubiquitination facilitates the recruitment of RabGAP-5, a Rab5-specific GAP, leading to Rab5 inactivation and subsequent release from the bacterial vacuole. Importantly, the K134 Rab5 mutant limits L. pneumophila replication within host cells. These findings reveal that Lpg2525-mediated Rab5 ubiquitination is a key survival strategy employed by L. pneumophila in infected host cells.
Introduction
Some pathogens that are internalized by eukaryotic cells can bypass host endosomal trafficking of their vacuoles to the lysosome. Vacuolar bacterial pathogen Legionella pneumophila is a suitable model organism for studies to investigate how bacterial evasion of phagocytic clearance occurs. After uptake by the host cell, L. pneumophila creates a specialized infectious niche called the Legionella-containing vacuole (LCV) and escapes from the route of phagocytic maturation (Horwitz, 1983; Isberg et al., 2009; Newton et al., 2010). Concomitantly, L. pneumophila intercepts trafficking of host endoplasmic reticulum–budded vesicles for conversion of the membrane composition of the LCV (Kagan and Roy, 2002; Kagan et al., 2004). Notably, bacterial proteins termed Legionella effectors, which are delivered to the cytosol of cells via the Dot/Icm type IV secretion apparatus, are indispensable for establishing the replication niche (Nagai et al., 2002; Luo and Isberg, 2004; de Felipe et al., 2008; Hubber and Roy, 2010; Qiu and Luo, 2017).
The host small GTPase Rab5 is specifically recruited to the early endocytic/phagocytic compartments, and the activation of Rab5 promotes subsequent recruitment of Rab7 to these compartments. After this, Rab7-positive compartments are delivered to the late endosome/lysosome, leading to degradation of their cargoes (Rink et al., 2005). Therefore, recruitment of Rab5 to the phagosome membrane is crucial for triggering initiation of phagocytic maturation (Huynh and Grinstein, 2008).
Previously, Clemens and colleagues reported that no signal of Rab5 could be detected on vacuoles harboring the WT L. pneumophila strain, whereas significant signals were detected on vacuoles containing the type IV secretion-deficient strain (Clemens et al., 2000). This implied that L. pneumophila uses functions of effector protein(s) to inhibit recruitment of Rab5 to the LCV or to remove Rab5 from the LCV after its recruitment. However, the molecular mechanism underlying Rab5 exclusion was not well understood.
Here, we show that L. pneumophila facilitates ubiquitination of Rab5 via the E3 ubiquitin (UB) ligase activity of S phase kinase-associated protein (SKP)–Cullin–F-box (SCF) complex containing Legionella F-box protein Lpg2525 (also known as MavK), and that the ubiquitination is linked to exclusion of Rab5 from the LCV. This implies a UB-mediated spatial regulation of cellular proteins. We further demonstrate that the ubiquitination of Rab5 leads to the recruitment of the Rab5 inactivator RabGAP-5 to the LCV, resulting in the release of inactivated Rab5 from the LCV. The Rab5 exclusion mediated by Lpg2525-dependent ubiquitination is shown to be crucial for intracellular replication of L. pneumophila.
Results
L. pneumophila facilitates Rab5 ubiquitination
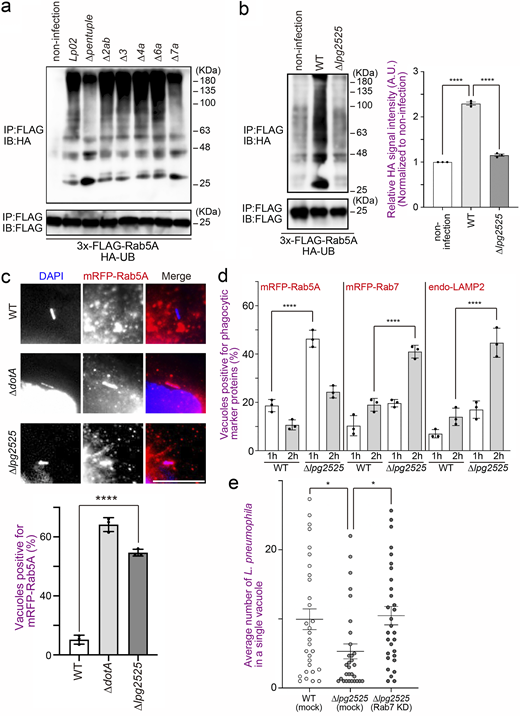
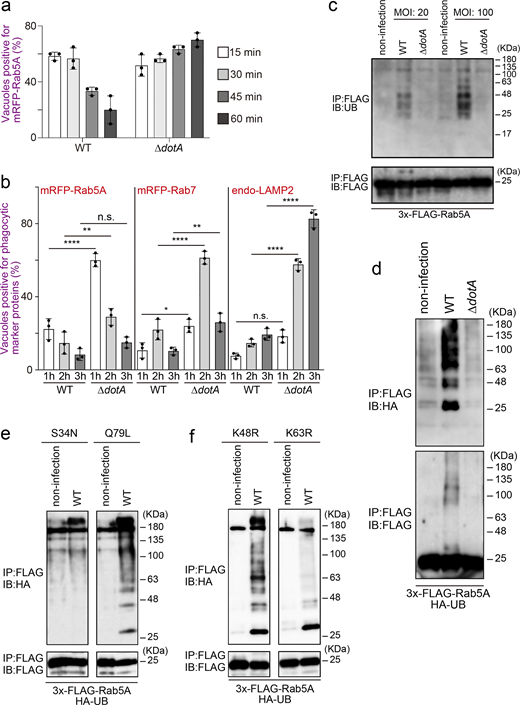
Two plausible mechanisms can be envisaged to explain why Rab5 is recruited to the vacuole containing the type IV secretion-deficient ∆dotA mutant but not the WT L. pneumophila strain (Clemens et al., 2000), that is, inhibition of Rab5 recruitment to the LCV and removal of Rab5 from the LCV after its recruitment. To distinguish between the two possibilities, we first analyzed the kinetics of Rab5A-positive LCVs in the early stage of infection (Fig. 1 a and Fig. S1 a). Rab5A-positive WT LCVs were significantly detected 15–30 min after infection, but their number quickly decreased to <20% of the total LCVs in 60 min. On the other hand, the number of Rab5A-positive LCVs increased over time in cells infected with the ∆dotA mutant strain (Fig. 1 a). We then examined the later stages of infection. The recruitment of phagocytic maturation markers Rab7 and LAMP2 to the vacuole-containing WT L. pneumophila was rarely detected throughout the examined time, while ∆dotA LCVs significantly recruit these proteins at 2–3 h after infection (Fig. 1 b and Fig. S1 b). These results indicate that L. pneumophila removes Rab5 from the LCV quickly after its recruitment by function(s) of effector protein(s) and that the Rab5 elimination leads to blocking the following phagocytic maturation events.
Rab5 ubiquitination upon L. pneumophila infection. (a) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with WT or ∆dotA L. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. (b) HeLa-FcγRII cells or the cells expressing mRFP-Rab5 or mRFP-Rab7 were infected with WT or ∆dotAL. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and the LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. **P < 0.01. *P < 0.05. (c) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A were infected without or with WT or ∆dotA L. pneumophila Lp01 strains for 1 h at the indicated MOI. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against UB or FLAG. (d–f) HEK293-FcγRII cells expressing HA-UB (WT) and 3x-FLAG-Rab5A (d), HA-UB (WT) and 3x-FLAG-Rab5A (S34N), or 3x-FLAG-Rab5A (Q79L) (e), or HA-UB (K48R) or HA-UB (K63R) and 3x-FLAG-Rab5A (f) were infected without or with WT or ∆dotA (d only) L. pneumophila for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. IB, immunoblotting. Source data are available for this figure: SourceData F1.
Rab5 ubiquitination upon L. pneumophila infection. (a) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with WT or ∆dotA L. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. (b) HeLa-FcγRII cells or the cells expressing mRFP-Rab5 or mRFP-Rab7 were infected with WT or ∆dotAL. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and the LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. **P < 0.01. *P < 0.05. (c) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A were infected without or with WT or ∆dotA L. pneumophila Lp01 strains for 1 h at the indicated MOI. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against UB or FLAG. (d–f) HEK293-FcγRII cells expressing HA-UB (WT) and 3x-FLAG-Rab5A (d), HA-UB (WT) and 3x-FLAG-Rab5A (S34N), or 3x-FLAG-Rab5A (Q79L) (e), or HA-UB (K48R) or HA-UB (K63R) and 3x-FLAG-Rab5A (f) were infected without or with WT or ∆dotA (d only) L. pneumophila for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. IB, immunoblotting. Source data are available for this figure: SourceData F1.
Kinetics of Rab5 on the LCV and characterization of Rab5 ubiquitination upon L. pneumophila infection. (a and b) Images showing typical vacuoles positive or negative for mRFP-Rab5A (a) or mRFP-Rab7 and endogenous LAMP2 (b), as related to Fig. 1. Bar, 5 µm. (c) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A were infected without or with the WT or ∆dotA L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were visualized by silver staining. Asterisks indicate precipitated 3x-FLAG-Rab5A. (d and e) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A (S34N) or (Q79L) (d) or 3x-FLAG-Rab5A (e) were infected without or with the WT or ∆dotA L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against UB and FLAG (d), or K48-specific UB, K63-specific UB, or FLAG (e). (f) HEK293-FcγRII cells were infected with the WT or ∆dotA L. pneumophila Lp01 strains with 10 µg/ml cycloheximide (CHX; 239763-M; Sigma-Aldrich) for the indicated times (hours) to inhibit de novo protein synthesis. After treatment, cell lysates were prepared, and equal amounts of proteins were analyzed by IB with antibodies against α-tubulin and Rab5A. IB, immunoblotting. Source data are available for this figure: SourceData FS1.
Kinetics of Rab5 on the LCV and characterization of Rab5 ubiquitination upon L. pneumophila infection. (a and b) Images showing typical vacuoles positive or negative for mRFP-Rab5A (a) or mRFP-Rab7 and endogenous LAMP2 (b), as related to Fig. 1. Bar, 5 µm. (c) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A were infected without or with the WT or ∆dotA L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were visualized by silver staining. Asterisks indicate precipitated 3x-FLAG-Rab5A. (d and e) HEK293-FcγRII cells expressing 3x-FLAG-Rab5A (S34N) or (Q79L) (d) or 3x-FLAG-Rab5A (e) were infected without or with the WT or ∆dotA L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against UB and FLAG (d), or K48-specific UB, K63-specific UB, or FLAG (e). (f) HEK293-FcγRII cells were infected with the WT or ∆dotA L. pneumophila Lp01 strains with 10 µg/ml cycloheximide (CHX; 239763-M; Sigma-Aldrich) for the indicated times (hours) to inhibit de novo protein synthesis. After treatment, cell lysates were prepared, and equal amounts of proteins were analyzed by IB with antibodies against α-tubulin and Rab5A. IB, immunoblotting. Source data are available for this figure: SourceData FS1.
We next tried to identify the effector(s) responsible for elimination of Rab5 from the LCV. After immunoprecipitation (IP) with the FLAG antibody to isolate ectopically expressed 3x-FLAG-Rab5A in the cells, we could not detect any specific proteins interacting with Rab5A upon infection with the WT strain (Fig. S1 c, middle lane). However, we detected high molecular weight smears in this lane, which were less prominent in the other lanes (Fig. S1 c). As polyubiquitination is generally detected as a smear-like pattern, we thought that Rab5 might be polyubiquitinated upon L. pneumophila infection, and this was confirmed by detection of endogenous (Fig. 1 c) and exogenous (Fig. 1 d, top panel) UB. Of note, the smear-like signal was also detected in an immunoblot with the FLAG antibody (Fig. 1 d, bottom panel), suggesting that L. pneumophila directly ubiquitinates Rab5.
Next, we characterized Legionella-mediated Rab5 ubiquitination. As shown in Fig. 1 e and Fig. S1 d, significant ubiquitination of Rab5 was observed in cells expressing 3x-FLAG-Rab5A Q79L (GTP-bound active form), but not those expressing 3x-FLAG-Rab5A S34N (GDP-bound inactive form). Furthermore, L. pneumophila infection facilitated K63-, but not K48-linked ubiquitination of Rab5 (Fig. S1 e). Consistently, K63R UB failed to assemble polyubiquitin chains on Rab5A, whereas K48R assembled (Fig. 1 f). No reduction in the protein level of Rab5A was observed following Legionella infection (Fig. S1 f). This suggests that L. pneumophila modulates the function of Rab5 rather than degrading it through the UB–proteasome pathway, as K63-linked ubiquitination typically plays a role in signal transduction and membrane traffic rather than in proteasomal degradation (Nelson et al., 2013).
The Legionella F-box protein Lpg2525 has a pivotal role in ubiquitination of Rab5
To identify factor(s) responsible for Rab5 ubiquitination, we infected cells with Legionella strains depleted of large genomic islands (O’Connor et al., 2011, Fig. S2 a) and examined ubiquitination of Rab5. In cells infected with the ∆pentuple (Fig. 2 a, third lane from left) or the ∆7a (Fig. 2 a, rightmost lane) strain, the ubiquitination signal of Rab5A was significantly reduced, and Rab5A was largely retained on the LCVs (Fig. S2 b). These results suggest that at least one Legionella effector encoded in the island 7a is responsible for Rab5 ubiquitination, which is associated with Rab5 release from the LCV.
Effect of large genomic islands depletion in the L. pneumophila genome on Rab5 recruitment to the LCV during infection. (a) Schematic representation of the ∆pentuple and single island-depleted strains (adopted from O’Connor et al. [2011]). (b) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with the indicated L. pneumophila Lp02 strains for 1 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. RFP-Rab5A–positive LCVs were quantified. Representative data of three independent experiments were presented (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) List of L. pneumophila effectors encoded in the island 7a. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Rab7. At 72 h after transfection, the efficacy of siRNA was assessed by IB with the Rab7 antibody. IB with α-tubulin antibody was conducted as an internal control. IB, immunoblotting. Source data are available for this figure: SourceData FS2.
Effect of large genomic islands depletion in the L. pneumophila genome on Rab5 recruitment to the LCV during infection. (a) Schematic representation of the ∆pentuple and single island-depleted strains (adopted from O’Connor et al. [2011]). (b) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with the indicated L. pneumophila Lp02 strains for 1 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. RFP-Rab5A–positive LCVs were quantified. Representative data of three independent experiments were presented (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) List of L. pneumophila effectors encoded in the island 7a. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Rab7. At 72 h after transfection, the efficacy of siRNA was assessed by IB with the Rab7 antibody. IB with α-tubulin antibody was conducted as an internal control. IB, immunoblotting. Source data are available for this figure: SourceData FS2.
Identification of Legionella effector(s) responsible for Rab5 ubiquitination. (a) HeLa-FcγRII cells expressing HA-UB and 3x-FLAG-Rab5A were infected without or with the indicated L. pneumophila Lp02 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. (b) HeLa-FcγRII cells expressing HA-UB and 3x-FLAG-Rab5A were infected with or without the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P value is determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (d) HeLa-FcγRII cells or the cells expressing mRFP-Rab5A or mRFP-Rab7 were infected with indicated L. pneumophila Lp01 strains for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (e) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Rab7. At 72 h after transfection, the cells were infected with indicated L. pneumophila Lp01 strains for 8 h at an MOI of 5. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. The graph shows the average number of L. pneumophila in a single vacuole. Data are representative of three independent experiments (30 vacuoles were scored in each experiment). Results are shown as means ± SEM. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. IB, immunoblotting. Source data are available for this figure: SourceData F2.
Identification of Legionella effector(s) responsible for Rab5 ubiquitination. (a) HeLa-FcγRII cells expressing HA-UB and 3x-FLAG-Rab5A were infected without or with the indicated L. pneumophila Lp02 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. (b) HeLa-FcγRII cells expressing HA-UB and 3x-FLAG-Rab5A were infected with or without the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P value is determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (d) HeLa-FcγRII cells or the cells expressing mRFP-Rab5A or mRFP-Rab7 were infected with indicated L. pneumophila Lp01 strains for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (e) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Rab7. At 72 h after transfection, the cells were infected with indicated L. pneumophila Lp01 strains for 8 h at an MOI of 5. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. The graph shows the average number of L. pneumophila in a single vacuole. Data are representative of three independent experiments (30 vacuoles were scored in each experiment). Results are shown as means ± SEM. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. IB, immunoblotting. Source data are available for this figure: SourceData F2.
Among 25 effectors encoded in the island 7a (Fig. S2 c), we focused on Lpg2525 which is known to contain an F-box domain (Angot et al., 2007; Ensminger and Isberg, 2010). Members of a family of F-box proteins form a complex with SKP1–Cullin1 through its F-box domain, and Cullin1 interacts with E3 UB ligase Ring-box protein 1 (Skaar et al., 2013). Therefore, F-box proteins are known to function as adaptors that link substrates with E3 UB ligase complexes (Skaar et al., 2013). Of note, L. pneumophila has five genes encoding plausible F-box proteins, including Lpg2525 (Angot et al., 2007; Ensminger and Isberg, 2010). We therefore examined whether Lpg2525 is required for Rab5 ubiquitination. As shown in Fig. 2 b, a significant reduction in Rab5A ubiquitination was observed in cells infected with the ∆lpg2525 strain. Furthermore, the number of Rab5A-positive LCVs was significantly increased in cells infected with the ∆lpg2525 strain (Fig. 2 c). Significant recruitment of Rab7 and LAMP2 to vacuoles containing the ∆lpg2525 strain was also observed at 2 h after infection (Fig. 2 d), suggesting that L. pneumophila promotes Rab5 ubiquitination on the vacuole through the function of Lpg2525, leading to Rab5 removal to avoid bacterial clearance by phagocytic maturation. The role of the Rab5-Rab7 axis targeted by Lpg2525 was further supported by data showing that the intracellular growth defect of L. pneumophila due to the loss of Lpg2525 was restored in Rab7-silenced cells, which inhibits phagocytic maturation (Fig. 2 e and Fig. S2 d).
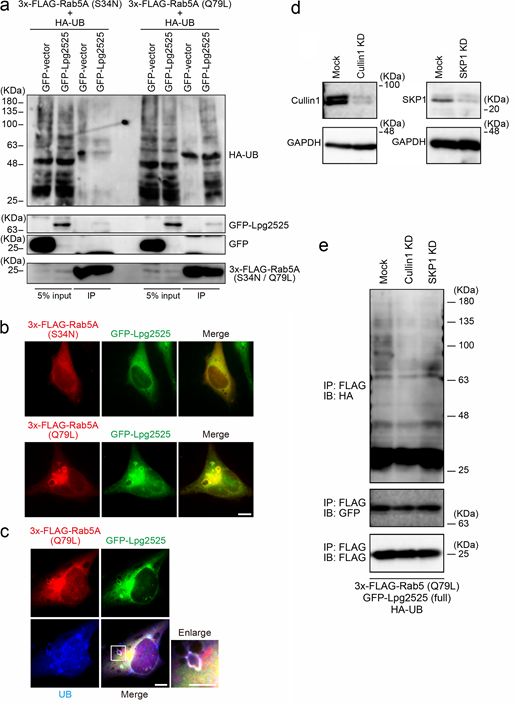
To validate the involvement of Lpg2525 as a component of the SCF complex, we conducted the interaction analysis using IP (Fig. 3 a). Contrary to a previous study that did not detect interaction between Lpg2525 and SKP1–Cullin1 (Ensminger and Isberg, 2010), we observed a slight but significant binding of Lpg2525 to Cullin1 and SKP1, specifically in cells expressing full-length (FL) Lpg2525, but not in those expressing the F-box deleted (∆FB) variant (Fig. 3 a). In a SCF complex, interaction between F-box proteins and Cullin1 is mediated by SKP1 (Skaar et al., 2013). The amount of Cullin1 co-precipitated with Lpg2525 was robustly reduced by silencing of SKP1 (Fig. 3 b), supporting that Lpg2525 functions as a physiological component of the SCF complex.
Lpg2525 forms a complex with SKP1–Cullin1, and the F-box domain of Lpg2525 is required for Rab5 ubiquitination. (a) HeLa-FcγRII cells were transfected with a plasmid-encoding 3x-FLAG vector, 3x-FLAG-Lpg2525 (FL), or 3x-FLAG-Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against Cullin1, SKP1, and FLAG. (b) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting SKP1. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding 3x-FLAG-Lpg2525 (FL) for 24 h, lysed, and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with the indicated antibodies. (c) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (Q79L), HA-UB, and GFP, GFP-Lpg2525, (FL) or GFP-Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against GFP, FLAG, and HA. (d) HeLa-FcγRII cells expressing 3x-FLAG-Rab5A and HA-UB were infected with the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. (e) HeLa-FcγRII cells or the cells expressing mRFP-Rab5A or Rab7 were infected with the indicated L. pneumophila Lp01 strains for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Phagocytic markers positive LCVs were quantified, and the representative data of three independent experiments were presented (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. ***P < 0.001. IB, immunoblotting. Source data are available for this figure: SourceData F3.
Lpg2525 forms a complex with SKP1–Cullin1, and the F-box domain of Lpg2525 is required for Rab5 ubiquitination. (a) HeLa-FcγRII cells were transfected with a plasmid-encoding 3x-FLAG vector, 3x-FLAG-Lpg2525 (FL), or 3x-FLAG-Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against Cullin1, SKP1, and FLAG. (b) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting SKP1. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding 3x-FLAG-Lpg2525 (FL) for 24 h, lysed, and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with the indicated antibodies. (c) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (Q79L), HA-UB, and GFP, GFP-Lpg2525, (FL) or GFP-Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against GFP, FLAG, and HA. (d) HeLa-FcγRII cells expressing 3x-FLAG-Rab5A and HA-UB were infected with the indicated L. pneumophila Lp01 strains for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. (e) HeLa-FcγRII cells or the cells expressing mRFP-Rab5A or Rab7 were infected with the indicated L. pneumophila Lp01 strains for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria or with DAPI and LAMP2 antibody for detection of intracellular bacteria and endogenous LAMP2. Phagocytic markers positive LCVs were quantified, and the representative data of three independent experiments were presented (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. ***P < 0.001. IB, immunoblotting. Source data are available for this figure: SourceData F3.
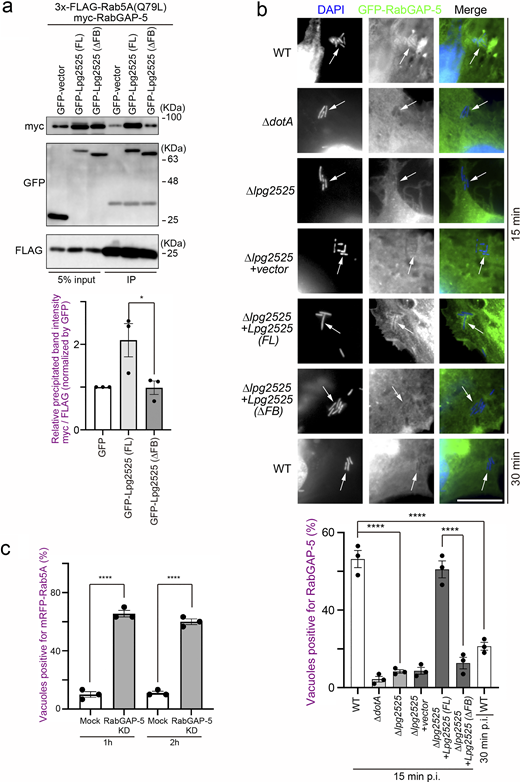
Consistent with that L. pneumophila preferentially ubiquitinates GTP-bound active Rab5A (Fig. 1 e), ectopically expressed Lpg2525-dependent Rab5A ubiquitination was detected in cells expressing 3x-FLAG-Rab5A (Q79L), but not in those expressing 3x-FLAG-Rab5A (S34N) (Fig. S3 a). Moreover, significant co-localization of GFP-Lpg2525 and 3x-FLAG-Rab5A (Q79L) was observed in the cell (Fig. S3 b), where UB was detected on the structure positive for both Rab5A (Q79L) and Lpg2525 (Fig. S3 c). Of note, the ubiquitination of Rab5A (Q79L) was detected only in cells expressing FL Lpg2525 but not in those expressing ∆FB Lpg2525 (Fig. 3 c, right panel), even though both proteins were found to interact with Rab5A (Q79L) (Fig. 3 c, left panel). Taken together, these results suggest that, within the SCF E3 ligase complex, the F-box protein Lpg2525 plays a role in recognizing Rab5A as a substrate and mediating its ubiquitination. These notions were confirmed by data showing that silencing of Cullin1 or SKP1 (Fig. S3 d) suppresses Lpg2525-dependent Rab5A ubiquitination (Fig. S3 e). Consistent with this, Rab5A ubiquitination was observed in cells infected with the ∆lpg2525 strain complemented with FL Lpg2525-expressing plasmid, but not with the ∆FB variant (Fig. 3 d). The number of Rab5A-positive LCVs was drastically reduced in cells infected with the ∆lpg2525 strain expressing FL Lpg2525 (Fig. 3 e). Furthermore, vacuoles containing the ∆lpg2525 strain expressing FL Lpg2525, but not the ∆FB variant, successfully evade phagocytic maturation (Fig. 3 e). Collectively, our data show that the SCF complex containing Lpg2525 specifically recognizes and ubiquitinates active Rab5 on the LCV, leading to its dissociation from the vacuole and evasion of the endocytic pathway.
Lpg2525 preferentially ubiquitinates GTP-bound active Rab5. (a) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (S34N) or (Q79L), HA-UB, and GFP or GFP-Lpg2525. At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA, GFP, and FLAG. (b) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (S34N) or (Q79L) and GFP-Lpg2525. At 24 h after transfection, cells were fixed and stained with an antibody against FLAG. Bar, 5 µm. (c) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (Q79L) and GFP-Lpg2525. At 24 h after transfection, cells were fixed and stained with antibodies against UB and FLAG. Bars, 5 µm. An enlarged image of the white inset box is shown. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Cullin1 or SKP1. At 72 h after transfection, the efficacy of siRNAs was assessed by IB with Cullin1 and SKP1 antibodies. IB with GAPDH antibody was conducted as an internal control. (e) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Cullin1 or SKP1. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding 3x-FLAG-Rab5A (Q79L), GFP-Lpg2525, and HA-UB for 24 h, lysed, and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with the indicated antibodies. IB, immunoblotting. Source data are available for this figure: SourceData FS3.
Lpg2525 preferentially ubiquitinates GTP-bound active Rab5. (a) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (S34N) or (Q79L), HA-UB, and GFP or GFP-Lpg2525. At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA, GFP, and FLAG. (b) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (S34N) or (Q79L) and GFP-Lpg2525. At 24 h after transfection, cells were fixed and stained with an antibody against FLAG. Bar, 5 µm. (c) HeLa-FcγRII cells were transfected with plasmids encoding 3x-FLAG-Rab5A (Q79L) and GFP-Lpg2525. At 24 h after transfection, cells were fixed and stained with antibodies against UB and FLAG. Bars, 5 µm. An enlarged image of the white inset box is shown. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Cullin1 or SKP1. At 72 h after transfection, the efficacy of siRNAs was assessed by IB with Cullin1 and SKP1 antibodies. IB with GAPDH antibody was conducted as an internal control. (e) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Cullin1 or SKP1. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding 3x-FLAG-Rab5A (Q79L), GFP-Lpg2525, and HA-UB for 24 h, lysed, and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with the indicated antibodies. IB, immunoblotting. Source data are available for this figure: SourceData FS3.
Lpg2525-mediated Rab5 ubiquitination promotes the association between active Rab5 and RabGAP-5
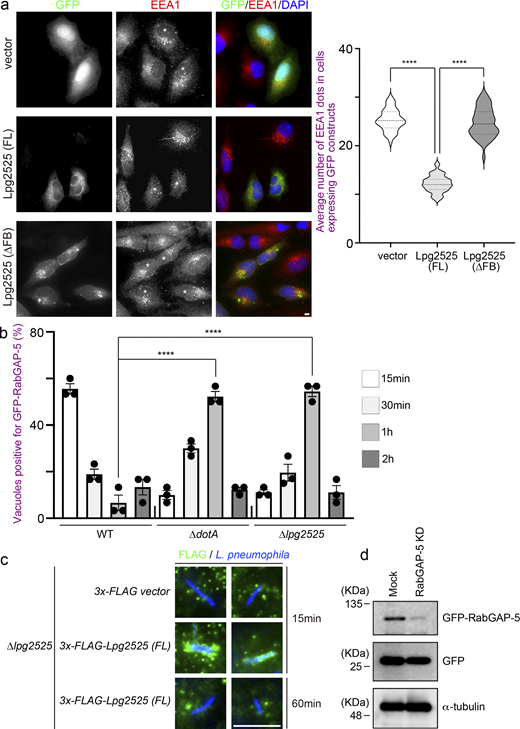
To explore the molecular mechanism by which Legionella-induced ubiquitination of Rab5 leads to its dissociation from the LCV, we examined the behavior of RabGAP-5, which inactivates Rab5 by hydrolyzing its GTP to GDP (Haas et al., 2005). Expression of Lpg2525 significantly increased the association between Rab5A and RabGAP-5 (Fig. 4 a). Furthermore, this enhancement was reduced upon deletion of the F-box domain of Lpg2525. Consistent with a previous report showing that the overproduction of RabGAP-5 leads to the dispersion of early endosome antigen 1 (EEA1) puncta due to Rab5 inactivation (Haas et al., 2005), a reduction in EEA1 puncta was observed in cells expressing GFP-Lpg2525 (FL), but not GFP-Lpg2525 (∆FB) (Fig. S4 a), suggesting that Lpg2525 facilitates inactivation of Rab5 through its ubiquitination and subsequent interaction with RabGAP-5. Significant recruitment of RabGAP-5 to vacuoles containing WT L. pneumophila was observed at 15 min after infection, but not at 30 min (Fig. 4 b, top and bottom rows), which explains the rapid release of Rab5 from the LCV (Fig. 1 a). However, the recruitment of RabGAP-5 to the LCV was suppressed in cells infected with the ∆dotA and ∆lpg2525 strains, even at 15 min after infection (Fig. 4 b, second to third rows), suggesting that Lpg2525 enhances the recruitment of RabGAP-5 to the LCV. Consistent with the delay in Rab5A dissociation from vacuoles occupied by ∆dotA or ∆lpg2525 (Fig. 1 b and Fig. 2 d), accumulation of RabGAP-5 on these vacuoles was observed at 1 h after infection (Fig. S4 b). It was further demonstrated that the recruitment of RabGAP-5 to the LCV requires the F-box domain of Lpg2525 (Fig. 4 b, fifth and sixth rows). Notably, translocated Lpg2525 was detected on the vacuoles in the early (15 min), but not the later (60 min), stage of infection (Fig. S4 c). Importantly, sustained localization of Rab5A was observed on the WT LCV in the RabGAP-5-silenced cells (Fig. 4 c and Fig. S4 d). These results support a model in which ubiquitinated Rab5 residing on the LCV is preferentially recognized by RabGAP-5, leading to its inactivation and subsequent removal from the vacuole.
Lpg2525 promotes the association of RabGAP-5 with Rab5, and its F-box domain is crucial in recruiting RabGAP-5 to the LCV. (a) HeLa-FcγRII cells were transfected with myc-RabGAP-5, 3x-FLAG Rab5A (Q79L), and GFP, GFP-Lpg2525 (FL) or GFP- Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against myc, GFP, and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P-value is determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. (b) HeLa-FcγRII cells expressing GFP-RabGAP-5 were infected with the indicated L. pneumophila Lp01 strains for the indicated time at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Arrows indicate the position of intracellular L. pneumophila. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting RabGAP-5. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding mRFP-Rab5A for 24 h, infected with the WT L. pneumophila Lp01 strain for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. IB, immunoblotting. Source data are available for this figure: SourceData F4.
Lpg2525 promotes the association of RabGAP-5 with Rab5, and its F-box domain is crucial in recruiting RabGAP-5 to the LCV. (a) HeLa-FcγRII cells were transfected with myc-RabGAP-5, 3x-FLAG Rab5A (Q79L), and GFP, GFP-Lpg2525 (FL) or GFP- Lpg2525 (∆FB). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against myc, GFP, and FLAG. Data are representative of three independent experiments. Results are shown as means ± SD. P-value is determined using one-way ANOVA with Tukey’s multiple comparisons. *P < 0.05. (b) HeLa-FcγRII cells expressing GFP-RabGAP-5 were infected with the indicated L. pneumophila Lp01 strains for the indicated time at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Arrows indicate the position of intracellular L. pneumophila. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting RabGAP-5. At 48 h after transfection, the cells were transfected with an additional plasmid-encoding mRFP-Rab5A for 24 h, infected with the WT L. pneumophila Lp01 strain for 1 and 2 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. IB, immunoblotting. Source data are available for this figure: SourceData F4.
The effect of Lpg2525 expression on early endosomal morphology and the kinetics of RabGAP-5 or translocated Lpg2525 on the LCV. (a) HeLa-FcγRII cells were transfected with a plasmid-encoding GFP, GFP-Lpg2525 (full), or GFP-Lpg2525 (∆FB). At 24 h after transfection, cells were fixed and stained with anti-EEA1 antibody and DAPI. Asterisks indicate cells expressing GFP or the GFP-fusion proteins. Bar, 5 µm. Data are representative of three independent experiments (50 cells expressing the GFP constructs were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (b) HeLa-FcγRII cells expressing GFP-RabGAP-5 were infected with the indicated L. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells were infected with the ∆lpg2525 L. pneumophila Lp01 strain complemented with 3x-FLAG- or 3x-FLAG-Lpg2525–expressing plasmids for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI and FLAG antibody for detection of intracellular bacteria and bacterially expressing FLAG-fusion proteins. Bar, 5 µm. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting RabGAP-5. At 48 h after transfection, the cells were transfected with the additional plasmids encoding GFP and GFP-RabGAP-5 for 24 h. The efficacy of siRNA was assessed by IB with GFP antibody. IB with α-tubulin antibody was conducted as an internal control. IB, immunoblotting. Source data are available for this figure: SourceData FS4.
The effect of Lpg2525 expression on early endosomal morphology and the kinetics of RabGAP-5 or translocated Lpg2525 on the LCV. (a) HeLa-FcγRII cells were transfected with a plasmid-encoding GFP, GFP-Lpg2525 (full), or GFP-Lpg2525 (∆FB). At 24 h after transfection, cells were fixed and stained with anti-EEA1 antibody and DAPI. Asterisks indicate cells expressing GFP or the GFP-fusion proteins. Bar, 5 µm. Data are representative of three independent experiments (50 cells expressing the GFP constructs were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (b) HeLa-FcγRII cells expressing GFP-RabGAP-5 were infected with the indicated L. pneumophila Lp01 strains for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) HeLa-FcγRII cells were infected with the ∆lpg2525 L. pneumophila Lp01 strain complemented with 3x-FLAG- or 3x-FLAG-Lpg2525–expressing plasmids for the indicated times at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria and then permeabilized and further stained with DAPI and FLAG antibody for detection of intracellular bacteria and bacterially expressing FLAG-fusion proteins. Bar, 5 µm. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting RabGAP-5. At 48 h after transfection, the cells were transfected with the additional plasmids encoding GFP and GFP-RabGAP-5 for 24 h. The efficacy of siRNA was assessed by IB with GFP antibody. IB with α-tubulin antibody was conducted as an internal control. IB, immunoblotting. Source data are available for this figure: SourceData FS4.
L. pneumophila induces ubiquitination of an evolutionarily conserved lysine residue of Rab5, and mutation of this residue suppresses the intracellular growth of L. pneumophila
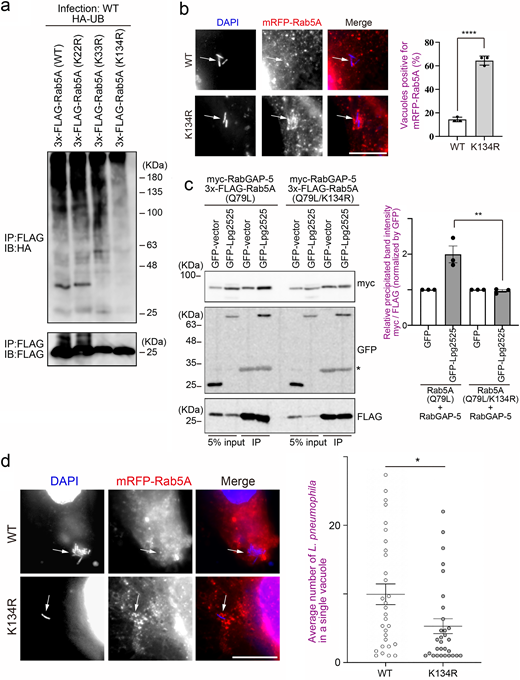
Generally, UB conjugates to the lysine residue(s) of a substrate protein. We next aimed to identify the lysine residue(s) of Rab5 that is targeted for Lpg2525-mediated ubiquitination. As the host range of L. pneumophila is quite diverse (Mintz and Shuman, 1988), we reasoned that L. pneumophila targets highly conserved lysine residue(s) of Rab5. Alignment of Rab5 sequences revealed that only three lysine residues (K22, K33, and K134 in human Rab5A) are conserved across most eukaryotic Rab5 sequences (Fig. S5 a). Therefore, we examined which of these were critical for ubiquitination by L. pneumophila. As shown in Fig. 5 a, a marked reduction in L. pneumophila–mediated ubiquitination was observed on the K134R mutant of Rab5A. Concomitantly, robust recruitment to the LCV was detected for Rab5A K134R (Fig. 5 b), but not Rab5A K22R or K33R (Fig. S5 b), compared with WT Rab5A (Fig. 5 b). Importantly, Lpg2525-enhanced interaction between Rab5A and RabGAP-5 was abolished in the K134R mutant of Rab5A (Fig. 5 c). Furthermore, intracellular replication of L. pneumophila was significantly restricted in cells expressing mRFP-Rab5A (K134R), a mutant that can not conjugate UB (Fig. 5 d). Taken together, these results suggest that the Lpg2525-mediated LCV recruitment of RabGAP-5 is the key strategy of L. pneumophila for intracellular survival via inactivation and subsequent elimination of the critical endocytic factor Rab5.
Alignment of Rab5 sequences across different spices. (a) Alignment of the aa sequences of human Rab5A and Rab5 proteins from Rattus norvegicus, Danio rerio, Caenorhabditis elegans, Drosophila melanogaster, Saccharomyces cerevisiae, Rhizoctonia solani, Dictyostelium discoideum, and Leishmania donovani. Identical and conserved substitution residues among all spices are highlighted in red and blue, respectively. Arrowheads indicate conserved lysine residues. (b) HeLa-FcγRII cells expressing mRFP-Rab5A (WT), (K22R), or (K33R) were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Arrows indicate the position of intracellular L. pneumophila. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. Values are determined using one-way ANOVA with Tukey’s multiple comparisons. n.s., not significance.
Alignment of Rab5 sequences across different spices. (a) Alignment of the aa sequences of human Rab5A and Rab5 proteins from Rattus norvegicus, Danio rerio, Caenorhabditis elegans, Drosophila melanogaster, Saccharomyces cerevisiae, Rhizoctonia solani, Dictyostelium discoideum, and Leishmania donovani. Identical and conserved substitution residues among all spices are highlighted in red and blue, respectively. Arrowheads indicate conserved lysine residues. (b) HeLa-FcγRII cells expressing mRFP-Rab5A (WT), (K22R), or (K33R) were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, and then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Arrows indicate the position of intracellular L. pneumophila. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. Values are determined using one-way ANOVA with Tukey’s multiple comparisons. n.s., not significance.
Mutation of K134 of Rab5, a residue identified as ubiquitinated upon infection with L. pneumophila, affects both Rab5 recruitment to the LCV and bacterial proliferation within cells. (a) HeLa-FcγRII cells expressing 3x-FLAG-Rab5A (WT, K22R, K33R, or K134R) and HA-UB were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. (b) HeLa-FcγRII cells expressing mRFP-Rab5A (WT) or (K134R) were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P value is determined using an unpaired two-tailed Student’s test. ****P < 0.0001. (c) HeLa-FcγRII cells were transfected with myc-RabGAP-5, GFP or GFP-Lpg2525, and 3x-FLAG-Rab5A (Q79L, left panel) or -Rab5A (Q79L/K134R, right panel). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against myc, GFP, and FLAG. An asterisk indicates nonspecific bands. Data are representative of three independent experiments. Results are shown as means ± SD. P value is determined using one-way ANOVA with Tukey’s multiple comparisons. **P < 0.01. (d) HeLa-FcγRII cells expressing mRFP-Rab5A (WT) or (K134R) were infected with the WT L. pneumophila Lp01 strain for 8 h at an MOI of 5. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. The graph shows the average number of L. pneumophila in a single vacuole. Data are representative of three independent experiments (30 vacuoles were scored in each experiment). Results are shown as means ± SEM. P value is determined using an unpaired two-tailed Student’s test. *P < 0.05. IB, immunoblotting. Source data are available for this figure: SourceData F5.
Mutation of K134 of Rab5, a residue identified as ubiquitinated upon infection with L. pneumophila, affects both Rab5 recruitment to the LCV and bacterial proliferation within cells. (a) HeLa-FcγRII cells expressing 3x-FLAG-Rab5A (WT, K22R, K33R, or K134R) and HA-UB were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 50. After infection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against HA and FLAG. (b) HeLa-FcγRII cells expressing mRFP-Rab5A (WT) or (K134R) were infected with the WT L. pneumophila Lp01 strain for 1 h at an MOI of 10. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. Data are representative of three independent experiments (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P value is determined using an unpaired two-tailed Student’s test. ****P < 0.0001. (c) HeLa-FcγRII cells were transfected with myc-RabGAP-5, GFP or GFP-Lpg2525, and 3x-FLAG-Rab5A (Q79L, left panel) or -Rab5A (Q79L/K134R, right panel). At 24 h after transfection, cell lysates were prepared and immunoprecipitated with the FLAG-M2 beads. The precipitated proteins were analyzed by IB with antibodies against myc, GFP, and FLAG. An asterisk indicates nonspecific bands. Data are representative of three independent experiments. Results are shown as means ± SD. P value is determined using one-way ANOVA with Tukey’s multiple comparisons. **P < 0.01. (d) HeLa-FcγRII cells expressing mRFP-Rab5A (WT) or (K134R) were infected with the WT L. pneumophila Lp01 strain for 8 h at an MOI of 5. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. Bar, 5 µm. The graph shows the average number of L. pneumophila in a single vacuole. Data are representative of three independent experiments (30 vacuoles were scored in each experiment). Results are shown as means ± SEM. P value is determined using an unpaired two-tailed Student’s test. *P < 0.05. IB, immunoblotting. Source data are available for this figure: SourceData F5.
Discussion
To summarize our present results, we propose a working model for the elimination of Rab5 from the LCV (Fig. 6). In the early stage of infection, GDP-bound inactive Rab5 is targeted to the LCV and activated by exchange of GDP for GTP (Fig. 6, I); this is implicated in a host defense mechanism that eliminates pathogens by inducing the endocytic pathway. Once Rab5 has been activated on the LCV, the SCF complex containing Lpg2525 recognizes it and catalyzes K63-linked ubiquitination on its residue K134 (Fig. 6 II). RabGAP-5 subsequently associates with ubiquitinated Rab5 (Fig. 6 III). Consequently, Rab5 is inactivated through RabGAP-5–dependent GTP hydrolysis and removed from the LCV (Fig. 6 IV), enabling the bacteria to evade the host response, leading to bacterial clearance.
Working model for the removal of Rab5 from the LCV. (I–IV) GDP-bound inactive Rab5 is targeted to the LCV and activated by the exchange of GDP for GTP (I). The SCF complex containing Lpg2525 recognizes Rab5 on the LCV and catalyzes ubiquitination on it (II). RabGAP-5 associates with Rab5, which is ubiquitinated on its K134 residue (III). RabGAP-5 inactivates Rab5 and removes it from the LCV (IV).
Working model for the removal of Rab5 from the LCV. (I–IV) GDP-bound inactive Rab5 is targeted to the LCV and activated by the exchange of GDP for GTP (I). The SCF complex containing Lpg2525 recognizes Rab5 on the LCV and catalyzes ubiquitination on it (II). RabGAP-5 associates with Rab5, which is ubiquitinated on its K134 residue (III). RabGAP-5 inactivates Rab5 and removes it from the LCV (IV).
Several studies have partially explored the molecular mechanisms by which L. pneumophila evades phagocytic maturation. L. pneumophila effector VipD was found to interact with Rab5 and disrupt endosomal trafficking (Ku et al., 2012). Machner and colleagues found that VipD is a phospholipase A1-like protein activated by binding to Rab5, which catalyzes the depletion of phosphatidylinositol-3 phosphate (PI3P) on endosomes. This depletion removes fusion factors from the endosomes, leading to L. pneumophila’s avoidance of endosomal maturation (Gaspar and Machner, 2014). Hilbi and colleagues reported that the Legionella effector RidL interacts with Vps29, a component of the retromer complex, and inhibits its function through the PI3P-binding domain of RidL, leading to evasion of endocytic maturation (Finsel et al., 2013). Furthermore, another PI3P-interacting Legionella effector, RavD, was identified as a suppressor of endolysosomal maturation (Pike et al., 2019). However, these studies did not address the compelling question of how L. pneumophila avoids the initial process of the host endocytic pathway that is achieved via the Rab5–Rab7 axis. Our results, demonstrating Rab5 exclusion through effector-mediated ubiquitination, provide valuable insight into the molecular mechanisms by which L. pneumophila evades endosomal maturation.
We show here that Lpg2525 prominently co-localizes with vacuolar compartments formed by the ectopic expression of GTP-locked Rab5A. A previous report suggests that Lpg2525 can associate with the endomembrane independently of the CAAX motif (lipidation motif by prenyltransferase) at its C terminus (Ivanov et al., 2010). These findings suggest that Lpg2525 may bind to phospholipid and/or phosphoinositide for its association with the endomembrane. Future studies could explore this possibility.
Identifying the specific lysine residue of Rab5 that is ubiquitinated upon L. pneumophila infection could provide further important insights into phagocytic avoidance of the LCV in other host species. L. pneumophila is known to manipulate Rab proteins such as Rab1, which is involved in interception of membrane trafficking, similarly in both human and Dictyostelium discoideum cells (Urwyler et al., 2009). However, it is not yet known whether the molecular mechanism of Rab5 manipulation by L. pneumophila is conserved across host species. To address this question, it would be necessary to investigate whether similar effector-induced Rab5 manipulation, such as UB-mediated association with RabGAP-5, occurs in D. discoideum upon infection. This could be achieved by mutating the lysine residue at position 162 of D. discoideum Rab5 (corresponding to K134 of human Rab5A). The detailed molecular mechanisms underlying the process of Rab5 exclusion remain elusive. To understand how RabGAP-5 recognizes ubiquitinated Rab5, a molecular dissection of RabGAP-5 to identify the responsible region(s) of the protein and/or potential adaptor/regulator proteins would be required. It is also worth noting that previous proteomic studies identified K134 as an endogenously polyubiquitinated residue of Rab5A (Wagner et al., 2012; Chen et al., 2014). A crucial role of K134 in structural integrity of Rab5 was suggested by its direct interaction with both GDP and GTP (Shin et al., 2017). These findings prompted us to hypothesize that ubiquitination of this lysine inactivates Rab5 function. RabGAP-5 may directly recognize the polyubiquitin chains on K134, making the GTP-bound form of Rab5 accessible to RabGAP-5. Alternatively, ubiquitination of K134 could induce a structural change in Rab5, enhancing its affinity for RabGAP-5. The precise molecular mechanism by which RabGAP-5 preferentially interacts with ubiquitinated Rab5 will need to be clarified through future structural analyses.
The involvement of the endosomal network in biogenesis of bacterial replicative vacuoles appears to be highly complex; Mycobacterium tuberculosis persistently recruits Rab5 to its phagosome and likely blocks phagosome maturation through events downstream of Rab5 acquisition (Clemens et al., 2000), while Rab7, recruited to the bacterial phagosome, has been shown to contribute toward the intracellular growth of L. pneumophila (Li et al., 2024). These studies suggest that host endosomal proteins, including Rab5 and Rab7, may play specific roles in LCV development, although their persistent presence could be detrimental to bacterial vacuoles. In this context, it is plausible that the temporal and spatial regulation of Rab5 involves additional bacterial and/or host factors. These aspects need to be explored further to fully understand the bacterial strategies for establishing their growth niches.
Materials and methods
Antibodies
Anti-FLAG rabbit polyclonal antibody, Cat# F7425; Sigma-Aldrich.
Anti-UB rabbit polyclonal antibody, Cat# 5839; Cell Signaling.
Anti-HA rabbit polyclonal antibody, Cat# 561; MBL.
Anti-Cullin1 rabbit polyclonal antibody, Cat# 12895-1-AP; Proteintech.
Anti-SKP1 rabbit polyclonal antibody, Cat# 10990-2-AP; Proteintech.
Anti-GFP rabbit polyclonal antibody, Cat# A-6455; Thermo Fisher Scientific.
Anti-Lamp2 mouse monoclonal antibody, Cat# Ab25631; Abcam.
Anti-myc rabbit polyclonal antibody, Cat# 562; MBL.
Anti-UB (K63) rabbit polyclonal antibody, Cat# 5621; Cell Signaling.
Anti-UB (K48) rabbit polyclonal antibody, Cat# 4289; Cell Signaling.
Anti–α-tubulin mouse monoclonal antibody, Cat# T6074; Sigma-Aldrich.
Anti-Rab5A mouse monoclonal antibody, Cat# 610725; BD Biosciences.
Anti-UB mouse monoclonal antibody, Cat# D058-3; MBL.
Anti-GAPDH mouse monoclonal antibody, Cat# H86504M; BioDesign.
Anti-EEA1 mouse monoclonal antibody, Cat# 610457; BD Biosciences.
Anti-Legionella rabbit antiserum (see “production of anti-Legionella antiserum”).
Anti-Legionella mouse antiserum (see “production of anti-Legionella antiserum”).
Peroxidase-conjugated goat anti-rabbit antibody, Cat# 170–6515; Bio-Rad.
Peroxidase-conjugated goat anti-mouse antibody, Cat# 170–6516; Bio-Rad.
Alexa Fluor 488–conjugated goat anti-rabbit antibody, Cat# A11034; Thermo Fisher Scientific.
Alexa Fluor 594–conjugated goat anti-rabbit antibody, Cat# A11037; Thermo Fisher Scientific.
Alexa Fluor 594–conjugated goat anti-mouse antibody, Cat# A11032; Thermo Fisher Scientific.
Alexa Fluor 350–conjugated donkey anti-mouse antibody, Cat# A10035; Thermo Fisher Scientific.
Bacterial strains and plasmids
The conditions for the growth of L. pneumophila strains (Lp01 WT and ∆dotA strains, thymidine auxotroph Lp02 pentuple and single island-depleting strains) were as described previously (Zuckman et al., 1999; Coers et al., 2000; Nagai et al., 2002; O’Connor et al., 2011; Choy et al., 2012). Briefly, the L. pneumophila strains were grown at 37°C in liquid N-(2-acetamido)-2-aminoethanesulfonic acid (Cat# 7365-82-4; Sigma-Aldrich)–buffered yeast extract media or on charcoal-yeast extract plates (Feeley et al., 1979) with or without appropriate antibiotics (100 µg/ml streptomycin, 10 µg/ml chloramphenicol, and 10 µg/ml kanamycin), as described previously (Berger et al., 1994). Thymidine (Cat# 203–19423; Wako Chemicals) was added to media at 100 µg/ml for the Lp02 and its derivative strains. (Berger and Isberg, 1993). The Lp01 ∆lpg2525 mutant was constructed by allelic exchanges as described previously (Zuckman et al., 1999). The deletion allele of lpg2525 was constructed using PCR; DNA fragments encoding regions of flanking homology that were immediately 5′ to the start codon and 3′ to the termination codon were generated. The two fragments were ligated and then inserted into the gene replacement vector pSR47S (Merriam et al., 1997). Recombination of the deletion allele onto the Lp01 chromosome was performed through conjugation between an Escherichia coli DH5αλpir strain (Zuckman et al., 1999) carrying a pSR47S-derived plasmid and the Lp01 strain (Roy and Isberg, 1997), The resulting deletion mutants were identified by the PCR analysis. To construct the L. pneumophila expression plasmids, pMMB207-3xmyc-lpg2525), the lpg2525 locus amplified from genomic DNA of Lp01 was cloned into the pMMB207-3xmyc vector (Kubori et al., 2022). For mammalian expression, lpg2525 DNA fragment or cDNAs encoding RabGAP-5, Rab5A, or Rab7 were inserted into the pEGFP (ClonTech), the pCMV-myc-N (Cat# 635689; ClonTech), or the pmRFP (ClonTech) vectors. For expression of 3x-FLAG fusion proteins, DNA locus encoding Lpg2525 or Rab5A was inserted into the 3x-FLAG vector (Ingmundson et al., 2007). Site-directed mutagenesis to construct S34N or Q79L of Rab5A and for truncation of the F-box domain (∆FB) of Lpg2525 (11–53 aa) was conducted by inverse PCR or by Gibson assembly. The sequences of primers for these cloning were as follows: Rab5A Fw (BamHI); 5′-CGCGGATCCATGGCTAATCGAGGAGCAACAAGACC-3′, Rab5A Rv (XhoI); 5′-CCGCTCGAGTCAGTTACTACAACACTGGCTTCTGG-3′, Rab5A (S34N) Fw; 5′-GTTGGCAAAAATAGCCTGGTTCTT-3′, Rab5A (S34N) Rv; 5′-AGCAGACTCTCCTAGAAGGACCAG-3′, Rab5A (Q79L) Fw; 5′-ACAGCTGGTCTGGAACGGTATCAT-3′, Rab5A (Q79L) Rv; 5′-ATCCCATATTTCAAATTTTACTGT-3′, Rab5A (K22R) Fw; 5′-TGCCAGTTCCGACTGGTACTT-3′, Rab5A (K22R) Rv; 5′-TATTTTATTTCCAGTATTTGG-3′, Rab5A (K33R) Fw; 5′-GCTGTTGGCCGATCAAGCCTA-3′, Rab5A (K33R) Rv; 5′-AGACTCTCCCAGAAGTACCAG-3′, Rab5A (K134R) Fw; 5′-TCAGGAAACCGGGCTGATCTT-3′, Rab5A (K134R) Rv; 5′-TAAAGCTATTACAATGTTAGG-3′, Rab7 Fw (EcoRV); 5′-CGCGATATCATGACCTCTAGGAAGAAAGTGTTGCT-3′, Rab7 Rv (XhoI); 5′-CCGCTCGAGCTAACAACTGCAGCTTTCTGCGGAGG-3′, Lpg2525 Fw (BglII); 5′-GGGAGATCTATGAAAGAAAAATACAAAAAG-3′, Lpg2525 Fw (SmaI); 5′-GGGCCCGGGTTATAAAATAGAACAAAAATT-3′, Lpg2525 (∆FB) Fw; 5′-TTTATCAAACGATTTTCCTATTCACCAACT-3′, Lpg2525 (∆FB) Rv; 5′-ATATAGACTCTTTTTGTATTTTTCTTTCAT-3′, RabGAP-5 Fw; 5′-ATGTCAGGAAGCCATACACCTGCC-3′, RabGAP-5 Rv, 5′-TCACCCGTCCACATCCCAGCTGAA-3′, lpg2525 d1; 5′-GCGAGCTCTAGCAGCCAAAGAACTGACTC-3′, lpg2525 d2; 5′-CTAATTTTACATATAGTTTAAAAAAAATC-3′, lpg2525 d3; 5′-ATATGTAAAATTAGAATAAATATATGAAC-3′, lpg2525 d4; 5′-GCTCTAGAGCCATCTCCAACTGTTAAAG-3′, lpg2525ΔF-box_1; 5′-ATATAGACTCTTTTTGTATTTTTCTTTCAT-3′, lpg2525ΔF-box_2; 5′-CAAAAAGAGTCTATATTTTATCAAACGATTTTCCTATTCAC-3′, lpg2525 Fw (BamHI); 5′-GCGGATCCTCATGAAAGAAAAATACAAAAAGAGTC-3′, lpg2525 Rv (PstI); 5′-CGCTGCAGTTATAAAATAGAACAAAAATTGTTTG-3′, RabGAP-5 Fw (EcoRI); 5′-CCCGAATTCGGATGTCAGGAAGCCATACACCTGCC-3′, RabGAP-5 Rv (XhoI); 5′-GCCTCGAGTCACCCGTCCACATCCCAGCTG-3′. HA-tagged UB constructs (WT, K48R, and K63R) were generous gifts from Dr. Shigeru Yanagi (Gakushuin University, Toshima, Tokyo, Japan).
Cell culture and transfection
HEK293 cells stably expressing FcγRII (HEK293-FcγRII cells) were grown in grown in DMEM supplemented with 50 IU/ ml penicillin, 50 μg/ml streptomycin, 10% fetal calf serum, and 400 μg/ml hygromycin (Cat# 080–07683; Wako Chemicals). HeLa cells stably expressing FcγRII (HeLa-FcγRII cells) were grown in α-MEM supplemented with 50 IU/ ml penicillin, 50 μg/ml streptomycin, 10% fetal calf serum, 2 mM L-glutamine, and 400 μg/ml hygromycin (Cat# 080-07683; Wako Chemicals) (Arasaki and Roy, 2010; Arasaki et al., 2017, 2018). Both HeLa- and HEK293-FcγRII cells were confirmed to be free from mycoplasma contamination. Plasmid transfection into the cells was performed using polyethylenimine (24765-2; Polysciences), and 1 µg of plasmid was used for transfection. The siRNAs were synthesized by Japan Bio Services, and the target sequences of the siRNAs were follows: Cullin1; 5′-CUAGAUACAAGAUUAUACAUGCGG-3′, SKP1; 5′-GCAAGUCAAUUGUAUUAGCAGAAU-3′, Rab7A; 5′-GGAUGACCUCUAGGAAGAA-3′, and RabGAP-5; purchased from SantaCruz (Cat# sc-76492). Transfection of siRNAs was performed using oligofectamine, according to the manufacture’s protocol.
Production of anti-Legionella antiserum
To produce anti-Legionella rabbit or mouse antiserum, heat-killed L. pneumophila were mixed with Freund’s incomplete adjuvant (Cat# 011–09551; Wako Chemicals) and injected into Japan White rabbits or BALB/c mice every 3 wk. Titer of the antiserums were assessed by internalization efficiency of L. pneumophila opsonized by these antiserums.
Preparation of L. pneumophila–infected cells
Both HeLa- and HEK293-FcγRII cells were seeded at a density of 1 × 106 cells per well of a 6-well plate on a previous day of infection or transfection. L. pneumophila opsonized with rabbit (dilution: 1/3,000) or mouse (dilution; 1/1,000) anti-L. pneumophila antisera was used for infection of the cells at the required MOI. At 10 min after infection, the plate was centrifuged at 200 g for 5 min at room temperature (centrifuge 5430; Eppendorf). After centrifugation, the plate was placed in a CO2 incubator for the required infection time, and the cells were extensively washed with phosphate-buffered saline prior to processing. In the case of infection of HEK 293-FcγRII cells, poly-l-lysine (MW 70,000–150,000; p1274; Sigma-Aldrich)–coated tissue culture plates were used.
Immunofluorescence
Cells were fixed with 4% paraformaldehyde/phosphate-buffered saline for 20 min at room temperature. After fixation, samples were stained with fluorescent-conjugated anti-rabbit or mouse secondary antibodies to detect the extracellular bacteria that were opsonized by rabbit or mouse antisera prior to permeabilization. After staining extracellular bacteria, cells were permeabilized with 0.2% Triton X-100 for 15 min at room temperature and further stained intracellular bacteria with DAPI (S36964; Invitrogen) and host proteins (Arasaki et al., 2017, 2018). The KEYENCE BZ-X810 microscope equipped with a 100× oil-immersion lens (NA 1.45) was used to obtain images. Microscopic observations were performed at room temperature.
IP
HEK293- or HeLa-FcγRII cells expressing tagged proteins were lysed in a buffer containing 25 mM HEPES-KOH (pH7.4) (342–01375; DOJINDO), 150 mM NaCl (195–01663; Wako), 2.5 mM ethylenediamine-N,N,N′,N′-tetraacetic acid (345–01865; DOJINDO), 1 µg/ml leupeptine (4041; Peptide Institute), 1 µM pepstatin A (4397; Peptide Institute), 2 µg/ml aprotinin (10236624001; Roche), 10 mM N-ethylmaleimide (058–02061; Wako), and 1% Triton X-100 (162–24755; Wako) for 15 min at 4°C. After lysis, samples were centrifuged, and the supernatants were subjected to IP with an anti-FLAG M2 affinity gel (A2220; Sigma-Aldrich). The bound proteins were eluted with 100 µg/ml 3x-FLAG peptides (F4799; Sigma-Aldrich) or SDS sample buffer, subjected to SDS-polyacrylamide gel electrophoresis, and analyzed by immunoblotting.
SDS-polyacrylamide gel electrophoresis, western blotting, and immunoblotting
Samples were loaded onto 12% polyacrylamide gels, and electrophoresis was conducted at 180 V for 60 min. After electrophoresis, proteins were transferred onto polyvinylidene fluoride membranes (IPVH00010; Merck Millipore) at 400 mA for 25 min. The membranes were blocked with 2% bovine serum albumin (017–15124; Wako) or 5% skim milk in TBS-T buffer (20 mM Tris-HCl, 137 mM NaCl, and 0.1% Tween-20) for 1 h at room temperature and then incubated with primary antibodies. After incubation, the membranes were washed with TBS-T buffer and incubated with secondary antibodies conjugated to horseradish peroxidase for 1 h at room temperature. The horseradish peroxidase signal on the membranes was detected by enhanced chemiluminescence (WBLUC0500; Merck Millipore). Images of immunoblots were obtained using LumiVision Pro image analyzer (TAITEC AISIN). The quantification of immunoblots was conducted using ImageJ/Fiji software.
Quantification and statistical analysis
GraphPad Prism version 10 was used to draw all graphs and conduct all statistical analyses.
Online supplemental material
Fig. S1 shows the representative immunofluorescent images of endocytic markers, indicating either LCV-positive or -negative staining, and the detection of endogenous ubiquitination of Rab5 during L. pneumophila infection. Fig. S2 shows the identification of the island-encoded Legionella effector responsible for ubiquitinating Rab5. Fig. S3 shows the nucleotide state-dependent recognition of Rab5 by Lpg2525 and the impact of Cullin1/SKP1 silencing on Lpg2525-mediated Rab5 ubiquitination. Fig. S4 shows the impact of ectopic expression of Lgp2525 on endosomal morphology and kinetic localization of RabGAP-5 and Lpg2525 to the LCV. Fig. S5 represents the sequence alignment of Rab5 from multiple species and examines the impact of substituting conserved lysine residues in Rab5 on its association with the LCV.
Data availability
All the data underlying this study are available in the published article and its online supplemental material.
Acknowledgments
We are grateful to Dr. Shigeru Yanagi (Gakushuin University, Toshima, Tokyo, Japan) for generous gifts of plasmids encoding HA-UB (WT), (K48R), and (K63R). We thank Misaki Tsuruoka, Tatsuyuki Oka, and Dr. Tomoe Kitao for technical assistance.
This work was supported in part by Grant-in-Aid for Scientific Research, #24790425, #26713016, #18H02656, and #20H05772 (to K. Arasaki); #19H03469 and #22H02867/23K24129 (to T. Kubori); and #23H02716/23K27407 (to H. Nagai) from the Ministry of Education, Culture, Sports, Science and Technology of Japan, and the Uehara Memorial Foundation (to K. Arasaki), the Naito Foundation (to K. Arasaki and H. Nagai), the Astellas Foundation for Research on Metabolic Disorders (to K. Arasaki), and the Takeda Science Foundation (to K. Arasaki and T. Kubori).
Author contributions: S. Tanaka: investigation and methodology. H. Oide: investigation and methodology. S. Ikeda: investigation and methodology. M. Tagaya: supervision and writing—review and editing. H. Nagai: conceptualization, funding acquisition, and writing—review and editing. T. Kubori: conceptualization, funding acquisition, investigation, resources, and writing—original draft, review, and editing. K. Arasaki: conceptualization, data curation, formal analysis, funding acquisition, investigation, methodology, project administration, resources, supervision, validation, visualization, and writing—original draft, review, and editing.
References
Author notes
S. Tanaka and H. Oide contributed equally to this paper.
Disclosures: The authors declare no competing interests exist.



![Effect of large genomic islands depletion in the L. pneumophila genome on Rab5 recruitment to the LCV during infection. (a) Schematic representation of the ∆pentuple and single island-depleted strains (adopted from O’Connor et al. [2011]). (b) HeLa-FcγRII cells expressing mRFP-Rab5A were infected with the indicated L. pneumophila Lp02 strains for 1 h at an MOI of 20. After infection, cells were fixed and stained with anti-Legionella antiserum for detection of extracellular bacteria, then permeabilized and further stained with DAPI for detection of intracellular bacteria. RFP-Rab5A–positive LCVs were quantified. Representative data of three independent experiments were presented (100 vacuoles were scored in each experiment). Results are shown as means ± SD. P values are determined using one-way ANOVA with Tukey’s multiple comparisons. ****P < 0.0001. (c) List of L. pneumophila effectors encoded in the island 7a. (d) HeLa-FcγRII cells were transfected without (mock) or with siRNA-targeting Rab7. At 72 h after transfection, the efficacy of siRNA was assessed by IB with the Rab7 antibody. IB with α-tubulin antibody was conducted as an internal control. IB, immunoblotting. Source data are available for this figure: SourceData FS2. Refer to the image caption for details.](https://cdn.rupress.org/rup/content_public/journal/jcb/224/4/10.1083_jcb.202406159/2/m_jcb_202406159_figs2.png?Expires=1781626478&Signature=sUFuamUvDqI1dFDKfhNltIhxMtbt3yVXMZUtzMGaxCAau5hl~T3fi~1UM6boRmqbcD7ul~tPV7fv7ySXei8Kh-jANLHWr31IbLBVsL~8SIdPjk3jZFdr9dNRVtRwE-UeWbzWmL5mkUm3PuSD5OoSlJeaPCc~Hy-hOSyhToz8Q09PDghGChw2SBIF3bfhBUf3ukiXiRS76o9G0Zs3WCuzHx2eQJacaINOclMCZyQ~KSYaQArZv54udk1g2pCI5JM3GAsy~wuOFq8kmAdRr0hb9UanERIC-SDe5oEJ0vPZQ9gPUDwpyWNRq7SlmNXUMCJoaApKmjCPxOLu-Ig-ZD53fA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)