Muscle nuclei are exposed to variable cytoplasmic strain produced by muscle contraction and relaxation, but their morphology remains stable. Still, the mechanism responsible for maintaining myonuclear architecture, and its importance, is currently elusive. Herein, we uncovered a unique myonuclear scaffold in Drosophila melanogaster larval muscles, exhibiting both elastic features contributed by the stretching capacity of MSP300 (nesprin) and rigidity provided by a perinuclear network of microtubules stabilized by Shot (spectraplakin) and EB1. Together, they form a flexible perinuclear shield that protects myonuclei from intrinsic or extrinsic forces. The loss of this scaffold resulted in significantly aberrant nuclear morphology and subsequently reduced levels of essential nuclear factors such as lamin A/C, lamin B, and HP1. Overall, we propose a novel mechanism for maintaining myonuclear morphology and reveal its critical link to correct levels of nuclear factors in differentiated muscle fibers. These findings may shed light on the underlying mechanism of various muscular dystrophies.
Introduction
Alterations in nuclear shape and positioning are often associated with changes in the behavior of cells (Starr and Fridolfsson, 2010). Recent studies proposed that tissue stiffness affects the levels of the nuclear lamina protein lamin A/C, resulting in altered transcriptional regulation (Simon and Wilson, 2011; Burke and Stewart, 2013; Zwerger et al., 2013; Fedorchak et al., 2014; Swift and Discher, 2014).
Muscles exhibit inherently altered biomechanical properties and therefore should develop a specific strategy to keep nuclear morphology intact. Myonuclei in striated muscles face variable cytoplasmic strain exerted by the tightly associated sarcolemma and the axial contractile forces. How the myonuclei resist these forces has yet to be elucidated.
Recent evidence has revealed the unique contribution of the microtubule (MT)-based cytoskeleton to nuclear shape and positioning in muscle fibers (Oddoux et al., 2013; Wilson and Holzbaur, 2015). In addition, various MT-associated proteins (MAPs), including dynein, kinesin, and MAP7/Esconsin, were shown to be essential for myonuclear positioning in striated muscles (Folker et al., 2012, 2014; Metzger et al., 2012; Wilson and Holzbaur, 2012). Furthermore, proteins of the linker of nucleoskeleton and cytoskeleton (LINC) complex, including the Klarsicht, ANC-1, Syne homology (KASH) proteins MSP300 and Klar and the Sad1p, UNC-84 (SUN) protein Klaroid (Elhanany-Tamir et al., 2012), are critical for maintenance of myonuclear positioning and shape. However, a direct connection between LINC proteins and the MT network has yet to be established.
In this work, we identify two additional cytoplasmic components, namely EB1 and Shot, that function together with MSP300 to maintain myonuclear morphology. Importantly, we show that the MSP300 protein exhibits elastic properties, helping the myonuclei to resist cytoplasmic strain. Finally, we demonstrate a link between the aberrant nuclear architecture detected in mutant muscles and altered levels of nuclear regulatory proteins.
Results and discussion
Identification of novel factors mediating myonuclear shape
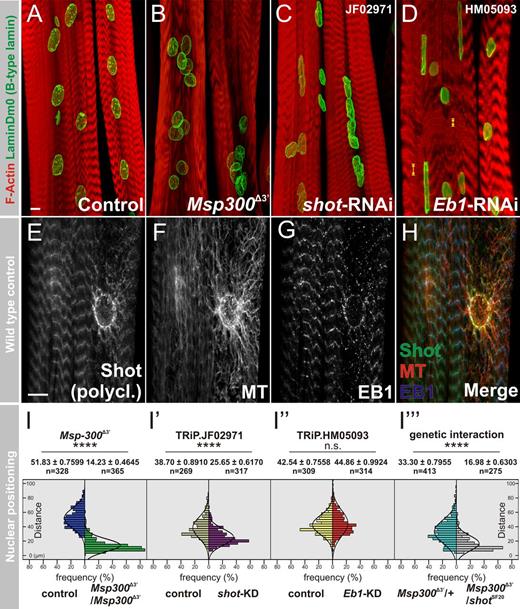
To identify novel elements mediating nuclear positioning and shape in Drosophila melanogaster larval muscles, an RNAi-based screen of 58 candidate genes expressed in myogenic tissues was performed using a muscle-specific driver. Results showed that relative to the control (Fig. 1 A), the myonuclei in both short stop (shot) and Eb1 knockdown (KD) larvae exhibited abnormal shape reminiscent of Msp300 mutants (Fig. 1, B–D). In shot-RNAi larvae, 90% (n = 80 muscles) of the ds-RNA line JF02971 and 86.2% (n = 80 muscles) of the ds-RNAi line GL01286 exhibited defective nuclear positioning, abnormal shape, or both. In Eb1-RNAi lines, 63.11% (n = 507 muscles) of ds-RNA line HM05093 and 51.5% (n = 240 muscles) of ds-RNAi line GL00559 exhibited abnormal nuclear shape phenotype. In comparison, a control using UAS-VALIUM10-luciferase displayed only 5.2% (n = 487 muscles) aberrant nuclear morphology.
Shot and EB1 are essential to maintain myonuclear morphology. (A–D) Larval muscles labeled with phalloidin (red) and laminDm0 (green) in control (A), Msp300Δ3′ (B), muscle-specific knockdown of shot (C) or Eb1 (D) using Mef2-Gal4 driver; maximal projections are shown. (E–H) Single optical sections of larval muscles triple-labeled with anti-Shot (E and H, green), anti–α-tubulin (F and H, red), and anti-EB1 (G and H). Bars, 10 µm. Yellow arrowheads indicate aberrant distance between Z-bands (see also Fig. S1 B). (I–I‴) Measurements of internuclear distances in Msp300Δ3′ (green versus control in blue), shot-KD (violet versus control in beige), Eb1-KD (red versus control in yellow), and Msp300Δ3/shotSF20 trans-heterozygotes (gray versus control Msp300Δ3/CyO heterozygotes in green) are given.
Shot and EB1 are essential to maintain myonuclear morphology. (A–D) Larval muscles labeled with phalloidin (red) and laminDm0 (green) in control (A), Msp300Δ3′ (B), muscle-specific knockdown of shot (C) or Eb1 (D) using Mef2-Gal4 driver; maximal projections are shown. (E–H) Single optical sections of larval muscles triple-labeled with anti-Shot (E and H, green), anti–α-tubulin (F and H, red), and anti-EB1 (G and H). Bars, 10 µm. Yellow arrowheads indicate aberrant distance between Z-bands (see also Fig. S1 B). (I–I‴) Measurements of internuclear distances in Msp300Δ3′ (green versus control in blue), shot-KD (violet versus control in beige), Eb1-KD (red versus control in yellow), and Msp300Δ3/shotSF20 trans-heterozygotes (gray versus control Msp300Δ3/CyO heterozygotes in green) are given.
Shot, the only Drosophila spectraplakin (Suozzi et al., 2012), and EB1 are both MAPs that were previously shown to form a protein complex where EB1 is an MT plus end–binding protein, and Shot contains N-terminal calponin homology (CH) domains and C-terminal MT and EB1 binding domains (Subramanian et al., 2003; Slep et al., 2005; Alves-Silva et al., 2012). The distance between myonuclei in shot-KD, but not in Eb1-KD muscles, was decreased so that the nuclei often formed aggregates (Fig. 1, C and I′; compare with Fig. 1 I″), which is reminiscent of Msp300 (Fig. 1 B, I) and klar mutant muscles (Elhanany-Tamir et al., 2012). In addition, whereas the sarcomeric organization of shot-KD muscles was relatively normal, Eb1-KD muscles exhibited irregularity in the distances between Z-bands (Fig. 1 D and Fig. S1, A and B).
Examination of the relative spatial distribution of Shot, EB1, and the MT network showed unequivocal colocalization of Shot and EB1 at the perinuclear region (Fig. 1, E–H). The MT exhibited an elaborate perinuclear network and both Shot and EB1 distributions overlapped along the nuclear envelope. These results implicated Shot and EB1 as essential components required for the maintenance of myonuclear shape in muscles.
Shot and EB1 are required to maintain the perinuclear MT network and to inhibit perinuclear F-actin distribution
Further analysis of the nuclear shape indicated that in contrast to the flat and oval shape of control myonuclei, both Eb1-KD and shot-KD muscles contained nuclei that on average were significantly more spherical and exhibited a high degree of variability in their dimensions (Fig. 2, A and B). Examination of the MT perinuclear network revealed that in both Eb1-KD and shot-KD muscles, MT distribution was disarranged (Fig. 2, C–E). In these muscles, fewer MT filaments were detected and individual MT filaments often appeared bent (Fig. 2, C–E and L). The disruption in the perinuclear MT network in both mutants correlated with the degree of abnormality in nuclear morphology.
Shot and EB1 recruit a perinuclear MT scaffold and inhibit perinuclear sarcomeric F-actin. (A and B) Quantification of nuclear sphericity (A) and nuclear dimensions (B) in control, Eb1-KD, and shot-KD muscles. (C–E) Perinuclear MT in control (C), muscle-specific Eb1-KD (D), or shot-KD (E) colabeled with anti–α-tubulin (white) and anti-lamin B/Dm0 (blue). Yellow arrows indicate aberrant MT distribution. (F–K) Ectopic sarcomeric F-actin labeled with phalloidin (white) in control (F and I), Eb1-KD (G and J), and shot-KD (H and K) muscles; the nuclei are labeled with anti-lamin B/Dm0 (blue). I, J, and K are orthogonal views of F, G, and H. Yellow arrowheads indicate ectopic actin. (L) Quantification of MT filaments per nuclear area. Bars, 10 µm. ***, P < 0.001.
Shot and EB1 recruit a perinuclear MT scaffold and inhibit perinuclear sarcomeric F-actin. (A and B) Quantification of nuclear sphericity (A) and nuclear dimensions (B) in control, Eb1-KD, and shot-KD muscles. (C–E) Perinuclear MT in control (C), muscle-specific Eb1-KD (D), or shot-KD (E) colabeled with anti–α-tubulin (white) and anti-lamin B/Dm0 (blue). Yellow arrows indicate aberrant MT distribution. (F–K) Ectopic sarcomeric F-actin labeled with phalloidin (white) in control (F and I), Eb1-KD (G and J), and shot-KD (H and K) muscles; the nuclei are labeled with anti-lamin B/Dm0 (blue). I, J, and K are orthogonal views of F, G, and H. Yellow arrowheads indicate ectopic actin. (L) Quantification of MT filaments per nuclear area. Bars, 10 µm. ***, P < 0.001.
This suggests that MTs play a dual role in controlling muscle nuclei: to maintain nuclear shape, which depends on both Shot and EB1 function, and to regulate nuclear position within the muscle fiber, requiring Shot but not EB1 function.
Interestingly, in both shot-KD and Eb1-KD muscles, intense accumulation of ectopic sarcomeric F-actin was observed in the vicinity of the myonuclei (Fig. 2, F–H; and Fig. S1 C). This indicates that Shot and EB1, as well as the perinuclear MT organization, inhibit sarcomeric actin distribution at the vicinity of the myonuclei.
Shot protein was previously shown to cross-link MTs with the actin-based cytoskeleton (Lee and Kolodziej, 2002; Applewhite et al., 2010). However, our analysis reveals that in muscles, Shot does not function as an MT-actin cross-linker, but rather contributes to recruiting MTs and excluding F-actin from the perinuclear region. A recent report demonstrated that Shot N-terminal CH-actin–binding domain is auto-inhibited by its C-terminal GAS2 domain when bound to EB1 (Applewhite et al., 2013). Consistent with this report, the perinuclear Shot–EB1–MT complex might inhibit interaction of Shot with F-actin at this region.
Collectively, these results indicate that the Shot–EB1 complex maintains nuclear morphology by mediating the assembly of the MT perinuclear network and repressing the assembly of sarcomeric F-actin in this region.
Perinuclear distribution of MSP300 depends on Shot function
MSP300 is essential for the organization of the MT network (Elhanany-Tamir et al., 2012). We therefore proceeded to examine whether a functional link between Shot and MSP300 exists. Co-staining of wild-type (WT) larval muscles with anti-Shot and anti-MSP300 antibodies showed partial overlap in the perinuclear region (Fig. 3, A–C). It is noteworthy that the monoclonal anti-Shot (mAbRod1; Lee et al., 2003) may only detect a subset of all 22 annotated isoforms (Fig. S2), explaining the discrepancy in staining patterns between polyclonal (Fig. 1 E) and monoclonal (Fig. 3 A) anti-Shot antibodies. Furthermore, muscles of double-heterozygous Msp300Δ3′/shotSF20 larvae showed specific defects in myonuclear shape and spacing relative to control heterozygous Msp300Δ3′/CyO or shotSF20/+ muscles (Fig. 3, D–F; and Fig. 1 I‴), which indicates functional cooperation between both gene products.
MSP300 perinuclear distribution depends on Shot function. (A–C) WT myonucleus double-labeled with Shot (A, green) and MSP300 (B, red) and their overlay (C) detected in a single optical section. (D–F) Larval muscles labeled with phalloidin (red) and Lamin B/Dm0 (green) of shot/CyO heterozygote (D), Msp300/CyO heterozygote (E), and shot/Msp300 trans-heterozygous larvae (F). (G–Q) Larval muscles labeled with anti-MSP300 (red; G, H, I, J, K, and L) and LaminDm0 (green), or with anti-Shot (red; M, N, O, P, Q, and R) and LaminDm0 (green) of control (G, H, I, I′, M, N, O, and O′), shot-muscle KD (J, K, L, and L′), or Msp300 hypomorphic alleles (Msp300MB00410; P, Q, R, and R′). Corresponding orthogonal sections are shown in H, K, N, and Q, respectively. (I–R’) Higher magnifications of single nucleus; black and white images are corresponding images of either MSP300 (I′ and L′) or Shot (O′ and R′). Bars, 10 µm.
MSP300 perinuclear distribution depends on Shot function. (A–C) WT myonucleus double-labeled with Shot (A, green) and MSP300 (B, red) and their overlay (C) detected in a single optical section. (D–F) Larval muscles labeled with phalloidin (red) and Lamin B/Dm0 (green) of shot/CyO heterozygote (D), Msp300/CyO heterozygote (E), and shot/Msp300 trans-heterozygous larvae (F). (G–Q) Larval muscles labeled with anti-MSP300 (red; G, H, I, J, K, and L) and LaminDm0 (green), or with anti-Shot (red; M, N, O, P, Q, and R) and LaminDm0 (green) of control (G, H, I, I′, M, N, O, and O′), shot-muscle KD (J, K, L, and L′), or Msp300 hypomorphic alleles (Msp300MB00410; P, Q, R, and R′). Corresponding orthogonal sections are shown in H, K, N, and Q, respectively. (I–R’) Higher magnifications of single nucleus; black and white images are corresponding images of either MSP300 (I′ and L′) or Shot (O′ and R′). Bars, 10 µm.
Further analysis revealed that the distribution of MSP300 at the nuclear ring and between proximal nuclei was severely impaired in shot-KD muscles (Fig. 3, G–L′), whereas its localization at the Z-bands was still prominent (Fig. 3 J and Fig. S1). This might reflect a differential sensitivity of distinct protein isoforms of MSP300 to Shot activity. In a reciprocal experiment, Shot perinuclear distribution was examined in mutant larvae of the hypomorphic Msp300MB00410 allele, which was used to eliminate possible secondary effects from the severely defective muscles of Msp300Δ3′. In these muscles, Shot perinuclear distribution was partially detected, but its typical perinuclear filamentous distribution vanished (Fig. 3, M–R′). Collectively, these results suggest that MSP300 perinuclear distribution is stabilized by Shot function, whereas Shot association with the nuclear membrane is partially independent of MSP300, and is possibly maintained by its interaction with the perinuclear MT and EB1.
Myonuclear morphology is maintained during application of external force on larval muscles
Muscles are exposed to variable cytoplasmic strain during contraction–relaxation waves. To address the contribution of MSP300 and Shot to myonuclear morphology under strain, we tested the effect of tensile stresses on nuclear morphology. For a stress test, intact larvae were stretched to up to 1.5-fold their normal length and analyzed (see Materials and methods for details).
Fig. 4 shows that the distribution of the perinuclear MT network in stretched muscles was similar to that of control muscles (Fig. 4, A′ and B′). In contrast, the perinuclear ring of MSP300 expanded in the direction of muscle stretching, reflecting the anisotropic strain applied on the nucleus (Fig. 4, A and B, arrowheads). This suggested that whereas perinuclear MSP300 responds to external strain, the perinuclear MT network retains its position. Comparison of nuclear dimensions of stretched versus nonstretched larvae (Fig. 4 C) revealed no significant alternations in the aspect ratio of myonuclei, although its distribution was slightly reduced in stretched larvae, possibly due to a limited degree of freedom induced by muscle stretching. Nuclear resistance to stretching implies the existence of a mechanism that protects and maintains nuclear morphology during the application of external force.
The effect of muscle stretching on nuclear shape and MSP300 conformation. (A–B″) Representative nuclei from nonstretched (A–A″) versus stretched (B–B″) larval muscle labeled with MSP300 (red; A, A″, B, and B″), anti–α-tubulin (green; A′, A″, B′, and B″), and lamin B/Dm0 (blue). Deconvolution was applied to A–B″. Arrowheads in A and B indicate the MSP300 nuclear ring. (C and D) Scatter plots of myonuclear dimensions of nonstretched (black/blue dots) and stretched (red/green dots) muscles of WT (C) and shot-KD (D). The slope (linear regression) of the dot plots indicates the aspect ratio: for control, neutral n = 334, stretched n = 335; for shot-KD, neutral n = 200, stretched n = 232. The assays were performed at least twice for each group. (G–O) High magnification of anti-MSP300 (white) staining pattern in nonstretched (G and H) versus stretched (I, J, and K) WT muscle. (F) Quantification of the distance between the dots of MSP300 spectrin repeat epitopes in neutral and stretched larvae (model in E). The mean distance increases from 634 ± 4.09 nm to 759 ± 4.76 nm in stretched larvae (mean ± SEM, n = 605 each, P < 0.0001). Centered lines show the medians, box limits indicate the 25th and 75th percentiles as determined by R software, whiskers extend to the 5th and 95th percentiles, and outliers are represented by dots. Notches indicate ∼95% confidence that two medians differ significantly. (L–O) Perpendicular MSP300 filaments emanate from the nuclear membrane to the peripheral MSP300 ring in WT (L and M) and shot-KD (N and O) myonuclei.
The effect of muscle stretching on nuclear shape and MSP300 conformation. (A–B″) Representative nuclei from nonstretched (A–A″) versus stretched (B–B″) larval muscle labeled with MSP300 (red; A, A″, B, and B″), anti–α-tubulin (green; A′, A″, B′, and B″), and lamin B/Dm0 (blue). Deconvolution was applied to A–B″. Arrowheads in A and B indicate the MSP300 nuclear ring. (C and D) Scatter plots of myonuclear dimensions of nonstretched (black/blue dots) and stretched (red/green dots) muscles of WT (C) and shot-KD (D). The slope (linear regression) of the dot plots indicates the aspect ratio: for control, neutral n = 334, stretched n = 335; for shot-KD, neutral n = 200, stretched n = 232. The assays were performed at least twice for each group. (G–O) High magnification of anti-MSP300 (white) staining pattern in nonstretched (G and H) versus stretched (I, J, and K) WT muscle. (F) Quantification of the distance between the dots of MSP300 spectrin repeat epitopes in neutral and stretched larvae (model in E). The mean distance increases from 634 ± 4.09 nm to 759 ± 4.76 nm in stretched larvae (mean ± SEM, n = 605 each, P < 0.0001). Centered lines show the medians, box limits indicate the 25th and 75th percentiles as determined by R software, whiskers extend to the 5th and 95th percentiles, and outliers are represented by dots. Notches indicate ∼95% confidence that two medians differ significantly. (L–O) Perpendicular MSP300 filaments emanate from the nuclear membrane to the peripheral MSP300 ring in WT (L and M) and shot-KD (N and O) myonuclei.
High-resolution analysis of MSP300 distribution before stretching revealed reiterated dots evenly spaced along thin filaments (Fig. 4, F, G, and H). Since the antibody for MSP300 was raised against a region with multiple spectrin repeats (Fig. S2; Volk, 1992), we reasoned that these dots might represent reiterated spectrin-repeat epitopes distributed along filaments of MSP300 oligomers (see scheme in Fig. 4 E). Strikingly, after larval stretching, the distance between these dots increased by 1.2-fold (Fig. 4, I, J, K, and F), indicating that the MSP300 filaments exhibit elastic properties, extending in response to strain (Fig. 4 E). Of note, mammalian Nesprin isoforms contain multiple spectrin repeats (Lu et al., 2012; Rajgor and Shanahan, 2013; Luxton and Starr, 2014), which might similarly exhibit elastic properties required for myonuclear protection during muscle contraction.
The MSP300 filaments were arranged as a tangential perinuclear ring (Fig. 4 K, yellow curve) as well as radial filaments emanating from the nuclear membrane toward the ring (Fig. 4, L and M, yellow arrowheads). This organization might provide force insulation for the myonuclei during muscle contraction (see model in Fig. 5).
Because Msp300Δ3′ mutant larvae were too fragile, it was technically unfeasible to assess the effect of muscle stretching on their myonuclear morphology. Instead, we analyzed this effect in shot-KD muscles. In these muscles, the perpendicular MSP300 filaments emerging from the nuclear envelope were partially missing (Fig. 4, N and O), and therefore the elastic shield of MSP300 was removed. The nuclear dimensions were variable even before larval stretching and were not significantly altered after stretching (Fig. 4 D, blue versus green dots). Collectively, these results indicate that the combination of Shot-dependent perinuclear distribution of elastic MSP300 filaments and the rigid MT network is indispensable for the maintenance of intact myonuclear morphology.
Aberrant myonuclear shape alters the nuclear distribution of Lamin A/C, Lamin B, and HP1
Next, we examined the relation between nuclear morphology and the levels of the nuclear factors lamin A/C, lamin B (Dm0), and Heterochromatin Protein 1 (HP1), all of which contribute to the chromatin state of the nucleus. To assess nuclear distribution of these factors, mean levels of fluorescent intensity were calculated per nuclear area. We analyzed homozygous klarΔ1–18 (null allele), strong (Msp300Δ3′) as well as mild Msp300 (Msp300MB00410) alleles, and shot-KD mutant muscles, all of which displayed variable degree of aberrant nuclear morphology. Representative examples are shown in Fig. 5 (A–H). Our measurements revealed a decrease of lamin A/C levels in Msp300 as well as in klar mutants. The most severe reduction was observed in Msp300Δ3′. Interestingly lamin A/C appeared in aggregates in klar mutant muscles, implicating aberrant distribution (Fig. 5 F). Lamin B levels dropped only in the Msp300 mutants, whereas in klar they were slightly elevated. HP1 nuclear levels were dramatically decreased in klar (∼60% of control) as well as in both Msp300 alleles (35% of control in the strong and 75% in the mild allele; Fig. 5 A, data for Msp300MB00410 is not shown). Interestingly, in shot-KD muscles the HP1 level remained unchanged, whereas Lamin A/C and B levels were slightly increased (Fig. 5 J).
Aberrant nuclear morphology correlates with changes in HP1 and Lamins. (A–H) Representative myonuclei of control (A and E), klar (B and F), Msp300 (C and G), and shot-KD (D and H) labeled with HP1 (red) and DAPI (blue; A–D), or with anti-lamin A/C (green; E–J). Bars, 10 µm. (I and J) Quantification of the nuclear fluorescent intensities of HP1, lamin A/C, and lamin Dm0 in control, klar, and Msp300 (I) or in control and shot-KD (J) mutant muscles. The values are as follows. For HP1, WT 138.2 ± 2.070, n = 320; klarΔ1-18 83.51 ± 1.803, n = 316; Msp300Δ3 48.93 ± 1.724, n = 319; P < 0.0001. For Lamin A/C, WT 142.8 ± 1.791, n = 287; klarΔ1-18 128.0 ± 2.756, n = 295; Msp300Δ3’ 94.72 ± 2.113, n = 298; P < 0.0001. For Lamin Dm0, WT 112.7 ± 2.147, n = 306; klarΔ1-18 130.1 ± 1.782, n = 300; Msp300Δ3’ 90.66 ± 1.532, n = 302; P < 0.0001. For HP1, WT 104.5 ± 1.526, n = 303; shot-KD 104.0 ± 1.523, n = 303; no significant difference. For Lamin A/C, WT 115.7 ± 1.444, n = 309; shot-KD 137.3 ± 2.070, n = 309; P < 0.0001; for Lamin Dm0, WT 113.2 ± 1.878, n = 303; shot-KD 139.5 ± 1.534, n = 304. P < 0.0001. ***, P < 0.0001. (K and L) A schematic model describing the cytoskeletal elements required for maintenance of myonuclear shape. (K) The lower half of the myonucleus facing the sarcomeric myofilaments, and surrounded by MSP300 nuclear ring attached to the nucleus through KASH–SUN interactions. (L) The upper half of the myonucleus, facing the sarcolemma, covered by a web of MTs associated with the outer nuclear membrane by Shot, EB1, and MSP300. The direction of contraction is indicated by the arrows.
Aberrant nuclear morphology correlates with changes in HP1 and Lamins. (A–H) Representative myonuclei of control (A and E), klar (B and F), Msp300 (C and G), and shot-KD (D and H) labeled with HP1 (red) and DAPI (blue; A–D), or with anti-lamin A/C (green; E–J). Bars, 10 µm. (I and J) Quantification of the nuclear fluorescent intensities of HP1, lamin A/C, and lamin Dm0 in control, klar, and Msp300 (I) or in control and shot-KD (J) mutant muscles. The values are as follows. For HP1, WT 138.2 ± 2.070, n = 320; klarΔ1-18 83.51 ± 1.803, n = 316; Msp300Δ3 48.93 ± 1.724, n = 319; P < 0.0001. For Lamin A/C, WT 142.8 ± 1.791, n = 287; klarΔ1-18 128.0 ± 2.756, n = 295; Msp300Δ3’ 94.72 ± 2.113, n = 298; P < 0.0001. For Lamin Dm0, WT 112.7 ± 2.147, n = 306; klarΔ1-18 130.1 ± 1.782, n = 300; Msp300Δ3’ 90.66 ± 1.532, n = 302; P < 0.0001. For HP1, WT 104.5 ± 1.526, n = 303; shot-KD 104.0 ± 1.523, n = 303; no significant difference. For Lamin A/C, WT 115.7 ± 1.444, n = 309; shot-KD 137.3 ± 2.070, n = 309; P < 0.0001; for Lamin Dm0, WT 113.2 ± 1.878, n = 303; shot-KD 139.5 ± 1.534, n = 304. P < 0.0001. ***, P < 0.0001. (K and L) A schematic model describing the cytoskeletal elements required for maintenance of myonuclear shape. (K) The lower half of the myonucleus facing the sarcomeric myofilaments, and surrounded by MSP300 nuclear ring attached to the nucleus through KASH–SUN interactions. (L) The upper half of the myonucleus, facing the sarcolemma, covered by a web of MTs associated with the outer nuclear membrane by Shot, EB1, and MSP300. The direction of contraction is indicated by the arrows.
These results indicate that aberrant nuclear morphology leads to altered nuclear levels of essential regulatory factors. Significantly, mutants of both KASH proteins Msp300 and klar exhibited a reduction in HP1 and lamin A/C levels, whereas in shot-KD muscles an opposite tendency was observed. As a possible explanation, based on observed nuclear phenotypes, we postulate that in both klar and Msp300 mutant muscles the myonuclei lose their association with the sarcomeres facing reduced cytoplasmic strain, leading to decreased lamin A/C and HP1 levels. However, in shot-KD muscles, the nuclei are partially attached to the sarcomeres facing elevated cytoplasmic strain, leading to the increased level of lamin A/C.
In summary, our results demonstrate a mechanistic strategy in which musculature adapts to protect its nuclei from strain-induced damage; this consists of two distinct but interdependent structural components (Fig. 5, K and L). The first is a rigid and dense nuclear-associated network of MTs that is stabilized by Shot-EB1, which physically wraps and protects each of the myonuclei and is less sensitive to extra nuclear strain. Such a network exhibits a limited capacity to change during contraction. The second exhibits elastic properties provided by perinuclear MSP300 filaments that emanate from the nuclear envelope and associate with the MSP300 perinuclear ring. Together, both components form a shield capable of absorbing the contractile cytoplasmic strain, thereby preserving nuclear morphology during muscle contractions. The maintenance and assembly of these integral components are interconnected, possibly through the MT network. In humans, the loss of either Nesprin 1 or 2 has been linked to several muscular dystrophies, including Emery-Dreifuss muscular dystrophy (EDMD; Zhang et al., 2007; Stewart-Hutchinson et al., 2008; Puckelwartz et al., 2009), cardiomyopathies (Puckelwartz et al., 2010; Banerjee et al., 2014), arthrogryposis (Attali et al., 2009), and autosomal recessive cerebellar ataxia (ARCA1; Gros-Louis et al., 2007). We suggest that a primary defect in these diseases originates from defective nuclear morphology, which leads to abnormal regulation of a variety of nuclear-related functions in the muscle fibers.
Materials and methods
Fly stocks and husbandry
All crosses were performed at 25°C and rose on cornmeal yeast agar. For the RNAi-based screen, we used the Mef2-GAL4 driver as described previously (Schnorrer et al., 2010). Progenies were kept at 25°C and then transferred to 29°C to achieve a higher degree of knockdown. For verification of each target gene, at least two stocks carrying independent dsRNA sequences were used.
All stocks used in this manuscript are listed in Table S1. The Msp300Δ3′ strain (also known as Msp3003prime, FBal0218044) was a gift from S. Roth (Cologne Biocenter, Institute of Developmental Biology, Cologne, Germany; Technau and Roth, 2008). KlarΔ1-18 (FBal0277658) has been described previously (Elhanany-Tamir et al., 2012). Immunostaining against Shot indicated a significant reduction of protein level in shot-KD larval muscles (Fig. S1 C).
Dissection of larval body wall muscles
For immunofluorescent staining of Drosophila larval body wall muscles, mobile third instar larvae were picked with a fine brush and rinsed in PBS. Then, the larva was pinned at the anterior and posterior ends with insect pins (0.10 mm diameter; Fine Science Tools) on a Sylgard dish (Electron Microscopy Sciences) and completely immersed in PBS. Next, the body cavity was opened with iridectomy scissors along the dorsal midline. All internal organs were removed and the body wall was spread apart and fixed with additional pins. The images of muscles 6 and 7 were analyzed.
Larval stretching
During the natural crawling process, Drosophila third instar larvae are highly flexible. As a reference, the longitudinal muscles change their length between 30 µm at maximal contraction and 50 µm at maximal relaxation (Heckscher et al., 2012). To determine the mechanical stability of the described nuclear scaffolds under extreme tensile stresses, intact larvae were pulled under a stereomicroscope to a maximal 150% of their neutral length, as measured by a steel precision ruler beneath. Subsequently, the larvae were dissected and chemically fixed under the stretched condition for further analysis. Although they are artificial, these procedures usually did not introduce visible damage to WT larval tissue.
Immunofluorescence
For most of the staining, especially costaining for HP1 and phalloidin, a standard protocol with paraformaldehyde (4% from 16% stock of EM grade; #15710; Electron Microscopy Sciences) and without methanol was applied in order to avoid destruction of native F-actin or chromosomal morphology. To minimize damage to MTs, fixation time was limited to 20 min, followed by several cycles of washing with PBS with 0.1% Triton X-100 on a horizontal shaker with gentle agitation.
To preserve MTs for costaining, dissected larvae were fixed with cold methanol (−20°C) twice for 5 and 10 min, respectively, then rehydrated in PBS and rinsed several times with PBS. Subsequently, the larvae were fixed with 4% paraformaldehyde. Image analyses were consistently performed on muscles 6 and 7.
All specimens were mounted in Thermo Scientific Shandon Immu-Mount for microscopy (Thermo Fisher Scientific).
Antibodies
The following primary antibodies were obtained from the Developmental Studies Hybridoma Bank (DSHB): anti-Shot mAbRod1 (1:200 concentration; Lee et al., 2003), anti-laminC (LC28.26, 1:200 concentration; Riemer et al., 1995), and anti-HP1 (C1A9, 1:10 supernatant; James and Elgin, 1986). Other antibodies used included guinea pig anti-MSP300 (Volk, 1992), 1:200 guinea pig anti-Shot (Strumpf and Volk, 1998), 1:200 monoclonal anti-laminDm0 (provided by Y. Gruenbaum, Hebrew University of Jerusalem, Jerusalem, Israel; Gruenbaum et al., 1988), 1:10 rabbit anti-DmEB1 (provided by H. Ohkura, University of Edinburgh, Edinburgh, Scotland, UK; Elliott et al., 2005), 1:100 rabbit anti-Sls (Burkart et al., 2007), and rat anti–α-tubulin (MCA78S; 1:10; AbD Serotec). All the details of the primary antibodies are presented in Table S2.
Secondary antibodies conjugated with Cy3, Cy5, and Cy2 raised against guinea pig, mouse, and rabbit were purchased from Jackson ImmunoResearch Laboratories. Alexa Fluor 488 Phalloidin (Invitrogen) was used at 1:200. DAPI (1 µg/ml; Sigma-Aldrich) was used for labeling of nuclei.
Microscopy and image analysis
Microscopic images were acquired at 23°C with confocal microscopes (LSM 710/780; Carl Zeiss) using ZEN 2012 software and the following lenses: Plan-Apochromat 20×/0.8 NA M27, C-Apochromat 40×/1.20 NA W Korr M27, and Plan-Apochromat 63×/1.40 NA oil differential interference contrast (DIC) M27. Immersion medium Immersol W 2010 (ne = 1.3339) and Immersion oil Immersol 518 F (ne = 1.518) were used, respectively. For quantitative analysis, photon-counting mode was applied. Images were analyzed with Fiji v1.48 (Schindelin et al., 2012). 3D deconvolution was performed with Huygens Professional 14.06 Beta (Scientific Volume Imaging). Figure panels were finally assembled using CorelDRAW X4 (Corel Corporation).
Statistical analysis
Statistical analysis was performed using SPSS software (IBM SPSS for Windows, Version 19) and GraphPad Prism 6 demo (GraphPad Software, Inc.). Measurements were evaluated using variance analysis followed by a two-tailed Student’s t test with Welch correction for post hoc comparison. Differences among multiple groups were analyzed by one-way ANOVA with a post hoc Bonferroni test. P-values <0.05 were considered statistically significant. Data are presented as mean ± SEM.
BoxPlotR was used to generate the notched box plots (Spitzer et al., 2014), in which the center lines show the medians, box limits indicate the 25th and 75th percentiles as determined by R software, whiskers extend to the 5th and 95th percentiles, outliers (data points beyond the 95% confidence intervals) are represented by dots, crosses represent sample means, and bars indicate 95% confidence intervals of the means. The notches are defined as ±1.58 × IQR/sqrt(n) and represent the 95% confidence interval for each median. Non-overlapping notches give roughly 95% confidence that two medians differ significantly. All experiments were performed at least twice independently and in at least six randomly selected larvae.
Quantification of fluorescent intensity
For assessment of nuclear factor distribution, overall levels of nuclear fluorescence intensity were calculated by Fiji in a single confocal section. Control and mutant samples were labeled simultaneously under identical conditions and visualized using the same microscopic setting; photon counting mode was chosen when applicable. Fluorescent intensity as detected by DAPI staining was measured per nuclear area in order to bypass nuclear size variation in the mutant muscles.
For each allele, ∼300 myonuclei were measured from at least six individual animals for each condition. The same UAS-RNAi line without Mef2-Gal4 was used as a control.
Quantification of nuclear shape
To measure the shape of muscle nuclei, we stained larval body wall muscles with LaminDm0 antibody along with phalloidin and selected confocal stacks with high resolution and low background. Then the LaminDm0 channel was converted into an 8-bit grayscale stack and automatic quantifications were performed by using the plugin “3D Objects Counter” of Fiji.
All nonmuscle nuclei and myonuclei on any edge of the stacks, including incomplete nuclei, were discarded. Nuclear aggregates were not taken into account. The surface (An) and the volume (Vn) of each nucleus, namely the number of pixels forming the structures and its surface, respectively, were used to calculate the sphericity (Ψ) according to the mathematical definition Ψ = π^(1/3)(6Vn)^(2/3)/An. For calculation of the aspect ratio, bounding box dimensions of single myonucleus were used.
Online supplemental material
Fig. S1 shows several aspects relevant to the results (for more details, see the figure legend). Fig. S2 illustrates the annotated isoforms of MSP300 and Shot, as well as antibody epitopes mentioned in this work. Table S1 lists information about all fly stocks that were used. Table S2 includes information about all the primary antibodies used in this study.
Acknowledgments
We thank the Bloomington Stock Centre for various fly lines and the DSHB. We thank the Transgenic RNAi Project at Harvard Medical School (National Institutes of Health/NIGMS R01-GM084947) for providing transgenic RNAi fly stocks used in this study, and FlyBase for important data. We also thank N. Konstantin for English editing, and E. Zelzer, B. Shilo, L. Gilboa, and H. Hou for critical comments.
This study was supported by grants from the Joint Lower Saxony–Israeli Research Project funding program by the Lower Saxony Ministry of Science and Culture (to T. Volk; 11-76251-99-15/12 [ZN2832]), and from MINERVA foundation, grant no. 711743 (to T. Volk).
The authors declare no competing financial interests.