MarvelD3 is a transmembrane component of tight junctions, but there is little evidence for a direct involvement in the junctional permeability barrier. Tight junctions also regulate signaling mechanisms that guide cell proliferation; however, the transmembrane components that link the junction to such signaling pathways are not well understood. In this paper, we show that MarvelD3 is a dynamic junctional regulator of the MEKK1–c-Jun NH2-terminal kinase (JNK) pathway. Loss of MarvelD3 expression in differentiating Caco-2 cells resulted in increased cell migration and proliferation, whereas reexpression in a metastatic tumor cell line inhibited migration, proliferation, and in vivo tumor formation. Expression levels of MarvelD3 inversely correlated with JNK activity, as MarvelD3 recruited MEKK1 to junctions, leading to down-regulation of JNK phosphorylation and inhibition of JNK-regulated transcriptional mechanisms. Interplay between MarvelD3 internalization and JNK activation tuned activation of MEKK1 during osmotic stress, leading to junction dissociation and cell death in MarvelD3-depleted cells. MarvelD3 thus couples tight junctions to the MEKK1–JNK pathway to regulate cell behavior and survival.
Introduction
Epithelial cells are joined to each other by junctional complexes that mediate cell–cell adhesion but also regulate cell proliferation and differentiation. Tight junctions, the most apical junctions, form the apical junctional complex together with adherens junctions. They form paracellular diffusion barriers required for functional epithelial tissues (Steed et al., 2010; Shen et al., 2011). Tight junctions are composed of transmembrane components and a complex submembrane plaque of proteins that link the junction to the cytoskeleton (Furuse and Tsukita, 2006; Van Itallie and Anderson, 2006; Balda and Matter, 2008). Tight junctions and components of the submembrane plaque have been linked to the regulation of signal transduction mechanisms that guide epithelial cell proliferation and differentiation (Balda and Matter, 2009). However, it is still poorly understood how junctional membrane proteins regulate these mechanisms and how they cross talk with the major signaling networks that guide cell behavior. Deregulation of expression of junctional transmembrane proteins has been reported for cancers, indicating that they may be important for tumorigenesis; however, it is not known whether up- or down-regulation is a consequence or cause of disease (Martin et al., 2011).
The three transmembrane proteins Occludin, Tricellulin, and MarvelD3 form the family of tight junction–associated Marvel domain proteins (Steed et al., 2010). Of the three, only Tricellulin seems to be directly required for the formation of functional paracellular diffusion barriers (Saitou et al., 2000; Ikenouchi et al., 2005; Krug et al., 2009; Steed et al., 2009; Raleigh et al., 2010). Hence, these proteins may be less important for barrier formation but may regulate junctional signaling mechanisms. Indeed, Occludin manipulation affects the permeability properties of tight junctions in different tissues and experimental systems, which is compatible with Occludin functioning as a regulatory protein (Balda et al., 1996; McCarthy et al., 1996; Chen et al., 1997; Hirase et al., 1997; Wong and Gumbiner, 1997; Antonetti et al., 1998, 1999; Matter and Balda, 1998). MarvelD3 is less well understood but may also have a modulatory role (Steed et al., 2009; Kojima et al., 2011). Expression of all three junctional Marvel domain proteins can be deregulated in different cancers or cancer cell lines; however, the pathological significance of these observations is not clear (Martin et al., 2010; Kojima et al., 2011; Korompay et al., 2012). Nevertheless, Occludin has been shown to cross talk with oncogenic Raf-1 signaling, as its expression is repressed by the kinase, and it can suppress junction dissolution induced by Raf-1 signaling if reexpressed ectopically (Li and Mrsny, 2000). The mechanism by which Occludin suppresses the effect of Raf-1 on cell–cell junctions is not clear.
Here, we demonstrate that MarvelD3 functions as a regulator of epithelial cell proliferation, migration, and survival. Our data show that MarvelD3 recruits MEKK1 to tight junctions to suppress the MEKK1–JNK pathway, leading to the suppression of JNK-regulated transcriptional mechanisms, inhibition of Cyclin D1 expression, and reduced cell proliferation and migration. We further show that interplay between dynamic MarvelD3 behavior and JNK signaling is important for the cellular response to osmotic stress.
Results
MarvelD3 regulates cell proliferation and migration
We first used a loss of function approach to ask whether MarvelD3 regulates epithelial cell migration and proliferation. As a model system, we used Caco-2 cells, a human intestinal cell line that spontaneously differentiates, and depleted MarvelD3 expression using specific siRNAs.
MarvelD3-targeting siRNAs efficiently depleted expression of the protein as described (Fig. 1 A; Steed et al., 2009). Wound-healing assays were then performed with confluent monolayers. Bright-field microscopy and subsequent quantifications revealed an increased rate of gap closure in monolayers depleted of MarvelD3, covering almost twice the space as controls in 26 h (Fig. 1, B and C). MarvelD3-depleted monolayers retained intact junctions, indicating that MarvelD3-depleted cells still migrated as cell sheets (Fig. 1 D). As the migration assays were performed in the presence of mitomycin C, enhanced wound closer was not caused by increased proliferation but faster migration. Nevertheless, proliferation assays indicated that MarvelD3 depletion also resulted in elevated cell numbers (Fig. 1 E). The functional effects observed upon depletion of MavelD3 could be complemented by expressing mouse MarvelD3, which is resistant to the siRNAs used, indicating that the observed phenotype was specific (Fig. 1, F–H). These data indicate that depletion of MarvelD3 stimulates cell migration and proliferation of Caco-2 cells.
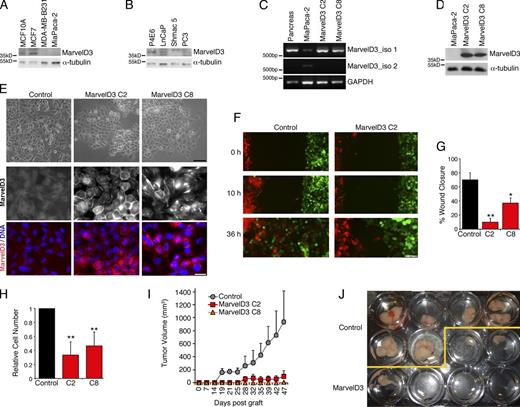
MarvelD3 regulates cell migration and proliferation in Caco-2 cells. (A–E) Caco-2 cells were transfected with siRNAs, and total cell extracts were analyzed by immunoblotting (A), or cells were assayed for migration or proliferation (B–E). B shows phase-contrast images, and D shows a maximal intensity projection of migrating cells labeled for tight and adherens junction proteins. The yellow lines label the wound edges at time 0. (C) Migration in to the cell-free space was measured after 26 h and normalized to controls. Shown are means ± 1 SD; n = 3. (E) Proliferation was analyzed by measuring the increase in cell numbers and normalizing the numbers to control cultures. Shown are means ± 1 SD; n = 3. (F–H) A complementation assay to test the specificity of MarvelD3 siRNAs was performed by repeating the assays as described in B–E using a Caco-2 cell line expressing mouse MarvelD3 (M-MD3) with a C-terminal VSV tag. G and H show means ± 1 SD; n = 6. *, P < 0.05; **, P < 0.01. Bars: (B) 100 µm; (D) 20 µm.
MarvelD3 regulates cell migration and proliferation in Caco-2 cells. (A–E) Caco-2 cells were transfected with siRNAs, and total cell extracts were analyzed by immunoblotting (A), or cells were assayed for migration or proliferation (B–E). B shows phase-contrast images, and D shows a maximal intensity projection of migrating cells labeled for tight and adherens junction proteins. The yellow lines label the wound edges at time 0. (C) Migration in to the cell-free space was measured after 26 h and normalized to controls. Shown are means ± 1 SD; n = 3. (E) Proliferation was analyzed by measuring the increase in cell numbers and normalizing the numbers to control cultures. Shown are means ± 1 SD; n = 3. (F–H) A complementation assay to test the specificity of MarvelD3 siRNAs was performed by repeating the assays as described in B–E using a Caco-2 cell line expressing mouse MarvelD3 (M-MD3) with a C-terminal VSV tag. G and H show means ± 1 SD; n = 6. *, P < 0.05; **, P < 0.01. Bars: (B) 100 µm; (D) 20 µm.
MarvelD3 has recently been shown to be down-regulated during snail-induced epithelial to mesenchymal transition in pancreatic epithelial cells (Kojima et al., 2011). Indeed, we found MarvelD3 expression to be strongly reduced in cell lines with invasive phenotypes derived from breast, pancreatic, and prostate tumors (i.e., MDA-MB-231, PC3, and MiaPaca-2; Fig. 2, A and B).
MarvelD3 regulates cell migration and proliferation of MiaPaca-2 cells. (A and B) Immunoblotting of extracts of cell lines derived from breast and pancreas (A) and prostate (B). (C) RT-PCR analysis of MarvelD3 isoform expression in healthy adult pancreas, and control and MarvelD3 isoform 1–transfected MiaPaca-2 cells. (D) Immunoblotting of control and MarvelD3 isoform 1–transfected MiaPaca-2 cells. (E) Morphology of control and MarvelD3-expressing MiaPaca-2 cells (two clones, C2 and C8, were analyzed in all experiments) was assessed by phase-contrast and immunofluorescence microscopy (see Fig. S2 for analysis of other junctional markers). (F and G) Migration assays were performed with GFP- and RFP-expressing pools of control and MarvelD3-expressing MiaPaca-2 cells over 2 d. The quantification was performed, and images were taken after 24 h and show means ± 1 SD; n = 3. Note, only control cells freely mix after they have filled the gap or when labeled cells are co-cultured (see Fig. S2 C for co-cultures). (H) Proliferation of control and MarvelD3-expressing cells was analyzed by measuring cell numbers after 3 d (shown are means ± 1 SD; n = 3). (I and J) Nude mice were injected with control or MarvelD3-expressing MiaPaca-2 cells, and tumor formation was assessed by measuring the size over 47 d. Shown are means ± 1 SD of five animals per cell line. *, P < 0.05; **, P < 0.01. Bars: (E, top) 50 µm; (E, bottom) 20 µm; (F) 100 µm.
MarvelD3 regulates cell migration and proliferation of MiaPaca-2 cells. (A and B) Immunoblotting of extracts of cell lines derived from breast and pancreas (A) and prostate (B). (C) RT-PCR analysis of MarvelD3 isoform expression in healthy adult pancreas, and control and MarvelD3 isoform 1–transfected MiaPaca-2 cells. (D) Immunoblotting of control and MarvelD3 isoform 1–transfected MiaPaca-2 cells. (E) Morphology of control and MarvelD3-expressing MiaPaca-2 cells (two clones, C2 and C8, were analyzed in all experiments) was assessed by phase-contrast and immunofluorescence microscopy (see Fig. S2 for analysis of other junctional markers). (F and G) Migration assays were performed with GFP- and RFP-expressing pools of control and MarvelD3-expressing MiaPaca-2 cells over 2 d. The quantification was performed, and images were taken after 24 h and show means ± 1 SD; n = 3. Note, only control cells freely mix after they have filled the gap or when labeled cells are co-cultured (see Fig. S2 C for co-cultures). (H) Proliferation of control and MarvelD3-expressing cells was analyzed by measuring cell numbers after 3 d (shown are means ± 1 SD; n = 3). (I and J) Nude mice were injected with control or MarvelD3-expressing MiaPaca-2 cells, and tumor formation was assessed by measuring the size over 47 d. Shown are means ± 1 SD of five animals per cell line. *, P < 0.05; **, P < 0.01. Bars: (E, top) 50 µm; (E, bottom) 20 µm; (F) 100 µm.
As an additional control for the specificity of the siRNAs used, MiaPaca-2 cells, which do not express detectable MarvelD3 protein, were transfected with the siRNAs, and the migration and proliferation experiments were repeated. Fig. S1 (A and B) shows that the siRNAs did indeed not affect the behavior of MiaPaca-2 cells. In contrast, in MCF7 cells, which express MarvelD3, depletion also stimulated migration and proliferation (Fig. S1, C–E).
To address the possibility that MarvelD3 can regulate cell behavior of invasive tumor cells, we expressed MarvelD3 in the pancreatic tumor cell line MiaPaca-2. Though there are two isoforms of MarvelD3, only isoform 1 could be detected in human pancreas by RT-PCR (Fig. 2 C; Steed et al., 2009). We therefore transfected MiaPaca-2 cells with a cDNA encoding isoform 1 and isolated two clones expressing comparable levels of MarvelD3 isoform 1 mRNA as normal pancreas (Fig. 2 C). Both clones expressed similar amounts of MarvelD3 protein (Fig. 2 D). Phase-contrast microscopy revealed that MarvelD3-expressing cells were less spindle shaped than control MiaPaca-2 cells and that the transfected protein was enriched at cell–cell contacts (Fig. 2 E). However, MarvelD3 reexpression did not induce a reversal of the epithelial to mesenchymal transition. Though ZO-1 was occasionally seen at cell–cell contacts, crucial adhesion proteins, such as E-cadherin and Occludin, were not recruited to these contacts sites, and their expression remained the same or was reduced (Fig. S2, A and B).
We next performed migration assays to determine whether reexpression indeed led to a corresponding gain-of-function phenotype. For each cell line, RFP- and GFP-expressing cells were used to follow intermixing of cells once the wound was closed. Fig. 2 (F and G) shows that control cells closed the cell-free space more quickly than MarvelD3-expressing clones. When RFP- and GFP-expressing MiaPaca-2 cells were co-cultured, MarvelD3-expressing cells formed distinct islands of green and red cells, whereas control cells formed randomly mixed monolayers, indicating that cell mobility within monolayers was also reduced by expression of MarvelD3 (Fig. S2 C). Similarly, reexpression of MarvelD3 repressed proliferation of MiaPaca-2 cells (Fig. 2 H).
Finally, we xenografted MiaPaca-2 cell lines into nude mice to determine the propensity of MarvelD3 to suppress proliferation in vivo. Control cells formed large tumors, and the tumor volume regularly increased until the end of the experiment (Fig. 2, I and J). In contrast, clones expressing MarvelD3 only formed very small tumors. MarvelD3 can thus suppress proliferation of tumor cells in vitro and in vivo. These data indicate that MarvelD3 regulates cell proliferation and cell migration of differentiating and dedifferentiated epithelial model cell lines, indicating that it functions as a signaling transmembrane component of tight junctions.
MarvelD3 suppresses JNK signaling
The MAPK signaling pathways are fundamental for the regulation of cell proliferation, migration, and survival. MAPKs are stimulated by kinase cascades that result in their phosphorylation. We therefore asked whether MarvelD3 expression affects phosphorylation of MAPKs using the MiaPaca-2–derived MarvelD3-expressing cell lines. MarvelD3-expressing cells exhibited strongly reduced levels of phosphorylated JNK in comparison to control cells, whereas the levels of phosphorylated p38 and Erk1/2 were not significantly affected, indicating that MarvelD3 specifically suppresses the JNK MAPK pathway (Fig. 3, A and B). Depletion of MarvelD3 in Caco-2 and MCF7 cells led to increased phosphorylation of JNK (Figs. 3, C and D; and S2, D and E), providing further support for a role of MarvelD3 in the regulation of JNK activation.
MarvelD3 regulates JNK signaling and cyclin D1 expression. (A and B) Control and MarvelD3-expressing MiaPaca-2 cells were analyzed by immunoblotting of whole-cell lysates to monitor phosphorylation of MAPKs. B shows a quantification of the ratios between phosphorylated and nonphosphorylated forms. (C and D) Caco-2 cells transfected with control and MarvelD3-targeting siRNAs were analyzed for MAPK activation as described in A and B. (E and F) Analysis of AP1 promoter activity using a dual-luciferase reporter assay in Caco-2 (E) and MiaPaca-2 (F) cells. Control cells were cotransfected with an empty expression vector, and the other cells were cotransfected as indicated with cDNAs encoding MarvelD3 isoforms or Occludin. (G) Immunoblot of Caco-2 cells transfected as in E using antibodies against MarvelD3 and Occludin. (H) Activities of the indicated promoters were analyzed as in E and F. (I–K) Analysis of cyclin D1 expression was analyzed by reporter gene assay in control and MarvelD3-targeting siRNA-transfected Caco-2 cells or by immunoblotting in MiaPaca-2 and Caco-2 cells after transfection or depletion of MarvelD3 (J and K). All graphs show means ± 1 SD; n = 3. *, P < 0.05; **, P < 0.01; ***, P < 0.001.
MarvelD3 regulates JNK signaling and cyclin D1 expression. (A and B) Control and MarvelD3-expressing MiaPaca-2 cells were analyzed by immunoblotting of whole-cell lysates to monitor phosphorylation of MAPKs. B shows a quantification of the ratios between phosphorylated and nonphosphorylated forms. (C and D) Caco-2 cells transfected with control and MarvelD3-targeting siRNAs were analyzed for MAPK activation as described in A and B. (E and F) Analysis of AP1 promoter activity using a dual-luciferase reporter assay in Caco-2 (E) and MiaPaca-2 (F) cells. Control cells were cotransfected with an empty expression vector, and the other cells were cotransfected as indicated with cDNAs encoding MarvelD3 isoforms or Occludin. (G) Immunoblot of Caco-2 cells transfected as in E using antibodies against MarvelD3 and Occludin. (H) Activities of the indicated promoters were analyzed as in E and F. (I–K) Analysis of cyclin D1 expression was analyzed by reporter gene assay in control and MarvelD3-targeting siRNA-transfected Caco-2 cells or by immunoblotting in MiaPaca-2 and Caco-2 cells after transfection or depletion of MarvelD3 (J and K). All graphs show means ± 1 SD; n = 3. *, P < 0.05; **, P < 0.01; ***, P < 0.001.
JNK activates the AP1 transcription factor complex by phosphorylation of c-Jun, one of its subunits (Davis, 2000). Therefore, we measured AP1 activity using a luciferase reporter gene assay with an AP1-responsive promoter. Transfection of either MarvelD3 isoform repressed luciferase expression in Caco-2 and MiaPaca-2 cells, indicating that MarvelD3 can suppress AP1 activity in cells with and without tight junctions (Fig. 3, E–G). In contrast, transfection of Occludin did not affect AP1 activity, suggesting that inhibition is not a general effect of Marvel domain–containing proteins. As expected, MarvelD3 expression also suppressed the activity of a serum response element (SRE)–driven promoter, in agreement with JNK’s regulatory role in Elk1 activation (Fig. 3 H; Cavigelli et al., 1995; Davis, 2000). ZO-1–associated nucleic acid binding protein (ZONAB) and T cell factor/β-catenin represent transcriptional pathways that are regulated by cell junctions, but their activities were not affected (Fig. 3 H).
To analyze the role of MarvelD3 in the regulation of physiologically relevant promoters, we used a cyclin D1 promoter construct (Watanabe et al., 1996; Sourisseau et al., 2006). The cyclin D1 promoter is regulated by multiple JNK-regulated transcription factors (Klein and Assoian, 2008). Indeed, activity of the cyclin D1 promoter was reduced upon MarvelD3 expression and was stimulated upon its depletion (Fig. 3, H and I). Analysis of protein expression in transfected MiaPaca-2 and depleted Caco-2 cells confirmed the effect of MarvelD3 on cyclin D1 expression. MarvelD3 signaling thus regulates the expression of this central cell cycle regulator, providing a mechanistic explanation for its proliferation suppressive activity (Fig. 3, J and K).
We next asked whether JNK activity is indeed important for MarvelD3-regulated cell migration and proliferation. Inhibition of JNK with the specific inhibitor SP600125 suppressed migration and proliferation of MiaPaca-2 cells, indicating that JNK signaling is indeed important for migration and proliferation of this tumor cell line (Fig. 4, A–C). Similarly, migration of MarvelD3-depleted Caco-2 cells was blocked by the JNK inhibitor (Fig. 4 D). The increase in cell numbers induced by MarvelD3 depletion was also strongly reduced by SP600125 (Fig. 4 E). If JNK expression was suppressed with siRNAs targeting JNK1/2, stimulation of proliferation and migration by MarvelD3 suppression was also inhibited, indicating that MarvelD3 is indeed acting on JNK (Fig. 4, F–H). Inhibition of JNK also inhibited increased expression of cyclin D1, indicating that regulation of expression of the cell cycle gene by MarvelD3 involves the JNK pathway (Fig. 4, I and J).
JNK signaling and regulation of proliferation and migration. (A) MiaPaca-2 cells were incubated with 20 and 50 µM SP600125, and JNK inhibition was then analyzed by blotting for phosphorylated c-Jun. All subsequent experiments were performed with 20 µm SP600125. (B–E) Migration and proliferation of MiaPaca-2 (B and C) and siRNA-transfected Caco-2 cells (D and E) were analyzed as in Fig. 1. No increase in cell death was observed when similarly treated cells were analyzed by DNA staining (Fig. 9). (F–H) Caco-2 cells were transfected with control and MarvelD3- and JNK1/2-targeting siRNAs as indicated. After 3 d, the cells were analyzed by immunoblotting (F), for cell migration (G), or cell proliferation (H). (I and J) Control and MarvelD3 siRNA–transfected Caco-2 cells were incubated with DMSO or SP600125 for 10 h before lysis and analysis by immunoblotting. J shows a quantification of cyclin D1 expression. All graphs show means ± 1 SD; n = 3 (B and C), n = 5 (G), n = 6 (H), and n = 4 (K). *, P < 0.05; **, P < 0.01; ***, P < 0.001.
JNK signaling and regulation of proliferation and migration. (A) MiaPaca-2 cells were incubated with 20 and 50 µM SP600125, and JNK inhibition was then analyzed by blotting for phosphorylated c-Jun. All subsequent experiments were performed with 20 µm SP600125. (B–E) Migration and proliferation of MiaPaca-2 (B and C) and siRNA-transfected Caco-2 cells (D and E) were analyzed as in Fig. 1. No increase in cell death was observed when similarly treated cells were analyzed by DNA staining (Fig. 9). (F–H) Caco-2 cells were transfected with control and MarvelD3- and JNK1/2-targeting siRNAs as indicated. After 3 d, the cells were analyzed by immunoblotting (F), for cell migration (G), or cell proliferation (H). (I and J) Control and MarvelD3 siRNA–transfected Caco-2 cells were incubated with DMSO or SP600125 for 10 h before lysis and analysis by immunoblotting. J shows a quantification of cyclin D1 expression. All graphs show means ± 1 SD; n = 3 (B and C), n = 5 (G), n = 6 (H), and n = 4 (K). *, P < 0.05; **, P < 0.01; ***, P < 0.001.
The N-terminal cytoplasmic domain regulates signaling
MarvelD3 is expressed as two isoforms with a common N-terminal cytoplasmic domain (Steed et al., 2009). As the two isoforms had comparable effects on the activity of transcriptional reporter constructs, we speculated that this N-terminal domain is responsible for the signaling activity. In support of this hypothesis, repetition of the AP1 reporter assay with constructs containing only the N-terminal domain (MarvelD3_NTD) or lacking it (MarvelD3_1ΔNTD) demonstrated that the N-terminal cytoplasmic domain was both necessary and sufficient to regulate AP1 activity (Fig. 5 A).
MarvelD3 regulates MEKK1 localization. (A) AP1 reporter gene assays were performed with MarvelD3 constructs consisting of the N-terminal domain only (MarvelD3_NTD) or lacking this domain (MarvelD3_1ΔNTD). Shown are means ± 1 SD; n = 3. (B) GST pull-down assays were performed using Caco-2 cell extracts and fusion proteins containing either the N-terminal domain (GST-MarvelD3_NTD) or either one of the two isoform C-terminal domains (GST-MarvelD3_CTD1 and GST-MarvelD3_CTD2). The precipitates were then analyzed by immunoblotting as indicated. (C) Schematic of MEKK1. Indicated are the RING finger and the kinase domains as well as the regions contained by the fusion proteins used for the recombinant protein-binding assay in D. IR, intervening region. (D). His6-tagged MEKK1 fusion proteins were incubated with GST-MarvelD3_NTD fusion protein bound to glutathione beads. After washing, bound fusion proteins were eluted analyzed by immunoblotting using antibodies against His6 and GST. (E) Control and MarvelD3-expressing MiaPaca-2 cells were stained for the VSV-tagged transfected protein and MEKK1. Arrowheads point to cell–cell contacts positive for MEKK1. (F and G) Immunoblotting of Caco-2 cells transfected with siRNAs as indicated. (H and I) Caco-2 cells transfected with the indicated siRNAs were stained with mouse anti-MEKK1 and rabbit anti-MarvelD3 antibodies. Arrowheads point to junctional fragments positive for MEKK1. Graphs in I show ratios of mean densities of junctional segments and cytoplasm (shown are means ± 1 SD; n = 10). Bars: (E) 20 µm; (H) 10 µm. ***, P < 0.001.
MarvelD3 regulates MEKK1 localization. (A) AP1 reporter gene assays were performed with MarvelD3 constructs consisting of the N-terminal domain only (MarvelD3_NTD) or lacking this domain (MarvelD3_1ΔNTD). Shown are means ± 1 SD; n = 3. (B) GST pull-down assays were performed using Caco-2 cell extracts and fusion proteins containing either the N-terminal domain (GST-MarvelD3_NTD) or either one of the two isoform C-terminal domains (GST-MarvelD3_CTD1 and GST-MarvelD3_CTD2). The precipitates were then analyzed by immunoblotting as indicated. (C) Schematic of MEKK1. Indicated are the RING finger and the kinase domains as well as the regions contained by the fusion proteins used for the recombinant protein-binding assay in D. IR, intervening region. (D). His6-tagged MEKK1 fusion proteins were incubated with GST-MarvelD3_NTD fusion protein bound to glutathione beads. After washing, bound fusion proteins were eluted analyzed by immunoblotting using antibodies against His6 and GST. (E) Control and MarvelD3-expressing MiaPaca-2 cells were stained for the VSV-tagged transfected protein and MEKK1. Arrowheads point to cell–cell contacts positive for MEKK1. (F and G) Immunoblotting of Caco-2 cells transfected with siRNAs as indicated. (H and I) Caco-2 cells transfected with the indicated siRNAs were stained with mouse anti-MEKK1 and rabbit anti-MarvelD3 antibodies. Arrowheads point to junctional fragments positive for MEKK1. Graphs in I show ratios of mean densities of junctional segments and cytoplasm (shown are means ± 1 SD; n = 10). Bars: (E) 20 µm; (H) 10 µm. ***, P < 0.001.
We next tested whether the N-terminal cytoplasmic domain interacts with upstream components that regulate JNK activation. We first performed GST pull-down assays from whole Caco-2 cell lysates using fusion proteins containing the cytosolic domains of MarvelD3 (NTD [N-terminal domain], CTD1 [C terminus of isoform 1], and CTD2 [C terminus of isoform 2]). Subsequent blotting with antibodies against MAPKKKs revealed that MEKK1 was pulled down specifically by the fusion protein containing the N-terminal domain. Other MAPKKKs, such as MEKK3 and MLK1, which can also signal to JNK, were not detected. Prominent junctional scaffolding proteins, such as ZO-1, -2, and -3, were also not detected in the pull-downs (Fig. 5 B). Reconstitution experiments with recombinant proteins indicated that the interaction between the N-terminal domain of MarvelD3 and MEKK1 can occur directly and involves the kinase domain (Fig. 5, C and D).
To confirm the relevance of this in vitro interaction, we tested whether MarvelD3 affects the subcellular distribution of MEKK1 using control and MarvelD3-expressing MiaPaca-2 cells. Fig. 5 E shows that MEKK1 was indeed partially recruited to cell–cell contacts when MarvelD3 was expressed in MiaPaca-2 cells but not in control cells, indicating that reexpression of the junctional membrane protein is sufficient to stimulate membrane recruitment of MEKK1. Similarly, MEKK1 antibodies stained cell junctions in control Caco-2 cells but not when MarvelD3 had been depleted (Fig. 5, F–I). In contrast, depletion of MEKK1 did not affect the distribution of the junctional membrane protein. MarvelD3 thus regulates the localization of MEKK1.
Our data indicate that MEKK1 can form complexes in vitro with MarvelD3 and that its subcellular distribution is regulated by the junctional membrane protein. However, we were unable to reliably coimmunoprecipitate MarvelD3 and MEKK1. Therefore, we made use of a recently published strategy to identify proteins in close proximity in live cells, which relies on a promiscuous biotin ligase that is fused to the protein of interest to transfer biotin to nearby lysine residues (Roux et al., 2012). We generated two MarvelD3 fusion proteins that carried the biotin ligase either at the N (Myc-BirA-MarvelD3) or the C terminus (MarvelD3-BirA-HA) and stably expressed them in MDCK cells. Both fusion proteins were efficiently expressed and transported to cell–cell contacts (Fig. 6, A–C). When biotin had been added to the cultures before fixation, a strong streptavidin staining was obtained in cells expressing the chimeric proteins but not in neighboring cells that lacked expression (Fig. 6, A and B). MarvelD3/biotin ligase chimeras are thus targeted to cell junctions and can mediate biotinylation of cellular proteins.
MarvelD3 and MEKK1 form complexes. (A and B) MDCK cell lines expressing MarvelD3 fusion proteins carrying the biotin ligase at the C terminus (A) or the N terminus (B) were incubated with biotin for 24 h as indicated before fixation and staining with fluorescently labeled streptavidin and anti-HA (A) or anti-Myc antibodies. The white lines demark the border between cells that do and do not express the fusion protein. (C) Immunoblotting of extracts of cells expressing the MarvelD3 biotin ligase fusion proteins. (D) Cell extracts of the cell lines shown in A and B were incubated with streptavidin beads, and the resulting precipitates were blotted with the indicated antibodies. (E) Caco-2 cells transiently transfected as indicated were analyzed as described in D. Bars, 10 µm.
MarvelD3 and MEKK1 form complexes. (A and B) MDCK cell lines expressing MarvelD3 fusion proteins carrying the biotin ligase at the C terminus (A) or the N terminus (B) were incubated with biotin for 24 h as indicated before fixation and staining with fluorescently labeled streptavidin and anti-HA (A) or anti-Myc antibodies. The white lines demark the border between cells that do and do not express the fusion protein. (C) Immunoblotting of extracts of cells expressing the MarvelD3 biotin ligase fusion proteins. (D) Cell extracts of the cell lines shown in A and B were incubated with streptavidin beads, and the resulting precipitates were blotted with the indicated antibodies. (E) Caco-2 cells transiently transfected as indicated were analyzed as described in D. Bars, 10 µm.
We then extracted whole-cell lysate and isolated biotinylated proteins with streptavidin immobilized on beads. Immunoblotting streptavidin precipitates with anti-MEKK1 antibodies demonstrated that the kinase was biotinylated in both types of cell lines (Fig. 6 D). Despite the lower levels, the fusion protein carrying the biotin ligase on the N terminus biotinylated MEKK1 more efficiently. Moreover, in transiently transfected Caco-2 cells, biotinylation of MEKK1 was only observed with the N-terminal fusion protein, possibly because of the low transfection efficiency of such cells (Fig. 6 E). These data thus support a model in which complex formation between MarvelD3 and MEKK1 involves the N-terminal cytoplasmic domain of the tight junction protein.
Our data thus indicate that MarvelD3 regulates the subcellular distribution of MEKK1. As MarvelD3 also suppresses JNK activation and its N terminus is sufficient and required to suppress AP1 activity, regulation of a MEKK1–JNK pathway seems a major mechanism by which MarvelD3 regulates epithelial cell behavior.
MarvelD3 is required for epithelial integrity during osmotic stress
To further address the significance of this mechanism, we investigated the osmotic shock response, in which JNK plays a crucial role (Seki et al., 2012). Strikingly, the subcellular distribution of MarvelD3 drastically changed in response to osmotic stress. Fig. 7 A shows that MarvelD3 was quickly internalized and, after an initial distribution in scattered vesicular structures, started to accumulate in perinuclear endosome-like structures before reappearing at the cell surface and cell–cell junctions. There was no apparent reduction in MarvelD3 expression during this time course even when protein synthesis was blocked with cycloheximide, suggesting that internalized protein did not get degraded but recycled (Figs. 7 B and S3 A). The perinuclear pool of MarvelD3 partially overlapped with transferrin receptor, suggesting that trafficking involved an endocytic recycling route (Fig. S3, B–D). In contrast, Occludin was more stably associated with the junctions and no endocytic burst was observed, but the staining became warped, irregular, and occasionally discontinuous (Fig. 7 A). Only after long osmotic shock times (>2 h), when MarvelD3 already started to reappear at the cell surface, a modest increase in perinuclear vesicular Occludin staining was evident (Figs. 7 A and S3 E). Thus, the junctional association of MarvelD3 is regulated by osmotic stress.
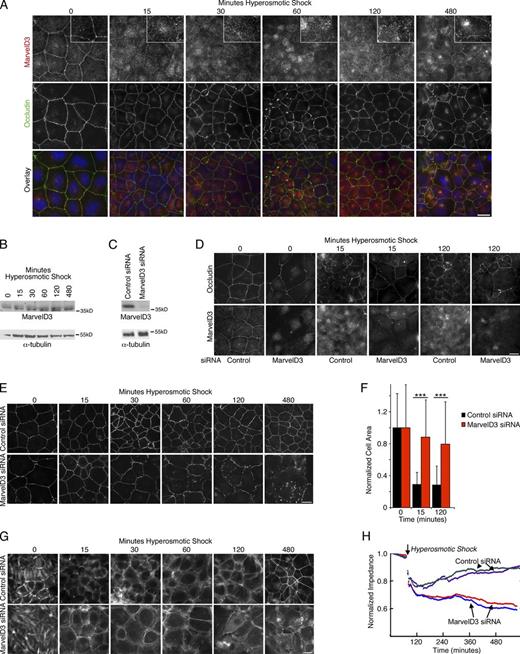
MarvelD3 is required for epithelial monolayer integrity during osmotic stress. (A and B) Caco-2 cells were osmotically stressed for the times indicated before analysis by immunofluorescence (A) or immunoblotting (B). The insets in A show twofold magnifications of the MarvelD3 staining. (C–H) Cells were transfected with control and MarvelD3 targeting siRNAs and were then osmotically stressed. The cells were then analyzed by immunoblotting (C), immunofluorescence (D–G), or by measuring electrical impedance (H). Cells were stained for Occludin and MarvelD3 (D), ZO-1 (E), or F-actin (G). For a quantification of Occludin and MarvelD3 localization and comparison with endocytic markers, see Fig. S3. F shows a quantification of the area covered by individual cells to assess cell contraction. Shown are means ± 1 SD of three experiments counting 20–40 cells per condition and experiment; ***, P < 0.001. Bars: (A, D, and E) 10 µm; (G) 20 µm.
MarvelD3 is required for epithelial monolayer integrity during osmotic stress. (A and B) Caco-2 cells were osmotically stressed for the times indicated before analysis by immunofluorescence (A) or immunoblotting (B). The insets in A show twofold magnifications of the MarvelD3 staining. (C–H) Cells were transfected with control and MarvelD3 targeting siRNAs and were then osmotically stressed. The cells were then analyzed by immunoblotting (C), immunofluorescence (D–G), or by measuring electrical impedance (H). Cells were stained for Occludin and MarvelD3 (D), ZO-1 (E), or F-actin (G). For a quantification of Occludin and MarvelD3 localization and comparison with endocytic markers, see Fig. S3. F shows a quantification of the area covered by individual cells to assess cell contraction. Shown are means ± 1 SD of three experiments counting 20–40 cells per condition and experiment; ***, P < 0.001. Bars: (A, D, and E) 10 µm; (G) 20 µm.
We next asked whether MarvelD3 is important for the cellular response to osmotic stress by making use of siRNAs to deplete its expression (Fig. 7 C). Repetition of the osmotic stress treatments followed by immunofluorescence for Occludin and ZO-1 revealed that MarvelD3 was indeed important for maintenance of the molecular junctional structure during the osmotic stress response, as the distribution of ZO-1 and, to a lesser extent, Occludin started to become discontinuous within 2 h in the absence of MarvelD3 (Figs. 7, D and E; and S3 F). Moreover, the surface area rapidly reduced in control cells in response to osmotic shock but not when MarvelD3 had been depleted (Fig. 7 F). These morphological changes were even more evident when cells were stained for F-actin, as the depleted cells became more spread and lost their normally regular cell shape (Fig. 7 G). Control cells reassembled a reinforced junctional F-actin ring after 480 min of osmotic treatment, but MarvelD3-depleted cells failed to do so. Instead, the F-actin network became increasingly disorganized. MarvelD3 is thus important for the maintenance of the junctional structure during osmotic stress.
MarvelD3 depletion does not affect epithelial barrier integrity in steady-state Caco-2 monolayers (Steed et al., 2009; Raleigh et al., 2010). Hence, we determined whether its depletion affected epithelial barrier integrity under stress. Transepithelial electrical impedance measurements showed that both control and MarvelD3-depleted cells responded with a rapid drop in electrical impedance when the osmotic shock was applied (Fig. 7 H). Control cells then started to recover, and impedance slowly increased again. However, impedance of MarvelD3-depleted monolayers continued to decrease and did not recover, indicating that the epithelial barrier was defective. MarvelD3 is thus required for epithelial barrier integrity during osmotic stress.
MarvelD3 regulates cell survival during osmotic stress
Because JNK signaling is an important component of the cellular stress response (Xia et al., 2007; Seki et al., 2012), we next asked whether MarvelD3 regulates JNK signaling during osmotic stress. To do so, we analyzed the activation profile of JNK in response to osmotic shock.
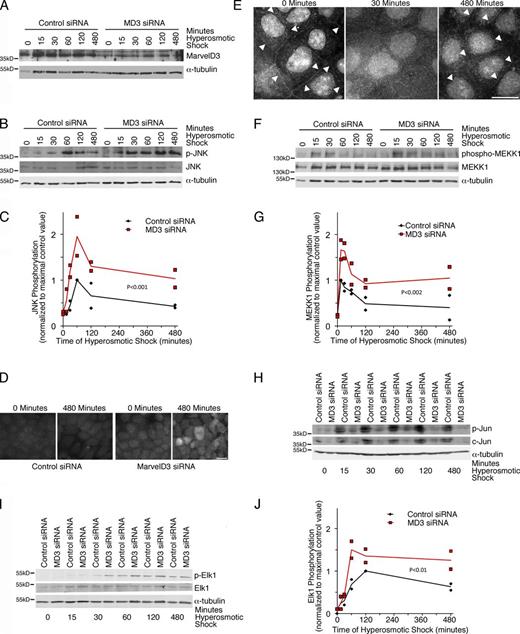
Levels of phosphorylated JNK increased within the first hour of hyperosmotic shock independent of whether control or MarvelD3-depleted cells were analyzed; however, phosphorylation levels were higher when the tight junction protein had been depleted and remained about twofold higher during the entire time course (Fig. 8, A–C). Increased phosphorylation of JNK was also evident when cells were analyzed by immunofluorescence (Fig. 8 D). MEKK1 was also affected. Upon application of an osmotic shock, the kinase dispersed before accumulating again at junctions at later time points (Fig. 8 E; a different fixation method was used compared with Fig. 5 H, which reduced the cytoplasmic background but increased nonspecific nuclear staining; Fig. S4, control siRNA experiments and quantification of MEKK1 distribution). Phosphorylation of MEKK1 was induced with similar kinetics in control and depleted cells, but again, phosphorylation was more intense and sustained in depleted cells (Fig. 8, F and G). The kinetics of activation and inactivation of p38 and Erk1/2 were not affected by MarvelD3 depletion (Fig. S5, A and B). These results further support the conclusion that MarvelD3 modulates the MEKK1–JNK pathway and not Erk1/2 and p38 signaling.
MarvelD3 and the MEKK1–JNK pathway during osmotic stress. (A–J) Caco-2 cells were transfected with control and MarvelD3 siRNAs and were then osmotically stressed for the times indicated before analysis by immunoblotting (A–C and F–J), staining for phospho-Jnk (p-JNK; D) or with mouse anti-MEKK1 antibodies (E). Arrowheads in E mark cell–cell contacts positive for MEKK1. The graphs in C, G, and J show individual determinations, and the lines were drawn through the corresponding means for each time point. See Fig. S4 for specificity controls for the MEKK1 staining and quantification of junction-associated MEKK1. Bars: (D) 20 µm; (E) 10 µm.
MarvelD3 and the MEKK1–JNK pathway during osmotic stress. (A–J) Caco-2 cells were transfected with control and MarvelD3 siRNAs and were then osmotically stressed for the times indicated before analysis by immunoblotting (A–C and F–J), staining for phospho-Jnk (p-JNK; D) or with mouse anti-MEKK1 antibodies (E). Arrowheads in E mark cell–cell contacts positive for MEKK1. The graphs in C, G, and J show individual determinations, and the lines were drawn through the corresponding means for each time point. See Fig. S4 for specificity controls for the MEKK1 staining and quantification of junction-associated MEKK1. Bars: (D) 20 µm; (E) 10 µm.
MEKK1 does not only induce activation of JNK and phosphorylation of c-Jun, it can also stimulate c-Jun degradation (Xia et al., 2007). Figs. 8 H and S5 C show that there was a modest increase in phosphorylation of c-Jun before osmotic shock despite lower total expression levels in MarvelD3-depleted cells. In control cells, osmotic shock caused an increase in phosphorylated c-Jun levels. However, if compared with the total expression level of c-Jun, phosphorylation remained higher in MarvelD3-depleted cells (Fig. S5 C). Reduced expression of c-Jun in MarvelD3-depleted cells only affected protein expression and was not observed at the mRNA level (Fig. S5, D and E) and was paralleled by a decrease in c-Jun protein stability (Fig. S5, F and G), suggesting that MarvelD3 depletion indeed led to increased MEKK1-stimulated c-Jun degradation. As we had observed that MarvelD3 also regulates the SRE promoter, we next asked whether MarvelD3 depletion also affected phosphorylation of Elk1, which can also be activated by JNK (Cavigelli et al., 1995; Davis, 2000). Indeed, Elk1 phosphorylation was also enhanced in MarvelD3-depleted cells in response to hyperosmotic shock, further supporting the conclusion that MarvelD3 depletion led to enhanced MEKK1–JNK signaling (Fig. 8, I and J).
Prolonged activation of MEKK1–JNK signaling and degradation of c-Jun leads to cell death (Xia et al., 2007; Seki et al., 2012); hence, we stained nuclei in osmotically stressed cells and quantified fragmented nuclei and apoptotic bodies, hallmarks of apoptosis. Fig. 9 A shows that fragmented nuclei were more commonly observed in MarvelD3-depleted than in control cells. Quantification confirmed this increase in fragmented nuclei (Fig. 9 B). Measurements of caspase 3/7 activity and TUNEL assays confirmed that MarvelD3 depletion indeed stimulated apoptosis in response to osmotic shock (Fig. 9, C–E). MarvelD3-mediated attenuation of MEKK1–JNK signaling is thus crucial for cell survival during osmotic stress.
MarvelD3 depletion induces apoptosis during osmotic stress. (A–E) Caco-2 cells were transfected with siRNAs and then osmotically stressed as indicated. The cells were then either labeled by nuclear staining (A and B), for determination of caspase 3/7 activation (C), or using a TUNEL assay (D and E). B shows a quantification of nuclear fragmentation as an indication of apoptotic cells. C shows a quantification of caspase 3/7 induction. E shows cells fluorescently labeled in the TUNEL assay. All graphs show means ± 1 SD; n = 3 (B), n = 5 (C), and n = 8 (E). *, P < 0.05; **, P < 0.01; ***, P < 0.001. Bars, 20 µm.
MarvelD3 depletion induces apoptosis during osmotic stress. (A–E) Caco-2 cells were transfected with siRNAs and then osmotically stressed as indicated. The cells were then either labeled by nuclear staining (A and B), for determination of caspase 3/7 activation (C), or using a TUNEL assay (D and E). B shows a quantification of nuclear fragmentation as an indication of apoptotic cells. C shows a quantification of caspase 3/7 induction. E shows cells fluorescently labeled in the TUNEL assay. All graphs show means ± 1 SD; n = 3 (B), n = 5 (C), and n = 8 (E). *, P < 0.05; **, P < 0.01; ***, P < 0.001. Bars, 20 µm.
JNK stimulates MarvelD3 internalization
Our data indicate that MarvelD3 expression at the cell surface is important for the attenuation of MEKK1–JNK signaling. Therefore, we asked whether initial MAPK stimulation triggers MarvelD3 internalization.
Fig. 10 A shows that inhibitors of extracellular signal–regulated kinase (Erk; UO126) and p38 (SB202190) did not affect internalization. However, the JNK inhibitor SP600125 prevented internalization, and MarvelD3 remained concentrated at cell–cell contacts (Fig. 10, A and B). Inhibitors of src tyrosine kinases had no effect on MarvelD3 redistribution. These data thus indicate that the initial JNK activity is important for stimulation of MarvelD3 internalization.
Regulation of MarvelD3 internalization by JNK. (A and B) Caco-2 cells were preincubated with the indicated inhibitors before a 15-min osmotic shock under the same condition. The cells were then fixed and stained for MarvelD3. B shows a quantification of junction-associated MarvelD3 fluorescence; provided are means ± 1 SD; n = 8. Note, only the JNK inhibitor SP600125 prevents MarvelD3 internalization. (C) Cell contraction was assessed as described in Fig. 6 F. Shown are means ± 1 SD of three experiments counting 20–40 cells per condition and experiment. (D) Cells were preincubated with the JNK inhibitor SP600125 or DMSO (control) and then osmotically stressed under the same conditions for 8 h before fixation and staining for the junctional markers ZO-1 and p120Catenin. (E) Cells transfected with the indicated siRNAs were hyperosmotically shocked for 8 h. The JNK inhibitor SP600125 was added either together with the hypertonic medium or 15 min after shock induction. Apoptosis was then assessed with a TUNEL assay. Shown are means ± 1 SD; n = 7. **, P < 0.01; ***, P < 0.001. Bars, 10 µm.
Regulation of MarvelD3 internalization by JNK. (A and B) Caco-2 cells were preincubated with the indicated inhibitors before a 15-min osmotic shock under the same condition. The cells were then fixed and stained for MarvelD3. B shows a quantification of junction-associated MarvelD3 fluorescence; provided are means ± 1 SD; n = 8. Note, only the JNK inhibitor SP600125 prevents MarvelD3 internalization. (C) Cell contraction was assessed as described in Fig. 6 F. Shown are means ± 1 SD of three experiments counting 20–40 cells per condition and experiment. (D) Cells were preincubated with the JNK inhibitor SP600125 or DMSO (control) and then osmotically stressed under the same conditions for 8 h before fixation and staining for the junctional markers ZO-1 and p120Catenin. (E) Cells transfected with the indicated siRNAs were hyperosmotically shocked for 8 h. The JNK inhibitor SP600125 was added either together with the hypertonic medium or 15 min after shock induction. Apoptosis was then assessed with a TUNEL assay. Shown are means ± 1 SD; n = 7. **, P < 0.01; ***, P < 0.001. Bars, 10 µm.
To test whether JNK activation is indeed important for the epithelial response to osmotic stress, we tested how inhibition of the kinase affects cell behavior. Fig. 10 C shows that inhibition of JNK not only prevented MarvelD3 internalization but also cell contraction. With increasing times of stress, the cells were not only more spread, but junctional staining started to become discontinuous when JNK was inhibited and the cells started to die, indicating that activation of JNK signaling during stress is functionally important (Figs. 10 D and S5 H). However, when the inhibitor was added only 15 min after the shock had been applied, no significant increase in cell death was observed and junction dissociation was attenuated (Figs. 10 E and S5 I). If this delayed addition of the JNK inhibitor was repeated with MarvelD3-depleted cells, it strongly attenuated cell death and junction dissociation (Figs. 10 E and S5 I). Hence, inhibition of prolonged JNK signaling rescued the stress-induced phenotype of MarvelD3-depleted cells. These data thus indicate that interplay between JNK signaling and MarvelD3 regulates epithelial integrity and cell survival.
Discussion
To maintain tissue integrity and function, cells need to be able to respond to changes in their environment in a controlled manner. Here, we report the discovery of a new mechanism that links cell–cell adhesion to the regulation of cell proliferation and migration and that controls signaling in response to osmotic stress to ensure cell survival and epithelial integrity. Our data indicate that MarvelD3 links tight junctions to the control of the MEKK1–JNK pathway to guide the extent and duration of pathway activation and that this mechanism is important for the regulation of cell behavior.
Manipulation of MarvelD3 strongly affected cell migration and proliferation. MEKK1 and the JNK pathway are well-known regulators of migration (Xia et al., 2000; Yujiri et al., 2000). Similarly, MEKK1 and JNK signaling stimulate the transcription factors AP1 and Elk1, and their transcriptional activities were regulated by MarvelD3. AP1 and Elk1 promote expression of genes that stimulate proliferation, such as cyclin D1 (Cavigelli et al., 1995; Davis, 2000). Expression of cyclin D1 was strongly affected by MarvelD3 expression and followed the effect on the cyclin D1 gene promoter, indicating that cyclin D1 is indeed a target of MarvelD3 signaling. Increased migration and proliferation induced by MarvelD3 depletion were dependent on JNK signaling; hence, the JNK pathway is an important route by which MarvelD3 regulates cell behavior.
MarvelD3 inhibits JNK signaling by regulating the localization of the upstream kinase MEKK1. Our data suggest that MarvelD3 inhibits the MAPK pathway by sequestration of the MAPKKK. In vitro, the N-terminal domain of MarvelD3 pulled down MEKK1 from cell extracts and interacted with the kinase domain in reconstitution experiments with recombinant proteins. Hence, junctional sequestration may involve formation of MarvelD3–MEKK1 complexes. However, we have not been able to coimmunoprecipitate MarvelD3 and MEKK1 from cell extracts, despite the biotinylation assay indicating that MEKK1 localizes in close proximity of MarvelD3. The reasons for this are not clear but may have sterical reasons, as the MarvelD3 antibodies had been generated against the N-terminal domain, which also binds MEKK1; unfortunately, the MEKK1 antibodies we tested immunoprecipitated the kinase only poorly or not at all. Alternatively, the interaction may not be stable as a result of its biochemical and/or functional properties. It is possible that the role of the interaction between MEKK1 and MarvelD3 is a regulatory one. MEKK1 has previously been shown to bind to α-actinin, and α-actinin isoforms localize to tight and adherens junctions (Christerson et al., 1999; Patrie et al., 2002). Hence, MEKK1 might be anchored to the junction by binding to α-actinin, and MarvelD3 might regulate kinase activation and dissociation from the junction.
A model in which MarvelD3 regulates junctional localization and, thereby, inactivation of MEKK1, is also supported by the osmotic shock experiments, as MarvelD3 was internalized and MEKK1 became dispersed. Whether removal of MarvelD3 from junctions by osmotic stress or siRNA-mediated depletion is the only factor that contributed to the release of MEKK1 is not known. MEKK1 is activated by RhoA signaling, which is stimulated by growth factors and osmotic shock; hence, it may also be involved in the release of MEKK1 from tight junctions (Gallagher et al., 2004).
Depletion of MarvelD3 not only affected JNK activation but also c-Jun expression levels. MEKK1 can function as a ubiquitin ligase that stimulates the degradation of c-Jun (Xia -et al., 2007). Hence, the decreased levels of c-Jun in MarvelD3-depleted cells are likely to be caused by deregulation of MEKK1. Both c-Jun degradation and deregulation of JNK can induce cell death (Davis, 2000; Seki et al., 2012). It is thus likely that the combined action of these two mechanisms caused the increase in apoptosis that we observed in MarvelD3-depleted cells in response to osmotic stress.
MarvelD3 belongs to a growing family of tight junction proteins that do not seem to play a fundamental structural role in forming the junctional diffusion barrier (Steed et al., 2009; Raleigh et al., 2010). Together with the data presented here, this suggests that the main function of MarvelD3 is a regulatory one, transmitting signals from the junction to the cell interior to guide MEKK1–JNK signaling. As indicated by the osmotic stress experiments, however, depletion of MarvelD3 can also lead to barrier failure caused by cell death and junction dissociation. Hence, the experimental conditions under which such experiments are performed and the type of model systems used can dramatically affect the results and may explain the variations observed in different studies on the role of MarvelD3 in junction formation (Steed et al., 2009; Raleigh et al., 2010; Kojima et al., 2011).
Another prominent example of this group of tight junction transmembrane proteins is Occludin (Furuse et al., 1993). Occludin does also not play a major structural role but functions as a regulator (Balda et al., 1996; McCarthy et al., 1996; Chen et al., 1997; Wong and Gumbiner, 1997; Antonetti et al., 1998, 1999; Matter and Balda, 1998; Saitou et al., 1998, 2000). Occludin and MarvelD3 are both Marvel domain proteins (Sánchez-Pulido et al., 2002; Steed et al., 2009; Raleigh et al., 2010), and they have some similarities in their biological function and behavior. Occludin has been shown to cross talk with the Raf-1–Erk signaling cascade (Li and Mrsny, 2000; Wang et al., 2005). Hence, MarvelD3 and Occludin both functionally interact with MAPK signaling; however, they do so in fundamentally different ways. Occludin performs a role downstream of Erk, as it does not affect activity of Erk but counteracts junction dissociation induced by oncogenic Raf-1 (Li and Mrsny, 2000; Wang et al., 2007). Hence, Occludin links the Raf-1–Erk pathway to junction regulation, whereas MarvelD3 links the junction to regulation of MEKK1–JNK signaling.
Another similarity between Occludin and MarvelD3 is that both are regulated by membrane traffic. Occludin is internalized via a caveolin-1–based mechanism in response to TNF signaling (Marchiando et al., 2010). MarvelD3 internalization is triggered by osmotic stress and requires JNK signaling. After initial internalization, MarvelD3 then recycles to the cell surface and becomes again concentrated at tight junctions. The mechanisms underlying internalization and recycling remain to be identified. However, the dependence on JNK activity for internalization indicates that MarvelD3 recycling functions as a regulatory feedback mechanism: initial removal from the cell surface allows efficient activation of the MEKK1–JNK pathway followed by inhibition upon reappearance and accumulation at tight junctions. Both steps seem to be important for the epithelial stress response, as inhibition of JNK not only inhibited MarvelD3 internalization but also led to increased cell death, whereas MarvelD3 depletion led to prolonged JNK signaling and increased cell death, and this could be rescued by inhibiting JNK after the initial 15 min of hypertonic stress. MarvelD3 thus functions as a negative regulator that tunes JNK signaling.
In this study, we demonstrate that MarvelD3 links tight junctions to the MEKK1–JNK pathway and exerts a negative feedback activity on the MAPK pathway that regulates cell proliferation, migration, and survival. The signaling activity of MarvelD3 may also be relevant in disease. Given the effect of MarvelD3 on cell migration and proliferation, it may have tumor suppressor activity. However, MEKK1 has recently been suggested to function as a tumor suppressor in breast cancer (Banerji et al., 2012; Cancer Genome Atlas Network, 2012; Ellis et al., 2012). An active MEKK1–JNK pathway may thus be beneficial for patients as tumor cells might die as a result of prolonged JNK signaling; hence, MarvelD3 expression may promote tumor cell survival by inhibiting MEKK1–JNK signaling. The role of the MarvelD3–MEKK1–JNK pathway and how it affects tumorigenesis and disease progression may thus depend on the type of tumor and the tumor microenvironment.
Materials and methods
Cell culture
Caco-2, MiaPaca-2, and MCF7 cells were grown in DMEM (Invitrogen) supplemented with 20% FBS (Invitrogen) for Caco-2 and 10% FBS for MiaPaca-2 and MCF7 cells. All cell lines were grown in the presence of 100 U/ml penicillin and 100 µg/ml streptomycin (Invitrogen) at 37°C in a 5% CO2 atmosphere. For hyperosmotic shock treatments, the medium was brought to 600 mOsm using NaCl.
siRNAs, cDNAs, and RT-PCR
For knockdown experiments, Caco-2 cells were transfected with individual or a pool of MarvelD3-specific siRNAs (Thermo Fisher Scientific) that were 5′-GAGAGGAGGTGGAATATTA-3′ and 5′-GCCGCAGACCCGAAAGTGA-3′ as previously described (Steed et al., 2009). MEKK1 was targeted with the following siRNAs: 5′-GAACAGAUGUGUCCUAUUU-3′, 5′-GAGCUCAACUCCAGUAUAUUG-3′, 5′-AGAAGAAGCUUUAGCAAUU-3′, and 5′-GCUGAAAGGUCAACAGAUA-3′. INTERFERin transfection reagent (Polyplus Transfection, Inc.) was used for transfections according to the manufacturer’s instructions. Effects of depletion were assayed 3 d after transfection. cDNAs encoding untagged full-length MarvelD3 isoforms and full-length Occludin were expressed using pcDNA4 and pCB6 expression vectors; both vectors have cytomegalovirus promoters for eukaryotic expression (Balda et al., 1996; Steed et al., 2009). Vesicular stomatitis virus (VSV)–tagged versions of full-length and truncated cDNAs were constructed as described by adding the VSV epitope tag (i.e., GYTDIEMNRLGK) to the C-terminal end of the full-length cDNAs and were then then expressed using pcDNA4 (Benais-Pont et al., 2003). The mouse MarvelD3 isoform 1 cDNA was amplified from total RNA isolated from mouse intestine using the primers 5′-CCGGAATTCCGATGAAAAATACTTCGGGGCAC-3′ and 5′-ATGCAGTGGCTAGCGAGCGTCCCGGACCACAGGTATTC-3′ and cloned into pCIN4-VSV. MarvelD3/biotin ligase fusion proteins were generated using the plasmids described by Roux et al. (2012) for N- and C-terminal fusions by cloning the full-length MarvelD3 isoform 1 cDNA into pcDNA3.1-mycBirA-R118G and pcDNA3.1-mycBirA-R118G. For the production of His6-tagged MEKK1 fusion proteins, fragments were amplified from a rat MEKK1 cDNA (provided by M. Cobb, University of Texas Southwestern Medical Center, Dallas, TX) using the primer pairs 5′-GAAGGCCTCGAGATGGCGGCGGCGGCGGGCGATC-3′ and 5′-GATCAGAATTCTACGCCTCCACCACGTCCCCTGAG-3′; 5′-GAAGGCCTCGAGGATGTTAGCGGGGCCCTGTTGTTG-3′ and 5′-GATCAGAATTCTAGAGCGCCTCCTCTTCCTCAGCCTC-3′; and 5′-GAAGGCCTCGAGGACGTCAATCACAATCAAAAGTGC-3′ and 5′-GATCAGAATTCTACCATGTTGTACGGAAGACAGGATG-3′. For expression experiments, plasmids were transfected into Caco-2 and MiaPaca-2 cells using transfection reagent (jetPEI; Polyplus Transfection, Inc.) according to the manufacturer’s instructions. For the generation of stable cell lines, MiaPaca-2 cells plated at 50% confluence in complete medium with 2.5% horse serum (Invitrogen) were transfected 24 h later with 20 µg pCIN4-MD3_1:VSV and incubated for 8 h at 37°C. Cells were then trypsinized, resuspended in 25 ml complete medium, and plated onto 15-cm plates and selected with 0.6 mg/ml G418. After 17 d, clones were picked and plated in complete medium containing 0.6 mg/ml G418. Transfection efficiency was determined by immunoblotting and immunofluorescence. Expression of MD3_1:VSV was established in two independent clones (C2 and C8). MD3–MiaPaca-2 C2 and C8 were cultured in DMEM containing 10% FBS and 3 mg/ml G418. All experiments were performed in the absence of G418. Fluorescent MiaPaca-2 cells were generated using lentiviral vectors expressing humanized GFP or RFP under the control of a spleen focus-forming virus promoter (Georgiadis et al., 2010). Viral packaging using pMD2VSVG was performed as previously described (Sourisseau et al., 2006). Caco-2 cell lines were generated with calcium phosphate transfections as previously described (Matter et al., 1992). Reverse transcription was performed on total RNA isolated from MiaPaca-2 cells and pancreatic RNA (Takara Bio Inc.) at 45°C using avian myoblastosis virus reverse transcription (Promega) for 1 h. The cDNAs were then amplified by PCR using polymerase (FastStart Taq; Roche) at an annealing temperature of 63°C using primers previously described (Steed et al., 2009). The following primers were used to detect expression of human MarvelD3 isoforms: 5′-AAAAATCTAGATCAAAGAGTTCCAGACCACAG-3′ and 5′-AAAAATCTAGATTAAAATTCAAACATTTCTGCTGG-3′ for reverse transcription and 5′-AAAAAGCTAGCAAATACTTGTGCACTGGGAGAG-3′, 5′-AAAAAGCTAGCTCGGTAGCTACGCAGGGC-3′, and 5′-AAAAAGCTAGCTCGGTAGCCCTTTATGGCCA-3′ for PCR. For the detection of c-Jun mRNA expression, total RNA was reverse transcribed with the primer 5′-TCAAAATGTTTGCAACTGCTGCG-3′, and a 676-bp fragment was amplified with the primer pair 5′-CGTTAGCATGAGTTGGCACCCACTG-3′ and 5′-CAAGAACGTGACAGATGAGCAGGAGG-3′. For quantitative PCR, the latter primer was replaced by 5′-AGCAGCAGCAGCAGCCGCCGCACCACC-3′, and reactions were performed using Power SYBR Green PCR Master Mix and a thermal cycler (7900HT; Applied Biosystems; Nie et al., 2012).
Antibodies
A rabbit anti-MarvelD3 antibody was generated against the complete N terminus of MarvelD3 (amino acids 1–198; Steed et al., 2009). The rabbit anti–phospho-MEKK1 antibody was generated against a phosphorylated peptide containing serine 1436 of mouse MEKK1 (serine 1455 of human MEKK1), which had previously been shown to become phosphorylated in T cells upon T cell receptor stimulation, and was generated and purified as previously described (Gallagher et al., 2002; Enzler et al., 2009). Antibodies against other proteins were as previously described or were obtained from commercial sources: rabbit anti-MEKK1 (Abcam); mouse anti-pJNK, rabbit anti-JNK, mouse anti-pErk, rabbit anti-Erk, and rabbit anti-pp38 and -p38 (Cell Signaling Technology); goat anti-p120Catenin, mouse anti-pELK1, rabbit anti-ELK1, mouse anti-MEKK1, and goat anti-MEKK3 and anti-MLK1 (Santa Cruz Biotechnology, Inc.); rabbit anti–cyclin D1 (MBL International); mouse anti-His6 (Sigma-Aldrich); mouse anti-Occludin and –ZO-1 (Life Technologies); mouse anti-EEA1 (BD); mouse anti–α-tubulin (antigen: a peptide containing the last 11 C-terminal amino acids of porcine α-tubulin) and mouse anti–α-VSV (antigen: the peptide N-CGYTDIEMNRLGK-C; Kreis and Lodish, 1986; Kreis, 1987); rabbit anti–ZO-1 (antigen: the peptide N-YTDQELDETLNDEVC-C), –ZO-2 (antigen: the peptide N-KMEGMDDDPEDRMSC-C), and –ZO-3 (antigen: the peptide N-CSDEDGYDWGPATDL-C; Benais-Pont et al., 2003); rabbit anti-GST (antigen: purified recombinant GST; Kavanagh et al., 2006); and mouse anti–transferrin receptor (antigen: isolated Golgi-enriched membranes from Caco-2 cells) and mouse anti-LAMP1 (antigen: isolated Golgi-enriched membranes from Caco-2 cells; Schweizer et al., 1988). Phalloidin was used to visualize F-actin. All secondary antibodies and fluorescently labeled streptavidin were obtained from Jackson ImmunoResearch Laboratories, Inc.
Immunofluorescence
Caco-2 or MiaPaca-2 cells were grown on glass coverslips for 3 d and then processed for immunofluorescence. Cells were fixed directly in methanol at −20°C for 10 min or with 3% paraformaldehyde for 20 min followed by permeabilization with 0.3% Triton X-100 as previously described (Matter and Balda, 2003; Steed et al., 2009). For the localization of EEA1, cells were fixed with 1% paraformaldehyde and for transferrin receptor with 1% paraformaldehyde followed by methanol before extracting for 10 min with 0.3% Triton X-100. Cells were then washed with PBS and blocked for 15 min with PBS containing 0.5% BSA (PBS-BSA) at room temperature. Coverslips were incubated with primary antibodies overnight, washed three times with PBS-BSA, and incubated with secondary antibody and Hoechst nuclear stain at room temperature. Coverslips were washed twice more with PBS-BSA and once in PBS before being mounted with mounting medium (ProLong Gold; Invitrogen) and stored at 4°C. Epifluorescent images were collected at ambient temperature with a fluorescent microscope (DMIRB; Leica) using a Apochromat 63×, 1.4 NA oil immersion objective fitted with a camera (C4742-95; Hamamatsu Photonics) and SimplePCI software (Hamamatsu Photonics) or an inverted microscope (Eclipse Ti-E; Nikon) using an Apochromat 60×,1.4 NA oil immersion objective, a camera (CoolSNAP HQ2; Photometrics), and Nikon software. Brightness and contrast were adjusted with Photoshop CS4 software (Adobe). ImageJ (National Institutes of Health) was used for quantifications of cell areas and fluorescent densities.
Immunoblotting
Whole-cell lysates were harvested by adding SDS sample buffer (3% SDS, 15% glycerol, 0.0015% bromophenol blue, and 0.25 M Tris HCl containing 150 mM DTT and 8 M urea) to samples washed once with PBS. Samples were homogenized through a 25-gauge needle and, after a 30-min incubation at room temperature, analyzed by SDS-PAGE. Proteins were transferred to nitrocellulose or polyvinylidene difluoride membranes, and bound primary antibodies were detected with horseradish peroxidase–conjugated secondary antibodies and an ECL detection system (ECL; GE Healthcare) or with IRDye680- and IRDye800-labeled secondary antibodies and an infrared scanner (Odyssey; LI-COR Biosciences; Elbediwy et al., 2012). Immunoblots were quantified with ImageJ, and the provided n numbers refer to analyzed experiments.
GST pull-downs and isolation of biotinylated proteins
Fragments encoding the N-terminal domain and isoform-specific C-terminal domains (CTD1 and CTD2) of MarvelD3 were cloned in to pGEX-4T-3 GST fusion vector. Fusion proteins were produced by transforming plasmids into competent BL21 pLysS cells, and production was induced by the addition of 1 mM isopropyl β-d-1-thiogalactopyranoside and incubation at 25°C for 2 h. Fusion proteins were then recovered by lysing cells in PBS containing 0.5% Triton X-100 (PBSTx) and a cocktail of protease and phosphatase inhibitors (50 µg/ml benzamidine, 10 µg/ml aprotinin, 10 µg/ml leupeptin, 10 µg/ml pepstatin A, and 1 mM PMSF) and 1 mM DTT. Samples were sonicated and centrifuged at 10,000 rpm, and the supernatant was stored at −80°C until use. To identify interaction partners and the responsible domains of MarvelD3, GST fusion proteins were conjugated to glutathione beads for 2 h at 4°C. Extracts of Caco-2 cells grown on 15-cm cell culture dishes until 80% confluent were generated with PBS containing 0.5% Triton X-100 (PBSTx) and preabsorbed with unconjugated Sepharose beads. The extracts were then incubated with the beads carrying the GST fusion proteins for 2 h at 4°C. The samples were washed once in PBSTx and twice in PBS before being separated by SDS-PAGE. For recombinant protein-binding assays, the His6-tagged protein was added to the GST fusion protein bound to beads. After an overnight incubation at 4°C, the beads were washed twice with PBSTx before immunoblotting. For the isolation of biotinylated proteins, cells were extracted with 10 mM Hepes, pH 7.4, containing 150 mM NaCl, 1% Triton X-100, 0.5% sodium deoxycholate, 0.2% sodium dodecylsulfate, and a cocktail of protease inhibitors. After centrifugation, the extracts were loaded on streptavidin–agarose beads and, after 2 h, washed four times with extraction buffer and once with PBS.
Migration assays
To assess cell migration, MiaPaca-2 and Caco-2 cells were applied to both 0.22-mm2 chambers of a 9 × 9 × 5–mm culture insert (µ-Dish; ibidi) in 70 µl complete medium. For knockdown experiments, cells were transfected in 24-well plates and transferred to culture inserts 24 h later. In all experiments, when confluent, inserts were gently removed to reveal a 500-µm cell-free space between monolayers, and the medium was replaced with fresh complete medium containing mitomycin c to inhibit proliferation. Phase-contrast images were taken at regular time points until the most rapidly migrating cells showed ∼80% wound closure. Quantification of wound closure was made using ImageJ software (percentage of wound closure = 100 − [(cell-free area at end point)/(cell-free area at time 0) × 100].
Proliferation and cell death assays
To assess proliferative behavior in vitro, MiaPaca-2 or Caco-2 cells (control and depleted of MarvelD3 by siRNA transfection) were plated in triplicate into 96-well plates at equal cell densities. Medium was removed after 3 d, and samples were frozen at −80°C. Cell numbers were quantified the next day using a CyQUANT Cell Proliferation Assay kit (Invitrogen) according to the manufacturer’s instructions. Fluorescence was determined using a microplate reader and converted into cell numbers against a reference standard curve for each cell type. For inhibition of JNK experiments, cells were incubated in medium containing 0.2% FBS for 24 h, plus 20 µM SP600125 or DMSO for the last 3 h. Complete medium containing SP600125 or DMSO was then added for an additional 24 h, and cell number was quantified. Induction of apoptosis and necrosis was assayed measuring caspase 3/7 activities and release of lactate dehydrogenase activity, respectively, using the respective fluorescent assay kits obtained from Promega (Nie et al., 2012). The DeadEnd Fluorometric TUNEL System obtained from Promega was used for the TUNEL analysis.
Xenografts
Mice were raised and treated according to the principles of laboratory animal care of the University Animal Welfare Committee. After a 7-d adaptation period, 6–8-wk-old female athymic nude mice (Harlan Laboratories) were injected subcutaneously into the right flank region with 5 × 106 MiaPaCa-2 cells suspended in PBS containing 50% growth factor–reduced Matrigel (BD; Kyriazis and Kyriazis, 1980). Tumor growth was monitored with a digital caliper, and their volume was estimated according to the following formula: V = (S2 × L)/2, in which S and L are the smaller and larger measures of the tumor, respectively (Tomayko and Reynolds, 1989). At the end of experiments, mice were sacrificed by cervical dislocation, and tumors were excised for imaging.
Impedance measurements
Impedance was measured in response to hyperosmotic shock using an Electric Cell-Substrate Impedance Sensing system (model 1600R; Applied BioPhysics). Control and MD3p siRNA–transfected Caco-2 cells were plated into an Electric Cell-Substrate Impedance Sensing electrode array containing 10 electrodes per well (8W10E; Applied BioPhysics). Array plates were first cleaned with 200 µl per well of 10 mM cysteine for 10 min and then washed three times with PBS. Cells were plated in quadruplicate and grown at 37°C for 48 h until confluent. Basal impedance across the monolayer was measure at 37°C for 1 h. Complete medium was then replaced with hypertonic medium (600 mOsm), and measurements of impedance were taken at 2-min intervals for a further 10 h.
Reporter assays
Cells were plated in 96-well plates in complete medium the day before transfection. For overexpression experiments, cells were cotransfected with cDNA encoding full-length MarvelD3 isoform 1, isoform 2, or Occludin or empty pCB6 vector (control) alongside a firefly luciferase reporter construct and Renilla luciferase control plasmid, with jetPEI reagent. The AP1 and SRE firefly luciferase reporter constructs were obtained from Takara Bio Inc., and the T cell factor/β-catenin reporter plasmid was obtained from Merck EMD Millipore. The ZONAB firefly reporter plasmid contained a minimal ErbB-2 promoter (−396 to −150 in pXb-1), and the Renilla control plasmid contained an analogous ErbB-2 promoter in pRL but in which the ZONAB binding site had been inactivated by substituting the inverted CCAAT box (Frankel et al., 2005). The cyclin D1 reporter plasmid contained a 1.7-kb promoter fragment in pXb1 (Sourisseau et al., 2006). Transfection mix was changed for fresh medium 24 h after transfection. Luciferase luminescent signals were determined 24 h later using the assay kit (Dual-Luciferase Reporter; Promega). Firefly and Renilla luciferase luminescent signals were measured sequentially using a microplate reader (FLUOstar OPTIMA; BMG Labtech). Relative promoter activity was calculated by making a ratio between luciferase and Renilla luminescent signals and then normalizing to control transfections. For knockdown experiments, cells were transfected with siRNA 24 h after plating, and medium was changed after 24 h and transfected with reporter constructs another 24 h later. Transfection mix was changed for fresh medium 24 h later, and luminescence signals were read after a further 8 h.
Statistical analysis
Means and SDs were calculated and provided in the graphs. Respective n values are provided in the figure legends. The indicated p-values were obtained with two-tailed Student’s t test.
Online supplemental material
Fig. S1 shows the effect of MarvelD3 siRNAs on proliferation and migration of MiaPaca-2 and MCF7 cells. Fig. S2 shows the effect of ectopic MarvelD3 expression in MiaPaca-2 cells and other junctional proteins by immunofluorescence and immunoblotting as well as intermixing of MiaPaca-2 cells in the presence or absence of MarvelD3 expression. Fig. S3 shows the effect of osmotic shock on MarvelD3 expression levels in the presence of cycloheximide, double immunofluorescence of MarvelD3 with endocytic markers, quantification of MarvelD3 and Occludin localization during osmotic shock, and the distribution of Occludin in MarvelD3-depleted cells after 2 h of hypertonic shock. Fig. S4 shows the localization of MEKK1 in paraformaldehyde-fixed cells and quantification of MEKK1 localization during osmotic shock. Fig. S5 shows the activation profiles of p38 and Erk in control and MarvelD3-depleted Caco-2 cells in response to osmotic shock, c-Jun mRNA levels and protein stability in control, and MarvelD3-depleted cells, quantification of TUNEL-positive cells in response to JNK inhibitor and osmotic shock, and attenuation of junction disassembly by JNK inhibition in MarvelD3-depleted cells.
Acknowledgments
This work was supported by the Biotechnology and Biological Sciences Research Council (BB/J015032/1), the Wellcome Trust (084678/Z/08/Z, 099173/Z/12/Z, and WT090939MA), Cancer Research UK (C26616/A12679), the Université Catholique de Louvain (Actions de Recherche Concertées), and the Fonds de la Recherche Scientifique (Belgium). S. Dupasquier was a postdoctoral fellow and C.E. Pierreux is senior research associate of the Fonds de la Recherche Scientifique (Belgium).
The authors declare no competing financial interests.
References
Author notes
M.S. Balda and K. Matter contributed equally to this paper.
E. Steed’s present address is Institut de Génétique et de Biologie Moléculaire et Cellulaire, 67404 Illkirch, Strasbourg, France.
A. Elbediwy’s present address is London Research Institute, London WC2A 3LY, England, UK.
S.A. Hemkemeyer’s present address is Institute of Molecular Cell Biology, Westphalian Wilhelms University, 48149 Münster, Germany.
A.C. Costa’s present address is Group of Molecular Microbiology, Institute for Molecular and Cell Biology, 4150-180 Porto, Portugal.