Maintenance of stem cell properties is associated with reduced proliferation. However, in mouse hematopoietic stem cells (HSCs), loss of quiescence results in a wide range of phenotypes, ranging from functional failure to extensive self-renewal. It remains unknown whether the function of human HSCs is controlled by the kinetics of cell cycle progression. Using human HSCs and human progenitor cells (HSPCs), we report here that elevated levels of CCND1–CDK4 complexes promoted the transit from G0 to G1 and shortened the G1 cell cycle phase, resulting in protection from differentiation-inducing signals in vitro and increasing human leukocyte engraftment in vivo. Further, CCND1–CDK4 overexpression conferred a competitive advantage without impacting HSPC numbers. In contrast, accelerated cell cycle progression mediated by elevated levels of CCNE1–CDK2 led to the loss of functional HSPCs in vivo. Collectively, these data suggest that the transition kinetics through the early cell cycle phases are key regulators of human HSPC function and important for lifelong hematopoiesis.
The continuous supply of de novo generated mature cells from adult stem cells is pivotal for the lifelong function of many organs, particularly tissues with high turnover rates such as the gut, skin, and blood. Continued tissue formation requires precise balancing of quiescence, self-renewal, and differentiation of stem cells over extended periods of time. Hematopoietic stem cells (HSCs) are routinely used in the clinic for the replacement of diseased blood tissues. Often, the limiting factor for successful clinical HSC transplantation is the availability of only low numbers of histocompatible donor cells, and understanding the regulation of HSC self-renewal and output may be a critical step toward overcoming this obstacle. Although considerable knowledge regarding cell cycle–mediated regulation of HSC function has been obtained over the last decade in mice (Pietras et al., 2011; Nakamura-Ishizu et al., 2014), very little information regarding cycle-associated regulatory circuits in human HSCs is presently available. Moreover, data suggest that human cell cycle kinetics and progenitor population dynamics are not well recapitulated in the mouse (Sykes and Scadden, 2013).
Although large fractions of progenitor populations divide, most immature long-term reconstituting HSCs are quiescent and thought to be protected from the accumulation of damage that contributes to leukemia and aging (Trumpp et al., 2010). Nevertheless, the HSC pool is maintained through self-renewing divisions tightly regulated by enzymatically active cyclin (CCN)/cyclin-dependent kinase (CDK) complexes that are controlled by CDK inhibitors (CKIs). However, how fate decisions between self-renewal versus differentiation are integrated in cycling activity is not known. The G1 phase of the cell cycle is divided into the mitogen-dependent early phase and a mitogen-independent late phase, and progression through these phases depends on CCND1,2,3/CDK4,6 and CCNE1,2/CDK2 complexes, respectively (Orford and Scadden, 2008). Signaling through growth factor receptors induces the expression of d-type cyclins, leading to the accumulation of active CCND1,2,3/CDK4,6 complexes that phosphorylate members of the retinoblastoma (Rb) tumor suppressor protein, resulting in the exit from quiescence (G0) and transition through G1 phase. Subsequent release of the E2F family of transcription factors from Rb results in transcription of CCNE1 followed by the transit from early to late G1 phase (Orford and Scadden, 2008; Pietras et al., 2011). Whereas the S, G2, and M phase lengths are comparable between cells of different origins, the entry and progression through the G1 cell cycle phase depend on the cell type and environmental context, suggesting that G1 transition is linked to functional decisions in stem cells (Massagué, 2004; Blomen and Boonstra, 2007; Orford and Scadden, 2008; Singh and Dalton, 2009; Pietras et al., 2011). Further, it has been proposed for embryonic stem cells and one adult stem cell type, neural stem cells, that a prolonged lack of cycling activity and extended time in G1 may allow the integration of signals necessary and sufficient for the initiation of differentiation, whereas a short retention time in G1 leads to the maintenance of self-renewal potential (Calegari and Huttner, 2003; Orford and Scadden, 2008; Singh and Dalton, 2009). Whether cell cycle phase length is a mechanism controlling hematopoietic stem cell function has been speculated on (Orford and Scadden, 2008) but not yet shown.
The effects on cycling activity and function of murine HSCs greatly differ in the absence of negative cell cycle regulators of the INK4 and CIP/KIP families and range from dramatic expansion to complete loss of functional HSCs (Orford and Scadden, 2008; Pietras et al., 2011). Further, it remains unclear whether exit from quiescence rather than progression through distinct periods of G1 or G1-to-S transition provides a regulatory platform for HSC function. To directly test this hypothesis, we enforced expression of functional CCND1–CDK4 or CCNE1–CDK2 complexes (together referred to as 4D or 2E) that are important for progression through early G1 and G1-to-S transition, respectively. We show that the progression kinetics through the early and late G1 phases of the cell cycle regulate the self-renewal of HSCs in vivo, providing a new regulatory platform for the regeneration of this highly dynamic tissue.
RESULTS
4D regulates G0-to-G1 transition and G1 phase length in human HSPCs
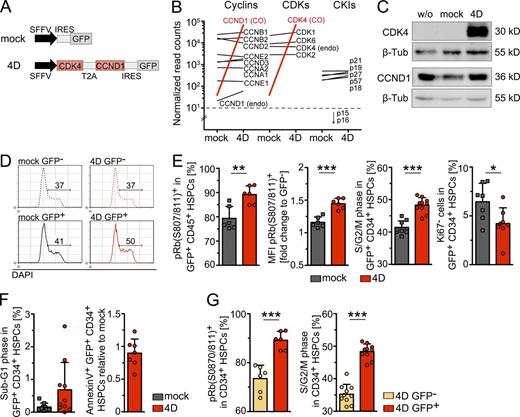
To stably manipulate early cell cycle transition of human hematopoietic stem and progenitor cells (HSPCs), CD34+ enriched cord blood (CB) cells were transduced with CCND1–CDK4 also encoding a GFP as reporter (Fig. 1 A). Overexpression of codon-optimized transcripts of CCND1 and CDK4 was verified by whole transcriptome sequencing (Fig. 1 B). Coverage of the codon-optimized sequences revealed expression of the complete construct (Fig. S1). The expression levels of other cell cycle regulatory factors, including endogenous CCND1 and CDK4, were not altered by overexpression of 4D (Fig. 1 B). Protein expression was verified in transduced NIH3T3 cells (Fig. 1 C). 4D was catalytically active in human CB CD34+ cells because cell cycle profiles shifted in 4D- but not in mock-transduced cells (Fig. 1 D,E) and increased numbers of transduced cells carried phosphorylated retinoblastoma protein (pRb, serine 807/811, Fig. 1 E). Further, the mean fluorescence intensity (MFI) of pRb was increased, corroborating its hyperphosphorylation in successfully 4D-transduced cells. Consistently, greater percentages of HSPCs were found in the S–G2–M phase of the cell cycle, suggesting a decrease in the proportion of quiescent cells in G0 and entry in the cell cycle. In fact, fewer HSPCs lacked the proliferation marker Ki-67 compared with controls (Fig. 1 E). The absence of apoptotic cells as measured by DNA fragmentation (Sub-G1 phase; Fig. 1 F; see also Fig. 5 B) or Annexin V staining (Fig. 1 F) indicated that 4D overexpression was not toxic. Rb phosphorylation and a shift in the cell cycle profile was exclusively detected in transduced, GFP+ HSPCs compared with nontransduced GFP− cells from the same transduction sample, providing a direct control for the specificity of GFP expression and activity of the 4D construct (Fig. 1 G). Collectively, these results indicate that ectopically expressed 4D complexes are functionally active in human HSPCs.
Elevated CCND1–CDK4 (4D) expression induces a G0-to-G1 transition and increased signaling in human HSPCs. (A) Scheme of the construct: CCND1 and CDK4 are linked by T2A, followed by an IRES-GFP-reporter sequence (4D). The control vector lacks CCND1 and CDK4 sequences (mock). (B) Expression analysis was performed on 4D- or mock-transduced human CD34-enriched cord blood (CB) cells by deep sequencing, and the plot shows the difference of the mean normalized read counts per condition (mock, 4D) of genes related to cell cycle (cyclins, CDKs and CKIs). WThe following criteria were applied to determine differential expression of genes (marked in red): false discovery rate (FDR) of 5% and a log2 fold change of >1, respectively ≤1. Read counts <10 were considered as background levels (dashed line). Due to usage of codon-optimized sequences CDK4 read counts were annotated to a different gene but read coverage of the codon-optimized (CO) CDK4 sequence revealed its expression in 4D- but not in mock-transduced samples (Fig. S1). Data were pooled from two independent biological replicates using two and three pooled CB samples. Endo, endogenous. (C) Western blot of CDK4 and CCND1 overexpressed in mouse NIH3T3 cells. The antibody recognizing CDK4 is specific for human protein, the antibody recognizing CCND1 is mouse–human cross-reactive. Data shown is representative of three independent experiments. (D) Cell cycle profiles of 4D- and mock-transduced human CD34+ CB HSPCs 3 d after transductions. 1 CB sample was used for the shown experiment. (E) Plots show the frequency (left) and MFI (middle left) of phosphorylated Rb at S807/811. Frequencies of cells in S–G2–M phase (middle right), and frequencies of Ki-67 negative cells (right) are shown. Graph bars depict independent values from 6–10 experiments. In two experiments, triplicates were measured to determine S–G2–M phase and Ki-67 expression, and the mean is depicted. Due to paucity of cells in 6 (pRb), 5 (Ki67), and 8 (S–G2–M) experiments, single measurements were performed and shown. Viability of GFP+ CD34+ HSPCs was determined before each assay using DAPI or Sytox negativity and found >97% in all cases. Two to five pooled CB samples were used for each experiment. (F) Frequencies of cells undergoing DNA fragmentation (sub-G1) are shown (left). Sample preparation as described in E. (right) The frequency of apoptotic cells after transduction with 4D relative to control-treated HSPCs 3 d after transduction. Data are pooled from three independent experiments. Single cord blood sample were used for each experiment. (G) Plots show frequencies of pRb(S807/811)+ cells and cells in the S–G2–M phase among GFP+ and GFP− cells of the same probe. Sample preparation as described in E. *, P = 0.05–0.01; **, P = 0.01–0.001; ***, P < 0.001.
Elevated CCND1–CDK4 (4D) expression induces a G0-to-G1 transition and increased signaling in human HSPCs. (A) Scheme of the construct: CCND1 and CDK4 are linked by T2A, followed by an IRES-GFP-reporter sequence (4D). The control vector lacks CCND1 and CDK4 sequences (mock). (B) Expression analysis was performed on 4D- or mock-transduced human CD34-enriched cord blood (CB) cells by deep sequencing, and the plot shows the difference of the mean normalized read counts per condition (mock, 4D) of genes related to cell cycle (cyclins, CDKs and CKIs). WThe following criteria were applied to determine differential expression of genes (marked in red): false discovery rate (FDR) of 5% and a log2 fold change of >1, respectively ≤1. Read counts <10 were considered as background levels (dashed line). Due to usage of codon-optimized sequences CDK4 read counts were annotated to a different gene but read coverage of the codon-optimized (CO) CDK4 sequence revealed its expression in 4D- but not in mock-transduced samples (Fig. S1). Data were pooled from two independent biological replicates using two and three pooled CB samples. Endo, endogenous. (C) Western blot of CDK4 and CCND1 overexpressed in mouse NIH3T3 cells. The antibody recognizing CDK4 is specific for human protein, the antibody recognizing CCND1 is mouse–human cross-reactive. Data shown is representative of three independent experiments. (D) Cell cycle profiles of 4D- and mock-transduced human CD34+ CB HSPCs 3 d after transductions. 1 CB sample was used for the shown experiment. (E) Plots show the frequency (left) and MFI (middle left) of phosphorylated Rb at S807/811. Frequencies of cells in S–G2–M phase (middle right), and frequencies of Ki-67 negative cells (right) are shown. Graph bars depict independent values from 6–10 experiments. In two experiments, triplicates were measured to determine S–G2–M phase and Ki-67 expression, and the mean is depicted. Due to paucity of cells in 6 (pRb), 5 (Ki67), and 8 (S–G2–M) experiments, single measurements were performed and shown. Viability of GFP+ CD34+ HSPCs was determined before each assay using DAPI or Sytox negativity and found >97% in all cases. Two to five pooled CB samples were used for each experiment. (F) Frequencies of cells undergoing DNA fragmentation (sub-G1) are shown (left). Sample preparation as described in E. (right) The frequency of apoptotic cells after transduction with 4D relative to control-treated HSPCs 3 d after transduction. Data are pooled from three independent experiments. Single cord blood sample were used for each experiment. (G) Plots show frequencies of pRb(S807/811)+ cells and cells in the S–G2–M phase among GFP+ and GFP− cells of the same probe. Sample preparation as described in E. *, P = 0.05–0.01; **, P = 0.01–0.001; ***, P < 0.001.
To determine whether 4D overexpression affects the length of the G1 phase, we fitted a mathematical cell cycle model to cumulative BrdU incorporation in vitro, identifying the lengths of all cell cycle phases in 4D- and control-transduced human HSPCs (Fig. 2, A and B; and Fig. S2). In agreement with a role for the catalytic activity of the 4D complex in promoting cell cycle progression, the length of the G1 phase was shortened. However, the total length of the cell cycle remained very similar due to slight prolongation of G2/M phase. To confirm the unaltered cell cycle length, we determined the cell growth of 4D-transduced and control cells in culture and found that leukocytes and CD34+ HSPCs exhibited the same growth rate (Fig. 2 C). Moreover, no further effect on colony-initiating potential was observed in a serial replating assay (Fig. 2 D), suggesting that 4D overexpression provides no advantage for progenitor self-renewal in human HSCPs in vitro. Fully consistent with previous reports on 4D activity in other somatic stem cell types (Lange et al., 2009), we conclude that 4D levels determine the timespan that human HSPCs spend in G1 without changing the total cell cycle length.
4D overexpression shortens G1 cell cycle phase. (A) Phase length calculations in human HSPCs based on cumulative BrdU incorporation 4 d after transduction. Phase lengths were calculated by determining the cell cycle transition rates of equation systems 1 and 2 (see Materials and methods). (top) Simultaneously, the equations of system 1 (lines) were fitted to time-course data of cells gated on BrdU and DNA content (dots), showing (top left) BrdU-positive, (top middle) G1 BrdU-negative, and (top right) G2/M BrdU-negative fractions, as well as (bottom) equations of system 2 (lines) to the same time-course data gated on DNA content only (dots), showing the steady-state distribution of (bottom left) S-phase, (bottom middle) G1-phase, and (bottom right) G2–M-phase fractions. Shaded regions indicated one standard deviation of the estimated measurement noise. (B) Plots show cell cycle phase lengths in human HSPCs after transduction with 4D calculated by the fitted values described in A. 95% confidence intervals are shown. Significant differences (**) between cell cycle parameters were evaluated at a confidence level of 99% by profile likelihood. Data shown is summarized from two independent experiments and two pooled CB samples were used each time. (C) GFP+ CD45+ (left) and GFP+ CD45+ CD34+ (right) cell numbers after cultivation of 4D- or mock-transduced HSPCs in medium supplemented with hSCF, hTPO, and mFlt3l. Before each plating, viability was measured and found comparable between 4D- and mock-transduced cells (mock: assay start day 4, >98 ± 0.8%; day 8, 95 ± 1.1%; day 13, 94 ± 0.5%; day 16, 97 ± 1.5%; 4D: assay start day 4, 99 ± 0.5%; day 8, 95 ± 1%; day 13, 95 ± 0.8%; day 16, 88 ± 7%). Plot shown is representative of two independent experiments using two or three pooled CB samples each. (D) Serial replating assay performed 3–5 d after transduction (left). Colonies were counted after 14 d and 5 × 104 CD45+ cells were replated. Colony types were determined after the first plating round (right). Each data point represents the mean of duplicates or triplicates (n = 7–8) and colony numbers were calculated to an input of 50,000 cells per well. Viability was comparable between mock- and 4D-transduced cells at each time point of analysis. Data are pooled from 7 independent experiments using two or three pooled CB samples. **, P = 0.01–0.001.
4D overexpression shortens G1 cell cycle phase. (A) Phase length calculations in human HSPCs based on cumulative BrdU incorporation 4 d after transduction. Phase lengths were calculated by determining the cell cycle transition rates of equation systems 1 and 2 (see Materials and methods). (top) Simultaneously, the equations of system 1 (lines) were fitted to time-course data of cells gated on BrdU and DNA content (dots), showing (top left) BrdU-positive, (top middle) G1 BrdU-negative, and (top right) G2/M BrdU-negative fractions, as well as (bottom) equations of system 2 (lines) to the same time-course data gated on DNA content only (dots), showing the steady-state distribution of (bottom left) S-phase, (bottom middle) G1-phase, and (bottom right) G2–M-phase fractions. Shaded regions indicated one standard deviation of the estimated measurement noise. (B) Plots show cell cycle phase lengths in human HSPCs after transduction with 4D calculated by the fitted values described in A. 95% confidence intervals are shown. Significant differences (**) between cell cycle parameters were evaluated at a confidence level of 99% by profile likelihood. Data shown is summarized from two independent experiments and two pooled CB samples were used each time. (C) GFP+ CD45+ (left) and GFP+ CD45+ CD34+ (right) cell numbers after cultivation of 4D- or mock-transduced HSPCs in medium supplemented with hSCF, hTPO, and mFlt3l. Before each plating, viability was measured and found comparable between 4D- and mock-transduced cells (mock: assay start day 4, >98 ± 0.8%; day 8, 95 ± 1.1%; day 13, 94 ± 0.5%; day 16, 97 ± 1.5%; 4D: assay start day 4, 99 ± 0.5%; day 8, 95 ± 1%; day 13, 95 ± 0.8%; day 16, 88 ± 7%). Plot shown is representative of two independent experiments using two or three pooled CB samples each. (D) Serial replating assay performed 3–5 d after transduction (left). Colonies were counted after 14 d and 5 × 104 CD45+ cells were replated. Colony types were determined after the first plating round (right). Each data point represents the mean of duplicates or triplicates (n = 7–8) and colony numbers were calculated to an input of 50,000 cells per well. Viability was comparable between mock- and 4D-transduced cells at each time point of analysis. Data are pooled from 7 independent experiments using two or three pooled CB samples. **, P = 0.01–0.001.
Enhanced 4D levels increase human leukocyte engraftment in vivo
To test whether 4D-mediated cell cycle progression affects HSPC maintenance and function in vivo, transduced human CD34-enriched CB cells were transplanted into optimized recipient mice (NSG KitWv/+; Cosgun et al., 2014). To normalize potential effects of 4D or mock overexpression on human cells, we transplanted a defined mixture of transduced GFP+ and nontransduced GFP− cells from the same CB preparation. Frequencies and absolute numbers of GFP+ cells within CD34+ CB donor cells were considered input and used to normalize the output activity of 4D or mock-transduced HSPCs. After transplantation, blood chimerism steadily increased over time (Fig. 3 A), and the frequency of GFP+ cells among human leukocytes remained constant throughout the observation period (Fig. 3 B), suggesting great stability of construct expression. At the time point of analysis (22–28 wk after transplantation), equal numbers of human donor-derived cells were found in the BM of recipient mice. However, although GFP expression in controls remained constant compared with the transplanted donor CD34+ CB HSPCs, the frequencies of GFP+ cells in 4D-transduced leukocytes was increased by 2.3-fold (Fig. 3, C and D), suggesting that overexpression of 4D conferred an advantage for engraftment. To determine at which stage of differentiation 4D overexpression provides an advantage, we compared the frequencies of GFP+ cells among HSCs, HSPCs, and mature lymphoid and myeloid cells (Fig. 3, E and F). The number of GFP+ cells dramatically increased in cells of the myeloid lineage but remained comparable to that in mock controls in all other cell types (Fig. 3 F). Increased myeloid output was confirmed by a shift in the composition of mature GFP+ human cell lineages (Fig. S3) in the BM (Fig. 3 G) and spleen (Fig. 3 H) of recipient mice. The numbers of myeloid progenitor cell types were unaffected by 4D overexpression (Fig. 3 I), and, consistently, similar frequencies of growth factor-responsive progenitor cells were found in the BM cells of recipient mice (Fig. 3 J). Within the stem cell compartment, 4D- and mock-transduced cells showed a comparable contribution to HSCs when the fold-change relative to donor cell mixture was determined (Fig. 3 F), suggesting that the pool size of stem cells is not affected by different levels of 4D. In line with that finding was a comparable cell cycle profile between 4D- and mock-transduced HSPCs in the mouse environment (Fig. 3 K), despite the continuous high expression of the 4D transcript in donor-derived GFP+ CD34+ HSPCs (Fig. 3 L). We conclude that increased 4D levels result in increased donor-leukocyte engraftment based on elevated output of cells of the myeloid lineage without expanding the HSC pool.
4D-mediated G1 transit improves engraftment in vivo. (A) 2–4 × 105 4D- or mock-transduced HSPCs were transplanted into unconditioned NSG-KitWv/+ recipient mice (Cosgun et al., 2014) and blood of recipients was periodically analyzed for the frequency of human CD45+ leukocytes for up to 28 wk. Data are pooled from three independent experiments; mock, n = 11; 4D, n = 10. Two to three pooled CB samples were used for each experiment. (B) Plot shows GFP expression in human peripheral blood (PB) leukocytes in samples described in A. (C) Dot plots show GFP expression in 4D- or mock-transduced hCD34+ donor CB HSPCs (left) and in hCD45+ BM cells (right) 28 wk after transplantation. Data are representative for 3 independent experiments using at least 2 pooled CB samples each. (D) Bar graphs show GFP+ cells in CD34+ donor CB cells (input, left) and in hCD45+ BM cells in recipient mice 22–28 wk after transplantation (1° recipient; middle) as outlined in A. Fold-change of GFP contribution to both populations (frequency in 1° recipient relative to CD34+ cell input’, right). (E) Dot plots show GFP expression in CD34+ CB donor cells before (Input) and 28 wk after transplantation in HSCs (hCD45+ CD34+ CD38− CD90+ CD45RA−), HSPCs (hCD45+ CD34+), mature lymphoid (hCD45+ CD3+ CD19+), and myeloid (hCD45+ CD33+ and/or hCD45+ CD16+) cells in the BM of recipient mice. (F) Plot shows changes of GFP+ cells in the indicated populations relative to GFP+ cells in donor CD34+ CB HSPCs. Absolute cell numbers were used for normalization. Samples as described in C. (G and H) Composition of mature donor-derived cells in the BM (G) and spleen (H). GFP− and GFP+ fractions from mock- and 4D-transduced donor cells are shown. Samples as described in C. (I) Plot shows changes of GFP+ cells in the indicated BM populations relative to GFP+ cells in donor CD34+ CB HSPCs. MPPs, CD34+ CD38− CD90− CD45RA−; MLPs, CD34+ CD38− CD90-/lo CD45RA+; B/NK/GMPs, CD34+ CD38+ CD45RA+; and CMPs, CD34+ CD38+ CD45RA− (van Galen et al., 2014). Samples as described in C. (J) BFU-E (left), CFU-G + GM (middle), and CFU-M (right) colony formation of 4D- or mock-transduced sorted GFP+ hCD45+ BM leukocytes in the presence of hSCF, hGM-CSF, hG-CSF, hIL-3, and hEPO. Colonies were scored after 14 d. Nontransduced GFP− human donor cells were used as control (w/o). Data are summarized from three independent experiments. In two experiments (∆ and ◇), BM cells from recipients of the same donor cells were pooled. Samples as described in C. (K) Frequency of Ki67− cells in mock- or 4D-transduced donor-derived hCD34+ GFP+ HSPCs 22–28 wk after transplantation (left). Frequency of hCD34+ GFP+ HSPCs in S–G2–M in the same samples (right). Data are summarized from two independent experiments, mock n = 8, 4D n = 7. (L) Plot shows the expression of the 4D construct in donor-derived hCD34+ GFP+ HSPCs 22 wk after transplantation in comparison to hCD34+ GFP− HSPC competitors. Data are summarized from 3 recipient mice using 3 pooled CB samples. *, P = 0.05–0.01; ***, P < 0.001.
4D-mediated G1 transit improves engraftment in vivo. (A) 2–4 × 105 4D- or mock-transduced HSPCs were transplanted into unconditioned NSG-KitWv/+ recipient mice (Cosgun et al., 2014) and blood of recipients was periodically analyzed for the frequency of human CD45+ leukocytes for up to 28 wk. Data are pooled from three independent experiments; mock, n = 11; 4D, n = 10. Two to three pooled CB samples were used for each experiment. (B) Plot shows GFP expression in human peripheral blood (PB) leukocytes in samples described in A. (C) Dot plots show GFP expression in 4D- or mock-transduced hCD34+ donor CB HSPCs (left) and in hCD45+ BM cells (right) 28 wk after transplantation. Data are representative for 3 independent experiments using at least 2 pooled CB samples each. (D) Bar graphs show GFP+ cells in CD34+ donor CB cells (input, left) and in hCD45+ BM cells in recipient mice 22–28 wk after transplantation (1° recipient; middle) as outlined in A. Fold-change of GFP contribution to both populations (frequency in 1° recipient relative to CD34+ cell input’, right). (E) Dot plots show GFP expression in CD34+ CB donor cells before (Input) and 28 wk after transplantation in HSCs (hCD45+ CD34+ CD38− CD90+ CD45RA−), HSPCs (hCD45+ CD34+), mature lymphoid (hCD45+ CD3+ CD19+), and myeloid (hCD45+ CD33+ and/or hCD45+ CD16+) cells in the BM of recipient mice. (F) Plot shows changes of GFP+ cells in the indicated populations relative to GFP+ cells in donor CD34+ CB HSPCs. Absolute cell numbers were used for normalization. Samples as described in C. (G and H) Composition of mature donor-derived cells in the BM (G) and spleen (H). GFP− and GFP+ fractions from mock- and 4D-transduced donor cells are shown. Samples as described in C. (I) Plot shows changes of GFP+ cells in the indicated BM populations relative to GFP+ cells in donor CD34+ CB HSPCs. MPPs, CD34+ CD38− CD90− CD45RA−; MLPs, CD34+ CD38− CD90-/lo CD45RA+; B/NK/GMPs, CD34+ CD38+ CD45RA+; and CMPs, CD34+ CD38+ CD45RA− (van Galen et al., 2014). Samples as described in C. (J) BFU-E (left), CFU-G + GM (middle), and CFU-M (right) colony formation of 4D- or mock-transduced sorted GFP+ hCD45+ BM leukocytes in the presence of hSCF, hGM-CSF, hG-CSF, hIL-3, and hEPO. Colonies were scored after 14 d. Nontransduced GFP− human donor cells were used as control (w/o). Data are summarized from three independent experiments. In two experiments (∆ and ◇), BM cells from recipients of the same donor cells were pooled. Samples as described in C. (K) Frequency of Ki67− cells in mock- or 4D-transduced donor-derived hCD34+ GFP+ HSPCs 22–28 wk after transplantation (left). Frequency of hCD34+ GFP+ HSPCs in S–G2–M in the same samples (right). Data are summarized from two independent experiments, mock n = 8, 4D n = 7. (L) Plot shows the expression of the 4D construct in donor-derived hCD34+ GFP+ HSPCs 22 wk after transplantation in comparison to hCD34+ GFP− HSPC competitors. Data are summarized from 3 recipient mice using 3 pooled CB samples. *, P = 0.05–0.01; ***, P < 0.001.
Elevated 4D levels confer a competitive advantage for engraftment in secondary recipient mice
To determine the effects of 4D overexpression on HSC function, we transplanted enriched human leukocytes from the BM of primary recipient mice into secondary recipients. Human leukocytes were engrafted in all secondary recipients of 4D-transduced cells, but only 3 out of 6 control recipients (Fig. 4 A), suggesting an advantage for 4D-transduced cells in HSC maintenance. Furthermore, the frequency of GFP+ cells increased in 4D-transduced leukocytes but decreased in mock-transduced cells, suggesting a competitive advantage for the engraftment in secondary recipient mice when 4D levels are increased (Fig. 4 B). The fold-change increase of GFP+ cells becomes highly significant compared with the CD34+ CB transplantation mixture and shows that the contribution of 4D-transduced cells increases by fourfold over serial transplantation rounds (Fig. 4 B, right). Only positively scored secondary recipients (hCD45 chimerism >1% in the BM; Fig. 4 A) showed engraftment of donor CD34+ HSPCs (Fig. 4 C). The donor human HSC pool size was found unaltered between secondary recipients of 4D and mock overexpressing cells (Fig. 4 D). In secondary recipient mice receiving 4D-transduced cells, there was a bias toward engraftment with cells of the myeloid lineage in the BM (Fig. 4 E) and spleen (Fig. 4 F). We conclude that increased 4D levels provide a competitive advantage for HSCs by promoting their maintenance in vivo.
4D provides a competitive advantage for HSPC engraftment. (A) Plot shows the frequency of successful engrafted secondary recipient mice that had received 4–6 × 106 hCD45+ enriched BM cells from primary recipient mice 23–24 wk earlier. Frequencies of hCD45+ cells <1% were considered as nonengrafted and excluded from subsequent analysis (dashed line). Data are summarized from two independent experiments using 2 or 3 donor and 3 recipient mice each. (B) GFP expression in hCD45+ BM leukocytes in primary (1°) and secondary (2°) recipient mice (dot plots). Bar graphs show the fold-change of GFP frequencies in hCD45+ BM cells relative to hCD45+ BM cells in the primary recipients (left) and to CD34+ CB donor HSPCs (right). Samples as described in A. (C) Dot plots show frequencies of CD34+ cells within human lineage negative leukocytes in the BM of secondary recipient mice. Data from one representative experiment are shown. (D) Plot shows changes of GFP+ cells in the indicated populations relative to GFP+ cells in donor hCD45+ CD34+ BM cells from primary recipients. Absolute cell numbers were used for normalization. Samples as described in A. (E) Composition of mature donor-derived GFP+ and GFP− cells in the BM of secondary recipients. (F) Composition of mature donor-derived GFP+ and GFP− cells in the spleen of secondary recipients. (G) Plot shows fold-change difference of human CD34+ HSPCs 6 d after cultivation with LPS relative to samples without LPS. Data are pooled from 5 experiments using 2–5 pooled and 1 experiment using 3 individual CB samples. Statistic was calculated based on mean values of each separate CB preparation. Different dot shapes indicate technical replicates from independent experiments. (H) Plot shows the expression of 4D transcript relative to mock-transduced cells before and after cultivation with LPS determined by qPCR. Data are pooled from two independent experiments using two pooled CB samples (triangles) and 3 individual CB samples (circles). (I and J) Plots show maintenance of human CD34+ HSPCs after 8 d of cultivation with hGM-CSF (I) or hIL-3 (J). Fold change in CD34+ GFP+ cells relative to mock control is shown. Data are pooled from three independent experiments using 5 (IL-3) and 7 (GM-CSF) single CB samples.
4D provides a competitive advantage for HSPC engraftment. (A) Plot shows the frequency of successful engrafted secondary recipient mice that had received 4–6 × 106 hCD45+ enriched BM cells from primary recipient mice 23–24 wk earlier. Frequencies of hCD45+ cells <1% were considered as nonengrafted and excluded from subsequent analysis (dashed line). Data are summarized from two independent experiments using 2 or 3 donor and 3 recipient mice each. (B) GFP expression in hCD45+ BM leukocytes in primary (1°) and secondary (2°) recipient mice (dot plots). Bar graphs show the fold-change of GFP frequencies in hCD45+ BM cells relative to hCD45+ BM cells in the primary recipients (left) and to CD34+ CB donor HSPCs (right). Samples as described in A. (C) Dot plots show frequencies of CD34+ cells within human lineage negative leukocytes in the BM of secondary recipient mice. Data from one representative experiment are shown. (D) Plot shows changes of GFP+ cells in the indicated populations relative to GFP+ cells in donor hCD45+ CD34+ BM cells from primary recipients. Absolute cell numbers were used for normalization. Samples as described in A. (E) Composition of mature donor-derived GFP+ and GFP− cells in the BM of secondary recipients. (F) Composition of mature donor-derived GFP+ and GFP− cells in the spleen of secondary recipients. (G) Plot shows fold-change difference of human CD34+ HSPCs 6 d after cultivation with LPS relative to samples without LPS. Data are pooled from 5 experiments using 2–5 pooled and 1 experiment using 3 individual CB samples. Statistic was calculated based on mean values of each separate CB preparation. Different dot shapes indicate technical replicates from independent experiments. (H) Plot shows the expression of 4D transcript relative to mock-transduced cells before and after cultivation with LPS determined by qPCR. Data are pooled from two independent experiments using two pooled CB samples (triangles) and 3 individual CB samples (circles). (I and J) Plots show maintenance of human CD34+ HSPCs after 8 d of cultivation with hGM-CSF (I) or hIL-3 (J). Fold change in CD34+ GFP+ cells relative to mock control is shown. Data are pooled from three independent experiments using 5 (IL-3) and 7 (GM-CSF) single CB samples.
Murine HSPCs respond to a challenge by LPSs by differentiating, which results in their depletion (Nagai et al., 2006). To test whether the 4D-mediated change in cell cycle progression supports HSPC maintenance by protecting them from stimulation-induced differentiation, we cultivated 4D-, CDK4-, CCND1-, and mock-transduced human HSPCs with LPS. A high frequency of 4D-transduced lineage-negative cells retained a HSC phenotype compared with controls (Fig. 4 G), which is consistent with a protective function upon overexpression of the G0/G1 phase regulators. In contrast, overexpression of CDK4 or CCND1 alone resulted in no rescue, suggesting that binding of CCND1 to endogenous CDK4 or CDK6 is not sufficient to protect HSPCs from differentiation-inducing signals. Continuous overexpression of 4D transcripts was confirmed before and after cultivation with the mitogenic stimulus LPS (Fig. 4 H). Expression levels were found to be in a similar range before and after this functional assay and comparable to the ones detected in donor-derived GFP+ CD34+ HSPCs isolated from the BM of primary recipient mice 22 wk after transplantation (Fig. 3 L), suggesting continuous overexpression of the transcript to similar levels. To test for an interconnection between cytokines and cell cycle dynamics we cultured 4D or mock transduced cells with human GM-CSF or IL-3 and also found higher numbers of remaining phenotypic HSPCs in 4D-transduced samples (Fig. 4, I and J). Collectively, our data suggest that 4D overexpression protects HSPCs from differentiation-inducing signals, resulting in an improved maintenance in vitro.
Increased levels of 2E confer a competitive disadvantage to human HSPCs
To test whether the second CCN–CDK complex important for G1-to-S transition also regulates human HSPC function, we overexpressed 2E in CD34-enriched CB cells (Fig. 5 A) and observed more severe perturbations of cell cycle kinetics (Fig. 5 B). First, a large proportion of cells contained fragmented DNA and underwent apoptosis after transduction (Fig. 5, B and C), suggesting that overexpression of 2E is not compatible with cellular survival. The absence of such an effect after 4D overexpression indicates the specificity of 2E (Fig. 5 B). Second, a large fraction of viable 2E-transduced cells exited G0 and shifted to the S–G2–M phases of the cell cycle (Fig. 5, B and D). This shift was induced by increased frequencies of pRb and Rb hyperphosphorylation in the majority of transduced HSPCs (Fig. 5 D). Consequently, G1 phase of the cell cycle was significantly shortened and here, in contrast to 4D, the time HSPCs spend in one cycle was also reduced (Fig. 5 E). After competitive transplantation performed in analogy to 4D samples, human donor cell engraftment in the BM was comparable between recipients of 2E- or mock-transduced HSPCs (Fig. 5 F). However, the proportion of 2E-transduced GFP+ cells therein was severely reduced compared with controls (Fig. 5 G). Nevertheless, GFP+ CD34+ HSPCs were still detectable at very low frequencies (Fig. 5 H), suggesting that elevated 2E expression is compatible with cellular survival but that 2E overexpression, contrary to 4D overexpression, provides a competitive functional disadvantage in vivo. Transplantation into secondary recipient mice further aggravated the phenotype, resulting in an almost complete loss of GFP+ 2E-transduced cells (Fig. 5 I). Collectively, our data suggest that increased 2E expression results in a more pronounced perturbation of quiescence leading to stem cell exhaustion, an outcome opposite to 4D-mediated cell cycle progression.
Elevated levels of CCNE1–CDK2 (2E) confer a competitive disadvantage to human HSC function in vivo. (A) Plot shows expression levels of the 2E construct relative to mock controls in GFP+ CD34+ CB samples 3 d after transduction. Data are summarized from two independent experiments using three single CB samples each. (B) Cell cycle profile 3 d after overexpression of mock, 4D, or 2E in human HSPCs gated on GFP+ CD34+ cells. Data are representative for 15 independent experiments. Single (12 expts) or 2–3 pooled (3 expts) CB samples were used. Data are quantified in (C and D). (C) Frequencies of GFP+ CD34+ HSPCs containing fragmented DNA (sub-G1, left) and apoptotic cells (right). (D) Frequencies (left) and MFI (second left) of phosphorylated pRb at S807/811 in mock- and 4D-tranduced HSPCs. Frequencies of GFP+ CD34+ HSPCs in S–G2–M phase (second right), and of Ki-67 negative cells (right). Experiments were done 3–5 d after transduction, and data are pooled from 3 (pRb), 2 (Ki67), and 15 (S–G2–M) independent experiments. Two to three pooled CB samples were used for each experiment. (E) Plots show cell cycle phase lengths in human HSPCs 3 d after overexpression of 2E, as outlined in Fig. 2 B. Data were pooled from two independent experiments and single CB samples were used each time. (F) Donor leukocyte numbers in the BM of NSG-KitWv/+ recipient mice 22–28 wk after transplantation of 2–4 × 105 HSPCs containing 2E- or mock-transduced cells. Data are summarized from two independent experiments using two pooled CB samples for the transplantation into three recipient mice each. (G) Dot plots show 2E-transduced (GFP+) human HSPCs in CD34+ cells of the transplantation mix before (Input) and in human CD45+ BM cells 28 wk after transplantation (1° recipient). Plots are representative for samples as described in F. Bar graphs summarize GFP frequencies in donor CD34+ HSPCs (left) and engrafted hCD45+ BM cells in primary recipients (middle). n = 6. The fold-change of GFP expression relative to the GFP frequencies in CD34+ CB donor cells is shown (right). n = 6. (H) Absolute numbers of GFP− and GFP+ engrafted hCD45+ CD34+ BM HSCPs in the BM of recipient mice. n = 6. (I) Bar graphs show the fold-change in GFP expression in human HSPCs in secondary recipient mice relative to that in hCD45+ BM cells in primary recipients (left) and relative to that of CD34+ donor CB HSPCs (middle) after transplantation of 4–6 × 106 hCD45+ enriched BM cells from primary recipient mice into secondary NSG-KitWv/+ recipients. Cell numbers of human GFP+ and GFP− CD34+ HSPCs in the BM of secondary recipient mice are shown (right). Data are pooled from two independent experiments with three mice each.
Elevated levels of CCNE1–CDK2 (2E) confer a competitive disadvantage to human HSC function in vivo. (A) Plot shows expression levels of the 2E construct relative to mock controls in GFP+ CD34+ CB samples 3 d after transduction. Data are summarized from two independent experiments using three single CB samples each. (B) Cell cycle profile 3 d after overexpression of mock, 4D, or 2E in human HSPCs gated on GFP+ CD34+ cells. Data are representative for 15 independent experiments. Single (12 expts) or 2–3 pooled (3 expts) CB samples were used. Data are quantified in (C and D). (C) Frequencies of GFP+ CD34+ HSPCs containing fragmented DNA (sub-G1, left) and apoptotic cells (right). (D) Frequencies (left) and MFI (second left) of phosphorylated pRb at S807/811 in mock- and 4D-tranduced HSPCs. Frequencies of GFP+ CD34+ HSPCs in S–G2–M phase (second right), and of Ki-67 negative cells (right). Experiments were done 3–5 d after transduction, and data are pooled from 3 (pRb), 2 (Ki67), and 15 (S–G2–M) independent experiments. Two to three pooled CB samples were used for each experiment. (E) Plots show cell cycle phase lengths in human HSPCs 3 d after overexpression of 2E, as outlined in Fig. 2 B. Data were pooled from two independent experiments and single CB samples were used each time. (F) Donor leukocyte numbers in the BM of NSG-KitWv/+ recipient mice 22–28 wk after transplantation of 2–4 × 105 HSPCs containing 2E- or mock-transduced cells. Data are summarized from two independent experiments using two pooled CB samples for the transplantation into three recipient mice each. (G) Dot plots show 2E-transduced (GFP+) human HSPCs in CD34+ cells of the transplantation mix before (Input) and in human CD45+ BM cells 28 wk after transplantation (1° recipient). Plots are representative for samples as described in F. Bar graphs summarize GFP frequencies in donor CD34+ HSPCs (left) and engrafted hCD45+ BM cells in primary recipients (middle). n = 6. The fold-change of GFP expression relative to the GFP frequencies in CD34+ CB donor cells is shown (right). n = 6. (H) Absolute numbers of GFP− and GFP+ engrafted hCD45+ CD34+ BM HSCPs in the BM of recipient mice. n = 6. (I) Bar graphs show the fold-change in GFP expression in human HSPCs in secondary recipient mice relative to that in hCD45+ BM cells in primary recipients (left) and relative to that of CD34+ donor CB HSPCs (middle) after transplantation of 4–6 × 106 hCD45+ enriched BM cells from primary recipient mice into secondary NSG-KitWv/+ recipients. Cell numbers of human GFP+ and GFP− CD34+ HSPCs in the BM of secondary recipient mice are shown (right). Data are pooled from two independent experiments with three mice each.
DISCUSSION
Our results showed that overexpression of cell cycle regulators important for the shift from G0-to-G1 and transition through G1 cell cycle phase results in increased cell cycle entry and accelerated progression through G1. Unexpectedly, increased levels of 4D and 2E complexes have opposite effects on HSC function in vivo. 4D confers a competitive advantage for engraftment in primary and secondary recipient mice, whereas ectopic expression of 2E results in poor repopulation and disappearance of transduced cells over time, likely due to the loss of the quiescent population of HSPCs acting as a reservoir for long-term reconstitution. Our data suggest that the progression kinetics through distinct cell cycle phases represent a novel regulatory platform for the function of human HSCs.
Overexpression of 4D is well tolerated by human HSPCs and results in increased phosphorylation of Rb at S807/811, which is required for cells to exit the G0 phase (Ren and Rollins, 2004), and consistently, human HSPCs exit G0 to enter G1. Cumulative BrdU labeling showed that the G1 phase was shortened upon 4D overexpression, whereas the G2–M phases were lengthened, resulting in an unaltered total cell cycle length of HSPCs. Consequently, there were no differences in cell growth in culture or with serial replating. In contrast, overexpression of 2E affects the cell cycle kinetics more profoundly, resulting in cell death, but a higher frequency of the remaining viable cells enter the cell cycle. Interestingly, the in vivo consequences of both manipulations were exactly opposite. 4D expression provided a competitive advantage for HSCs by maintaining their functionality, resulting in increased engraftment in primary and secondary recipient mice. 4D-mediated cell cycle progression may exert its effects by protecting HSPCs from the accumulation of differentiation-inducing signals as a consequence of the shortened G1 phase length. Cells in early G1 were suggested to be sensitive to extrinsic signals, and consistently, human HSPCs became partially resistant to LPS-induced differentiation after 4D overexpression in vitro. In contrast, increased 2E levels conferred a functional disadvantage, resulting in the loss of GFP+ HSCs, perhaps by exhaustion of the stem cell pool. These findings suggest that early and late G1 cell cycle phases may serve very different purposes. In line with this interpretation is the finding that targeted deletion of early G1 phase CKIs p18INK4c (Yuan et al., 2004) and p16INK4a (Janzen et al., 2006) result in improved functionality of murine HSCs. In contrast, loss of other CKIs, including other members of the INK4 family and the entire CIP/KIP family that affect late G1 phase progression, results in different degrees of impaired functionality in murine HSCs (Cheng et al., 2000a,b; van Os et al., 2007; Matsumoto et al., 2011; Zou et al., 2011; Hilpert et al., 2014). Furthermore, a lack of Rb family genes Rb, p107, and p130 in murine HSCs results in constitutive expression of E2F target genes, conveying cell cycle entry and subsequent exhaustion of HSCs (Viatour et al., 2008). This may be mediated by accelerated progression through late G1 phase, comparable to the effects seen after 2E overexpression. Collectively, we suggest that a shortened transition time through the mitogen-dependent early G1 phase by the overexpression of 4D provides an advantage for the maintenance of human HSC function, whereas shortened transit time through the mitogen-independent late G1 phase mediated by 2E overexpression is disadvantageous for HSC maintenance. In addition, loss of the potential to enter quiescence seems to further contribute to the loss of a reservoir pool of long-term HSPCs resulting in their depletion upon 2E overexpression.
Adult HSCs mainly rest in G0 and are thought to preserve their functionality by preventing the accumulation of potential threats that may promote the development of cancer (Trumpp et al., 2010). In agreement with previous studies (Yuan et al., 2004; Janzen et al., 2006), our data suggest that improved function of human HSPCs is not inevitably connected to quiescence, but rather that the timing of cell cycle progression is an important modulator of HSC fitness. This interpretation is consistent with several works, indicating that the undifferentiated state of stem cells is favored by a short G1 phase; the phase during which cell fate decisions are made (Massagué, 2004; Blomen and Boonstra, 2007; Orford and Scadden, 2008; Singh and Dalton, 2009; Pietras et al., 2011). However, our data may also be compatible with a potential role for alterations in G2–M transition affecting HSC functionality. The cell cycle length hypothesis was originally investigated in neural stem cells (NSCs) during development (Calegari and Huttner, 2003), but it was soon speculated that a similar mechanism may also hold true in embryonic pluripotent cells (Singh and Dalton, 2009) and adult HSCs (Orford and Scadden, 2008). Although validation of this model has been achieved in the developing (Lange et al., 2009) and adult central nervous system (Artegiani et al., 2011), to the best of our knowledge, our data are the first to validate and significantly extend the role of the G0-to-G1 transition and G1 length beyond murine NSCs to, specifically, human HSCs.
CDK4 and 6 are structurally very similar and functionally redundant cell cycle components important for G1 transition in HSCs (Pietras et al., 2011; Nakamura-Ishizu et al., 2014). Most recently a functional delimitation of CDK6 from CDK4 was proposed for murine (Scheicher et al., 2015) and human (Laurenti et al., 2015) HSCs, and CDK6 was assigned an important role during the exit of quiescence. Overexpression of CDK6 induced extensive G0-to-G1 transition resulting in expansion of human HSPCs upon transplantation and, consequently, improved repopulation of secondary recipient mice (Laurenti et al., 2015). In contrast, we show that the overexpression of CCND1 and CDK4 results in a minor G0-to-G1 transition but highly significantly affects other cell cycle parameters including Rb signaling and shortening of the G1 phase resulting in a shift of the cell cycle profile. As a consequence we find improved HSC function in the absence of expansion. Lack of alterations in the expression of other cell cycle regulators by our manipulation and the need for the coexpression of CCND1 and CDK4 to see a functional effect, strongly suggests that shortened G1 phase rather than increased G0-to-G1 transition is the underlying cause for improved HSC maintenance seen in our study and further underlines functional secession between CDK4 and CDK6 in human HSCs.
Collectively, our results provide evidence that the kinetics of early and late G1 phases of the cell cycle serve different purposes and result in a competitive advantage or disadvantage for adult human HSC function in vivo. In future studies, it will be interesting to determine whether the regulatory role of the transition time through early and late G1 is a feature that is conserved in other adult and embryonic stem cell types.
MATERIALS AND METHODS
Mice
C57BL/6 (B6) mice were purchased from The Jackson Laboratory. NSG KitWv/+ mice were generated as previously described (Cosgun et al., 2014). All mice were bred and maintained under specific pathogen–free conditions in the animal facility of the TU Dresden. Animal experiments were performed in accordance with German animal welfare legislation and were approved by the relevant authorities, the Landesdirektion Dresden, Referat 24.
Constructs and viral preparations
To generate transfer vectors for the overexpression of human CCN–CDK complexes, codon-optimized (GeneOptimizer; Life Technologies) cDNA sequences of CDK4 and CCND1 or CDK2 and CCNE1 were linked by a T2A sequence. A Kozak consensus sequence and an additional stop codon were introduced at the 5′ or 3′ end, respectively, and genes were synthesized by Life Technologies GeneArt Synthesis with flanking EcoRI and BamHI restriction sites that were used for ligation into the HIV-1–based transfer vector puc2CL6IEGwo. This lentiviral transfer vector is based on puc2CL6 (Nakano et al., 2012) and expresses a spleen focus–forming virus, U3 promoter–driven bicistronic transcript harboring a 5′ transgenic sequence and a 3′ EMCV-IRES–controlled EGFP ORF, followed by an optimized Woodchuck hepatitis virus posttranscriptional regulatory element (WPRO). Transfer vectors for single CDK4 or CCND1 expression were generated by TA cloning of the CCND1 or CDK4 codon-optimized cDNA sequence into pCR™2.1 (Invitrogen; primers for PCR amplification are as follows: CCND1 forward, 5′-ATGGAACATCAGCTGCTGT-3′, and reverse, 5′-TCAGATGTCCACATCCCGC-3′; CDK4 forward, 5′-ATGGCTACAAGCAGATATGAG-3′, and reverse, 5′-TCACTCGGGGTTGCCCTC-3′) and subsequently cloned into puc2CL6IEG using EcoRI restriction sites. puc2CL6IEGwo was used as mock control. Lentiviral PFV Env pseudotypes were generated by cotransfection of 293T cells with the respective transfer vector together with the HIV-1 Gag-Pol expressing packaging vector pCD/NL-BH (Mochizuki et al., 1998) and the mutant PFV Env packaging vector pcoPE01 (Hamann et al., 2014). Lentiviral supernatant was harvested 48 h after transfection and stored at –80°C.
Gene transfer into HSPCs and cell lines
Human umbilical CB samples were provided by the DKMS Cord Blood Bank Dresden and were used in accordance with the guidelines approved by the Ethics Committee of the Dresden University of Technology. CB probes were prepared as described previously (Cosgun et al., 2014). CD34-enriched HSPCs were prestimulated overnight in medium supplemented with 1% penicillin/streptomycin (P/S), 10% FCS, 50 ng/ml recombinant human thrombopoietin (rhTPO), 50 ng/ml recombinant human stem cell factor (rhSCF; both from PeproTech), and 100 ng/ml recombinant human angiopoietin-like protein 5 (rhAngptl5; Miltenyi Biotec). The next day, lentiviral supernatant was concentrated to yield a multiplicity of infection (MOI) of 10–30 and added to a nontissue culture-treated 24-well plate (BD) that was previously coated with RetroNectin (8 µg/cm2; Takara Bio). After 1 h, cells were added and medium was changed 24 h after transduction. Transplantations and experiments were performed 2–5 d after transduction. For the transduction of NIH3T3 cells, the cells were cultured with viral supernatant overnight, and the medium was changed the next day. Before each experiment, the viability of all cells and CD34+ GFP+ cells was assessed by DAPI and Sytox negativity and was found to be >97% in all experiments.
Transplantation
For the initial transplantation, 2.5–3 × 105 transduced CB cells (HSPCs) were intravenously transplanted into NSG KitWv/+ adult recipient mice (4–12 wk of age) without prior conditioning. To have controls of the same genotype transduced GFP+ and nontransduced GFP− cells from the same donor samples were cotransplanted. GFP− control cells were treated identically. For secondary transplantations, BM of 2–3 pooled primary recipients was enriched for hCD45+ cells (Miltenyi Biotec), and 4–6 × 106 cells were transplanted intravenously into secondary NSG KitWv/+ mice. After transplantations, mice were given antibiotic-containing drinking water for 3 wk.
Flow cytometry and cell sorting
Cell suspensions were prepared, counted, and stained as previously described (Arndt et al., 2013; von Bonin et al., 2013; Cosgun et al., 2014). Human cells were stained using antibodies specific for the following antigens (clones in brackets): CD11c (3.9), CD16 (3G8), CD45 (HI30), CD117 (A3C6E2), CD133 (EMK08; BioLegend); CD3 (OKT3, HIT3A), CD10 (CB-CALLA, SN5c), CD11b (ICRF44), CD14 (61D3), CD15 (HI98), CD16 (CB16), CD19 (HIB19), CD33 (WM-53), CD38 (HIT2), CD45RA (HI100), CD235a (HIR2, eBioscience); and CD34 (581/CD34), CD38 (HIT2), CD90 (5E10; BD). Lineage-negative cells were identified by the lack of CD11b, CD235a, CD19, CD3, CD16, CD14, CD10, and CD15 expression. Mouse lineage-positive cells were stained using antibodies specific for the following antigens: CD3 (2C11), CD11b (M170), CD19 (eBio1D3), CD45 (30-F11), CD45R (RA3-6B2), Gr-1 (RB6-8C5), NK1.1 (PK136), and Ter119 (Ter-119). For cell cycle analysis, cell surface markers were stained and intranuclear staining was performed according to the manufacturer’s protocol (BD). The following antibodies were used: anti–human Ki-67 (20Raj1; eBioscience), anti–human/mouse Rb (pS807/pS811, J112-906; BD), and anti-BrdU (3D4’ BD). For DNA staining, cells were resuspended in PBS containing 10% dimethyl sulfoxide, 0.1% Tween 20, 10 µg/ml DAPI, and 5 µg/ml RNase A (Roche) or 7AAD-containing PBS (BD). Samples were acquired and sorted using an LSRII and AriaIII cytometer (BD), respectively, and the data were analyzed using FlowJo software (Tree Star).
In vitro assays
Proliferation assay.
4 d after transduction, 8 × 104 human CD34+ enriched cells were seeded into Iscove’s modified Dulbecco’s medium (IMDM) containing 10% FCS, hSCF, hTPO, and mFlt3l (100 ng/ml). Cells were counted and analyzed for cell surface marker expression every 4 d.
Colony assay.
At 3–5 d after transduction, 4,000–5,000 GFP+ CD34+ cells were sorted and plated into a 6-well plate with methylcellulose-containing medium (MethoCult GF H84434; StemCell Technologies). After 14 d, the number of colonies was analyzed, and 5 × 104 CD45+ cells were replated into a second methylcellulose assay. Colony types were determined after the first plating round. To determine the colony formation of 1° recipient mice, 2.5 × 104 – 1.25 × 105 hCD45+ GFP+ BM cells were sorted, and colony assays were performed. Duplicate or triplicate wells were set up for each condition.
LPS stimulation.
4–5 d after transduction, 5 × 104 cells were cultivated in medium supplemented with SCF, TPO, and hAngptl5 and then stimulated with LPS (50 µg/ml), hIL-3 (20 ng/ml), or hGM-CSF (20 ng/ml). After 6 d (LPS) or 8 d (IL-3, GM-CSF), the cells were analyzed by flow cytometry.
Apoptosis assay.
To assess for ongoing apoptosis, we performed Annexin V staining as outlined by the manufacturer (ImmunoTools). In brief, 2 × 104 cells were incubated with Annexin V APC for 30 min and cells were acquired on a flow cytometer.
Next generation sequencing
RNA of sorter-purified CD34+ GFP+ cells (3 d after transduction) was isolated using the RNeasy MicroKit (QIAGEN). Complete cDNA was synthesized from mRNA of mock- or 4D-transduced human CD34-enriched CB cells using the SMARTer Ultra HV kit (Takara Bio), which was followed by amplification of the purified cDNA with the Advantage 2 DNA Polymerase (12 cycles). After ultrasonic shearing of the amplified cDNA (Covaris S2), samples were subjected to standard Illumina fragment library preparation using the NEBNext chemistries (New England Biolabs) as previously described (Grinenko et al., 2014). Sequencing was performed on the Illumina NextSeq 500 platform, resulting in 75–90 million 75-bp single-end reads per sample. The two codon-optimized cDNA sequences of CDK4 and CCND1 were added as additional chromosomes to the human reference (hg38). Reads were aligned to this reference using the GSNAP alignment program (version 2014–12-17) and we used the gene annotation of Ensembl version 78 to detect splice sites. The uniquely aligned reads were counted with featureCounts (v1.4.6) and the same Ensembl annotation to which the codon-optimized sequences of CDK4 and CCND1 were added. We used DESeq2 (v1.6.3) and the counts table to test for differential gene expression between mock and 4D.
Expression analysis
qPCR.
cDNA synthesis from sorted CD34+ GFP+ HSPCs was performed after RNA isolation (RNeasy MicroKit; QIAGEN) using the SuperScript II Reverse transcription (Invitrogen). PCR reactions were performed on a Light cycler 480 (Roche) using the Quantifast SYBR Green kit (QIAGEN). Expression levels were normalized with the ΔΔCt method using the following primers: ACTB forward, 5′-AGCGAGCATCCCCCAAAGTT-3′ and reverse 5′-GGGCACGAAGGCTCATCATT-3′; 4D cDNA forward, 5′-CTACCTGCACAAGGACGAGG-3′, and reverse, 5′-ATGGTTTCCACCTCGCAACA-3′; 2E cDNA forward, 5′-ACTACGACCCCAACAAGCG-3′, and reverse, 5′-TTCTTTGGCGTCCCTCTCTC-3′.
Western blot.
Protein lysates (lysis buffer with 20 mM Hepes pH 7.4, 100 mM NaCl, 5 mM MgCl2, 1% TX-100, 1 mM phenylmethylsulfonyl fluoride [Sigma-Aldrich] and Protease Inhibitor Cocktail [Roche]) of sorter-purified transduced NIH-3T3 cells were separated on 12% sodium dodecyl sulfate-PAGE gels and transferred to Whatman Protran nitrocellulose membranes (Sigma-Aldrich). Immunoblots were probed with antibodies against β-tubulin (rabbit; Cell Signaling Technology; 1:1,000), CDK4 (rabbit mAb, D9G3E; Cell Signaling Technology; 1:1,000), or CCND1 (rabbit mAb, 17H3L3; Life Technologies; 1:500). Proteins of interest were detected with HRP-conjugated goat anti–rabbit IgG antibody (Jackson ImmunoResearch Laboratories; 1:10,000) and visualized with the Pierce ECL Western blotting substrate (Thermo Fisher Scientific) using the Amersham Imager 600 (GE Healthcare).
Statistical analysis
Two-tailed Student’s t tests were performed using Prism 5 for MacOSX software for all analyses (*, P = 0.05–0.01; **, P = 0.01–0.001; ***, P < 0.001). Unless indicated otherwise, standard deviations are depicted in all graphs.
Cell cycle phase length calculations
Cumulative BrdU incorporation.
4 d after transduction, 5–7 × 105 human cells were seeded into 96-well plates in IMDM supplemented with 10% FCS, 1% P/S, 50 ng/ml hSCF, 50 ng/ml hTPO, and 100 ng/ml mFlt3l (PeproTech). BrdU (Sigma-Aldrich) was resuspended in PBS and added at a final concentration of 10 µM for pulses in the time frame of 30 min to 28 h. Cells were fixed, and GFP+ CD34+ cells were sorted and followed via the detection of BrdU by flow cytometry.
Cell cycle model.
The lengths of individual cell cycle phases, G0–1, S, and G2–M, were calculated based on the observed cell cycle distributions measured as relative fractions with g1(t) + s(t) + g2(t) = 1. The transition of cells through the cell cycle was described by a system of ordinary differential equations that contain the transition rates ai for i = {g1, s, g2}, at which cells enter and exit the three cycle phases; these rates are related to the mean time cells reside in each phase, T = 1/ai. Because the cells divide at the end of G2/M, two new cells enter the next G1 phase. Therefore, the observed distribution is not directly proportional to the phase lengths. As derived previously (Toettcher et al., 2009), cell cycle progression is described by:
In the cumulative BrdU incorporation experiments, cells were gated by DNA content and BrdU staining, dividing the population into g1 BrdU-negative (g1b-), g2 BrdU-negative (g2b-), and BrdU-positive (brd) fractions and simplifying equation system 1 to:
Model parameters were estimated simultaneously for the two equation systems using the Data 2 Dynamics software package implemented in MATLAB (Raue et al., 2013). This implementation performs a maximum likelihood estimation using the trust-region algorithm and user-supplied derivatives. In addition, the variance of the measurement noise is estimated together with the model dynamics, giving improved optimization at small numbers of replicates. Parameter identifiability and confidence bounds were determined by profile likelihood estimation at a confidence level of 95% using the same software (Raue et al., 2009). The 95% confidence bounds of the total cell cycle length, Ttotal = Tg1 + Ts + Tg2, were computed by bootstrapping with 1,000–10,000 repeats. To determine differences between mock and C4, mock parameters ai,m were related to the C4 parameters by a scaling factor κi, ai,c4 = ai,m/κi. A significant difference in parameter values was observed if the subsequent profile likelihood estimates of κi excluded the value of one, rejecting ai,c4 = ai,m at a confidence level of 95% (*) and 99% (**). At each bootstrap repeat, Ttotal,m − Ttotal,c4 was calculated. Cell cycles were treated as significantly different if the resulting distribution excluded zero at the given confidence level.
Online supplemental material
Fig. S1 shows the alignments for codon-optimized constructs for the overexpression of CCND1 and CDK4. Fig. S2 shows the gating strategy used for the analysis of BrdU incorporation for the calculation of cell cycle phase lengths. Fig. S3 shows the gating strategy for the analysis of mature human blood cell populations used in this study.
Acknowledgments
The authors thank Christiane Ruedl and Chengcheng Zhang for providing the pMyc-IRES-GFP transfer vector and pcDNA3.1(-)-Flag-hAngptl2, respectively. Further thanks go to Susann Rahmig, Melanie Portz, Isabella Kutschick, Sindy Böhme, and Tania Araujo Ramos for expert technical assistance and help.
This work was supported by the German Research Foundation (DFG) through WA2837, SFB 655-B9, FOR2033-A03, and TRR127-A3 and by funding from the EU’s Seventh Programme for research, technological development, and demonstration under grant agreement No 261387 (CELL-PID) to C. Waskow and by the Federal Ministry for Education and Research (BMBF) grant no. 0316076 (MYC-NET/CancerSys) to T. Höfer.
The authors declare no competing financial interests.
Author contributions: N. Mende performed all experiments on human cells; M. Lesche and A. Dahl performed deep sequencing analysis and analyzed the data; K.D. Kokkaliaris, T. Grinenko, and F. Calegari advised the study; E.E. Kuchen and T. Höfer calculated the lengths of cell cycle phases and interpreted data; H. Hanenberg, D. Lindemann, and A. Platz provided crucial reagents; N. Mende and C. Waskow conceived the study, designed experiments, analyzed data, and wrote the manuscript.
References
Author notes
T. Grinenko’s present address is Dept. of Clinical Pathobiochemistry, TU Dresden, 01307 Dresden, Germany.