Immunoglobulin (Ig) class switch recombination (CSR) is initiated by the transcription-coupled recruitment of activation-induced cytidine deaminase (AID) to switch regions and by the subsequent generation of double-stranded DNA breaks (DSBs). These DNA breaks are ultimately resolved through the nonhomologous end joining (NHEJ) pathway. We show that during CSR, AID associates with subunits of cohesin, a complex previously implicated in sister chromatid cohesion, DNA repair, and the formation of DNA loops between enhancers and promoters. Furthermore, we implicate the cohesin complex in the mechanism of CSR by showing that cohesin is dynamically recruited to the Sμ-Cμ region of the IgH locus during CSR and that knockdown of cohesin or its regulatory subunits results in impaired CSR and increased usage of microhomology-based end joining.
During immune responses, B cells diversify their receptors through somatic hypermutation (SHM) and class switch recombination (CSR). SHM introduces mutations in Ig variable regions that modify the affinity of the receptor for its cognate antigen (Di Noia and Neuberger, 2007). CSR replaces the antibody isotype expressed (from IgM to IgG, IgE, or IgA), providing novel antibody effector functions (Chaudhuri et al., 2007). Mechanistically, SHM and CSR are initiated by activation-induced cytidine deaminase (AID), an enzyme which deaminates cytosines in both strands of transcribed DNA substrates (Petersen-Mahrt et al., 2002; Basu et al., 2011). AID-induced DNA deamination is then processed to trigger mutations in variable regions during SHM or to generate double-stranded DNA break (DSB) intermediates in switch (S) regions during CSR (Chaudhuri et al., 2007; Di Noia and Neuberger, 2007). These breaks activate the DNA damage response (Ramiro et al., 2007) and are resolved through classical and alternative nonhomologous end joining (NHEJ; Stavnezer et al., 2010).
CSR is a transcription-dependent, long-range recombination that occurs at the Ig heavy chain (IgH) locus and that involves the joining of two S regions, which may be separated by several hundreds of kilobase pairs. For CSR to succeed, donor and acceptor S regions must be brought into close proximity. This is believed to occur through three-dimensional conformational changes involving the generation of transcription-coupled DNA loops (Kenter et al., 2012). Nevertheless, the precise mechanisms controlling these conformational changes remain to be elucidated.
The cohesin complex has been described to play a prominent role in sister chromatid cohesion during cell division, in favoring DNA repair by homologous recombination (Nasmyth and Haering, 2009), in modulating gene expression (Dorsett, 2009), and in promoting the transcription-coupled formation of long-range DNA loop structures (Kagey et al., 2010). In addition, cohesin and the transcriptional insulator CTCF (Dorsett, 2009; Nasmyth and Haering, 2009) have been shown to control the RAG1/2-dependent rearrangement of antigen receptor genes during early B and T lymphocyte development by mechanisms involving the regulation of transcription and formation of long-range in cis DNA interactions (Degner et al., 2011; Guo et al., 2011; Seitan et al., 2011). Here, we have examined the role of cohesin in mature B cells undergoing CSR.
RESULTS AND DISCUSSION
Nuclear and chromatin-bound AID associate with cohesin
We have previously shown that nuclear AID exists in a large molecular weight complex containing proteins that are required for CSR (Jeevan-Raj et al., 2011). To further characterize this complex and investigate the functional role of novel AID partners in CSR, we have performed additional coimmunoprecipitation experiments coupled to identification by mass spectrometry. Nuclear and chromatin extracts prepared from CH12 cells expressing a full-length N-terminally tagged AID protein (AIDFlag-HA) or the epitope tags alone (Flag-HA) as negative controls were immunoprecipitated using an anti-Flag antibody. Eluted proteins were submitted for identification by mass spectrometry. Among the proteins identified, we found multiple AID partners previously implicated in CSR and/or SHM (Table S1). In addition, we found several proteins with no known function in CSR (Table S2), including subunits of the cohesin, condensin, Smc5/6 complex and Ino80 complexes. Given the described role for cohesin in mediating long-range recombination during B cell and T cell differentiation, we focused on the potential role of cohesin in CSR. The association between AID and the cohesin complex subunits (Smc1, Smc3, Nipbl, and Wapal) was confirmed by reciprocal coimmunoprecipitations and Western blotting in the nuclear (Fig. 1 A) and chromatin (Fig. 1 B) fractions and was specific, as they did not coprecipitate with an irrelevant tagged protein (EGFPFlag-HA; Fig. 1 C). Importantly, these interactions were not mediated by nonspecific nucleic acid binding, as extracts and immunoprecipitations were done in the presence of the benzonase nuclease. We conclude that endogenous subunits of the cohesin complex associate with a fraction of nuclear and chromatin-bound tagged AID through interactions that do not involve nonspecific nucleic acid binding.
Nuclear AID associates with cohesin subunits. Nuclear extracts (A and C) and chromatin fractions (B) prepared from CH12 cells expressing AIDFlag-HA (A and B) or EGFPFlag-HA (C) were immunoprecipitated and blotted with antibodies specific for Flag, AID, Smc1, Smc3, Wapal, and Nipbl. Note that the Nipbl antibody works only on immunoprecipitation. Input represents 1% of material used. Theoretical molecular masses in kilodaltons are indicated. Data are representative of three independent experiments.
Nuclear AID associates with cohesin subunits. Nuclear extracts (A and C) and chromatin fractions (B) prepared from CH12 cells expressing AIDFlag-HA (A and B) or EGFPFlag-HA (C) were immunoprecipitated and blotted with antibodies specific for Flag, AID, Smc1, Smc3, Wapal, and Nipbl. Note that the Nipbl antibody works only on immunoprecipitation. Input represents 1% of material used. Theoretical molecular masses in kilodaltons are indicated. Data are representative of three independent experiments.
Smc1 and Smc3 are dynamically recruited to the IgH locus during CSR
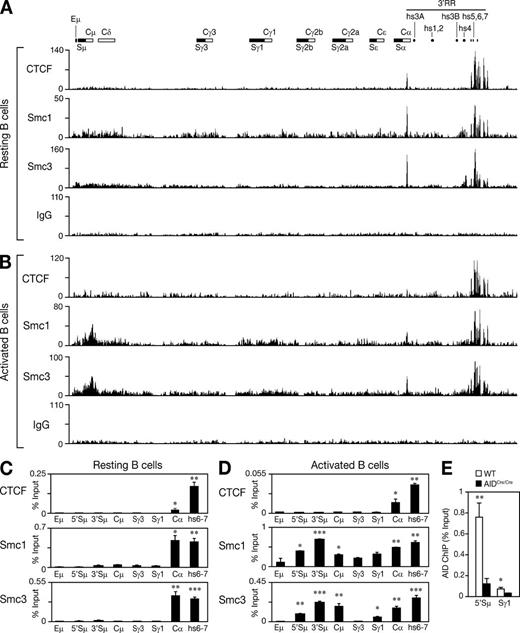
To determine whether cohesin is recruited to the IgH locus in B cells undergoing CSR, we performed ChIP-Seq experiments on chromatin prepared from resting or activated splenic B cells isolated from wild-type mice and using antibodies specific for Smc1, Smc3, and CTCF (Fig. 2). In resting B cells, we found that Smc1, Smc3, and CTCF are co-recruited to the 3′ regulatory region (3′RR; Fig. 2 A). This is consistent with published ChIP data on CTCF (Chatterjee et al., 2011) in mature B cells and ChIP-Seq results for CTCF and cohesin (Rad21) in Rag1-deficient pro–B cells (Degner et al., 2011). A sharp peak of CTCF, Smc1, and Smc3 binding was observed at Cα. This peak occurred over a region containing a predicted DNaseI hypersensitive site and a CTCF consensus motif (Nakahashi et al., 2013). No significant enrichment was observed at the Eμ enhancer Sμ or Sγ1 (Fig. 2 A). After stimulation, under conditions that induce CSR to IgG1, we found that Smc1 and Smc3 are significantly co-recruited, independently of CTCF, to a region spanning from the 5′ end of the donor switch region (Sμ) to the 3′ end of the Cμ constant region that did not comprise the Eμ enhancer (Fig. 2 B). Surprisingly, we failed to detect a reproducible recruitment of Smc1 or Smc3 over the Sγ1 switch region (Fig. 2 B), suggesting that Smc1 and Smc3 are not recruited to the acceptor switch region upon activation. It is possible, however, that our cell culture conditions (in which ∼15–20% of the cells switch to IgG1) are not robust enough to detect a specific enrichment. Consistent with this, we were unable to reproducibly detect a specific enrichment of AID at Sγ1 by ChIP-qPCR (Fig. 2 E).
Smc1 and Smc3 are dynamically recruited to the IgH locus during CSR. UCSC genome browser screenshots showing the ChIP-Seq binding profiles of CTCF, Smc1, Smc3, and IgG (negative control) at the IgH locus (chr12:114,438,857–114,669,149) in resting (A) and activated (B; with LPS + IL-4) B cells isolated from wild-type mice. A schematic map of the IgH locus indicates the switch regions (black boxes), the constant region exons (white boxes), the Eμ enhancer, and the DNaseI hypersensitive sites (hs) located in the 3′RR. Similar ChIP-Seq profiles were observed in an additional biological replicate experiment for Smc3 that was conducted in resting and activated B cells (not depicted). Chip-Seq results were verified by analytical-scale ChIP-qPCR experiments performed on chromatin prepared from 107 splenic resting (C) and activated (D) B cells. qPCR was performed at several locations across the IgH locus using primers listed in Table S4. Results are expressed as percent input and are representative of two independent biological replicate experiments. Mean of triplicate samples (+SD) is shown. Statistical significance versus Sγ3 (two-tailed Student’s t test) is indicated: *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001. Additional statistical analyses across the locus and between resting and activated B cells are shown in Table S5. (E) ChIP analysis for AID occupancy at the Sμ and Sγ1 switch regions in wild-type and AIDCre/Cre B cells cultured in vitro with LPS + IL-4 for 60 h. Results are expressed as percent input. Mean of triplicate samples (+SD) is shown. Statistical significance versus AIDCre/Cre was determined by a two-tailed Student’s t test. *, P ≤ 0.05; **, P ≤ 0.01. Results are representative of four independent experiments.
Smc1 and Smc3 are dynamically recruited to the IgH locus during CSR. UCSC genome browser screenshots showing the ChIP-Seq binding profiles of CTCF, Smc1, Smc3, and IgG (negative control) at the IgH locus (chr12:114,438,857–114,669,149) in resting (A) and activated (B; with LPS + IL-4) B cells isolated from wild-type mice. A schematic map of the IgH locus indicates the switch regions (black boxes), the constant region exons (white boxes), the Eμ enhancer, and the DNaseI hypersensitive sites (hs) located in the 3′RR. Similar ChIP-Seq profiles were observed in an additional biological replicate experiment for Smc3 that was conducted in resting and activated B cells (not depicted). Chip-Seq results were verified by analytical-scale ChIP-qPCR experiments performed on chromatin prepared from 107 splenic resting (C) and activated (D) B cells. qPCR was performed at several locations across the IgH locus using primers listed in Table S4. Results are expressed as percent input and are representative of two independent biological replicate experiments. Mean of triplicate samples (+SD) is shown. Statistical significance versus Sγ3 (two-tailed Student’s t test) is indicated: *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001. Additional statistical analyses across the locus and between resting and activated B cells are shown in Table S5. (E) ChIP analysis for AID occupancy at the Sμ and Sγ1 switch regions in wild-type and AIDCre/Cre B cells cultured in vitro with LPS + IL-4 for 60 h. Results are expressed as percent input. Mean of triplicate samples (+SD) is shown. Statistical significance versus AIDCre/Cre was determined by a two-tailed Student’s t test. *, P ≤ 0.05; **, P ≤ 0.01. Results are representative of four independent experiments.
The ChIP-Seq signal obtained in resting and activated B cells for Smc1 and Smc3 (Fig. 2, A and B) is consistent with the fact that they are known to exist as a heterodimer and was reproducible and specific, as we did not observe any significant enrichment at the IgH locus when using an IgG antibody as a negative control (Fig. 2, A and B). The recruitment of Smc1 and Smc3 at the IgH locus only partially correlated with that reported for AID (Yamane et al., 2011) and is consistent with the fact that only a fraction of chromatin-bound AID associates with the cohesin complex (Fig. 1 B). This suggests that cohesin is not a targeting factor for AID. The recruitment of Smc1, Smc3, and CTCF in resting and activated B cells observed by ChIP-Seq (Fig. 2, A and B) was confirmed by additional independent analytical-scale ChIP-qPCR experiments, using primer pairs at individual locations across the IgH locus (Fig. 2, C and D). We conclude that Smc1 and Smc3 are dynamically recruited, independently of CTCF, to the IgH locus (at the Sμ-Cμ region) during CSR. As Eμ is not bound by cohesin in resting B cells, the constitutive long-range interactions between Eμ and the 3′RR that take place in resting B cells (Wuerffel et al., 2007) are most likely cohesin-independent. Nevertheless, given the dynamic recruitment of Smc1 and Smc3 at Sμ-Cμ (and possibly Sγ1) in activated B cells, we speculate that cohesin may play a role in supporting the structural changes occurring at the IgH locus upon B cell activation.
Cohesin is required for efficient CSR
To determine the functional relevance of the cohesin complex in CSR, we undertook knockdown experiments in CH12 cells, a B cell line which can be induced to undergo CSR from IgM to IgA in vitro and which allows the study of the role of specific factors in CSR (Pavri et al., 2010; Willmann et al., 2012). CH12 cells were transduced with lentiviruses expressing a GFP reporter together with shRNAs specific for AID (as a positive control), the core subunits of the cohesin complex (Smc1 and Smc3), the cohesin loader/unloader subunits (Nipbl and Wapal), and a Non-Target shRNA as a negative control. Knockdown efficiencies were determined by Western blotting or by quantitative RT-PCR (qRT-PCR) on GFP+ sorted cells (Fig. 3 A). Transduced cells were stimulated for 48 h, and their ability to undergo CSR to IgA was determined by flow cytometry (Fig. 3, B and C). As expected, knockdown of AID resulted in a robust reduction in the efficiency of CSR relative to the Non-Target shRNA control (Fig. 3, B and C). Interestingly, we found that knockdown of Smc1, Smc3, Nipbl, and Wapal resulted in a significant reduction in the efficiency of CSR (18–41%) in GFP+ cells (Fig. 3, B and C). This reduction was more pronounced (30–52%) when the analysis was performed by gating on cells expressing high levels of GFP (Fig. 3 C). The effect on CSR after cohesin knockdown was not due to decreased survival (Topro-3 staining; unpublished data), strong defects in proliferation (CFSE dilution; unpublished data), significant activation of the DNA damage response and cell cycle checkpoints (Western blot for γ-H2AX and p-Chk1; unpublished data), or defective cell cycle progression (flow cytometry; unpublished data).
CSR is impaired by the knockdown of cohesin subunits. (A) CH12 cells were transduced with a lentivirus expressing a GFP reporter and shRNAs specific for AID, Smc1, Smc3, Nipbl, Wapal, or a Non-Target control. Transduced cells were stimulated for 48 h and sorted for GFP expression. Protein extracts and cDNAs were prepared and knockdown was determined by Western blotting or qPCR. Western blot for β-actin, Smc1, Smc3, and Wapal and qRT-PCR for Nipbl transcripts are shown. Expression was normalized to Cd79b and is presented relative to the Non-Target control, set as 1. Mean of triplicate samples (+SD) is shown. Statistical significance versus the Non-Target control (two-tailed Student’s t test): P = 0.0023. Data are representative of three experiments. (B) CH12 cells treated as in A were analyzed for surface IgA and GFP expression by flow cytometry. Representative plots from four to eight independent experiments are shown. (C) CH12 cells treated as in A were gated on cells expressing GFP (GFP+; white bars) or high levels of GFP (GFPHigh; black bars). The percentage (+SD) of CSR relative to the Non-Target shRNA control from four to eight independent experiments is shown. CSR in cells expressing the Non-Target shRNA control was set to 100%. The difference in CSR efficiency relative to the Non-Target control (Δ) is indicated below. Statistical significance versus the Non-Target control (two-tailed Student’s t test) is indicated: ***, P ≤ 0.001. (D and E) cDNA was prepared from CH12 cells treated as in A and qRT-PCR for μ (D) and α (E) germline transcripts was performed. Expression was normalized to HPRT mRNA abundance and is presented relative to the Non-Target control, set as 1 (black line). Mean of triplicate samples (+SD) is shown. Statistical significance versus the Non-Target control (two-tailed Student’s t test) is indicated: *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001. (F) Proteins extracts were prepared from CH12 cells treated as in A. Western blots for β-actin and AID are shown. Data are representative of three independent experiments. Theoretical molecular masses in kilodaltons are indicated.
CSR is impaired by the knockdown of cohesin subunits. (A) CH12 cells were transduced with a lentivirus expressing a GFP reporter and shRNAs specific for AID, Smc1, Smc3, Nipbl, Wapal, or a Non-Target control. Transduced cells were stimulated for 48 h and sorted for GFP expression. Protein extracts and cDNAs were prepared and knockdown was determined by Western blotting or qPCR. Western blot for β-actin, Smc1, Smc3, and Wapal and qRT-PCR for Nipbl transcripts are shown. Expression was normalized to Cd79b and is presented relative to the Non-Target control, set as 1. Mean of triplicate samples (+SD) is shown. Statistical significance versus the Non-Target control (two-tailed Student’s t test): P = 0.0023. Data are representative of three experiments. (B) CH12 cells treated as in A were analyzed for surface IgA and GFP expression by flow cytometry. Representative plots from four to eight independent experiments are shown. (C) CH12 cells treated as in A were gated on cells expressing GFP (GFP+; white bars) or high levels of GFP (GFPHigh; black bars). The percentage (+SD) of CSR relative to the Non-Target shRNA control from four to eight independent experiments is shown. CSR in cells expressing the Non-Target shRNA control was set to 100%. The difference in CSR efficiency relative to the Non-Target control (Δ) is indicated below. Statistical significance versus the Non-Target control (two-tailed Student’s t test) is indicated: ***, P ≤ 0.001. (D and E) cDNA was prepared from CH12 cells treated as in A and qRT-PCR for μ (D) and α (E) germline transcripts was performed. Expression was normalized to HPRT mRNA abundance and is presented relative to the Non-Target control, set as 1 (black line). Mean of triplicate samples (+SD) is shown. Statistical significance versus the Non-Target control (two-tailed Student’s t test) is indicated: *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001. (F) Proteins extracts were prepared from CH12 cells treated as in A. Western blots for β-actin and AID are shown. Data are representative of three independent experiments. Theoretical molecular masses in kilodaltons are indicated.
To determine whether switch region transcription is affected by the knockdown of cohesin subunits, we measured the level of donor (Sμ) and acceptor (Sα) switch region transcripts by qRT-PCR in activated CH12 cells. We found that the level of Sµ and Sα transcripts was increased after knockdown of AID and cohesin (relative to the Non-Target control), with the exception of Sα transcripts after knockdown of Wapal (Fig. 3, D and E), as expected from cells in which CSR is compromised and that continue to transcribe the switch regions. As no significant reduction in the level of these transcripts after Smc1, Smc3, and Nipbl knockdown was observed, we conclude that switch regions continue to be efficiently transcribed and that they are accessible for DNA deamination by AID. Therefore, cohesin appears not to be involved in the transcriptional regulation of switch regions during CSR. Importantly, we excluded a potential reduction in AID expression levels by Western blot (Fig. 3 F). We conclude that the cohesin complex is required for efficient CSR in CH12 cells. The role of cohesin in CSR appears to be independent of regulating switch region transcription and/or AID accessibility. Concerning a potential more global effect on transcription, we cannot exclude the possibility that the expression of additional genes required for CSR (other than AID) is affected by the knockdown of cohesin.
Knockdown of cohesin affects NHEJ
DSBs triggered by AID in switch regions during CSR are resolved through the NHEJ pathway, and the resulting switch junctions display small insertions and short stretches of microhomology (Stavnezer et al., 2010). In the absence of core NHEJ components, an increase in the usage of microhomology is observed concomitantly with a complete loss of direct joining (Yan et al., 2007). To determine whether cohesin knockdown affects the resolution of DSBs generated during CSR, we cloned and sequenced Sμ/Sα switch junctions from stimulated CH12 transduced with lentiviruses expressing shRNAs for Smc1, Smc3, Nipbl, Wapal, and a Non-Target negative control (Fig. 4) and sorted for GFP expression. Sequence analysis (Stavnezer et al., 2010) revealed that knockdown of cohesin subunits resulted in a significant increase in the usage of microhomology when compared with the Non-Target control (Fig. 4). Although the mean length of overlap (excluding insertions) was of 1.58 bp for the Non-Target control, it was increased to 3.22 bp for Smc1 (P = 0.0001), 2.60 bp for Smc3 (P = 0.0139), and 2.90 bp for Nipbl (P = 0.0066). The switch junctions obtained after Wapal knockdown displayed an overlap of 2.04 bp that was not statistically different from the Non-Target control (P = 0.6125). The increase in microhomology was due to sequences bearing >7 bp of microhomology at the junction and a reduction in those bearing short insertions (Fig. 4), similar to what has been described in human patients with deficiency in DNA ligase IV (Du et al., 2008), Artemis (Du et al., 2008), or ATM (Pan-Hammarström et al., 2006). In contrast to deficiency in core NHEJ components (Stavnezer et al., 2010), we did not find a reduction in the frequency of direct joining events (Fig. 4). We conclude that switch recombination junctions generated after Smc1, Smc3, and Nipbl knockdown (but not Wapal) are biased toward the usage of longer microhomologies. Given the role of Wapal in releasing cohesin from chromatin (Kueng et al., 2006), this suggests that cohesin is recruited but not released from the IgH locus and that NHEJ proceeds unaffected. Therefore, it appears that the loading of cohesin is sufficient to determine the outcome of DSB repair and that cohesin participates in the resolution of AID-induced DNA breaks.
Knockdown of cohesin affects NHEJ. (A) CH12 cells were transduced with a lentivirus expressing a GFP reporter and shRNAs specific for AID, Smc1, Smc3, Nipbl, Wapal, or a Non-Target control. 48 h after stimulation, GFP-expressing cells were sorted. Sμ-Sα switch junctions were amplified by PCR, cloned, and sequenced. Bar graphs show the percentage of switch junction sequences with indicated nucleotide overlap. Number of junctions analyzed (n), mean length of overlap (OL), and p-values relative to the Non-Target control (Mann-Whitney test) are indicated. White bars indicate the percentage of sequences with small (1–4 nucleotides) insertions. Overlap was determined by identifying the longest region of perfect uninterrupted donor/acceptor identity. Sequences with insertions were not included in the calculation of the mean length of overlap. Significant differences relative to the Non-Target control (χ2 test) are indicated: **, P ≤ 0.01; ***, P ≤ 0.0001. Data are from three independent experiments. (B) Cumulative percentage of sequences with a given length of microhomology (bp) and obtained from CH12 cells transduced with lentiviruses expressing shRNAs specific for Smc1 (red squares), Smc3 (green squares), Nipbl (blue squares), Wapal (gray squares), or a Non-Target negative control (black squares) and sorted for GFP expression. Data are from three independent experiments.
Knockdown of cohesin affects NHEJ. (A) CH12 cells were transduced with a lentivirus expressing a GFP reporter and shRNAs specific for AID, Smc1, Smc3, Nipbl, Wapal, or a Non-Target control. 48 h after stimulation, GFP-expressing cells were sorted. Sμ-Sα switch junctions were amplified by PCR, cloned, and sequenced. Bar graphs show the percentage of switch junction sequences with indicated nucleotide overlap. Number of junctions analyzed (n), mean length of overlap (OL), and p-values relative to the Non-Target control (Mann-Whitney test) are indicated. White bars indicate the percentage of sequences with small (1–4 nucleotides) insertions. Overlap was determined by identifying the longest region of perfect uninterrupted donor/acceptor identity. Sequences with insertions were not included in the calculation of the mean length of overlap. Significant differences relative to the Non-Target control (χ2 test) are indicated: **, P ≤ 0.01; ***, P ≤ 0.0001. Data are from three independent experiments. (B) Cumulative percentage of sequences with a given length of microhomology (bp) and obtained from CH12 cells transduced with lentiviruses expressing shRNAs specific for Smc1 (red squares), Smc3 (green squares), Nipbl (blue squares), Wapal (gray squares), or a Non-Target negative control (black squares) and sorted for GFP expression. Data are from three independent experiments.
Increased usage of microhomology at the junctions is reminiscent of what is observed in B cells defective for core components of the NHEJ pathway (Yan et al., 2007). Nevertheless, deficiency in XRCC4 or DNA ligase IV also results in a complete loss of sequences repaired through a direct joining (Yan et al., 2007). Therefore, it is unlikely that the cohesin complex is, by itself, part of the NHEJ machinery. As cohesin has been implicated in the recruitment of 53BP1 to γ-irradiation–induced foci (Watrin and Peters, 2009) and 53BP1 deficiency leads to defective CSR, increased DNA end resection, and preferential usage of microhomology (Bothmer et al., 2010), we speculate that cohesin could participate in the recruitment of 53BP1 to AID-induced DSBs and that defective 53BP1 recruitment could account for the increased usage of microhomology observed. Overall, our results implicate the cohesin complex in the mechanism of CSR and provide evidence for its involvement in regulating the repair of programmed DSBs.
MATERIALS AND METHODS
Nuclear extracts and coimmunoprecipitation.
Nuclear extracts and chromatin fractions were prepared using standard techniques (in the presence of 100 U/ml Benzonase; Novagen) from CH12F3 cells stably expressing AIDFLAG-HA, EGFPFLAG-HA, or the tags alone (Jeevan-Raj et al., 2011). Coimmunoprecipitations (in the presence of 100 U/ml Benzonase; Novagen) and Western blot analysis were performed as previously described (Jeevan-Raj et al., 2011). See Table S3 for antibodies used.
Mass spectrometry analysis.
20 mg nuclear extract were immunoprecipitated with Flag M2-agarose beads, washed, and eluted with Flag peptide as previously described (Jeevan-Raj et al., 2011). Flag eluates were fractionated by one-dimensional electrophoresis and processed as previously described (Jeevan-Raj et al., 2011) for identification by nano-LC-MS/MS or directly submitted to Multidimensional Protein Identification Technology (MudPIT). MudPIT analyses were performed as previously described (Washburn et al., 2001; Florens et al., 2006). In brief, protein mixtures were TCA-precipitated, urea-denatured, reduced, alkylated, and digested with endoproteinase Lys-C (Roche), followed by modified trypsin digestion (Promega). Peptide mixtures were loaded onto a triphasic 100 µm inner diameter fused silica microcapillary column. Loaded columns were placed in-line with a Dionex Ultimate 3000 nano-LC (Thermo Fisher Scientific) and an LTQ Velos linear ion trap mass spectrometer equipped with a nano-LC electrospray ionization source (Thermo Fisher Scientific). A fully automated 12-step MudPIT run was performed as previously described (Florens et al., 2006), during which each full MS scan (from 300 to 1,700 m/z range) was followed by 20 MS/MS events using data-dependent acquisition. Proteins were identified by database searching using SEQUEST (Thermo Fisher Scientific) with Proteome Discoverer 1.3 software (Thermo Fisher Scientific) against the mouse Swissprot database (2011–02 release). Peptides were filtered with Xcorr versus charge state 1.5–1, 2.5–2, 3–3, 3.2–4, and peptides of at least 7 amino acids in length.
shRNA-mediated knockdown.
The lentiviral vectors (pLKO.1 and pLKO.1-puro-CMV-TurboGFP) expressing shRNAs specific for AID (TRCN0000112031), Smc1 (TRCN0000109034), Smc3 (TRCN0000109007), Nipbl (TRCN0000124037), and Wapal (TRCN0000177268) or a Non-Target control (SHC002) were obtained from Sigma-Aldrich. The lentiviral vectors were transiently transfected into Lenti-X 293T cells (Takara Bio Inc.) to produce infectious viral particles as previously described (Willmann et al., 2012). 2 d later, CH12 cells were spin-infected with viral supernatants supplemented with 10 µg/ml polybrene (Sigma-Aldrich). Cells were selected for 5 d with 1 µg/ml puromycin before CSR induction.
Real-time quantitative (q) RT-PCR.
RNA and cDNA were prepared using standard techniques. qPCR was performed in triplicates using the Universal Probe Library (UPL) system (Roche) or SyberGreen (QIAGEN) and a LightCycler 480 (Roche). Transcript quantities were calculated relative to standard curves and normalized to β-actin, CD79b, or HPRT mRNA. See Table S4 for primers and probes.
Cell culture and flow cytometry.
Lentivirally transduced CH12 cells were cultured with 5 ng/ml IL-4 (Sigma-Aldrich), 1 ng/ml TGF-β (R&D System), 200 ng/ml monoclonal anti-CD40 antibody (eBioscience), and 1 µg/ml puromycin and analyzed after 48–72 h for cell surface expression of IgA by flow cytometry as previously described (Robert et al., 2009). Resting splenic B cells were isolated from 8–12-wk-old C57BL/6 mice using CD43 microbeads (Miltenyi Biotec) and cultured for 60 h with 50 µg/ml LPS (Sigma-Aldrich) and 5 ng/ml IL-4 (Sigma-Aldrich) as previously described (Jeevan-Raj et al., 2011). All animal work was performed under protocols approved by the Direction des Services Vétérinaires du Bas-Rhin, France (Authorization No. 67–343).
Switch junction analysis.
Sμ-Sα switch junctions were amplified using previously described primers (Ehrenstein and Neuberger, 1999; Schrader et al., 2002) and conditions (Robert et al., 2009) from genomic DNA prepared from lentivirally transduced CH12 cells stimulated for 72 h and sorted for GFP expression. PCR products were cloned using TOPO-TA cloning kit (Invitrogen) and sequenced using T7 universal primer. Sequence analysis was performed as previously described (Robert et al., 2009).
ChIP-Seq.
Resting or activated B cells were cross-linked for 10 min at 37°C with 1% (vol/vol) formaldehyde, followed by quenching with glycine (0.125 M final concentration). Cross-linked samples were then sonicated to obtain DNA fragments 200–500 bp in length using a sonicator (Covaris). Chromatin (from 10 × 107 cells) was precleared with protein A magnetic beads prewashed with PBS 0.05% tween, 5% BSA, and immunoprecipitated in ChIP dilution buffer (0.06% SDS, 20 mM Tris, pH 8.1, 2 mM EDTA, 160 mM NaCl, 1.045% Triton X-100, and 0.05× proteinase inhibitor cocktail) overnight at 4°C with protein A magnetic beads (Invitrogen) coupled to 100 µg Smc1 or Smc3 antibodies and processed according to the Millipore protocol. Cross-links were reversed for 4 h at 65°C in Tris-EDTA buffer with 0.3% (wt/vol) SDS and 1 mg/ml proteinase K. ChIP DNA was extracted with an IPure kit (Diagenode). Libraries were prepared for sequencing according to the manufacturer’s protocol (Illumina) and sequenced on the Illumina Genome Analyzer IIx as single-end 50 base reads according to manufacturer instructions. Image analysis and base calling were performed using the Illumina Pipeline and sequence reads were mapped to reference genome mm9/NCBI37 using Bowtie v0.12.7. Peak calling was performed using MACS (Zhang et al., 2008) with default parameters. Global comparison of samples and clustering analysis were performed using seqMINER (Ye et al., 2011).
ChIP-qPCR.
Analytical-scale ChIP was performed on chromatin prepared from 107 (resting or activated) splenic B cells isolated from a pool of five mice. qPCR was performed at several locations across the IgH locus using primers listed in Table S4. Results are expressed as percent input and represent the mean of three qPCR technical replicates. Error bars represent the corresponding standard deviation.
Accession codes.
ChIP-Seq data for CTCF, Smc1, and Smc3 on resting and activated B cells was submitted to GEO (GSE43594).
Online supplemental material.
Table S1 lists known AID partner proteins found. Table S2 lists novel AID partner proteins found. Table S3 lists antibodies. Table S4 lists primers. Table S5 shows ChIP-qPCR statistical analysis.
Acknowledgments
We thank members of the Reina-San-Martin laboratory for discussions; E. Soutoglou, I. Sumara, and M. Nussenzweig for comments on the manuscript; S. Pattabhiraman, M. Mendoza Parra, S. Legras, and B. Jost for assistance with ChIP-Seq analyses; V. Chavant and F. Ruffenach for assistance in mass spectrometry analysis; C. Ebel for cell sorting; and A. Gazumyan for advice on lentiviral infections.
A.-S. Thomas-Claudepierre was supported by the Ministère de l’Enseignement Supérieur et de la Recherche, France and the Fondation ARC. E. Schiavo was supported by the IGBMC International PhD Program and the Fondation ARC. This work was supported by grants to B. Reina-San-Martin from the Agence Nationale pour la Recherche (ANR-Blanc), the Fondation ARC (Program ARC), and the Institut National de la Santé et de la Recherche Médicale (Avenir-INSERM).
The authors have no conflicting financial interests.
References
Author notes
A.-S. Thomas-Claudepierre and E. Schiavo contributed equally to this paper.