The T cell receptor (TCR) pathway receives, processes, and amplifies the signal from pathogenic antigens to the activation of T cells. Although major components in this pathway have been identified, the knowledge on how individual components cooperate to effectively transduce signals remains limited. Phase separation emerges as a biophysical principle in organizing signaling molecules into liquid-like condensates. Here, we report that phospholipase Cγ1 (PLCγ1) promotes phase separation of LAT, a key adaptor protein in the TCR pathway. PLCγ1 directly cross-links LAT through its two SH2 domains. PLCγ1 also protects LAT from dephosphorylation by the phosphatase CD45 and promotes LAT-dependent ERK activation and SLP76 phosphorylation. Intriguingly, a nonmonotonic effect of PLCγ1 on LAT clustering was discovered. Computer simulations, based on patchy particles, revealed how the cluster size is regulated by protein compositions. Together, these results define a critical function of PLCγ1 in promoting phase separation of the LAT complex and TCR signal transduction.
Introduction
Mesoscale signaling clusters have been frequently observed in a variety of immune receptor pathways, including the T cell receptor (TCR; Campi et al., 2005), B cell receptor (Wang et al., 2017), Fcγ receptor (Lin et al., 2016), engulfment receptor (Draper; Williamson and Vale, 2018), chimeric antigen receptor (Dong et al., 2020), T cell coreceptor CD28 (Yokosuka et al., 2008), and PD1 (Hui et al., 2017). These signaling clusters, ranging from tens of nanometers to microns in size, share common features of complex composition, heterogeneous size, and dynamic assembly (Dustin and Groves, 2012). Because of these features, it remains a challenge to understand the mechanism and functional consequences of the clusters in transducing immune signaling.
The T cell microcluster represents a good example of this scenario. Following TCR activation, downstream signaling molecules self-organize into micron or submicron-sized clusters. These clusters are enriched of TCR, adaptor proteins, including linker for activation of T cells (LAT), Grb2, Gads, SLP76, Nck, and effectors, including kinase ZAP70, RasGEF Sos1, phospholipase Cγ1 (PLCγ1), E3 ubiquitin-protein ligase CBL, and actin regulator WASP (Balagopalan et al., 2015; Bunnell, 2010; Bunnell et al., 2002). Because the majority of early TCR signaling components reside in these microclusters, they are considered a hub for transducing TCR signals (Balagopalan et al., 2015; Choudhuri and Dustin, 2010). Previous work showed that LAT, a transmembrane protein essential for TCR signal transduction (Zhang et al., 1998), serves as a scaffold to form a macromolecular signaling complex (Houtman et al., 2006; Zhang et al., 2000). Mathematical modeling suggested that multivalent protein–protein interactions play a critical role in forming the LAT complex (Nag et al., 2012; Nag et al., 2009). Using a supported lipid bilayer–based system, our previous work showed that LAT forms near-micron-sized, membrane-embedded clusters through a mechanism of liquid-liquid phase separation (Su et al., 2016). The liquid-like LAT microclusters enrich kinase ZAP70 but exclude phosphatase CD45, thus promoting tyrosine phosphorylation. LAT microclusters also increase downstream Ras activation (Huang et al., 2019), actin polymerization, and extracellular signal–regulated kinase (ERK) activation (Su et al., 2016). These works revealed a critical role of LAT microclusters in promoting TCR signaling. However, the regulatory mechanism of LAT microclusters was not fully understood.
Following TCR activation, PLCγ1 is recruited to the LAT microclusters, which further hydrolyzes phosphatidylinositol 4,5-bisphosphate (PIP2) to generate inositol trisphosphate (IP3) and diacylglycerol (DAG), triggering calcium influx and PKC activation, respectively (Balagopalan et al., 2015; Courtney et al., 2018). PLCγ1 is a multidomain lipase (Hajicek et al., 2019) that contains an N-terminal PH domain, two pairs of calcium-sensing EF hands, a catalytic core (TIM barrel) interspaced by a split PH domain and structural SH2 and SH3 domains, and a C-terminal C2 domain (Fig. 1 A). As a unique feature of the γ subfamily of phospholipase C, PLCγ1 contains two SH2 domains and one SH3 domain that serve structural or regulatory roles (Braiman et al., 2006; Manna et al., 2018). Its N-terminal SH2 domain (nSH2) binds specifically to phosphotyrosine (Y132) on LAT (Braiman et al., 2006), whereas its C-terminal SH2 domain (cSH2) is involved in releasing autoinhibition of PLCγ1 (Gresset et al., 2010; Hajicek et al., 2019). The SH3 domain of PLCγ1 directly interacts with the proline-rich motifs (PRMs) on Sos1 (Kim et al., 2000), a RasGEF that is also enriched in the LAT microclusters. The multiple binary interactions between PLCγ1 and other components in the LAT complex mediate a synergistic assembly of the LAT complex (Braiman et al., 2006; Hartgroves et al., 2003; Manna et al., 2018). However, given the complex protein–protein interactions involved, the exact mechanism of how PLCγ1 regulates LAT microcluster formation remains elusive. This is mainly because traditional biochemical assays were performed in solution, which did not recapitulate two important features of cellular LAT microclusters: membrane association and giant size (several hundred nanometers).
PLCγ1 promotes LAT cluster formation in vitro. (A) Domain structure of PLCγ1. (B) Schematics of the assay. (C) Top: TIRF microscopy revealed that both Grb2 and PLCγ1 (full-length) promote LAT microcluster formation. Alexa Fluor 488–labeled LAT at 300 molecules/µm2 was incubated with 125 nM Sos1 and 250 nM Grb2 or PLCγ1 for 0.5 h before imaging. Scale bar, 5 µm. Bottom: Quantification of Grb2- or PLCγ1-driven LAT microclusters. LAT clustering was quantified as normalized variance (Su et al., 2016). Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; **, P < 0.01. (D) FRAP analysis revealed that PLCγ1-driven microclusters are less dynamic than Grb2-driven LAT microclusters. Shown are mean ± SD; n = 10 clusters.
PLCγ1 promotes LAT cluster formation in vitro. (A) Domain structure of PLCγ1. (B) Schematics of the assay. (C) Top: TIRF microscopy revealed that both Grb2 and PLCγ1 (full-length) promote LAT microcluster formation. Alexa Fluor 488–labeled LAT at 300 molecules/µm2 was incubated with 125 nM Sos1 and 250 nM Grb2 or PLCγ1 for 0.5 h before imaging. Scale bar, 5 µm. Bottom: Quantification of Grb2- or PLCγ1-driven LAT microclusters. LAT clustering was quantified as normalized variance (Su et al., 2016). Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; **, P < 0.01. (D) FRAP analysis revealed that PLCγ1-driven microclusters are less dynamic than Grb2-driven LAT microclusters. Shown are mean ± SD; n = 10 clusters.
We have recently developed a supported lipid bilayer–based system to reconstitute near-micron-sized LAT microclusters on synthetic membranes (Su et al., 2017). In the present study, using our biochemical reconstitution approach together with live-cell microscopy and computer modeling, we delineated the role of PLCγ1 in regulating LAT microcluster formation. We found that the SH2 and SH3 domain of PLCγ1 can directly bridge LAT to Sos1 to form microclusters. PLCγ1 also protects LAT from CD45-dependent dephosphorylation. Therefore, PLCγ1 stabilizes LAT microclusters by both physical cross-linking and chemical protection. Moreover, we found that the PLCγ1 concentration influences the sizes of LAT microclusters in a nonmonotonic way, pointing to a novel mechanism for the size control of liquid condensates. Together, these results expand the traditional view that PLCγ1 acts as an enzyme downstream of LAT; instead, PLCγ1 plays a more upstream structural role in promoting LAT cluster assembly and regulating the stability of LAT microclusters.
Results
PLCγ1 promotes LAT cluster formation in vitro
To determine how PLCγ1 regulates LAT microcluster formation, we implemented a supported lipid bilayer–based reconstitution assay that allows quantitative monitoring of microcluster assembly (Su et al., 2017). The cytoplasmic fragment of LAT that contains the four C-terminal tyrosines that are both necessary and sufficient for TCR signaling (Zhu et al., 2003) was purified, phosphorylated, and labeled with maleimide–Alexa Fluor 488 on a C-terminal cysteine residue. It was then attached to the Ni2+-NTA (nitrilotriacetic acid) functionalized supported lipid bilayer via a polyhistidine tag on the N terminus. Total internal reflection fluorescence (TIRF) microscopy was used to visualize the formation of LAT clusters. As reported before (Su et al., 2016), LAT formed microclusters when Grb2 and Sos1 were added to the reaction mixture (Fig. 1, B and C). Intriguingly, when Grb2 was replaced with full-length PLCγ1, LAT still formed clusters, although in higher numbers but of smaller sizes, as compared with the Grb2-mediated cluster formation (Fig. 1 C). Furthermore, FRAP analysis revealed that PLCγ1-induced LAT clusters are less dynamic than Grb2-induced LAT clusters, as indicated by the lower recovery frequency and longer half-recovery time (Fig. 1 D). We also tested a fragment of PLCγ1 that contains the SH2 and SH3 domains (PLCγ1-SH; illustrated in Fig. 1 A) and found that it also robustly induced LAT microcluster formation, even at a higher level as compared with the full-length PLCγ1 (Fig. S1, A and B).
Both the full-length and SH fragment of PLCγ1 drive LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with the full-length or SH2-SH2-SH3 domain of PLCγ1. Alexa Fluor 488–labeled, phosphorylated LAT at 300 molecules/µm2 was incubated with 250 nM Sos1 and indicated concentrations of PLCγ1 or fragment. Scale bar, 5 µm. (B) Quantification of PLCγ1-driven LAT microclusters. Shown are mean ± SD; n = 3 independent experiments. (C) Recombinant proteins used in this study. Purified proteins were applied to SDS-PAGE, followed by Coomassie blue staining.
Both the full-length and SH fragment of PLCγ1 drive LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with the full-length or SH2-SH2-SH3 domain of PLCγ1. Alexa Fluor 488–labeled, phosphorylated LAT at 300 molecules/µm2 was incubated with 250 nM Sos1 and indicated concentrations of PLCγ1 or fragment. Scale bar, 5 µm. (B) Quantification of PLCγ1-driven LAT microclusters. Shown are mean ± SD; n = 3 independent experiments. (C) Recombinant proteins used in this study. Purified proteins were applied to SDS-PAGE, followed by Coomassie blue staining.
To understand if the ability to drive LAT cluster formation is a general feature of proteins containing the SH2 and SH3 domains, we replaced PLCγ1 with other LAT-associated proteins that play a role in TCR signal transduction. These include (1) Gads, an adaptor protein that binds LAT on overlapping sites with Grb2 (Zhang et al., 2000), (2) Vav1, a RhoGEF that closely associates with LAT (Sherman et al., 2016), and (3) Nck1, an adaptor protein that promotes actin polymerization (Wunderlich et al., 1999). We found that Gads, as reported before (Su et al., 2016), drove LAT microcluster formation, whereas Vav1 or Nck1 did not (Fig. 2, A and B). This could be potentially explained by the binding preference of individual SH2 domains for the sequence context around phosphotyrosines. LAT contains phosphotyrosine motifs that are preferentially recognized by the SH2 domain of Grb2, Gads, and PLCγ1, but not Vav1 or Nck1 (Tinti et al., 2013). Nevertheless, we do not exclude other factors, for example, the linker length and flexibility between the SH2 and SH3 domain, that could affect clustering ability. Together, those data demonstrate that PLCγ1, together with Sos1, can specifically induce LAT microcluster formation.
Specific SH2-SH3–containing proteins promote LAT cluster formation. (A) TIRF microscopy revealed LAT microcluster formation with different SH2 and SH3 domain proteins. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 250 nM Sos1 (labeled with Alexa Fluor 647) and indicated proteins at 500 nM. Scale bar, 5 µm. (B) Quantification of LAT clustering and membrane recruitment of Sos1. Shown are mean ± SD; n = 3 independent experiments.
Specific SH2-SH3–containing proteins promote LAT cluster formation. (A) TIRF microscopy revealed LAT microcluster formation with different SH2 and SH3 domain proteins. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 250 nM Sos1 (labeled with Alexa Fluor 647) and indicated proteins at 500 nM. Scale bar, 5 µm. (B) Quantification of LAT clustering and membrane recruitment of Sos1. Shown are mean ± SD; n = 3 independent experiments.
PLCγ1 cross-links LAT through two SH2 domains
Next, we determined the mechanism by which PLCγ1 drives LAT microcluster formation. PLCγ1 contains an nSH2, a cSH2, and an SH3 domain. We produced PLCγ1 truncation mutants that lack either the nSH2, cSH2, or SH3 domain (Fig. 3 A). We found that mutants lacking either nSH2 or cSH2 lost the ability to drive LAT cluster formation, whereas mutants lacking the SH3 domain drove attenuated LAT cluster formation (Fig. 3, B and C). Because the SH3 domain interacts with the PRM on Sos1, the SH3-independent clustering suggested that Sos1 might be dispensable for PLCγ1-driven LAT cluster formation. Indeed, the nSH2-cSH2 fragment of PLCγ1 drove LAT cluster formation in the absence of Sos1 (Fig. S2, A and B). It has to be noted that in this assay, PLCγ1 and mutants were used at 500 nM. We titrated PLCγ1 concentration and found that at the physiologically relevant concentration of PLCγ1 (∼50 nM), Sos1 is still required for LAT microcluster formation (Fig. 4, A and B). The binding affinities of the SH3 domain to the individual PRMs on Sos1 were measured by surface plasmon resonance (Fig. 4 C), which supports a model in which Sos1 cross-links PLCγ1 to promote LAT clustering (Fig. 1 B). Interestingly, the SH3 domain of PLCγ1 was also reported to interact with the PRM on SLP76, another component of the LAT complex (Deng et al., 2005). This raises the possibility that SLP76 competes with Sos1 in binding PLCγ1 and inhibits PLCγ1-mediated clustering. Indeed, we found a mild inhibition of SLP76 on LAT/PLCγ1/Sos1 clustering (Fig. 4, D and E). Together, these assays comprehensively revealed an interaction network that regulates LAT cluster formation.
PLCγ1 cross-links LAT by two SH2 domains. (A) Domains of the proteins used in the study. (B) TIRF microscopy revealed that both nSH2 and cSH2 are required for PLCγ1-driven LAT microcluster formation. SH3 domain promotes cluster formation. Alexa Fluor 488–labeled LAT at 300 molecules/µm2 was incubated with 300 nM Sos1 and 50 nM PLCγ1 for 0.5 h before imaging. Scale bar, 5 µm. (C) Quantification of PLCγ1-driven LAT microclusters. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05. (D) Schematics of the assay of testing SH2 domain binding sites. (E) PLCγ1 nSH2 binds LAT Y132. Phosphopeptides were synthesized, biotinylated at the N terminus, and attached to the biotin-functionalized supported lipid bilayers by streptavidin. The SH2 domains were labeled with fluorescent dye (Maleimide-Ax647) and incubated with the individual phosphopeptides. The membrane-associated SH2 domain was measured by TIRF microscopy. Scale bar, 5 µm. (F) PLCγ1 cSH2 binds LAT Y171. Same settings as in E.
PLCγ1 cross-links LAT by two SH2 domains. (A) Domains of the proteins used in the study. (B) TIRF microscopy revealed that both nSH2 and cSH2 are required for PLCγ1-driven LAT microcluster formation. SH3 domain promotes cluster formation. Alexa Fluor 488–labeled LAT at 300 molecules/µm2 was incubated with 300 nM Sos1 and 50 nM PLCγ1 for 0.5 h before imaging. Scale bar, 5 µm. (C) Quantification of PLCγ1-driven LAT microclusters. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05. (D) Schematics of the assay of testing SH2 domain binding sites. (E) PLCγ1 nSH2 binds LAT Y132. Phosphopeptides were synthesized, biotinylated at the N terminus, and attached to the biotin-functionalized supported lipid bilayers by streptavidin. The SH2 domains were labeled with fluorescent dye (Maleimide-Ax647) and incubated with the individual phosphopeptides. The membrane-associated SH2 domain was measured by TIRF microscopy. Scale bar, 5 µm. (F) PLCγ1 cSH2 binds LAT Y171. Same settings as in E.
Domains required for PLCγ1-driven LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with a high concentration of PLCγ1 fragments. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 125 nM Sos1 and 500 nM of indicated PLCγ1 fragments. Scale bar, 5 µm. (B) Quantification of LAT clustering in A. Shown are mean ± SD; n = 3 independent experiments. (C) TIRF microscopy revealed LAT microcluster formation in the presence or absence of PLCγ1. Alexa Fluor 488 LAT at 1,000 molecules/µm2 was incubated with 500 nM Sos1 and 1,000 nM Grb2 with or without 100 nM full-length PLCγ1. Scale bar, 5 µm. (D) Quantification of LAT clustering. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. ***, P < 0.001. (E) FRAP analysis revealed that PLCγ1 decreases the recovery of LAT signal in clusters after photobleaching. Shown are mean ± SD; n = 10 clusters.
Domains required for PLCγ1-driven LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with a high concentration of PLCγ1 fragments. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 125 nM Sos1 and 500 nM of indicated PLCγ1 fragments. Scale bar, 5 µm. (B) Quantification of LAT clustering in A. Shown are mean ± SD; n = 3 independent experiments. (C) TIRF microscopy revealed LAT microcluster formation in the presence or absence of PLCγ1. Alexa Fluor 488 LAT at 1,000 molecules/µm2 was incubated with 500 nM Sos1 and 1,000 nM Grb2 with or without 100 nM full-length PLCγ1. Scale bar, 5 µm. (D) Quantification of LAT clustering. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. ***, P < 0.001. (E) FRAP analysis revealed that PLCγ1 decreases the recovery of LAT signal in clusters after photobleaching. Shown are mean ± SD; n = 10 clusters.
Sos1 facilitates PLCγ1-driven LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with titrated PLCγ1. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with or without 250 nM Sos1 and the indicated concentration of PLCγ1 nSH2-cSH2-SH3 domains. Scale bar, 5 µm. (B) Quantification of LAT clustering in A. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. *, P < 0.05; **, P < 0.01. (C) Binding affinity (Kd) of the SH3 domain of PLCγ1 to the PRM on Sos1 as measured by surface plasmon resonance. Shown are mean ± SEM. (D) TIRF microscopy revealed that SLP76 inhibits LAT microcluster formation driven by PLCγ1 and Sos1. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 300 nM Sos1, 50 nM PLCγ1 (nSH2-cSH2-SH3 domains), and the indicated concentration of SLP76. Proteins were used at the physiologically relevant concentration. The cellular concentration of SLP76 was estimated as 150 nM (by MaxQB Database). Scale bar, 5 µm. (E) Quantification of LAT clustering in D. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. *, P < 0.05; ns, not significant.
Sos1 facilitates PLCγ1-driven LAT clustering. (A) TIRF microscopy revealed LAT microcluster formation with titrated PLCγ1. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with or without 250 nM Sos1 and the indicated concentration of PLCγ1 nSH2-cSH2-SH3 domains. Scale bar, 5 µm. (B) Quantification of LAT clustering in A. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. *, P < 0.05; **, P < 0.01. (C) Binding affinity (Kd) of the SH3 domain of PLCγ1 to the PRM on Sos1 as measured by surface plasmon resonance. Shown are mean ± SEM. (D) TIRF microscopy revealed that SLP76 inhibits LAT microcluster formation driven by PLCγ1 and Sos1. Alexa Fluor 488 LAT at 300 molecules/µm2 was incubated with 300 nM Sos1, 50 nM PLCγ1 (nSH2-cSH2-SH3 domains), and the indicated concentration of SLP76. Proteins were used at the physiologically relevant concentration. The cellular concentration of SLP76 was estimated as 150 nM (by MaxQB Database). Scale bar, 5 µm. (E) Quantification of LAT clustering in D. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test. *, P < 0.05; ns, not significant.
Next, we determined the binding sites on LAT that interact with the SH2 domain of PLCγ1. The four C-terminal tyrosines on LAT are necessary and sufficient to transduce the TCR signaling (Zhu et al., 2003). Synthesized peptides containing each one of these four phosphotyrosines of LAT were attached to the supported lipid bilayer. Recombinant nSH2 or cSH2 of PLCγ1 was purified, labeled with CoA-647 on an N-terminal ybbR tag, and incubated with individual phosphopeptides on the membrane (Fig. 3 D). The binding of the SH2 domain to phosphopeptides on the membrane was revealed by TIRF microscopy. We found that the nSH2 strongly interacted with LAT Y132 (Fig. 3 E), which is consistent with previous reports (Zhang et al., 2000). The cSH2 robustly bound to LAT Y171, although the binding was slightly lower than PLCγ1 Y783 (Fig. 3 F), the previously reported site that interacts with the cSH2 in cis (Hajicek et al., 2013; Poulin et al., 2005). Together, those data suggested a model in which PLCγ1 cross-links LAT through two interaction pairs: nSH2 preferentially with Y132, and cSH2 preferentially with Y171.
PLCγ1 cooperates with Grb2 to regulate LAT clustering
Next, we investigated how PLCγ1 cooperates with Grb2 and Sos1 to induce LAT microcluster formation. Because cluster formation depends on the concentration of individual components, we adopted concentrations of proteins that were measured in T cells (Nag et al., 2009; Voisinne et al., 2019). We incubated LAT at a density of 300 molecules/µm2 with 3,000 nM Grb2 and 300 nM Sos1. PLCγ1 (SH2-2-3 domain) was additionally included in the clustering assay. Surprisingly, we found that PLCγ1, at 50 nM (concentration in Jurkat T cells), could significantly increase LAT cluster formation and recruitment of Sos1 to the membrane. This clustering-promoting effect is robust even at a concentration as low as 5 nM PLCγ1 (Fig. 5, A and B). Notably, this concentration (5 nM) is orders of magnitude lower than that of Grb2 (3,000 nM), the other SH2-SH3 adaptor in the system. Furthermore, FRAP analysis revealed that PLCγ1 significantly reduces the exchange of LAT molecules between inside and outside of the clusters (Fig. S2, C–E), supporting the idea that PLCγ1 serves as a cross-linker orthogonal to Grb2, to stabilize LAT clusters. To understand how PLCγ1 affects the kinetics of LAT cluster formation, we performed time-lapse imaging. LAT was attached to the membrane, and Grb2 and Sos1 were added into the system at time 0. We observed cluster formation as usual (Video 1). Intriguingly, when PLCγ1 (SH2-2-3 domains) was additionally included in the system, clustering was significantly enhanced from the very beginning (Video 2 and Fig. 5 C). Together, these data suggest that PLCγ1, by serving as a bridging protein besides Grb2, could dramatically increase LAT microcluster formation at physiologically relevant concentrations.
PLCγ1 cooperates with Grb2 to regulate LAT microcluster formation. (A) TIRF microscopy revealed that PLCγ1 regulates LAT microcluster formation in a nonmonotonic manner. Physiologically relevant concentrations of proteins were used in the assay: LAT at 300 molecules/µm2, Grb2 at 3 µM, Sos1 at 0.3 µM, and PLCγ1 at 50 nM. LAT was labeled with Alexa Fluor 488, PLCγ1 (SH2-2-3 domains) was labeled with DY547, and Sos1 was labeled with Alexa Fluor 647. Scale bar, 5 µm. (B) Quantification of LAT clustering, membrane recruitment of Sos1. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. (C) PLCγ1 accelerates LAT cluster formation. TIRF microscopy revealed the time course of LAT microcluster formation in the presence or absence of PLCγ1. LAT–Alexa Fluor 488 at 1,000 molecules/µm2 was incubated with 1,000 nM Grb2 and 500 nM Sos1 and/or 50 nM PLCγ1 at time 0. Shown are mean ± SEM; n = 3 independent experiments.
PLCγ1 cooperates with Grb2 to regulate LAT microcluster formation. (A) TIRF microscopy revealed that PLCγ1 regulates LAT microcluster formation in a nonmonotonic manner. Physiologically relevant concentrations of proteins were used in the assay: LAT at 300 molecules/µm2, Grb2 at 3 µM, Sos1 at 0.3 µM, and PLCγ1 at 50 nM. LAT was labeled with Alexa Fluor 488, PLCγ1 (SH2-2-3 domains) was labeled with DY547, and Sos1 was labeled with Alexa Fluor 647. Scale bar, 5 µm. (B) Quantification of LAT clustering, membrane recruitment of Sos1. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. (C) PLCγ1 accelerates LAT cluster formation. TIRF microscopy revealed the time course of LAT microcluster formation in the presence or absence of PLCγ1. LAT–Alexa Fluor 488 at 1,000 molecules/µm2 was incubated with 1,000 nM Grb2 and 500 nM Sos1 and/or 50 nM PLCγ1 at time 0. Shown are mean ± SEM; n = 3 independent experiments.
LAT cluster formation with Grb2 and Sos1. TIRF microscopy revealed LAT cluster formation. His-pLAT–Alexa Fluor 488 was attached to Ni-functionalized supported lipid bilayers at 1,000 molecules/µm2. Grb2 (1 µM) and Sos1 (0.5 µM) were added at 0 s to trigger LAT cluster assembly. Shown is a field view of 30 × 30 μm. The video was recorded at a rate of 10 s/frame.
LAT cluster formation with Grb2 and Sos1. TIRF microscopy revealed LAT cluster formation. His-pLAT–Alexa Fluor 488 was attached to Ni-functionalized supported lipid bilayers at 1,000 molecules/µm2. Grb2 (1 µM) and Sos1 (0.5 µM) were added at 0 s to trigger LAT cluster assembly. Shown is a field view of 30 × 30 μm. The video was recorded at a rate of 10 s/frame.
LAT cluster formation with Grb2, Sos1, and PLCγ1. Same condition as in Video 1 except that PLCγ1 (50 nM) was added together with Grb2 and Sos1 at 0 s. The video was recorded at a rate of 10 s/frame.
LAT cluster formation with Grb2, Sos1, and PLCγ1. Same condition as in Video 1 except that PLCγ1 (50 nM) was added together with Grb2 and Sos1 at 0 s. The video was recorded at a rate of 10 s/frame.
Computer simulations of PLCγ1-mediated LAT clustering
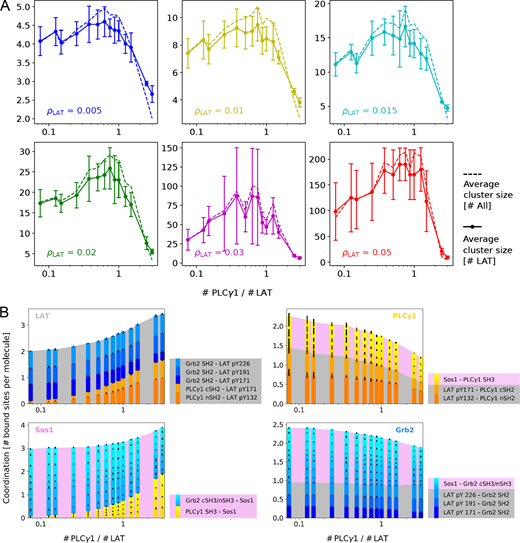
Intriguingly, a nonmonotonic effect of PLCγ1 on LAT clustering was revealed. PLCγ1, at low concentrations, promotes LAT clustering and increases cluster sizes, but this effect is diminished at high concentrations (Fig. 5, A and B). This points to the fact that PLCγ1 not only promotes LAT clustering, but also regulates the cluster size. To understand the physical mechanism underlying this size regulation, we developed a minimal coarse-grained computer model in which the proteins are described as spherical particles decorated with patches that represent binding domains and are simulated via molecular dynamics. The model, described quantitatively in Materials and methods and in Simulation details, consists of four types of 2D particles, representing LAT, PLCγ1, Sos1, and Grb2; these can bind mutually, respecting biochemical valence and bond specificity (Fig. 6 A). In simulations, the pool of such 2D particles readily aggregates into clusters that grow either via addition of individual proteins or via merging with other clusters until they reach steady-state sizes. The average cluster size in simulations exhibits a nonmonotonic dependence on the concentration of PLCγ1 (Fig. 6 B; and Videos 3, 4, 5, 6, 7, and 8), indeed recapitulating what was observed in wet experiments (Fig. 5 B). The nonmonotonic effect was robustly revealed in a wide range of LAT densities, with cluster size measured either by the number of LAT or by the total number of proteins (Fig. 7 A). To understand the origin of this nonmonotonic phenomenon, we computed, at a given time, the number of possible bonds between free binding sites that can make whichever two clusters merge into a bigger one; this parameter, that we named coalescence likelihood, is a measure of how easy it is for the clusters to merge at a certain PLCγ1 concentration. We found that the coalescence likelihood closely captures the nonmonotonicity of the cluster size (Fig. 6 B). A breakdown of the coalescence likelihood by bond type (Fig. 6 C), together with an analysis of the average coordination per molecule (Simulation details, section SI 3), showed the following. At low concentrations, PLCγ1 particles provide a binding site for the otherwise unbound pY132 in LAT particles and increase binding of the pY171 site and of all Sos1 sites, through an SH3-PRM bond. The availability of these additional binding sites increases the coalescence likelihood (yellow and orange bars in Fig. 6 C); at the same time, it increases the average coordination (number of sites that interact with other molecules) of both LAT and Sos1 particles, making clusters more connected (Figs. 6 E and 7 B). At high concentrations, though, PLCγ1 ends up saturating PRMs on Sos1 and, more slowly, phosphotyrosines on LAT particles, with the help of Grb2 (Fig. 7 B). As this occurs, free binding sites become rare, and the coalescence likelihood decreases drastically. Clusters are still more connected and compact than without PLCγ1, but they are smaller: indeed, Sos1 and LAT are almost fully bound and it is unlikely for new bonds to form upon random collisions (pink and gray bands in Fig. 6 C restrict by more than half; Video 8).
A coarse-grained model explains how PLCγ1 nonmonotonically regulates LAT clustering. (A) Sketch of the model in which the proteins are represented as 2D particles decorated by interaction patches. All bonds possible in the system, based on biochemical data, are illustrated with colored lines. (B) Top: The average cluster size displays nonmonotonic dependence on the PLCγ1 concentration (gray circles). This behavior is well captured by the likelihood for cluster coalescence (black squares). Error bars represent statistical errors on the average size over 10 different realizations of the simulation, shown are mean ± SEM. Bottom: Snapshots of typical clusters in simulations, for relative PLCγ1:LAT concentrations of 0.075, 0.75, and 3 (these clusters contain, respectively, 19, 30, and 10 LAT molecules, and with reference to D, their compactness is 0.23, 0.39, and 0.48). (C) Breakdown of the coalescence likelihood per type of possible bond. The gray and pink areas represent available bonds involving a LAT or a Sos1 molecule, respectively; blue and yellow-orange bars represent bonds involving Grb2 and PLCγ1, respectively. (D) Compactness (gray circles, see Materials and methods) and fraction of terminal nodes (black squares), as a function of PLCγ1 concentration. Shown are mean ± SD; n = 10 realizations. (E) Fraction of LAT, PLCγ1, Sos1, and Grb2 molecules per cluster, as a function of PLCγ1 concentration, shaded according to the number of other molecules they are bound to.
A coarse-grained model explains how PLCγ1 nonmonotonically regulates LAT clustering. (A) Sketch of the model in which the proteins are represented as 2D particles decorated by interaction patches. All bonds possible in the system, based on biochemical data, are illustrated with colored lines. (B) Top: The average cluster size displays nonmonotonic dependence on the PLCγ1 concentration (gray circles). This behavior is well captured by the likelihood for cluster coalescence (black squares). Error bars represent statistical errors on the average size over 10 different realizations of the simulation, shown are mean ± SEM. Bottom: Snapshots of typical clusters in simulations, for relative PLCγ1:LAT concentrations of 0.075, 0.75, and 3 (these clusters contain, respectively, 19, 30, and 10 LAT molecules, and with reference to D, their compactness is 0.23, 0.39, and 0.48). (C) Breakdown of the coalescence likelihood per type of possible bond. The gray and pink areas represent available bonds involving a LAT or a Sos1 molecule, respectively; blue and yellow-orange bars represent bonds involving Grb2 and PLCγ1, respectively. (D) Compactness (gray circles, see Materials and methods) and fraction of terminal nodes (black squares), as a function of PLCγ1 concentration. Shown are mean ± SD; n = 10 realizations. (E) Fraction of LAT, PLCγ1, Sos1, and Grb2 molecules per cluster, as a function of PLCγ1 concentration, shaded according to the number of other molecules they are bound to.
Early phase of simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 15 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Early phase of simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 15 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 15 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 15 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Early phase of simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 150 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Early phase of simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 150 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 150 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 150 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Early phase of simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 600 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Early phase of simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. The simulation involves 200 LAT, 600 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 10 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 8 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 600 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Full-length simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. High-resolution video for visualizing individual chemical bonds. The simulation involves 200 LAT, 600 PLCγ1, 400 Grb2, and 200 Sos1 molecules, all in a monomeric state. The simulation starts at second 2.00, corresponding to timestep 0, and ends at timestep 50 × 106. The interval between two frames is 0.2 × 106 timesteps, and the frame rate is 16 s−1. Particle scheme is the same as in Fig. 6 A: gray, LAT; yellow, PLCγ1; blue, Grb2; and pink, Sos1.
Simulating PLCγ1’s effect on LAT cluster size and bond type.(A) The effect of PLCγ1 on LAT clustering is independent of LAT density ρLAT. Coarse-grained model simulating LAT clustering as a function of PLCγ1 concentration. In a wide range of LAT densities tested, PLCγ1 regulates LAT clustering in a nonmonotonic manner. LAT clusters are quantified by the number of LAT in each cluster (solid line) or the total number of molecules (LAT, Grb2, PLCγ1, or Sos1) in each cluster (dashed line, scaled down by a factor of 4 to fit in the same plot). Surface densities ρLAT are in units of σ−2, where σ is the diameter of a particle, of the order of a few nanometers, and our experiments correspond roughly to ρLAT = 0.02σ−2 (see Materials and methods and Simulation details). Shown are mean ± SEM. (B) Average coordination number for all four kinds of particles, as a function of ratio of PLCγ1:LAT, broken down to the contribution of each specific bond. Yellow-orange bars represent bonds involving PLCγ1, and blue bars, Grb2; a gray background represents bonds involving LAT, and a pink background, Sos1. Here, as throughout Fig. 6, ρLAT = 0.02 σ−2. See Simulation details, section SI 3, for a complete analysis. Shown are mean ± SEM.
Simulating PLCγ1’s effect on LAT cluster size and bond type.(A) The effect of PLCγ1 on LAT clustering is independent of LAT density ρLAT. Coarse-grained model simulating LAT clustering as a function of PLCγ1 concentration. In a wide range of LAT densities tested, PLCγ1 regulates LAT clustering in a nonmonotonic manner. LAT clusters are quantified by the number of LAT in each cluster (solid line) or the total number of molecules (LAT, Grb2, PLCγ1, or Sos1) in each cluster (dashed line, scaled down by a factor of 4 to fit in the same plot). Surface densities ρLAT are in units of σ−2, where σ is the diameter of a particle, of the order of a few nanometers, and our experiments correspond roughly to ρLAT = 0.02σ−2 (see Materials and methods and Simulation details). Shown are mean ± SEM. (B) Average coordination number for all four kinds of particles, as a function of ratio of PLCγ1:LAT, broken down to the contribution of each specific bond. Yellow-orange bars represent bonds involving PLCγ1, and blue bars, Grb2; a gray background represents bonds involving LAT, and a pink background, Sos1. Here, as throughout Fig. 6, ρLAT = 0.02 σ−2. See Simulation details, section SI 3, for a complete analysis. Shown are mean ± SEM.
A further analysis of the compactness of clusters was performed by computing their inertia tensor, as well as by graph-theoretical means (see Materials and methods and Simulation details). The compactness index showed that clusters become monotonically more compact as PLCγ1 concentration increases (Fig. 6 D). Nonetheless, at high concentrations, clusters exhibit a large amount of “terminal nodes,” i.e., particles bound to one other particle only. This is due to the overwhelming amount of PLCγ1 particles, which tend to cap any binding site available to them, competing with themselves and with Grb2 for LAT and Sos1 sites. At the same time, as expected, LAT and Sos1 molecules become more and more bound (Figs. 6 E and 7 B). This suggests that the stabilizing effect of PLCγ1 observed in FRAP experiments is due to increased compactness of clusters and increased complexity of the LAT network therein. In short, PLCγ1 concentration emerges as a possible regulator of both cluster size and cluster stability.
PLCγ1 promotes LAT clustering in T cells
We then investigated how PLCγ1 regulates LAT cluster formation in T cells. A LAT-mCherry construct was introduced into wild-type or PLCγ1-null Jurkat T cells by lentiviral transduction. Those T cells were activated by cover glass–coated TCR-activating antibody OKT3. The formation of LAT microclusters was monitored by TIRF microscopy. We found that LAT clustering in PLCγ1-null cells was significantly reduced as compared with the wild-type cells (Fig. 8 A). This suggested a positive role of PLCγ1 in promoting LAT clusters, which is consistent with aforementioned in vitro results (Fig. 4 A). To understand how the SH2 and SH3 domains of PLCγ1 contribute to LAT clustering, we reconstituted PLCγ1-null cells with full-length PLCγ1 or PLCγ1 lacking the nSH2 or the SH3 domain. These cells also express LAT-mCherry, as an indicator for LAT clusters. Unfortunately, PLCγ1-null cells that are reconstituted with PLCγ1 lacking the cSH2 do not grow, potentially because of the hyperactivity and toxicity resulting from the deletion of the inhibitory cSH2. We found that LAT microcluster formation was significantly higher in the wild-type cells, as compared with the ΔnSH2 or ΔSH3 cells (Fig. 8 B).
PLCγ1 promotes LAT clustering, SLP76 phosphorylation, and ERK activation in Jurkat T cells. (A) Diminished LAT microcluster formation in PLCγ1-null cells. Wild-type or PLCγ1-null Jurkat T cells expressing LAT-mCherry were plated on OKT3-coated cover glass. LAT microcluster formation was revealed by TIRF microscopy. Images showed clustering 90 s after cell landing on the glass. Scale bar, 5 µm. Shown are mean ± SEM; n = 25 or 26 cells. Unpaired two-tailed t test was used. **, P < 0.01. (B) The nSH2 and SH3 domain of PLCγ1 promotes LAT cluster formation. PLCγ1-null Jurkat T cells expressing LAT-mCherry were reconstituted with the GFP-tagged wild-type, ΔnSH2, or ΔSH3 PLCγ1. Those cells were plated on OKT3-coated cover glass. Images showed clustering 90 s after cell landing on the glass. LAT microcluster formation was revealed by TIRF microscopy. Scale bar, 5 µm. Shown are mean ± SEM; n = 22–30 cells. Unpaired two-tailed t test was used. **, P < 0.01. (C) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild-type, ΔnSH2, or ΔSH3 PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (D) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; **, P < 0.01.
PLCγ1 promotes LAT clustering, SLP76 phosphorylation, and ERK activation in Jurkat T cells. (A) Diminished LAT microcluster formation in PLCγ1-null cells. Wild-type or PLCγ1-null Jurkat T cells expressing LAT-mCherry were plated on OKT3-coated cover glass. LAT microcluster formation was revealed by TIRF microscopy. Images showed clustering 90 s after cell landing on the glass. Scale bar, 5 µm. Shown are mean ± SEM; n = 25 or 26 cells. Unpaired two-tailed t test was used. **, P < 0.01. (B) The nSH2 and SH3 domain of PLCγ1 promotes LAT cluster formation. PLCγ1-null Jurkat T cells expressing LAT-mCherry were reconstituted with the GFP-tagged wild-type, ΔnSH2, or ΔSH3 PLCγ1. Those cells were plated on OKT3-coated cover glass. Images showed clustering 90 s after cell landing on the glass. LAT microcluster formation was revealed by TIRF microscopy. Scale bar, 5 µm. Shown are mean ± SEM; n = 22–30 cells. Unpaired two-tailed t test was used. **, P < 0.01. (C) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild-type, ΔnSH2, or ΔSH3 PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (D) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; **, P < 0.01.
Because LAT clustering activates the downstream signaling pathway including SLP76 and MAPK/ERK, we decided to determine if PLCγ1 affects these pathways. Jurkat T cells were activated by anti-CD3 and anti-CD28 antibodies before being harvested for immunoblot analysis. Indeed, as compared with cells expressing the wild-type PLCγ1, cells expressing PLCγ1ΔnSH2 or PLCγ1ΔSH3 displayed defects in SLP76 phosphorylation and MAPK activation (as indicated by pERK1/2; Fig. 8, C and D). In the concern that deleting the nSH2 domain might result in a defect in folding or conformation change of other parts of PLCγ1, we introduced a point mutation into the nSH2 domain (R586K) that abolishes its interaction with LAT (Stoica et al., 1998). We showed that, similar to the PLCγ1ΔnSH2, PLCγ1 R586K mediated attenuated LAT clustering and reduced SLP76 and ERK phosphorylation (Fig. S3). To rigorously test the enzyme-independent, structural function of PLCγ1 in promoting LAT signaling, we expressed PLCγ1 H380F, which loses most of the lipase activity (Smith et al., 1994), in PLCγ1-null cells. We found that PLCγ1 H380F can still mediate robust SLP76 phosphorylation as compared with wild-type PLCγ1 (Fig. S4, A and B). Notably, the ERK activation in PLCγ1 H380F cells is significant, but it is much lower than in wild-type cells (Fig. S4, A and B). This can be explained by the fact that the enzymatic activity of PLCγ1 is required for PIP2 hydrolysis, which generates diacylglycerol, and diacylglycerol-induced RasGRP1 activation is required for priming Sos1 activation and dominates over Sos1 to induce ERK phosphorylation (Roose et al., 2007). Together, our data delineated the structural function of PLCγ1 in promoting LAT clustering and LAT downstream pathways in T cells.
The nSH2–pY interaction is required for PLCγ1-mediated LAT clustering and signaling. (A) Diminished LAT microcluster formation in cells expressing PLCγ1 R586K. R586K abolishes the nSH2 interaction with LAT pY132. PLCγ1-null Jurkat T cells that express LAT-mCherry were reconstituted with the GFP-tagged wild type or R586K PLCγ1. They were plated on OKT3-coated cover glass. LAT microcluster formation was revealed by TIRF microscopy. Left: Images showed clustering 60 s after cell landing on the glass. Scale bar, 5 µm. Right: Quantification of clustering. Shown are mean ± SEM; n = 23–29 cells. Unpaired two-tailed t test was used. *, P < 0.05. (B) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild type or R586K PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (C) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; ***, P < 0.001.
The nSH2–pY interaction is required for PLCγ1-mediated LAT clustering and signaling. (A) Diminished LAT microcluster formation in cells expressing PLCγ1 R586K. R586K abolishes the nSH2 interaction with LAT pY132. PLCγ1-null Jurkat T cells that express LAT-mCherry were reconstituted with the GFP-tagged wild type or R586K PLCγ1. They were plated on OKT3-coated cover glass. LAT microcluster formation was revealed by TIRF microscopy. Left: Images showed clustering 60 s after cell landing on the glass. Scale bar, 5 µm. Right: Quantification of clustering. Shown are mean ± SEM; n = 23–29 cells. Unpaired two-tailed t test was used. *, P < 0.05. (B) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild type or R586K PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (C) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. *, P < 0.05; ***, P < 0.001.
The lipase-independent signaling role of PLCγ1. (A) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild type or H380F PLCγ1. H380F abolishes most of the enzymatic activity of PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (B) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. (C) Cells as indicated were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. (D) The level of LAT pY171, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments.
The lipase-independent signaling role of PLCγ1. (A) Immunoblot analysis of LAT-null Jurkat T cells reconstituted with the GFP-tagged wild type or H380F PLCγ1. H380F abolishes most of the enzymatic activity of PLCγ1. Cells were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. MW, molecular weight. (B) Quantification of the level of indicated proteins, after being normalized to the expression level of GAPDH. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. (C) Cells as indicated were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. (D) The level of LAT pY171, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments.
PLCγ1 protects LAT from dephosphorylation
The formation of LAT microclusters is phosphorylation dependent. Antigen engagement with TCR triggers the phosphorylation of LAT by the kinase ZAP70, which is antagonized by phosphatases (Su et al., 2016). LAT, once phosphorylated, can recruit SH2-containing proteins to form microclusters. Therefore, PLCγ1 has traditionally been viewed as a “passenger” that is passively recruited to phosphorylated LAT. Interestingly, we found that PLCγ1 also regulates the phosphorylation of LAT. As compared with wild-type cells, PLCγ1-null cells had significantly lower phosphorylation on Y132, but not on Y191 (Fig. 9 A) or Y171 (Fig. S4, C and D), the two binding sites for Grb2 and Gads (Zhang et al., 2000). Deleting Grb2 or Gads, the other major SH2 domain–containing proteins in the LAT complex, did not alter the phosphorylation of Y132 (Fig. 9 A). This is consistent with the fact that Y132 binds only PLCγ1 but not Grb2 or Gads. Intriguingly, overexpressing a fragment of PLCγ1 that contains the SH2 and SH3 domains significantly increased phosphorylation on Y132 (Fig. 9 A), suggesting PLCγ1 as a two-way (both up and down) regulator of Y132 phosphorylation. Because PLCγ1 is the only identified binding partner of LAT Y132, we hypothesized that the SH2 domain of PLCγ1 binds to LAT Y132, protecting it from being dephosphorylated. Supporting that, when the cellular phosphatase activity was inhibited by vanadate, the difference in phosphorylation level of Y132 between the wild-type and PLCγ1-null cells was abolished (Fig. 9 B). To directly test the hypothesis that PLCγ1 protects LAT Y132 from dephosphorylation, we set up an in vitro dephosphorylation assay. LAT-Grb2-Sos1 microclusters were assembled in the presence or absence of PLCγ1. Then CD45, the most abundant phosphatase on T cell membranes, was added to dephosphorylate LAT (Fig. 9 C). Intriguingly, PLCγ1 significantly suppressed the dephosphorylation of LAT Y132 (Fig. 9 D). Together, these data suggest that PLCγ1 specifically stabilizes the phosphorylation on LAT Y132 by protecting it from being dephosphorylated by CD45.
PLCγ1 protects LAT from dephosphorylation by CD45. (A) Reduced phosphorylation at LAT Y132 in PLCγ1-null cells. Cells as indicated were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. The level of indicated proteins, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. MW, molecular weight. (B) PLCγ1 prevents LAT Y132 from being dephosphorylated. Cells as indicated were pretreated with 0.1 mM vanadate (pan-phosphatase inhibitor) before being stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. The level of indicated proteins, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01; ns, not significant. (C) Schematics of the in intro dephosphorylation assay. (D) PLCγ1 prevents LAT Y132 from being dephosphorylated by CD45 in vitro. pLAT, at 1,000 molecules/µm2, was incubated with 1 µM Grb2, 0.5 µM Sos1, and/or 100 nM full-length PLCγ1 for 0.5 h. CD45 was then added to dephosphorylate pLAT for 5 min. The reaction was terminated by adding SDS-PAGE loading buffer with 2 mM vanadate. The level of phosphorylated LAT, after being normalized to total LAT, was quantified. Shown are mean ± SD; n = 3 independent experiments.
PLCγ1 protects LAT from dephosphorylation by CD45. (A) Reduced phosphorylation at LAT Y132 in PLCγ1-null cells. Cells as indicated were stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. The level of indicated proteins, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01. MW, molecular weight. (B) PLCγ1 prevents LAT Y132 from being dephosphorylated. Cells as indicated were pretreated with 0.1 mM vanadate (pan-phosphatase inhibitor) before being stimulated with 2 µg/ml anti-CD3 and anti-CD28 antibodies for 2 min, lysed, and applied for Western blot analysis. The level of indicated proteins, after being normalized to the level of GAPDH, was quantified. Shown are mean ± SD; n = 3 independent experiments. Unpaired two-tailed t test was used. **, P < 0.01; ns, not significant. (C) Schematics of the in intro dephosphorylation assay. (D) PLCγ1 prevents LAT Y132 from being dephosphorylated by CD45 in vitro. pLAT, at 1,000 molecules/µm2, was incubated with 1 µM Grb2, 0.5 µM Sos1, and/or 100 nM full-length PLCγ1 for 0.5 h. CD45 was then added to dephosphorylate pLAT for 5 min. The reaction was terminated by adding SDS-PAGE loading buffer with 2 mM vanadate. The level of phosphorylated LAT, after being normalized to total LAT, was quantified. Shown are mean ± SD; n = 3 independent experiments.
Discussion
PLCγ1 was traditionally considered an enzyme that acts downstream of LAT. Following TCR activation, PLCγ1 is recruited to LAT microclusters and is activated by Itk1-triggered phosphorylation. Activated PLCγ1 hydrolyzes PIP2 to generate IP3 and DAG, which triggers downstream calcium and PKC pathways, respectively. Here, we report that PLCγ1 can also promote LAT cluster formation. This is achieved in two ways: PLCγ1 cross-links LAT either with or without the assistance of Sos1; and PLCγ1 protects LAT from being dephosphorylated by CD45. Therefore, PLCγ1 and LAT reciprocally regulate each other in a positive manner.
The protection of LAT phosphorylation could be achieved by either a traditional chemical occupancy mechanism (the SH2 domain covers phosphotyrosines and limits the access of local phosphatase) and/or a spatial exclusion mechanism in which LAT clusters exclude CD45 from the clusters. Indeed, we found that the PLCγ1-mediated LAT clusters can exclude CD45 (Fig. S5, C and D), similarly to what happens for Grb2-mediated LAT clusters as we reported before (Su et al., 2016). To exclusively test if the spatial exclusion mechanism contributes to phosphotyrosine protection in LAT clusters, we need to examine a phosphotyrosine that is not recognized by the SH2 domain that mediates LAT clustering. The chemical occupancy mechanism does not apply to this phosphotyrosine, and therefore, any clustering-dependent protection effect can be attributed to the spatial exclusion mechanism. The only eligible phosphotyrosine in our system is pY132, which does not bind the SH2 domain of Grb2. Therefore, we assembled the LAT clusters in the presence of Grb2 and Sos1 and treated them with CD45. To control for the chemical occupancy in this assay, we increased the input of Grb2 in the nonclustered condition so that an equal amount of Grb2 was associated with membrane in the clustered versus nonclustered condition (Fig. S5 A). We found that pY132 is still protected by LAT clustering (Fig. S5 B), supporting the spatial exclusion mechanism.
Mechanism of phosphotyrosine protection by LAT clustering. (A) TIRF microscopy revealed clustered and unclustered LAT and Grb2. Alexa Fluor 488–labeled LAT at 1,000 molecules/µm2 was incubated with 1 µM Grb2 + 500 nM Sos1 or 6 µM Grb2. 20% of Grb2 was labeled with Alexa Fluor 568. Similar Grb2 was recruited to the membrane in the two indicated conditions. Shown are mean ± SD; n = 3 independent experiments. Scale bar, 5 µm. (B) LAT clustering by Grb2 prevents LAT Y132 from being dephosphorylated by CD45 in vitro. pLAT, at 1,000 molecules/µm2, was incubated with 1 µM Grb2, 0.5 µM Sos1, or 1 µM Grb2. CD45 was added to dephosphorylate pLAT for 5 min. The reaction was terminated by adding SDS-PAGE loading buffer with 2 mM vanadate. The level of LAT pY132, after being normalized to total LAT, was quantified. Shown are mean ± SD; n = 3 independent experiments. MW, molecular weight. (C) CD45 is excluded by Grb2- or PLCγ1-mediated LAT clustering. pLAT–Alexa Fluor 488 (1,000 molecules/μm2) was incubated with 1 µM Sos1 and 1 µM Grb2 or 1 µM PLCγ1-SH2-2-3 fragment. The cytoplasmic domain of CD45-TMR (4 nM, with an N-terminal His10 tag) was added, and its localization was visualized by TIRF microscopy. Scale bar, 5 µm. (D) Quantification of fluorescence intensity of pLAT and CD45 along the line scan indicated by a white line in the top merged image.
Mechanism of phosphotyrosine protection by LAT clustering. (A) TIRF microscopy revealed clustered and unclustered LAT and Grb2. Alexa Fluor 488–labeled LAT at 1,000 molecules/µm2 was incubated with 1 µM Grb2 + 500 nM Sos1 or 6 µM Grb2. 20% of Grb2 was labeled with Alexa Fluor 568. Similar Grb2 was recruited to the membrane in the two indicated conditions. Shown are mean ± SD; n = 3 independent experiments. Scale bar, 5 µm. (B) LAT clustering by Grb2 prevents LAT Y132 from being dephosphorylated by CD45 in vitro. pLAT, at 1,000 molecules/µm2, was incubated with 1 µM Grb2, 0.5 µM Sos1, or 1 µM Grb2. CD45 was added to dephosphorylate pLAT for 5 min. The reaction was terminated by adding SDS-PAGE loading buffer with 2 mM vanadate. The level of LAT pY132, after being normalized to total LAT, was quantified. Shown are mean ± SD; n = 3 independent experiments. MW, molecular weight. (C) CD45 is excluded by Grb2- or PLCγ1-mediated LAT clustering. pLAT–Alexa Fluor 488 (1,000 molecules/μm2) was incubated with 1 µM Sos1 and 1 µM Grb2 or 1 µM PLCγ1-SH2-2-3 fragment. The cytoplasmic domain of CD45-TMR (4 nM, with an N-terminal His10 tag) was added, and its localization was visualized by TIRF microscopy. Scale bar, 5 µm. (D) Quantification of fluorescence intensity of pLAT and CD45 along the line scan indicated by a white line in the top merged image.
Our domain truncation analysis of PLCγ1 in T cells revealed that the SH2 and SH3 domains contribute to SLP76 and ERK activation. This could be explained by two nonexclusive mechanisms: (1) the SH3 domain of PLCγ1 directly binds Sos1 (Kim et al., 2000) and recruits Sos1 to the membrane, which facilitates Ras and ERK activation; similarly, the SH3 domain directly binds and recruits SLP76 to the membrane (Yablonski et al., 2001) in preparation for being phosphorylated by ZAP70; (2) the SH2 and SH3 domains promote LAT cluster formation, which further enhances the recruitment of Grb2 and Gads to LAT. Grb2 and Gads are constitutive binding partners for Sos1 and SLP76, respectively. Therefore, the SH2 and SH3 domain of PLCγ1 can contribute to downstream signaling both directly or indirectly through LAT.
Through both biochemical reconstitution and computational approaches, we revealed a nonmonotonic mechanism of regulating LAT clustering by PLCγ1. Compositional control emerges as an important mechanism for regulating the physical features and chemical activities of liquid-like condensates (Banani et al., 2016; Ditlev et al., 2019; Riback et al., 2020). The concentration of PLCγ1 may have been tuned to control cluster size and stability, with the purpose of transducing signaling to the physiological needs. Interestingly, PLCγ1 is up-regulated in acute myeloid leukemia (Mahmud et al., 2017), colorectal carcinoma (Noh et al., 1994), and squamous cell carcinoma (Xie et al., 2010). It remains an interesting question whether PLCγ1 misregulates signaling cluster formation in these pathological conditions.
PLCγ1 is involved in a variety of membrane receptor signaling pathways. The natural killer cell and mast cell receptor (FcεR1) pathways share almost the same machinery of LAT clustering with the T cell. We reasoned that PLCγ1 could play a similar role in promoting receptor signaling by enhancing LAT clusters in natural killer and mast cells. In the pathways outside immune responses, such as EGF receptor, FGF receptor, or HER2, LAT is absent. However, similar scaffold proteins are present that contain multivalent interaction sites (e.g., EGF receptor and Shc) that can interact with PLCγ1 and Grb2-Sos1. The cooperation between PLCγ1 and Grb2 in promoting receptor clustering, as revealed in this work, could serve as a general mechanism for regulating membrane receptor signaling.
Materials and methods
Resources used (cell lines, plasmids, primers, and antibodies) are listed in Table 1.
Key resources
| Reagent type | Designation | Source or reference | Additional information |
|---|---|---|---|
| Cell line | HEK293T | University of California, San Francisco, Cell Culture Facility | Lentivirus package |
| Cell line | Jurkat T cell E6.1 | University of California, San Francisco, Cell Culture Facility | Stable expression cell line construction |
| Cell line | PLCG1 null | This study (J74B) | PLCG1 knockout by CRISPR |
| Cell line | GRB2 null | This study (J81) | GRB2 knockout by CRISPR |
| Cell line | GADS null | This study (J82) | GADS knockout by CRISPR |
| Plasmid | pMD2.G | Vale Laboratory (XSB395) | Viral packaging, plasmid |
| Plasmid | poPAX | Vale Laboratory (XSB396) | Viral packaging, plasmid |
| Plasmid | pSpCas9(BB)-PLCG1 234F-2A-GFP | This study (XSB717) | CRISPR-cas9 vector for knocking out PLCG1 |
| Plasmid | pSpCas9(BB)-Grb2 42F-2A-GFP | This study (XSB740) | CRISPR-cas9 vector for knocking out GRB2 |
| Plasmid | pSpCas9(BB)-Gads 97F-2A-GFP | This study (XSB742) | CRISPR-cas9 vector for knocking out GADS |
| Plasmid | pHR-sfGFP | This study (XSB401) | Lentiviral vector with a superfold GFP tag |
| Plasmid | pHR-PLCγ1-sfGFP | This study (XSB434) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 1–1,291) fused with an sfGFP tag |
| Plasmid | pHR-PLCγ1ΔnSH2-sfGFP | This study (XSB838) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 1–549, 658–1,291) fused with an sfGFP tag |
| Plasmid | pHR-PLCγ1ΔSH3-sfGFP | This study (XSB840) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 1–790, 852–1,291) fused with an sfGFP tag |
| Plasmid | pHR-PLCγ1 H380F-sfGFP | This study (LZB74) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 1–1,291, H380F) fused with an sfGFP tag |
| Plasmid | pHR-PLCγ1 R586K-sfGFP | This study (LZB75) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 1–1,291, R586K) fused with an sfGFP tag |
| Plasmid | pHR-PLCγ1 nSH2-cSH2-SH3-sfGFP | This study (XSB801) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: P08487, aa 550–851) fused with an sfGFP tag |
| Plasmid | pHR-LAT-mCherry | This study (XSB376) | Lentiviral vector; human LAT (UniProt accession no.: O43561-2, aa 1–233) fused with an mCherry tag |
| Plasmid | pFastBac-Bv PLCγ1-SNAPf | This study (XSB490) | Protein expression; His10-TEV-Bovine PLCγ1 (UniProt accession no.: P08487, aa 1–1,291) fusing SNAPf tag |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-cSH2-SH3 | This study (XSB406) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 550–851) |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-cSH2 | This study (LZB14) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 550–790) |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-SH3 | This study (LZB13) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 550–667, 757–851) |
| Plasmid | pGEX-ybbR-BV PLCγ1 cSH2-SH3 | This study (LZB16) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 658–851) |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2 | This study (LZB15) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 550–667) |
| Plasmid | pGEX-ybbR-BV PLCγ1 cSH2 | This study (LZB17) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: P08487, aa 658–790) |
| Plasmid | pET-His6-BV PLCγ1 SH3 | This study (LZB86) | Protein expression; His6-TEV-Bovine PLCg1 (UniProt accession no.: P08487, aa 791–851) |
| Plasmid | pET-MBP-His8-LAT 48-233-His6 | This study (XSB714) | Protein expression; MBP-TEV-His8-Human LAT (UniProt accession no.: O43561-2, aa 488–233)-TEV-His6 |
| Plasmid | pFastBac-His10-CD45 598-1304-SNAP | This study (XSB482) | Protein expression; His10-TEV-Human CD45 (UniProt accession no.: P08575, aa 598–1,304)-SNAP |
| Plasmid | pGEX-Grb2 | This study (XSB369) | Protein expression; GST-PreScission cleavage site-Human Grb2 (UniProt accession no.: P62993, aa 1–217) |
| Plasmid | pGEX-Sos1 1117-1319-Cys | This study (XSB370) | Protein expression; GST-PreScission cleavage site-Human Sos1 (UniProt accession no.: Q07889, aa 1,117–1,319)-Cys |
| Primer | pU6 F | This study (XSP130) | To verify plasmids for CRISPR knockout, 5′-GAGGGCCTATTTCCCATGATTCC-3′ |
| Primer | pHR F | This study (XSP79) | To verify plasmids for lentiviral infection, 5′-AGTCCTCCGACAGACTGAGT-3′ |
| Primer | pHR R | This study (XSP80) | To verify plasmids for lentiviral infection, 5′-CTTTCACAAATTTTGTAATCCAGAGGTTG-3′ |
| Primer | GST F | This study (XSP29) | To verify bacteria plasmids for recombinant protein production, 5′-ATGGCCTTTGCAGGGCTGG-3′ |
| Primer | PH F | This study (XSP5) | To verify baculoviral plasmids for recombinant protein production, 5′-AAATGATAACCATCTCGC-3′ |
| Primer | Fastback R | This study (XSP6) | To verify baculoviral plasmids for recombinant protein production, 5′-CAAGTAAAACCTCTACAAATGTG-3′ |
| Antibody | Anti-human CD3 | eBioscience | CD3 monoclonal antibody (OKT3), Cat#16-0037-85 |
| Antibody | Anti-human CD28 | eBioscience | Cat#16-0289-85 |
| Antibody | Anti-human pERK | Cell Signaling | Phospho-p44/42 MAPK (Erk1/2; Thr202/Tyr204), Cat#9101, Western blot (1:4,000) |
| Antibody | Anti-human PLCγ1 pY783 | Cell Signaling | Cat#2821, Western blot (1:3,000) |
| Antibody | Anti-human PLCγ1 | Cell Signaling | Cat#5690, Western blot (1:4,000) |
| Antibody | Anti-human pSLP76 | Abcam | Anti-SLP76 (phospho Y145), Cat#ab75829, Western blot (1:2,000) |
| Antibody | Anti-human LAT pY132 | Invitrogen | Cat#44-224, Western blot (1:2,000) |
| Antibody | Anti-human LAT pY171 | Cell Signaling | Cat#3581, Western blot (1:2,500) |
| Antibody | Anti-human LAT pY191 | Cell Signaling | Cat#3584, Western blot (1:2,500) |
| Antibody | Anti-human LAT | Millipore | Cat#05-770, Western blot (1:500) |
| Antibody | Anti-GAPDH | BioLegend | Cat#649202, Western blot (1:4,000) |
| Antibody | Anti-mouse IgG | Invitrogen | Goat anti-mouse IgG (H+L) secondary antibody, Cat#31430, Western blot (1:15,000) |
| Antibody | Anti-rabbit IgG | Invitrogen | Goat anti-rabbit IgG (H+L) secondary antibody, Cat#31460, Western blot (1:10,000) |
| Chemical | Vanadate | New England Biolabs | Sodium orthovanadate (Vanadate), Cat#P0758S |
| Reagent type | Designation | Source or reference | Additional information |
|---|---|---|---|
| Cell line | HEK293T | University of California, San Francisco, Cell Culture Facility | Lentivirus package |
| Cell line | Jurkat T cell E6.1 | University of California, San Francisco, Cell Culture Facility | Stable expression cell line construction |
| Cell line | PLCG1 null | This study (J74B) | PLCG1 knockout by CRISPR |
| Cell line | GRB2 null | This study (J81) | GRB2 knockout by CRISPR |
| Cell line | GADS null | This study (J82) | GADS knockout by CRISPR |
| Plasmid | pMD2.G | Vale Laboratory (XSB395) | Viral packaging, plasmid |
| Plasmid | poPAX | Vale Laboratory (XSB396) | Viral packaging, plasmid |
| Plasmid | pSpCas9(BB)-PLCG1 234F-2A-GFP | This study (XSB717) | CRISPR-cas9 vector for knocking out PLCG1 |
| Plasmid | pSpCas9(BB)-Grb2 42F-2A-GFP | This study (XSB740) | CRISPR-cas9 vector for knocking out GRB2 |
| Plasmid | pSpCas9(BB)-Gads 97F-2A-GFP | This study (XSB742) | CRISPR-cas9 vector for knocking out GADS |
| Plasmid | pHR-sfGFP | This study (XSB401) | Lentiviral vector with a superfold GFP tag |
| Plasmid | pHR-PLCγ1-sfGFP | This study (XSB434) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-PLCγ1ΔnSH2-sfGFP | This study (XSB838) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-PLCγ1ΔSH3-sfGFP | This study (XSB840) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-PLCγ1 H380F-sfGFP | This study (LZB74) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-PLCγ1 R586K-sfGFP | This study (LZB75) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-PLCγ1 nSH2-cSH2-SH3-sfGFP | This study (XSB801) | Lentiviral vector; bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pHR-LAT-mCherry | This study (XSB376) | Lentiviral vector; human LAT (UniProt accession no.: |
| Plasmid | pFastBac-Bv PLCγ1-SNAPf | This study (XSB490) | Protein expression; His10-TEV-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-cSH2-SH3 | This study (XSB406) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-cSH2 | This study (LZB14) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2-SH3 | This study (LZB13) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 cSH2-SH3 | This study (LZB16) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 nSH2 | This study (LZB15) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pGEX-ybbR-BV PLCγ1 cSH2 | This study (LZB17) | Protein expression; GST-PreScission cleavage site-ybbR tag-Bovine PLCγ1 (UniProt accession no.: |
| Plasmid | pET-His6-BV PLCγ1 SH3 | This study (LZB86) | Protein expression; His6-TEV-Bovine PLCg1 (UniProt accession no.: |
| Plasmid | pET-MBP-His8-LAT 48-233-His6 | This study (XSB714) | Protein expression; MBP-TEV-His8-Human LAT (UniProt accession no.: |
| Plasmid | pFastBac-His10-CD45 598-1304-SNAP | This study (XSB482) | Protein expression; His10-TEV-Human CD45 (UniProt accession no.: |
| Plasmid | pGEX-Grb2 | This study (XSB369) | Protein expression; GST-PreScission cleavage site-Human Grb2 (UniProt accession no.: |
| Plasmid | pGEX-Sos1 1117-1319-Cys | This study (XSB370) | Protein expression; GST-PreScission cleavage site-Human Sos1 (UniProt accession no.: |
| Primer | pU6 F | This study (XSP130) | To verify plasmids for CRISPR knockout, 5′-GAGGGCCTATTTCCCATGATTCC-3′ |
| Primer | pHR F | This study (XSP79) | To verify plasmids for lentiviral infection, 5′-AGTCCTCCGACAGACTGAGT-3′ |
| Primer | pHR R | This study (XSP80) | To verify plasmids for lentiviral infection, 5′-CTTTCACAAATTTTGTAATCCAGAGGTTG-3′ |
| Primer | GST F | This study (XSP29) | To verify bacteria plasmids for recombinant protein production, 5′-ATGGCCTTTGCAGGGCTGG-3′ |
| Primer | PH F | This study (XSP5) | To verify baculoviral plasmids for recombinant protein production, 5′-AAATGATAACCATCTCGC-3′ |
| Primer | Fastback R | This study (XSP6) | To verify baculoviral plasmids for recombinant protein production, 5′-CAAGTAAAACCTCTACAAATGTG-3′ |
| Antibody | Anti-human CD3 | eBioscience | CD3 monoclonal antibody (OKT3), Cat#16-0037-85 |
| Antibody | Anti-human CD28 | eBioscience | Cat#16-0289-85 |
| Antibody | Anti-human pERK | Cell Signaling | Phospho-p44/42 MAPK (Erk1/2; Thr202/Tyr204), Cat#9101, Western blot (1:4,000) |
| Antibody | Anti-human PLCγ1 pY783 | Cell Signaling | Cat#2821, Western blot (1:3,000) |
| Antibody | Anti-human PLCγ1 | Cell Signaling | Cat#5690, Western blot (1:4,000) |
| Antibody | Anti-human pSLP76 | Abcam | Anti-SLP76 (phospho Y145), Cat#ab75829, Western blot (1:2,000) |
| Antibody | Anti-human LAT pY132 | Invitrogen | Cat#44-224, Western blot (1:2,000) |
| Antibody | Anti-human LAT pY171 | Cell Signaling | Cat#3581, Western blot (1:2,500) |
| Antibody | Anti-human LAT pY191 | Cell Signaling | Cat#3584, Western blot (1:2,500) |
| Antibody | Anti-human LAT | Millipore | Cat#05-770, Western blot (1:500) |
| Antibody | Anti-GAPDH | BioLegend | Cat#649202, Western blot (1:4,000) |
| Antibody | Anti-mouse IgG | Invitrogen | Goat anti-mouse IgG (H+L) secondary antibody, Cat#31430, Western blot (1:15,000) |
| Antibody | Anti-rabbit IgG | Invitrogen | Goat anti-rabbit IgG (H+L) secondary antibody, Cat#31460, Western blot (1:10,000) |
| Chemical | Vanadate | New England Biolabs | Sodium orthovanadate (Vanadate), Cat#P0758S |
Recombinant proteins
The panel of recombinant proteins used in this study is shown in Fig. S1 C. Full-length PLCγ1 was purified from insect cells by a baculovirus expression system. PLCγ1 fragments containing individual or combined SH2 and SH3 domains were expressed and purified from bacteria. Peptides including each of the four LAT C-terminal phosphotyrosine residues (pY132, pY171, pY191, and pY226), PLCγ1 pY783, and each of the four Sos1 PRMs were synthesized by Peptide 2.0. The peptides are modified with biotin at the N terminus. The exact protein sequences of PLCγ1, truncations, and synthesized peptides are listed in Table S1. Gads, Nck1, Grb2, Sos1, CD45, SLP76, and LAT were purified as previously described (Su et al., 2016).
Protein purification
Full-length PLCγ1
The full-length bovine PLCγ1 with an N-terminal His10 tag, a TEV cleavage site, and a C-terminal SNAP tag was expressed in SF9 cells using the Bac-to-Bac baculoviral expression system (Life Technologies). Cells were harvested by centrifugation and lysed by Dounce homogenizer in 50 mM Hepes, pH 7.4, 300 mM NaCl, 30 mM imidazole, 5% glycerol, 5 µg/ml DNase, 0.5% Triton X-100, 0.5 mM Tris(2-carboxyethyl)phosphine (TCEP), 1 mM PMSF, and protease inhibitor cocktail. Centrifuge-cleared lysate was applied to Ni sepharose (GE Healthcare), washed with 50 mM Hepes, pH 7.4, 300 mM NaCl, 5% glycerol, 15 mM imidazole, and 0.5 mM TCEP, and eluted with the same buffer with additional 400 mM imidazole. The eluted protein was further purified by size-exclusion chromatography using a Superdex 200 prepgrade column (GE Healthcare) in 50 mM Hepes, pH 7.5, 150 mM NaCl, 10% glycerol, and 1 mM TCEP. The N-terminal His 10 tag was further cleaved off by TEV protease.
PLCγ1 nSH2-cSH2-SH3 and purification
The plasmids encoding GST-PLCγ1 nSH2-cSH2-SH3 or truncations were transformed into an Escherichia coli BL21 strain. Bacteria were grown at 18°C in LB medium with 100 µg/ml ampicillin. After cells reached a density of OD600 0.4, protein expression was induced overnight by 0.1 mM IPTG at 18°C. The culture was harvested and resuspended in PBS and lysed using a high-pressure homogenizer. The crude lysate was centrifuged at 30,000 rpm for 45 min in a Ti-70 rotor. The supernatant was supplemented with 1 mM DTT, applied to Glutathione Sepharose 4B (GE Healthcare), and washed with 50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, and 5% glycerol. The PLCγ1 fragments were cleaved off from the Sepharose by PreScission protease in cleavage buffer (50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, and 5% glycerol) at 4°C overnight. The cleaved products were pooled, concentrated, and further purified by gel filtration using a Superdex 200 size-exclusion column (GE Healthcare) equilibrated in 50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, and 10% glycerol.
Protein labeling
Full-length PLCγ1 was labeled with SNAP-Surface Alexa Fluor 647 (S9136S; NEB). Briefly, 10 µM PLCγ1 was mixed with 20 µM SNAP-Ax647 in 500 µl reaction buffer (50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, 1 mM DTT, and 10% glycerol). Following incubation in the dark for 2 h at 37°C, the labeled protein was separated from the solution by size-exclusion chromatography using PD MiniTrap G25 (GE Healthcare).
PLCγ1 nSH2-cSH2-SH3 and truncations were labeled with CoA 647 (S9350; NEB) via an N-terminal ybbR tag. 10 µM ybbR-fusion proteins were mixed with 20 µM CoA-647 and 2 µM GST-Sfp in 200 µl reaction buffer (50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, 1 mM DTT, 10 mM MgCl2, and 10% glycerol). After incubation for 2 h at room temperature, 20 µl Glutathione Sepharose 4B was added to deplete GST-Sfp. The supernatant was further applied to size-exclusion chromatography using PD MiniTrap G25 and stored in 50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, and 10% glycerol.
Biochemical reconstitution assay on supported lipid bilayers
The reconstitution of LAT microclusters was performed according to a previous protocol (Su et al., 2017) with minor modification. 1-Palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy (polyethylene glycol)-5000] (PEG-5000 PE), and 1,2-dioleoyl-sn-glycero-3-[(N-(5-amino-1-carboxypentyl)iminodiacetic acid)succinyl] (DOGS-NTA) were purchased from Avanti. A lipid mix composed of 98% POPC, 2% DOGS-NTA, and 0.1% PEG-5000 was dried with a stable flow of argon gas, followed by complete drying in a vacuum desiccator. The dried lipids were resuspended in PBS followed by 30 cycles of freezing and thawing to break down multivesicular bodies into small unilamellar vesicles (SUVs). The remaining multivesicular bodies were depleted by a centrifugation at 35,000 rpm for 45 min at 4°C using a TLA120 rotor. SUVs were stored under argon at 4°C.
The clustering assay was performed in a 96-well glass-bottom plate (MGB096-1-2-LG-L; Matriplate). The plate was washed with 5% Hellmanex III (Sigma-Aldrich) overnight and with 5 M NaOH three times, thoroughly rinsed with ultrapure water, and equilibrated with a basic buffer (50 mM Hepes, pH 7.4, 150 mM NaCl, and 1 mM TCEP). SUVs were added to each well and incubated for 1 h at 37°C to form the supported lipid bilayers. Excess SUVs were washed out afterward. The bilayers were blocked with clustering buffer (50 mM Hepes, pH 7.4, 150 mM NaCl, 1 mM TCEP, and 1 mg/ml BSA) for 30 min at 37°C. His-tagged LAT-Ax488 was incubated with the supported lipid bilayers for 2 h, and unbound LAT was washed out. Adaptor proteins (Grb2, Sos1, PLCγ1, etc.) diluted in an oxygen scavenger mix (0.2 mg/ml glucose oxidase, 0.035 mg/ml catalase, 25 mM glucose, and 70 mM β-mercaptoethanol in clustering buffer) were added to the well to form LAT clusters. Clusters were imaged by TIRF microscopy.
Cell culture and cell line construction
Jurkat T cell lines were maintained in RPMI 1640 (#11875-093; Gibco) supplemented with 10% FBS (Invitrogen) and 1% PenStrep-Glutamine in a humidified incubator with 5% CO2 at 37°C. The CRISPR-cas9 gene editing system was used to generate the PLCγ1 knockout Jurkat T cell line.
Jurkat T cell lines stably expressing a fluorescence reporter was generated by lentiviral transduction. HEK293T cells were transfected with the pHR plasmids encoding the genes of interest and the viral packaging plasmids pMD2.G and psPAX2 using Genejuice (EMD Millipore). 72 h after plasmid transfection, cell culture medium containing viral particles was harvested and added into Jurkat cells for infection in RPMI 1640 overnight. Jurkat cells expressing fluorescent proteins were sorted by FACS to generate a stable and homogeneous expression population.
Live-cell imaging of microcluster formation in Jurkat T cells
A 96-well glass-bottom imaging plate was coated with 5 µg/ml anti-CD3ε antibody (OKT3) in PBS overnight at room temperature. Unbound OKT3 was washed out the next morning. The imaging plate was equilibrated with image medium (RPMI 1640 without phenol red and 20 mM Hepes, pH 7.4). 100,000 Jurkat T cells were added to each well, and microclusters were imaged by TIRF microscopy at 37°C.
Immunoblot analysis
Jurkat T cells were washed with PBS three times, incubated at 37°C for 30 min, and stimulated with 2 µg/ml OKT3 (#16-0037-85; eBioscience) and 2 µg/ml anti-CD28 antibody (#16-0289-85; eBioscience). The reaction was stopped by directly adding 2× SDS-PAGE loading buffer (#1610737; Bio-Rad) containing protease inhibitor cocktail (#11873580001; Roche). The lysates were boiled for 10 min at 95°C and clarified by centrifugation at 12,000 rpm for 15 min at 4°C. The supernatants were loaded onto a 4–20% protein gel (#4568096; Bio-Rad) for SDS-PAGE analysis, followed by a transferring onto PVDF membrane (#1620177; Bio-Rad). The membrane was blocked with TBST buffer containing 5% nonfat milk for 1 h at room temperature and blotted with indicated primary antibodies overnight at 4°C. The next day, the membrane was further blotted with HRP-conjugated secondary antibody for 1 h at room temperature. Target proteins were detected with a chemiluminescent HRP substrate (#34577; Thermo Fisher Scientific) and visualized by a Bio-Rad ChemiDoc imaging system. Images were quantified by ImageJ.
Microscopy and image analysis
TIRF microscopy, with FRAP assay, was performed on a Nikon Ti2-E inverted motorized microscope stand, motorized stage with stagetop Piezo, LApps System with XY Miniscanner for 405-nm FRAP, Nikon H-TIRF, Agilent laser combiner with four lines, 405, 488, 561, and 640 nm, scientific CMOS camera Photometrics Prime 95B, 100× TIRF objective with an NA of 1.49, and Oko Lab temperature control system set at 37°C. Images were acquired using Nikon Elements and analyzed in Fiji (ImageJ). LAT clustering was quantified as normalized variance, which equals the square of SD divided by mean (after subtracting background). Graphs in the same panel were displayed with the same brightness and contrast settings. The half recovery time of FRAP was obtained by fitting a one-phase association model in GraphPad Prism 7.00 software.
Statistical analysis
Unless specified, unpaired two-tailed Student’s t tests were performed on data from at least three individual experiments. Data distribution was assumed to be normal, but this was not formally tested.
Computer simulations
The computational model consists of 2D particles of circular shape and diameter σ, with patches playing the role of binding sites (Fig. 6 A). All particles interact with each other through Weeks–Chandler–Anderson potential, of strength 10 kBT, kB being the Boltzmann constant and T being the temperature, representing volume exclusion. On top of that, two particles can attract each other if two interacting patches on their surfaces come close: in this case, we say that they form a bond. LAT particles have four binding sites, representing pY132, pY171, pY191, and pY226; PLCɣ1 particles have three binding sites, representing nSH2, cSH2, and the SH3 domain; Sos1 particles have four identical binding sites, representing PRMs; and Grb2 particles have three binding sites, representing the SH2 domain, nSH3, and cSH3.
2D molecular dynamics simulations are performed using LAMMPS software (Plimpton, 1995). There, patches are implemented by means of ghost particles of diameter equal to 0.05 σ and positioned at fixed distance, 0.475 σ, from the center of the proper particle. Two interacting patches (as per Fig. 6 A) interact via a cosine-squared potential with range of attractiveness 0.15 σ between the centers of the patches. The maximum depth ε of the cosine-squared potential is set to 30 kBT, rendering the bonds practically unbreakable within simulation time. Each molecule, i.e., a volume-excluded particle together with its patches representing binding sites, is treated as a rigid body. The angular distance between patches belonging to the same molecule is chosen in such a way to (1) make binding between two patches exclusive and (2) allow a Sos1 and a Grb2 molecule to bind through either one or two pairs of patches. 2D Brownian dynamics is ensured by an overdamping Langevin thermostat, enforcing room temperature at every step.
In simulations, the number of LAT molecules is fixed at 200. The ratio of LAT/Sos1/Grb2 particles is kept constant at 1:1:2, whereas the proportion PLCγ1/LAT is varied from 0 to 3. Our goal is not to capture the exact experimental value of these ratios, both because in experiments the surface density of cytosolic cross-linkers at the membrane is unknown and because it can only affect the results quantitatively, but not qualitatively. Different surface densities ρLAT are chosen for LAT, from 0.005 to 0.05 σ−2, with no noticeable difference in the mechanism of clustering (see Fig. 7 A and Simulation details). In the text, we show results for ρLAT = 0.02 σ−2.
The compactness parameter mentioned in the text is based on the analysis of the gyration tensor of each cluster, using as pole the cluster’s center of mass (the gyration tensor is equal to the inertia tensor for unit masses). The tensor is diagonalized to obtain the three principal moments of inertia: one of them, Iz, represents the inertia of in-plane rotations about the axis z, perpendicular to the plane and going through the center of mass; the remaining two principal moments I1 and I2, represent the squares of the gyration radii of the cluster along two perpendicular directions in the plane. It is possible to compute the moment of inertia Iz,min along z of an ellipse of gyration radii and made of close-packed circles (continuous bulk density for hexagonal close-packing is assumed): this represents the inertia of a fictional cluster of approximately the same shape as the real one, made of as many circular particles as can fit. The compactness parameter is then defined as the ratio Iz,min/Iz and tends to 0 for a maximally sparse cluster and 1 for a maximally dense ellipse with no void regions (see Fig. 6 B for examples and Simulation details, section SI 4).
Online supplemental material
Fig. S1 shows that both the full-length and SH fragment of PLCγ1 drive LAT clustering. Fig. S2 shows domains required for PLCγ1-driven LAT clustering. Fig. S3 shows that the nSH2–pY interaction is required for PLCγ1-mediated LAT clustering and signaling. Fig. S4 shows the lipase-independent signaling role of PLCγ1. Fig. S5 shows the mechanism of phosphotyrosine protection by LAT clustering. Video 1 shows LAT cluster formation with Grb2 and Sos1. Video 2 shows LAT cluster formation with Grb2, Sos1, and PLCγ1. Video 3 shows the early phase of simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. Video 4 shows a full-length simulation of LAT cluster formation at low PLCγ1-to-LAT ratio. Video 5 shows the early phase of simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. Video 6 shows a full-length simulation of LAT cluster formation at intermediate PLCγ1-to-LAT ratio. Video 7 shows the early phase of simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. Video 8 shows a full-length simulation of LAT cluster formation at high PLCγ1-to-LAT ratio. Simulation details provide detailed information on computer simulation of LAT microcluster assembly in the presence of PLCγ1.
Acknowledgments
We thank Christian Vanhille Campos and Johannes Krausser for providing the basis for the simulation code and for discussion on this project. Surface plasmon resonance was performed at the Yale Keck Biotechnology Resource Laboratory.
The T200 Biacore instrumentation was supported by National Institutes of Health Award S10RR026992-0110. L. Zeng is supported by a Cancer Research Institute Irvington Postdoctoral Fellowship (CRI3516). X. Su has received support from an American Cancer Society Institutional Research Grant, the Charles H. Hood Foundation Child Health Research Awards, the Andrew McDonough B+ Foundation Research Grant, the Gilead Sciences Research Scholars Program in Hematology/Oncology, the Rally Foundation and Bear Necessities Foundation—a Collaborative Pediatric Cancer Research Awards Program, the Yale University Specialized Program of Research Excellence (SPORE) in Skin Cancer DRP Award, the Yale University DeLuca Pilot Award, and the National Institutes of Health Maximizing Investigators’ Research Award (R35) program (GM138299). I. Palaia and A. Šarić received funding from the European Research Council (grant StG 802960) and the Royal Society.
The authors declare no competing financial interests.
Author contributions: L. Zeng and X. Su conceived the projects and wrote the manuscript with contributions from I. Palaia and A. Šarić. L. Zeng performed wet experiments and analyzed the data. I. Palaia and A. Šarić conceived the computer model. I. Palaia performed computer simulations and analyzed the data. X. Su and A. Šarić supervised the project.