Phospholipases cleave phospholipids, major membrane constituents. They are thus essential for many developmental processes, including male gamete development. In flowering plants, mutation of phospholipase NOT-LIKE-DAD (NLD, also known as MTL or ZmPLA1) leads to peculiar defects in sexual reproduction, notably the induction of maternal haploid embryos. Contrary to previous reports, NLD does not localize to cytosol and plasma membrane of sperm cells but to the pollen endo-plasma membrane (endo-PM), a specific membrane derived from the PM of the pollen vegetative cell that encircles the two sperm cells. After pollen tube burst, NLD localizes at the apical region of the egg apparatus. Pharmacological approaches coupled with targeted mutagenesis revealed that lipid anchoring together with electrostatic interactions are involved in the attachment of NLD to this atypical endo-PM. Membrane surface-charge and lipid biosensors indicated that phosphatidylinositol-4,5-bisphosphate is enriched in the endo-PM, uncovering a unique example of how membrane electrostatic properties can define a specific polar domain (i.e., endo-PM), which is critical for plant reproduction and gamete formation.
Introduction
Sexual reproduction represents an evolutionary success and is thus widespread among eukaryotes (Otto and Lenormand, 2002). It allows the mixing of genetic information from parents of opposite sex. Phospholipases are ubiquitous enzymes that hydrolyze phospholipids, a major component of cellular membranes, and are implicated in multiple cellular processes (Park et al., 2012; Wang, 2001; Burke and Dennis, 2009). Thus it is not surprising to find evidence that phospholipases are important for sexual reproduction in numerous organisms (Kelliher et al., 2017; Gilles et al., 2017a; Roldan and Shi, 2007; Fry et al., 1992; Barman et al., 2018). Unlike animals, a double fertilization event is required in flowering plants to achieve sexual reproduction, giving rise to the embryo and an adjacent nourishing tissue, the endosperm (Walbot and Evans, 2003; Dresselhaus et al., 2016). A second feature distinct from animal systems is that flowering plants do not have motile sperm cells: the sperm cells are passively carried to the egg apparatus (a group of three cells consisting of the egg cell and two synergids) by the pollen tube (Dresselhaus et al., 2016; Sprunck, 2020). Finally, plant sperm cells are not the direct product of meiosis but stem from two successive mitoses of the initial haploid cell (microspore; McCormick, 2004; Zhou et al., 2017a; Hackenberg and Twell, 2019). The first postmeiotic division (pollen mitosis I) is asymmetric and leads to the formation of the vegetative cell, which does not divide further, and a smaller generative cell. This generative cell goes through an additional mitosis (pollen mitosis II) to produce two genetically identical sperm cells. Sexually mature pollen (the male gametophyte) is thus composed of three haploid cells: a large vegetative cell that engulfs two sperm cells. The vegetative nucleus and the two sperm cells are connected in a stable association named the male germ unit (MGU; Fig. 1 A; Sprunck, 2020). Although small in size, the male gametophyte thus exhibits a unique topology, having cells within a cell (Dumas et al., 1984; McCue et al., 2011; Sprunck, 2020). Yet how this configuration of two small sperm cells within a large vegetative cell contributes to fertilization in flowering plant is largely unknown.
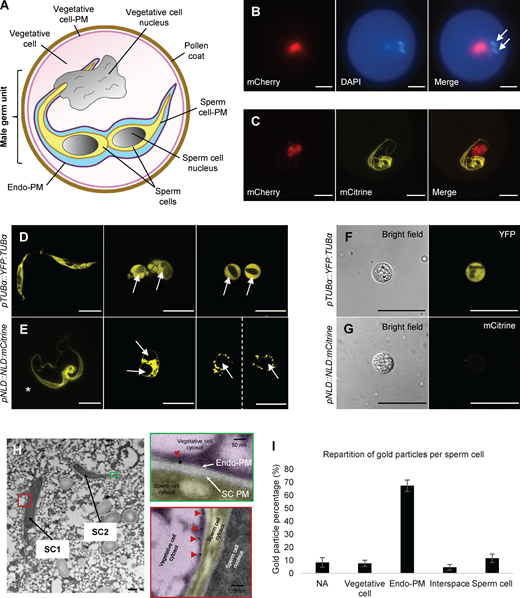
NLD is expressed in the vegetative cell of maize pollen, and protein localizes to the endo-PM of the MGU. (A) Schematic representation of the mature pollen grain organization, focusing on the MGU structure. The two sperm cells (yellow) are surrounded by an endo-PM (dark purple) originating from the PM of the vegetative cell. Light blue indicates the extracellular space (polysaccharide) of the MGU. (B) Characterization of NLD promoter activity (pNLD::H2B:mCherry) in mature pollen. Promoter activity (red signal) is found in the vegetative cell nucleus and not in the sperm cells nuclei. Nuclei are stained with DAPI (blue signal); white arrows indicate the sperm cells nuclei. (C) Subcellular localization of NLD protein (yellow signal) together with NLD promoter activity (red signal) in mature pollen. Confocal imaging (Z-stack projection) of mature pollen expressing pNLD::NLD:mCitrine and pNLD::H2B:mCherry constructs shows that NLD:mCitrine protein localizes at the MGU, whereas the promoter activity is found in the vegetative cell. (D–G) Sperm cells are released (white arrows) from mature pollen of pTUBα::YFP:TUBα (D and F) and pNLD::NLD:mCitrine (E and G) lines after osmotic shock (D and E) and sperm cells isolation using Percoll gradient (F and G). YFP:TUBα fusion protein is localized inside the sperm cells, whereas NLD:mCitrine fusion protein is not detected inside isolated sperm cells (white arrows). Asterisk in E shows an MGU before pollen burst (Z-stack projection). Scale bars in B–G = 20 µm. (H and I) Immunogold labeling of NLD:citrine protein in the MGU. (H) TEM pictures of mature pollen grain of the pNLD::NLD:mCitrine line focusing on the ultrastructure of the two sperm cells (SC1 and SC2). The red and green boxes are close-up views to visualize gold particles (red arrowheads) that recognize anti-citrine antibody and localize to the endo-PM of the MGU, and not the PM of sperm cells (SC PM). Scale bar in H = 0.5 µm. (I) Quantification of gold particles supporting preferential localization of NLD:mCitrine fusion protein at the endo-PM. Data are mean particle repartition (± SD) obtained from six different sperm cells, and quantification is detailed in Table S3. Vegetative cell, particles found within the vegetative cell; endo-PM, particles found in the endo-PM; interspace, particles found within the space between sperm cells PM and endo-PM; sperm cell, particles found in sperm cells (cytoplasm, nucleus, and sperm cells PM); NA, particles that could not be assigned to a particular compartment.
NLD is expressed in the vegetative cell of maize pollen, and protein localizes to the endo-PM of the MGU. (A) Schematic representation of the mature pollen grain organization, focusing on the MGU structure. The two sperm cells (yellow) are surrounded by an endo-PM (dark purple) originating from the PM of the vegetative cell. Light blue indicates the extracellular space (polysaccharide) of the MGU. (B) Characterization of NLD promoter activity (pNLD::H2B:mCherry) in mature pollen. Promoter activity (red signal) is found in the vegetative cell nucleus and not in the sperm cells nuclei. Nuclei are stained with DAPI (blue signal); white arrows indicate the sperm cells nuclei. (C) Subcellular localization of NLD protein (yellow signal) together with NLD promoter activity (red signal) in mature pollen. Confocal imaging (Z-stack projection) of mature pollen expressing pNLD::NLD:mCitrine and pNLD::H2B:mCherry constructs shows that NLD:mCitrine protein localizes at the MGU, whereas the promoter activity is found in the vegetative cell. (D–G) Sperm cells are released (white arrows) from mature pollen of pTUBα::YFP:TUBα (D and F) and pNLD::NLD:mCitrine (E and G) lines after osmotic shock (D and E) and sperm cells isolation using Percoll gradient (F and G). YFP:TUBα fusion protein is localized inside the sperm cells, whereas NLD:mCitrine fusion protein is not detected inside isolated sperm cells (white arrows). Asterisk in E shows an MGU before pollen burst (Z-stack projection). Scale bars in B–G = 20 µm. (H and I) Immunogold labeling of NLD:citrine protein in the MGU. (H) TEM pictures of mature pollen grain of the pNLD::NLD:mCitrine line focusing on the ultrastructure of the two sperm cells (SC1 and SC2). The red and green boxes are close-up views to visualize gold particles (red arrowheads) that recognize anti-citrine antibody and localize to the endo-PM of the MGU, and not the PM of sperm cells (SC PM). Scale bar in H = 0.5 µm. (I) Quantification of gold particles supporting preferential localization of NLD:mCitrine fusion protein at the endo-PM. Data are mean particle repartition (± SD) obtained from six different sperm cells, and quantification is detailed in Table S3. Vegetative cell, particles found within the vegetative cell; endo-PM, particles found in the endo-PM; interspace, particles found within the space between sperm cells PM and endo-PM; sperm cell, particles found in sperm cells (cytoplasm, nucleus, and sperm cells PM); NA, particles that could not be assigned to a particular compartment.
The lack of certain phospholipases impairs pollen development (Kelliher et al., 2017; Gilles et al., 2017a; Kim et al., 2011; Liu et al., 2017) and, in the case of a particular maize mutant, leads to the formation of haploid embryos with only the chromosomes from the female parent, lacking the ones from the mutant male parent (Coe, 1959; Jacquier et al., 2020). This maize mutant, named maternal haploid inducer, represents not only an interesting resource to study pollen development and plant reproduction, but also a powerful plant breeding tool (Gilles et al., 2017b; Jacquier et al., 2020, 2021). Maize breeders draw benefits from this reproductive anomaly, combining haploid induction with genome doubling of resulting haploid embryos/plantlets to produce doubled haploid plants. Doubled haploid plants are homozygous at all loci in only two generations, compared with the traditional method taking a minimum of seven generations (Kalinowska et al., 2019; Jacquier et al., 2020, 2021; Widiez, 2021). Mutational inactivation of the gene MATRILINEAL (MTL) /NOT LIKE DAD (NLD) /ZmPHOSPHOLIPASE-A (ZmPLA1) (for simplicity, NLD hereafter) was found to cause the induction of maternal haploid embryos (Kelliher et al., 2017; Gilles et al., 2017a; Liu et al., 2017). Sequence analysis revealed that NLD belongs to the patatin-like phospholipase family (Gilles et al., 2017a). The patatin catalytic domain is widely spread in bacteria, yeast, plants, and animals, and members of this group are involved in diverse cellular functions (Scherer et al., 2010; Baulande and Langlois, 2010; Wilson et al., 2006; Kienesberger et al., 2009). How NLD functions is not understood yet. NLD was reported to localize at different places of the sperm cells: either the plasma membrane (PM; Gilles et al., 2017a) or the cytoplasm (Kelliher et al., 2017). In addition, single sperm cell sequencing documented chromosome fragmentation in sperm cells of the nld mutant (Li et al., 2017), suggesting a direct or indirect effect of NLD on genome stability. The original nld mutant carries a 4-bp insertion in the last exon of NLD. This mutation causes a frameshift that replaces the last 49 amino acids of the WT protein by 20 unrelated amino acids, followed by a premature STOP codon (Gilles et al., 2017a; Kelliher et al., 2017). This C-terminal truncation is the cause of haploid embryo induction. The resulting truncated NLD protein is unstable and does not accumulate in maize pollen, whereas in the heterologous Arabidopsisthaliana (designated Arabidopsis hereafter) root cell system, it loses PM localization (Gilles et al., 2017a). These results indicate a critical role of the C-terminus of NLD, but the mechanism through which the C-terminal truncation of NLD leads to loss of protein stability and haploid induction is not known.
Here we demonstrate that NLD does not localize to sperm cells PM, nor in sperm cells cytosol, but, instead, to the endo-PM that surrounds the sperm cells. This endo-PM is produced by the adjacent vegetative cell (Sprunck, 2020). We demonstrate that the C-terminal part of the NLD protein is not sufficient by itself to ensure subcellular localization, but it comprises important amino acid residues, which, together with additional amino acid residues in the N-terminus, are key to target NLD to the endo-PM. Pharmacological and mutagenesis approaches demonstrated the involvement of both lipid anchoring and electrostatic interactions to properly address NLD to the pollen endo-PM. Imaging during the pollination process indicates the presence of NLD along the pollen tube path to the ovule, until being delivered at the apex of the egg apparatus after pollen tube rupture.
Results
NLD promoter is active in the vegetative cell of maize pollen
In sexually mature maize pollen, the vegetative cell fully encloses the two sperm cells, forming a complex cellular structure, the MGU, which regroups all the genetic material of the pollen grain (Fig. 1 A; Dumas et al., 1984; Sprunck, 2020). The sperm cells are thus “cells within a cell.” Previous studies reported the subcellular localization of the NLD protein, expressed under its own promoter, in both PM and cytoplasm of sperm cells (Gilles et al., 2017a; Kelliher et al., 2017). To refine the NLD spatial and temporal expression pattern during maize pollen development, we developed a new promoter reporter line, pNLD::H2B:mCherry. It contains the same 2.6-kbp NLD promoter fragment used in our previous study (Gilles et al., 2017a), driving the expression of two mCherry fluorescent proteins that are fused to histone H2B in order to target fluorescence to nuclei. Imaging of mature pollen grains surprisingly revealed promoter activity in the vegetative cell, and not in the sperm cells, as one would have expected based on previous observations of NLD protein localization (Fig. 1 B). This observation was confirmed using nine independent transgenic lines (Table S1). It is further supported by NLD expression data in a transcriptomic dataset from isolated maize sperm cells (Chen et al., 2017), which indicates the absence of NLD transcripts from sperm cells. Time course experiments during microgametogenesis demonstrated that the NLD promoter activity starts during the late bicellular stage, after the DNA condensation of the generative cell, just before or during the second pollen mitosis, which leads to the formation of two sperm cells after mitotic division of the generative cell (Fig. S1). The start of NLD expression at the late bicellular stage is consistent with previous observations based on both β-glucuronidase promoter activity (but lacking subcellular resolution; Gilles et al., 2017a) and RNA sequencing kinetics on whole pollen grain (Liu et al., 2017).
NLD promoter activity in the vegetative cell starts at the late bicellular stage, just before formation of the two sperm cells. (A) Calibration of anther size according to pollen development stages in A188 background. 35 anthers were measured for each stage. (B) Tassel branches with male flowers at late bicellular stage (top) when NLD promoter activity is starting, and mature pollen stage (bottom). (C and D) Temporal analysis of NLD promoter activity (pNLD::H2B:mCherry) at different stages of pollen development, between early bicellular and mature pollen stages. NLD promoter activity (red signal) starts at late bicellular stage, probably just before or during the second pollen mitosis. White arrows point at the vegetative nucleus, red arrowheads point at the generative cell nucleus, and white arrowheads point at the sperm cell nuclei. The red asterisk indicates a second pollen mitosis in progress. Determination of pollen stages by epifluorescence imaging of pollen after DAPI staining (D). Scale bars = 500 µm (B); 50 µm (C and D).
NLD promoter activity in the vegetative cell starts at the late bicellular stage, just before formation of the two sperm cells. (A) Calibration of anther size according to pollen development stages in A188 background. 35 anthers were measured for each stage. (B) Tassel branches with male flowers at late bicellular stage (top) when NLD promoter activity is starting, and mature pollen stage (bottom). (C and D) Temporal analysis of NLD promoter activity (pNLD::H2B:mCherry) at different stages of pollen development, between early bicellular and mature pollen stages. NLD promoter activity (red signal) starts at late bicellular stage, probably just before or during the second pollen mitosis. White arrows point at the vegetative nucleus, red arrowheads point at the generative cell nucleus, and white arrowheads point at the sperm cell nuclei. The red asterisk indicates a second pollen mitosis in progress. Determination of pollen stages by epifluorescence imaging of pollen after DAPI staining (D). Scale bars = 500 µm (B); 50 µm (C and D).
NLD protein localizes specifically to the endo-PM of the vegetative cell that surrounds the sperm cells
Pollen grains expressing both the pNLD::H2B:mCherry promoter fusion and the pNLD::NLD:mCitrine protein fusion underline an original situation in which the NLD promoter is active in the vegetative cell (red color in Fig. 1 C), whereas the NLD protein seems to be localized in the adjacent sperm cells (yellow color in Fig. 1 C). Complementation assays demonstrated that the NLD:mCitrine protein fusion is functional, since it fully abolished the haploid induction phenotype of the nld mutant (Table S2). Two possible explanations of this pattern were that (1) NLD transcripts (or NLD proteins) are synthesized in the vegetative cell and then move to the sperm cells, as reported for ABA-hypersensitive germination 3 (AHG3) transcript in Arabidopsis (Jiang et al., 2015), or (2) NLD protein is not localized within the sperm cells, but rather at the endo-PM of the vegetative cell, which tightly surrounds the sperm cells (Fig. 1 A; Dumas et al., 1984; Sprunck, 2020; Hafidh and Honys, 2021).
To test these hypotheses, we simplified the experimental system by disconnecting the sperm cells from the MGU including the endo-PM. Both the pNLD::NLD:mCitrine line and a sperm cell marker line used as control, for which α-tubulin-YFP (pTUBα::YFP:TUBα) had been demonstrated to be expressed in sperm cells and not in the vegetative cell (Kliwer and Dresselhaus, 2010), were subjected to osmotic shock directly on a microscope slide and imaged during the process of sperm cell release (Fig. 1, D and E). The α-tubulin-YFP line evenly marked the sperm cell cytosol and allowed clear visualization of the isolated sperm cell (Fig. 1 D). In contrast, the application of hyperosmotic shock to pNLD::NLD:mCitrine pollen gave rise to an mCitrine fluorescent signal, which first surrounded the sperm cells and was then progressively lost when sperm cells individualized (Fig. 1 E). These observations were confirmed by a complementary approach of sperm cell isolation using a Percoll gradient (Fig. 1, F and G; Dupuis et al., 1987). Sperm cells derived from the α-tubulin-YFP reporter pollen produced clearly distinguishable individualized sperm cells based on YFP fluorescence signal, whereas no fluorescent signal was detected in sperm cells derived from pNLD::NLD:mCitrine pollen. These results are in favor of the hypothesis that NLD localizes to the endo-PM of the vegetative cell. In mature pollen, the close proximity of vegetative endo-PM and sperm cell PM, the twisted nature of the MGU, and the strength of the signal likely gave the erroneous impression of localization within the sperm cells.
To fully confirm this hypothesis, immuno-EM experiments were performed on ultrathin sections (100 nm) of pollen grains containing functional pNLD::NLD:mCitrine construct. NLD was labeled using an antibody against mCitrine followed by antibody detection with a protein A–gold electron-dense probe. Nontransgenic mature pollen grains followed the same protocol as an internal control to assess background level in transmission electron microscopy (TEM; Table S3). Observation of the immuno-EM staining of transgenic (pNLD::NLD:mCitrine) and control pollen grains revealed the ultrastructure of MGU and an enrichment of gold particles in the endo-PM surrounding the sperm cells and not in the PM of the sperm cells or the cytosol of the sperm cells (Fig. 1 H). Quantification of gold particles from different pollen grains demonstrated that the NLD:mCitrine fusion protein localizes predominantly at the endo-PM, clarifying NLD subcellular localization (Fig. 1 I andTable S3).
Endo-PM NLD localization is conserved in Arabidopsis pollen
To further investigate NLD association to the endo-PM, the Arabidopsis plant model was used as a heterologous system (1) because of the absence of efficient transient expression for maize pollen and (2) to save time, since the development of transgenic maize lines takes 1 yr and the life cycle is more than twice as long as that of Arabidopsis. Since there was no clearly identified NLD orthologue in Arabidopsis (Gilles et al., 2017a), maize NLD gene was expressed in Arabidopsis pollen. To drive gene expression in the vegetative cell and not in the sperm cells within the pollen grain, similarly to the endogenous maize NLD promoter in pollen, the promoter of the POLYUBIQUITIN 10 gene (At4G05320, AtUBQ10) was chosen. Although AtUBQ10 is transcribed in nearly all plant tissues, it has been reported to show high expression in pollen grain (containing both vegetative and sperm cells), but low expression in isolated sperm cells (Borges et al., 2008). In addition, the high promoter activity in roots allowed the use of the same Arabidopsis lines to carry out confocal observations in a more conventional plant cell system without endo-PM. The expected AtUBQ10 expression pattern was experimentally validated by confocal imaging of a pUBQ10::VENUS-N7 transgenic line targeting the VENUS fluorescent protein to the nucleus (Fig. S2). In the pollen context, an expression pattern specific to the vegetative cell was demonstrated, and no fluorescence was detected in sperm cells (Fig. S2 A), similarly to what had been found for the NLD promoter in maize pollen (Fig. 1 B). In addition, the promoter was indeed found to be active in root cells (Fig. S2 B), as reported earlier (Geldner et al., 2009; Grefen et al., 2010).
Validation of the pUBQ10 promoter expression pattern in Arabidopsis pollen. (A and B) Confocal imaging of mature pollen (A) and root tip cells (B) of the pUBQ10::Venus-N7 line (yellow signal). DAPI stains nuclei (blue signal), white arrowheads indicate the sperm cells nuclei, and red arrow points at the vegetative nucleus. Scale bars = 20 µm.
Validation of the pUBQ10 promoter expression pattern in Arabidopsis pollen. (A and B) Confocal imaging of mature pollen (A) and root tip cells (B) of the pUBQ10::Venus-N7 line (yellow signal). DAPI stains nuclei (blue signal), white arrowheads indicate the sperm cells nuclei, and red arrow points at the vegetative nucleus. Scale bars = 20 µm.
Previous work in the heterologous Arabidopsis root expression system had shown that NLD full-length protein (NLDfull; Fig. 2 A) localizes predominantly to the PM, but also to intracellular compartments and cytosol, whereas the truncated NLD protein (NLDtrunc; Fig. 2 A), which is responsible for the haploid induction phenotype, is not recruited at the PM (Gilles et al., 2017a). We confirmed these previous results in root cells (Fig. 2, B–D) and quantified them by measuring the ratio of fluorescence found at the PM versus fluorescence found in the cytosol (PM/cyto fluorescence ratio; Fig. 2 L). Whereas NLDfull:mCitrine had a PM/cyto fluorescence ratio of 2.39, indicating PM enrichment over cytosol, NLDtrunc:mCitrine had a low PM/cyto fluorescence ratio of 0.71, illustrating that NLDtrunc:mCitrine loses PM localization. In Arabidopsis pollen grains, NLDfull:mCitrine displays a strong fluorescent signal at the MGU and was not detected at the delimiting PM of the vegetative cell (Fig. 2 E). This signal likely corresponds to the endo-PM, as for maize. The truncation of NLD (NLDtrunc:mCitrine) provoked loss of this subcellular localization pattern, with fluorescent signal found only in the cytosol and in intracellular compartments of the vegetative cell (Fig. 2 F). Of note, the weaker fluorescent signal surrounding the pollen grain was likely due to autofluorescence from the pollen coat, since similar autofluorescence was observed in pollen grain of the nontransgenic Columbia-0 control line (Fig. 2 G). Taken together, these observations suggest that NLD protein subcellular localization is highly conserved between maize and Arabidopsis pollen and that NLD truncation leads to a loss of membrane localization to both the PM in roots and the endo-PM in pollen, suggesting a role of the NLD C-terminus in protein subcellular localization.
NLD protein localization is conserved in Arabidopsis pollen, and the C-terminal part of the NLD protein is not sufficient to ensure subcellular localization. (A) Schematic representation of NLDfull and NLDtrunc proteins. Since the NLD C-terminal region contains both a predicted lipid anchor posttranslational modification site (C423) and a stretch of positively charged amino acids (in red in the sequence), targeting of NLD to membrane could be achieved via lipid anchoring and/or electrostatic interactions between the anionic lipids present at the membrane surface and the cationic amino acid present at the NLD C-terminus. (B–K) Confocal imaging of different forms of NLD protein in Arabidopsis root cells and pollen. Confocal imaging of root tips (B–D, H, and I) and mature pollen grains (E–G, J, and K) from nontransformed Columbia 0 (Col 0; D and G) and lines expressing NLDfull:mCitrine (B and E), NLDtrunc:mCitrine (C and F), the C-terminal part of NLDfull (mCitrine:NLDfull-Cter; H and J) fused to mCitrine, and the C-terminal part of NLDtrunc (mCitrine:NLDtrunc-Cter; I and K) fused to mCitrine. Scale bars = 20 µm. (L) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio) in root cells (mean ± SD). ***, significant difference between means as compared with NLDfull:mCitrine (P < 0.001; Wilcoxon test, n = cell number analyzed).
NLD protein localization is conserved in Arabidopsis pollen, and the C-terminal part of the NLD protein is not sufficient to ensure subcellular localization. (A) Schematic representation of NLDfull and NLDtrunc proteins. Since the NLD C-terminal region contains both a predicted lipid anchor posttranslational modification site (C423) and a stretch of positively charged amino acids (in red in the sequence), targeting of NLD to membrane could be achieved via lipid anchoring and/or electrostatic interactions between the anionic lipids present at the membrane surface and the cationic amino acid present at the NLD C-terminus. (B–K) Confocal imaging of different forms of NLD protein in Arabidopsis root cells and pollen. Confocal imaging of root tips (B–D, H, and I) and mature pollen grains (E–G, J, and K) from nontransformed Columbia 0 (Col 0; D and G) and lines expressing NLDfull:mCitrine (B and E), NLDtrunc:mCitrine (C and F), the C-terminal part of NLDfull (mCitrine:NLDfull-Cter; H and J) fused to mCitrine, and the C-terminal part of NLDtrunc (mCitrine:NLDtrunc-Cter; I and K) fused to mCitrine. Scale bars = 20 µm. (L) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio) in root cells (mean ± SD). ***, significant difference between means as compared with NLDfull:mCitrine (P < 0.001; Wilcoxon test, n = cell number analyzed).
The C-terminal part of the NLD protein is not sufficient to ensure subcellular localization
The reversible addition of a fatty acid with 16 carbon atoms (palmitate) to a protein is called S-palmitoylation or, more generally, S-acylation. This posttranslational modification is emerging as a ubiquitous mechanism to control membrane affinity, trafficking, maturation, and function of proteins (Chamberlain and Shipston, 2015; Hurst and Hemsley, 2015). Two cysteine residues (C10 and C423) of the NLD protein sequence were predicted in silico to be S-acylated (Fig. 2 A; Gilles et al., 2017a). In addition, a stretch of positively charged amino acid residues (arginine [R]), was found in proximity to the predicted C423 lipid anchor site (Fig. 2 A). These in silico observations fed the working hypothesis that NLD was specifically addressed to the endo-PM via a lipid anchor and/or electrostatic interactions between positively charged amino acids of NLD and negatively charged membrane lipids (Fig. 2 A). Similar protein/membrane interactions have been demonstrated to operate in other cellular contexts in plants (Hurst and Hemsley, 2015; Alassimone et al., 2016; Simon et al., 2016; Platre et al., 2018; Barbosa et al., 2016) and animals (Chamberlain and Shipston, 2015; Zhou et al., 2017b; Liu et al., 2007; He et al., 2007). Since NLDtrunc loses membrane localization (PM in root cells and endo-PM in pollen), and since all amino acids concerned in this hypothesis are located within the C-terminus of NLDfull (absent in NLDtrunc) except the residue C10, we investigated whether the C-terminus of NLDfull is sufficient to ensure membrane localization. The NLD sequence corresponding to the last 49 amino acids that are absent in NLDtrunc was fused to mCitrine (mCitrine:NLDfull-Cter), as was the unrelated sequence of 20 amino acids replacing it in NLDtrunc in a second fusion (mCitrine:NLDtrunc-Cter). Confocal imaging of pollen grains of both lines showed fluorescent signal in the cytosol and nucleus of the vegetative cell, but absence of fluorescence in sperm cells (Fig. 2, J and K). Cytosolic fluorescence was also observed in root cells (Fig. 2, H and I), and the PM/cyto fluorescence ratios (0.66 and 0.68, respectively) substantiated quantitatively cytosolic enrichment as compared with PM (Fig. 2 L). These observations suggest that the C-terminus of NLDfull is not sufficient to address the mCitrine fluorescent protein to membranes, indicating that other regions of NLD contribute to its PM and endo-PM localization in root cells and pollen grains, respectively. The presence of eight positively charged amino acids in the unrelated C-terminus of NLDtrunc (Fig. 2 A) further underlined that other features than solely positive charged amino acids are required for proper NLD localization.
Membrane localization of NLD protein is mediated by S-acylation
Multiple reports, in both animals and plants, show that a lipid anchor allows membrane attachment for proteins that do not contain transmembrane domains (Chamberlain and Shipston, 2015; Hurst and Hemsley, 2015). Thus, the involvement of the two predicted lipid anchor sites, C10 and C423 (Fig. 2 A), was first examined by a pharmacological approach. 2-Bromopalmitate (2-BP), an inhibitor of S-acyl transferase that catalyzes the S-acylation reaction (Webb et al., 2000), was used to treat 7-d-old Arabidopsis seedlings expressing NLDfull:mCitrine (Fig. 3). Confocal imaging of root cells after a 2-h treatment with 50 µM 2-BP revealed a partial removal of NLDfull:mCitrine from the PM (Fig. 3 B) as compared with mock treatment (Fig. 3 A). Fluorescent intensity quantification confirmed this observation: the PM/cyto fluorescence ratio significantly decreased in root cells after 2-BP treatment as compared with mock treatment (2.79 to 0.99; Fig. 3 E). To ensure that the 2-BP treatment had no major effects on PM integrity, similar 2-BP treatment was applied to seedlings expressing the PM-localized GFP:LTI6b construct, where the fusion protein is inserted in the PM by two transmembrane domains (Fig. 3, C and D; Platre et al., 2018; Cutler et al., 2000). The 2-BP treatment had no effect on GFP:LTI6b subcellular localization, the signal being maintained at the PM (Fig. 3, C, D and F). Pollen grains expressing NLDfull:mCitrine in the vegetative cell were also treated with 2-BP. Similar treatment as done for root cells (50 µM for 2 h) did not affect NLD localization in pollen, most likely because the protective pollen coat, which forms an impermeable barrier, partially impedes drug penetration. Increasing the 2-BP concentration to 150 µM resulted in significantly fewer pollen grains having NLDfull:mCitrine localized to the endo-PM (13.4% in 2-BP–treated versus 80.8% in mock-treated pollen), and more pollen grains were found to have either spotty or cytosolic NLDfull:mCitrine signal as compared with the mock treated pollen grains (Fig. S3). Altogether, these results demonstrate the involvement (direct or indirect) of S-acylation in addressing NLD to the PM in root cells and to the endo-PM in pollen.
Lipid anchors are necessary for NLD targeting at membranes. (A–F) Pharmacological approach showing the involvement of S-palmitoylation in NLD addressing to the PM of root cells. Confocal imaging of root cells expressing NLDfull:mCitrine (A and B) or transmembrane GFP:LTI6B control (C and D) after 2-h treatment with 50 µM 2-BP, an inhibitor of palmitoylation (B and D) or mock treatment (EtOH; A and C). (E and F) Quantification of drug treatment effect on NLD subcellular localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio) in root cells (mean ± SD). (G–L) Targeted mutagenesis of predicted NLD lipid anchor sites C10 and C423. Confocal imaging of root cells and mature pollen from the following lines: NLDC10S:mCitrine (C10 mutated to S; G and H), NLDC423S:mCitrine (I and J), and NLDC10S,423S:mCitrine (K and L). (M and N) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol in root cells (PM/cyto fluorescence ratio; M) and in pollen grain (endo-PM/cyto fluorescence ratio; mean ± SD; N). ***, significant difference as compared with NLDfull:mCitrine (P < 0.001; Wilcoxon test, n = cell number analyzed). Scale bars = 20 µm.
Lipid anchors are necessary for NLD targeting at membranes. (A–F) Pharmacological approach showing the involvement of S-palmitoylation in NLD addressing to the PM of root cells. Confocal imaging of root cells expressing NLDfull:mCitrine (A and B) or transmembrane GFP:LTI6B control (C and D) after 2-h treatment with 50 µM 2-BP, an inhibitor of palmitoylation (B and D) or mock treatment (EtOH; A and C). (E and F) Quantification of drug treatment effect on NLD subcellular localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio) in root cells (mean ± SD). (G–L) Targeted mutagenesis of predicted NLD lipid anchor sites C10 and C423. Confocal imaging of root cells and mature pollen from the following lines: NLDC10S:mCitrine (C10 mutated to S; G and H), NLDC423S:mCitrine (I and J), and NLDC10S,423S:mCitrine (K and L). (M and N) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol in root cells (PM/cyto fluorescence ratio; M) and in pollen grain (endo-PM/cyto fluorescence ratio; mean ± SD; N). ***, significant difference as compared with NLDfull:mCitrine (P < 0.001; Wilcoxon test, n = cell number analyzed). Scale bars = 20 µm.
Pharmacological approach showing the involvement of S-palmitoylation in NLD targeting the endo-PM in Arabidopsis pollen. (A–D) Confocal imaging of mature pollen expressing NLDfull:mCitrine after 2-h treatment with mock (EtOH; A) or with 150 µM 2-BP (B–D). After mock treatment, NLDfull:mCitrine mainly localizes at the endo-PM (A), whereas 2-BP treatment leads to three different types of localization: endo-PM (B), noncontinuous (“spotty”) fluorescence signal at endo-PM (C), and low fluorescent signal in cytosol (D). Scale bars = 20 µm. (E) Quantification of drug treatment effect on NLD subcellular localization by counting the three different classes of localization patterns after 2-BP treatment. Data were obtained from five independent experiments (mean ± SD). ***, significant difference as compared with the mock treatment (P < 0.001; χ2 test, n = total number of counted mature pollens). (F and G) Verification of transgene expression for the three protein fusions harboring mutation on C10 or/and C423 residues (NLDC10S:mCitrine, NLDC423S:mCitrine, and NLDC10S,C423S:mCitrine; G) together with the two fusions (NLDfull:mCitrine and NLDtrunc:mCitrine; F). RT-PCR was done on one or two selected lines for each construct, and three independent biological replicates were used per line. Minus RT samples were RT reactions conducted without RT enzyme to evaluate possible gDNA contaminations in RNA extractions.
Pharmacological approach showing the involvement of S-palmitoylation in NLD targeting the endo-PM in Arabidopsis pollen. (A–D) Confocal imaging of mature pollen expressing NLDfull:mCitrine after 2-h treatment with mock (EtOH; A) or with 150 µM 2-BP (B–D). After mock treatment, NLDfull:mCitrine mainly localizes at the endo-PM (A), whereas 2-BP treatment leads to three different types of localization: endo-PM (B), noncontinuous (“spotty”) fluorescence signal at endo-PM (C), and low fluorescent signal in cytosol (D). Scale bars = 20 µm. (E) Quantification of drug treatment effect on NLD subcellular localization by counting the three different classes of localization patterns after 2-BP treatment. Data were obtained from five independent experiments (mean ± SD). ***, significant difference as compared with the mock treatment (P < 0.001; χ2 test, n = total number of counted mature pollens). (F and G) Verification of transgene expression for the three protein fusions harboring mutation on C10 or/and C423 residues (NLDC10S:mCitrine, NLDC423S:mCitrine, and NLDC10S,C423S:mCitrine; G) together with the two fusions (NLDfull:mCitrine and NLDtrunc:mCitrine; F). RT-PCR was done on one or two selected lines for each construct, and three independent biological replicates were used per line. Minus RT samples were RT reactions conducted without RT enzyme to evaluate possible gDNA contaminations in RNA extractions.
To further support NLD membrane targeting by S-acylation of cysteine residues, targeted mutagenesis was performed on C10 and C423 residues (Fig. 2 A). The C residues were separately or simultaneously substituted with serine (S) residues, leading to the following protein fusions: NLDC10S:mCitrine, NLDC423S:mCitrine, and NLDC10S,C423S:mCitrine (Fig. 3, G–N). Confocal imaging of root cells revealed drastic relocalization of NLDC10S:mCitrine from the PM to the cytosol and intracellular compartments, with a significant decrease of the PM/cyto fluorescent ratio from 2.39 for NLDfull:mCitrine to 0.57 for NLDC10S:mCitrine (Fig. 3, G and M). In contrast, the NLDC423S:mCitrine fusion protein remained accumulated at the PM, with no significant change in the PM/cyto fluorescent ratio as compared with NLDfull:mCitrine (2.25 versus 2.39, respectively; Fig. 3, I and M). The simultaneous mutation of both C10 and C423 resulted in a loss of mCitrine fluorescent signal (Fig. 3, K and M) in 15 independent lines (Table S1), although the transgene was expressed (Fig. S3, F and G). These results underline the importance of these two cysteine residues in promoting NLD membrane localization, which is likely important to stabilize the protein based on evidence gained in other biological contexts (Valdez-Taubas and Pelham, 2005; Abrami et al., 2008; Chamberlain and Shipston, 2015; Hurst and Hemsley, 2015). Similar conclusions on the importance of both C10 and C423 for correct NLD localization and stability could be drawn based on observations of these three mutated versions in pollen (Fig. 3, H, J, L, and N). Interestingly, NLDC10S:mCitrine showed only partial localization of fluorescent signal at the pollen endo-PM (Fig. 3 H), whereas it was almost fully removed from PM in root cells (Fig. 3 G), indicating that NLD is probably more strongly attached to the pollen endo-PM than the root cell PM. Altogether, both pharmacological and targeted mutagenesis approaches demonstrated the importance of a lipid-anchoring mechanism to stabilize NLD at membranes: at the PM in root cells and specifically at the endo-PM in pollen grain.
NLD targeting to membranes involves electrostatic interactions
If S-acylation is often essential for anchoring proteins within lipid bilayers (membranes), the specific attachment of proteins to particular membranes can be fine-tuned through electrostatic interactions with lipids (Simon et al., 2016; Platre et al., 2018; Zhou et al., 2017b). To address the importance of membrane electrostatics for the subcellular localization of NLD, the electronegativity of the membrane surface in cells was indirectly decreased by phenylarsine oxide (PAO) treatment. PAO is an inhibitor of PI4-kinases (PI4Ks), involved in the biosynthesis of highly electronegative PI4P phospholipids, which are predominantly present on the PM surface in root cells (Simon et al., 2016, 2014). Confocal imaging of root cells from NLDfull:mCitrine seedlings after 30 min of PAO treatment at 60 µM showed a clear relocalization of the fluorescent signal from PM to cytosol and endomembrane compartments (Fig. 4, A and B), although low fluorescent signal was maintained at PM. Quantification of PM/cyto fluorescence ratio corroborated a significant reduction of this ratio in PAO-treated cells (0.98), as compared with mock treatment (2.39; Fig. 4 I). The effectiveness of short-term PAO treatment was established for control lines expressing either a “PI4P biosensor” (mCitrine:1x-PHFAPPI) or a surface charge biosensor (mCitrine:FARN6+; Simon et al., 2016, 2014). After PAO treatment, a strong PI4P biosensor delocalization and a partial charge biosensor delocalization were observed in root cells (Fig. 4, D and F) as compared with mock treatment (Fig. 4, C and E). In addition, the absence of a negative effect of short-term PAO treatment on PM integrity was established using GFP:LTI6b seedlings (Fig. 4, G, H, and J). In pollen, PAO treatment also led to the relocalization of NLD from the endo-PM but necessitated longer incubation time (1 h) and higher concentration (180 µM), again likely due to the protective effect of the pollen coat. Significantly fewer pollen grains had NLDfull:mCitrine localized to the endo-PM (12.4% in PAO-treated versus 80.4% in mock-treated pollen grains), and more pollen grains were found to have either spotty or cytosolic NLDfull:mCitrine signal as compared with the mock treated pollen grains (Fig. S4). Altogether, this pharmacological approach confirmed the importance of membrane electronegativity for correct NLD targeting.
Electrostatic interactions between NLD and negatively charged membranes are necessary for correct targeting. (A–H) Pharmacological approach showing the importance of membrane electronegativity in targeting NLD to PM of root cells. Confocal imaging of root cells from lines expressing NLDfull:mCitrine (A and B), the PI4P biosensor mCitrine:1XPHFAPPI (C and D), the biosensor of surface membrane charge mCitrine:FARN6+ (E and F) or transmembrane GFP:LTI6b (G and H) after 30-min treatment with 60 µM PAO, an inhibitor of PI4Ks, or mock treatment (DMSO; A, C, E, and G). (I and J) Quantification of drug treatment effect in root cells on NLD subcellular localization (I) and GFP:LTI6b (J) using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio; mean ± SD). (K–N) Targeted mutagenesis of the nine positively charged amino acids located in the NLD C-terminus impaired NLD subcellular localization. Confocal imaging of root cells and mature pollen from lines NLD9R→9K:Citrine (9 R residues mutated to K residues; K and L), or NLD9R→9Q:Citrine (9 R residues mutated to Q residues; M and N). (O and P) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol in root cells (PM/cyto fluorescence ratio; O) and in pollen grain (endo-PM/cyto fluorescence ratio; mean ± SD; P). ***, significant difference as compared with the mock treatment (I) or as compared with NLDfull:mCitrine (O and P; P < 0.001; Wilcoxon test, n = cell number analyzed). Scale bars = 20 µm.
Electrostatic interactions between NLD and negatively charged membranes are necessary for correct targeting. (A–H) Pharmacological approach showing the importance of membrane electronegativity in targeting NLD to PM of root cells. Confocal imaging of root cells from lines expressing NLDfull:mCitrine (A and B), the PI4P biosensor mCitrine:1XPHFAPPI (C and D), the biosensor of surface membrane charge mCitrine:FARN6+ (E and F) or transmembrane GFP:LTI6b (G and H) after 30-min treatment with 60 µM PAO, an inhibitor of PI4Ks, or mock treatment (DMSO; A, C, E, and G). (I and J) Quantification of drug treatment effect in root cells on NLD subcellular localization (I) and GFP:LTI6b (J) using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol (PM/cyto fluorescence ratio; mean ± SD). (K–N) Targeted mutagenesis of the nine positively charged amino acids located in the NLD C-terminus impaired NLD subcellular localization. Confocal imaging of root cells and mature pollen from lines NLD9R→9K:Citrine (9 R residues mutated to K residues; K and L), or NLD9R→9Q:Citrine (9 R residues mutated to Q residues; M and N). (O and P) Quantification of mutagenesis effect on localization using the ratio of the fluorescence intensity at PM compared with the fluorescence intensity in cytosol in root cells (PM/cyto fluorescence ratio; O) and in pollen grain (endo-PM/cyto fluorescence ratio; mean ± SD; P). ***, significant difference as compared with the mock treatment (I) or as compared with NLDfull:mCitrine (O and P; P < 0.001; Wilcoxon test, n = cell number analyzed). Scale bars = 20 µm.
Pharmacological approach showing the involvement of electrostatic interactions between NLD and endo-PM in pollen. (A–E) Confocal imaging of mature pollen from line expressing NLDfull:mCitrine after 1-h treatment with mock (DMSO; A) or with 180 µM PAO (B–D), an inhibitor of PI4Ks. After DMSO treatment, NLDfull:mCitrine localized to the endo-PM (A and E), whereas PAO treatment led to three different types of localization: endo-PM (B), noncontinuous (“spotty”) fluorescence signal at endo-PM (C), and low fluorescent signal in cytosol (D). Scale bars = 20 µm. (E) Quantification of drug treatment effect on NLD subcellular localization by counting the three different classes of localization patterns after PAO treatment. Data were obtained from five independent experiments (mean ± SD). ***, significant difference as compared with the mock treatment (P < 0,001; χ2 test, n = total number of counted mature pollens).
Pharmacological approach showing the involvement of electrostatic interactions between NLD and endo-PM in pollen. (A–E) Confocal imaging of mature pollen from line expressing NLDfull:mCitrine after 1-h treatment with mock (DMSO; A) or with 180 µM PAO (B–D), an inhibitor of PI4Ks. After DMSO treatment, NLDfull:mCitrine localized to the endo-PM (A and E), whereas PAO treatment led to three different types of localization: endo-PM (B), noncontinuous (“spotty”) fluorescence signal at endo-PM (C), and low fluorescent signal in cytosol (D). Scale bars = 20 µm. (E) Quantification of drug treatment effect on NLD subcellular localization by counting the three different classes of localization patterns after PAO treatment. Data were obtained from five independent experiments (mean ± SD). ***, significant difference as compared with the mock treatment (P < 0,001; χ2 test, n = total number of counted mature pollens).
Mechanistically, positively charged parts of NLD could interact with negative membrane surface charges. Since the loss of NLDtrunc membrane localization correlated with the loss of a stretch of positively charged amino acid residues in the missing NLD C-terminus (Fig. 2, C and F), targeted mutagenesis was performed to assess the role of this stretch in addressing NLD to membranes. In the resulting NLD9R→9Q:mCitrine line, the nine positively charged R residues were substituted, in NLDfull, by nine neutral glutamine (Q) residues. Confocal imaging of NLD9R→9Q:mCitrine root cells showed mainly cytosolic fluorescence (Fig. 4 M), with a significant relocalization from PM to cytosol as compared with nonmutated NLDfull:mCitrine root cells (PM/cyto fluorescence ratio of 0.56 versus 2.39, respectively; Fig. 4 O). In pollen grains, a similar effect was observed, with a drastic loss of endo-PM localization accompanied by a significant increase of fluorescent signal in cytosol for NLD9R→9Q:mCitrine as compared with NLDfull:mCitrine (Fig. 4, N and P). In a control experiment, the nine R residues were replaced by nine lysine (K) residues. This conservative change maintaining the positive charges gave rise to the NLD9R→9K:mCitrine line. In pollen, NLD9R→9K:mCitrine still localized to the endo-PM as NLDfull:mCitrine, indicating that maintaining the same number of positively charged amino acids on NLD C-terminus was sufficient to maintain NLD at the endo-PM (Fig. 4, L and P). Unexpectedly, NLD9R→9K:mCitrine was not targeted to the PM in root cells and rather localized in cytosol and endomembrane compartments (Fig. 4, K and O). Although there is a trend of the different NLD mutated protein fusions to be more “sticky” to the endo-PM as compared with the root cell PM (Fig. 2, B and E, Fig. 3, G and H, and Fig. 3, I and J), the situation for NLD9R→9K:mCitrine could be due to the fact that the mutations of the nine amino acids impaired more drastically protein conformation and/or interactions with other proteins in root cells as compared with the vegetative cell. Altogether, both the pharmacological and the targeted mutagenesis approaches demonstrated the implication of electrostatic interactions between NLD and negatively charged membranes to correctly target NLD, especially to pollen endo-PM.
The endo-PM of the pollen vegetative cell is enriched for PI(4,5)P2 lipid sensor
Although the above results on lipid anchoring and electrostatic interactions provided insights into the type of interactions between NLD and membranes, they did not answer the question why NLD was targeted specifically to the endo-PM and not to both endo-PM and vegetative cell PM. To investigate the lipid composition of the poorly characterized endo-PM, we took advantage of a lipid biosensor collection that had been shown to recognize phosphatidic acid (PA), phosphatidylserine (PS), phosphatidylinositol 3-phosphate (PI3P), phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2), or phosphatidylinositol 4-phosphate (PI4P) lipids in Arabidopsis root cells and other cellular contexts (Simon et al., 2014, 2016; Platre et al., 2018). All biosensors presented the advantage to be expressed under the pUBQ10 promoter that confers expression in the vegetative cell but not in the sperm cells in the pollen grain (Fig. S2 A). Four (PA, PS, PI4P, and charge) of the six tested biosensors localized to both PM and endo-PM (Fig. 5, A, B, E, and F). The PIP3 biosensor localized to intracellular compartments of the vegetative cell (Fig. 5 C). This pattern is similar to the one reported for the root cell type, where these intracellular compartments have been identified as late endosomes (Simon et al., 2014, 2016; Platre et al., 2018). Interestingly, the PI(4,5)P2 sensor showed strong enrichment at the endo-PM as compared with the PM (Fig. 5 D). This enrichment of PI(4,5)P2 at the endo-PM, which could also be seen as a depletion of PI(4,5)P2 at the PM, is an appealing candidate to represent a signature that allows distinction between the PM and the endo-PM in the vegetative cell context.
Characterization of endo-PM properties using genetically encoded fluorescent lipid sensors. (A–E) Localization of different lipids in the vegetative cell of Arabidopsis mature pollen through lipid biosensors. Confocal imaging allowed visualization of different lipid biosensors (Simon et al., 2014; Platre et al., 2018) expressed in the vegetative cell of pollen under the pUBQ10 promoter. PA, PS, PI3P, Pi(4,5)P2, and PI4P. (F) Confocal imaging of Arabidopsis mature pollen showing subcellular localization of a charge biosensor (KA1Kcc4p) which consists in a folded domain that lacks stereospecificity and associates nonspecifically with anionic lipids (Simon et al., 2016). (G) Quantification of fluorescence intensity for biosensors at the endo-PM as compared with the fluorescence intensity at PM of vegetative cell (endo-PM/PM fluorescence ratio; mean ± SD). n = number of mature pollens analyzed. All scale bars = 20 µm.
Characterization of endo-PM properties using genetically encoded fluorescent lipid sensors. (A–E) Localization of different lipids in the vegetative cell of Arabidopsis mature pollen through lipid biosensors. Confocal imaging allowed visualization of different lipid biosensors (Simon et al., 2014; Platre et al., 2018) expressed in the vegetative cell of pollen under the pUBQ10 promoter. PA, PS, PI3P, Pi(4,5)P2, and PI4P. (F) Confocal imaging of Arabidopsis mature pollen showing subcellular localization of a charge biosensor (KA1Kcc4p) which consists in a folded domain that lacks stereospecificity and associates nonspecifically with anionic lipids (Simon et al., 2016). (G) Quantification of fluorescence intensity for biosensors at the endo-PM as compared with the fluorescence intensity at PM of vegetative cell (endo-PM/PM fluorescence ratio; mean ± SD). n = number of mature pollens analyzed. All scale bars = 20 µm.
Destiny of the endo-PM during fertilization
To track this poorly characterized endo-PM during pollen tube growth to the ovule, WT maize female reproductive tissues that received transgenic pollen expressing pNLD::NLD:mCitrine were imaged (Figs. 6 and S5 and Videos 1, 2, 3, 4, 5, 6, 7, 8, and 9). Due to deep embedding of maize embryo sac within maternal tissues, the recently improved ClearSeeAlpha clearing method compatible with fluorescent reporter imaging was applied (Kurihara et al., 2021). During pollen tube elongation in maize silk tissues (transmitting tract), confocal imaging showed NLD:mCitrine signal surrounding both sperm cells (Fig. 6 A and Video 1), consistent with previous visualization of NLD using in vitro pollen tube germination (Gilles et al., 2017a; Kelliher et al., 2017). NLD:mCitrine fluorescent signal was also clearly visible before pollen tube discharge in the embryo sac (Fig. 6 B and Video 2). After pollen rupture, which could be monitored thanks to autofluorescence of the degenerating synergid (Leydon et al., 2015; Figs. 6 D and S5 A, Video 5, and Video 6), NLD:mCitrine signal was observed within the embryo sac (Fig. 6 C), consistent with previous observations (Kelliher et al., 2017). Z-stacks and 3D projections showed the position of NLD:mCitrine signal at the apical region of the egg apparatus (Fig. 6 C; Fig. S5, B and C; Video 3; Video 4; Video 7; and Video 8), which was shown to be the place of sperm cells discharge in Arabidopsis (Hamamura et al., 2011; Sprunck, 2020). One may speculate that the dotty nature of the signal observed on the top of the egg apparatus and degenerating synergid (most clearly visible on Video 8) could indicate fragmentation of the endo-PM to allow sperm cells liberation, as suggested by Sprunck (2020). Consistent with this hypothesis, later observations (44 h after pollination) of fertilized embryo sacs failed to identify NLD:mCitrine signal (Fig. S5 D and Video 9), indicating loss of the endo-PM. A diagram summarizing the localization of the pollen endo-PM within embryo sac is depicted in Fig. 6 E.
Destiny of endo-PM during maize fertilization. (A–D) Tracking of NLD:mCitrine marking the endo-PM during pollen tube growth to the maize ovule. Confocal imaging (Z-projections) after pollination of WT female plants with NLD:mCitrine pollen (A–C) or control cross with WT instead of transgenic pollen (D). Calcofluor-white (CFW), mCitrine, and autofluorescence (AF) channels are used to visualize the cell walls, NLD:mCitrine, and autofluorescence, respectively. (A) NLD:mCitrine location during pollen tube elongation inside of the silk (SI) tissues (pollen tube transmitting tract). Arrows show the NLD:mCitrine signal. Sperm cells (SC) could be identified (white lines). Video 1 shows the Z-stack. (B) NLD:mCitrine location in a pollen tube before pollen tube discharge into a synergid cell. Arrows show the NLD:mCitrine signal. Video 2 shows the Z-stack. (C) NLD:mCitrine location within the embryo sac, after pollen tube discharge into a synergid cell. After pollen tube discharge (monitored thanks to the autofluorescence of the degenerating synergid cell, see D), NLD:mCitrine protein localizes at the apical region of the egg cell apparatus (arrow). The arrowhead represents NLD:mCitrine signal from another arriving pollen tube. Videos 3 and 4 show Z-stack and 3D projection, respectively. (D) Control pollination with WT ovule receiving WT pollen. It allows evaluation of autofluorescence (in both mCitrine and AF channels) of the degenerating synergid (DS) without NLD:mCitrine signal. Video 5 shows Z-stack. (E) Schematic illustration of endo-PM localization within the embryo sac after pollen rupture. AP, antipodal cells; CC, central cell; DS, degenerating synergid; EA, egg apparatus cell (egg cell or intact synergid); ES, embryo sac; I, integument; Nc, nuclei central cell; NU, nucellus; OV, ovary tissue. Scale bars = 50 µm.
Destiny of endo-PM during maize fertilization. (A–D) Tracking of NLD:mCitrine marking the endo-PM during pollen tube growth to the maize ovule. Confocal imaging (Z-projections) after pollination of WT female plants with NLD:mCitrine pollen (A–C) or control cross with WT instead of transgenic pollen (D). Calcofluor-white (CFW), mCitrine, and autofluorescence (AF) channels are used to visualize the cell walls, NLD:mCitrine, and autofluorescence, respectively. (A) NLD:mCitrine location during pollen tube elongation inside of the silk (SI) tissues (pollen tube transmitting tract). Arrows show the NLD:mCitrine signal. Sperm cells (SC) could be identified (white lines). Video 1 shows the Z-stack. (B) NLD:mCitrine location in a pollen tube before pollen tube discharge into a synergid cell. Arrows show the NLD:mCitrine signal. Video 2 shows the Z-stack. (C) NLD:mCitrine location within the embryo sac, after pollen tube discharge into a synergid cell. After pollen tube discharge (monitored thanks to the autofluorescence of the degenerating synergid cell, see D), NLD:mCitrine protein localizes at the apical region of the egg cell apparatus (arrow). The arrowhead represents NLD:mCitrine signal from another arriving pollen tube. Videos 3 and 4 show Z-stack and 3D projection, respectively. (D) Control pollination with WT ovule receiving WT pollen. It allows evaluation of autofluorescence (in both mCitrine and AF channels) of the degenerating synergid (DS) without NLD:mCitrine signal. Video 5 shows Z-stack. (E) Schematic illustration of endo-PM localization within the embryo sac after pollen rupture. AP, antipodal cells; CC, central cell; DS, degenerating synergid; EA, egg apparatus cell (egg cell or intact synergid); ES, embryo sac; I, integument; Nc, nuclei central cell; NU, nucellus; OV, ovary tissue. Scale bars = 50 µm.
Destiny of endo-PM during maize fertilization. Tracking of NLD:mCitrine marking the endo-PM during pollen tube growth to the maize ovule by confocal imaging (Z-projections). Calcofluor-white (CFW), mCitrine, and auto-fluorescence (AF) channels are used to visualize the cell walls, NLD:mCitrine, and autofluorescence, respectively. (A) Control pollination of a WT ovule with WT pollen. It allows evaluation of autofluorescence of the degenerating synergid (DS) without NLD:mCitrine signal. Video 6 shows the Z-stack. (B and C) NLD:mCitrine location within the embryo sac, after pollen tube rupture (which could be monitored thanks to the autofluorescence of the degenerating synergid cell). After pollen tube rupture, the NLD:mCitrine protein locates at the apical region of the egg cell apparatus. Arrows show the NLD:mCitrine signal specifically detected in the mCitrine channel. Arrowhead in B represents NLD:mCitrine signal from another arriving pollen tube. Videos 7 and 8 show Z-stack and 3D projection, respectively. (D) WT embryo sac observed 44 h after pollination with NLD:mCitrine pollen; no mCitrine signal was detected. Video 9 shows 3D projection. ES, embryo sac; NU, nucellus; OV, ovary tissue. Scale bars = 50 µm.
Destiny of endo-PM during maize fertilization. Tracking of NLD:mCitrine marking the endo-PM during pollen tube growth to the maize ovule by confocal imaging (Z-projections). Calcofluor-white (CFW), mCitrine, and auto-fluorescence (AF) channels are used to visualize the cell walls, NLD:mCitrine, and autofluorescence, respectively. (A) Control pollination of a WT ovule with WT pollen. It allows evaluation of autofluorescence of the degenerating synergid (DS) without NLD:mCitrine signal. Video 6 shows the Z-stack. (B and C) NLD:mCitrine location within the embryo sac, after pollen tube rupture (which could be monitored thanks to the autofluorescence of the degenerating synergid cell). After pollen tube rupture, the NLD:mCitrine protein locates at the apical region of the egg cell apparatus. Arrows show the NLD:mCitrine signal specifically detected in the mCitrine channel. Arrowhead in B represents NLD:mCitrine signal from another arriving pollen tube. Videos 7 and 8 show Z-stack and 3D projection, respectively. (D) WT embryo sac observed 44 h after pollination with NLD:mCitrine pollen; no mCitrine signal was detected. Video 9 shows 3D projection. ES, embryo sac; NU, nucellus; OV, ovary tissue. Scale bars = 50 µm.
Z-stack of NLD:mCitrine observation during pollen tube elongation inside of the silk (from Fig. 6 A). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine observation during pollen tube elongation inside of the silk (from Fig. 6 A). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine observation in a pollen tube before pollen tube discharge into a synergid cell (from Fig. 6 B). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine observation in a pollen tube before pollen tube discharge into a synergid cell (from Fig. 6 B). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture (from Fig. 6 C). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture (from Fig. 6 C). Scale bar = 50 µm. Frame rate = 2 fps.
3D projection of Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture (from Fig. 6 C). Arrow shows NLD:mCitrine signal (endo-PM). Scale bar = 50 µm. Frame rate = 2 fps.
3D projection of Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture (from Fig. 6 C). Arrow shows NLD:mCitrine signal (endo-PM). Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of the embryo sac pollinated with WT pollen, from Fig. 6 D. Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of the embryo sac pollinated with WT pollen, from Fig. 6 D. Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of the embryo sac pollinated with WT pollen, from Fig. S5 A. Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of the embryo sac pollinated with WT pollen, from Fig. S5 A. Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture, from Fig. S5 B. Scale bar = 50 µm. Frame rate = 2 fps.
Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture, from Fig. S5 B. Scale bar = 50 µm. Frame rate = 2 fps.
3D projection of Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture, from Fig. S5 B. Arrow shows NLD:mCitrine signal (endo-PM). Scale bar = 50 µm. Frame rate = 1.75 fps.
3D projection of Z-stack of NLD:mCitrine location within the embryo sac, after pollen tube rupture, from Fig. S5 B. Arrow shows NLD:mCitrine signal (endo-PM). Scale bar = 50 µm. Frame rate = 1.75 fps.
3D projection of Z-stack of 44-h fertilized ovule, from Fig. S5 D. Scale bar = 50 µm. Frame rate = 2.5 fps.
3D projection of Z-stack of 44-h fertilized ovule, from Fig. S5 D. Scale bar = 50 µm. Frame rate = 2.5 fps.
Discussion
Distinct lipid signature of the pollen endo-PM
There is rapidly growing evidence for an important role of spatiotemporal phospholipid patterns in plant development at the cell, tissue, and organ level (Colin and Jaillais, 2020). Here we establish such a pattern for the endo-PM, which encircles the germ cells (generative cell or sperm cells) in pollen, based on the specific subcellular localization of the phospholipase NLD to this membrane and the demonstration of PI(4,5)P2 enrichment (Figs. 1 and 5). The endo-PM is extremely poorly characterized, and even often omitted from textbooks. To our knowledge, only three proteins, two small GTP-binding rho of plants (PsROP1 and AtROP9) and la and related protein 6c (AtLARP6c), had previously been demonstrated to have endo-PM localization (Lin et al., 1996; Li et al., 2013; Billey et al., 2021). Among these endo-PM–localized proteins, NLD is the only one to be specifically localized to the endo-PM, because both ROPs and AtLARP6c were also reported to localize in other places in the pollen tube and/or pollen grain (Lin et al., 1996; Li et al., 2013; Billey et al., 2021), which is not the case for NLD (this study; Gilles et al., 2017a; Kelliher et al., 2017). In this study, the importance of two putatively S-acylated cysteine residues was demonstrated for proper NLD localization to the endo-PM (Fig. 3). Interestingly, for AtROP9, three cysteine residues were also shown to be critical for proper endo-PM localization (Li et al., 2013), pinpointing the implication of lipid anchors to attach nontransmembrane proteins to the endo-PM. In addition, the contribution of electrostatic interactions between the endo-PM and NLD was shown to be necessary but not sufficient to correctly target NLD to the endo-PM, based on pharmacological and targeted mutagenesis evidence (Figs. 2 and 4).
It was stunning to not observe NLD targeting to the pollen or pollen tube PM, indicating an efficient targeting mechanism to the endo-PM rather than PM. The observed enrichment of PI(4,5)P2 at the endo-PM presumably plays a role in this NLD targeting, and the use of recently engineered tools such as the inducible depletion of PI(4,5)P2 will help to test this hypothesis (Doumane et al., 2021). A first element in favor of this hypothesis comes from observations in root cells, where NLD is found at the PM just like the PI(4,5)P2 sensor (Simon et al., 2014). It is thus probably not a coincidence that the NLD:mCitrine and PI(4,5)P2 sensors show very similar localization in two quite different cell types (root cell and vegetative cell). These observations may also explain why NLD is found in the PM of root cells, whereas it localizes at the endo-PM in the pollen vegetative cell.
Parallels between pollen endo-PM and other invaginated PM
During infection of plant cells by biotrophic pathogens, a feeding structure, called a haustorium, forms inside the plant host cell. Consequently, an extrahaustorial membrane (EHM) derived from the host PM surrounds the haustorium (Hückelhoven and Panstruga, 2011; Gil and Gay, 1977; Roberts et al., 1993). Thus, a parallel can be drawn between this EHM in infected cells and the endo-PM in pollen, because both enclose structures within the cell and have a PM origin. The EHM was found to differ in its molecular composition from the conventional PM (Hückelhoven and Panstruga, 2011; Gil and Gay, 1977; Roberts et al., 1993; Micali et al., 2011; Koh et al., 2005). Remarkably, two recent studies reported that the EHM is enriched in PI(4,5)P2 (Shimada et al., 2019; Qin et al., 2020), as described in this study for the pollen endo-PM. An analogous situation is found in the case of arbuscular mycorrhiza symbiosis, in which the fungal hyphae, located inside the plant cell, are entirely enveloped by a plant membrane, the periarbuscular membrane (PAM; Parniske, 2008). Interestingly, strong accumulation of a PI(4,5)P2 reporter was found at discrete regions of the PAM, together with specific targeting of certain proteins to this PAM (Pumplin et al., 2012; Liu et al., 2020). Altogether, these examples represent three distinct systems in which a membrane derived from the plant cell PM forms a special “internal” PM that has different enrichments in terms of lipids and proteins as compared with the “normal” PM surrounding the cell, PI(4,5)P2 enrichment being a common signature.
Roles of pollen endo-PM in haploid embryo induction and fertilization
Contrary to previous reports, we demonstrate that NLD is not localized inside but outside of the sperm cells. Indeed, the discovery of NLD localization on the endo-PM refines initial findings by our own group (Gilles et al., 2017a) and another laboratory (Kelliher et al., 2017) and raises a new question about the instructive role of the vegetative cell in reproduction. This revision of NLD localization is an essential step forward to correctly understand the biological mechanism underlying NLD function, knowing that the absence of NLD in pollen (male) causes the induction of haploid embryos with solely the maternal (female) genome in the process of in planta haploid induction (Jacquier et al., 2020). In fact, the cellular insights of this study raise the challenge to explain mechanistically the astonishing fact that the loss of NLD outside the sperm cells (on endo-PM) leads to a drastic phenotype on chromosome stability within sperm cells (Li et al., 2017), together with double fertilization defects (Jacquier et al., 2020, 2021). The revised NLD localization lends support to the hypothesis regarding the importance of communications between the vegetative cell and the enclosed generative or sperm cells (McCue et al., 2011; Sprunck, 2020) during plant reproduction. Future work on the pollen endo-PM needs to track the genesis of this special interface during male gamete differentiation and investigate its functional significance, in particular the dependence on local membrane properties of a unique cell–cell communication route between the vegetative pollen cell and the enclosed sperm cells.
A time course of the fluorescence generated by NLD:mCitrine allowed tracking of endo-PM behavior during the fertilization process within the female embryo sac (Figs. 6 and S5). After pollen tube rupture, NLD:mCitrine localizes at the apical region of the egg apparatus. Although the lack of a compatible female fluorescent marker in maize did not allow attribution of the signal to either the egg cell or the persistent synergid, the endo-PM definitely localized at the interface between the egg apparatus and the central cell, the strategic position of sperm cells discharge and double fertilization (Hamamura et al., 2011; Sprunck, 2020). One may speculate that the particular properties in charge and/or composition of the endo-PM might facilitate its positioning in the appropriate apoplastic space between egg cell and central cell, thanks to interactions with specific cell wall components. It would thus allow or facilitate the correct delivering of the two precious sperm cells at the right place. Overall, this study reveals new insights on the pollen endo-PM that will help to understand how paternal genome integrity is maintained during gamete development and how double fertilization is properly coordinated.
Materials and methods
Plant material and plant growth conditions
Maize (Zea mays) plants were cultivated in equipment that fulfill the French S2 safety standards for the culture of transgenic plants: either in an S2 greenhouse (16-h illumination period at 24/19°C [day/night] and without control of relative humidity; Rousseau et al., 2015) or in S2 growth chambers (16-h light and 8-h darkness at 26/18°C [day/night] and relative humidity controlled to 55/65% [day/night]; Doll et al., 2020, 2019). Nontransgenic plants were also cultivated in a field located in Lyon. Haploid induction rate was evaluated using a hybrid female tester line (F564 × DH7) homozygous for the glossy1 mutation. The lines to be tested for haploid induction were crossed as male parent, and haploid induction rate was determined as the percentage of glossy plantlets among germinated plantlets (Gilles et al., 2017a). Arabidopsis plants were grown in soil under long-day conditions at 21°C and 70% humidity. The plant in vitro experiments were done on Murashige and Skoog (MS) basal medium supplemented with 0.8% plant agar (pH 5.7) in continuous light conditions at 21°C for 6–9 d. All lines generated in this study and used are presented in Table S1. For generation and growth of all transgenic plants, we followed national guidelines and legislation.
Plant transformation and selection
Maize transformation was performed in inbred line A188 (Gerdes and Tracy, 1993). Agrobacterium-mediated transformation was executed according to a published protocol (Ishida et al., 1996), and transfer DNA integrity was checked (Gilles et al., 2017a). Arabidopsis ecotype Columbia-0 was transformed using the floral-dip method, which consists of incubating the flowers in a bacterial solution (Clough and Bent, 1998), and a modified procedure for Agrobacterium preparation (Logemann et al., 2006). T1 plants were selected in vitro on the appropriate antibiotic/herbicide, and construction integrity was double-checked by genotyping with specific primers (Table S4). Approximately 10 independent T1 plants were transplanted in soil. In the T2 generation, at least two independent lines were selected according to the following criteria: a single insertion of the transgene (Mendelian segregation on selection agent), no obvious abnormal developmental phenotypes, good expression of the transgene for detection by confocal imaging, fluorescence clearly visible, and a uniform expression pattern.
Construct generation for transgenic lines
Plasmids used for plant transformations were generated using the multisite gateway three-fragment strategy (Life Technologies), according to the recommendations of the supplier, or by site-directed mutagenesis using PCR amplification of preexisting plasmids with specific primers (Table S4 and Table S5). DNA fragments compatible with entry vectors were generated by PCR amplification of the fragments of interest from genomic DNA, with specific primers compatible with Gateway technology, or synthesized by Genewiz (Table S5). Amplifications were done with the high-fidelity enzyme Phusion Hot Start II (Thermo Fisher Scientific). PCR products were purified using NucleoSpin kit (Macherey-Nagel). Plasmids generated, and the corresponding methods, are presented in Table S5.
Pollen observations
Arabidopsis fresh pollen was obtained after crushing of young open flowers in liquid solution (water or specific treatments) between blade and coverslip. Fresh maize mature pollen was collected and observed between slide and coverslip in appropriate medium to conserve pollen grain integrity (10% sucrose [wt/vol], 5 mM CaCl2, 1 mM MgSO4, 5 mM KCl, and 0.01% H3BO4 [wt/vol], pH 8 with KOH). Nuclear coloration was done using 1 µg/ml DAPI solution (wt/vol) diluted in citrate-phosphate buffer (0.1 M C6H8O7, 0.2 M Na2HPO4 2H2O, pH 4 adjustment and addition of 0.1% Triton X-100 [vol/vol]). The kinetics of maize pollen development was calibrated on A188 background (Fig. S1 A). Anthers were collected at different developmental stages and classified according to their size and position on the tassel. Anthers were imaged with a binocular loupe to measure length and then were crushed in DAPI solution between blade and coverslip to evaluate pollen stage development by epifluorescence microscopy (nuclei observation). After calibration, the same experiment was done with the NLD reporter line for temporal analysis of promotor activity.
Maize sperm cells isolation
Sperm cells were isolated either by osmotic shock or by use of a Percoll gradient, whenever a better purity was needed. For sperm cells isolation by osmotic shock, fresh pollen was placed between slide and coverslip at room temperature in shock medium containing sucrose and low ion concentration (10% sucrose [wt/vol], 5 µM CaCl2, 1 µM MgSO4, 5 µM KCl, and 0.01% H3BO4 [wt/vol], without pH adjustment). For sperm cells isolation by Percoll gradient, 500 mg of fresh pollen was collected in a 15-ml tube containing 6 ml of 1× BK medium (100× BK: 0.1% H2BO3 [wt/vol], 20 mM Ca(NO3)2, 20 mM MgSO4, and 10 mM KNO3) coupled with 0.52 M mannitol to burst pollen grains. Tubes were vortexed quickly and then gently agitated on a rotary agitator for 20 min (4°C). After this step, all processes were done on ice. To remove large debris, the solution was filtered (40 and 70 µm), and the filters were rinsed with an equivalent volume of 80% Percoll in BK-S15-MOPS medium (1× BK medium, 30% sucrose [wt/vol], and 20 mM MOPS (3-(N-Morpholino)propane sulfonic acid), pH 7.5 with NaOH [wt/vol]). Sperm cells purification was done using 5 ml of filtrate in a new 15-ml tube. Above this volume, 1 ml of 20% Percoll in BK-S15-MOPS medium (wt/vol) was gently deposited, then 5 ml of BK-Man-MOPS medium (1× BK medium, 0.52 M mannitol, and 10 mM MOPS, pH 7.5 with NaOH) was added. Tubes were centrifuged for 40 min at 1,400 rpm (4°C). After this step, two bands containing sperm cells were formed and were gently removed and individually deposited in new tubes. Six equivalent volumes of BK-Man-MOPS medium were added, and tubes were centrifuged for 30 min at 1,200 rpm (4°C). Supernatants were mostly removed to leave 500 µl of solution in the tube, and the pellets were resuspended in this volume. This suspension containing isolated sperm cells was observed between slide and coverslip.
Maize embryo sac imaging
WT ovules from the inbred A188 cultivated in growth chambers were harvested either 18–23 h after pollination to have access to the pollen tube rupture time point or 42–46 h to have ovules with fertilized embryo sacs. Ovules were individually removed from the ear and longitudinally dissected with a razor blade (sliced on both sides of the silk) to obtain ∼1-mm-width slices. Those slices were immediately fixed in 4% PFA by vacuum infiltration on ice (−0.06 mPA, vacuum broken 5 times every 15 min) and incubated in a new 4% PFA solution overnight at 4°C. Samples were then washed three times during 1 min with 1% PBS and transferred into a ClearSeeAlpha solution to make ovules transparent according to recently published protocol (Kurihara et al., 2021). This clearing step was done at room temperature during 1–3 wk (baths renewed 2–3 times per week), and samples were stored at 4°C in clearing solution until imaging (imaging occurred 9–55 d after the beginning of clearing). Samples were stained with 0.1% (vol/vol) Calcofluor-white (Ursache et al., 2018) and observed using a Zeiss AxioImager M2 microscope with a dry 10× objective (EC Plan –NEOFLUAR 10×/0.3; Zeiss) to preselect slices and determine which side of the slice was the closest to the embryo sac. Selected slices were mounted in ClearSee medium and surrounded by a rectangle of vacuum grease, which allowed adjustments of the mounting system to the different samples. The slices were then observed with a water-immersion objective (HC FLUOTAR L 25×/0.95 W; Leica) on a Leica SP8 laser scanning confocal upright microscope (DM6000) and using LAS X software v3.5.7.23225. Three channels were sequentially acquired for the Calcofluor-white signal, the mCitrine signal, and the autofluorescence, with respective excitation at 405, 514, and 552 nm and detection at 413–504, 520–600, and 564–660 nm, respectively. Laser intensities and gain of the detectors were adjusted independently to collect the best signal. The images were then processed with Fiji (Schindelin et al., 2012) to adjust brightness and contrast, and to create maximum-intensity projections as well as 3D reconstitution. For some videos, a bleaching correction was applied on the Z-stack on the Calcofluor-white channel only to correct, and therefore equalize, the signal throughout the Z-stack.
Microscopy observations
Maize pollen development analyses were performed on epifluorescence microscope, AxioImager M2 (Zeiss), harboring a lamp LED X-Cite 120LED (Excelitas). Objectives Plan-Apochromat 20×/0.8 (Zeiss) and Plan-Apochromat 40×/0.95 Korr (Zeiss) were used, and images were acquired on a Zeiss camera Axiocam 503 color with the Zen software (Zen 2, v2.0.0.0; Zeiss). DAPI staining and mCherry signal were acquired with fluorescence filter sets excG365-emLP420 and excBP587/25-emBP647/70, respectively. Confocal imaging of maize pollen marked with both mCitrine and mCherry fluorescent reporters was done at 20–22°C on a Leica SP8 upright confocal microscope, with a water-immersion objective (HCX IRAPO L 25×/0.95 W), and using LAS X software v3.5.7.23225. Fluorophores were excited using diode lasers (Leica Microsystems) emitting at wavelengths of 514 nm for mCitrine and 552 nm for mCherry. Images were collected at 521–550 nm for mCitrine and 610–650 nm for mCherry. Root cell and pollen grain observations for both Arabidopsis and maize were done on an inverted Zeiss LSM710 confocal microscope mounted on AxioObserver. Samples were observed using a 40× oil objective (Plan-Apochromat 40×/1.4 oil differential interference contrast M27; Zeiss). Dual-color images were acquired by sequential line switching, allowing the separation of channels by both excitation and emission. Depending on the fluorophores observed, different wavelengths of excitation and bandpass filters were used: mCitrine, YFP, VENUS (514/520–580 nm; excitation/bandpass), and DAPI (405/410–480 nm; excitation/band pass). Immunogold labeling imaging was done with TEM: Philips CM120 at 120 kV using a charge-coupled device camera, Gatan Orius 200.
Immunogold labeling
Pollen was fixed in 2% PFA/0.2% glutaraldehyde (vol/vol) in 0.1 M phosphate buffer (PB), pH 7.4, with 3% sucrose (wt/vol) under vacuum for 2 h at room temperature and overnight at 4°C. The next day, pollen was rinsed three times with 0.1 M PB with 3% sucrose, washed twice in 50 mM glycine in 0.1 M PB, pH 7.4, with 3% sucrose, embedded in 12% gelatin in 0.1 M PB with 3% sucrose, and placed at 4°C until solid. Small cubes (∼1 mm3) were cut and placed in 2.3 M sucrose in 0.1 M PB, pH 7.4, overnight at 4°C. For cutting, the samples were put on an aluminum pin and snap frozen in liquid nitrogen, and ultrathin sections (100 nm) were cut with a cryo-ultramicrotome (Leica UC S; Leica Microsystems) at −100°C with a glass knife. Sections were picked up with a mixture of 2% methylcellulose/2.3 M sucrose (vol/vol), collected on Formvar-carbon–coated copper grids (Electron Microscopy Sciences), and immunostained using standard protocols. Briefly, sections were treated with 50 mM glycine for aldehyde quenching, and epitopes were blocked with 0.1 M PBS/1% Cold Water Fish Skin Gelatin/0.2% BSA with 3% sucrose, incubated with the primary antibody anti-GFP (1:250; A-6455; Thermo Fisher Scientific), and washed. The reaction was revealed using protein-A-gold (10 nm, 1:70; Cell Microscopy Core, University of Utrecht).
PAO and 2-BP drug treatments
Stock solutions of 60 mM PAO in DMSO and 50 mM 2-BP in EtOH were ordered from Sigma-Aldrich. 7-d-old seedlings grown in vitro vertically (root tip cells analyses) and fresh open flowers (pollen grains analyses) were incubated in the dark and under agitation in liquid 1/2 MS medium complemented with appropriate drug treatment: for roots, 60 µM PAO for 30 min or 50 µM 2-BP for 2 h; for open flowers, 180 µM PAO for 1 h or 50 µM 2-BP for 2 h; for pollen treatment, flowers were wounded with a pincer to facilitate the drug penetration, and after treatment they were crushed on the blades to release the pollen grains. The mock conditions correspond to tissue incubation in liquid 1/2 MS medium supplemented with a volume of DMSO and EtOH, respectively, equivalent to the PAO and 2-BP volumes used and for the same time as the actual treatment. The effect of drug treatment was evaluated using confocal image analysis for roots. For pollen, the number of pollen grains from different classes defined according to the subcellular localization of the protein fusion were counted directly under the microscope. Three classes were observed: endo-PM (continuous fluorescent signal at the endo-PM), spotty (noncontinuous fluorescent signal at the endo-PM), and cytosolic (low fluorescent signal in the cytosol). For the pollen analyses, a minimum of five independent treatments were done at the same time.
RNA extractions and RT-PCR
For RNA extractions, newly opened flowers of Arabidopsis were harvested and snap frozen in liquid nitrogen. For each line, three biological replicates were done, each replicate containing a pool of ∼30 flowers of 3–4 different plants. Tissues were ground to powder thanks to Qiagen TissueLyser II. Total RNA was extracted as previously described (Creff et al., 2019). Briefly, the total RNA was extracted using the Spectrum Plant Total RNA Kit (Sigma-Aldrich), and DNA removed thanks to the TURBO DNA-free Kit (Invitrogen) according to the manufacturer’s instructions. RNA was transcribed using a SuperScript IV VILO Kit (Invitrogen) according to the suppliers’ instructions. RT–PCR was performed using the two primers listed in Table S4, which give an expected product size of 823 bp for all NLD:mCitrine constructs, except for NLDtrunc:mCitrine, in which the truncation leads to PCR amplification of 736 bp.
Imaging quantification
Image analyses were performed using Fiji (Schindelin et al., 2012). Maize anthers were measured using the measure function of Fiji. For Arabidopsis confocal imaging, different ratios were defined to characterize the subcellular localization of proteins according to the repartition of the fluorescent signal intensity in cells (Platre et al., 2018). Fluorescent intensity of a region of interest in pictures was evaluated by the measure of the mean gray value of pixels using the mean gray value function of Fiji. The PM/cyto fluorescence ratio was measured on the root cell tips and corresponds to the ratio between the intensity of the fluorescent signal at the level of the plasma membrane (PM) compared with the intensity of the fluorescent signal at the level of the cytosol (cyto). Per cell, two linear region-of-interest measurements were done (one per cell side) at the level of the PM and at the level of the cytosol, using the Measure function of Fiji, and the mean of measurements was taken. For the transgenic line analyses, the measurements were performed on ≥5 independent lines with the measurement of 20 cells per root tip on ≥3 different roots per line. For the pharmacological analyses, the measurements were performed in 3 independent experiments, with the measurement of 20 cells per root tips on ≥5 roots per experiment. The fluorescence ratio endo-PM/PM, measured in mature pollen, corresponds to a ratio between the intensity of fluorescent signal at the level of the endo-PM compared with the intensity of the fluorescent signal at the PM level. All pictures for the same experiment were acquired in same microscope setup. For the Immunogold labeling quantifications, full sperm cells pictures were reconstituted using Adobe Photoshop software. Gold particles were then quantified in three independent experiments in a triple-blind setup by three independent operators (Table S3).
Statistical analyses
Statistical analyses were performed with RStudio software. Comparisons of the fluorescence intensity between different samples were performed using nonparametric Wilcoxon–Mann–Whitney test. Comparisons of two different proportions were performed using the χ2 test.
Online supplemental material
Fig. S1 shows NLD promoter activity during pollen development. Fig. S2 shows atUBQ10 promoter expression pattern in Arabidopsis pollen. Fig. S3 shows 2-BP drug effects on pollen grains as well as verification of transgene expression for constructs expressing a mutated version of NLD. Fig. S4 shows PAO drug effects on pollen grains. Fig. S5 shows localization of NLD:mCitrine within a maize embryo sac, complementary to Fig. 6. Table S1 lists maize and Arabidopsis transgenic lines used. Table S2 shows the complementation of maize haploid induction phenotype by pNLD::NLD:mCitrine. Table S3 details quantification of Immunogold labeling of NLD:mCitrine in maize pollen grains. Table S4 lists all primers used in this study. Table S5 details the plasmids used. Videos 1, 2, 3, 4, 5, 6, 7, 8, and 9 are Z-stacks or 3D reconstitutions of some images presented in Figs. 6 and S5.
Acknowledgments
We thank M. Simon, Y. Jaillais, and M. Platre for discussions and comments; G. Brunoud (Laboratoire Reproduction et Développement des Plantes, Lyon, France), T. Dresselhaus (University of Regensburg, Regensburg, Germany), and Y. Jaillais (Laboratoire Reproduction et Développement des Plantes, Lyon, France) for providing transgenic lines; and O. Hamant and Y. Jaillais for critical reading of the manuscript. We are grateful to G. Gendrot, M.-F. Gérentes, and J. Laplaige for maize transformation; J. Berger, P. Bolland, and A. Lacroix for maize culture; I. Desbouchages and H. Leyral for buffer and media preparation; J. Guichard and S. Boeuf for technical assistance; and F. Rozier for help in maize ovule dissection. Sample preparation, immuno-EM experiments, and TEM observations were performed at the Centre Technologique des Microstructures at Université Lyon 1. We acknowledge the contribution of the green house technical platform of FR3728 BioEnviS at Université Lyon 1, and SFR Biosciences (UMS3444, Centre national de la recherche scientifique; US8/INSERM; École normale supérieure de Lyon, Université Claude Bernard Lyon) facilities, and in particular LBI-PLATIM-MICROSCOPY for assistance with imaging.
This research was supported by an Agence Nationale de la Recherche grant (ANR-19-CE20-0012, “Not-Like-Dad”) to T. Widiez and by “pack ambition recherche” from the Région Auvergne-Rhône-Alpes (HD-INNOV) to T. Widiez. N.M.A. Jacquier and L.M. Gilles were supported by Convention industrielle de formation par la recherche PhD fellowships from Association Nationale de la Recherche et de la Technologie funding agency (grants 2019/0771 and 2015/0777, respectively).
N.M.A. Jacquier, L.M. Gilles, and J.P. Martinant are employees of LIMAGRAIN Europe. P.M. Rogowsky is part of the "Groupement d’Intérêt Scientifique Biotechnologies Vertes.” The authors declare no additional competing financial interests.
Author contributions: L.M. Gilles and T. Widiez conceived and designed the experiments; L.M. Gilles, A.R.M. Calhau, V. La Padula, N.M.A. Jacquier, C. Lionnet, and T. Widiez conducted experiments; V. La Padula did immuno-EM and TEM imaging, which were then analyzed by L.M. Gilles, N.M.A. Jacquier, and T. Widiez; A.R.M. Calhau and C. Lionnet imaged and analyzed the behavior of NLD:mCitrine during fertilization; A.R.M. Calhau did RT-PCR experiments; L.M. Gilles performed all other remaining experiments presented; L.M. Gilles prepared tables and figures with the exception of Fig. 6 and its corresponding supplemental figures, which were prepared by A.R.M. Calhau and C. Lionnet; L.M. Gilles and T. Widiez led the writing of the manuscript, and V. La Padula and P.M. Rogowsky contributed to the critical reading of the manuscript; P.M. Rogowsky, J-P. Martinant, and T. Widiez were involved in project management; T. Widiez obtained funding; T. Widiez initiated and coordinated the project.