Keratin intermediate filaments (KIFs) protect the epidermis against mechanical force, support strong adhesion, help barrier formation, and regulate growth. The mechanisms by which type I and II keratins contribute to these functions remain incompletely understood. Here, we report that mice lacking all type I or type II keratins display severe barrier defects and fragile skin, leading to perinatal mortality with full penetrance. Comparative proteomics of cornified envelopes (CEs) from prenatal KtyI−/− and KtyII−/−K8 mice demonstrates that absence of KIF causes dysregulation of many CE constituents, including downregulation of desmoglein 1. Despite persistence of loricrin expression and upregulation of many Nrf2 targets, including CE components Sprr2d and Sprr2h, extensive barrier defects persist, identifying keratins as essential CE scaffolds. Furthermore, we show that KIFs control mitochondrial lipid composition and activity in a cell-intrinsic manner. Therefore, our study explains the complexity of keratinopathies accompanied by barrier disorders by linking keratin scaffolds to mitochondria, adhesion, and CE formation.
Introduction
The mammalian epidermis is a stratified epithelium that protects the body against mechanical force, dehydration, and infections. These functions rely on a series of differentiation events that give rise to the expression of distinct sets of epidermal keratins and to the formation of the cornified envelope (CE), which together with tight junctions and epidermal Langerhans cells represent the protective epidermal barrier (Roop, 1995; Fuchs, 2008; Watt, 2014).
The keratin intermediate filament (KIF) cytoskeleton is essential for the maintenance of epithelial integrity by virtue of its unique biochemical and biophysical properties, its interaction with cell adhesion complexes (Herrmann and Aebi, 2004). KIFs assemble from heterodimers of 28 keratin type I (KtyI) and 26 type II (KtyII) keratins expressed in cell- and differentiation-specific patterns (Schweizer et al., 2006; Homberg and Magin, 2014). The basal, proliferative compartment expresses the keratin pair K5/K14, which is replaced during terminal differentiation by K1/K10, whereas K6/K16/K17 are induced during barrier breach and in disorders, including atopic dermatitis and psoriasis (Fuchs and Green, 1980; Coulombe and Lee, 2012; Roth et al., 2012b). Loss or disruption of keratins causes blistering and hyperkeratotic skin disorders, accompanied by cell fragility, diminished adhesion, barrier defects, growth alterations, and inflammation (Schmuth et al., 2001; Lane and McLean, 2004; Segre, 2006; Arin et al., 2011; Coulombe and Lee, 2012). The molecular basis for keratin anchorage at hemidesmosomes and desmosomes is relatively well understood (Simpson et al., 2011; Kröger et al., 2013; Seltmann et al., 2013b, 2015). In contrast, the mechanisms by which keratins contribute to CE formation and barrier function, which are crucial for understanding the pathomechanisms underlying atopic dermatitis, remain largely unknown. Based on cryo-electron microscopy, it has been proposed that KIFs are arranged in cube-like rods to form the template for the membrane of corneocytes (Norlén and Al-Amoudi, 2004). In fact, a conserved lysine residue in the head domain of type II keratins mediates the attachment of KIF to the CE through isodipeptide cross-linking mediated by transglutaminases (Candi et al., 1998).
The CE represents the outermost epidermal layer composed of transglutaminase-cross-linked proteins involucrin, loricrin, filaggrins, small proline-rich proteins (SPRRs), late cornified envelope proteins (LCEs), S100 protein family members, and protein-bound ω-hydroxyceramides (Kypriotou et al., 2012). CE formation proceeds in three major steps, beginning with transglutaminase 1–dependent cross-linking of envoplakin, periplakin, involucrin, and filaggrin, with desmosomal proteins underneath the plasma membrane in a Ca2+-dependent manner (Kalinin et al., 2001; Candi et al., 2005; Matsui and Amagai, 2015). In a second step, cross-linking of loricrin, Sprr, and Lce by transglutaminase 3 provides reinforcement. The third step comprises release of lipids from lamellar granules into the extracellular space, fusion with the plasma membrane, and their cross-linking to CE proteins (Candi et al., 2005).
The analysis of genetic defects in major genes coding for CE proteins, including involucrin, envoplakin, periplakin, filaggrin, and loricrin, has led to the concept of compensatory redundancy to sustain barrier integrity (Koch et al., 2000; Sevilla et al., 2007; Huebner et al., 2012; Kawasaki et al., 2012; Matsui and Amagai, 2015). In this direction, upregulation of Sprr2d and Sprr2h via the bZIP antioxidant transcription factor Nrf2 has been suggested to correct a transient barrier defect in mice lacking loricrin (Koch et al., 2000; Huebner et al., 2012). Although type II keratins were identified among cross-linked CE proteins, their significance for CE assembly and barrier function has not been investigated so far (Candi et al., 2005).
In addition, certain keratins interact with mitochondria (Nishizawa et al., 2005; Duan et al., 2009) to mediate their intracellular distribution and possibly to modulate mitochondrial energy metabolism (Tao et al., 2009; Helenius et al., 2015). Moreover, dysmorphic mitochondria were detected in Krt5−/− and Krt16−/− mice (Takahashi et al., 1994; Alvarado and Coulombe, 2014), indicating an involvement of keratins in mitochondrial function. Given that epidermal differentiation also depends on mitochondrial inner membrane dynamics and activity of the electron transport chain (ETC) (Baris et al., 2011; Hamanaka et al., 2013; Kloepper et al., 2015), we set out to investigate the scaffolding function of keratins for CE formation and mitochondrial activity. To this end, we generated mice lacking all type I keratin (KtyI−/−) genes in the epidermis and compared them with previously reported KtyII deficiency in mice that were rescued from embryo mortality by transgenic Krt8 expression (KtyII−/−K8; Bär et al., 2014). This revealed an unprecedented reliance of CE assembly and barrier function on KIF, identifying them as essential CE components. Despite persistence of loricrin and some keratin proteins, the strong activation of the Nrf2 pathway failed to rescue barrier defects in the absence of KIF. Further, we discovered cell-intrinsic and keratin-dependent changes in mitochondrial protein and lipid composition, leading to increased oxygen consumption and elevated ATP levels.
Results
Keratin pairs K8/K19 or K8/K18 sustain morphogenesis of simple epithelia until birth
To identify novel functions of type I and II keratins critical for epidermal integrity, barrier function, and metabolism, newly generated mice lacking the entire KtyI gene cluster on chromosome 11 were compared with recently established KtyII−/−K8 mice (Bär et al., 2014). In KtyI−/− mice, TyI Krt18, located in the KtyII gene cluster (Fig. S1 A), enables KIF formation in all simple epithelia together with TyII Krt8 to overcome embryo mortality of KtyII−/− mice at embryonic day 9.5 (E9.5) (Vijayaraj et al., 2009; Kröger et al., 2011). The absence of all type I keratin genes in stratified epithelia was predicted to disable KIF formation in all stratified epithelia.
Floxed KtyI−/− mice were analyzed by PCR and Southern blotting (Fig. S1, B–D). Crossing of homozygous KtyI floxed 129S1hprt Cre deleter mice resulted in KtyI−/− at approximate Mendelian ratios (Fig. S1 B). Similar to KtyII−/−K8 (Bär et al., 2014), KtyI−/− embryos developed to term, displayed shiny and extremely trauma-sensitive skin, and died at birth (Fig. S1 E for E18.5 embryos). Therefore, both K8/K18 and K8/K19 rescued the embryonic growth defect reported before, rendering loss of miRNAs or ncRNAs unlikely to cause the previously reported phenotype (Vijayaraj et al., 2009).
Given the expression of up to seven keratins in developing simple epithelia (Owens and Lane, 2004; Habtezion et al., 2005; Langbein et al., 2015; Loschke et al., 2015; Fig. 1 F), key adhesion molecules, differentiation markers, and remaining keratins were examined in the small intestine and stomach of E18.5 KtyI−/− and, for some specimens, in KtyII−/−K8 embryos. In both mouse strains, the distribution of K8/K18 and of K8/K19 in the intestine was highly similar to that in controls (Fig. 1 A). The strong decrease of Krt8 protein amount in KtyI−/− intestine (Fig. 1 C), which lacks Krt19, likely results from the low amount of its residual dimerization partner Krt18, which limits the amount of heterodimeric intermediate filaments. This indicates preferential KIF formation between K8/K19 in wild-type (WT) intestinal epithelia. In support, K8/K19 amounts in KtyII−/−K8, which has no Krt18 gene (Figs. S1 A and S2, A and B), were unaltered, and K8/K18 in the stomach of KtyI−/− embryos were reduced (Fig. S2 E). Histologic sections from the stomach and small intestine showed intact morphology without alterations in length of the crypt-villus axis, despite reduced keratin expression (Fig. 1 E and Fig. S2, C and E). Further, based on periodic acid–Schiff staining, intestinal goblet cell number and integrity appeared unchanged (Fig. 1 E). In KtyI−/− intestinal epithelia, intercellular junctions and the polarity of internal epithelia until birth in KtyI−/− intestinal epithelia are collectively supported by the filament formation proficiency of K8/K18 (Fig. 1 C) or K8/K19 in KtyII−/−K8. Throughout these stages, cell proliferation and localization of actin, E-cadherin, and desmoplakin remained unaltered (Fig. 1 B and Fig. S2 D).
K8/K18 filaments sustain normal development of intestine simple epithelia until birth in KtyI−/− mice. (A) Immunofluorescence analysis of Krt8, Krt18, and Krt19 in embryonic intestine of KtyI+/+ and KtyI−/− E18.5 embryos. Bars, 20 µm. (B) Immunofluorescence of actin (phalloidin), E-cadherin (E-Cad), and desmoplakin (DSP) showed unaltered localization in E18.5 KtyI+/+ and KtyI−/− embryos. Bars, 20 µm. (C) Western blotting of Krt8, Krt18, Krt19, and α-tubulin as loading control of intestine of KtyI+/+ and KtyI−/− E18.5 embryos. (D) Unaltered proliferation rate in the intestine of E18.5 KtyI+/+ and KtyI−/− embryos based on Ki-67 labeling. Related to Fig. S2 D. (E) Hematoxylin/eosin and periodic acid–Schiff (PAS) stained sections of intestine from E18.5 KtyI+/+ and KtyI−/− embryos revealing normal morphology. (F) Expression of simple epithelial keratins in different organs of WTs and KtyI−/− and KtyII−/−K8 E18.5 embryos.
K8/K18 filaments sustain normal development of intestine simple epithelia until birth in KtyI−/− mice. (A) Immunofluorescence analysis of Krt8, Krt18, and Krt19 in embryonic intestine of KtyI+/+ and KtyI−/− E18.5 embryos. Bars, 20 µm. (B) Immunofluorescence of actin (phalloidin), E-cadherin (E-Cad), and desmoplakin (DSP) showed unaltered localization in E18.5 KtyI+/+ and KtyI−/− embryos. Bars, 20 µm. (C) Western blotting of Krt8, Krt18, Krt19, and α-tubulin as loading control of intestine of KtyI+/+ and KtyI−/− E18.5 embryos. (D) Unaltered proliferation rate in the intestine of E18.5 KtyI+/+ and KtyI−/− embryos based on Ki-67 labeling. Related to Fig. S2 D. (E) Hematoxylin/eosin and periodic acid–Schiff (PAS) stained sections of intestine from E18.5 KtyI+/+ and KtyI−/− embryos revealing normal morphology. (F) Expression of simple epithelial keratins in different organs of WTs and KtyI−/− and KtyII−/−K8 E18.5 embryos.
Extensive adhesion defects and cytolysis in the epidermis of KtyI−/− and KtyII−/−K8 embryos
Recent genetic and biochemical data established a major role of keratins in cytoarchitecture, cell adhesion, cell growth, and regulation of distinct immune functions (Kim et al., 2006; Vijayaraj et al., 2009; Kim and Coulombe, 2010; Wallace et al., 2012; Roth et al., 2012a,b; Lessard et al., 2013; Chung et al., 2015).
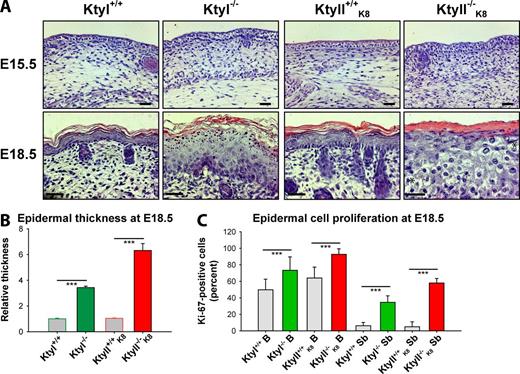
To examine the respective contribution of type I and type II keratins to epidermal differentiation, tissue morphology and proliferation were analyzed in histologic sections at the onset of stratification at E15.5 in KtyI−/− and KtyII−/−K8 embryos. At this developmental stage, the epidermis of both knockout (KO) strains of mice was intact and of similar thickness (Fig. 2 A), but it was more fragile during dissection than WT (Bär et al., 2014) and showed no significant difference in proliferation rates (Fig. S3 B). In contrast, at E18.5, extensive cytolysis, predominantly between the upper spinous and cornified layers, and skin hyperthickening of both KtyI−/− and KtyII−/−K8 became apparent (3.5-fold increase in the former and sixfold in the latter) (Fig. 2, A and B; and Fig. S3 A; Bär et al., 2014). Moreover, KO keratinocytes appeared larger. At the same time, cell-matrix contacts were less severely affected (Seltmann et al., 2013b, 2015). In line with extensive cell damage and epidermal hyperthickening, proliferation rates were significantly increased in basal and mostly in suprabasal keratinocytes of KtyI−/− and KtyII−/−K8, respectively, when compared with WT as inferred by the percentage of Ki-67 positive cells (Fig. 2 C and Fig. S3 C). Unexpectedly, filaggrin-containing keratohyalin granules were detectable in the granular layer of KtyI−/− but absent in KtyII−/−K8 epidermis (Fig. 2 A). Finally, stratum corneum compactness appeared decreased in KtyI−/− and increased in KtyII−/−K8 skin samples, with respect to controls. This suggested keratin-dependent CE defects.
KtyI and KtyII ablation causes epidermal thickening, hyperproliferation, cytolysis, and distinct structural abnormalities of the granular and stratum layers. (A) Hematoxylin/eosin-stained skin sections of E15.5 and E18.5 mouse embryos. Note apparently normal epidermis at E15.5 but hyperthickening and cytolysis (asterisk) in KtyI−/− and KtyII−/−K8 at E18.5. Large keratohyalin granules in KtyI−/− and their absence in KtyII−/−K8 epidermis, which shows a compact stratum corneum. Bars, 20 µm. (See also different magnification of other pictures in Fig. S3 A). (B) Measurements showing distinct increase of epidermal thickness at E18.5 in KtyI−/− and KtyII−/−K8. (C) Ki-67–based quantification of proliferation rates in basal (B) and suprabasal (Sb) cells of the epidermis of WT and KtyI−/− and KtyII−/−K8 embryos at E18.5 (see pictures in Fig. S3 C), showing significant increase in the absence of KIF. ***, P ≤ 0.001.
KtyI and KtyII ablation causes epidermal thickening, hyperproliferation, cytolysis, and distinct structural abnormalities of the granular and stratum layers. (A) Hematoxylin/eosin-stained skin sections of E15.5 and E18.5 mouse embryos. Note apparently normal epidermis at E15.5 but hyperthickening and cytolysis (asterisk) in KtyI−/− and KtyII−/−K8 at E18.5. Large keratohyalin granules in KtyI−/− and their absence in KtyII−/−K8 epidermis, which shows a compact stratum corneum. Bars, 20 µm. (See also different magnification of other pictures in Fig. S3 A). (B) Measurements showing distinct increase of epidermal thickness at E18.5 in KtyI−/− and KtyII−/−K8. (C) Ki-67–based quantification of proliferation rates in basal (B) and suprabasal (Sb) cells of the epidermis of WT and KtyI−/− and KtyII−/−K8 embryos at E18.5 (see pictures in Fig. S3 C), showing significant increase in the absence of KIF. ***, P ≤ 0.001.
To unravel the origin for the distinct changes in KtyI−/− and KtyII−/−K8 compared with control epidermis, immunofluorescence and electron microscopic analysis were performed. This confirmed absence of KIF in both genotypes (Figs. 3 A and 4, B, F, and H) and presence of Krt5 and Krt1 aggregates in KtyI−/− and of Krt14 aggregates in KtyII−/−K8 epidermis (Fig. 3 A), reminiscent of those reported in single keratin gene KO mice and in patients suffering from keratinopathies (Magin, 2004; Coulombe et al., 2009; Arin et al., 2011). On KtyI deletion, Krt1 and Krt5 protein amounts strongly decreased, whereas Krt6 increased (Fig. 3 B). Similarly, Krt10 quantity was decreased in KtyII−/−K8 skin (Fig. 3 B). Krt14 quantity was, however, not affected (Fig. 3 B). Most remarkably, Krt17, known to regulate mTOR and involved in immune responses (Kerns et al., 2010), was elevated only threefold, whereas the Nrf2 target genes Krt16 (Huebner et al., 2012) and its KtyII partner Krt6 were induced 500-fold at the mRNA level in corresponding KO mice (Fig. S3, D and E), suggesting elevated Nrf2 activity because of barrier defects (Huebner et al., 2012). Krt6 was also detected by Western blot (Fig. 3 B) and formed aggregates in suprabasal keratinocytes (Fig. 3 B).
Isotype-specific keratin expression and localization in KtyI −/− and KtyII −/− K8 epidermis. (A) Immunofluorescence analysis of keratin localization in WTs and KtyI−/− and KtyII−/−K8 epidermis at E18.5. Note absence of KIF in both KOs, suprabasal aggregates of Krt1 and Krt5 in KtyI−/−, and widespread aggregates of Krt14, but not of Krt10, in KtyII−/−K8. Basement membrane is indicated by white dotted lines. Bars, 20 µm. (B) Western blot analysis of Krt1, Krt5, Krt6, Krt10, Krt14, and α-tubulin as loading control in skin extracts of WT and keratin-deficient embryos at E18.5. Note positive correlation with A, showing a significant increase in Krt6 in KtyI−/− but unaltered presence of Krt14 in KtyII−/−K8 compared with KtyII+/+K8.
Isotype-specific keratin expression and localization in KtyI −/− and KtyII −/− K8 epidermis. (A) Immunofluorescence analysis of keratin localization in WTs and KtyI−/− and KtyII−/−K8 epidermis at E18.5. Note absence of KIF in both KOs, suprabasal aggregates of Krt1 and Krt5 in KtyI−/−, and widespread aggregates of Krt14, but not of Krt10, in KtyII−/−K8. Basement membrane is indicated by white dotted lines. Bars, 20 µm. (B) Western blot analysis of Krt1, Krt5, Krt6, Krt10, Krt14, and α-tubulin as loading control in skin extracts of WT and keratin-deficient embryos at E18.5. Note positive correlation with A, showing a significant increase in Krt6 in KtyI−/− but unaltered presence of Krt14 in KtyII−/−K8 compared with KtyII+/+K8.
Electron microscopy analysis of skin from KtyI+/+ and KtyI −/− E18.5 embryos depicting structural changes. (A and B) Hemidesmosomes (black arrowheads) of the stratum basale are readily detectable in KtyI−/− animals. Note, however, the absence of keratin filaments (KF) and occurrence of cytolysis in the basal cell layer of KtyI−/− epidermis (B, asterisk). Insets show higher magnification of hemidesmosomes together with the adjacent basal lamina (BL). (E and F) Desmosomes (black arrows) in the suprabasal layers of KtyI−/− epidermis are smaller and less frequent compared with KtyI+/+. Note the wide intercellular space in the stratum spinosum of KtyI−/− epidermis and intramitochondrial inclusions (white arrowheads). Insets in E and F show higher magnification of desmosomes. (G and H) Corneodesmosomes (white arrows) in the stratum corneum can be easily distinguished in KtyI−/− animals. Insets show higher magnification of corneodesmosomes. Bars: (insets) 100 nm; (A, B, and E–H) 1 µm; (C and D) 5 µm.
Electron microscopy analysis of skin from KtyI+/+ and KtyI −/− E18.5 embryos depicting structural changes. (A and B) Hemidesmosomes (black arrowheads) of the stratum basale are readily detectable in KtyI−/− animals. Note, however, the absence of keratin filaments (KF) and occurrence of cytolysis in the basal cell layer of KtyI−/− epidermis (B, asterisk). Insets show higher magnification of hemidesmosomes together with the adjacent basal lamina (BL). (E and F) Desmosomes (black arrows) in the suprabasal layers of KtyI−/− epidermis are smaller and less frequent compared with KtyI+/+. Note the wide intercellular space in the stratum spinosum of KtyI−/− epidermis and intramitochondrial inclusions (white arrowheads). Insets in E and F show higher magnification of desmosomes. (G and H) Corneodesmosomes (white arrows) in the stratum corneum can be easily distinguished in KtyI−/− animals. Insets show higher magnification of corneodesmosomes. Bars: (insets) 100 nm; (A, B, and E–H) 1 µm; (C and D) 5 µm.
Our recent analysis of KtyII−/−K8 mice and keratinocytes has shown a major role of keratins in desmosome maintenance and levels of desmosomal proteins (Kröger et al., 2013; Bär et al., 2014). By electron microscopy, hemidesmosomes remained intact in KtyI−/− epidermis, despite localized cytolysis (Fig. 4, A–D). Desmosomes were smaller, as previously reported (Fig. 4, C–F), and corneodesmosomes appeared fully formed (Fig. 4, G and H). Remarkably, a subset of mitochondria showed dark inclusions but appeared otherwise intact (Fig. 4 D), indicating an involvement of keratins in their integrity (Alvarado and Coulombe, 2014). Collectively, expression and localization of desmosomal proteins in KtyI−/− epidermis was similar to that in KtyII−/−K8 mice (Fig. S4, A–C). Notably, desmoplakin and desmogleins 1 and 2 proteins were significantly reduced on extraction with Laemmli buffer (Fig. S4 C). At the same time, plakoglobin protein expression and distribution remained unaffected by the keratin status in SDS extracts (Fig. S4, A and C).
To better understand the impact of keratin loss on the epidermis, we analyzed the transcriptome profiles of E18.5 embryo skin. Gene set enrichment analysis, followed by gene ontology classification and grouping according to function, revealed numerous expression changes after keratin locus deletion, notably in genes involved in oxidative response, epidermal differentiation, immune response, lipid metabolism, and apoptosis/proliferation (Fig. S3 F), suggesting epidermal barrier defects.
Barrier integrity, CE composition, and assembly depend on filamentous keratins
The ill-defined role of keratins in CE assembly and barrier function (Candi et al., 2005), together with known compensatory mechanisms restoring the barrier on genetic or pharmacologic interference (Segre et al., 1999; Yu et al., 2006; Sevilla et al., 2007), prompted us to investigate a mechanistic link between keratins, barrier composition, and function. Examination of the outside-in barrier in E18.5 embryos using a dye-penetration assay detected a more severe barrier defect in KtyII−/−K8 than in KtyI−/− compared with respective controls (Fig. 5 A). At E17.5, the difference was less pronounced between the two strains of mice (Fig. 5 A). To resolve this, CEs were isolated. Those from KtyII−/−K8 embryos were completely ruptured, whereas ∼15% remained intact in KtyI−/− preparations, suggesting alterations in protein composition or cross-linking activity (Fig. 5, B and C). This difference was reflected by the toluidine blue assay, indicating outside-in barrier defects. Western blotting and immunofluorescence analysis of E18.5 skin showed that in KtyI−/− epidermis, expression of loricrin, involucrin, and filaggrin appeared unaltered (Fig. 5 D). Surprisingly, loricrin was absent in KtyII−/−K8 but remained unaltered in KtyI−/− samples (Fig. 5, D and E). Profilaggrin is a large keratin-associated protein of >400 kD processed by caspase-14 and matriptase to filaggrin monomers contributing to barrier function (Hoste et al., 2011). Its deficiency is a major cause of atopic dermatitis (Sandilands et al., 2007; Kawasaki et al., 2012). Similar to loricrin, filaggrin was absent in KtyII−/−K8 skin, and profilaggrin to filaggrin processing was nearly abrogated in KtyI−/− skin, as indicated by a strong reduction of filaggrin monomers (Fig. 5 E). In contrast, protein levels of involucrin, another prominent CE protein, were unaltered in KtyI−/− but reduced by ∼50% in KtyII−/−K8 skin extracts (Fig. 5 E). To understand the mechanisms underlying the altered expression of those CE genes encoded in the epidermal differentiation complex (EDC) locus (Kypriotou et al., 2012), illumina arrays and quantitative RT-PCR were performed from dorsal skin RNA of E18.5 keratin-deficient and control embryos. Of the 34 selected genes, 20 were upregulated twofold or more, and one was downregulated twofold or more in KtyI−/−, whereas 14 genes were upregulated twofold or more, and 14 were downregulated twofold or more in the KtyII−/−K8 (Fig. 5 F). 15 genes were similarly regulated in both mouse strains. Focusing on differentially regulated CE genes, mRNAs for filaggrin, hornerin, loricrin, and Lce3b (Fig. 5 G) were absent in KtyII−/−K8 compared with KtyI−/− mice, suggesting that this contributed to the more severe barrier defect in the former. The molecular basis for the transcriptional dysregulation of EDC genes in KtyII−/−K8 will be described elsewhere (unpublished data).
Distinct epidermal barrier defects of KtyI −/− and KtyII −/− K8 mouse embryos. (A) Toluidine blue assay indicating moderately delayed acquisition of dye-impermeable barrier in KtyI−/− and a severe delay in KtyII−/−K8 compared with WT embryos. Bars, 5 mm. (B and C) 85% of CEs from E18.5 embryos were disrupted in KtyI−/− compared with 99% in KtyII−/−K8 CE compared with WTs. Bars, 100 µm. (D) Immunofluorescence analysis of E18.5 epidermis revealing highly similar distribution of involucrin, loricrin, and filaggrin in KtyI+/+ and KtyI−/−. Note reduced filaggrin staining in KtyI−/− samples. The basement membrane is indicated by white dotted lines. Bars, 20 µm. (E) Western blotting of skin extracts from E18.5 embryos showing a decrease in involucrin (Ivl) and absence of loricrin (Lor) and filaggrin (Flg) in KtyII−/−K8, together with decreased processing of profilaggrin (Pro-flg, upper bands) into filaggrin (bottom bands) in KtyI−/− skin; α-tubulin (tub) served as loading control. (F) Transcriptome analysis exposing distinct changes in the expression of barrier protein-encoding genes in KtyI−/− and KtyII−/−. Note strong downregulation of a few barrier protein-encoding genes in KtyII−/−K8 only (in red); n = 3. (G) Quantitative RT-PCR analysis confirming the absence of transcription of loricrin, filaggrin, and hornerin genes in KtyII−/−K8 and upregulation of involucrin and loricrin mRNA in KtyI−/− skin. ***, P ≤ 0.001.
Distinct epidermal barrier defects of KtyI −/− and KtyII −/− K8 mouse embryos. (A) Toluidine blue assay indicating moderately delayed acquisition of dye-impermeable barrier in KtyI−/− and a severe delay in KtyII−/−K8 compared with WT embryos. Bars, 5 mm. (B and C) 85% of CEs from E18.5 embryos were disrupted in KtyI−/− compared with 99% in KtyII−/−K8 CE compared with WTs. Bars, 100 µm. (D) Immunofluorescence analysis of E18.5 epidermis revealing highly similar distribution of involucrin, loricrin, and filaggrin in KtyI+/+ and KtyI−/−. Note reduced filaggrin staining in KtyI−/− samples. The basement membrane is indicated by white dotted lines. Bars, 20 µm. (E) Western blotting of skin extracts from E18.5 embryos showing a decrease in involucrin (Ivl) and absence of loricrin (Lor) and filaggrin (Flg) in KtyII−/−K8, together with decreased processing of profilaggrin (Pro-flg, upper bands) into filaggrin (bottom bands) in KtyI−/− skin; α-tubulin (tub) served as loading control. (F) Transcriptome analysis exposing distinct changes in the expression of barrier protein-encoding genes in KtyI−/− and KtyII−/−. Note strong downregulation of a few barrier protein-encoding genes in KtyII−/−K8 only (in red); n = 3. (G) Quantitative RT-PCR analysis confirming the absence of transcription of loricrin, filaggrin, and hornerin genes in KtyII−/−K8 and upregulation of involucrin and loricrin mRNA in KtyI−/− skin. ***, P ≤ 0.001.
Keratin-dependent epidermal barrier defects persist despite upregulation of Nrf2
The CE defects prompted the question whether Nrf2-mediated upregulation of Sprr2d and Sprr2h, rescuing transient barrier defects in loricrin−/− mice (Huebner et al., 2012; Singla et al., 2012), was compromised in keratin-deficient embryos. Transcriptome profiling, followed by quantitative RT-PCR, established strong upregulation of 14 major Nrf2 target genes in keratin-deficient skin. mRNAs encoding the CE constituents Sprr2d, Sprr2h, Krt6, and Krt16, and Slpi, a protease inhibitor displaying antimicrobial activity (Schäfer et al., 2014), were induced 200-fold (Fig. 6, A and B; and Fig. S3, D and E). mRNAs for Nqo1, Gsta3, and additional Nrf2 target genes mediating an antioxidant response were also upregulated, albeit to a lower extent (Fig. 6, A and B). Moreover, Nrf2 mRNA and proteins were upregulated in the epidermis of both strains of KO mice (Fig. 6, C and D), whereas the Nrf2 regulator Keap1 remained unaltered. The strong and persistent nuclear Nrf2 accumulation supported its role in expression of the aforementioned target genes (Fig. 6 E). Therefore, in the absence of KIF, significant upregulation of several Sprr genes, including Sprr2d and Sprr2h, failed to restore a functional barrier. This was irrespective of the absence or presence of loricrin (Fig. 5, D–G). To further dissect KIF requirements for barrier formation, the proteome profile of highly insoluble CEs from KtyI−/− and KtyII−/−K8 in comparison with WT epidermis was analyzed.
Induction of Nrf2 in KtyI −/− and KtyII −/− K8 prenatal epidermis. (A) Strong upregulation of selected Nrf2 target genes from transcriptome profiles of E18.5 KtyI−/− and KtyII−/−K8 skin RNA. (B) Confirmation of data in A, using quantitative RT-PCR analysis. (C) Transcriptome profile and quantitative RT-PCR showing increase in Nrf2 mRNA in the skin of keratin-deficient embryos. (D) Western blot establishing a strong increase of Nrf2 protein in KtyI−/− and KtyII−/−K8 at E17.5 and E18.5, but no difference in Keap1. (E) Persistent upregulation and nuclear localization of Nrf2 in the epidermis of both KO mice, based on immunohistochemistry of E16.5–E18.5 embryos. Bars: (main) 50 µm; (insets) 10 µm. *, P ≤ 0.05.
Induction of Nrf2 in KtyI −/− and KtyII −/− K8 prenatal epidermis. (A) Strong upregulation of selected Nrf2 target genes from transcriptome profiles of E18.5 KtyI−/− and KtyII−/−K8 skin RNA. (B) Confirmation of data in A, using quantitative RT-PCR analysis. (C) Transcriptome profile and quantitative RT-PCR showing increase in Nrf2 mRNA in the skin of keratin-deficient embryos. (D) Western blot establishing a strong increase of Nrf2 protein in KtyI−/− and KtyII−/−K8 at E17.5 and E18.5, but no difference in Keap1. (E) Persistent upregulation and nuclear localization of Nrf2 in the epidermis of both KO mice, based on immunohistochemistry of E16.5–E18.5 embryos. Bars: (main) 50 µm; (insets) 10 µm. *, P ≤ 0.05.
A keratin scaffold regulates CE formation at distinct stages
Previous analysis of CE constituents by gene deletion suggested a considerable redundancy because of upregulation of alternative constituents (Segre, 2006; Matsui and Amagai, 2015). Although genetic evidence specifically indicated an involvement of Krt1 in CE integrity (Candi et al., 2005; Roth et al., 2012b), the overall requirement of keratins during primary CE formation at E18.5 has not been investigated so far. Mass spectrometry of CE tryptic peptides from WTs epidermis revealed that Krt1 and Krt10, together with threefold lower amounts of Krt77, were major CE constituents, in addition to moderate amounts of Krt75, Krt78, and Krt5 (Fig. 7 A; Rorke et al., 2015). Additional keratins were very minor constituents according to mass spectrometry (Fig. 7 A). We note that incorporation of Krt1, Krt10, and Krt77 coincides with their known distribution in the epidermis and that the precursor cell keratin Krt15 was excluded from the CE (Fuchs and Green, 1980). Analysis of CEs from KtyI−/− and KtyII−/−K8 pups revealed substantial KIF-dependent alterations. Although not detected by antibody analysis, a decreased amount of Krt10 was present, as revealed by mass spectrometry of CE preparations of KtyII−/−K8 (Fig. 3, A and B; and Fig. 7C). The residual amount of type II proteins in KtyI−/− was two- to threefold lower than that of type I proteins in KtyII−/− (Fig. 7, B and C). In the former, only suprabasally expressed keratins were present (Fig. 7 B), whereas cross-linking of the basal Krt14 was not altered in the latter. In agreement with mRNA levels of Krt16 being much higher than for Krt17, both stress keratins were represented accordingly among CE proteins. Therefore, keratins are major CE constituents by mass. The extensive CE defects in both strains of KO mice, despite persistence of significant amounts of cross-linked individual keratins, for the first time show that KIFs are a prerequisite for CE function.
Keratins as major CE constituents at E18.5. Proteomic profiling of enriched CE preparations from WTs and KtyI−/− and KtyII−/−K8 mouse embryos and subsequent mass spectrometry. (A) Spectral counts of major CE keratins of mouse skin are presented. Type I keratins are underlined in red, and type II are underlined in green. In addition to Krt1 and Krt10, Krt77, Krt75, and Krt78 represent major CE constituents. (B) Same as in A but for type II keratins in KtyI+/+ and KtyI−/− mice. Note significant decrease of type II keratins except Krt6 among KtyI−/− cross-linked samples. (C) Remarkably, only the type I keratin Krt10 is significantly decreased in KtyII−/−K8, whereas Krt14 and Krt17 remain unaltered and more Krt16 is cross-linked in KtyII−/−K8. *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
Keratins as major CE constituents at E18.5. Proteomic profiling of enriched CE preparations from WTs and KtyI−/− and KtyII−/−K8 mouse embryos and subsequent mass spectrometry. (A) Spectral counts of major CE keratins of mouse skin are presented. Type I keratins are underlined in red, and type II are underlined in green. In addition to Krt1 and Krt10, Krt77, Krt75, and Krt78 represent major CE constituents. (B) Same as in A but for type II keratins in KtyI+/+ and KtyI−/− mice. Note significant decrease of type II keratins except Krt6 among KtyI−/− cross-linked samples. (C) Remarkably, only the type I keratin Krt10 is significantly decreased in KtyII−/−K8, whereas Krt14 and Krt17 remain unaltered and more Krt16 is cross-linked in KtyII−/−K8. *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
To substantiate this further, we classified nonkeratin CE constituents into four groups according to their function and assembly into the CE (Candi et al., 2005; Matsui and Amagai, 2015), namely, (a) initiation and reinforcement (corneodesmosome, transglutaminase, and Sprr), (b) major CE constituent proteins, (c) proteases and protease inhibitors, and (d) alarmins (Fig. 8). In contrast with lower amounts in epidermal SDS-soluble extracts (Fig. S4 C; Bär et al., 2014), constituent proteins of corneodesmosomes were significantly more cross-linked in both KtyI−/− and KtyII−/−K8, with the noticeable exception of desmoglein 1, which was decreased in both KO strains (Fig. 8 A). More periplakin and envoplakin, known early CE constituents, together with desmosomal proteins plakoglobin and plakophilins 1 and 3, were cross-linked in the absence of both keratin families. On the contrary, enzymes such as Tgm 1, 3, or Alox12b, critical for protein cross-linking and the synthesis of corneocyte lipids, respectively (Matsui and Amagai, 2015), were underrepresented in the CE–cross-linked fraction (Fig. 8 A). This was accompanied by a significant increase of most Sprr, including the Nrf2 targets Sprr2d, Sprr2f, and Sprr2h. Despite their upregulation, barrier function was not restored (Figs. 5 and 8A). Therefore, early steps of CE assembly were altered but not prevented in the absence of KIF. At the same time, with the exception of the S100 family member repetin, which was increased, the major known CE components loricrin, hornerin, filaggrin, and filaggrin-2 were decreased in both KOs (Fig. 8 B). Residual loricrin detected in the CE of E18.5 KtyII−/−K8 animals most probably resulted from its biosynthesis during earlier embryonic stages, indicating slow turnover (Fig. 5 E and Fig. S5 B).
CE proteome profile is keratin dependent. Grouping of CE constituents according to function in the presence and absence of type I and type II keratins. (A) CE initiation and reinforcement. Decrease in desmoglein-1 (underlined in blue), but increase in corneodesmosome and CE initiation and reinforcement proteins in KtyI−/− and KtyII−/−K8 compare d with WT. Less transglutaminase (tgm-1 and -3) and arachidonate 12-lipoxygenase (alox12b) are cross-linked in both KtyI−/− and KtyII−/−K8 compared with WT. (B) Except repetin, fewer major CE structural proteins are cross-linked in both KO strains compared with WT. (C) Cross-linking of proteases and protease inhibitors involved in terminal differentiation and barrier function is similarly changed in KtyI−/− and KtyII−/−K8 compared with WTs. Cathepsin-D (Ctsd) and Spink5 are underlined in orange and are only altered in KtyII−/−K8. (D) Enhanced cross-linking of alarmins in KtyI−/− and KtyII−/−K8 compared with WTs (red hash marks hspa2, which is upregulated in KtyII−/−K8 only). *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
CE proteome profile is keratin dependent. Grouping of CE constituents according to function in the presence and absence of type I and type II keratins. (A) CE initiation and reinforcement. Decrease in desmoglein-1 (underlined in blue), but increase in corneodesmosome and CE initiation and reinforcement proteins in KtyI−/− and KtyII−/−K8 compare d with WT. Less transglutaminase (tgm-1 and -3) and arachidonate 12-lipoxygenase (alox12b) are cross-linked in both KtyI−/− and KtyII−/−K8 compared with WT. (B) Except repetin, fewer major CE structural proteins are cross-linked in both KO strains compared with WT. (C) Cross-linking of proteases and protease inhibitors involved in terminal differentiation and barrier function is similarly changed in KtyI−/− and KtyII−/−K8 compared with WTs. Cathepsin-D (Ctsd) and Spink5 are underlined in orange and are only altered in KtyII−/−K8. (D) Enhanced cross-linking of alarmins in KtyI−/− and KtyII−/−K8 compared with WTs (red hash marks hspa2, which is upregulated in KtyII−/−K8 only). *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
The proteases kallikreins 6 and 7 were slightly increased, whereas the reverse was observed for calpain 1 and caspase 14 in KIF-deficient CE (Fig. 8 C). Dysregulation of the protease inhibitors serpins B2, B5, and B12 and calpain 1 suggested spatial control of these enzymatic activities by the keratin scaffold to ensure CE assembly and homeostasis.
Genome-wide association and gene expression profiling studies have linked a subset of keratins to skin inflammation (Quigley et al., 2009; Roth et al., 2012b; Lessard et al., 2013). The increase in alarmins, a group of proteins involved in the pathogenesis of inflammatory diseases (Bianchi, 2007; Chan et al., 2012), has also been associated with tissue damage and immune activation after the loss of Krt16 (Nestle et al., 2009; Lessard et al., 2013). Elevated alarmin levels (Fig. 8 D), including nucleolin and S100A9 proteins (Sakaguchi et al., 2003; Lessard et al., 2013) and the anti-inflammatory annexin A1 (Lim and Pervaiz, 2007) or annexin A2, which interacts with filaggrin and has chaperone activity (Bunick et al., 2015), indicate a barrier rescue response in utero in the absence of KIF. Unlike predicted by the analysis of Krt16−/− mice (Lessard et al., 2013), upregulation of alarmins coincided with a strong increase in Krt16 (Fig. 8 D and Fig. S3, D and E). Moreover, several chaperones, including Hspb1 involved in epidermal differentiation, wound healing, and barrier formation, were upregulated in both KO mouse models. Similarly, cross-linked galectins 3 and 7 were increased in both KO CEs and are known to play a role in re-epithelialization on wound healing (Gendronneau et al., 2008; Panjwani, 2014). General activation of alarmins in KIF-deficient skin was also detected at transcript levels (Fig. S5 C).
These findings for the first time establish KIF as essential partners of a protein network that coordinates the spatiotemporal assembly of important CE proteins (loricrin, hornerin, and filaggrin 2), alarmins, and chaperones that cannot be compensated by upregulation of other constituents.
Keratin-dependent changes in mitochondrial lipids and proteins mediate elevated oxygen consumption and increased cellular ATP levels
Previous studies have documented a dependence of proper mitochondrial function on simple epithelial keratins, desmin, and vimentin and suggested context-dependent functions (Duan et al., 2009; Tao et al., 2009; Alvarado and Coulombe, 2014; Capetanaki et al., 2015; Chernoivanenko et al., 2015). Moreover, skin architecture and barrier function are severely impaired in murine genetic models of mitochondrial dysfunction (Baris et al., 2011; Hamanaka et al., 2013). In view of these data, we asked whether keratin scaffolding functions documented for the CE extended to organization and activity of mitochondria. Immunofluorescence analysis using Hsp60 antibodies revealed a keratin-dependent distribution of mitochondria. The increased immunofluorescence intensity in KtyI−/− suggested an increased quantity of mitochondria (Fig. 9 A). Moreover, mitochondria were scattered throughout the cytoplasm in granular layer keratinocytes of KtyI−/−, resulting in a punctate pattern, whereas they were predominantly aligned along cell borders in the granular layer of KtyI+/+ epidermis (Fig. 9 A, insets).
Cell-intrinsic and keratin-dependent changes in mitochondrial composition and activity. (A) Irregular distribution with punctate cytoplasmic localization (red arrowhead) of mitochondria in the epidermis of KtyI−/− E18.5 embryos. In WT controls, mitochondria are enriched in a diffuse pattern surrounding the nuclei in the spinous layer (red asterisk) and aligned to cell borders in the granular layer. The basement membrane is indicated by white dotted lines. Bars, 10 µm. (B) Western blotting of mitochondrial respiratory chain subunits and β-actin as loading control from extracts of KtyI+/+, KtyI−/−, and KtyI−/−K17 keratinocytes. (C) Quantification of B, normalized to loading control. Note the specific increase of NdufA9, NdufV2 (complex I), and Cox1 (complex IV) but not of CORE2 (complex III), SDHA (complex II), and α-subunit (complex V) in the absence of keratins. (D) Quantitative lipid analysis of purified mitochondria showing increased levels of PE and CL and a decrease of PC and PS in KtyI−/− compared with KtyI+/+ and KtyI−/−K14 rescue keratinocytes. (E) Increased oxygen consumption in KtyI−/− compared with KtyI+/+ and KtyI−/−K14 keratinocytes. (F) Cellular ATP levels are increased in KtyI−/− compared with KtyI+/+ and Krt14 re-expressing cells. *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
Cell-intrinsic and keratin-dependent changes in mitochondrial composition and activity. (A) Irregular distribution with punctate cytoplasmic localization (red arrowhead) of mitochondria in the epidermis of KtyI−/− E18.5 embryos. In WT controls, mitochondria are enriched in a diffuse pattern surrounding the nuclei in the spinous layer (red asterisk) and aligned to cell borders in the granular layer. The basement membrane is indicated by white dotted lines. Bars, 10 µm. (B) Western blotting of mitochondrial respiratory chain subunits and β-actin as loading control from extracts of KtyI+/+, KtyI−/−, and KtyI−/−K17 keratinocytes. (C) Quantification of B, normalized to loading control. Note the specific increase of NdufA9, NdufV2 (complex I), and Cox1 (complex IV) but not of CORE2 (complex III), SDHA (complex II), and α-subunit (complex V) in the absence of keratins. (D) Quantitative lipid analysis of purified mitochondria showing increased levels of PE and CL and a decrease of PC and PS in KtyI−/− compared with KtyI+/+ and KtyI−/−K14 rescue keratinocytes. (E) Increased oxygen consumption in KtyI−/− compared with KtyI+/+ and KtyI−/−K14 keratinocytes. (F) Cellular ATP levels are increased in KtyI−/− compared with KtyI+/+ and Krt14 re-expressing cells. *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001.
To investigate whether mitochondrial activity depended on keratins, we performed loss and gain of function studies using keratinocyte cell lines established from KtyI+/+ and KtyI−/− mice, in addition to KtyI−/− keratinocytes re-expressing different keratin pairs at levels representing ∼35% of the WT (KtyI−/−K14 and KtyI−/−K17; Homberg et al., 2015). Western blotting of mitochondrial ETC proteins revealed that complex I proteins NdufA9 and NdufV2 and complex IV protein Cox1 were increased in KtyI−/− cells (Fig. 9, B and C). In contrast, Sdha, Uqcrc2, and α-subunit, constituents of complexes II, III, and V, respectively, remained unaffected. Quantitative RT-PCR showed that these changes occurred at the posttranscriptional level (data not shown). Given that ETC activity is dependent on the lipid composition of mitochondria (Ellis et al., 2005; Böttinger et al., 2012; Tasseva et al., 2013), the phospholipid content of mitochondria isolated from KtyI+/+ and KtyI−/− keratinocytes was analyzed by mass spectrometry for phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylinositol (PI), phosphatidylserine (PS), phosphatidylglycerol (PG), phosphatidic acid (PA), and TLC for cardiolipin (CL; Tatsuta et al., 2014). This revealed that CL and PE were increased in KtyI−/− mitochondria when compared with KtyI+/+ (Fig. 9 D), whereas PC and PS were decreased.
Because ETC activity relies on PE and CL levels (Epand et al., 2007; Böttinger et al., 2012), oxygen consumption and ATP levels in KtyI+/+ and KtyI−/− cells, two major parameters of mitochondrial activity, were determined (Sabharwal and Schumacker, 2014). Oxygen consumption was increased by ∼25% (Fig. 9 E), and the ATP level increased even by ∼2.5-fold (Fig. 9 F) in KtyI−/− keratinocytes. Re-expression of Krt14 or Krt17 (KtyI−/−K14 and KtyI−/−K17) reduced both oxygen consumption and ATP content to levels similar to WT cells (Fig. 9, E and F). Therefore, mitochondrial activity is regulated in a cell-intrinsic and keratin-dependent manner, probably to supply energy for increased mechanical stress resistance and migration and invasive potential in absence of keratins (Seltmann et al., 2013a,b).
In summary, the comparative analysis of KtyI−/− and KtyII−/−K8 mice has uncovered an unanticipated role of a filamentous keratin scaffold in barrier formation and mitochondrial activity.
Discussion
Epithelia line body surfaces to provide structural support and serve as barriers against pathogens, oxidative stress, and dehydration. Failure to restore barriers results in inflammatory and immune disorders (Nestle et al., 2009; Schäfer et al., 2012). Here, we reveal an unprecedented and crucial role of filamentous keratins in epidermal barrier formation and show that known compensatory mechanisms fail to restore a functional barrier in their absence. The major defects in mice lacking individual keratins or the entire protein family are presented in Fig. 10 A. At the cellular level, we have also identified a KIF-dependent reliance of mitochondrial activity (Fig. 10 B). We propose a model whereby localized KIF scaffolds are necessary for the ordered CE assembly and for the correct composition and activity of mitochondria (Fig. 10 B).
Summary of major findings. (A) Comparison of major phenotypes in keratin-deficient mice (Reichelt et al., 2001, 2004; Reichelt and Magin, 2002; Kröger et al., 2011; Roth et al., 2012b; Bär et al., 2014). (B) Model integrating keratin filaments as scaffolds crucial for CE formation and mitochondrial composition/activity.
Summary of major findings. (A) Comparison of major phenotypes in keratin-deficient mice (Reichelt et al., 2001, 2004; Reichelt and Magin, 2002; Kröger et al., 2011; Roth et al., 2012b; Bär et al., 2014). (B) Model integrating keratin filaments as scaffolds crucial for CE formation and mitochondrial composition/activity.
A single keratin pair sustains embryonic development of simple epithelia
Previous work has established placental development, coinciding with an increased demand for nutrients, as a major developmental stage that depends on keratins from E9.5 (Hesse et al., 2000; Tamai et al., 2000; Jaquemar et al., 2003; Vijayaraj et al., 2009, 2010; Kröger et al., 2011). We demonstrate that expression of keratin pairs K8/K19 and K8/K18 in KtyII−/−K8 and KtyI−/− mice, respectively, overcomes this restriction point and maintains the formation of simple epithelia until birth, without noticeable adhesion and proliferation defects. Whether the considerable redundancy among simple epithelial keratins extends into adult life, when organ-specific immune and metabolic functions become important (Alam et al., 2013; Mor-Vaknin et al., 2013), remains to be studied. The finding that Krt8, which lacks Cys residues, fails to rescue skin fragility in Krt5−/− mice supports the idea that Cys-dependent cross-linking of selected epidermal keratins is necessary for keratin organization to provide strain resilience of stratified epithelia (Alvarado and Coulombe, 2014).
KIFs are crucial constituents and organizers of CE assembly
Previous data have suggested the concept of compensatory redundancy underlying CE composition aimed to maintain the epidermal barrier at several levels. For instance, a triple KO for periplakin, envoplakin, and involucrin is necessary to disrupt the barrier (Sevilla et al., 2007). On deletion of loricrin, an important CE constituent, a transient upregulation of Sprr2d, Sprr2h, and repetin, mediated by the antioxidant transcription factor Nrf2, was sufficient to restore the barrier (Koch et al., 2000; Huebner et al., 2012). Similarly, filaggrin KO mice have a mild alteration of their barrier integrity (Kawasaki et al., 2012). Even failure of tgm 1, causing perinatal death because of dehydration in mice, can be partly compensated on grafting neonatal tgm 1–deficient skin onto adult nude mice (Matsuki et al., 1998; Kuramoto et al., 2002). Finally, dysregulation of transcription factors and chromatin regulators, including Satb1, Klf4, Arnt, and AP1, are connected to hyperkeratosis and epidermal barrier defects (Segre et al., 1999; Geng et al., 2006; Fessing et al., 2011; Rorke et al., 2015).
The proteomic comparison of KtyI−/− and KtyII−/−K8 CEs demonstrates an absolute requirement of KIF, but not of nonfilamentous keratin isotypes, for distinct stages of CE assembly and barrier formation. This is underlined by the severe CE defects despite the presence of individual keratins Krt1, Krt77, and Krt75 in KtyI−/− or Krt10, Krt14, and Krt16 in KtyI−/−K8 mice (Fig. 7). Overall, absence of KIF leads to remarkably similar alterations of most CE constituents in both KO mice (Fig. 8), identifying loricrin, hornerin, and filaggrin 2, together with keratins, as essential CE components (Fig. 8 B). In their absence and contrary to the lack of loricrin (Huebner et al., 2012), upregulation of Nrf2 target genes in prenatal keratin KO epidermis failed to restore epidermal homeostasis (Figs. 5 and 6), narrowing down the Nrf2 protective activity on amniotic fluid metabolite exposure to a short window of time. Given that epidermal morphology of KtyI−/− and KtyII−/−K8 embryos is intact at E15.5 but distinctly altered at E18.5 (Fig. 2 A), increased Nrf2 could however contribute to the observed hyperkeratosis and barrier defects through several mechanisms (Schäfer et al., 2012, 2014).
The concomitant increased cross-linking of early CE components (desmoplakin, periplakin, envoplakin, plakophilins, and plakoglobin), which is in contrast with their downregulation or unaltered presence among total epidermal proteins (Fig. 8 A and Fig. S4 C) in both strains of keratin deficient mice, indicates that KIFs normally limit their cross-linking before the ordered incorporation of other CE components (Fig. 8 B and Fig. S5 A).
KIF-dependent CE deficiency implicates keratin involvement in barrier diseases
Terminal differentiation is regulated by cascades of proteases and protease inhibitors involved in the processing of tgm 1 and 3, profilaggrin, desmosomal constituents, and additional proteins (Zeeuwen, 2004; Samuelov and Sprecher, 2015). Remarkably, CE incorporation of distinct proteases and protease inhibitors was dysregulated in both strains of KO mice, suggesting that KIFs are involved in the balance between CE formation and desquamation. With the exception of kallikrein 6 and 7 involved in corneodesmosome degradation (Matsui and Amagai, 2015), other proteases like calpain 1, caspase 14, and cathepsin D were significantly decreased in absence of KIF (Fig. 8 C). Interestingly, animals deficient in the latter enzymes also showed epidermal barrier defects with altered processing of profilaggrin and tgm 1 (Egberts et al., 2004; Matsui and Amagai, 2015). These and our results provide a rationale to investigate mechanisms underlying hyperkeratosis in epidermolytic ichthyosis caused by Krt1, Krt2e, Krt9, and Krt10 mutations (Szeverenyi et al., 2008). Conversely, upregulation of the protease inhibitor Spink 5, of which genetic polymorphisms are associated with atopic dermatitis (Kato et al., 2003), implicates an involvement of keratins in barrier-related skin diseases.
Mutations of filaggrin and corneodesmosomal proteins, such as DSG1, are linked to inflammatory disorders, such as atopic dermatitis or multiple allergies and metabolic wasting (SAM syndrome), most probably via barrier defects allowing entry of environmental agents and provoking innate and adaptive immune responses (Palmer et al., 2006; Smith et al., 2006; Nestle et al., 2009; Oji et al., 2010; Samuelov et al., 2013; Matsui and Amagai, 2015; McAleer et al., 2015). Remarkably, SAM syndrome shares a cytokine profile similar to that reported for mice deficient in Krt1, Krt16, and Krt17, known to mediate innate immune responses through distinct pathways (Kim and Coulombe, 2010; Roth et al., 2012b; Lessard et al., 2013; Chung et al., 2015). The decrease of Dsg1 among cross-linked proteins of both KO strains (Fig. 8 A and Fig. S5 A) supports the hypothesis that keratin mutations also predispose to inflammatory skin conditions, including atopic dermatitis (Roth et al., 2012b). In this direction, the upregulation of alarmins may represent an attempt to restore barrier function and initiate inflammatory response in the absence of KIF (Fig. 8 D and Fig. S5, A and C). Among them, hspb1 and hsp90a1 have been linked to allergic contact hypersensitivity, inflammation, or wound healing (Crowe et al., 2013; Jayaprakash et al., 2015). Future work has to clarify how upregulation of alarmins depends on the status of Krt16 (Lessard et al., 2013).
KIFs regulate mitochondrial glycerophospholid content and restrain mitochondrial activity
Epidermal homeostasis also results from a balance between growth and differentiation, which are influenced by the activity of mitochondria (Baris et al., 2011; Hamanaka et al., 2013; Kloepper et al., 2015). Our localization and functional data indicate a link between keratins, mitochondrial localization, and activity (Fig. 9). We show a KIF-dependent regulation of mitochondrial ETC proteins with a notable increased amount of Ndufa9 and Ndufv2, which are essential for the catalytic function of the ETC complex I (Hoefs et al., 2012), and of the mitochondrial DNA-encoded Cox1, the central subunit of the catalytic core of complex IV acting as the terminal part of the ETC driving ATP synthesis by complex V (Fontanesi et al., 2008). Most interestingly, keratin re-expression in KtyI−/− cells restored ETC protein levels (Fig. 9, B and C), and keratin expression also affected the lipid composition of mitochondrial membranes (Fig. 9 D). CL and PE lipids were increased in absence of KIF and are known to stabilize mitochondrial ETC complexes, particularly for the larger supercomplexes (I and IV), and are therefore able to increase mitochondria activity (Haines and Dencher, 2002; Böttinger et al., 2012; Tasseva et al., 2013; Paradies et al., 2014; Tatsuta et al., 2014).
Although most mitochondrial lipids are synthesized in the endoplasmic reticulum and transported to the mitochondria (Rowland and Voeltz, 2012; Tatsuta et al., 2014), PE and CL are synthesized in the mitochondrial membrane (Tatsuta et al., 2014). We hypothesize that keratins affect the trafficking between the endoplasmic reticulum and mitochondria, with an impact on lipid distribution.
In summary, we have shown that KIF is essential for the formation of the CE and for the control of mitochondrial activity. Together with the dysregulation of a small subset of genes, including desmoglein 1, hornerin, and filaggrins, which are involved in inflammatory skin and systemic disorders, our study strongly supports an involvement of keratins in barrier disorders.
Materials and methods
Generation of KtyI−/− mice
Targeting the 3′ end of the keratin type I cluster
The hprt vector clone MHPN347p11 (Adams et al., 2004) was used. This contained an insert of 8.1 kb spanning 101, 202, 450–101, 208, and 417 bp on chromosome 11. The insert was flipped using AscI restriction sites and re-ligated to change the orientation of the LoxP site. A gap of 1.3 kb was generated with the double-cutter EcoNI restriction enzyme, re-ligated to restore the EcoNI restriction site, and yielded 1.5- and 1.6-kb arms of homology. The plasmid was linearized with EcoNI before targeting and transfected (200 µg) into AB2.2 cells (gift from A. Bradley, Wellcome Trust Sanger Institute, Cambridge, England, UK) at 3 µF and 800 V by electroporation. G418 selection was initiated 24 h after targeting at 350 µg/ml. Neomycin-resistant colonies were screened for homologous recombination; the plasmid without the gap served as a positive control. PCR primers spanned the gap region and vector backbone (Table S1). By PCR, nine clones were positive for the homologous recombination and were further analyzed by Southern blotting with a 497-bp probe specific to the gap region (Table S1). This identified a 5.8-kb band in the WT and one of 14.2-kb targeted NheI restriction fragments (Fig. S1 C). A clone positive for the homologous recombination was used to target the 5′ end of the keratin type I cluster.
Targeting the 5′ end of the keratin type I cluster
The mutagenic insertion and chromosome engineering resource hprt clone MHPP65n22 (Adams et al., 2004) was excised from the vector at the AscI sites flanking the insert and flipped to change the orientation of the LoxP site. This contained an 8.1-kb insert spanning 101,876,016–101,882,964 bp on chromosome 11. A gap of 2.4 kb was generated using MfeI and SnaB1 restriction sites to yield 0.8- and 5.1-Kb arms of homology (Fig. S1 C). The construct was re-ligated to restore the MfeI restriction site. Before targeting, the vector was linearized using MfeI and targeted into the positive clone of the previous section and selected in 3 µg/ml puromycin 24 h after electroporation. Homologous recombination was screened by PCR, with primers spanning the gap region and vector backbone (Table S1). Positive clones were confirmed by Southern blot analysis with a 433-bp probe designed within the gap region (Fig. S1 C) that identified 4.5-kb WT and 16.7-kb targeted HpaI restricted fragments.
Cre-mediated deletion of the keratin type I cluster
Positive clones for the deleted cluster were used to generate male chimeras by blastocyst injections (gift from R. Maniu, Universität Bonn, Bonn, Germany). Type I keratin floxed male C57BL/6 mice were crossed with 129S1hprt Cre hemizygous females. Littermates were screened using PCR for the type I keratin heterozygous mice, and resulting heterozygotes were in turn crossed to obtain 25% KtyI−/− mice (Fig. S1 B). Animal care and experimental procedures were in accordance with the institutional and governmental guidelines.
Generation of KtyII−/−K8 mice
Generation and genotyping of these mice were previously described as KtyII−/− mice (Bär et al., 2014). In brief, Krt8 transgenic FVB/N mice (Nakamichi et al., 2005) were backcrossed five times to C57BL/6 (KtyII+/+K8). Then, C57BL/6 KtyII+/− mice (Vijayaraj et al., 2009) were mated with KtyII+/+K8 mice to generate KtyII−/−K8 mice after two generations (Fig. S1 B).
RNA extraction, real-time PCR, and transcriptome profiling and gene set enrichment analysis
Total RNA was extracted from E18.5 embryo skin using TRIzol according to the manufacturer’s instructions (15596-026; Life Technologies), phenol/chloroform extracted, and precipitated, followed by DNaseI treatment (18047-019; Life Technologies) and purification with the RNeasy MinElute Cleanup kit (74204; QIAGEN). cDNA synthesis was performed with 2 µg total RNA using RevertAid H Minus First Strand cDNA Synthesis kit (K1631; Life Technologies), according to the supplier’s instructions. Semiquantitative PCR was performed using Taq polymerase (A600X; Promega). After reverse transcription, real-time PCR was performed with a Maxima SYBR Green/ROX qPCR Master Mix (K0221; Life Technologies), run on ABI 7500 instrument (Applied Biosystems), and analyzed with 7500 software v2.0.1. GAPDH was used as a reference. Primer sequences are listed in Table S2.
Using similarly prepared cDNA, skin transcriptome profiling was performed as described previously (Roth et al., 2012b) with the MouseWG-6v2.0 Expression BeadChip kit (Illumina) in triplicate from E18.5 embryos. Data analysis was based on the R Statistical language (2.8.0; R Development Core Team) and Beadstudio 3.1.1.0 software (Illumina). Data were quantile-normalized. A fold-change/p-value filter was used to select differentially expressed genes; P values <0.05, expression changes >twofold, and a difference between mean intensity signals greater than background were considered statistically significant. The false discovery rate of P values was adjusted by the Benjamini-Hochberg method. The full transcriptome profile will be published along with the characterization of corresponding mice.
For the gene set enrichment analysis, the Molecular Signatures Database (MSigDB database v3.0; Broad Institute) was used. To minimize the noise, the enrichment score of each set was calculated using root square means. P values were calculated using t tests.
Tissue preparation and histochemistry
Pregnant female mice with E18.5-d-old embryos were sacrificed, and dorsal skin, stomach, and intestine samples of embryos were either embedded in Tissue-Tek (Sakura) freezing medium and snap-frozen in isopentane precooled at −80°C for cryosections or fixed overnight in freshly prepared 4% formaldehyde in PBS and processed for routine paraffin embedding (Roth et al., 2012b). Sections were cut at 7–14 µm depending on the type of tissue and method. For hematoxylin/eosin staining, sections were deparaffinized for 30 min at 60°C, washed two times in xylol, progressively rehydrated in 2-min baths of ethanol solutions (2× 100%, 1× 95%, 1× 90%, 1× 80%, 1× 70%, and 1× 50%), and washed three times in distilled water before staining. Slides were incubated for 15 s in a hematoxylin solution (1:6; 109249; Merck), shortly washed in distilled water twice, rinsed for 10 min in cold tap water, incubated for 45 s in 0.25% eosin solution (E4382; Sigma-Aldrich), and shortly washed in distilled water twice. Sections were dehydrated again progressively in ethanol baths with increasing concentrations (1× 30 s 70%, 1× 30 s 95%, and 1× 10 s 100%) and 2× 5 min in xylol. Sections were then mounted in DPX-mounting medium (44581; Fluka).
Immunostaining of tissue samples
Frozen and paraffin-embedded tissues were cut with CM3050 S cryotome and RM2255 microtome (Leica), respectively. Sections were processed and stained as previously described (Vijayaraj et al., 2009; Kröger et al., 2013). The antibody and dyes used are listed in Table S3. For immunohistochemistry of NRF2 in the epidermis, 4-µm paraffin-embedded tissue sections were processed as previously described (Roth et al., 2012b). Secondary antibodies (HK-9KT; BioGenex) and revelation reagents (QP900-9; BioGenex; and K3468; Dako) were used according to the manufacturer’s protocol. Antibodies are listed in Table S3. Cell proliferation in the intestine and the epidermis was assessed by determining the percentage of Ki-67–stained cells among the total DAPI-stained epithelial cells.
Microscopy
Histologic analysis and immunohistochemistry were documented using Axioplan2 (Carl Zeiss) with 25/0.8 NA, 40/1.3 NA, or 63/1.25 NA oil immersion objectives. ApoTome.2 Imager.Z2, equipped with AxioCam 506 Color and AxioCam Mrm cameras (Carl Zeiss), was used for z-stacks of Ki-67 immunofluorescently Ki-67–labeled tissue sections using 25/0.8 NA and 40/1.3 NA oil immersion objectives. z-stacks of fluorescently labeled cell and tissues were collected with a Zeiss LSM 780 confocal microscope equipped with 40/1.3 NA or 63/1.46 NA oil immersion. Image analysis and processing were performed using ZEN software 2010 and 2011 (Carl Zeiss), AxioVision Rel.4.8 (Carl Zeiss), NIS-Elements (Nikon), and Photoshop CS5.1 (Adobe). For quantitation of epidermal thickness, between 12 and 20 randomly chosen areas per animal were measured using the AxioVision linear measurement tool.
Electron microscopy
Skin samples from E18.5 mice were excised and directly fixed in 4% formaldehyde/1% glutaraldehyde for 2 h and in 1% OsO4 for 1 h. Fixed samples were treated with 0.5% uranyl acetate in 0.05 M sodium maleate buffer, pH 5.2, for 2 h in the dark and thereafter dehydrated and embedded in araldite using acetone as the intermedium. Polymerization was performed at 60°C for 48 h. Semi- and ultrathin sections were prepared with an ultramicrotome (Leica) using a diamond knife. To enhance contrast, sections were treated with 3% uranyl acetate for 5 min and with 0.08 M lead citrate solution for 3 min. Images were taken on an EM10 (Carl Zeiss) with a digital camera (Olympus) using iTEM software (Olympus).
Toluidine blue dye skin permeability assay and CE isolation for microscopy
Skin permeability of E18.5 embryos was investigated by toluidine blue dye penetration assay (Hardman et al., 1998). Small pieces of the embryo tails were cut for positive control of the tests. Pups were then gently placed in sequential PBS-based baths with various methanol concentrations of 1 min each: 0%, 25%, 50%, 75%, 100% (2 min), 75%, 50%, 25%, and 0%. Embryos were then placed for 5 min in filtered 0.0125% toluidine blue solution before single wash in PBS and image acquisition with a stereomicroscope (SMZ1500; Nikon).
CEs were isolated as described previously (Jarnik et al., 1996). Total skin was incubated at 95°C for 10 min in CE extraction buffer (100 mM Tris-HCl, pH 8.5, 0.2% SDS, 20 mM DTT, and 5 mM EDTA). CEs were collected by centrifugation at 5,000 g. CEs were further washed in CE extraction buffer. Pelleted CEs were dissolved in 0.2% SDS solution and counted using a hemocytometer. Equal numbers of CEs from all specimens were placed on coverslips, and images were taken. The number of intact and altered CEs was counted.
CE extraction for mass spectrometry
For protein profiling, mouse epidermis was isolated by heat treatment (Macdiarmid and Wilson, 2001). After decapitation and appendage amputation, skins of E18.5 mouse embryos were isolated via dorsal longitudinal excision, heated 2 min at 55°C, placed 3 min in ice-cold PBS, and spread (after a gentle dab with a tissue to remove PBS) on a Petri dish with the epidermal side down. The dermis was then removed by gentle teasing using forceps, and the translucent epidermis was collected. Epidermis was placed in 1.5 ml 2% SDS and 0.1 M sodium phosphate, pH 7.8, solution (CE extraction buffer II [CE-II]) and vortexed and heated 10 min at 100°C. CEs were collected after addition of 5 ml CE-II and centrifugation at 5,000 g for 15 min at room temperature. CEs were washed two more times in 1.5 ml CE-II and heated at 70°C overnight. CEs were then washed three times, collected by centrifugation, and dried in a Speedvac (Univapo 100H; UniEquip). Pellets were then extracted four times at 90°C for 15 min in 2% sodium dodecanoate, 50 mM ammonium bicarbonate, and 25 mM dithiothreitol and were then alkylated with 50 mM iodoacetamide at room temperature in the dark with stirring (Rice, 2011). The digest was acidified with trifluoroacetic acid, extracted three times with ethyl acetate, brought to pH 8.0 with ammonia and ammonium bicarbonate buffer, digested with reductively methylated bovine trypsin (Rice et al., 2012), and analyzed using a Q-Exactive mass spectrometer (Thermo Fisher Scientific). Searching of the UniProt Database (20140310; 87,012 entries) was performed with X! Tandem (version CYCLONE, 2013.02.01.1; GPM), and Scaffold software (version 4.4.1.1; Avontus Software) was used to validate MS/MS-based peptide and protein identifications as previously described (Rice et al., 2013; Laatsch et al., 2014). False discovery rates were estimated as 0.2% for peptide and 2.6% for protein identifications based on a decoy peptide library. Results are illustrated as normalized weighted spectral counts after verification of the protein presence by exclusive counts in protein clusters with shared peptides. In samples with low Krt10, illustrated values are maximal.
Cell culture
Keratinocytes were isolated from the epidermis of E18.5 WTs and KtyI−/− and KtyII−/−K8 embryos and cultivated (Kröger et al., 2013). In brief, epidermal sheets were obtained from the E18.5 embryos by placing the skin in dispase (5 mg/ml) overnight at 4°C, followed by manual dissection to separate the epidermis from dermis. Keratinocytes were isolated by placing the epidermal sheet in PBS 0.025% trypsin and 0.02% EDTA for 5 min at 37°C. Cells were cultivated as described previously (Kröger et al., 2013; Seltmann et al., 2013b). KtyI−/− keratinocytes stably expressing mouse K14 or K17 (KtyI−/−K14 or KtyI−/−K17) were cultured in the same condition but with selecting agent in the media and were generated as described previously (Homberg et al., 2015).
Western blotting
Lysed tissue samples were kept in 5× Laemmli buffer containing protease inhibitor (78439; Thermo Fisher Scientific), homogenized with T10 basic ULTRA-TURRAX (IKA), and treated with repeated cycles of heating (95°C) and sonication to extract total proteins. Subsequent SDS-PAGE and Western blotting were performed as previously described (Vijayaraj et al., 2009). Primary and secondary antibodies are listed in Table S3.
Mitochondrial lipid quantitation
Mitochondria were isolated from cultured keratinocytes using magnetic beads coupled to anti-TOM22 antibodies as previously described (Hornig-Do et al., 2009). In brief, cells (107) were lysed in 1 ml of ice-cold PBS, including Complete Protease Inhibitor Cocktail Tablets (Roche), by shearing through a 29G needle ∼20 times. The crude cell lysate was incubated with 25 µl anti-TOM22 MicroBeads for 60 min at 4°C for magnetic labeling. The suspension was loaded onto a pre-equilibrated MACS Column (Miltenyi Biotec), which was placed in the magnetic field of a MACS Separator (Miltenyi Biotec). Columns were washed three times with 3 ml PEB buffer (PBS, pH 7.2, 2 mM EDTA, and 0.5% BSA). After removing the column from the magnetic field, retained mitochondria were eluted with 5 ml of PEB buffer. After centrifugation at ∼13,000 g for 1 min, the mitochondrial pellet was washed twice with 0.32 M sucrose, 1 mM EDTA, and 10 mM Tris-HCl and stored at –80°C. Protein concentration of the mitochondrial pellet was estimated using Bradford assay. For the analysis of PC, PE, PI, PS, PG, and PA by mass spectrometry, the equivalent of 75 µg of protein was suspended in 500 µl of water. Lipids were extracted using the “One-Step Extraction” described in Özbalci et al. (2013), a method modified from Bligh and Dyer (1959). Lipid extraction was done in the presence of 120–165 pmol of each of the following internal standards (Avanti Polar Lipids): PC 17:0-14:1, PC 17:0-20:4, PE 17:0-14:1, PE 17:0-20:4, PI 17:0-14:1, PI 17:0-20:4, PS 17:0-14:1, PS 17:0-20:4, PG 17:0-14:1, PG 17:0-20:4, PA 17:0-14:1, and PA 17:0-20:4. Dried lipid extracts were dissolved in 300 µl of methanol. 20 µl of the lipid extract in methanol were loaded into 96-well plates and diluted with 20 µl of 20 mM ammonium acetate in methanol. Lipid infusion and ionization was conducted using Nano-ESI chips with the TriVersa NanoMate operated by the ChipSoft Software (Advion) under the following settings: sample infusion volume: 14 µl, volume of air to aspirate after sample: 1 µl, air gap before chip: enabled, aspiration delay: 0 s, prepiercing: with mandrel, spray sensing: enabled, cooling temperature: 14°C, gas pressure: 0.5 psi, ionization voltage: 1.4 kV, and vent headspace: enabled. Prewetting was done once.
Mass spectrometric analysis was performed using the QTRAP 6500 (SCIEX) operated by Analyst 1.6.2. The following instrument-dependent settings were used: curtain gas, 20 psi; CAD gas, medium; and interface heater temperature, 100°C. PC analysis was performed in the positive ion mode by scanning for precursors of m/z 184 at a collision energy of 35 eV. PE, PS, PG, PI, and PA measurements were performed in the positive ion mode by scanning for neutral losses of 141, 185, 189, 277, and 115 D at CE of 25 eV. The value for the declustering potential was 100 V (Özbalci et al., 2013). Scanning was performed in a mass range of m/z 650–900 D and at a scan rate of 200 D/s. 61 MCA spectra were accumulated. Mass spectra were processed by the LipidView Software Version 1.2 (SCIEX) for identification and quantification of glycerophospholipids. Endogenous glycerophospolipids were quantified by normalizing their peak areas to those of the internal standards.
The analysis of CL was performed by TLC.
TLC analysis
Two suspensions of mitochondria in 1 ml of water (containing the equivalent of ∼200 µg of protein), 2 ml of methanol, and 1 ml of chloroform were added. Lipids were extracted for 24 h at 37°C. The liquid phase was separated by filtration, and the solvent was evaporated in a stream of nitrogen. The lipid residues were purified using a modification of the Bligh-Dyer procedure as previously described (Signorelli and Hannun, 2002). The lipid extract was applied to 20- × 10-cm-high performance TLC Silica Gel 60 plates (Merck), which were prewashed twice with chloroform/methanol 1:1 (vol/vol) and air dried for 30 min. Each lane of the TLC plate was loaded with the equivalent of 80 µg of protein (n = 3). Glycerophospholipids were separated using chloroform/methanol/glacial acetic acid 65:28:8 (vol/vol/vol) as the solvent system. For quantitative analytical TLC determination, increasing amounts of a CL standard (Sigma-Aldrich) were applied to the TLC plates in addition to the lipid samples. For detection of lipid bands, the TLC plates were sprayed with a phosphoric acid/copper sulfate reagent (15.6 g of CuSO4(H2O)5 and 9.4 ml of H3PO4 [85%, wt/vol] in 100 ml of water) and charred at 180°C for 10 min (Yao and Rastetter, 1985). Lipid bands were then quantified by densitometry using the TLC-Scanner 3 (CAMAG) at a wavelength of 595 nm.
Measurements of endogenous and uncoupled respiration
Keratinocytes were exponentially grown with medium change 24 h before measurement. Cells were resuspended after trypsinization at a density of 1–2 × 106 per ml in PBS at 37°C. Oxygen consumption was monitored using a Clark-type oxygen electrode (Hansatech Instruments) in PBS.
Measurement of cellular ATP content
ATP levels in viable cells were quantified using CellTiter-GloTM Luminescent Cell Viability assay kit (Promega) according to the manufacturer’s instructions. In brief, lyophilized enzyme/substrate mixtures (100 µl) were transferred to opaque 96-well microplates containing cell lysates. The microplates were then incubated at room temperature for 10 min to stabilize relative luminescence signals, which were then measured using a GENios Microplate Reader (Tecan). The concentration of ATP corresponding to relative luminescence signals is determined from an ATP standard curve. Protein concentration in the cell extract was estimated using BCA assay (Thermo Fisher Scientific). Cellular ATP levels were expressed as nanomoles per milligram of cell protein.
Statistical analysis
All the experiments were performed at least three times. Mean, SD, and t test were calculated using Excel software (Microsoft; *, P ≤ 0.05; **, P ≤ 0.01; ***, P ≤ 0.001). Graphs were generated using SigmaPlot software.
Online supplemental material
Fig. S1 describes the murine KtyI and KtyII gene clusters, the strategies used for the generation of the keratin KO mice, and the molecular characterization and the phenotypic appearance of KtyI+/+ and KtyI−/− animals at E18.5. Fig. S2 presents the analysis of keratins expression and localization in KtyII+/+K8 and KtyII−/−K8 intestine, similar histologic appearance of intestine and stomach simple epithelia in KtyI and/or KtyII animals at E18.5, and this way shows that keratin pairs K8/K18 or K8/K19 can sustain their development until birth. Fig. S3 displays epidermal histologic appearance and cell proliferation at E15.5 and E18.5, keratin mRNA levels in KtyI and KtyIIK8 WT and KO skin embryos at E18.5 (with upregulation of Krt6 and Krt16 in KOs), and major upregulated gene functions in the skin of KO animals according to transcriptome profiles at E18.5. Fig. S4 exhibits the similar desmosomal defects in KtyI−/− and KtyII−/−K8 skin at E18.5 via their analysis by immunofluorescence, transcriptome profiling, and Western blotting. Fig. S5 presents normalized data of the proteomic profiling in CEs from keratin WT and KO mice at E18.5 (allowing comparison of KtyI−/− and KtyII−/−K8), the detection of loricrin expression in KtyI+/+ and KtyI−/− mouse epidermis at E15.5, and the transcription level of several alarmins in the skin of keratin WT and KO mice at E18.5. Table S1 lists the primers used for the generation and the analysis of KtyI mice. Table S2 lists the primers used for the different quantitative PCR performed in this study. Table S3 presents all the dye and antibodies (with their source and working dilutions) used for the immunofluorescence and Western blot analysis of this study.
Acknowledgments
We thank Gabriele Baumbach, Maria Bust, Sabine Eisner, Romina Kühne, and Miriam Richter for technical assistance, and Michelle Salemi and Dr. Brett S. Phinney for mass spectrometry. We are also kindly indebted to Prof. Omary Bishr for providing K8 transgenic mice.
Work in the Magin laboratory is supported by the Deutsche Forschungsgemeinschaft (DFG; MA1316-15, MA1316-17, MA1316-19, MA1316-21, and INST 268/230-1) and the Translational Center for Regenerative Medicine Leipzig (No. 0315883). Work in the Leube laboratory is supported by the DFG (LE566-18-1) and by the START-Program of the Faculty of Medicine, Rheinisch-Westfälische Technische Hochschule Aachen. Work in the Wiesner laboratory is supported by the DFG (Wi 889/6-2). Mass spectrometry analysis was supported by National Institutes of Health grant 2 P42 ES04699 (R.H. Rice).
The authors declare no competing financial interests.
References
- CE
cornified envelope
- CL
cardiolipin
- E
embryonic day
- ETC
electron transport chain
- KIF
keratin intermediate filament
- KO
knockout
- Krt
keratin
- Kty
keratin type
- LCE
late cornified envelope protein
- PA
phosphatidic acid
- PC
phosphatidylcholine
- PE
phosphatidylethanolamine
- PG
phosphatidylglycerol
- PI
phosphatidylinositol
- PS
phosphatidylserine
- SPRR
small proline-rich protein
- WT
wild type
Author notes
P. Vijayaraj’s present address is Dept. of Pediatrics, University of California, Los Angeles, Los Angeles, CA 90095.
V. Kumar and J.-E. Bouameur contributed equally to this paper.