Maintenance of epithelial cell adhesion is crucial for epidermal morphogenesis and homeostasis and relies predominantly on the interaction of keratins with desmosomes. Although the importance of desmosomes to epidermal coherence and keratin organization is well established, the significance of keratins in desmosome organization has not been fully resolved. Here, we report that keratinocytes lacking all keratins show elevated, PKC-α–mediated desmoplakin phosphorylation and subsequent destabilization of desmosomes. We find that PKC-α activity is regulated by Rack1–keratin interaction. Without keratins, desmosomes assemble but are endocytosed at accelerated rates, rendering epithelial sheets highly susceptible to mechanical stress. Re-expression of the keratin pair K5/14, inhibition of PKC-α activity, or blocking of endocytosis reconstituted both desmosome localization at the plasma membrane and epithelial adhesion. Our findings identify a hitherto unknown mechanism by which keratins control intercellular adhesion, with potential implications for tumor invasion and keratinopathies, settings in which diminished cell adhesion facilitates tissue fragility and neoplastic growth.
Introduction
The interaction of keratin intermediate filaments (IFs) with desmosomes protects epithelia against mechanical injury, dehydration, and infections (Simpson et al., 2011). Desmosomes connect neighboring cells through desmosomal cadherins that associate with plakoglobin (PG), plakophilins 1–3 (PKP), and desmoplakin (DP). DP binds keratins via its C terminus, facilitating strong intercellular adhesion necessary to maintain tissue morphogenesis and architecture (Simpson et al., 2011).
Perturbation of the keratin–desmosome complex severely compromises cell and tissue integrity, exemplified by missense or loss-of-function mutations in corresponding genes of humans and mice (McMillan and Shimizu, 2001; Magin et al., 2004; Jonkman et al., 2005; Simpson et al., 2011). While K5 or K14 mutations affect the cytoskeleton, giving rise to epidermolysis bullosa simplex (EBS), mutations in DP and PKP1 disrupt intercellular adhesion, resulting in skin fragility (McMillan and Shimizu, 2001; Jonkman et al., 2005; Simpson et al., 2011). In several keratin knockout mice, desmosomes are affected to a limited extent (Magin et al., 1998; Hesse et al., 2000; Vijayaraj et al., 2009; Wallace et al., 2012). In contrast, loss of DP in extraembryonic tissues or in the epidermis causes a collapse of the keratin cytoskeleton and weakened intercellular adhesion (Gallicano et al., 1998).
In these settings, the respective contribution of keratins and desmosomal proteins and mechanisms by which they maintain architectural and signaling functions are not well understood. To complicate the issue, DP-deficient keratinocytes have fewer desmosomes and display alterations in keratin and actin organization (Vasioukhin et al., 2001). Thus, the role of keratins in desmosome maintenance and their contribution to desmosome-dependent adhesive strength remains unknown. The down-regulation of desmosomes preceding loss of E-cadherin during epithelial–mesenchymal transition (EMT) in some tumors (Dusek and Attardi, 2011) suggests an important role of the keratin–desmosome complex in the maintenance of an epithelial phenotype.
Epithelial remodeling during morphogenesis, wound healing, and hyperproliferative conditions requires transient down-regulation and redistribution of keratins and desmosomes to allow cell migration and tissue repair (Wallis et al., 2000; Coulombe, 2003; Thomason et al., 2012). Desmosome remodeling involves protein kinase C α (PKC-α), mediating DP phosphorylation and modification of desmosomal adhesion. PKC-α is necessary for both desmosome formation (Sheu et al., 1989; Godsel et al., 2005; Bass-Zubek et al., 2008) and disassembly during wound healing (Wallis et al., 2000; Thomason et al., 2012). The mechanisms upstream of PKC-α activation remain only partially understood.
To investigate the role of keratins in desmosome maintenance and intercellular epidermal adhesion, we analyzed murine keratinocytes lacking the entire set of keratins (KtyII−/−; Vijayaraj et al., 2009) or KtyII−/− cells reexpressing the single keratin pair K5/14. We found that keratins maintain desmosomes and adhesive strength. Without keratins, DP became phosphorylated and accumulated in the cytosol in a PKC-dependent manner, rendering epithelial sheets susceptible to mechanical stress. Our data support a sequestration model whereby keratins sequester RACK1, which can bind PKC-α and keeps it away from DP to limit its phosphorylation, thereby promoting desmosome stability/maintenance and intercellular adhesive strength.
Results and discussion
Altered desmosomes and reduced intercellular adhesion in KtyII−/− keratinocytes
Recently, we reported that absence of all keratins diminished the number of desmosomes, accompanied by cytosolic accumulation of DP in yolk sac and placental trophoblast cells (Vijayaraj et al., 2009; Kröger et al., 2011). To examine the contribution of keratins to cell integrity and desmosome maintenance in keratinocytes, KtyII−/− mice (Vijayaraj et al., 2009; Kröger et al., 2011) were mated with K8 transgenic mice (Toivola et al., 2008). This fully rescued the embryonic phenotype. The epidermis of these mice is completely devoid of keratin IF, as K8 remained restricted to simple epithelia. Similar to embryonic tissues, DP was reduced at the plasma membrane (PM) in E18.5 epidermis lacking all keratin IF, and accumulated throughout the cytosol (Fig. 1, A–B′).
Keratin-dependent mislocalization of desmosomes in vivo and in vitro. (A–B′) Mislocalization of DP in E18.5 KtyII−/− mouse skin compared with WT. K5 (C–C′′′) and K14 staining (D–D′′′) confirmed keratin in WT and rescue cells and its absence in KtyII−/− and GFP-transfected cells. (E–E′′′) Dsg and DP are reduced and irregular along the PM in KtyII−/− (E′ and E′′) compared with WT and rescue cells (E and E′′′). Cytosolic accumulation of desmosomal proteins in KtyII−/− cells (E′ and E′′). (F–F′′′) Double labeling of E-cadherin and α-catenin showed no obvious difference between WT (F and F′′′) and KtyII−/− (F′ and F′′). Bars: (A and B) 20 µm; (A′, B′, and C–F′′′) 10 µm.
Keratin-dependent mislocalization of desmosomes in vivo and in vitro. (A–B′) Mislocalization of DP in E18.5 KtyII−/− mouse skin compared with WT. K5 (C–C′′′) and K14 staining (D–D′′′) confirmed keratin in WT and rescue cells and its absence in KtyII−/− and GFP-transfected cells. (E–E′′′) Dsg and DP are reduced and irregular along the PM in KtyII−/− (E′ and E′′) compared with WT and rescue cells (E and E′′′). Cytosolic accumulation of desmosomal proteins in KtyII−/− cells (E′ and E′′). (F–F′′′) Double labeling of E-cadherin and α-catenin showed no obvious difference between WT (F and F′′′) and KtyII−/− (F′ and F′′). Bars: (A and B) 20 µm; (A′, B′, and C–F′′′) 10 µm.
To further analyze how keratins regulate desmosome maintenance, we isolated KtyII−/− and control keratinocyte lines expressing the normal set of keratins (WT) from corresponding strains of mice. To confirm keratin dependence, two additional cell lines, reexpressing the keratin pair K5/14 by reintroduction of K5 (“rescue cells”) or introducing GFP (“negative rescue control”), were generated by lentiviral transduction. K5/14 levels in rescue cells reached ∼13% of the controls (Seltmann et al., 2013). Genetically, KtyII−/− cells lacked all type II and maintained type I keratin genes (Fig. S1, G and H); at the protein level, no type I keratin was detectable (Vijayaraj et al., 2009; Seltmann et al., 2013). Irrespective of their keratin content, these cells showed a surprisingly normal morphology and did not display obvious growth differences. Immunofluorescence staining confirmed the absence of keratins and reexpression of K5/14 in rescue cells (Fig. 1, C–D′′′; Fig. S1, A–C′′′). At elevated Ca2+ concentrations, WT cells formed extensive keratin IFs connected to cell junctions. IFs formed between K5/14 in rescue cells resembled the controls with minor exceptions (Fig. 1, C′′′ and D′′′). This might result from cross-linking subsets of K14 molecules (Lee et al., 2012). No compensatory up-regulation of vimentin or other IFs, based on immunofluorescence, Western blot (WB), or transcriptome profiling was found (Fig. S1, D–D′′′; (Vijayaraj et al., 2009; unpublished data). Cortical actin fibers were significantly reduced in the absence of keratins. Conversely, cytosolic stress fibers were significantly increased in KtyII−/− cells (Fig. S1 E′) and decreased by the reexpression of K5/14 (Fig. S1 E′′′). Microtubule organization appeared very similar in all keratinocyte lines (Fig. S1, F–F′′′).
Confocal microscopy of KtyII−/− cells revealed a significant decrease in Dsg, DP, and PKP1 at the PM and their concomitant accumulation in the cytosol. Colocalization of Dsg/DP and DP/PKP1 suggested accumulation of half-desmosomes in the cytosol (Fig. S3, A–A′). Re-expression of K5/14 (Fig. 1, E–E′′′; Fig. S2, D–D′′′) reverted this phenotype. Considering the context, occurrence of half-desmosomes is likely to be different from endocytosis of individual desmosomal components (Green et al., 2010; Kitajima, 2013). Possibly, absence of keratins reduces the strength by which desmosomal cadherins associate, promoting internalization of half-desmosomes under steady-state conditions. The distribution of PG, a protein of adherens junctions (AJ) and desmosomes (Lewis et al., 1997; Fig. S2, C–C′′′), was not obviously affected by the cells’ keratin status. WB uncovered a down-regulation of desmosome-specific proteins, ranging from 79% for Dsg1/2 to 32% for DP and 36% for PKP1, but not for PG (Fig. S2, E and F; Löffek et al., 2012). PKP2 is expressed at extremely low levels in all keratinocytes described here (Fig. S2 E). The notion that keratinocytes with severe K5 and K14 mutations show a decrease of desmosomal proteins (Liovic et al., 2009) supports a mechanistic role of keratins in desmosomal protein regulation. This decline was not rescued by reexpression of K5/14. This might be due to (1) a need for a threshold level of keratins engaged in multiple desmosomal protein interactions to stabilize them; (2) the keratin isotypes present; or (3) a potential involvement of posttranslational keratin modifications that may not be present in rescue cells. Alternatively, a potential cross talk between the type II keratin gene locus and genes controlling/encoding desmosomal proteins might be disturbed (Ferone et al., 2013). To address a possible cross talk between desmosomes and AJs (Vasioukhin et al., 2001), distribution and expression of E-cadherin, α-catenin, β-catenin, and p120, involved in AJ maintenance and linkage to the actin cytoskeleton (Niessen et al., 2011), were studied. No major difference in the distribution of these proteins was found (Fig. 1, F–F′′′). The unaltered levels of AJ proteins (Fig. S2 F) indicated that keratins affect primarily desmosomal proteins. Surface biotinylation established a 50% reduction of Dsg2 at the PM, restored by K5/14 reexpression (Fig. 2 A).
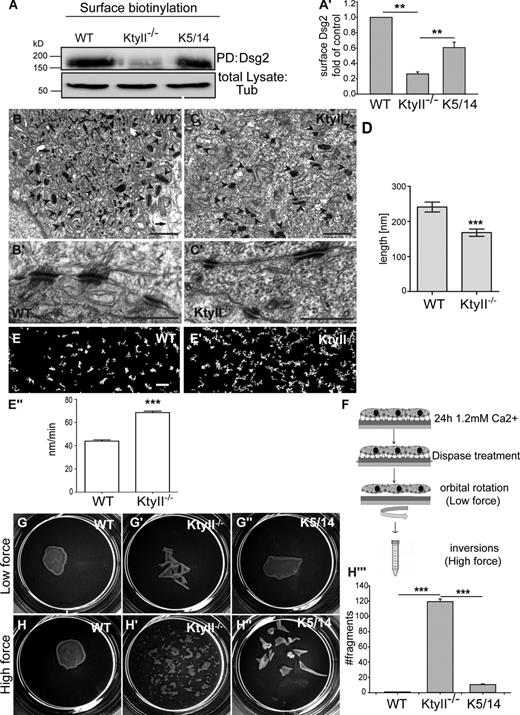
Desmosome size and epithelial sheet stability depend on keratins. (A–A′) Surface biotinylation and WB showed a 60% reduction of Dsg2 at the PM in KtyII−/− compared with WT and rescue cells. Data (mean ± SEM, n = 3) are fold change relative to WT. (B–D) EM revealed smaller and fewer desmosomes (arrowheads) in KtyII−/− (C–C′) compared with controls (D–D′). Arrows mark keratins. (D) 30% reduced desmosome size in KtyII−/− (n = 46) compared with WT (n = 52). (E–E′′) Transfected Dsg2-eGFP demonstrated elevated Dsg motility in the PM of KtyII−/− cells. Graphs show Dsg2 tracks in WT and KtyII−/− cells (Fig. S1). (E′′) Histogram: motion of desmosomes within PM elevated by 48% in KtyII−/− (79 nm/min, n = 449) compared with WT cells (53 nm/min, n = 319). (F) Model of dispase assay. (G–H′′′) Epithelial sheets of WT (G and H) and rescue cells (G′′ and H′′) resisted mechanical stress, not so KtyII−/− cells (G′ and H′). (G′′′) Quantification of stress assay (n = 3). Bars: (B and C) 1 µm; (B′ and C′) 500 nm; (E–E′) 2 µm.
Desmosome size and epithelial sheet stability depend on keratins. (A–A′) Surface biotinylation and WB showed a 60% reduction of Dsg2 at the PM in KtyII−/− compared with WT and rescue cells. Data (mean ± SEM, n = 3) are fold change relative to WT. (B–D) EM revealed smaller and fewer desmosomes (arrowheads) in KtyII−/− (C–C′) compared with controls (D–D′). Arrows mark keratins. (D) 30% reduced desmosome size in KtyII−/− (n = 46) compared with WT (n = 52). (E–E′′) Transfected Dsg2-eGFP demonstrated elevated Dsg motility in the PM of KtyII−/− cells. Graphs show Dsg2 tracks in WT and KtyII−/− cells (Fig. S1). (E′′) Histogram: motion of desmosomes within PM elevated by 48% in KtyII−/− (79 nm/min, n = 449) compared with WT cells (53 nm/min, n = 319). (F) Model of dispase assay. (G–H′′′) Epithelial sheets of WT (G and H) and rescue cells (G′′ and H′′) resisted mechanical stress, not so KtyII−/− cells (G′ and H′). (G′′′) Quantification of stress assay (n = 3). Bars: (B and C) 1 µm; (B′ and C′) 500 nm; (E–E′) 2 µm.
EM-based quantitation confirmed a 30% reduction in desmosome size without keratins (Fig. 2, B–D). The morphology of KtyII−/− and WT desmosomes was unaltered (Fig. 2, B′ and C′). Video microscopy of transgenic Dsg2-eGFP in WT and KtyII−/− cells uncovered an elevated motility of desmosomes in the PM where it colocalized with endogenous DP (Fig. 2, E–E′′; Fig. S3 B′′). Accumulation of desmosomal proteins in the cytoplasm and elevated motility of desmosomes in KtyII−/− cells suggested a dependence on keratins, prompting the question whether they affected desmosomal adhesive strength and integrity of epithelial sheets upon mechanical stress (Vasioukhin et al., 2000; Huen et al., 2002). 24 h after calcium exposure an intact epithelial sheet of WT cells lifted off the dish upon mechanical stress application (Fig. 2, G, H′, and H′′′). In contrast, KtyII−/− cells displayed a ∼100× increase in fragmented sheets (Fig. 2, G′, H′, and H′′′), rescued to a significant extent by ∼13% of WT keratin content (Fig. 2, G′′, H′′, and H′′′). This suggests the usefulness of EBS therapy approaches aimed at partial restoration of intact keratins (Roth et al., 2012a).
Desmosomes assemble with similar kinetics in KtyII−/− and in WT cells
To dissect mechanisms underlying altered distribution and functionality of desmosomes in KtyII−/− cells, we monitored junction assembly after CaCl2 addition over 48 h. This revealed that desmosomes assembled with similar speed to WT cells up to 12 h after calcium treatment (Fig. 3, A–B′′). However, 24 h into junction formation, DP had redistributed to a significant extent into a cytosolic pool (Fig. 3, A′′′–B′′′ and C), suggesting that keratins might be primarily involved in desmosome maintenance.
Assembly and disassembly of cell junctions. (A–B′′′) Similar assembly rates in WT and KtyII−/− cells up to 12 h. Increased cytosolic DP in KtyII−/− cells after 24 h (B′′′). (C) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (D–F′) DP staining upon DMSO or Gö6976 treatment before and after Ca2+ depletion. Re-localization of DP to PM and decreased desmosome internalization to WT levels upon Gö6976. (G) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (H) Surface biotinylation and WB showed increased surface localization of Dsg2 after Gö6976 treatment in KtyII−/− cells. The first two control lanes (WT and KtyII−/− with no drug treatment) are the same as shown in Fig. 5 G because the treated samples were loaded on the same blot. (I–I′′) Blocking PKC activity in KtyII−/− cells stabilized cell–cell contacts against mechanical stress after dispase treatment (n = 3). Bars, 10 µm.
Assembly and disassembly of cell junctions. (A–B′′′) Similar assembly rates in WT and KtyII−/− cells up to 12 h. Increased cytosolic DP in KtyII−/− cells after 24 h (B′′′). (C) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (D–F′) DP staining upon DMSO or Gö6976 treatment before and after Ca2+ depletion. Re-localization of DP to PM and decreased desmosome internalization to WT levels upon Gö6976. (G) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (H) Surface biotinylation and WB showed increased surface localization of Dsg2 after Gö6976 treatment in KtyII−/− cells. The first two control lanes (WT and KtyII−/− with no drug treatment) are the same as shown in Fig. 5 G because the treated samples were loaded on the same blot. (I–I′′) Blocking PKC activity in KtyII−/− cells stabilized cell–cell contacts against mechanical stress after dispase treatment (n = 3). Bars, 10 µm.
PKC-α–mediated phosphorylation of DP is enhanced in KtyII−/− cells and induces desmosome disassembly
PKC-α regulates DP phosphorylation in an unknown manner (Hobbs and Green, 2012). Phosphorylation of DP at Ser2849, a site close to the keratin-binding domain of DP (Choi et al., 2002), is involved in desmosome assembly and remodeling in response to wound healing (Wallis et al., 2000; Green and Simpson, 2007). Although the phospho-deficient mutant DPS2849G impairs desmosome formation by retaining DP on IFs (Stappenbeck et al., 1994; Godsel et al., 2005; Bass-Zubek et al., 2008), once incorporated into desmosomes this mutant confers strong intercellular adhesion on keratinocyte cell sheets (Hobbs and Green, 2012). To test whether keratins are involved in PKC-α–mediated destabilization of desmosomes, we treated KtyII−/− with the PKC inhibitor Gö6976 (Wallis et al., 2000). This resulted in stabilization of DP at the PM in KtyII−/− cells, whereas vehicle-treated cells retained DP in the cytosol (Fig. 3, D–G). Given the dual function of PKC-α in desmosome assembly and disassembly (Hobbs and Green, 2012), the impact of elevated DP phosphorylation on desmosomal protein endocytosis was studied. To test this, junction formation was allowed for 24 h after Ca2+ addition, followed by EGTA-mediated Ca2+ depletion to induce internalization of Ca2+-dependent cell junction proteins (Kartenbeck et al., 1982). 1 h after EGTA addition, desmosomal proteins accumulated further in the cytoplasm of KtyII−/− (Fig. 3, E′ and G); in WT cells, however, internalization was much slower (Fig. 3, D′ and G). Co-stainings of Dsg and EEA1 or Lamp2 revealed accumulation of internalized desmosomal proteins in a compartment different from early endosomes or lysosomes (Fig. S3, A′′–A′′′). Treatment of KtyII−/− cells with Gö6976 slowed down desmosome internalization to WT levels (Fig. 3, F′ and G), in agreement with surface biotinylation (Fig. 3 H), and rescued epithelial sheet integrity to almost WT levels in the mechanical stress assay (Fig. 3, I–I′′).
For further evidence of PKC-α–mediated phosphorylation of DP in the absence of keratins, DP was immunoprecipitated from KtyII−/− and WT cells using a P-Ser antibody. Substantial amounts of P-DP were enriched in total protein lysates from KO cells, but hardly any P-DP was detected in WT lysates (Fig. 4 A). To confirm PKC-α involvement, the experiment was repeated using Gö6976 (Fig. 4 A), and by PKC-α knockdown (Fig. 4 B), both of which reduced P-DP to control levels. In addition, coimmunoprecipitation (Co-IP) revealed elevated levels of PKC-α relative to DP in KtyII−/− cells compared with WT cells (Fig. 4 C). Moreover, PKC-α was significantly enriched in PM fractions from KtyII−/− compared with WT cells. PKC-α inhibition diminished its PM localization (Fig. 4, D–D′). In the presence of keratins, PKC-α–mediated DP phosphorylation is low, resulting in stable desmosomes. To substantiate P-DP–dependent desmosome disassembly, KtyII−/− cells were transfected either with wild-type DP-GFP or mutant DPS2849G-GFP (Stappenbeck et al., 1994). Upon Ca2+ depletion DPS2859G-GFP was more stable at the PM compared with DP-GFP in KtyII−/− cells (Fig. 4, E–G). Full reversion may depend on additional DP phosphorylation sites to be characterized.
PKC-α–mediated phosphorylation of DP causes desmosome disruption in KtyII−/− cells. (A–B) IP with α-pSer antibodies followed by WB demonstrated PKC-α–mediated phosphorylation of DP in KtyII−/− cells. (C) PKC-α and DP interaction in absence of keratins (coIP, n = 2). (D–D′) Cell fractionation and WB showed increased PKC-α in PM fractions in KtyII−/− cells. Data (mean ± SEM, n = 3) are fold change relative to WT. (E–F′) Transfection of KtyII−/− cells with DPWTGFP or DPS2849GGFP. DPS2849GGFP showed increased stability at PM compared with DPWTGFP. (G) Ratios of average pixel intensities for DP at PM and cytosol. (H–I′) Double labeling of K5 (green) and Rack1 (red) showed partial colocalization in WT cells (H–H′′), in contrast to diffuse cytosolic localization of Rack1 in KtyII−/− cells (I). (J–K) PLA showed interaction between K5 and Rack1. PLA signals overlapped with keratins in WT cells (J–J′′). Very few PLA signals were observed in KtyII−/− cells (K). Bars, 10 µm. (L) Co-IP of PKC-α and Rack1 revealed significant association in both cell lines and interaction of K5 and Rack1 in WT cells (n = 2).
PKC-α–mediated phosphorylation of DP causes desmosome disruption in KtyII−/− cells. (A–B) IP with α-pSer antibodies followed by WB demonstrated PKC-α–mediated phosphorylation of DP in KtyII−/− cells. (C) PKC-α and DP interaction in absence of keratins (coIP, n = 2). (D–D′) Cell fractionation and WB showed increased PKC-α in PM fractions in KtyII−/− cells. Data (mean ± SEM, n = 3) are fold change relative to WT. (E–F′) Transfection of KtyII−/− cells with DPWTGFP or DPS2849GGFP. DPS2849GGFP showed increased stability at PM compared with DPWTGFP. (G) Ratios of average pixel intensities for DP at PM and cytosol. (H–I′) Double labeling of K5 (green) and Rack1 (red) showed partial colocalization in WT cells (H–H′′), in contrast to diffuse cytosolic localization of Rack1 in KtyII−/− cells (I). (J–K) PLA showed interaction between K5 and Rack1. PLA signals overlapped with keratins in WT cells (J–J′′). Very few PLA signals were observed in KtyII−/− cells (K). Bars, 10 µm. (L) Co-IP of PKC-α and Rack1 revealed significant association in both cell lines and interaction of K5 and Rack1 in WT cells (n = 2).
To further investigate how PKC-α activation involves keratins, Rack1, involved in the spatio-temporal regulation of PKC isoforms, was analyzed (Adams et al., 2011). Confocal imaging revealed significant colocalization with keratin IF in WT cells (Fig. 4, H–H′′), but a scattered distribution in KtyII−/− cells (Fig. 4, I–I′). Next, a proximity ligation assay (PLA) was performed. This confirmed extensive direct Rack1–K5 interactions in WT (Fig. 4, J–J′′), absent in KtyII−/− (Fig. 4 K) or in control cells (Fig. 3, E–E′). Moreover, Rack1 and PKC-α were coimmunoprecipitated from both KtyII−/− and WT cells (Fig. 4 L). Together with the presence of K5 in IPs from WT lysates (Fig. 4 L), this suggests that keratins sequester PKC-α through Rack1, thereby limiting DP phosphorylation necessary for stable desmosomes.
Keratins control dynamin-dependent endocytosis of desmosomal proteins
Dsg is predominantly internalized through caveolin-dependent (Resnik et al., 2011; Brennan et al., 2012) or raft-dependent mechanisms (Delva et al., 2008). As caveolin-dependent pathways rely on dynamin for vesicle fission, blocking its activity by the inhibitor Dynasore (Macia et al., 2006) should prevent endocytosis of desmosomes. In fact, Dynasore treatment reestablished the distribution of desmosomes in KtyII−/− cells even under conditions of cell adhesion or after Ca2+ depletion (Fig. 5, A–C′ and E). Crucially, Dynasore blocked endocytosis to such an extent in KtyII−/− cells that Dsg1/2 levels were restored at the PM nearly to those determined in WT cells (Fig. 5 G). Most importantly, inhibition of endocytosis restored desmosome functionality during the mechanical stress assay, whereas vehicle-treated cell sheets remained susceptible to mechanical stress (Fig. 5, H–H′′). To examine whether enhanced endocytosis and protein degradation with subsequent consequences on cell adhesion were keratin dependent, experiments were performed in K5/14-expressing rescue cells. As expected, K5/14 stabilized desmosomes under elevated calcium conditions and reverted Dsg1/2 internalization to WT levels (Fig. 5, D–D′ and E). In support, reexpression of K5/14 in the presence of CHX extended Dsg2 half-life time compared with KtyII−/− cells (Fig. 5 F; Fig. S3 C). Dsg2 degradation was blocked both by lysosomal (ELP) and proteasomal (MG132) inhibitors in KtyII−/− cells, suggesting complex degradation mechanisms of desmosomes (Fig. 3 D). IP with a P-Ser antibody after Dynasore treatment revealed a reduction of P-DP to control levels in KtyII−/− cells (Fig. 5 I), suggesting either limited susceptibility by PKC-α or rapid de-phosphorylation of DP at the PM upon blocked endocytosis. Thus, PKC-α–mediated phosphorylation of DP is followed by rapid internalization of desmosomal proteins in the absence of keratins.
Keratins affect dynamin-dependent endocytosis. (A–D′) DP staining revealed almost complete desmosome internalization in KtyII−/− cells after EGTA treatment (B′), whereas internalization is slower in controls (A′). Blocking endocytosis or reexpression of K5/14 in KtyII−/− cells induced relocalization of junctional proteins to PM in high Ca2+ conditions (C and D) as well as 1 h after EGTA incubation (C′ and D′). (E) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (F) WB of total protein lysates after EGTA treatment demonstrated accelerated Dsg2 degradation in KtyII−/− cells, reverted by reexpression of K5/14. Data (mean ± SEM, n = 3) are fold change relative to control. (G) Surface biotinylation and WB showed increased surface localization of Dsg2 after Dynasore treatment in KtyII−/− cells. The first two control lanes (WT and KtyII−/− with no drug treatment) are the same as shown in Fig. 3 H because the treated samples were loaded on the same blot. (H–H′′) Blocking endocytosis in KtyII−/− cells restored mechanical strength after dispase treatment (n = 3). (I) IP with pSer antibodies followed by WB: reduction of P-DP in KtyII−/− cells after Dynasore treatment (n = 2). (J) Model suggesting keratins regulate PKC-α–mediated phosphorylation of DP via RACK1.
Keratins affect dynamin-dependent endocytosis. (A–D′) DP staining revealed almost complete desmosome internalization in KtyII−/− cells after EGTA treatment (B′), whereas internalization is slower in controls (A′). Blocking endocytosis or reexpression of K5/14 in KtyII−/− cells induced relocalization of junctional proteins to PM in high Ca2+ conditions (C and D) as well as 1 h after EGTA incubation (C′ and D′). (E) Ratios of average pixel intensities for DP at PM and cytosol (n = 50). (F) WB of total protein lysates after EGTA treatment demonstrated accelerated Dsg2 degradation in KtyII−/− cells, reverted by reexpression of K5/14. Data (mean ± SEM, n = 3) are fold change relative to control. (G) Surface biotinylation and WB showed increased surface localization of Dsg2 after Dynasore treatment in KtyII−/− cells. The first two control lanes (WT and KtyII−/− with no drug treatment) are the same as shown in Fig. 3 H because the treated samples were loaded on the same blot. (H–H′′) Blocking endocytosis in KtyII−/− cells restored mechanical strength after dispase treatment (n = 3). (I) IP with pSer antibodies followed by WB: reduction of P-DP in KtyII−/− cells after Dynasore treatment (n = 2). (J) Model suggesting keratins regulate PKC-α–mediated phosphorylation of DP via RACK1.
The discovery of PKC-α–mediated phosphorylation of DP, dependent on Rack1–keratin interactions, revealed a hitherto unknown function of keratins in maintenance of desmosomes and cohesion of epithelial tissues (Fig. 5 J). Further, our data strongly support a major role of the keratin-dependent DPS2859 state in hyperadhesion of desmosomes (Hobbs and Green, 2012; Thomason et al., 2012). It remains to be shown how hyperadhesion and DP phosphorylation are regulated in the presence of keratins and whether keratin isotypes act in a specific manner as one might expect in view of highly regulated keratin expression during epidermal differentiation. Our findings raise a number of issues, including a potential role of the desmosome–keratin complex as a force sensor, a topic now addressable in our new cell models. Our data suggest that spatiotemporal control of DP phosphorylation involves keratins and is crucial for desmosome maintenance and internalization during wound healing and tissue regeneration. Further, the data implicate that reduced cell adhesion contributes to pathomechanisms in keratinopathies (Coulombe and Lee, 2012). Given that reduction of desmosomes precedes loss of E-cadherin in some tumors (Dusek and Attardi, 2011), our findings support the hypothesis that expression and organization of keratins maintain a barrier against EMT through several mechanisms including maintenance of intercellular adhesion.
Materials and methods
Cell culture
Keratinocytes were isolated from the epidermis of newborn C57Bl6 KtyIIfl/− and KtyIIfl/+ mice using 0.025% trypsin/0.02% EDTA/PBS and manual dissection, and spontaneous immortalization was permitted (see chapter 5 in Turksen, 2005. Cells were cultivated in complete FAD media (0.05 mM CaCl2) on collagen I–coated (rat tail; BD) 6-well plates (coated for 1 h at room temperature and then washed with 1× PBS) at 32°C, 5% CO2 on a mitomycin C–inactivated 3T3 Swiss NIH feeder layer. Upon confluence, keratinocytes were split 1:2. After passage 4, they were passaged without feeders and after 5–6 passages, cells immortalized spontaneously. C57Bl6 KtyIIfl/− keratinocytes were transfected using the Amaxa Nucleofector kit for primary endothelial cells according to the manufacturer’s protocol program T27: per transfection 106 cells were used and 4 µg of 1 µg/µl super-coil plasmid DNA of pcCre+ Hygro (Magin laboratory). After selection (20 µg/ml hygromycin solution) single colonies were picked and tested for keratin expression using K5 immunofluorescences. Keratin-free keratinocytes stably expressing K5/14 or GFP were generated by lentiviral transduction essentially as described previously (Stöhr et al., 2012; Seltmann et al., 2013). Lentiviruses for transduction of cDNAs were generated using the Lenti-X lentiviral expression system (Takara Bio Inc.). For virus production, HEK293T cells were cotransfected by CaPO4 precipitation with pMD2.G (no. 12259; Addgene), psPAX2 (no. 12260; Addgene), and pLVX-puro plasmids containing either eGFP or mK5 cDNA. Lentiviruses were purified 48 h after transfection by Lentivirus Concentrator (Takara Bio Inc.) according to the manufacturer’s instructions. Keratinocytes were incubated with viruses for 24 h. Resulting cell lines represent pools of puromycin-selected lentiviral transfectants. For all experiments, if not stated otherwise, cells were switched to high calcium medium (FAD plus 1.2 mM CaCl2) for at least 48 h. cDNAs encoding DPWTGFP or DPS2849GGFP (gifts of K. Green, Northwestern University, Chicago, IL) were transfected using Xfect Transfection reagent (Takara Bio Inc.) following the manufacturer’s protocol. SiPKC-α and control siRNA were purchased from Santa Cruz Biotechnology, Inc. siRNA were transfected using Xfect siRNA Transfection reagent (Takara Bio Inc.) following the manufacturer’s protocol.
Genomic PCR
The primers and PCR conditions have been described previously (Tonack et al., 2004; Lu et al., 2005).
Immunofluorescence analysis
Coverslips were placed into culture dishes before collagen coating. For keratin staining, cells were fixed for 5 min in −20°C methanol, 30 s in −20°C acetone, otherwise cells were fixed for 15 min in 4% formalin/PBS, made freshly from PFA, at 4°C, washed in TBS, and then permeabilized for 5 min in 0.25% Triton/PBS. After a TBS wash, they were incubated for 1 h or overnight at 4°C with primary antibodies diluted in TBS/1% BSA. Secondary antibodies at appropriate dilutions were applied and incubated for 30–60 min. Nuclei were counterstained using 1:1,000 diluted DAPI. F-actin was stained with Promofluor-488 phalloidin (PromoKine). Coverslips were mounted with Prolong Gold (Invitrogen).
Immunofluorescence microscopy and data processing
Image stacks were collected with a confocal microscope (LSM 780; Carl Zeiss) with 40×/1.3 NA or 63×/1.46 NA oil immersion objectives or an AxioImager (Carl Zeiss) equipped with Apotome2 with 40×/1.3 NA or 63×/1.4 NA oil immersion objectives and an AxioCam MRm (Carl Zeiss). Image analysis and processing were performed using Zen Software 2010 (Carl Zeiss), Zen 2011 Blue software (Carl Zeiss), and Photoshop CS4 (Adobe) software. LUT (lookup table; brightness and gamma) was adjusted using Photoshop. Fluorescence pixel intensity of DP was determined by measuring the average pixel intensity in a region of interest relative to the area of this region using Zen 2011 Blue software. To compare DP intensity at the PM relative to the cytosol, the ratio between the PM intensity and the cytosol of the same cell was calculated. Raw unsaturated 12-bit images were used for all quantifications.
Cell surface biotinylation
Cells were seeded on collagen I–coated 10-cm dishes. 24 h after plating, the medium was switched to high calcium medium for 48 h. Cells were surface labeled with 2 mg/ml cell-impermeable EZ-Link Sulfo-NHS-SS-Biotin (Thermo Fisher Scientific) following the manufacturer’s instructions. Cells were washed twice with PBS and lysed with 500 µl of cold RIPA buffer (10 mM Na2HPO, 150 mM NaCl, 5 mM EDTA, 2 mM EGTA, 1% Triton X-100, 0.1% SDS, 0.5% sodium deoxycholate, pH 7.5, and protease and phosphatase inhibitor cocktails; Thermo Fisher Scientific). Lysates were incubated with 100 µl streptavidin beads for 2 h. Biotinylated PM proteins were finally eluted by SDS-PAGE buffer and subsequently analyzed by WB. The amount of PM-bound Dsg2 was calculated as the ratio between biotinylated proteins to total amount of Dsg2.
Electron microscopy
2 × 105 cells were seeded on collagen cell carriers (Viscofan BioEngineering, Weinheim, Germany) and grown to confluency. 48 h before fixation, the medium was supplemented with calcium to a final concentration of 1.2 mM. Cells were washed twice with PBS and incubated in fixative (4% formaldehyde, 1% glutaraldehyde, and 50 mM K-cacodylate, pH 7.2) for 2 h and 1 h in 1% OsO4. Cells were then treated with 0.5% uranyl acetate in 0.05 M sodium maleate buffer (pH 5.2) for 1.5 h in the dark and thereafter dehydrated and embedded in araldite using acetone as medium. Polymerization was performed at 60°C for 48 h. Ultrathin sections were prepared with an ultramicrotome (Leica) equipped with a diamond knife. Sections were treated with 3% uranyl acetate for 5 min and with 0.08 M lead citrate solution for 3 min. Images were taken on a microscope (model EM10; Carl Zeiss) with a digital camera (Olympus) using iTEM software (Olympus).
Analysis of desmosomal protein motility
To quantify the lateral motility of desmosomes within the PM, desmoglein 2–labeled desmosomes were tracked in cells grown on top of each other in adjoining cell regions, thereby facilitating a top view of adhesion sites. WT and KtyII−/− cells expressing Dsg2-eGFP were recorded every 30 s with a confocal microscope (LSM 710; Carl Zeiss) using a 488-nm laser. At each time point, 10 focal planes were recorded and projections were used for further analysis. Using Fiji (http://fiji.sc) routines for smoothing and thresholding, plaques of desmosomal fluorescence were identified with the “Analyze Particles” tool. To correct for cell movement the centers of identified desmosomal plaques were used for registration with Fiji`s “StackReg” tool. In these corrected datasets, movement of desmosomes was traced and transformed into tracks using the “MTrack2” tool. A total of 2,679 desmosomes were tracked in 5 cells and the mean speed for each desmosome was calculated in WT and KtyII−/− cells.
Dispase assay
The assay was performed as described in Hobbs et al. (2011). In brief, cells were plated in triplicate on 6-well plates. 24 h after plating, confluent cell layers were switched to high calcium for 48 h, cells were washed with PBS and incubated with 2 ml/well of dispase II (Roche), 2.4 U/ml diluted in FAD medium with 2 mM CaCl2, for 30 min at 37°C. Plates were then subjected to orbital rotation (150 rpm) for 5 min at 37°C. Then, detached monolayers were exposed to mechanical stress by 50 inversions in 4 ml PBS in a 15-ml Falcon tube and the fragments were counted. Inhibitors were preincubated for 1 h. Inhibitors were added into dispase-containing medium. Images were acquired using a dissecting microscope (SMZ1500; Nikon) with a digital sight camera (DS-F12; Nikon). Image analysis and processing were performed using NIS-ELEMENTS (Nikon) and Photoshop CS4 (Adobe) software. LUT (lookup table; brightness and gamma) was adjusted using Photoshop.
EGTA and drug treatment
EGTA treatment was performed as described in Kartenbeck et al. (1982). EGTA from a 0.5-M stock solution was added to a final concentration of 2 mM and the resulting pH shift was corrected by addition of a few drops of 0.1 N NaOH. Cells were then incubated at 32°C for different time points (0, 1, 2, 4, 8, and 16 h) before processing for immunofluorescence and protein analysis. Dynasore (Sigma-Aldrich) was added from a 100-mM stock solution to the medium to obtain a final concentration of 50 µM. Cells were incubated for 1 h before further analysis. Gö6976 (Sigma-Aldrich) was diluted in DMSO and added to the medium to obtain a final concentration of 100 nM. Cells were incubated for 1 h. To investigate the degradation of desmoglein the cells were preincubated with 150 µg/ml of cycloheximide (CHX). Desmosome disassembly and degradation was induced by EGTA treatment for 2–8 h in the presence of DMSO alone or 10 µM MG132 (Sigma-Aldrich) as proteasomal inhibitor or 10 µg/ml E64D (Sigma-Aldrich), 10 µg/ml leupeptin (Peptanova), and 10 µg/ml pepstatin A (ELP; Peptanova) as lysosomal inhibitor cocktail.
Western blotting
SDS-PAGE was performed as described in Vijayaraj et al. (2009). In brief, total proteins were extracted in SDS-PAGE sample buffer under repeated heating (95°C) and sonication cycles. Total protein concentration was determined by the Bio-Rad Laboratories protein quantification kit and equal amounts of protein were loaded. Separation of total protein extracts was performed by standard procedures (8–10% SDS-PAGE). The WB was performed as described earlier (Vijayaraj et al., 2009).
Immunoprecipitation
For immunoprecipitation experiments, 2 × 106 cells were cultured in a 10-cm tissue culture dish. 24 h after plating, the medium was switched to high calcium medium for 48 h. The cells were washed twice with PBS and lysed with 500 µl of cold RIPA buffer (10 mM Na2HPO, 150 mM NaCl, 5 mM EDTA, 2 mM EGTA, 1% Triton X-100, 0.25% SDS, 1% sodium deoxycholate, 0.1 mM DTT, pH 7.5, and protease and phosphatase inhibitor cocktails; Thermo Fisher Scientific). Lysates were transferred to 1.5-ml tubes, incubated 30 min on ice, and clarified by centrifugation at 16,000 g for 10 min. Supernatants were incubated with appropriate primary antibodies overnight at 4°C. A 25-µl volume of protein G–Agarose beads (Thermo Fisher Scientific) was added and tubes were rotated at 4°C for 2 h. Beads were washed with 800 µl buffer containing 50 mM Tris-HCl, 150 mM NaCl, 0.1 mM EDTA, and 0.5% Tween 20, pH 7.5; and 800 µl wash buffer containing 100 mM Tris-HCl, 200 mM NaCl, 2 M urea, and 0.5% Tween 20, pH 7.5. Finally, beads were washed with 1% Triton X-100 in PBS and boiled in 30 µl Laemmli sample buffer. Released proteins were separated on 8–10% SDS-PAGE gels.
Cell fractionation
To separate cytosolic from PM proteins cells were washed twice with ice-cold PBS and then collected using a rubber scraper in 500 µl of lysis buffer (20 mM Tris, pH 7.5, 5 mM EDTA, 2 mM EGTA, and 1× protease inhibitor cocktail; Sigma-Aldrich). Lysates were sonicated for 5 s and subjected to centrifugation (1 h, 20,000 g). Supernatant represents the soluble cytosolic fraction. The pellet was solubilized in resuspension buffer (1% Triton X-100, 20 mM Tris, pH 7.5, 5 mM EDTA, 2 mM EGTA, and 1× protease inhibitor cocktail; Sigma-Aldrich), incubated for 1 h on ice, and centrifuged again (1 h, 20,000 g). This supernatant represents the PM protein pool. Laemmli sample buffer was added to all samples before loading onto gels for electrophoresis.
Proximity ligation assay
All reagents used for the PLA analysis were from Olink Bioscience (Uppsala Science Park, Uppsala, Sweden). The PLA assay was performed following the manufacturer’s protocol using anti-K5 (rabbit, diluted 1:800; Magin laboratory) and anti-Rack1 (mouse monoclonal, diluted 1:100; Santa Cruz Biotechnology, Inc.) as primary antibodies.
Transcriptome analysis of KtyII−/− and control skin RNA
This was performed as described in Roth et al. (2012b). The MouseWG-6v2.0 Expression BeadChip kit (Illumina, Inc.) was used to probe triplicate WT and KtyII−/− skin samples from E18.5 embryos. Data analysis was based on the R Statistical language (R Development Core Team 2007, 2.8.0) and Beadstudio 3.1.1.0 software. Data were quantile-normalized. A fold-change/P-value filter was used to select differentially expressed genes; P values smaller than 0.05, expression changes higher than twofold, and a difference between mean intensity signals greater than background were considered statistically significant. The false discovery rate (FDR) of P values was adjusted by the Benjamini–Hochberg method.
KtyII+/− x mKrt8 mice
KtyII+/− mice contain one copy of the type II keratin gene cluster flanked by loxP sites at the 5′- and 3′-ends, respectively, and one copy in which the cluster has been deleted upon cre recombination in ES cells (Vijayaraj et al., 2009). These mice were mated to mouse Krt8 transgenic mice (FVB/N; Nakamichi et al., 2005) and crossed further to receive litter expressing Krt8 but lacking the entire KtyII gene cluster. According to prediction, transgenic mKrt8 should form IFs in all embryonic and simple epithelia with Krt19 encoded by the KtyI locus, rescuing the previously described embryonic phenotype. Surviving litter was genotyped as described previously (Vijayaraj et al., 2009), using the primers (5′-CTGAGCCTTCTGGAGCTATTT-3′, 5′-AAACATGTTGTCCATTTGCT-3′) for detection of mKrt8. Litters developed to term at the expected Mendelian ratio. Skin from E18.5 pups lacking all KtyII+/− genes and expressing mKrt8 in simple epithelia but not in the epidermis was used for the isolation of keratinocytes as described above.
Online supplemental material
Fig. S1 shows characterization of keratinocyte lines. Fig. S2 shows analysis of junction proteins. Fig. S3 shows analysis of keratin desmosome interactions. Table S1 is a list of antibodies.
Acknowledgments
We thank K. Green for DP vectors. We thank S. Eisner for expert electron microscopy. We thank B. Omary for mK8 transgenic mice, and M. Hatzfeld and S. Hauschildt for discussions and critical reading of the manuscript. Microarray analysis was carried out at the Core Unit DNA-Technology, IZKF Leipzig (K. Krohn).
Work in the Magin lab is partially supported by the DFG (SFB832 TP05, MA1316-9/3, MA1316-15, and INST 268/230-1) and the Translational Center for Regenerative Medicine, TRM, Leipzig (no. 0315883); and the Leube/Windoffer labs by the DFG (LE566/18-1, WI1731/6-1, and WI1731/8-1).
References
Author notes
C. Kröger and F. Loschke contributed equally to this paper.