On rigid surfaces, the cytoskeleton of migrating cells is polarized, but tissue matrix is normally soft. We show that nonmuscle MIIB (myosin-IIB) is unpolarized in cells on soft matrix in 2D and also within soft 3D collagen, with rearward polarization of MIIB emerging only as cells migrate from soft to stiff matrix. Durotaxis is the tendency of cells to crawl from soft to stiff matrix, and durotaxis of primary mesenchymal stem cells (MSCs) proved more sensitive to MIIB than to the more abundant and persistently unpolarized nonmuscle MIIA (myosin-IIA). However, MIIA has a key upstream role: in cells on soft matrix, MIIA appeared diffuse and mobile, whereas on stiff matrix, MIIA was strongly assembled in oriented stress fibers that MIIB then polarized. The difference was caused in part by elevated phospho-S1943–MIIA in MSCs on soft matrix, with site-specific mutants revealing the importance of phosphomoderated assembly of MIIA. Polarization is thus shown to be a highly regulated compass for mechanosensitive migration.
Introduction
Cell migration on rigid substrates, such as coverslips, has revealed the potential for polarization of key cytoskeletal components, including myosin-II (Kolega, 2003; Vicente-Manzanares et al., 2008; Barnhart et al., 2011). On soft substrates and in 3D matrix, however, the morphologies of migrating cells and their phosphoprotein profiles appear distinct from those on rigid 2D substrates (Pelham and Wang, 1997; Doyle et al., 2009). Within a soft tissue, such as the developing brain, cytoskeletal polarization also shows no clear relation to the direction of migration, whereas cells cultured on rigid substrates polarize in the direction of migration (Distel et al., 2010). The impact of soft matrix microenvironments on cytoskeletal polarization and migration appears understudied as are the effects of gradients in matrix elasticity. Durotaxis is the tendency of a cell to crawl from soft matrix to stiff matrix in the absence of any gradients in ligand density or chemotactic factors (Lo et al., 2000; Cheung et al., 2009; Isenberg et al., 2009), and durotaxis has been speculated to result in part from an increase in the stability of adhesions to stiff matrix as cells migrate from soft matrix (Lo et al., 2000). However, the molecular mechanisms of durotaxis have remained unexplored.
A cell typically moves forward by detaching its adherent tail with contractile forces exerted by nonmuscle myosin-II on the matrix (Kolega, 2003). Myosin-II forces have also been found critical to sensing matrix elasticity E (Discher et al., 2005), although any specific role for myosin-II in sensing gradients in stiffness remains unclear. Of the A, B, and C isoforms of nonmuscle myosin-II, the A isoform (MIIA) is most abundant in mesenchymal tissues based on mass spectrometry (MS) estimates of tryptic peptide abundance (Ma et al., 2010), and it proves essential to any differentiation of embryos (Conti et al., 2004). Importantly, MIIA also contributes the majority of traction force exerted by mesodermal cells, such as embryo-derived fibroblasts (Cai et al., 2006). Nonmuscle MIIB (myosin-IIB) knockout mice exhibit select, but critical, defects in formation of heart and other tissues, and MIIB knockdown (KD) fibroblasts in culture exhibit extended tails that fragment, leading to a frequent change in direction and faster migration (Lo et al., 2004; Swailes et al., 2006). In cells crawling on rigid coverslips, MIIB is more enriched or polarized toward the cell rear (Saitoh et al., 2001; Sandquist et al., 2006), whereas MIIA appears more uniform. The isoform localization difference is caused, surprisingly, by a more stable cytoskeletal assembly mediated by the coiled-coil tail of MIIB (Vicente-Manzanares et al., 2008). On the other hand, phosphorylation of MIIA’s tail promotes disassembly of this traction-critical isoform, impacting epithelial cell migration on rigid substrates (Dulyaninova et al., 2007). We hypothesized therefore that MIIB could be important to the persistent migration of cells on matrix gradients and that the levels of MIIA phosphorylation could impact both durotaxis and cytoskeletal polarization.
Polarization of myosin-II and perhaps phosphoregulated states of the tails could be keys to understanding how mesenchymal stem cells (MSCs) traffic to sites of scarring and wounding in collagen-rich tissues, such as the heart (Orlic et al., 2001; Quevedo et al., 2009). In such sites, these cells have immunomodulatory functions that limit formation of a collagen-I–rich scar (Salem and Thiemermann, 2010; Shi et al., 2010), which is perhaps why these cells are being widely used in clinical trials today even though we know very little about their motility. MSCs have a fibroblast-like cytoskeleton with MIIA and MIIB that contribute to various cellular processes, including matrix elasticity sensing (Engler et al., 2006; Johnson et al., 2007). Using an atomic force microscope (AFM), we have previously measured the elasticity of an infarct scar in heart to be Escar = 30–70 kPa (Berry et al., 2006), which is stiffer than normal muscle and most other soft tissues (E = ∼0.1–30 kPa; Engler et al., 2006). A scar is far softer than glass or plastic, and its stiffness can be mimicked with collagen-coated gels of polyacrylamide that are in wide use to understand matrix elasticity effects on cells. Such gels are used here in 2D and 3D to understand the roles of MIIB and phospho-MIIA in polarization and durotaxis of MSCs. Various perturbations to the myosins reveal that both durotaxis and polarization are maximal at wild-type (WT) levels of MIIB and phosphodynamic MIIA. Cytoskeletal polarization thus appears to be a highly regulated, mechanosensitive compass in directed migration.
Results
MIIB progressively polarizes as MSCs migrate from soft to stiff matrix
Gradients in matrix elasticity were reproducibly made by pipetting two drops with different concentrations of polyacrylamide solution onto a coverslip 2 mm apart and compressing the drops together with a removable upper coverslip, in a method adapted from Lo et al. (2000). After the polymerization reactions ended, collagen-I was covalently attached and verified by immunofluorescence to be uniform across the entire gel. We then measured the elasticity of the gel surface by AFM using a submicrometer probe tip: one side of the gel is made to be soft with Esoft = 1 kPa, which is typical of soft tissue (Discher et al., 2005). The elasticity gradually increased over ∼2,000 µm to a stiff gel with scarlike elasticity of Escar = 34 kPa (Fig. 1 A, inset). The broad plateaus on either side of the gradient allowed for facile comparisons of the various polarization and motility measurements that follow. The gradient region was large relative to cell sizes and proved similar to stiffness gradients measured near myocardial infarct scars in which MSCs are injected (Berry et al., 2006). Within hours of plating MSCs sparsely on these substrates, the cells attached, spread, and then migrated. Regardless of whether the cells were on the soft matrix, stiff matrix, or gradient matrix, most cells exhibited the classical fan shape of migrating cells with a broad leading edge and a narrow tail. To obtain 3D cultures, we polymerized an overlay of collagen-I with E = ∼1 kPa. Cells in these 3D cultures typically exhibited multiple pseudopods rather than the single broad lamellipodium in 2D cultures. Nonetheless, >90% of MSCs were similarly motile in both 2D and 3D matrix systems.
MSCs migrate from soft matrix toward scarlike, stiff matrix with increasing polarization of MIIB. (A) Human MSCs durotax across a gradient in stiffness on collagen-I–coated polyacrylamide hydrogels, even with a 3D overlay gel of collagen-I. MSCs also migrate upward into the soft overlay but only from the soft gel side of the gradient as indicated by arrows. (inset plot) Elasticity, E (±SD), determined by AFM and the gradient in E (red curve). Error bars show SEMs. (B) MSCs were immunostained for nonmuscle MIIB and with rhodamine-phalloidin for F-actin and imaged by fluorescence microscopy. MIIB localizes to the rear of the cell body only on the stiff matrix in 3D and 2D (2D lacks the collagen-I overlay). On soft matrix, MIIB is diffuse in the main cell body but is still absent from the farthest extensions in lamellipodia. Arrows show migration direction. Bars, 50 µm.
MSCs migrate from soft matrix toward scarlike, stiff matrix with increasing polarization of MIIB. (A) Human MSCs durotax across a gradient in stiffness on collagen-I–coated polyacrylamide hydrogels, even with a 3D overlay gel of collagen-I. MSCs also migrate upward into the soft overlay but only from the soft gel side of the gradient as indicated by arrows. (inset plot) Elasticity, E (±SD), determined by AFM and the gradient in E (red curve). Error bars show SEMs. (B) MSCs were immunostained for nonmuscle MIIB and with rhodamine-phalloidin for F-actin and imaged by fluorescence microscopy. MIIB localizes to the rear of the cell body only on the stiff matrix in 3D and 2D (2D lacks the collagen-I overlay). On soft matrix, MIIB is diffuse in the main cell body but is still absent from the farthest extensions in lamellipodia. Arrows show migration direction. Bars, 50 µm.
On stiff matrix in 3D as well as 2D systems, MIIB was most abundant toward the rear of fixed and immunostained MSCs (Fig. 1 B, right). Cells migrating on rigid glass showed a similar polarization of MIIB (Vicente-Manzanares et al., 2008). MIIB in MSCs was seen within stress fibers and sometimes appeared striated. On soft matrix, however, MIIB within the main cell body was surprisingly uniform, nonstriated, and diffuse (Fig. 1 B, left). In the leading lamella of an MSC and in the most distal trailing tail, MIIB was depleted relative to F-actin on soft as well as stiff matrix. This edge depletion of myosin-II depends on the extensive actin polymerization in the leading lamellipodia, which is independent of myosin-II motor activity (Vicente-Manzanares et al., 2008; Mogilner and Keren, 2009).
Our gradient gels ensured consistent immunostaining of cells across soft and stiff matrices so that cytoskeletal polarization within the main cell body could be reliably quantified (Fig. 2, A and B). The ratio of fluorescent signal for the rear half of the migrating cell relative to the front half was calculated as rear/front. F-actin always appeared uniform throughout the cell (within 10%) with rear/front = 1. In cells that spread for 2 h on the gradient region, vinculin in focal adhesions as well as F-actin showed no asymmetry. MIIA showed a slight, persistent polarization of rear/front ∼1.3 independent of matrix elasticity (Fig. 2 C). MIIB appeared strongly polarized toward the rear of the cell on stiff matrix and on collagen-coated glass as reported with other cells on rigid substrates (Kolega, 2003; Vicente-Manzanares et al., 2008). However, MIIB was consistently unpolarized on soft matrix (Fig. 2 D). Hyperbolic fitting of rear/front as a function of E showed that MIIB polarization varies by fourfold with a transition from soft to stiff matrix at a midpoint stiffness of Em = 6.9 kPa. The projected area of MSCs versus E fits a similar hyperbolic form with similar Em (Rehfeldt et al., 2012), which reflects in some broad manner the interrelated tendency of cells to actively spread and polarize in response to stiffer matrix.
MIIB polarization is suppressed in cells on soft matrix, but both the total density and insoluble fraction of MIIB remain constant with matrix stiffness. (A) F-actin and MIIB (gray) were stained in MSCs on soft or stiff matrix. The presence of the lamellipodium was used to identify the cell front. (bottom left) The front half of the cell and the rear half were marked as shown in the image. Migrating MSCs on soft or stiff matrix show uniform F-actin, but MIIB localizes to the cell rear only on stiff matrix. (B) Line scans of 6–10 µm width from the cell rear to front (of cells in A) were used to analyze the distributions of MIIB and F-actin. Each graph consists of data for one representative cell. Such data taken from three experiments are averaged in C and D. (C) Ratios of intensity from the front half of the cell to the rear half show that F-actin is always uniform. Inset image is representative of MIIA, which is throughout the cell body, although it tends to be depleted from the front lamellipodium. MIIA distribution is independent of matrix elasticity. (D) On soft matrix, MIIB is nearly homogeneous (except for the lamellipodium), whereas on stiff matrix, MIIB is more concentrated in the cell rear per Fig. 1 B. (E) Total MIIB density (black, open circles) is independent of matrix elasticity, and increasing matrix stiffness minimally affects the fraction of insoluble MIIB determined by Triton X-100 extraction (green triangles). Shaded regions indicate range of means. Means ± SEM for n ≥ 20 cells among three experiments per data point. au, arbitrary unit; Coll., collagen. Bars, 50 µm.
MIIB polarization is suppressed in cells on soft matrix, but both the total density and insoluble fraction of MIIB remain constant with matrix stiffness. (A) F-actin and MIIB (gray) were stained in MSCs on soft or stiff matrix. The presence of the lamellipodium was used to identify the cell front. (bottom left) The front half of the cell and the rear half were marked as shown in the image. Migrating MSCs on soft or stiff matrix show uniform F-actin, but MIIB localizes to the cell rear only on stiff matrix. (B) Line scans of 6–10 µm width from the cell rear to front (of cells in A) were used to analyze the distributions of MIIB and F-actin. Each graph consists of data for one representative cell. Such data taken from three experiments are averaged in C and D. (C) Ratios of intensity from the front half of the cell to the rear half show that F-actin is always uniform. Inset image is representative of MIIA, which is throughout the cell body, although it tends to be depleted from the front lamellipodium. MIIA distribution is independent of matrix elasticity. (D) On soft matrix, MIIB is nearly homogeneous (except for the lamellipodium), whereas on stiff matrix, MIIB is more concentrated in the cell rear per Fig. 1 B. (E) Total MIIB density (black, open circles) is independent of matrix elasticity, and increasing matrix stiffness minimally affects the fraction of insoluble MIIB determined by Triton X-100 extraction (green triangles). Shaded regions indicate range of means. Means ± SEM for n ≥ 20 cells among three experiments per data point. au, arbitrary unit; Coll., collagen. Bars, 50 µm.
Because of the stiffness-dependent distribution of MIIB in cells on gels, we examined the MIIB density per cell and found that this showed no significant dependence on matrix stiffness (Fig. 2 E). In addition, because of the distinctly diffuse distribution of MIIB in cells on the soft matrices compared with its stress fiber association in cells on stiff matrices, we then extracted the cells with Triton X-100 to image by immunofluorescence just the insoluble myosin-II in the cytoskeleton. Surprisingly, the amount of MIIB left in these extracted cytoskeletons showed no significant difference between soft and stiff matrix (Fig. 2 E). The lack of polarization of MIIB in cells on soft matrix is thus not a result of differences in either MIIB abundance or integration into the cytoskeleton.
MSCs durotax in 2D and 3D
Time-lapse imaging showed that cells crawled at the same mean speed of ∼0.5 µm/min independent of matrix elasticity (Fig. 3 A). On the soft matrix and also within the 3D overlay, cells visibly deformed the matrix (Fig. S1, A and B), whereas cell-induced deformations of stiff matrix were not easily resolved. Cells migrated frequently from the soft side of the gel into the soft overlay, but cells did not migrate from the stiff matrix into the overlay (as indicated schematically in Fig. 1 A), consistent with 3D durotaxis. Durotaxis on the gel surface was evident in the gradient regions in both 2D and 3D with “flower” plot trajectories showing more endpoints (Fig. 3 A, red dots) to the right (stiff) side of the starting point compared with the left (soft) side (Fig. 3 A). For cells on the gradients in both 2D and 3D, a durotaxis index (defined in the legend of Fig. 3 B) proved statistically different from zero after ∼1–3 h of observation (which is approximately one to three cell lengths), indicative of durotaxis.
MSCs crawl from soft to stiff matrix, with durotaxis appearing more sensitive to MIIB KD than MIIA KD. (A) Trajectories of cells with final positions in red after 12 h of imaging. Speed is determined from the cell’s contour length along the trajectory divided by time. (B) Durotaxis index versus time averaged for all cells. We define a step as the net displacement of the cell every 15 min, and we quantify the bias from the time-dependent number of steps to the right (R, stiff) and to the left (L, soft) as durotaxis index = (R − L)/(R + L). If 100% of steps are toward the stiff side, the durotaxis index = 1, whereas if there is no bias to cell migration, the durotaxis index = 0. A positive durotaxis index (P < 0.01) found only on gradient gels indicates biased migration toward stiff matrix after the indicated time constant. On homogeneous soft and stiff matrices, durotaxis index is not statistically different from zero, indicating a random direction. In all plots, means ± SEM for n ≥ 12 cells per data point among three independent experiments are given. (C) Durotaxis index vanishes for both low and high knockdown (KD) of MIIB (red), but a 50% KD of MIIA (blue) still yields a significantly nonzero durotaxis index, whereas a higher KD causes the durotaxis index to vanish. The error bars in the KD level reflect the variation determined from quantitative immunofluorescence imaging. Means ± SEM for n ≥ 20 cells per data point. For shallow MIIB KD, P < 0.05 compared with WT. For shallow MIIA KD, P < 0.05 when compared with both WT and to durotaxis index = 0. (D) Western blots against MIIB and MIIA for graded KDs of MIIA and MIIB, with actin used as a loading control. Bar graph summarizes three independent experiments in showing that minor KD of MIIB has minimal impact on MIIA. Error bars show SEMs. (E) MSCs with MIIB KD have extended tails and increased migration speed compared with WT. MIIA KD cells crawl more slowly than WT. Cells do not durotax with sufficient KD of either isoform, as indicated by the flower plots for cells on the gradient. The cell front and rear were determined from migration paths in time-lapse imaging. The arrow points to the extended tail. au, arbitrary unit; ctl, control. Bars, 50 µm.
MSCs crawl from soft to stiff matrix, with durotaxis appearing more sensitive to MIIB KD than MIIA KD. (A) Trajectories of cells with final positions in red after 12 h of imaging. Speed is determined from the cell’s contour length along the trajectory divided by time. (B) Durotaxis index versus time averaged for all cells. We define a step as the net displacement of the cell every 15 min, and we quantify the bias from the time-dependent number of steps to the right (R, stiff) and to the left (L, soft) as durotaxis index = (R − L)/(R + L). If 100% of steps are toward the stiff side, the durotaxis index = 1, whereas if there is no bias to cell migration, the durotaxis index = 0. A positive durotaxis index (P < 0.01) found only on gradient gels indicates biased migration toward stiff matrix after the indicated time constant. On homogeneous soft and stiff matrices, durotaxis index is not statistically different from zero, indicating a random direction. In all plots, means ± SEM for n ≥ 12 cells per data point among three independent experiments are given. (C) Durotaxis index vanishes for both low and high knockdown (KD) of MIIB (red), but a 50% KD of MIIA (blue) still yields a significantly nonzero durotaxis index, whereas a higher KD causes the durotaxis index to vanish. The error bars in the KD level reflect the variation determined from quantitative immunofluorescence imaging. Means ± SEM for n ≥ 20 cells per data point. For shallow MIIB KD, P < 0.05 compared with WT. For shallow MIIA KD, P < 0.05 when compared with both WT and to durotaxis index = 0. (D) Western blots against MIIB and MIIA for graded KDs of MIIA and MIIB, with actin used as a loading control. Bar graph summarizes three independent experiments in showing that minor KD of MIIB has minimal impact on MIIA. Error bars show SEMs. (E) MSCs with MIIB KD have extended tails and increased migration speed compared with WT. MIIA KD cells crawl more slowly than WT. Cells do not durotax with sufficient KD of either isoform, as indicated by the flower plots for cells on the gradient. The cell front and rear were determined from migration paths in time-lapse imaging. The arrow points to the extended tail. au, arbitrary unit; ctl, control. Bars, 50 µm.
In contrast to cells on the gradient region, cells on the soft or stiff matrix regions showed no significant durotaxis index. However, cells on the soft matrix did not appear to migrate as far as on stiff matrix, even though cell speeds were similar (Fig. 3 A). A quantitative analysis of the trajectories (Fig. S1 C) indeed established that MSCs on both regions started with directed migration that decayed to random migration on soft matrix (time constant of 2.3 h) more rapidly than on stiff matrix (6.3 h). On the gradient, the decay was not monotonic and revealed a statistical tendency for many cells to be more persistent and directed (up the gradient) rather than random in their migration. The less persistent migration on soft matrix versus stiff matrix was consistent with other recent observations (Fischer et al., 2009) and provides a simple explanation for directed migration toward stiff matrix. Moreover, because MIIB knockout fibroblasts crawl with less directional persistence on rigid glass coverslips (Lo et al., 2004), we hypothesized that MIIB would be critical to migration from soft to stiff matrix.
Durotaxis is more sensitive to the minor isoform, MIIB, than to abundant MIIA
Graded KD of both MIIB and MIIA was then used to increasingly perturb physiological levels. Quantitative immunofluorescence of single cells was used to determine the percentage of KD to account for cell-to-cell variations (e.g., Fig. S2 A) as represented by x axis error bars in Fig. 3 C’s measurements of the durotaxis index for randomly chosen cells. KD of MIIB by 28% was sufficient to decrease the durotaxis index to zero, eliminating MSC durotaxis (Fig. 3 C). In comparison, a 50% KD of MIIA led to only a 50% reduction in the durotaxis index. A deeper KD of MIIA by 75% was required to significantly block durotaxis.
Levels of MIIA and MIIB were further assessed by Western immunoblotting (Fig. 3 D), quantitative immunofluorescence (Fig. S2, A and E; Fig. 2 E; and Fig. 3 C), and quantitative liquid chromatography (LC)–coupled tandem MS (LC-MS/MS; Fig. S3). Immunoblotting verified the mean KD in the cell populations (Fig. 3 D). We used LC-MS/MS to additionally quantify KDs of MIIA and MIIB together with potential changes in other proteins (Shin et al., 2011) as validated by immunoblotting for seven proteins (Fig. 3 D and Fig. S3 E). The most important case is the minimal KD of MIIB that blocks durotaxis, for which nearly all cytoskeletal proteins changed minimally (Fig. 3 D, plot; and Fig. S3 A). The decrease in MIIB is therefore the likely explanation for the durotaxis defect. On the other hand, deep KD of MIIA led to significant changes in several other proteins detected by MS, including an increase in MIIB (by 33%) as seen also in immunoblots (Fig. 3 D). The effects of ectopic overexpression of myosin-II on cytoskeletal polarization and durotaxis are described in a later section of the Results.
The ratio of MIIB to MIIA isoforms was also determined from LC-MS/MS using ion currents for tryptic peptides common to both isoforms and also unique to each isoform. MIIB proves to be the minor isoform at only 3–9% of total myosin-II (Fig. S3 B). The elimination of biased migration by a minimal and specific KD of the minor isoform, MIIB, shows that durotaxis is far more sensitive to MIIB than to MIIA.
We hypothesized that blebbistatin inhibition of myosin-II ATPase/motor activity could inhibit durotaxis much faster than could be studied by KD. Blebbistatin was pulsed into established cultures on gradients during time-lapse imaging, and although cells continued to migrate everywhere, the durotaxis index decreased to zero with time constant τ < 15 min (Fig. S4 A, i and ii). Because MIIB polarization typically accompanies durotaxis in the aforementioned experiments, we examined MIIB polarization after this brief treatment with blebbistatin and found that the polarization of MIIB decreased only slightly within the first hour of drug treatment (Fig. S4 A, i, inset bar graph) but is then lost several hours later in blebbistatin. It is not surprising that blebbistatin did not immediately disrupt MIIB polarization because cellular localization of myosin-II isoforms is caused primarily by the coiled-coil tails rather than the heads with the ATPase/motor activity (Vicente-Manzanares et al., 2008). Importantly, the blebbistatin results here indicate that MIIB polarization is not sufficient for durotaxis and can decouple from durotaxis, whereas myosin-II motor activity is necessary for durotaxis.
The treatment with blebbistatin initially slowed down MSCs on the soft matrix and slightly sped up MSCs on the stiff matrix (Fig. S4 A, iii), but by 6 h after drug exposure, cell shapes and crawling speeds became almost independent of matrix elasticity (Fig. S4 B). Sustained blebbistatin treatment and KD of the MII isoforms generally altered the shapes of MSCs, but the most striking morphological change followed MIIB KD, which led to highly extended tails (Fig. 3 E), with migration direction verified by time-lapse imaging. The observations are similar to those for myoblasts on rigid coverslips after MIIB KD (Swailes et al., 2006) and are consistent with a role of MIIB in tail cohesion and/or retraction during in vitro migration on coverslips (Kolega, 2003). Surprisingly, on soft matrix, the trailing tails of MIIB KD cells were significantly longer compared with either control cells or KD cells on stiff matrix (Fig. S2 B). MIIB in untreated MSCs therefore has an important function in cohesion and/or deadhesion of the cell tail on soft matrix despite its diffuse distribution (Fig. 1 B).
In addition to knocking down the two myosin-II isoforms, we overexpressed both as GFP fusion constructs. Overexpression of GFP-MIIB by 3–20-fold (mean of eightfold) showed integration into the cytoskeleton but suppressed polarization of MIIB to the cell rear on stiff substrates (Fig. S2 D). These cells also did not migrate significantly and could not durotax. In comparison, overexpression of WT GFP-MIIA by ≤1.4-fold (i.e., 40% overexpression) had no significant impact on migration, durotaxis, or polarization of MIIB (as elaborated in a later section of the Results). It is also worth noting that the mean level of MIIB overexpression, combined with the low endogenous levels, gives a total level of myosin-II (MIIA + MIIB) that is about the same as with the MIIA overexpression, and yet the former MIIB-overexpressing cells do not crawl, polarize, or durotax, whereas the latter MIIA-overexpressing cells do crawl, polarize, and durotax. This hints at the functionally distinct roles of MIIA and MIIB isoforms and suggests that MIIB must be expressed at relatively low levels in WT MSCs in order for them to functionally durotax.
Molecular mobility is highest for MIIA and for soft matrix, with phosphodynamics of MIIA contributing to mechanism
For insight into physical differences in molecular regulation, FRAP was used to study molecular mobility of GFP fusions. GFP-MIIB had been shown to be less mobile than GFP-MIIA in cells on rigid coverslips (Vicente-Manzanares et al., 2007), and similar results were seen here with cells on the stiff but compliant 34-kPa matrix (Fig. 4 A, i). The result is consistent with a general similarity of cell responses to stiff gels and rigid glass (Fig. 2 D). Although the results also indicate more stable integration of MIIB, additional FRAP experiments showed that MIIA is even more mobile on soft matrix (Fig. 4 A, ii), which is consistent with a past study showing that MIIB is more mobile in cells on soft matrix versus stiff matrix (Fischer et al., 2009).
MIIA is more assembled on stiff matrix than soft, and the phosphomimetic S1943D is more soluble and mobile than WT or S1943A. (A, i) MSCs transfected with GFP-MIIA or GFP-MIIB on stiff matrix were photobleached along actin bundles and allowed to recover. MIIA recovered faster and thus was more mobile than MIIB. (ii) MSCs expressing GFP-MIIA on either soft 1-kPa matrix or stiff 34-kPa matrix were similarly analyzed, and MIIA in cells on soft matrix recovered more quickly. (iii) Phosphomimetic GFP-MIIA S1943D mutant recovered faster than GFP-MIIA S1943A, which was less than or equal to the behavior of GFP-MIIA on stiff matrix. Data are means ± SEM for at least five cells. (B) Fraction of insoluble MIIA is quantified by immunofluorescence of cell ghosts that were derived from Triton X-100 extraction of cells on matrices for 4 min (±SEM; n = 3). Western blotting of the insoluble fraction of the cells on matrices was also performed. Representative blot for MIIA is shown, normalized to total protein. (C) MSCs transfected with either GFP only, GFP-MIIA (WT), S1943D, or S1943A mutants were Triton X-100 extracted, and the extracted amount of MIIA (soluble) was compared with the insoluble fraction. Immunoblots show anti-MIIA blotting with β-actin as a loading control. When the amount of MIIA is normalized to β-actin, the only significant difference in the insoluble/soluble MIIA ratio is between S1943D and S1943A (P ≤ 0.05; ±SEM; n = 3). Endog., endogenous.
MIIA is more assembled on stiff matrix than soft, and the phosphomimetic S1943D is more soluble and mobile than WT or S1943A. (A, i) MSCs transfected with GFP-MIIA or GFP-MIIB on stiff matrix were photobleached along actin bundles and allowed to recover. MIIA recovered faster and thus was more mobile than MIIB. (ii) MSCs expressing GFP-MIIA on either soft 1-kPa matrix or stiff 34-kPa matrix were similarly analyzed, and MIIA in cells on soft matrix recovered more quickly. (iii) Phosphomimetic GFP-MIIA S1943D mutant recovered faster than GFP-MIIA S1943A, which was less than or equal to the behavior of GFP-MIIA on stiff matrix. Data are means ± SEM for at least five cells. (B) Fraction of insoluble MIIA is quantified by immunofluorescence of cell ghosts that were derived from Triton X-100 extraction of cells on matrices for 4 min (±SEM; n = 3). Western blotting of the insoluble fraction of the cells on matrices was also performed. Representative blot for MIIA is shown, normalized to total protein. (C) MSCs transfected with either GFP only, GFP-MIIA (WT), S1943D, or S1943A mutants were Triton X-100 extracted, and the extracted amount of MIIA (soluble) was compared with the insoluble fraction. Immunoblots show anti-MIIA blotting with β-actin as a loading control. When the amount of MIIA is normalized to β-actin, the only significant difference in the insoluble/soluble MIIA ratio is between S1943D and S1943A (P ≤ 0.05; ±SEM; n = 3). Endog., endogenous.
When we had imaged MIIA to quantify what turned out to be a minimal polarization of MIIA in cells on both soft and stiff matrices (Fig. 2 C), we noticed that MIIA appeared much more diffuse and less integrated in filaments in cells on soft matrix, consistent with its increased mobility in FRAP (Fig. 4 A, ii). Given that MIIA is the major isoform (Fig. S3 B) that contributes the majority of traction forces used to sense soft versus stiff matrix (Lo et al., 2004), we thought that the dependence of MIIA assembly and filamentation on matrix rigidity would ultimately prove critical to MIIB polarization as well as durotaxis. Our MS analyses had detected phospho-Serine1943 (pS1943) in MIIA (unpublished data), and pS1943 is known to favor disassembly of MIIA from stress fibers (Dulyaninova et al., 2007). Our working hypothesis therefore became that pS1943 would be high on soft matrix and low on stiff matrix with a significant change in phosphorylation at this site being a key part of the molecular mechanism for how matrix stiffness promotes MIIA assembly. Note that this is a hypothesis on dynamics because our myosin-II KDs had already indicated that overall levels of MIIA do not greatly impact durotaxis. We sought to test the phosphodynamics hypothesis with FRAP and then extractability methods applied to suitable mutants. A phosphomimetic Asp1943 (S1943D) mutant and a nonphosphorylatable Ala1943 (S1943A) mutant were first studied as GFP fusions by FRAP of MSCs on stiff matrix. Phosphomimetic S1943D-MIIA proved more mobile than WT GFP-MIIA (Fig. 4 A, iii), consistent with the cited disassembly from stress fibers. In contrast, S1943A-MIIA had a mobility less than or equal to that of WT GFP-MIIA. Because S1943A favors assembly into stress fibers, the results indicate stable integration of WT GFP-MIIA into stress fibers on stiff matrix, consistent with the working hypothesis.
Extraction methods were used to expand on the FRAP findings. Cells on soft and stiff matrices were prepermeabilized with Triton X-100 to extract soluble protein from the cells, and then, the cytoskeletons were fixed, immunostained for insoluble MIIA, and imaged in quantitative fluorescence microscopy. Stiff matrix showed 50% insoluble MIIA, and this decreased significantly to 32% on soft matrix (Fig. 4 B) with confirmation of the trend by Western blotting. This is consistent with the FRAP results in indicating that stiff matrix promoted MIIA assembly into stress fibers. The results indicate that the insoluble/soluble ratio of MIIA is only ∼1:2 on soft matrix, and this doubles to ∼1:1 on stiff matrix. This began to indicate elevated pS1943 in cells on soft matrix as analyzed directly in the next section.
Validation of FRAP findings with the different myosin GFP mutants (and a monomeric GFP control) was performed by once again treating with Triton X-100 and then analyzing the soluble fraction and insoluble pellet by Western blotting with anti-MIIA. GFP-myosins were seen at a distinctly higher molecular weight than endogenous MIIA. The quantity of myosin-II in the insoluble pellet relative to the soluble extract is shown in Fig. 4 C as the insoluble/soluble ratio for each transfection. For endogenous MIIA, this ratio was similar for all transfections. Ratios of insoluble/soluble for both endogenous myosin and GFP-MIIA WT fit within the broad error bars of Fig. 4 B on the stiffest substrate, but direct comparisons of endogenous to GFP-myosins are tenuous because of the proximity of the two bands. Whereas the S1943A construct did not differ significantly from WT, the S1943D construct appeared the most soluble, consistent with the high mobile fraction in FRAP studies of single cells (Fig. 4 A, iii). In a later section of the Results, the results prompted detailed analyses of stress fibers, MIIB polarization, and durotaxis after transfections.
MIIA phosphorylation at S1943 decreases as matrix stiffness increases
Differences in FRAP mobility and solubility of WT MIIA in cells on 1-kPa matrix versus 34-kPa matrix and the difference between S1943A-MIIA and S1943D-MIIA led us to hypothesize that S1943 would be more phosphorylated on soft matrix than on stiff matrix. Quantitative immunofluorescence of endogenous pSer1943-MIIA indeed shows significantly higher intensity in cells on soft matrix compared with stiff matrix after normalization for total MIIA (Fig. 5 A, bar graph). Western blotting of lysates derived from MSCs cultured on soft or stiff matrix confirmed a greater amount of pS1943 in cells on soft matrix (Fig. 5 B). Moreover, pS1943 staining appeared more diffuse and less prominent in stress fibers than total MIIA (Fig. 5 A, images). This was quantified with an algorithm that first calculates the orientation of each stress fiber structure referenced to a mean orientation angle (Zemel et al., 2010). The sample images showed myosin mostly oriented above a low background, whereas pS1943 exhibited a high background (Fig. 5 C). The ratio oriented/isotropic subtracts the isotropic background from the oriented signal. Thus, on stiff matrix, MIIA was more apparent in ordered filaments than is the case for pS1943 (P < 0.05). Soft matrix strongly suppressed orientational order of MIIA (Fig. 5 C, inset), which is consistent with MIIA being more phosphorylated and more mobile in cells on soft matrix. Triton X-100 extractions examined in a later section of the Results are consistent with these conclusions.
S1943 of MIIA is matrix elasticity E regulated and impacts structural integration into filaments. (A) MSC on stiff matrix stained with anti-MIIA and with anti-pS1943. pS1943 is more diffuse compared with MIIA and less integrated into stress fibers. The total intensity of pS1943 normalized to MIIA signal always appeared higher for MSCs on soft than stiff matrix (n ≥ 20 cells). (B) Representative Western blot for pS1943 from MSCs cultured on soft or stiff matrix with MIIA as a loading control shows relatively more pS1943 per MIIA (representative of Western blots from n = 3 independent experiments). (C) The orientation of the fluorescence signal of MIIA was compared with that of pS1943 on both soft and stiff matrix conditions and plotted as a normalized abundance as a function of the angle. The peak of the fitted curve marks the oriented signal, whereas the baseline represents the isotropic structure. (inset) MIIA was less structured on soft matrix compared with signal on stiff matrix. (D) Orientation of myosin structure with GFP-MIIA and MIIA mutants. GFP-MIIA S1943D was less structured than MIIA WT or S1943A. Endogenous pS1943 was immunostained and again showed less structure on soft matrix in all transfects but showed more fiber integration with S1943A. We numbered all conditions in a table and then calculated the p-value in comparing all possible datasets. Squares filled in with yellow indicate a P < 0.05. Data are means ± SEM with ≥35 cells. endog., endogenous. Bar, 50 µm.
S1943 of MIIA is matrix elasticity E regulated and impacts structural integration into filaments. (A) MSC on stiff matrix stained with anti-MIIA and with anti-pS1943. pS1943 is more diffuse compared with MIIA and less integrated into stress fibers. The total intensity of pS1943 normalized to MIIA signal always appeared higher for MSCs on soft than stiff matrix (n ≥ 20 cells). (B) Representative Western blot for pS1943 from MSCs cultured on soft or stiff matrix with MIIA as a loading control shows relatively more pS1943 per MIIA (representative of Western blots from n = 3 independent experiments). (C) The orientation of the fluorescence signal of MIIA was compared with that of pS1943 on both soft and stiff matrix conditions and plotted as a normalized abundance as a function of the angle. The peak of the fitted curve marks the oriented signal, whereas the baseline represents the isotropic structure. (inset) MIIA was less structured on soft matrix compared with signal on stiff matrix. (D) Orientation of myosin structure with GFP-MIIA and MIIA mutants. GFP-MIIA S1943D was less structured than MIIA WT or S1943A. Endogenous pS1943 was immunostained and again showed less structure on soft matrix in all transfects but showed more fiber integration with S1943A. We numbered all conditions in a table and then calculated the p-value in comparing all possible datasets. Squares filled in with yellow indicate a P < 0.05. Data are means ± SEM with ≥35 cells. endog., endogenous. Bar, 50 µm.
GFP-MIIA in transfected cells (Fig. 5 D) showed the same trends in orientational order as endogenous MIIA and pS1943 on soft and stiff gels (Fig. 5 C, inset), with statistical tests establishing the expected similarities and differences in showing that overexpression of MIIA (by ∼30%) is not introducing structural artifacts in actomyosin filaments. However, the S1943D mutant of GFP-MIIA always produced a more diffuse, isotropic signal than WT, which is consistent with the idea that a negatively charged 1,943 site promotes filament destabilization and disassembly (Dulyaninova et al., 2007). Conversely, the orientation of GFP-S1943A filaments in cells on soft matrix appeared statistically the same as GFP-WT and GFP-1943A in cells on stiff matrix, which indicates that S1943A promotes oriented filaments on soft matrix. Staining of the S1943A mutant MIIA cells with anti-pS1943 (which binds only to endogenous MIIA per Western blots) also showed for stiff matrix an increase in filament-oriented pS1943 compared with S1943D. The various effects of this tail phosphosite on assembly into stress fibers thus suggest a role in the matrix mechanosensitivity of MSCs.
On stiff matrix, cells assembled more stress fibers and achieved a greater spread area than cells on soft matrix (Lo et al., 2000; Discher et al., 2005; Rehfeldt et al., 2012). MSCs expressing the stress fiber–stable S1943A-MIIA mutant showed more oriented stress fiber signal compared with WT MIIA (Fig. 5 D) and also spread more on soft matrix compared with MSCs transfected with WT MIIA or S1943D (Fig. S5 A). These phenotypic results are consistent with endogenous MIIA being less phosphorylated and more assembled into stress fibers on stiff matrix (Fig. 5). MSCs transfected with MIIA WT cells spread more on stiff matrix than on soft matrix, as is typical of a mechanosensitive response, and which the aforementioned data indicate should lead to a decrease in pS1943. The S1943D mutant mimics unalterable phosphorylation, and this mutant seemed to spread more at 24 h than WT MIIA or S1943A, which is consistent with previous measurements of MDA-MB-231 cells that also spread 20% more with S1943D on rigid surfaces (Dulyaninova et al., 2007). The difference in spreading appeared time dependent in our observations and has a likely basis in retrograde flow per the Discussion section.
Dephosphorylation of MIIA pS1943 is essential for durotaxis and for MIIB polarization but not for migration
Next, we studied the impact of the MIIA phosphosite mutants on durotaxis and MIIB polarization. MSCs transfected with GFP-MIIA WT on gels with gradients of stiffness exhibited a durotaxis index of ∼0.22 (Fig. 6 A), which is statistically the same as nontransfected MSCs (Fig. 3, B and C). However, MSCs transfected with the S1943A mutant showed a 50% reduction in the durotaxis index, and the S1943D mutant showed no significant durotaxis. Crawling speeds were similar (differences within <20%), and so, transfections did not cause overall motility defects.
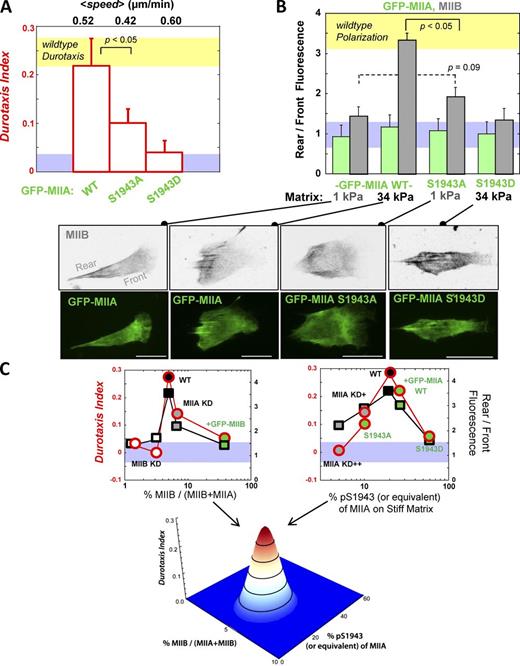
Durotaxis and MIIB polarization are disrupted by MIIA S1943D overexpression and are maximal for WT levels of MIIB expression and pS1943. (A) MSCs on gradient gels transfected with WT GFP-MIIA show a durotaxis index similar to nontransfected, WT cells (yellow band). The S1943A mutant shows a reduced durotaxis index, and the S1943D mutant shows no significant durotaxis above background. Data are means ± SEM for ≥12 cells. (B) The same transfected MSCs were immunostained for MIIB, which polarizes in GFP-MIIA cells on stiff matrix (34 kPa) but does not on soft matrix (1 kPa) similar to nontransfected, WT cells (Fig. 1). The S1943A mutant shows a modest increase in MIIB polarization even on soft matrix, where these cells tend to spread more than any other cell (Fig. S5), as is found on stiff matrix. The S1943D mutant shows no significant MIIB polarization on soft or stiff matrix, even though these cells tend to spread on stiff matrix more so than any other cell (Fig. S5). The blue band is the range of durotaxis index when there is no durotaxis per Fig. 3 C. (C) Summary of all durotaxis index and polarization data from WT, KD, and overexpression experiments shows that WT cells are optimized for durotaxis and polarization. For only the data point involving transfection with S1943D, the S1943D is considered equivalent to pS1943 and considered part of the percentage of pS1943 in the graph. The surface plot illustrates the sensitivity of durotaxis and MIIB polarization to both MIIB expression level and the percentage of MIIA that is phosphorylated at S1943 within the cell. Circles are red for the durotaxis index, whereas squares are black for rear/front fluorescence. Data are means ± SEM for ≥20 cells among three independent experiments. Bars, 50 µm.
Durotaxis and MIIB polarization are disrupted by MIIA S1943D overexpression and are maximal for WT levels of MIIB expression and pS1943. (A) MSCs on gradient gels transfected with WT GFP-MIIA show a durotaxis index similar to nontransfected, WT cells (yellow band). The S1943A mutant shows a reduced durotaxis index, and the S1943D mutant shows no significant durotaxis above background. Data are means ± SEM for ≥12 cells. (B) The same transfected MSCs were immunostained for MIIB, which polarizes in GFP-MIIA cells on stiff matrix (34 kPa) but does not on soft matrix (1 kPa) similar to nontransfected, WT cells (Fig. 1). The S1943A mutant shows a modest increase in MIIB polarization even on soft matrix, where these cells tend to spread more than any other cell (Fig. S5), as is found on stiff matrix. The S1943D mutant shows no significant MIIB polarization on soft or stiff matrix, even though these cells tend to spread on stiff matrix more so than any other cell (Fig. S5). The blue band is the range of durotaxis index when there is no durotaxis per Fig. 3 C. (C) Summary of all durotaxis index and polarization data from WT, KD, and overexpression experiments shows that WT cells are optimized for durotaxis and polarization. For only the data point involving transfection with S1943D, the S1943D is considered equivalent to pS1943 and considered part of the percentage of pS1943 in the graph. The surface plot illustrates the sensitivity of durotaxis and MIIB polarization to both MIIB expression level and the percentage of MIIA that is phosphorylated at S1943 within the cell. Circles are red for the durotaxis index, whereas squares are black for rear/front fluorescence. Data are means ± SEM for ≥20 cells among three independent experiments. Bars, 50 µm.
MSCs transfected with GFP-MIIA exhibited polarization of the MIIB isoform to the cell rear on stiff matrix, but rear/front polarization was weak and insignificant on soft matrix (Fig. 6 B), just as observed with nontransfected cells (Fig. 1 B). MIIA polarization was consistently unaffected for all constructs. The S1943D mutant suppressed polarization of MIIB on stiff matrix and also showed no polarization on soft matrix. The S1943A mutant showed a slight but not statistically significant (P = 0.09) increase in MIIB polarization on soft matrix relative to WT, but S1943A did not differ from WT for stiff matrix (unpublished data); the trends are consistent with the orientational order of GFP-S1943A being sufficiently enhanced in cells on soft matrix to show no statistical difference when compared with either GFP-WT or GFP-S1943A in cells on stiff matrix (Fig. 5 D). MIIA’s pS1943 can thus affect MIIB polarization either through direct or indirect interactions.
Discussion
Many observations here are summarized in the plots of Fig. 6 C, which show that durotaxis and rigidity-induced polarization operate only within a narrow range of two independent variables: the percentage of MIIB of total cellular myosin-II and the percentage of MIIA that is phosphorylated at S1943. For polarization and durotaxis of MSCs, only 6–12% of total myosin-II can be the MIIB isoform (Fig. 6 C, top left plot), whereas MIIA can be knocked down and overexpressed to change total myosin-II amounts by about ±50% without affecting durotaxis (Fig. 3 C and Fig. 6 A). Although this corresponds to a 5–10-fold greater tolerance for changes in MIIA levels compared with changes in MIIB levels, matrix-driven changes in phosphorylation of S1943-MIIA also prove critical.
To provide an absolute scale for this MIIA phosphoregulation rather than just a relative scale, we estimated the stoichiometry of pS1943-MIIA in WT MSCs on rigid substrates (as a reference). Immunodepletion of pS1943-MIIA was followed by immunoblotting for the remaining MIIA that is not phosphorylated at S1943 while controlling for nonspecific antibody binding (unpublished data), with the result being that ∼20% of MIIA has pS1943 in WT MSCs. Because overexpression of S1943D-MIIA effectively enhanced endogenous pS1943 as a more mobile, isotropic, and soluble pool of MIIA (Fig. 4 and Fig. 5), S1943D is treated in Fig. 6 C as an “equivalent” in adding to pS1943 when plotting the suppression of durotaxis and polarization by this mutant. Conversely, overexpression of the S1943A mutant decreased the net fraction of pS1943-MIIA (because endogenous pS1943 levels did not appear statistically different after transfection; Fig. S5 B), and this mutant also recruited more pS1943 into filaments (Fig. 5, C and D). This mutant limited the usual phosphorylation changes from soft to stiff matrix (Fig. 5, A and B) and reduced the usual differences in cytoskeletal assembly between cells on soft and stiff matrix, with the ultimate result being that S1943A-MIIA partially suppressed durotaxis and polarization. Given the results for all of these perturbations, durotaxis and polarization prove maximal at WT levels of pS1943 (Fig. 6 C, top right plot).
Soft matrix suppresses MIIA actomyosin assembly and orientation, impacting MIIB
The reasons why organization and polarization of MIIB is lacking in WT MSCs on soft matrix are in need of explanation. In cells on rigid substrates where MIIB is rearward polarized, Vicente-Manzanares et al. (2007) concluded that MIIA initiates formation of the actomyosin filaments, and MIIB subsequently binds and stabilizes these filaments. In MSCs on soft matrix, the major isoform MIIA is less assembled and more disorganized compared with cells on stiff matrix (Fig. 4 and Fig. 5), and the minor isoform MIIB in cells on soft matrix is likewise more disorganized and homogeneous compared with cells on stiff matrix (Fig. 1 B and Fig. 2 D).
Why MIIB polarization is maximal in WT MSCs on stiff matrix is another key question. In the case of MIIB KD, we could see a decreased assembly of MIIB into filamentous structures, thus producing more isotropic than oriented MIIB. This could result from MIIB being closer to or below its critical concentration needed for assembly (Pollard, 1982). In the case of MIIB overexpression, the higher concentration of MIIB tends to come off the stress fibers and into the cytosol, and once again produces more isotropic than oriented MIIB (Fig. S2 D, inset). MIIA did not polarize on stiff matrix for similar reasons of high mobility (Fig. 4 A). KD of MIIA considerably reduces actomyosin mass, and overexpression of S1943D-MIIA suppresses MIIA organization (Fig. 5 D), and both treatments therefore suppress MIIB polarization. The S1943A-MIIA mutant slightly enhances both oriented assembly and MIIB polarization on soft matrix but not stiff matrix (Fig. 5 D and Fig. 6 B); this suppresses the soft–stiff contrast needed for durotaxis as does the inability to phosphorylate–dephosphorylate S1943A, and the results thus confirm that MIIB polarization does not exceed that seen in WT MSCs on stiff matrix. Overall, the various perturbations explained here suggest that the peak in polarization for WT states of myosin-II is reasonable.
Durotaxis, myosin-II polarization, and implications for nonmesenchymal cell types
The surface plot of Fig. 6 C and the schematic of Fig. 7 further summarize the findings here that WT MSCs possess an optimally polarizable, phosphodynamic actomyosin cytoskeleton for biased migration from soft to stiff matrix. Durotaxis has been reported previously for the NIH 3T3 line of mouse embryonic fibroblasts (Lo et al., 2000) and primary bovine aortic vascular smooth muscle cells (Isenberg et al., 2009). Such cells of diverse mesenchymal origin from different species are expected to express predominantly MIIA and similarly low fractions of MIIB (Ma et al., 2010) when compared with the human MSCs derived from bone marrow as studied here. The surface plot here might therefore generalize to other cells of mesenchymal origin. Interestingly, durotaxis of other cell types has not yet been reported to our knowledge and could be limited by stringent requirements revealed here for myosin-II isoforms and phosphorylation. Human neutrophils, for example, express only MIIA (Maupin et al., 1994) and migrate into many tissues. On the other hand, MIIA in the neutrophil does polarize to the cell rear in the uropod on rigid substrates (Xu et al., 2003), and neutrophils on soft matrices are less persistent in migration and less contractile (Oakes et al., 2009). Neutrophils and other cell types might therefore also polarize their cytoskeletons as they durotax by similar mechanisms from soft to stiff matrix.
Schematic summary of matrix elasticity–dependent assembly of MIIA, polarization of MIIB, and persistence in migration of single cells.
Schematic summary of matrix elasticity–dependent assembly of MIIA, polarization of MIIB, and persistence in migration of single cells.
Durotaxis has been described as resulting in part from an increase in the stability of adhesions to stiff matrix as cells migrate from soft matrix (Lo et al., 2000), which would make durotaxis similar to haptotaxis. In the present observations, however, 30% KD of the minor isoform MIIB not only eliminated durotaxis but also led to highly extended, adherent tails that demonstrate that tail adhesion is relatively unperturbed by MIIB KD (Fig. S2 B). The majority of traction forces in similar mesenchymal cell types derives from MIIA (Cai et al., 2006), but durotaxis still occurs here after MIIA is knocked down 50% (Fig. 3 C). The remaining MIIA is 10-fold above the molar amount of MIIB. The effect of MIIB KD is therefore not a simple matter of reduced force generation. Proteomics analyses (Fig. S3 A) further suggested only one protein, calpain-2, was consistently up-regulated upon KD or inhibition of myosin-IIs, but such proteases undermine adhesion, which is not consistent with the adherent tails of MIIB KD cells. Durotaxis is thus not primarily a result of differences in cell adhesion, which distinguishes durotaxis from haptotaxis.
Trafficking of MSCs to wounds and scars might involve durotaxis
MSCs are known to mobilize from soft bone marrow (Orlic et al., 2001) and are also routinely injected into tissue (Burt et al., 2008), and in either case, they are somehow home to wounds and scars as they traverse tissues of locally varied rigidity. Such complex migratory processes seem to be a highly evolved function in higher species, and durotaxis of MSCs certainly seems specialized given the sensitivity to the myosin-II isoform ratio and phosphoregulation (Fig. 6 C). The present findings with multiple batches of low passage MSCs from multiple donors nonetheless suggest the importance of rapidly reversible phosphopathways to durotaxis, polarization, and matrix elasticity sensing. A seminal study of substrate flexibility effects on cells have described overall increases in pTyr levels in cells on stiff matrices (Pelham and Wang, 1997), but the opposite trend is found here with this particular pS1943 site in MIIA. A deep understanding of regulatory kinases and phosphatases might one day be exploited to better mobilize MSCs or perhaps optimize the trafficking and retention of these cells at sites of injury or repair.
Lastly, cells in wounds and fibrotic scars engage matrix all around the cell in 3D, but durotaxis has previously been studied only on 2D substrates (Lo et al., 2000; Gray et al., 2003; Cheung et al., 2009; Hadjipanayi et al., 2009; Isenberg et al., 2009; Tse and Engler, 2011). Second messenger signaling in cell migration within 3D model matrices has been reported very recently to exhibit matrix elasticity–dependent polarization (Petrie et al., 2012), but gradients and transitions in the myosin-II motors that generate critical forces have been unexplored. 3D gels with gradients in collagen density have been examined (Hadjipanayi et al., 2009) but involve changes in ligand density, and so the 3D results here with controlled gels (Fig. 1) eliminate haptotaxis effects. More importantly, the results provide a visible molecular signature of likely relevance to any stiffness sensing and durotaxis in various systems, including tissue.
Materials and methods
Polyacrylamide gels with gradient in stiffness
25-mm circular glass coverslips were treated first with ethanol and then RCA solution (1:1:3 for 15 N NH4OH/30% H2O2/distilled H2O) and functionalized with 1% allyltrichlorosilane and 1% triethylamine in chloroform solution. To control the gel’s stiffness, N,N′methylene-bis-acrylamide (Sigma-Aldrich) and the acrylamide solution (40%; Sigma-Aldrich) were mixed at the final concentrations in PBS. Solution was polymerized on a coverslip with 0.1% ammonium persulfate and 0.1% N,N,N′,N′-tetramethylethylenediamine. During polymerization, gels were covered with another coverslip that had been pretreated with dichlorodimethylsilane. Gels with gradients in stiffness were prepared using a method modified from Lo et al. (2000). We made gradient gels by juxtaposing two drops (13 µl for each drop) of different acrylamide and bisacrylamide concentrations on a 25-mm circular coverslip. The drop for the stiffer side contained 8% acrylamide and 0.3% bisacrylamide (∼34 kPa), and the drop for the softer side had 3% acrylamide and 0.11% bisacrylamide (∼1 kPa). FluoSpheres (Invitrogen) 1 µm in diameter were supplemented into the stiffer drop to a 0.005% by weight in the gel solution. This ensured that any possible stiffening effects of the beads would be on the stiff side of the gel. After waiting 5 min for gel polymerization, the two drops were mixed by applying a 25-mm square coverslip precoated with dichlorodimethylsilane. Gradient gels were allowed to polymerize for 45 min followed by removing the dichlorodimethylsilane coverslip and washing twice with PBS. Sulfo-SANPAH (Thermo Fisher Scientific) was diluted in Hepes buffer, pH 8.0, to 0.5 mg/ml, and 300 µl was added to the gel and then reacted using a 365-nm UV light for 10 min. Excess sulfo-SANPAH was removed by two washes of Hepes buffer. Type I rat tail collagen was diluted to 0.2 mg/ml in cold Hepes buffer, pH 8.0, and incubated with the gel at 4°C for 4 h. Collagen was removed, and the gel was washed and equilibrated with PBS. MSCs were plated onto gels within 24 h of collagen attachment. The spatial resolution of AFM used here is ≤1 µm, whereas much larger probes of a diameter of 640 µm (Lo et al., 2000) had suggested a steeper, steplike change in elasticity for similar gradient gel systems used to first describe durotaxis.
Cells, culture, pharmacological perturbations, and transfections
At least three different batches of human bone marrow–derived MSCs were purchased from Lonza, and at least three batches of MSCs were isolated from human donor bone marrows (University of Pennsylvania School of Medicine) by standard methods (Orlic et al., 2001) so that cells from at least six different human donors were used here. We did not detect major donor or batch variability in generating the reasonably consistent set of results here. Cells were used at passages 3–5 for all experiments and cultured in low glucose Dulbecco’s minimum essential medium (DMEM) supplemented with 10% FBS, 100 µg/ml penicillin, and 100 µM streptomycin. Racemic blebbistatin (EMD) was used at 5 or 50 µM as specified. Plasmid constructs for GFP–nonmuscle MIIB and GFP–nonmuscle MIIA were obtained from Addgene, GFP-MIIA S1943A was constructed by standard methods, and GFP-MIIA S1943D was a gift from A. Bresnick (Albert Einstein College of Medicine, Bronx, NY). Constructs were transfected via electroporation following the recommended procedure of the kit for MSCs (Nucleofector; Lonza) or by using Lipofectamine LTX (Invitrogen) with plus reagent using 0.5 g DNA per well in a 6-well plate. Transfection levels were similar for the three constructs (within 20%) based on GFP intensities and densitometry of Western blots.
FRAP
Confocal time-lapse imaging was acquired at 37°C with 5% CO2 in a humidified chamber with an inverted spinning-disk microscope (IX-81; Olympus) with a 14-bit high-resolution charge-coupled device (CCD) camera (HQ2; Photometrics) with MetaMorph software (Molecular Devices). Images were acquired with a 60× water immersion lens, NA 1.2, every 5 s for ≥150 s after 25 s of prebleached acquisition. Cells were imaged in low glucose DMEM (Invitrogen) with 10% FBS (Sigma-Aldrich). GFP-tagged myosin constructs were photobleached using a region of interest line drawn perpendicular to the actin bundles. Image sequences were analyzed with Fiji with prewritten Jythons script algorithm (National Institutes of Health) to quantify fluorescence recovery kinetics along myosin-labeled actin bundles.
Western blotting
Unless otherwise stated, cells were trypsinized, pelleted, and resuspended in PBS. Then, equal volume of 50 mM Tris-HCl, pH 8.0, with 10% SDS, 1 mM EDTA, 15% sucrose, 5% β-mercaptoethanol, and bromophenol blue was vortexed and boiled for 5 min followed by shearing through a 25-gauge needle. Samples were loaded onto 3–8% Tris-acetate gels (NuPAGE Novex; Life Technologies) and then transferred to blotting paper using a blotting system (iBlot; Life Technologies) with the settings of P3 for 9 min. Blots were blocked with 5% milk in TBST (TBS with Tween) for 1 h at RT, and then, antibodies in TBS were added overnight at 4°C. Secondary antibodies with conjugated HRP in TBST were added for 1 h at RT. ChromoSensor (GenScript) was added for the ECL reaction. We routinely performed Western blotting with different loads, and established linearity between signal versus sample load.
siRNA KD of MIIA and MIIB
Lipofectamine 2000 (Invitrogen) with 30 nM siRNA was used to knock down MIIA and MIIB. Different levels of KD were assessed by waiting different times after adding the complex and then quantified by immunofluorescence, Western blotting, and MS. For MIIB siRNA duplexes, the sequences were obtained from Bao et al. (2005) and were synthesized by Thermo Fisher Scientific along with scrambled siRNA. MIIA siRNA sequence was 5′-GGCCAAACCUGCCGAAUAAAUU-3′ with complement sequence 5′-UUUAUUCGGCAGGUUUGGCCUU-3′. Scramble siRNA was siGENOME nontargeting siRNA #1 (Thermo Fisher Scientific). For double KD of MIIA and MIIB, 30 nM of each duplex was used with Lipofectamine.
Fixation, immunostaining, and microscopy
Cells were fixed with 3.7% formaldehyde (Sigma-Aldrich) in PBS for 10 min at RT followed by PBS washing 2× for 5 min. Blocking and antibody staining was performed in 1% BSA in PBS. Rabbit anti-MIIA antibody (Sigma-Aldrich) was used at 1:100, anti-MIIB antibody (Cell Signaling Technology) was used at 1:150, anti-pS1943 was used at 1:100 (Cell Signaling Technology), and all primary antibodies were incubated at RT for 1 h or overnight at 4°C. All donkey secondary antibodies (Alexa Fluor dyes 488, 564, and 647) were stained for 1 h at RT at 1:300 dilution in PBS in 1% BSA. TRITC-phalloidin (Sigma-Aldrich) was used with the donkey secondary antibodies at a concentration of 100 ng/ml, and Hoechst 33342 (Invitrogen) was used to stain DNA at a concentration of 1 µg/ml for 10 min at RT. Imaging for quantification of myosin-II levels and localization was performed using an inverted microscope (IX-71; Olympus) with a 40× LUCPlanFLN objective, NA 0.60, and a CCD camera (Cascade; Photometrics). Image acquisition was performed with Image Pro software (Media Cybernetics), and subsequently, background was subtracted, and image analysis was performed using ImageJ (National Institutes of Health) according to JCB guidelines.
Image analysis
For quantitative immunofluorescence, samples that were directly compared were imaged at the same sitting, and the same gain and exposure time were used. If different exposure times were necessary for quantification between samples, several exposure times were used on the same image to ensure that images were in the linear range of exposure time to fluorescence intensity. For orientation analysis performed in Fig. 5, the directionality analysis was used in Fiji with the local gradient orientation method selected and using 90 bins for data segmentation. Curve fits of the histograms of data were used if there were high goodness of fits, and the ratio of the peak of the curve fit to the value of the baseline was taken as the oriented/isotropic parameter. Front/rear polarization of F-actin and myosins was determined by drawing a line 6–10 µm in width from the cell rear to front and plotting the fluorescence versus length. The ratio is the fluorescence in the front half over the back half of the cell.
AFM indentation
A force microscope (1-D; Asylum Research) was used quantify the elastic modulus of the gradient gels. A pyramid-tipped probe with spring constant of 30–100 pN/nm (Veeco) was used for measuring gradient gels, and the AFM head was placed over an inverted microscope (TE300; Nikon) to detect the beads with fluorescent microscopy using a 4× objective. A micromanipulator was used to move across the gradient during AFM probing and ensured high accuracy determinations of the elasticity gradients when compared with the methods of Lo et al. (2000).
Time-lapse cell imaging
Phase-contrast imaging was performed in a humidified chamber at 37°C and 5% CO2 using an inverted microscope (IX-71) with a 10× objective, NA 0.3, using a 300-W xenon lamp illumination and a high-resolution CCD camera (CoolSNAP HQ; Photometrics). softWoRx software (DeltaVision) was used for acquiring images. Cells were imaged in low glucose DMEM with 10% FBS. For cell speed measurements, images were acquired in 3-min intervals for 2 h. The center of the cell nucleus was used as a reference point in cell tracking. For imaging at longer times (≥12 h), images were acquired at 15-min intervals. ImageJ was used to track the center of the nuclei of video sequences, and the summed contour distance traveled divided by the time was used as our measurement of speed. This measurement is independent of how persistent the cells are and instead reflects the total distance traveled. Cells that began to divide or were in contact and cells that migrated out of the trackable region were included until such events required exclusion. In time-lapse imaging for several hundred cells on soft, stiff, and gradient matrices, we did not observe cell detachment or cell loss, and we did observe about an equal number of cell division events on all matrices, which largely eliminates any bias in the measurements here.
Triton X-100 extraction of MSCs
MSCs on either plastic or gels were treated with 0.1% Triton X-100 in PEM buffer (0.1 M Pipes, pH 6.95, 2 mM EGTA, and 1 mM MgSO4) with 2 µM phalloidin (Sigma-Aldrich) and protease inhibitors (Sigma-Aldrich) for 4 min, and then, the cell ghosts were fixed and immunostained to determine the fraction of insoluble myosin. For Western blots of soluble and insoluble lysates, cells with transfected myosin constructs were Triton X-100 extracted as previously described (Breckenridge et al., 2009). In brief, 600,000 transfected MSCs (using nucleofection) were plated onto a 10-cm dish, and after 24 h, cells were rinsed with PBS, and 300 µl lysis buffer (50 mM Tris-HCl, 5 mM NaCl, 140 mM Na-acetate, 0.6% Triton X-100, 5 mM EGTA, and 1 mM EDTA with protease inhibitors) was added for 4 min during which the plate was swirled for liquid to cover cells. The plate rested on ice between swirling. Then, the extracted cell ghosts were scraped, and the total solution was collected, sheared through a 25-gauge syringe needle, and then centrifuged for 10 min at 4°C at 10,000 g. The pelleted fraction is considered to contain the insoluble cytoskeleton, and the supernatant is the soluble portion. 4× lithium dodecyl sulfate (LDS) buffer was added to the supernatant, and LDS (diluted in PBS) was added to the pellet to create a volume of 50 µl. Samples were boiled for 5 min and then loaded onto the gel.
Collagen-I overlay
MSCs were seeded onto collagen-coated polyacrylamide gels and allowed to adhere for 1–2 h. Media were removed, and 1.65 mg/ml collagen in 50:50 neutral buffered solution/complete growth media was added to the polyacrylamide gel surface and allowed to gel for 1 h at 37°C followed by the gentle addition of media. Cells were fixed and stained as previously described for this collagen overlay culture system (Fischer et al., 2009) with minor alterations. In brief, samples were fixed with 3.7% formaldehyde in PBS with 0.1% Triton X-100 for 30 min at RT and then rinsed with PBS with 0.25% Triton X-100 for 30 min. PBS with 0.5 mg/ml NaBH4 was added to reduce autofluorescence for 10 min at RT twice. Samples were subsequently blocked with PBS with 2% BSA, 1% goat serum, and 0.25% Triton X-100 overnight at 4°C. Primary antibodies were diluted in PBS with 0.25% Triton X-100 with 2% BSA and 1% goat serum at 4°C overnight. After rinsing for 30 min in PBS, secondary antibodies at 1:300 were added and incubated for 2.5 h at RT.
Calculation of cell migration exponent
The root–mean squared displacement is assumed to scale with time as RMSD(t) = A × tα(t), with A being a constant and α determined from α = d[log(RMSD)]/d[log(t)] as calculated in this paragraph. Cell motility is perfectly persistent and directed if α = 1, but the cell is executing a random walk if α = 0.5. For an accelerating cell, α > 1, and for a stalled cell, α = 0. In each region of a substrate, this “migration exponent” α is determined for every cell at each time point and then averaged among all cells. The time dependence of α leads to an additional but small term upon differentiation:
MS sample preparation
For proteomic experiments (Fig. S3), cultured MSCs subjected to MIIA KD, MIIB KD, or treatment with scrambled siRNA were harvested with 0.05% trypsin and washed three times by successive resuspension in ice-cold PBS. Proteins were solubilized by cellular disruption with a probe sonicator in ice-cold radioimmunoprecipitation assay buffer with 0.1% protease inhibitor cocktail (∼5,000 cells/µl). NuPAGE LDS sample buffer (Invitrogen) with 1% β-mercaptoethanol was added to a 1× concentration followed by heating to 80°C for 10 min. Proteins were separated on SDS-PAGE gels (NuPAGE 4–12% Bis-Tris) and run at 100 V for 10 min followed by 25 min at 160 V. Sections of excised polyacrylamide gel (cut in two molecular mass ranges: 55–160 and ≥160 kD) were washed (50% of 0.2 M ammonium bicarbonate [AB] solution and 50% acetonitrile for 30 min at 37°C), dried by lyophilization, incubated with a reducing agent (20 mM tris(2-carboxyethyl)phosphine in 25 mM AB solution, pH 8.0, for 15 min at 37°C) and alkylated (40 mM iodoacetamide in 25 mM AB solution, pH 8.0, for 30 min at 37°C). The gel sections were dried by lyophilization before in-gel trypsinization (20 µg/ml of sequencing grade modified trypsin in buffer as described in the manufacturer’s protocol [Promega] for 18 h at 37°C with gentle shaking). Before analysis, peptide solutions were acidified by addition of 50% digest dilution buffer (60 mM AB solution with 3% methanoic acid).
MS protocol
Peptide separations (5-µl injection volume) were performed on a 15-cm column (75-µm inner diameter; PicoFrit; New Objective) packed with 5-µm C18 reversed-phase resin (Magic; Michrom Bioresources) using a nanoflow high-pressure LC system (Eksigent Technologies), which was coupled online to a hybrid mass spectrometer (LTQ Orbitrap XL; Thermo Fisher Scientific) via a nanoelectrospray ion source. Chromatography was performed with solvent A (water [Milli-Q; Millipore] with 0.1% formic acid) and solvent B (acetonitrile with 0.1% formic acid). Peptides were eluted at 200 nl/min for 3–28% B over 42 min, 28–50% B over 26 min, 50–80% B over 5 min, and 80% B for 4.5 min before returning to 3% B over 0.5 min. To minimize sample carryover, a fast blank gradient was run between each sample. The LTQ Orbitrap XL was operated in the data-dependent mode to automatically switch between full-scan MS (mass per charge [m/z] = 350–2,000) in the Orbitrap analyzer (with resolution of 60,000 at m/z = 400) and the fragmentation of the six most intense ions by collision-induced dissociation in the ion trap mass analyzer.
Raw mass spectroscopy data were processed using Elucidator (version 3.3; Rosetta Biosoftware). The software was set up to align peaks in data from samples derived from the same ranges of molecular weight. Peptide and protein annotations were made using SEQUEST (Thermo Fisher Scientific) with full tryptic digestion and up to two missed cleavage sites. Peptide masses were selected between 800 and 4,500 amu with peptide mass tolerance of 1.1 amu and fragment ion mass tolerance of 1.0 amu. Peptides were searched against a database compiled from UniRef100 human (for proteomic experiments; downloaded on 5 November, 2010) plus contaminants and a reverse decoy database. Search results were selected with a Δ cross-correlation filter of 0.01 and a mass error better than 20 ppm. Ion currents of oxidized peptides (Δ = 15.995 D) were summed with their parent peptide; posttranslational modification sites of phosphorylation (Δ = 79.966 D), acetylation (Δ = 42.011 D), and methylation (Δ = 14.016 D) were considered in the search. In matrix experiments, we additionally looked for hydroxylation of asparagine, aspartic acid, proline, and lysine (Δ = 15.995 D).
The two molecular mass ranges of the proteomic dataset were analyzed separately. In the mid–molecular mass range (55–160 kD), the false-positive detection rate was estimated to be 11.4% (based on search hits of the decoy database), and only proteins with two or more peptides/protein were considered for further analysis (2,015 peptides from 231 unique proteins). High–molecular mass range (≥160 kD) resulted in a false-positive rate of 11.3% with subsequent analysis of 1,223 peptides from 55 unique proteins. Label-free relative peptide quantitation was performed with in-house software coded for Mathematica (Wolfram Research). Datasets were normalized against optimized housekeeping peptide sets that were found to be invariant between experimental conditions. A peptide set optimization algorithm (peptide ratio fingerprint [PRF]; Shin et al., 2011) was used to select peptides that show a similar “fingerprint” behavior between samples, and these peptides were used for the basis of quantification and normalization. We report only quantification of proteins with at least three PRF peptides/protein (total of 178 proteins). Peptides from regions common to several proteins or isoforms were treated distinctly. Standard errors were calculated from at least two technical repetitions. As a further check of the peptide selection algorithm, ratio comparisons were made between all datasets and checked for consistency (for example, when considering data A, B, and C, the ratio A/B should be consistent with A/C × C/B).
Online supplemental material
Fig. S1 shows differences in cell migration on soft and stiff gels. Fig. S2 illustrates phenotypical differences observed with myosin-II KD experiments. Fig. S3 quantifies results from MS on cells with myosin-II KD. Fig. S4 summarizes effects from blebbistatin on cell migration and shape. Fig. S5 quantifies differences in cell area and pS1943 levels with myosin mutant transfections.
Acknowledgments
The authors thank Dr. Anne Bresnick for kindly providing the construct for S1943D-MIIA, Dr. Thomas Egelhoff (Cleveland Clinic, Cleveland, OH) for helpful discussions and experimental suggestions, Dr. Alex Mogilner (University of California, Davis, Davis, CA) for discussions of polarization modeling, and Dr. Julie Theriot (Stanford University, Stanford, CA) for discussions of keratocyte and neutrophil polarization. Careful reading of the revised manuscript by Dr. Fei Wang (University of Illinois, Chicago, IL), Dr. Tatiana Svitkina (University of Pennsylvania, Philadelphia, PA), Dr. Ryan Petrie (National Institutes of Health, Bethesda, MD), and Dr. Wenting Shih (University of California, Davis) is very gratefully acknowledged. We also thank Andrea Stout and the Cell and Developmental Biology Microscopy Core at the Perelman School of Medicine, where time-lapse imaging was performed under precisely controlled temperature, humidity, and carbon dioxide conditions.
Support from the National Institutes of Health (grants P01DK032094 and R01HL062352) and National Science Foundation (University of Pennsylvania’s Nano Science and Engineering Center-Nano Bio Interface Center) is very gratefully acknowledged.