Accurate DNA replication requires proper regulation of replication licensing, which entails loading MCM-2–7 onto replication origins. In this paper, we provide the first comprehensive view of replication licensing in vivo, using video microscopy of Caenorhabditis elegans embryos. As expected, MCM-2–7 loading in late M phase depended on the prereplicative complex (pre-RC) proteins: origin recognition complex (ORC), CDC-6, and CDT-1. However, many features we observed have not been described before: GFP–ORC-1 bound chromatin independently of ORC-2–5, and CDC-6 bound chromatin independently of ORC, whereas CDT-1 and MCM-2–7 DNA binding was interdependent. MCM-3 chromatin loading was irreversible, but CDC-6 and ORC turned over rapidly, consistent with ORC/CDC-6 loading multiple MCM-2–7 complexes. MCM-2–7 chromatin loading further reduced ORC and CDC-6 DNA binding. This dynamic behavior creates a feedback loop allowing ORC/CDC-6 to repeatedly load MCM-2–7 and distribute licensed origins along chromosomal DNA. During S phase, ORC and CDC-6 were excluded from nuclei, and DNA was overreplicated in export-defective cells. Thus, nucleocytoplasmic compartmentalization of licensing factors ensures that DNA replication occurs only once.
Introduction
Faithful error-free DNA replication is essential to maintain genome integrity. To prevent rereplication of DNA segments, eukaryotes divide DNA replication into two nonoverlapping phases (Blow and Dutta, 2005; DePamphilis et al., 2006). During the first phase, which takes place in late mitosis and early G1, replication origins are licensed for replication by loading Mcm2–7 double hexamers. During S phase, CDKs and Dbf4-dependent kinases activate the Mcm2–7 helicase by promoting its interaction with Cdc45 and the GINS complex (Moyer et al., 2006; Ilves et al., 2010). This Cdc45-MCM-GINS helicase moves ahead of the replication fork to unwind the template DNA, allowing DNA polymerases access to single-stranded DNA. Preventing licensing during S phase ensures that no DNA is rereplicated.
Licensing occurs by the stepwise assembly of prereplicative complex (pre-RC) proteins at origins. Origin recognition complex (ORC), Cdc6, Cdt1, and Mcm2–7 are the minimal components required for origin licensing in vitro (Gillespie et al., 2001; Evrin et al., 2009; Remus et al., 2009). Current data suggest that the ORC complex binds first and recruits Cdc6 and Cdt1, which together then load Mcm2–7 complexes onto chromatin, thereby licensing the origin (Blow and Dutta, 2005; DePamphilis et al., 2006). This involves ring-shaped Mcm2–7 hexamers being clamped around origin DNA (Evrin et al., 2009; Remus et al., 2009). It is essential that a large enough number of origins is licensed before entry into S phase and that these licensed origins are distributed along chromosomal DNA without excessively large gaps. It is unclear how this distribution is achieved but may involve a dynamic interaction between the pre-RC proteins and DNA.
To prevent rereplication, it is essential that the ability to license new origins is shut down before entry into S phase. This is achieved by a range of mechanisms, which vary between different organisms and cell types (Blow and Dutta, 2005; Drury and Diffley, 2009). In yeasts, phosphorylation of pre-RC proteins by S phase CDKs promotes either their inactivation, degradation, or nuclear export. In animal cells, different regulatory mechanisms predominate, particularly the inhibition of Cdt1 by geminin and proliferating cell nuclear antigen–dependent degradation of Cdt1. In Caenorhabditis elegans, for instance, GMN-1 (geminin) has been shown to bind and inhibit CDT-1 (Yanagi et al., 2005), CDC-6 is excluded from nuclei (Kim et al., 2007), and CUL-4 promotes CDT-1 degradation (Zhong et al., 2003).
Although licensing has been extensively studied by biochemical and genetic approaches (Blow and Dutta, 2005; DePamphilis et al., 2006), we know very little about the dynamics of licensing proteins in vivo. The C. elegans life cycle provides a unique opportunity to study the dynamics of origin licensing during meiotic and early embryonic cell divisions. Within the gonad of an adult worm, the differentiation of a mature oocyte from a mitotic germ cell takes ∼1 d, during which time genes can be inactivated by RNAi. Shortly after fertilization, the embryo resumes meiosis, completing the two meiotic divisions (meiosis I and II). After meiosis, the oocyte and sperm chromatin decondenses, and the two pronuclei grow in size at opposite poles of the embryo. During this time, the bulk of DNA replication occurs (Edgar and McGhee, 1988). The pronuclei then migrate toward each other and center in the embryo while chromosomes condense. After nuclear envelope breakdown, the first mitosis gives rise to two blastomeres of different sizes and fates. This sequence of events is shown in Video 1. Analysis of these cell cycles is facilitated by the speed of meiotic and early zygotic divisions, their invariant nature, and the relatively weak cell cycle checkpoints, which allow continued cycling even when essential cell cycle processes are defective (Encalada et al., 2000; Brauchle et al., 2003).
Video of an embryo expressing GFP–MCM-3 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Images were acquired every 30 s with a widefield fluorescent microscope (DeltaVision DV3; Applied Precision). The animated diagram (right) shows GFP–MCM-3 chromatin or nuclear enrichment over time, and the timeline helps the reader to recognize the cell cycle stages. Time is indicated in minutes and seconds. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Video of an embryo expressing GFP–MCM-3 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Images were acquired every 30 s with a widefield fluorescent microscope (DeltaVision DV3; Applied Precision). The animated diagram (right) shows GFP–MCM-3 chromatin or nuclear enrichment over time, and the timeline helps the reader to recognize the cell cycle stages. Time is indicated in minutes and seconds. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Here, we provide a comprehensive analysis of replication licensing in vivo, using time-lapse video microscopy and antibody staining of C. elegans early embryos. We show that chromatin licensing during late M phase alters the dynamics of licensing factors to promote the distribution of licensed origins throughout the genome. When cells exit M phases, nuclear export excludes licensing factors from the nucleus, switching off origin licensing to prevent rereplication.
Results
Licensing in the early embryo
To analyze origin licensing in vivo, we generated worms expressing functional GFP–MCM-3 (see Materials and methods) and stained with pan-MCM antibodies that recognize all six MCM-2–7 subunits (see Fig. S1). To visualize chromatin, GFP–MCM-3–expressing strains were crossed with worms expressing mCherry–Histone H2B (McNally et al., 2006). The resulting embryos were then used to examine the localization of GFP–MCM-3 from anaphase of meiosis I, through meiosis II, and to the end of the first embryonic cell cycle (Fig. 1, A [arrowheads] and E; and Video 1). GFP–MCM-3 became enriched on chromosomes during anaphase of meiosis II and during metaphase and anaphase of mitosis. GFP–MCM-3 was also massively accumulated in pronuclei and nuclei during interphase and prophase. The ratio of chromatin-associated GFP–MCM-3 signal relative to the cytoplasmic signal is shown in the diagram of Fig. 1 A and in the graph of Video 1. Worms expressing GFP–MCM-2 showed an identical localization to that of GFP–MCM-3 during the first embryonic cell cycle (Video 2), consistent with both of them being part of the hexameric MCM-2–7 complex. The chromatin binding of GFP–MCM-3 during meiosis and mitosis reflects the expected behavior of MCM-2–7 proteins participating in origin licensing, whereas during interphase, the chromatin association of GFP–MCM-3 appears to be masked by nuclear accumulation of soluble protein.
Video of an embryo expressing GFP–MCM-2 progressing throughout the first embryonic cell cycle. Images were acquired every 10 s with a widefield fluorescent microscope (DeltaVision DV3).
Video of an embryo expressing GFP–MCM-2 progressing throughout the first embryonic cell cycle. Images were acquired every 10 s with a widefield fluorescent microscope (DeltaVision DV3).
Visualizing origin licensing by GFP–MCM-3. (A–D) Images taken from time-lapse sequences of embryos progressing through the first (I) and the second (II) meiotic division (left panels) and the first embryonic cell cycle (right panels). Embryos are orientated with the anterior pointing to the top left corner with GFP–MCM-3 (left images in each panel) and mCherry-Histone (right images in each panel). Meiotic divisions are shown at 3x higher magnification. Asterisks indicate polar bodies. Arrowheads point to GFP–MCM-3 chromatin binding in anaphase of meiosis II and mitosis. Diagrams show the enrichment of GFP–MCM-3 signal on chromatin or in pronuclei and nuclei relative to the cytoplasm. Values are plotted over time. Each tick on the x axis represents 5 min. Cell cycle stages are indicated. Bars, 10 µm. (A) Wild-type embryo. (B) mcm-7 RNAi embryo. (C) cdc-6 RNAi embryo. (D) gmn-1 (geminin) RNAi embryo. (E) Mean GFP–MCM-3 chromatin enrichment of five embryos according to cell cycle stage. Error bars are SDs. See also Video 1 for wild type.
Visualizing origin licensing by GFP–MCM-3. (A–D) Images taken from time-lapse sequences of embryos progressing through the first (I) and the second (II) meiotic division (left panels) and the first embryonic cell cycle (right panels). Embryos are orientated with the anterior pointing to the top left corner with GFP–MCM-3 (left images in each panel) and mCherry-Histone (right images in each panel). Meiotic divisions are shown at 3x higher magnification. Asterisks indicate polar bodies. Arrowheads point to GFP–MCM-3 chromatin binding in anaphase of meiosis II and mitosis. Diagrams show the enrichment of GFP–MCM-3 signal on chromatin or in pronuclei and nuclei relative to the cytoplasm. Values are plotted over time. Each tick on the x axis represents 5 min. Cell cycle stages are indicated. Bars, 10 µm. (A) Wild-type embryo. (B) mcm-7 RNAi embryo. (C) cdc-6 RNAi embryo. (D) gmn-1 (geminin) RNAi embryo. (E) Mean GFP–MCM-3 chromatin enrichment of five embryos according to cell cycle stage. Error bars are SDs. See also Video 1 for wild type.
To test whether the localization of GFP–MCM-3 depends on other MCM-2–7 subunits, MCM-7 was knocked down by RNAi. In mcm-7 RNAi embryos, GFP–MCM-3 did not localize to DNA during M phases and did not accumulate in pronuclei and nuclei (Fig. 1, B and E). This result is consistent with the idea that GFP–MCM-3 is part of a hexameric MCM-2–7 complex and that complex formation is required for origin licensing and nuclear accumulation. Embryos stained with pan-MCM antibodies that recognize all six MCM-2–7 subunits showed a similar pattern to that observed in live experiments (Fig. S1 A), consistent with a previous study (Korzelius et al., 2011). The pan-MCM antibodies also showed additional centrosome staining (indicated by an asterisk in Fig. S1 A), which might either be unspecific or a result of the location of MCM subunits other than MCM-3 and MCM-7. Like GFP–MCM-3, the pan-MCM antibody staining on anaphase chromatin and interphase nuclei was abolished by mcm-7 RNAi (Fig. S1 B).
Origin licensing is expected to occur before entry in S phase and be dependent on the pre-RC proteins ORC, CDC-6, and CDT-1. To confirm that the binding of MCM-2–7 to chromatin during anaphase indeed represents origin licensing, we inactivated other pre-RC proteins and examined the localization of GFP–MCM-3. RNAi of cdc-6 and cdt-1 resulted in embryonic lethality, cell cycle delay, and appearance of anaphase chromosome bridges, whereas the inactivation of orc-5 caused a partial embryonic lethality consistent with previous studies (Zhong et al., 2003; Sönnichsen et al., 2005; Kim et al., 2007). In cdc-6, cdt-1, and orc-5 RNAi embryos, GFP–MCM-3 and MCM-2–7 were strongly reduced on chromatin during anaphase of meiosis II and mitosis, whereas the nuclear localization during interphase was not affected (Figs. 1 [C and E] and S1 [C and D]). Inactivation of Y47D3A.29, the DNA polymerase-α (pol-α) catalytic subunit, did not affect GFP–MCM-3 localization (Fig. 1 E). In summary, our results suggest that the observed binding of MCM-2–7 to chromatin during anaphase of meiosis II and late mitosis represents origin licensing.
C. elegans geminin
Geminin negatively regulates origin licensing by binding and inhibiting CDT-1 (Wohlschlegel et al., 2000; Tada et al., 2001). C. elegans geminin, GMN-1, binds CDT-1 and is required for proper cell division in intestinal and germ cells of adult worms (Yanagi et al., 2005). In gmn-1 RNAi embryos, GFP–MCM-3 localized to chromatin earlier than in wild type, becoming visible from anaphase of meiosis I and from prometaphase of mitosis (Fig. 1, D and E). Therefore, GMN-1 inhibits licensing during anaphase of meiosis I, metaphase of meiosis II, and prometaphase of mitosis. This suggests that ORC, CDC-6, and CDT-1 can be active at these periods (see Figs. 3 [A and F] and 4 A). However, we saw no obvious cell cycle or chromosome defects in gmn-1 RNAi embryos.
Visualizing S phase in the early embryo
To demonstrate that origin licensing is required for replication, we sought to visualize replication forks by GFP-tagging fork proteins. GFP–SLD-5 (a component of the GINS complex), GFP–RPA-1, and GFP–DIV-1 (primase) all accumulated in the nucleus both during S phase and prophase, as was described for GFP–PCN-1 (proliferating cell nuclear antigen; Brauchle et al., 2003), and therefore do not reflect ongoing DNA replication. In contrast, GFP–CDC-45 was enriched in pronuclei during S phase but not during prophase (Fig. 2 A and Video 3). The nuclear GFP–CDC-45 colocalized with chromatin, consistent with its association with replication forks (Fig. 2 A, arrowhead). Furthermore, blocking licensing by mcm-7 RNAi abolished GFP–CDC-45 nuclear enrichment (Fig. 2 A). These observations suggest that GFP–CDC-45 is enriched in nuclei because of its binding to DNA rather than being retained in the nucleoplasm. The relative amount of GFP–CDC-45 engaged in replication can be estimated by multiplying the nuclear volume by the nuclear enrichment (Fig. 2 A, bottom diagram). By this measure, GFP–CDC-45 persists at >20% of its maximum level for 8.5 min, which is our estimate for S phase length (Fig. 2 B). In pol-α RNAi embryos, GFP–CDC-45 remained enriched in pronuclei for longer than in wild type (Fig. 2, A and B), consistent with S phase being extended as a consequence of compromised replication fork progression. These data are consistent with chromatin-bound MCM-2–7 proteins being required for the assembly of active replisomes during S phase.
Video of an embryo expressing GFP–CDC-45 (left) and mCherry-Histone (middle) progressing throughout meiosis II and the first embryonic cell cycle. The display is similar to Video 1, except that images were acquired every 10 s. Images were taken with a spinning-disk confocal microscope. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Video of an embryo expressing GFP–CDC-45 (left) and mCherry-Histone (middle) progressing throughout meiosis II and the first embryonic cell cycle. The display is similar to Video 1, except that images were acquired every 10 s. Images were taken with a spinning-disk confocal microscope. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Visualizing replication forks by GFP–CDC-45 localization. (A) Images from time-lapse sequences taken by a spinning-disk confocal microscope of wild-type, mcm-7 RNAi, and pol-α RNAi embryos expressing GFP–CDC-45 (left images in each panel) and mCherry-Histone (right images in each panel). Cell cycle stages are indicated. Wild-type, mcm-7 RNAi, and pol-α RNAi embryos are shown in the left, middle, and right panels, respectively. Asterisks indicate the position of the polar body. Red arrowhead shows GFP–CDC-45 colocalizing with chromatin. Bar, 10 µm. The top diagrams immediately below the time-lapse series show enrichment of nuclear GFP–CDC-45 relative to the cytoplasm. The bottom diagrams indicate the relative amount of nuclear GFP–CDC-45 (fluorescence intensity multiplied by nuclear volume). The blue lines are smoothings of the data by averaging 10 successive time points. (B) Estimate of S phase length in wild-type and pol-α RNAi embryos by quantification of the period in which GFP–CDC-45 is enriched on chromatin by >20% of its maximum. n = 5. Error bars represent SD. See also Video 3 for wild type.
Visualizing replication forks by GFP–CDC-45 localization. (A) Images from time-lapse sequences taken by a spinning-disk confocal microscope of wild-type, mcm-7 RNAi, and pol-α RNAi embryos expressing GFP–CDC-45 (left images in each panel) and mCherry-Histone (right images in each panel). Cell cycle stages are indicated. Wild-type, mcm-7 RNAi, and pol-α RNAi embryos are shown in the left, middle, and right panels, respectively. Asterisks indicate the position of the polar body. Red arrowhead shows GFP–CDC-45 colocalizing with chromatin. Bar, 10 µm. The top diagrams immediately below the time-lapse series show enrichment of nuclear GFP–CDC-45 relative to the cytoplasm. The bottom diagrams indicate the relative amount of nuclear GFP–CDC-45 (fluorescence intensity multiplied by nuclear volume). The blue lines are smoothings of the data by averaging 10 successive time points. (B) Estimate of S phase length in wild-type and pol-α RNAi embryos by quantification of the period in which GFP–CDC-45 is enriched on chromatin by >20% of its maximum. n = 5. Error bars represent SD. See also Video 3 for wild type.
The ORC
ORC binds origins and is essential for origin licensing. ORC contains six subunits, Orc1 to Orc6. The ORC core complex contains Orc2–5, whereas Orc1, whose level fluctuates during cell cycle progression, associates with Orc2–5 and promotes its binding to DNA (DePamphilis et al., 2006). Orc6 only weakly associates with Orc1–5 in metazoans. We performed reciprocal blast searches with human ORC sequences to identify the corresponding C. elegans genes, confirming the identity of ORC-1 (Y39A1A.12), ORC-2 (F59E10.1), and ORC-5 (ZC168.3). Then, we created transgenic worms expressing GFP–ORC-1, GFP–ORC-2, and GFP–ORC-5. To confirm that ORC-2 and ORC-5 associate together and to identify other subunits of the ORC complex, we used high-affinity GFP antibodies (Rothbauer et al., 2008) to purify proteins associated with GFP–ORC-2 and GFP–ORC-5. GFP-tagged PCN-1 and SLD-5 and nontransgenic wild-type worms were used as specificity controls. Of the 66 proteins enriched >10 fold in both ORC-2 and ORC-5 precipitations, the four most abundant were ORC-2, ORC-5, Y39A1A.13, and Y113D3B.11 (Table S1). Y39A1A.13 is encoded in the same operon as ORC-1 and appears to be the C. elegans orthologue of ORC-4 (Fig. S2 A), whereas Y113D3B.11 appears to be C. elegans ORC-3 (Fig. S2 B).
GFP–ORC-2 was enriched on chromatin during anaphase of meiosis I, metaphase and anaphase of meiosis II, and during all of mitosis (Fig. 3, A [red arrowheads] and E; and Video 4). We observed a similar localization pattern using ORC-2 antibodies whose specificity was confirmed by RNAi (Fig. S3, A and C; Hadwiger et al., 2010). Therefore, binding of ORC to chromatin precedes that of MCM-2–7 but coincides with the loading of MCM-2–7 in gmn-1 RNAi embryos. These results are consistent with the known role of ORC in origin licensing. However, during interphase, GFP–ORC-2 was rapidly released from DNA and was partially excluded from the pronuclei and nuclei (Fig. 3 A [green arrowheads] and Video 4), which is in contrast to the reported behavior of Orc2 in other organisms.
Video of an embryo expressing GFP–ORC-2 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Video of an embryo expressing GFP–ORC-2 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
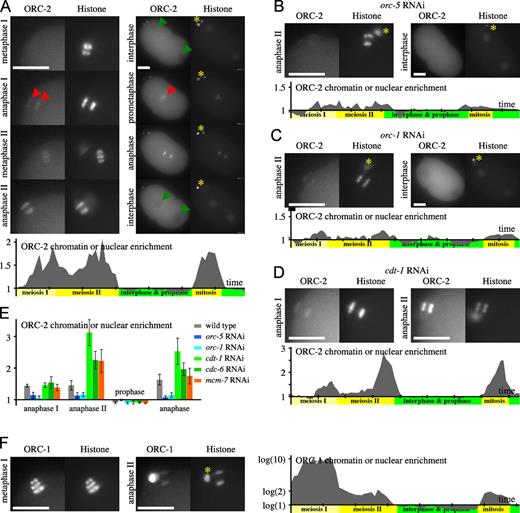
GFP–ORC-2 and replication origins. (A–D) Images from time-lapse sequences of embryos expressing GFP–ORC-2 (left images in each panel) and mCherry-Histone (right images in each panel), presented as for Fig. 1. (A) Wild-type embryo. Red arrowheads indicate the enrichment of GFP–ORC-2 on chromatin in anaphase of meiosis I and prometaphase of first mitosis. Green arrowheads show the position of interphase nuclei. (B) orc-5 RNAi embryo. (C) orc-1 RNAi embryo. (D) cdt-1 RNAi embryo. (E) GFP–ORC-2 chromatin enrichment (the mean of five embryos at various cell cycle stages). Error bars are SDs. (F) Embryos expressing GFP–ORC-1 and mCherry-Histone. Asterisks (A–C, F) indicate polar bodies. Bars, 10 µm. See also Videos 4 and 5.
GFP–ORC-2 and replication origins. (A–D) Images from time-lapse sequences of embryos expressing GFP–ORC-2 (left images in each panel) and mCherry-Histone (right images in each panel), presented as for Fig. 1. (A) Wild-type embryo. Red arrowheads indicate the enrichment of GFP–ORC-2 on chromatin in anaphase of meiosis I and prometaphase of first mitosis. Green arrowheads show the position of interphase nuclei. (B) orc-5 RNAi embryo. (C) orc-1 RNAi embryo. (D) cdt-1 RNAi embryo. (E) GFP–ORC-2 chromatin enrichment (the mean of five embryos at various cell cycle stages). Error bars are SDs. (F) Embryos expressing GFP–ORC-1 and mCherry-Histone. Asterisks (A–C, F) indicate polar bodies. Bars, 10 µm. See also Videos 4 and 5.
In orc-1, orc-3, orc-4, and orc-5 RNAi embryos, ORC-2 binding to DNA was reduced during meiosis and mitosis (Figs. 3 [B, C, and E] and S4 [C–F]). This suggests that ORC-2 DNA binding depends on the formation of an ORC-1–5 complex. Furthermore, GFP–ORC-2 was not excluded from pronuclei and nuclei upon orc-5 inactivation (Fig. 3, B and E) but remained excluded upon orc-1 inactivation (Fig. 3, C and E), indicating that the core ORC-2–5 complex is excluded from nuclei independently of ORC-1. In summary, we have identified ORC-3 and ORC-4 and provide evidence that the core ORC-2–5 complex is present in the cytoplasm and that it associates with DNA in an ORC-1–dependent manner.
The association of mammalian Orc1 with the Orc2–5 core complex and DNA is regulated by CDKs (Li et al., 2004; Siddiqui and Stillman, 2007; Baldinger and Gossen, 2009). We observed that GFP–ORC-1 was strongly associated with chromatin during metaphase of meiosis I (Figs. 3 F and S5 A and Video 5). A more modest binding was observed during meiosis II and mitosis, and it was excluded from pronuclei and nuclei. Thus, GFP–ORC-1 localization resembled that of GFP–ORC-2, apart from its massive binding to DNA during meiosis I and its more pronounced nuclear exclusion.
Video of an embryo expressing GFP–ORC-1 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Video of an embryo expressing GFP–ORC-1 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
In orc-5 and orc-2 RNAi embryos, but not cdt-1 or cdc-6 RNAi embryos, the protein level of GFP–ORC-1 was reduced on DNA and in the cytoplasm by ∼70% (Fig. S4, B and D), suggesting a potential role for the ORC-2–5 complex in ORC-1 stability. The residual GFP–ORC-1 in orc-5 RNAi embryos was enriched on DNA and excluded from pronuclei and nuclei, though the quantification of GFP–ORC-1 enrichment on DNA is probably affected by the reduced protein level (Figs. S4 B and S5 E). Therefore, whereas GFP–ORC-1 stability depends on other ORC subunits, GFP–ORC-1 binds DNA independently of other ORC subunits during metaphase of meiosis I.
Together, our results suggest that ORC localization appears more dynamic in C. elegans as compared with previously examined eukaryotes in which ORC associates with DNA throughout interphase (Blow and Dutta, 2005; DePamphilis et al., 2006). We show evidence for the nuclear exclusion of both ORC-2–5 and ORC-1 in C. elegans commencing with the formation of the nuclear envelope. Unlike other organisms in which the binding of Orc1 to DNA is strictly dependent on the core ORC-2–5 complex, C. elegans ORC-1 can bind chromosomal DNA in the absence of other ORC subunits.
The licensing factors CDC-6 and CDT-1
In eukaryotes, the two licensing factors Cdc6 and Cdt1 are recruited to chromatin by ORC and are required to load Mcm2–7 complexes. In C. elegans, CDC-6 and CDT-1 are required for DNA replication, and their overexpression induces rereplication (Zhong et al., 2003; Kim et al., 2007). We generated GFP–CDC-6–expressing worms and showed that GFP–CDC-6 was enriched on DNA during anaphase of meiosis I, throughout meiosis II, and during mitosis (Fig. 4, A [red arrowheads] and D; and Video 6). During interphase, GFP–CDC-6 was excluded from pronuclei and nuclei (Fig. 4 A, green arrowheads). The dynamic localization of CDC-6 was confirmed by staining embryos with CDC-6 antibodies (Fig. S3, A and B; Kim et al., 2007). Thus, the localization of CDC-6 resembles that of ORC-2. Interestingly, GFP–CDC-6 localization was not significantly affected by orc-5 RNAi (Fig. 4, B and D), even though GFP–MCM-3, GFP–ORC-2, and GFP–ORC-1 chromatin binding was strongly reduced under these conditions (Figs. 1 E, 3 B, and S5 B). To provide further evidence that the chromatin binding of CDC-6 does not depend on ORC, we simultaneously stained embryos with ORC-2 and CDC-6 antibodies. When orc-2, orc-3, orc-4, or orc-5 was inactivated, chromatin binding of ORC-2 was abolished, whereas CDC-6 remained enriched on DNA (Fig. S3, C–F). Our results suggest that the cell cycle–regulated chromatin binding of C. elegans CDC-6 can occur in the absence of chromatin-bound ORC. This is unlike other eukaryotes studied in which Cdc6 chromatin binding is ORC dependent (Blow and Dutta, 2005; DePamphilis et al., 2006).
Video of an embryo expressing GFP–CDC-6 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
Video of an embryo expressing GFP–CDC-6 (left) and mCherry-Histone (middle) progressing throughout meiotic divisions and the first embryonic cell cycle. Acquisition and display of the sequence were similar to that of Video 1. M, mitosis; MI, meiosis I; MII, meiosis II; Pro, Prophase; S, S phase.
GFP–CDC-6 and CDT-1 localization. (A–C) Images from time-lapse sequences of embryos expressing GFP–CDC-6 (left images in each panel) and mCherry-Histone (right images in each panel), presented as for Fig. 1. Asterisks indicate polar bodies. (A) Wild-type embryo. Red arrowheads indicate the enrichment of GFP–CDC-6 on chromatin in anaphase of meiosis I and prometaphase of first mitosis. Green arrowheads show the position of interphase nuclei. (B) orc-5 RNAi embryo. (C) cdt-1 RNAi embryo. (D) Mean chromatin enrichment in five embryos according to cell cycle stage. Error bars are SDs. See also Video 6. (E and F) CDT-1 localization during metaphase and anaphase of meiosis II. Wild-type (E) and mcm-7 RNAi (F) embryos stained with anti–CDT-1 and antitubulin antibodies are shown. The arrow indicates chromatin-bound CDT-1. Insets to the left show a 2.5-fold magnification of maternal chromosomes. Bars, 10 µm, except for the insets in E and F, in which the bars are 5 µm.
GFP–CDC-6 and CDT-1 localization. (A–C) Images from time-lapse sequences of embryos expressing GFP–CDC-6 (left images in each panel) and mCherry-Histone (right images in each panel), presented as for Fig. 1. Asterisks indicate polar bodies. (A) Wild-type embryo. Red arrowheads indicate the enrichment of GFP–CDC-6 on chromatin in anaphase of meiosis I and prometaphase of first mitosis. Green arrowheads show the position of interphase nuclei. (B) orc-5 RNAi embryo. (C) cdt-1 RNAi embryo. (D) Mean chromatin enrichment in five embryos according to cell cycle stage. Error bars are SDs. See also Video 6. (E and F) CDT-1 localization during metaphase and anaphase of meiosis II. Wild-type (E) and mcm-7 RNAi (F) embryos stained with anti–CDT-1 and antitubulin antibodies are shown. The arrow indicates chromatin-bound CDT-1. Insets to the left show a 2.5-fold magnification of maternal chromosomes. Bars, 10 µm, except for the insets in E and F, in which the bars are 5 µm.
We could not generate worms expressing GFP–CDT-1, possibly as a consequence of toxicity caused by rereplication associated with elevated levels of GFP–CDT-1 (Zhong et al., 2003). Therefore, we examined CDT-1 localization using antibodies. CDT-1 associated with chromatin during anaphase of meiosis II as well as during the metaphase (11 of 15 embryos) and anaphase of mitosis (Figs. 4 E and S5 A). The specificity of the signal was confirmed by staining cdt-1 RNAi embryos (Fig. S5 B; Zhong et al., 2003). Therefore, chromatin binding of CDT-1 is restricted to the licensing period and commences later than ORC and CDC-6. Consistent with this, in mcm-7 RNAi embryos, the chromatin binding of CDT-1 was abolished (Fig. 4 F), indicating that MCM-2–7 and CDT-1 binding to DNA are interdependent. In contrast to MCM-2–7, however, the chromatin binding of CDT-1 did not increase during anaphase (compare Fig. S5 A with Fig. S1 A). This is consistent with a model in which CDT-1 helps to bring MCM-2–7 to origin DNA during the licensing reaction but does not stably associate with MCM-2–7 once it has been clamped around DNA.
The dynamics of pre-RC components
Biochemical studies using Xenopus laevis egg extracts have suggested that origin licensing destabilizes ORC on DNA, potentially creating a negative feedback loop that helps to ensure that all origins are licensed (Rowles et al., 1999; Harvey and Newport, 2003; Oehlmann et al., 2004). To investigate whether this destabilization of ORC occurs in vivo, we examined the localization of pre-RC components when licensing was blocked. When licensing was blocked by RNAi against cdt-1, cdc-6, or mcm-7, the levels of ORC-2 on DNA were greater than in wild type during anaphase of meiosis II and late mitosis, periods in which licensing normally occurs (Figs. 3 [D and E] and S4 [B and G]). Similarly, cdt-1 RNAi also increased GFP–ORC-1 and CDC-6 chromatin binding at these cell cycle stages (Figs. 4 [C and D], S3 C, and S4 G). Interestingly, the hyperloading of CDC-6 did not occur when ORC subunits were knocked down (Figs. 4 [B and D] and S4 [C–F]).
Next, we used FRAP to investigate whether the kinetics of pre-RC proteins in wild-type embryos can explain the hyperaccumulation of ORC and CDC-6 on chromatin that occurs when licensing is blocked. We took advantage of the fact that licensing occurs before nuclear envelope assembly to investigate the mobility of the ORC-1, ORC-2, CDC-6, and MCM-3 reporters during the first mitosis when the mitotic spindle is large and chromosomes are in the center of the cell. The anterior chromatids were photobleached 15–30 s after anaphase onset, whereas the posterior chromatids served as a control. Consistent with our previous results, the levels of GFP–ORC-1, –ORC-2, and –CDC-6 remained fairly constant on unbleached chromatids, whereas levels of GFP–MCM-3 increased steadily, indicating an accumulation of GFP–MCM-3 on chromatin as origins became licensed (Fig. 5, A and B [dashed lines]). After photobleaching, GFP–CDC-6 showed a complete and rapid recovery, whereas GFP–ORC-1, –ORC-2, and –MCM-3 recovered only partially (Fig. 5 B, solid lines). Release of ORC and CDC-6 from chromatin that occurs after nuclear assembly prevented us from following recovery over longer periods of time.
Dynamic chromatin binding of licensing factors. (A) Time-lapse images of embryos progressing through anaphase, expressing GFP–CDC-6, –ORC-1, –ORC-2, or –MCM-3 as indicated. The anterior (left) mass of DNA was photobleached 20–30 s after anaphase onset. The posterior (right) mass of DNA was used as control. Embryos are shown at −1, 1, 4, and 8 s after photobleaching. Bar, 10 µm. (B) Normalized fluorescence intensity of bleached (solid lines) or unbleached control (dashed lines) DNA (intensity prephotobleaching = 1, and cytoplasmic intensity = 0). (C) Turnover rates obtained by subtracting the accumulation of GFP on chromatin (given by the unbleached control DNA) from the normalized fluorescence intensity. (D and E) Mobile fractions of chromatin-bound licensing proteins (left graphs) and half-lives of protein recovery (right graphs) for wild-type (D) and cdt-1 RNAi (E) embryos. The mobile fraction is the fraction of GFP recovered 20 s after photobleaching. The half-life is the time to recover half of the mobile fraction. Error bars represent SD. n = 5.
Dynamic chromatin binding of licensing factors. (A) Time-lapse images of embryos progressing through anaphase, expressing GFP–CDC-6, –ORC-1, –ORC-2, or –MCM-3 as indicated. The anterior (left) mass of DNA was photobleached 20–30 s after anaphase onset. The posterior (right) mass of DNA was used as control. Embryos are shown at −1, 1, 4, and 8 s after photobleaching. Bar, 10 µm. (B) Normalized fluorescence intensity of bleached (solid lines) or unbleached control (dashed lines) DNA (intensity prephotobleaching = 1, and cytoplasmic intensity = 0). (C) Turnover rates obtained by subtracting the accumulation of GFP on chromatin (given by the unbleached control DNA) from the normalized fluorescence intensity. (D and E) Mobile fractions of chromatin-bound licensing proteins (left graphs) and half-lives of protein recovery (right graphs) for wild-type (D) and cdt-1 RNAi (E) embryos. The mobile fraction is the fraction of GFP recovered 20 s after photobleaching. The half-life is the time to recover half of the mobile fraction. Error bars represent SD. n = 5.
Fluorescence recovery on chromatin could be a result of the exchange of GFP between chromatin and cytoplasm or a result of accumulation of GFP on chromatin. To distinguish between these two possibilities, the accumulated fluorescence intensity of the unbleached chromatid was subtracted from that of the photobleached sample (Fig. 5, C and D). This subtraction showed that for GFP–MCM-3, recovery and accumulation occurred at a similar rate. Thus, there is very little, if any, unloading of GFP–MCM-3 during mitosis. In contrast, virtually all of the GFP–CDC-6 was exchangeable with a half-life of ∼3 s. GFP–ORC-1 and GFP–ORC-2 showed an intermediate behavior, with ∼50% of the protein being exchangeable with a half-life of ∼5 s.
Our previous experiments showed that inhibition of licensing increases the amount of GFP–ORC-2, –ORC-1, and –CDC-6 on chromatin. Therefore, we investigated whether the mobility of ORC and CDC-6 changed when licensing was inhibited by cdt-1 RNAi (Fig. 5 E). The mobile fraction of both GFP–ORC-1 and GFP–ORC-2 showed a significant decrease in cdt-1 RNAi embryos, suggesting a stabilization of ORC on DNA in the absence of origin licensing. The relatively small amount of GFP–ORC-1 or GFP–ORC-2 in the mobile fraction made it difficult to establish an accurate half-life, though it appeared relatively unchanged. The mobile fraction of GFP–CDC-6 was also reduced by cdt-1 RNAi, and the half-life of the exchangeable fraction was increased from ∼3 to ∼5 s. The increased amount of ORC and CDC-6 on DNA, the reduction of the mobile fraction of ORC, and the slower half-life of CDC-6 are consistent with ORC and CDC-6 having a higher affinity for unlicensed DNA than licensed DNA as a result of a reduction in the off-rate. This provides a feedback loop that may help to distribute licensed origins throughout chromosomal DNA (see Discussion).
Preventing rereplication
Licensing must be switched off during S phase to prevent rereplication. We have previously shown that in C. elegans embryos, ORC-1, ORC-2, and CDC-6 are excluded from nuclei during S phase and prophase. Indeed, GFP–ORC-2 is rapidly released from DNA as chromosomes decondense after the end of anaphase, both in meiosis II (Fig. 6 A and Video 7) and at the end of the anaphase of the first embryonic cell cycle. Therefore, we tested whether the nuclear envelope acts as a physical barrier preventing the binding of ORC to DNA. To test this hypothesis, we took advantage of the finding that the nuclear envelope fails to assemble when certain components of the nuclear pore such as NPP-8/Nup155 are depleted by RNAi (Galy et al., 2003). In npp-8 RNAi embryos, chromosomes failed to decondense, and GFP–ORC-2 remained associated with chromatin during interphase (Fig. 6 [A and B] and Video 7). Similar results were obtained with GFP–ORC-1 and GFP–CDC-6. These results indicate that the nuclear envelope prevents ORC and CDC-6 association with DNA during interphase. As ORC and CDC-6 release from DNA begins before nuclei massively increase in size, dissociation appears to depend on an active process rather than simply as a result of a dispersal of chromatin-bound ORC. To test this hypothesis, we blocked nuclear protein export by depleting the XPO-1/Exportin-1 homologue. xpo-1 RNAi caused a delay in GFP–ORC-1, –ORC-2, and –CDC-6 nuclear exclusion during the first and second embryonic S phases (Fig. 6 [A–C] and Video 7).
Composite of three videos showing a wild-type embryo (top), an xpo-1 RNAi embryo (middle), and an npp-8 RNAi embryo (bottom). Embryos are expressing GFP–ORC-2 (left) and mCherry-Histone (right) and progress throughout meiotic divisions and the first embryonic cell cycle. Images were acquired every 10 s with a widefield fluorescent microscope (DeltaVision Spectris; Applied Precision). Time is indicated in minutes and seconds.
Composite of three videos showing a wild-type embryo (top), an xpo-1 RNAi embryo (middle), and an npp-8 RNAi embryo (bottom). Embryos are expressing GFP–ORC-2 (left) and mCherry-Histone (right) and progress throughout meiotic divisions and the first embryonic cell cycle. Images were acquired every 10 s with a widefield fluorescent microscope (DeltaVision Spectris; Applied Precision). Time is indicated in minutes and seconds.
Nuclear export prevents rereplication. (A) Time-lapse images of embryos at 3, 4, and 5 min after anaphase onset of meiosis II. Embryos are expressing GFP–ORC-2 (left images in each panel) and mCherry-Histone (right images in each panel). Wild type (left), npp-8 RNAi (middle), and xpo-1 RNAi (right) are shown. Bar, 5 µm. See also Video 7. (B) Chromatin enrichment of GFP–ORC-2 in wild-type and xpo-1 RNAi embryos. (C) Mean time for GFP–ORC-2 chromatin enrichment to persist after anaphase of meiosis and mitosis (mean of five embryos). (D) Accumulation of mCherry-Histone during metaphase of the first mitosis in wild-type (top) and xpo-1 RNAi (bottom) embryos. The mean mCherry fluorescence intensity of five embryos is shown. (A and D) Asterisks indicate polar bodies. Arrows point to the position of the chromatin used for quantification. (E) Visualization of chromatin in ∼20 cell stage embryos fixed and stained with antitubulin and a DNA dye. (C and D) Error bars represent SD. (D and E) Bars, 10 µm.
Nuclear export prevents rereplication. (A) Time-lapse images of embryos at 3, 4, and 5 min after anaphase onset of meiosis II. Embryos are expressing GFP–ORC-2 (left images in each panel) and mCherry-Histone (right images in each panel). Wild type (left), npp-8 RNAi (middle), and xpo-1 RNAi (right) are shown. Bar, 5 µm. See also Video 7. (B) Chromatin enrichment of GFP–ORC-2 in wild-type and xpo-1 RNAi embryos. (C) Mean time for GFP–ORC-2 chromatin enrichment to persist after anaphase of meiosis and mitosis (mean of five embryos). (D) Accumulation of mCherry-Histone during metaphase of the first mitosis in wild-type (top) and xpo-1 RNAi (bottom) embryos. The mean mCherry fluorescence intensity of five embryos is shown. (A and D) Asterisks indicate polar bodies. Arrows point to the position of the chromatin used for quantification. (E) Visualization of chromatin in ∼20 cell stage embryos fixed and stained with antitubulin and a DNA dye. (C and D) Error bars represent SD. (D and E) Bars, 10 µm.
We investigated whether the persistence of nuclear ORC and CDC-6 during S phase might have functional consequences. Defects in DNA replication typically leads to an excessive asynchrony in the cell cycle timing of the two blastomeres in the C. elegans two-cell embryo, which depends on the preferential engagement of cell cycle checkpoints in the posterior blastomere (Encalada et al., 2000; Brauchle et al., 2003). Indeed, the asynchrony of xpo-1 RNAi embryos was of 6 ± 2.4 min instead of 2 ± 0.3 min in the wild type. To determine whether preventing nuclear protein export leads to overreplication, we used the intensity of mCherry-Histone on the metaphase plate of the first zygotic cell division as a marker for the amount of chromosomal DNA. The fluorescence intensity of metaphase-associated mCherry-Histone approximately doubled as a result of xpo-1 depletion, indicative of excessive DNA replication (Fig. 6 D). Importantly, we have not observed cytokinesis failure or additional rounds of chromosome condensation during the first or the second embryonic cell cycle, and we have followed similar embryos from meiotic divisions (Video 7). Therefore, the embryos that have an increased histone content have completed only one S phase. Codepletion of the regulatory subunit of DNA polymerase-α by RNAi in conjunction with xpo-1 RNAi reduced the level of mCherry-Histone on mitotic chromosomes, indicating that mCherry-Histone accumulation is indeed a consequence of DNA replication. To further support the notion that excessive replication occurs upon xpo-1 depletion, we stained DNA with Hoechst 33258. xpo-1 RNAi induced the formation of large nuclei and a massively increased amount of DNA in embryos, which seem to arrest at the 15–20-cell stage (Fig. 6 E). These results are consistent with the idea that nuclear export of licensing factors such as ORC and CDC-6 is required to prevent rereplication in the C. elegans embryo.
Discussion
Origins must be licensed by loading Mcm2–7 complexes before entry into S phase. Despite the importance of this process for replicating the genome, relatively little is known about its dynamics in vivo. We have used live-cell imaging of GFP-tagged pre-RC proteins to investigate the dynamics of origin licensing in the early C. elegans embryo. Several considerations strongly support the idea that the GFP fusions we use for live imaging reflect the behavior of their endogenous counterparts. This conclusion is supported by the overall dynamic behavior of pre-RC proteins, by their biochemical interactions, and by the genetic dependencies for their dynamic chromatin localization. Furthermore, antibodies against endogenous CDC-6 and ORC-2 confirmed the localization of the respective GFP fusions. GFP–MCM-3 localization was confirmed by anti–pan-MCM antibodies, and the dynamic behavior of GFP–MCM-3 was identical to GFP–MCM-2. Also, the lethality caused by RNAi against the 3′ untranslated region (UTR) of MCM-3 could be partially rescued by GFP–MCM-3 driven from the pie-1 promoter (see Materials and methods).
Our results show several differences in the way that origin licensing is regulated in the C. elegans early embryo as compared with other metazoan cell types. These differences may partly be because of the short duration of the early embryonic cell cycle and because the C. elegans early embryo is transcriptionally silent. The independent binding to DNA of ORC and CDC-6 that we observe might help in speeding up pre-RC formation, whereas the nuclear export of these proteins provides a way of preventing rereplication without the need for their complete degradation and resynthesis in each cell cycle. These ideas are consistent with previous studies showing that geminin and Cdt1 protein levels remain high throughout the rapid cell cycles in Xenopus and Drosophila melanogaster early embryos (Quinn et al., 2001; Kisielewska and Blow, 2012).
Origin licensing
Our live-cell imaging showed that GFP–MCM-3 loading on DNA occurred during anaphase of meiosis II and late mitosis. This observation was confirmed by staining the MCM-2–7 protein. Consistent with previous biochemical and genetic analysis, MCM-2–7 loading was dependent on the other pre-RC proteins ORC, CDC-6, and CDT-1 (Blow and Dutta, 2005; DePamphilis et al., 2006). It was also dependent on MCM-7, consistent with origin licensing requiring all six Mcm2–7 proteins (Prokhorova and Blow, 2000). Knockdown of MCM-7 abolished the enrichment of the replication fork protein GFP–CDC-45 on S phase chromatin, consistent with MCM-2–7 playing an essential role in the establishment of replication forks during S phase.
We could not determine whether MCM-2–7 chromatin loading continued into early interphase, as chromatin association was obscured by its nuclear import. However, ORC and CDC-6 were released from chromatin when chromosomes decondensed at the start of interphase, which likely marks the end of the licensing period. FRAP experiments showed that GFP–MCM-3 accumulated steadily on chromatin during anaphase, and this rate of accumulation was essentially unchanged after photobleaching. This suggests that there is no significant unloading of MCM-3 proteins during this period and that the licensing reaction is essentially unidirectional. This is consistent with a recent study showing that the association of human Mcm2–7 with DNA is exceptionally stable over a period of >12 h (Kuipers et al., 2011). The stable association of MCM-2–7 proteins with chromatin is as expected from the licensing model of replication control: because licensing cannot continue once S phase has started, the previously established association of MCM-2–7 proteins with origins must be stable enough to persist until the completion of DNA replication. This firm, irreversible association is plausibly explained by the licensing reaction representing the clamping of ring-shaped Mcm2–7 double hexamers around DNA (Fig. 7, A and B; Evrin et al., 2009; Remus et al., 2009; Gambus et al., 2011).
A model for the efficient distribution of licensed origins. Events occurring on DNA (solid black line) as origins are licensed during anaphase and replicate during S phase. (A) ORC and Cdc6 bind to an unlicensed origin. (B) The origin is licensed by loading an Mcm2–7 double hexamer (M). (C) The binding of Mcm2–7 destabilizes ORC, Cdc6, and Cdt1 at that site so they can be displaced to a new site on the chromosome. (D) ORC and Cdc6 are exported out of the interphase nucleus, whereas Cdc45 is recruited to Mcm2–7 when replication forks initiate.
A model for the efficient distribution of licensed origins. Events occurring on DNA (solid black line) as origins are licensed during anaphase and replicate during S phase. (A) ORC and Cdc6 bind to an unlicensed origin. (B) The origin is licensed by loading an Mcm2–7 double hexamer (M). (C) The binding of Mcm2–7 destabilizes ORC, Cdc6, and Cdt1 at that site so they can be displaced to a new site on the chromosome. (D) ORC and Cdc6 are exported out of the interphase nucleus, whereas Cdc45 is recruited to Mcm2–7 when replication forks initiate.
The pre-RC components ORC, Cdc6, and Cdt1 are required to load Mcm2–7 onto DNA and license replication origins. Here, we show that in worm embryos, the association of ORC subunits and CDC-6 with chromatin is more complex than has been described in other systems. We have identified C. elegans homologues of the major Orc1–5 subunits and provided evidence that ORC-2–5 form a core complex, which requires ORC-1 for binding chromatin. ORC-2, and probably the other components of the ORC-2–5 complex, binds to chromatin from anaphase of meiosis I through to anaphase of meiosis II, and this binding is dependent on ORC-1. In contrast, ORC-1 binds to chromatin, earlier in metaphase of meiosis I. Like ORC-1, CDC-6 also binds chromatin in the absence of ORC-2. The ability of ORC-1 to bind chromatin independently of the other ORC subunits is feasible, given the elongated nature of the Orc1–5 complex (Speck et al., 2005; Clarey et al., 2006, 2008; Chen et al., 2008). Cdc6 shares significant homology with regions of certain ORC subunits, particularly Orc1, and Archaea typically contain only a single Cdc6/Orc1 homologue (Liu et al., 2000). Therefore, it is possible that worm ORC and CDC-6 polypeptides associate with DNA in a manner more similar to their archaeal counterparts. The strong binding of ORC-1 to chromosomes during metaphase of meiosis I, a cell cycle stage when origin licensing does not occur, may reflect another function at this stage of development that is independent of replication licensing. The independent binding of ORC-1, ORC-2–5, and CDC-6 to chromatin could also help speed up pre-RC assembly, which has to occur very rapidly in the C. elegans early embryo.
Analysis of Cdt1 in other metazoans has shown that Cdt1 can associate with ORC on DNA in the absence of Mcm2–7 (Gillespie et al., 2001). Here, we show that the association of worm CDT-1 with chromosomes occurs only during periods when licensing is taking place and is dependent on MCM-2–7. This is reminiscent of the behavior of Cdt1 in budding yeast, which forms a physical complex with Mcm2–7 and is consistent with the idea that the biochemical function of Cdt1 is to bring Mcm2–7 together with an ORC–Cdc6 complex on DNA. The chromatin binding of this hypothetical Cdt1–Mcm2–7 complex might be faster than the sequential binding of Cdt1 and Mcm2–7 observed in somatic cells, which might be important to increase the rate of origin licensing in the early embryo.
Licensing-dependent origin inactivation
It is currently unclear how eukaryotic replication origins are recognized and efficiently licensed by pre-RC proteins in vivo. It is essential that before entry into S phase, a large enough number of origins is licensed and that these licensed origins are appropriately distributed along chromosomal DNA. The appropriate distribution of licensed origins is particularly important in early embryos, which have a very short time to license and replicate all their DNA. In C. elegans embryos, most DNA replication occurs in ∼8.5 min (our data; Edgar and McGhee, 1988). At a typical metazoan fork rate of 1 kb/min, this would suggest that throughout the 100-Mbp genome, adjacent replication origins can be spaced no more than 17 kb apart (8.5 min × 1 kb/min × two replication forks), and a minimum of 5,800 origins (100 Mbp/17 kb) must be licensed during M phase. Therefore, the embryo must possess a mechanism to ensure that there are no excessively large gaps between origins that cannot be replicated before cells progress into mitosis.
Our live-cell imaging data suggest that C. elegans ORC interacts with chromosomal DNA in a highly dynamic fashion. First, we and others showed that in C. elegans embryos, ORC-1, ORC-2, CDT-1, and CDC-6 show cell cycle–dependent changes in chromatin association during progression through meiotic divisions and are released from chromatin during interphase (Zhong et al., 2003; Kim et al., 2007). Second, FRAP experiments showed a rapid turnover of GFP–ORC-1, –ORC-2, and –CDC-6 on chromatin with half-lives ranging from 3 to 5 s. This dynamic behavior is in line with previous live-cell imaging of GFP-Orc4 in CHO cells (McNairn et al., 2005). Our data suggest that in the C. elegans early embryo, the licensing period takes ∼150 s between the end of metaphase and chromosome decondensation. The rapid off-rate of GFP–ORC-1, –ORC-2, and –CDC-6 could allow them to repeatedly load MCM-2–7 complexes ∼30 times, which is sufficient to yield the expected excess of chromatin-bound MCM-2–7 over ORC.
We have also observed evidence of a negative feedback loop operating on ORC and CDC-6 that could prevent excessively large gaps between adjacent origins. We show that inhibition of licensing results in an increased localization of ORC and CDC-6 to chromatin. FRAP experiments showed that in the absence of licensing, most ORC-1 and ORC-2 no longer exchange with the cytoplasmic pool, whereas CDC-6 also has a reduced exchange time. These results are consistent with a model in which the dissociation of ORC and Cdc6 from chromatin is significantly decreased in the absence of licensing, but, when Mcm2–7 complexes are loaded, dissociation is promoted (Fig. 7 C). Our data also show that hyperloading of CDC-6 does not occur when ORC is inactivated, suggesting that ORC stabilization on DNA is required for CDC-6 to be stabilized. Biochemical studies in Xenopus and yeast are also consistent with a change in ORC affinity for DNA once Mcm2–7 has been loaded (Rowles et al., 1999; Oehlmann et al., 2004; Randell et al., 2006; Waga and Zembutsu, 2006). By destabilizing ORC at sites where Mcm2–7 are bound, this behavior would have the overall effect of distributing Mcm2–7 loading to different sites throughout the genome and thus helping to ensure complete genome replication during S phase.
Preventing rereplication
To prevent replicated DNA from being relicensed and thereby rereplicated, it is crucial that the ability to license origins ceases before S phase starts. Several overlapping mechanisms that down-regulate pre-RC proteins on entry into S phase have been described in other experimental systems (Blow and Dutta, 2005; DePamphilis et al., 2006). In metazoans, geminin plays a central role in preventing rereplication by inhibiting Cdt1 during S phase and G2. Geminin also restricts licensing in unfertilized Xenopus eggs (Tada et al., 2001; Li and Blow, 2004). Here, we show that the C. elegans homologue GMN-1 (Yanagi et al., 2005) is required to prevent licensing during anaphase of meiosis I, metaphase of meiosis II, and prometaphase of mitosis. However, it seems unlikely that geminin is essential for preventing rereplication in the early C. elegans embryo because most gmn-1 RNAi embryos are viable.
Here, we also show that ORC, CDC-6, and CDT-1 are excluded, either fully or partially, from interphase nuclei (Fig. 7 D). Although nuclear exclusion of Cdc6 is common in a range of cell types, including C. elegans somatic cells (Kim et al., 2007), to our knowledge, nuclear exclusion of ORC has not been described before. ORC nuclear exclusion appears to occur in at least two different ways because exclusion of GFP–ORC-2 depended on ORC-5 but not on ORC-1, whereas nuclear exclusion of GFP–ORC-1 (and GFP–CDC-6) did not require ORC-5. The removal of ORC from chromatin during interphase required a functional nuclear envelope and the nuclear export factor XPO-1. The high rate of ORC turnover on chromatin revealed by FRAP, coupled with active nuclear export, may be sufficient to explain the release of ORC from DNA at the end of anaphase. Importantly, inhibition of nuclear protein export by knocking down the XPO-1 exportin caused a rapid and severe increase in cellular DNA content, consistent with overreplication of the DNA. This suggests that nuclear export of ORC and CDC-6 is required to prevent relicensing (and hence rereplication) of DNA during interphase. It is noteworthy that the licensing system of the early Xenopus embryo, although regulated in a different way to C. elegans, also depends on differential partition of licensing proteins between the nucleus and cytoplasm (Blow and Laskey, 1988; Li and Blow, 2005). A similar strategy for controlling the licensing system may also occur in mammalian embryonic stem cells (Yang et al., 2011). The use of nucleocytoplasmic compartmentalization that we demonstrate here may be generally suited for early embryos and other cell types with rapid cell cycles because proteins are recycled rather than being resynthesized in each cell cycle.
Materials and methods
Worms
C. elegans were maintained according to standard procedure (Brenner, 1974). Worms were raised, fed with bacteria expressing double-strand RNA (dsRNA), and recorded at 23–25°C. Transgenic worms were generated by particle bombardment (Praitis et al., 2001). To generate the GFP-tagged fusion protein, the respective full-length cDNAs were amplified from N2 worms and cloned into pie-1 regulatory element (Wallenfang and Seydoux, 2000) in a pIC26 vector (Cheeseman and Desai, 2005). Worms expressing mCherry-Histone were derived from OD57 (also see Table S2; McNally et al., 2006). To determine whether GFP–MCM-3 is functional, we inactivated the endogenous mcm-3 using RNAi against the 3′ UTR of mcm-3. This led to 0% embryonic viability (0/95 embryos hatch). In contrast, 10/85 GFP–MCM-3–expressing embryos hatched and gave rise to adult worms. The same experiment could not be performed for orc-2 and cdc-6 because their 3′ UTR RNAi does not confer embryonic lethality.
RNAi
Bacterial clones of orc-5, mcm-7, and Y47D3A.29 (DNA polymerase-α catalytic subunit) dsRNA feeding strains were obtained from a commercial library (Kamath et al., 2003). Their identity was verified by examining the insert sizes. orc-1, orc-2, cdc-6, cdt-1, and gmn-1 RNAi feeding strains were generated by cloning cDNAs into L4440 (Timmons and Fire, 1998). Bacteria were grown to OD600 = 1, supplemented with 1 mM IPTG, and spread on 10-ml nematode growth media plates. dsRNA was induced 4–12 h at room temperature. L4 worms were then added to plates and fed for 24–32 h before analysis.
Microscopy
To record meiotic divisions, embryos were dissected in a previously described isotonic growth medium for blastomeres containing 35% bovine FCS (Shelton and Bowerman, 1996). The described medium composition is 1 ml of 5 mg/ml inulin (dissolved with brief autoclaving; Sigma-Aldrich), 50 mg of tissue culture–grade polyvinylpyrrolidone powder (Sigma-Aldrich), and 100 µl each of Basal Medium Eagle vitamins (Invitrogen), and chemically defined lipid concentrate (Invitrogen) and 100× concentrated penicillin-streptomycin (Invitrogen) were added to 9 ml of Drosophila Schneider’s Medium (Invitrogen). The final volume was 10.3 ml. Before use, bovine FCS (heat treated for 30 min at 56°C; Invitrogen) was added. Embryos were mounted on 2% agarose pads, and Vaseline patches on the slide reduced the pressure of the coverslip on the embryo. Images were captured every 30 s using a widefield deconvolution microscope (Personal DeltaVision; Applied Precision) mounted on an inverted microscope (TE200; Nikon) with a 100×/1.40 Plan Apochromat oil immersion lens (Nikon), a camera (CoolSNAP HQ2; Roper Scientific), and softWoRx software (Applied Precision). The exposure time was 0.25 s, and binning was 2 × 2. There was a 1-s time interval between acquisition of GFP and mCherry images (except for Fig. 6, in which a DeltaVision Core microscope [Applied Precision] was used; see details below). Embryos expressing GFP–CDC-45 (Fig. 2) were analyzed using a spinning-disk confocal microscope (MAG Biosystems) mounted on a microscope (IX81; Olympus) with a 100×/1.45 Plan Apochromat oil immersion lens (Olympus), a camera (Cascade II; Photometrics), spinning-disk head (CSU-X1; Yokogawa Electric Corporation), and MetaMorph software (Molecular Devices). Embryos dissected after meiosis completion (Figs. 5 and 6 D) were mounted in M9 buffer on 2% agarose pads, and images were produced using a widefield DeltaVision Core microscope mounted on a microscope (IX71; Olympus) with a 100×/1.40 Plan Apochromat oil immersion lens (Olympus), a camera (CoolSNAP HQ; Photometrics), and softWoRx software. The exposure time was 0.25 s, and binning was 2 × 2. For FRAP experiments, GFP images were captured every 400 ms with 4 × 4 binning. During anaphase, the anterior mass of DNA was bleached with a laser pulse of 400 ms using a 488-nm diode laser (Point Souce; Qioptiq) at 100% laser power (10-mW laser). Image analysis and video processing were performed in ImageJ software (National Institutes of Health). For immunostaining, embryos were fixed in methanol at −20°C and stained using standard procedures with rabbit antibodies for CDC-6 (1:2,000; Kim et al., 2007), CDT-1 (1:1,000; Zhong et al., 2003), or MCM-2–7 (1:1,000; BD) and with the mouse antibodies for ORC-2 (1:100; Hadwiger et al., 2010) or α-tubulin (1:400, DM1A; Sigma-Aldrich). CDT-1 and CDC-6 antibodies were a gift from E. Kipreos (University of Georgia, Athens, GA). Secondary antibodies were donkey anti–rabbit conjugated to Alexa Fluor 568 (Invitrogen) and donkey anti–mouse conjugated with Alexa Fluor 488 (Invitrogen). DNA was visualized with Hoechst 33258. Embryos were imaged using a confocal laser-scanning microscope (SP2; Leica) using a 63×/1.40 Plan Apochromat oil immersion lens (Leica), except for Fig. 6 E, in which a DeltaVision Core microscope was used (see details above). Each embryo shown is representative of >10 embryos observed at a similar cell cycle stage.
Quantification of GFP signal
An area of interest was drawn around the oocyte-derived DNA (excluding polar bodies). The 100 brighter pixels were recognized as DNA, whereas 100 dark pixels were recognized as cytoplasm. The corresponding GFP image was then used to determine mean values for DNA and cytoplasm. Background intensity was taken outside the embryo. The chromatin/cytoplasm ratio was determined as follows: ratio = (DNA − background)/(cytoplasm − background). This procedure was applied to all images of the video. Macros were written using ImageJ to proceed in an automated manner.
ORC purification
Approximately 0.5 ml of adult-enriched liquid cultures was washed, snap frozen in liquid nitrogen, and stored at −80°C. Pellets were resuspended in 0.75 ml of lysis buffer (10 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.5 mM EDTA, 0.5% NP-40, and protease inhibitor cocktail [Complete, Mini; Roche]) and an addition of 300 µl × 0.7-mm Zirconia beads followed by bead beating (three times for 20 s with 20-s intervals). Lysates were incubated on ice for 30 min, transferred to a new tube, clarified at 18,000 g for 10 min at 4°C. 40 mg of protein (corresponding to 3–5 ml of worm extract) was used for immunoprecipitations, and the volume was adjusted to 500 µl with dilution buffer (10 mM Tris-Cl pH 7.5, 150 mM NaCl, 0.5 mM EDTA, and 1× protease inhibitor cocktail). Lysates were cleared with a 1:1 slurry of 50 µl protein G–Sepharose for 1 h at 4°C, and beads were removed. The supernatant was incubated with a 1:1 slurry of 30 µl GFP-Trap A beads (ChromoTek) for 1 h. Beads were washed once with 0.5 ml of dilution buffer and once with 0.5 ml of wash buffer (10 mM Tris-Cl pH 7.5, 300 mM NaCl, 0.5 mM EDTA, and 1× protease inhibitor cocktail) for 30 min at 4°C, and beads were pelleted. Proteins were eluted with 20 µl SDS sample buffer, separated on SDS gels, and subjected to mass spectroscopy.
Online supplemental material
Fig. S1 shows that MCM-2–7 localization to chromatin mimics that of GFP–MCM-3 and depends on MCM-7, ORC-5, and CDT-1. Fig. S2 shows Orc3 and Orc4 amino acid conservation in eukaryotes. Fig. S3 shows that ORC-2 and CDC-6 localization to chromatin mimics that of GFP–ORC-2 and GFP–CDC-6, ORC-2 accumulates in cdc-6 RNAi and cdt-1 RNAi embryos, CDC-6 accumulates in cdt-1 RNAi embryos, and ORC-2, but not CDC-6, DNA binding depends on ORC-3, ORC-4, and ORC-5. Fig. S4 shows that GFP–ORC-1 localization on chromatin depends on ORC-5, but GFP–ORC-1 overall quantity is reduced in orc-2 RNAi and orc-5 RNAi embryos, and GFP–ORC-1 accumulates on DNA in cdt-1 RNAi embryos. Fig. S5 shows that CDT-1 localizes to chromatin during anaphase of meiosis II, metaphase, and anaphase of mitosis and that the amount of CDT-1 on chromatin does not seem to increase from metaphase to anaphase of mitosis. Video 1 shows the chromatin or nuclear localization of GFP–MCM-3 during meiotic divisions and the first embryonic cell cycle. Video 2 shows the chromatin or nuclear localization of GFP–MCM-2 during the first embryonic cell cycle. Video 3 shows the chromatin or nuclear localization of GFP–CDC-45 during the second meiotic divisions and the first embryonic cell cycle. Video 4 shows the chromatin localization or nuclear exclusion of GFP–ORC-2 during meiotic divisions and the first embryonic cell cycle. Video 5 shows the chromatin localization or nuclear exclusion of GFP–ORC-1 during meiotic divisions and the first embryonic cell cycle. Video 6 shows the chromatin localization or nuclear exclusion of GFP–CDC-6 during meiotic divisions and the first embryonic cell cycle. Video 7 shows that GFP–ORC-2 release from chromatin is slower in xpo-1 RNAi embryos and is abolished in npp-8 RNAi embryos; in addition, the higher intensity of mCherry-Histone on chromatin during the first mitosis observed in xpo-1 RNAi embryos is not a consequence of failed mitosis or successive S phases. Table S1 shows that proteins associated with GFP–ORC-2, GFP–ORC-5, GFP–PCN-1, and GFP–SLD-5 were precipitated and identified by mass spectrometry; the number of different peptides identified are indicated, and nonexpressing N2 worms were used as control. Table S2 shows the genotype and construction of GFP fusion worm strains.
Acknowledgments
We thank Sam Swift and the College of Life Sciences microscopy facility for help with imaging and Sara ten Have for help with proteomics analysis.
This work was supported by Cancer Research UK grants C303/A7399 (to J.J. Blow) and C11852/A4500 (to A. Gartner), by the Association for International Cancer Research grant 06-57 and a Wellcome Senior Research Fellowship (to A. Gartner), and a European Molecular Biology Organization long-term fellowship (to R. Sonneville). During this work, M. Querenet was a Master’s student at Ecole Normale Supérieure de Lyon.